Divergent mutational processes distinguish hypoxic and normoxic tumours
Abstract
Many primary tumours have low levels of molecular oxygen (hypoxia), and hypoxic tumours respond poorly to therapy. Pan-cancer molecular hallmarks of tumour hypoxia remain poorly understood, with limited comprehension of its associations with specific mutational processes, non-coding driver genes and evolutionary features. Here, as part of the ICGC/TCGA Pan-Cancer Analysis of Whole Genomes (PCAWG) Consortium, which aggregated whole genome sequencing data from 2658 cancers across 38 tumour types, we quantify hypoxia in 1188 tumours spanning 27 cancer types. Elevated hypoxia associates with increased mutational load across cancer types, irrespective of underlying mutational class. The proportion of mutations attributed to several mutational signatures of unknown aetiology directly associates with the level of hypoxia, suggesting underlying mutational processes for these signatures. At the gene level, driver mutations in TP53, MYC and PTEN are enriched in hypoxic tumours, and mutations in PTEN interact with hypoxia to direct tumour evolutionary trajectories. Overall, hypoxia plays a critical role in shaping the genomic and evolutionary landscapes of cancer.
Article type: Research Article
Keywords: Cancer genomics, Cancer microenvironment, Tumour heterogeneity
Affiliations: grid.17063.330000 0001 2157 2938Department of Medical Biophysics, University of Toronto, Toronto, ON Canada; grid.419890.d0000 0004 0626 690XOntario Institute for Cancer Research, Toronto, Canada; grid.19006.3e0000 0000 9632 6718Department of Human Genetics, University of California, Los Angeles, USA; grid.5379.80000000121662407Division of Cancer Sciences, Faculty of Biology, Health and Medicine, University of Manchester, Manchester, UK; grid.412917.80000 0004 0430 9259The Christie NHS Foundation Trust, Manchester, UK; CRUK Manchester Institute and Centre, Manchester, UK; grid.17063.330000 0001 2157 2938Department of Pharmacology and Toxicology, University of Toronto, Toronto, Canada; grid.494618.6Vector Institute for Artificial Intelligence, Toronto, Canada; grid.19006.3e0000 0000 9632 6718Department of Urology, University of California, Los Angeles, USA; grid.19006.3e0000 0000 9632 6718Jonsson Comprehensive Cancer Centre, University of California Los Angeles, Los Angeles, USA; grid.19006.3e0000 0000 9632 6718Institute for Precision Health, University of California Los Angeles, Los Angeles, USA; grid.7737.40000 0004 0410 2071Applied Tumor Genomics Research Program, Research Programs Unit, University of Helsinki, Helsinki, Finland; grid.10306.340000 0004 0606 5382Wellcome Sanger Institute, Wellcome Genome Campus, Hinxton, UK; grid.51462.340000 0001 2171 9952Memorial Sloan Kettering Cancer Center, New York, NY USA; grid.26999.3d0000 0001 2151 536XGenome Science Division, Research Center for Advanced Science and Technology, University of Tokyo, Tokyo, Japan; grid.170205.10000 0004 1936 7822Department of Surgery, University of Chicago, Chicago, IL USA; grid.414067.00000 0004 0647 8419Department of Surgery, Division of Hepatobiliary and Pancreatic Surgery, School of Medicine, Keimyung University Dongsan Medical Center, Daegu, South Korea; grid.256155.00000 0004 0647 2973Department of Oncology, Gil Medical Center, Gachon University, Incheon, South Korea; grid.257022.00000 0000 8711 3200Hiroshima University, Hiroshima, Japan; grid.240145.60000 0001 2291 4776Department of Bioinformatics and Computational Biology, The University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.240145.60000 0001 2291 4776University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.415310.20000 0001 2191 4301King Faisal Specialist Hospital and Research Centre, Al Maather, Riyadh, Saudi Arabia; grid.7719.80000 0000 8700 1153Bioinformatics Unit, Spanish National Cancer Research Centre (CNIO), Madrid, Spain; grid.13648.380000 0001 2180 3484Bioinformatics Core Facility, University Medical Center Hamburg, Hamburg, Germany; grid.418481.00000 0001 0665 103XHeinrich Pette Institute, Leibniz Institute for Experimental Virology, Hamburg, Germany; grid.419890.d0000 0004 0626 690XOntario Tumour Bank, Ontario Institute for Cancer Research, Toronto, ON Canada; grid.240145.60000 0001 2291 4776Department of Pathology, The University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.48336.3a0000 0004 1936 8075Laboratory of Pathology, Center for Cancer Research, National Cancer Institute, Bethesda, MD USA; grid.266100.30000 0001 2107 4242Department of Cellular and Molecular Medicine and Department of Bioengineering, University of California San Diego, La Jolla, CA USA; grid.516081.b0000 0000 9217 9714UC San Diego Moores Cancer Center, San Diego, CA USA; grid.434706.20000 0004 0410 5424Canada’s Michael Smith Genome Sciences Centre, BC Cancer, Vancouver, BC Canada; grid.1008.90000 0001 2179 088XSir Peter MacCallum Department of Oncology, Peter MacCallum Cancer Centre, University of Melbourne, Melbourne, VIC Australia; grid.11794.3a0000000109410645Centre for Research in Molecular Medicine and Chronic Diseases (CiMUS), Universidade de Santiago de Compostela, Santiago de Compostela, Spain; grid.11794.3a0000000109410645Department of Zoology, Genetics and Physical Anthropology, (CiMUS), Universidade de Santiago de Compostela, Santiago de Compostela, Spain; grid.6312.60000 0001 2097 6738The Biomedical Research Centre (CINBIO), Universidade de Vigo, Vigo, Spain; grid.416177.20000 0004 0417 7890Royal National Orthopaedic Hospital – Bolsover, London, UK; grid.240145.60000 0001 2291 4776Department of Genomic Medicine, The University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.39382.330000 0001 2160 926XQuantitative and Computational Biosciences Graduate Program, Baylor College of Medicine, Houston, TX USA; grid.249880.f0000 0004 0374 0039The Jackson Laboratory for Genomic Medicine, Farmington, CT USA; grid.419890.d0000 0004 0626 690XGenome Informatics Program, Ontario Institute for Cancer Research, Toronto, ON Canada; grid.9764.c0000 0001 2153 9986Institute of Human Genetics, Christian-Albrechts-University, Kiel, Germany; grid.410712.10000 0004 0473 882XInstitute of Human Genetics, Ulm University and Ulm University Medical Center, Ulm, Germany; grid.1003.20000 0000 9320 7537Queensland Centre for Medical Genomics, Institute for Molecular Bioscience, University of Queensland, St. Lucia, Brisbane, QLD Australia; grid.412346.60000 0001 0237 2025Salford Royal NHS Foundation Trust, Salford, UK; grid.411475.20000 0004 1756 948XDepartment of Surgery, Pancreas Institute, University and Hospital Trust of Verona, Verona, Italy; grid.5288.70000 0000 9758 5690Molecular and Medical Genetics, OHSU Knight Cancer Institute, Oregon Health and Science University, Portland, OR USA; grid.248762.d0000 0001 0702 3000Department of Molecular Oncology, BC Cancer Research Centre, Vancouver, BC Canada; grid.4367.60000 0001 2355 7002The McDonnell Genome Institute at Washington University, St. Louis, MO USA; grid.83440.3b0000000121901201University College London, London, UK; grid.272242.30000 0001 2168 5385Division of Cancer Genomics, National Cancer Center Research Institute, National Cancer Center, Tokyo, Japan; DLR Project Management Agency, Bonn, Germany; grid.410818.40000 0001 0720 6587Tokyo Women’s Medical University, Tokyo, Japan; grid.51462.340000 0001 2171 9952Center for Molecular Oncology, Memorial Sloan Kettering Cancer Center, New York, NY USA; grid.148313.c0000 0004 0428 3079Los Alamos National Laboratory, Los Alamos, NM USA; grid.417184.f0000 0001 0661 1177Department of Pathology, University Health Network, Toronto General Hospital, Toronto, ON Canada; grid.240404.60000 0001 0440 1889Nottingham University Hospitals NHS Trust, Nottingham, UK; grid.7497.d0000 0004 0492 0584Epigenomics and Cancer Risk Factors, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.419890.d0000 0004 0626 690XComputational Biology Program, Ontario Institute for Cancer Research, Toronto, ON Canada; grid.17063.330000 0001 2157 2938Department of Molecular Genetics, University of Toronto, Toronto, ON Canada; grid.494618.6Vector Institute, Toronto, ON Canada; grid.9764.c0000 0001 2153 9986Hematopathology Section, Institute of Pathology, Christian-Albrechts-University, Kiel, Germany; grid.10698.360000000122483208Department of Pathology and Laboratory Medicine, School of Medicine, University of North Carolina at Chapel Hill, Chapel Hill, NC USA; grid.55325.340000 0004 0389 8485Department of Cancer Genetics, Institute for Cancer Research, Oslo University Hospital, The Norwegian Radium Hospital, Oslo, Norway; grid.5841.80000 0004 1937 0247Pathology, Hospital Clinic, Institut d’Investigacions Biomèdiques August Pi i Sunyer (IDIBAPS), University of Barcelona, Barcelona, Spain; grid.5335.00000000121885934Department of Veterinary Medicine, Transmissible Cancer Group, University of Cambridge, Cambridge, UK; grid.4367.60000 0001 2355 7002Alvin J. Siteman Cancer Center, Washington University School of Medicine, St. Louis, MO USA; grid.8756.c0000 0001 2193 314XWolfson Wohl Cancer Research Centre, Institute of Cancer Sciences, University of Glasgow, Glasgow, UK; grid.10698.360000000122483208Lineberger Comprehensive Cancer Center, University of North Carolina at Chapel Hill, Chapel Hill, NC USA; grid.66859.340000 0004 0546 1623Broad Institute of MIT and Harvard, Cambridge, MA USA; grid.511177.4Dana-Farber/Boston Children’s Cancer and Blood Disorders Center, Boston, MA USA; grid.38142.3c000000041936754XDepartment of Pediatrics, Harvard Medical School, Boston, MA USA; grid.443984.60000 0000 8813 7132Leeds Institute of Medical Research @ St. James’s, University of Leeds, St. James’s University Hospital, Leeds, UK; grid.411475.20000 0004 1756 948XDepartment of Pathology and Diagnostics, University and Hospital Trust of Verona, Verona, Italy; grid.412744.00000 0004 0380 2017Department of Surgery, Princess Alexandra Hospital, Brisbane, QLD Australia; grid.1003.20000 0000 9320 7537Surgical Oncology Group, Diamantina Institute, University of Queensland, Brisbane, QLD Australia; grid.67105.350000 0001 2164 3847Department of Population and Quantitative Health Sciences, Case Western Reserve University School of Medicine, Cleveland, OH USA; grid.443867.a0000 0000 9149 4843Research Health Analytics and Informatics, University Hospitals Cleveland Medical Center, Cleveland, OH USA; grid.413144.70000 0001 0489 6543Gloucester Royal Hospital, Gloucester, UK; grid.225360.00000 0000 9709 7726European Molecular Biology Laboratory, European Bioinformatics Institute (EMBL-EBI), Cambridge, UK; grid.419890.d0000 0004 0626 690XDiagnostic Development, Ontario Institute for Cancer Research, Toronto, ON Canada; grid.10097.3f0000 0004 0387 1602Barcelona Supercomputing Center (BSC), Barcelona, Spain; grid.22072.350000 0004 1936 7697Arnie Charbonneau Cancer Institute, University of Calgary, Calgary, AB Canada; grid.22072.350000 0004 1936 7697Departments of Surgery and Oncology, University of Calgary, Calgary, AB Canada; grid.55325.340000 0004 0389 8485Department of Pathology, Oslo University Hospital, The Norwegian Radium Hospital, Oslo, Norway; grid.419890.d0000 0004 0626 690XPanCuRx Translational Research Initiative, Ontario Institute for Cancer Research, Toronto, ON Canada; grid.21107.350000 0001 2171 9311Department of Oncology, Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins University School of Medicine, Baltimore, MD USA; grid.430506.40000 0004 0465 4079University Hospital Southampton NHS Foundation Trust, Southampton, UK; grid.439344.d0000 0004 0641 6760Royal Stoke University Hospital, Stoke-on-Trent, UK; grid.419890.d0000 0004 0626 690XGenome Sequence Informatics, Ontario Institute for Cancer Research, Toronto, ON Canada; grid.459583.60000 0004 4652 6825Human Longevity Inc, San Diego, CA USA; grid.1018.80000 0001 2342 0938Olivia Newton-John Cancer Research Institute, La Trobe University, Heidelberg, VIC Australia; grid.9227.e0000000119573309Computer Network Information Center, Chinese Academy of Sciences, Beijing, China; grid.440163.40000 0001 0352 8618Genome Canada, Ottawa, ON Canada; grid.473715.30000 0004 6475 7299CNAG-CRG, Centre for Genomic Regulation (CRG), Barcelona Institute of Science and Technology (BIST), Barcelona, Spain; grid.5612.00000 0001 2172 2676Universitat Pompeu Fabra (UPF), Barcelona, Spain; grid.272799.00000 0000 8687 5377Buck Institute for Research on Aging, Novato, CA USA; grid.189509.c0000000100241216Duke University Medical Center, Durham, NC USA; grid.10423.340000 0000 9529 9877Department of Human Genetics, Hannover Medical School, Hannover, Germany; grid.50956.3f0000 0001 2152 9905Center for Bioinformatics and Functional Genomics, Cedars-Sinai Medical Center, Los Angeles, CA USA; grid.50956.3f0000 0001 2152 9905Department of Biomedical Sciences, Cedars-Sinai Medical Center, Los Angeles, CA USA; grid.9619.70000 0004 1937 0538The Hebrew University Faculty of Medicine, Jerusalem, Israel; grid.4868.20000 0001 2171 1133Barts Cancer Institute, Barts and the London School of Medicine and Dentistry, Queen Mary University of London, London, UK; grid.9647.c0000 0004 7669 9786Department of Computer Science, Bioinformatics Group, University of Leipzig, Leipzig, Germany; grid.9647.c0000 0004 7669 9786Interdisciplinary Center for Bioinformatics, University of Leipzig, Leipzig, Germany; grid.9647.c0000 0004 7669 9786Transcriptome Bioinformatics, LIFE Research Center for Civilization Diseases, University of Leipzig, Leipzig, Germany; grid.65499.370000 0001 2106 9910Department of Medical Oncology, Dana-Farber Cancer Institute, Boston, MA USA; grid.65499.370000 0001 2106 9910Department of Cancer Biology, Dana-Farber Cancer Institute, Boston, MA USA; grid.38142.3c000000041936754XHarvard Medical School, Boston, MA USA; grid.42505.360000 0001 2156 6853USC Norris Comprehensive Cancer Center, University of Southern California, Los Angeles, CA USA; grid.411475.20000 0004 1756 948XDepartment of Diagnostics and Public Health, University and Hospital Trust of Verona, Verona, Italy; grid.7048.b0000 0001 1956 2722Department of Mathematics, Aarhus University, Aarhus, Denmark; grid.154185.c0000 0004 0512 597XDepartment of Molecular Medicine (MOMA), Aarhus University Hospital, Aarhus N, Denmark; Instituto Carlos Slim de la Salud, Mexico City, Mexico; grid.1005.40000 0004 4902 0432Cancer Division, Garvan Institute of Medical Research, Kinghorn Cancer Centre, University of New South Wales (UNSW Sydney), Sydney, NSW Australia; grid.1005.40000 0004 4902 0432South Western Sydney Clinical School, Faculty of Medicine, University of New South Wales (UNSW Sydney), Liverpool, NSW Australia; grid.411714.60000 0000 9825 7840West of Scotland Pancreatic Unit, Glasgow Royal Infirmary, Glasgow, UK; grid.484013.a0000 0004 6879 971XCenter for Digital Health, Berlin Institute of Health and Charitè – Universitätsmedizin Berlin, Berlin, Germany; grid.7497.d0000 0004 0492 0584Heidelberg Center for Personalized Oncology (DKFZ-HIPO), German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.189509.c0000000100241216The Preston Robert Tisch Brain Tumor Center, Duke University Medical Center, Durham, NC USA; grid.32224.350000 0004 0386 9924Massachusetts General Hospital, Boston, MA USA; grid.410872.80000 0004 1774 5690National Institute of Biomedical Genomics, Kalyani, West Bengal India; grid.5510.10000 0004 1936 8921Institute of Clinical Medicine and Institute of Oral Biology, University of Oslo, Oslo, Norway; grid.10698.360000000122483208University of North Carolina at Chapel Hill, Chapel Hill, NC USA; grid.411475.20000 0004 1756 948XARC-Net Centre for Applied Research on Cancer, University and Hospital Trust of Verona, Verona, Italy; grid.18886.3fThe Institute of Cancer Research, London, UK; grid.428397.30000 0004 0385 0924Centre for Computational Biology, Duke-NUS Medical School, Singapore, Singapore; grid.428397.30000 0004 0385 0924Programme in Cancer and Stem Cell Biology, Duke-NUS Medical School, Singapore, Singapore; grid.4514.40000 0001 0930 2361Division of Oncology and Pathology, Department of Clinical Sciences Lund, Lund University, Lund, Sweden; grid.411327.20000 0001 2176 9917Department of Pediatric Oncology, Hematology and Clinical Immunology, Heinrich-Heine-University, Düsseldorf, Germany; grid.509459.40000 0004 0472 0267Laboratory for Medical Science Mathematics, RIKEN Center for Integrative Medical Sciences, Yokohama, Japan; grid.509459.40000 0004 0472 0267RIKEN Center for Integrative Medical Sciences, Yokohama, Japan; Department of Internal Medicine/Hematology, Friedrich-Ebert-Hospital, Neumünster, Germany; grid.47100.320000000419368710Departments of Dermatology and Pathology, Yale University, New Haven, CT USA; grid.473715.30000 0004 6475 7299Centre for Genomic Regulation (CRG), The Barcelona Institute of Science and Technology, Barcelona, Spain; grid.4991.50000 0004 1936 8948Radcliffe Department of Medicine, University of Oxford, Oxford, UK; grid.14709.3b0000 0004 1936 8649Canadian Center for Computational Genomics, McGill University, Montreal, QC Canada; grid.14709.3b0000 0004 1936 8649Department of Human Genetics, McGill University, Montreal, QC Canada; grid.19006.3e0000 0000 9632 6718Department of Human Genetics, University of California Los Angeles, Los Angeles, CA USA; grid.17063.330000 0001 2157 2938Department of Pharmacology, University of Toronto, Toronto, ON Canada; grid.412330.70000 0004 0628 2985Faculty of Medicine and Health Technology, Tampere University and Tays Cancer Center, Tampere University Hospital, Tampere, Finland; grid.415967.80000 0000 9965 1030Haematology, Leeds Teaching Hospitals NHS Trust, Leeds, UK; grid.418116.b0000 0001 0200 3174Translational Research and Innovation, Centre Léon Bérard, Lyon, France; grid.249335.a0000 0001 2218 7820Fox Chase Cancer Center, Philadelphia, PA USA; grid.17703.320000000405980095International Agency for Research on Cancer, World Health Organization, Lyon, France; grid.421605.40000 0004 0447 4123Earlham Institute, Norwich, UK; grid.8273.e0000 0001 1092 7967Norwich Medical School, University of East Anglia, Norwich, UK; grid.5590.90000000122931605Department of Molecular Biology, Faculty of Science, Radboud Institute for Molecular Life Sciences, Radboud University, Nijmegen, HB The Netherlands; grid.17063.330000 0001 2157 2938Department of Radiation Oncology, University of Toronto, Toronto, ON Canada; grid.5379.80000000121662407Division of Cancer Sciences, Manchester Cancer Research Centre, University of Manchester, Manchester, UK; grid.415224.40000 0001 2150 066XRadiation Medicine Program, Princess Margaret Cancer Centre, Toronto, ON Canada; grid.38142.3c000000041936754XDepartment of Pathology, Brigham and Women’s Hospital, Harvard Medical School, Boston, MA USA; grid.21107.350000 0001 2171 9311Department of Surgery, Division of Thoracic Surgery, The Johns Hopkins University School of Medicine, Baltimore, MD USA; grid.430814.a0000 0001 0674 1393Division of Molecular Pathology, The Netherlands Cancer Institute, Oncode Institute, Amsterdam, CX The Netherlands; grid.205975.c0000 0001 0740 6917Department of Biomolecular Engineering, University of California Santa Cruz, Santa Cruz, CA USA; grid.205975.c0000 0001 0740 6917UC Santa Cruz Genomics Institute, University of California Santa Cruz, Santa Cruz, CA USA; grid.7497.d0000 0004 0492 0584Division of Applied Bioinformatics, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.7497.d0000 0004 0492 0584German Cancer Consortium (DKTK), German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.461742.20000 0000 8855 0365National Center for Tumor Diseases (NCT) Heidelberg, Heidelberg, Germany; grid.5170.30000 0001 2181 8870Center for Biological Sequence Analysis, Department of Bio and Health Informatics, Technical University of Denmark, Lyngby, Denmark; grid.5254.60000 0001 0674 042XNovo Nordisk Foundation Center for Protein Research, University of Copenhagen, Copenhagen, Denmark; grid.1003.20000 0000 9320 7537Institute for Molecular Bioscience, University of Queensland, St. Lucia, Brisbane, QLD Australia; grid.5288.70000 0000 9758 5690Biomedical Engineering, Oregon Health and Science University, Portland, OR USA; grid.7497.d0000 0004 0492 0584Division of Theoretical Bioinformatics, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.7700.00000 0001 2190 4373Institute of Pharmacy and Molecular Biotechnology and BioQuant, Heidelberg University, Heidelberg, Germany; grid.5586.e0000 0004 0639 2885Federal Ministry of Education and Research, Berlin, Germany; grid.1013.30000 0004 1936 834XMelanoma Institute Australia, University of Sydney, Sydney, NSW Australia; grid.16149.3b0000 0004 0551 4246Pediatric Hematology and Oncology, University Hospital Muenster, Muenster, Germany; grid.21107.350000 0001 2171 9311Department of Pathology, Johns Hopkins University School of Medicine, Baltimore, MD USA; grid.21107.350000 0001 2171 9311McKusick-Nathans Institute of Genetic Medicine, Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins University School of Medicine, Baltimore, MD USA; grid.418158.10000 0004 0534 4718Foundation Medicine, Inc, Cambridge, MA USA; grid.168010.e0000000419368956Department of Biomedical Data Science, Stanford University School of Medicine, Stanford, CA USA; grid.168010.e0000000419368956Department of Genetics, Stanford University School of Medicine, Stanford, CA USA; grid.266102.10000 0001 2297 6811Bakar Computational Health Sciences Institute and Department of Pediatrics, University of California, San Francisco, CA USA; grid.5510.10000 0004 1936 8921Institute of Clinical Medicine, Faculty of Medicine, University of Oslo, Oslo, Norway; grid.94365.3d0000 0001 2297 5165National Cancer Institute, National Institutes of Health, Bethesda, MD USA; grid.5072.00000 0001 0304 893XRoyal Marsden NHS Foundation Trust, London and Sutton, UK; grid.4709.a0000 0004 0495 846XGenome Biology Unit, European Molecular Biology Laboratory (EMBL), Heidelberg, Germany; grid.5335.00000000121885934Department of Oncology, University of Cambridge, Cambridge, UK; grid.5335.00000000121885934Li Ka Shing Centre, Cancer Research UK Cambridge Institute, University of Cambridge, Cambridge, UK; grid.14925.3b0000 0001 2284 9388Institut Gustave Roussy, Villejuif, France; grid.24029.3d0000 0004 0383 8386Cambridge University Hospitals NHS Foundation Trust, Cambridge, UK; grid.5335.00000000121885934Department of Haematology, University of Cambridge, Cambridge, UK; grid.5841.80000 0004 1937 0247Anatomia Patológica, Hospital Clinic, Institut d’Investigacions Biomèdiques August Pi i Sunyer (IDIBAPS), University of Barcelona, Barcelona, Spain; grid.451322.30000 0004 1770 9462Spanish Ministry of Science and Innovation, Madrid, Spain; grid.412590.b0000 0000 9081 2336University of Michigan Comprehensive Cancer Center, Ann Arbor, MI USA; grid.5734.50000 0001 0726 5157Department for BioMedical Research, University of Bern, Bern, Switzerland; grid.5734.50000 0001 0726 5157Department of Medical Oncology, Inselspital, University Hospital and University of Bern, Bern, Switzerland; grid.5734.50000 0001 0726 5157Graduate School for Cellular and Biomedical Sciences, University of Bern, Bern, Switzerland; grid.8982.b0000 0004 1762 5736University of Pavia, Pavia, Italy; grid.265892.20000000106344187University of Alabama at Birmingham, Birmingham, AL USA; grid.417184.f0000 0001 0661 1177UHN Program in BioSpecimen Sciences, Toronto General Hospital, Toronto, ON Canada; grid.59734.3c0000 0001 0670 2351Department of Urology, Icahn School of Medicine at Mount Sinai, New York, NY USA; grid.1009.80000 0004 1936 826XCentre for Law and Genetics, University of Tasmania, Sandy Bay Campus, Hobart, TAS Australia; grid.7700.00000 0001 2190 4373Faculty of Biosciences, Heidelberg University, Heidelberg, Germany; grid.28046.380000 0001 2182 2255Department of Biochemistry, Microbiology and Immunology, Faculty of Medicine, University of Ottawa, Ottawa, ON Canada; grid.66875.3a0000 0004 0459 167XDivision of Anatomic Pathology, Mayo Clinic, Rochester, MN USA; grid.94365.3d0000 0001 2297 5165Division of Cancer Epidemiology and Genetics, National Cancer Institute, National Institutes of Health, Bethesda, MD USA; grid.417154.20000 0000 9781 7439Illawarra Shoalhaven Local Health District L3 Illawarra Cancer Care Centre, Wollongong Hospital, Wollongong, NSW Australia; BioForA, French National Institute for Agriculture, Food, and Environment (INRAE), ONF, Orléans, France; grid.21107.350000 0001 2171 9311Department of Biostatistics, Bloomberg School of Public Health, Johns Hopkins University, Baltimore, MD USA; grid.266100.30000 0001 2107 4242University of California San Diego, San Diego, CA USA; grid.66875.3a0000 0004 0459 167XDivision of Experimental Pathology, Mayo Clinic, Rochester, MN USA; grid.1013.30000 0004 1936 834XCentre for Cancer Research, The Westmead Institute for Medical Research, University of Sydney, Sydney, NSW Australia; grid.413252.30000 0001 0180 6477Department of Gynaecological Oncology, Westmead Hospital, Sydney, NSW Australia; PDXen Biosystems Inc, Seoul, South Korea; grid.37172.300000 0001 2292 0500Korea Advanced Institute of Science and Technology, Daejeon, South Korea; grid.36303.350000 0000 9148 4899Electronics and Telecommunications Research Institute, Daejeon, South Korea; grid.455095.80000 0001 2189 059XInstitut National du Cancer (INCA), Boulogne-Billancourt, France; grid.265892.20000000106344187Department of Genetics, Informatics Institute, University of Alabama at Birmingham, Birmingham, AL USA; grid.410724.40000 0004 0620 9745Division of Medical Oncology, National Cancer Centre, Singapore, Singapore; grid.411475.20000 0004 1756 948XMedical Oncology, University and Hospital Trust of Verona, Verona, Italy; grid.412468.d0000 0004 0646 2097Department of Pediatrics, University Hospital Schleswig-Holstein, Kiel, Germany; grid.231844.80000 0004 0474 0428Hepatobiliary/Pancreatic Surgical Oncology Program, University Health Network, Toronto, ON Canada; grid.9654.e0000 0004 0372 3343School of Biological Sciences, University of Auckland, Auckland, New Zealand; grid.1008.90000 0001 2179 088XDepartment of Surgery, University of Melbourne, Parkville, VIC Australia; grid.416107.50000 0004 0614 0346The Murdoch Children’s Research Institute, Royal Children’s Hospital, Parkville, VIC Australia; grid.1042.70000 0004 0432 4889Walter and Eliza Hall Institute, Parkville, VIC Australia; grid.412541.70000 0001 0684 7796Vancouver Prostate Centre, Vancouver, Canada; grid.416166.20000 0004 0473 9881Lunenfeld-Tanenbaum Research Institute, Mount Sinai Hospital, Toronto, ON Canada; grid.8273.e0000 0001 1092 7967University of East Anglia, Norwich, UK; grid.240367.40000 0004 0445 7876Norfolk and Norwich University Hospital NHS Trust, Norwich, UK; grid.433802.e0000 0004 0465 4247Victorian Institute of Forensic Medicine, Southbank, VIC Australia; grid.38142.3c000000041936754XDepartment of Biomedical Informatics, Harvard Medical School, Boston, MA USA; grid.5335.00000000121885934Department of Chemistry, Centre for Molecular Science Informatics, University of Cambridge, Cambridge, UK; grid.38142.3c000000041936754XLudwig Center at Harvard Medical School, Boston, MA USA; grid.39382.330000 0001 2160 926XHuman Genome Sequencing Center, Baylor College of Medicine, Houston, TX USA; grid.1008.90000 0001 2179 088XPeter MacCallum Cancer Centre, University of Melbourne, Melbourne, VIC Australia; grid.32224.350000 0004 0386 9924Physics Division, Optimization and Systems Biology Lab, Massachusetts General Hospital, Boston, MA USA; grid.39382.330000 0001 2160 926XDepartment of Medicine, Baylor College of Medicine, Houston, TX USA; grid.6190.e0000 0000 8580 3777University of Cologne, Cologne, Germany; grid.450294.e0000 0004 0641 0756International Genomics Consortium, Phoenix, AZ USA; grid.419890.d0000 0004 0626 690XGenomics Research Program, Ontario Institute for Cancer Research, Toronto, ON Canada; grid.439436.f0000 0004 0459 7289Barking Havering and Redbridge University Hospitals NHS Trust, Romford, UK; grid.1013.30000 0004 1936 834XChildren’s Hospital at Westmead, University of Sydney, Sydney, NSW Australia; grid.411475.20000 0004 1756 948XDepartment of Medicine, Section of Endocrinology, University and Hospital Trust of Verona, Verona, Italy; grid.51462.340000 0001 2171 9952Computational Biology Center, Memorial Sloan Kettering Cancer Center, New York, NY USA; grid.5801.c0000 0001 2156 2780Department of Biology, ETH Zurich, Zürich, Switzerland; grid.5801.c0000 0001 2156 2780Department of Computer Science, ETH Zurich, Zurich, Switzerland; grid.419765.80000 0001 2223 3006SIB Swiss Institute of Bioinformatics, Lausanne, Switzerland; grid.5386.8000000041936877XWeill Cornell Medical College, New York, NY USA; grid.5335.00000000121885934Academic Department of Medical Genetics, University of Cambridge, Addenbrooke’s Hospital, Cambridge, UK; grid.415041.5MRC Cancer Unit, University of Cambridge, Cambridge, UK; grid.10698.360000000122483208Departments of Pediatrics and Genetics, University of North Carolina at Chapel Hill, Chapel Hill, NC USA; grid.492568.4Seven Bridges Genomics, Charlestown, MA USA; Annai Systems, Inc, Carlsbad, CA USA; grid.5608.b0000 0004 1757 3470Department of Pathology, General Hospital of Treviso, Department of Medicine, University of Padua, Treviso, Italy; grid.9851.50000 0001 2165 4204Department of Computational Biology, University of Lausanne, Lausanne, Switzerland; grid.8591.50000 0001 2322 4988Department of Genetic Medicine and Development, University of Geneva Medical School, Geneva, CH Switzerland; grid.8591.50000 0001 2322 4988Swiss Institute of Bioinformatics, University of Geneva, Geneva, CH Switzerland; grid.451388.30000 0004 1795 1830The Francis Crick Institute, London, UK; grid.5596.f0000 0001 0668 7884University of Leuven, Leuven, Belgium; grid.10392.390000 0001 2190 1447Institute of Medical Genetics and Applied Genomics, University of Tübingen, Tübingen, Germany; grid.418377.e0000 0004 0620 715XComputational and Systems Biology, Genome Institute of Singapore, Singapore, Singapore; grid.4280.e0000 0001 2180 6431School of Computing, National University of Singapore, Singapore, Singapore; grid.4991.50000 0004 1936 8948Big Data Institute, Li Ka Shing Centre, University of Oxford, Oxford, UK; grid.451388.30000 0004 1795 1830Biomedical Data Science Laboratory, Francis Crick Institute, London, UK; grid.83440.3b0000000121901201Bioinformatics Group, Department of Computer Science, University College London, London, UK; grid.17063.330000 0001 2157 2938The Edward S. Rogers Sr. Department of Electrical and Computer Engineering, University of Toronto, Toronto, ON Canada; grid.418119.40000 0001 0684 291XBreast Cancer Translational Research Laboratory JC Heuson, Institut Jules Bordet, Brussels, Belgium; grid.5596.f0000 0001 0668 7884Department of Oncology, Laboratory for Translational Breast Cancer Research, KU Leuven, Leuven, Belgium; grid.473715.30000 0004 6475 7299Institute for Research in Biomedicine (IRB Barcelona), The Barcelona Institute of Science and Technology, Barcelona, Spain; grid.5612.00000 0001 2172 2676Research Program on Biomedical Informatics, Universitat Pompeu Fabra, Barcelona, Spain; grid.415224.40000 0001 2150 066XDivision of Medical Oncology, Princess Margaret Cancer Centre, Toronto, ON Canada; grid.5386.8000000041936877XDepartment of Physiology and Biophysics, Weill Cornell Medicine, New York, NY USA; grid.5386.8000000041936877XInstitute for Computational Biomedicine, Weill Cornell Medicine, New York, NY USA; grid.415596.a0000 0004 0440 3018Department of Pathology, UPMC Shadyside, Pittsburgh, PA USA; Independent Consultant, Wellesley, USA; grid.8993.b0000 0004 1936 9457Department of Cell and Molecular Biology, Science for Life Laboratory, Uppsala University, Uppsala, Sweden; grid.4367.60000 0001 2355 7002Department of Medicine and Department of Genetics, Washington University School of Medicine, St. Louis, St. Louis, MO USA; grid.256896.60000 0001 0395 8562Hefei University of Technology, Anhui, China; grid.5284.b0000 0001 0790 3681Translational Cancer Research Unit, GZA Hospitals St.-Augustinus, Center for Oncological Research, Faculty of Medicine and Health Sciences, University of Antwerp, Antwerp, Belgium; grid.61971.380000 0004 1936 7494Simon Fraser University, Burnaby, BC Canada; grid.25879.310000 0004 1936 8972University of Pennsylvania, Philadelphia, PA USA; grid.440820.aFaculty of Science and Technology, University of Vic—Central University of Catalonia (UVic-UCC), Vic, Spain; grid.52788.300000 0004 0427 7672The Wellcome Trust, London, UK; grid.42327.300000 0004 0473 9646The Hospital for Sick Children, Toronto, ON Canada; grid.511123.50000 0004 5988 7216Department of Pathology, Queen Elizabeth University Hospital, Glasgow, UK; grid.1049.c0000 0001 2294 1395Department of Genetics and Computational Biology, QIMR Berghofer Medical Research Institute, Brisbane, QLD Australia; grid.5335.00000000121885934Department of Oncology, Centre for Cancer Genetic Epidemiology, University of Cambridge, Cambridge, UK; grid.5335.00000000121885934Department of Public Health and Primary Care, Centre for Cancer Genetic Epidemiology, University of Cambridge, Cambridge, UK; grid.453281.90000 0004 4652 6665Prostate Cancer Canada, Toronto, ON Canada; grid.5335.00000000121885934University of Cambridge, Cambridge, UK; grid.4514.40000 0001 0930 2361Department of Laboratory Medicine, Translational Cancer Research, Lund University Cancer Center at Medicon Village, Lund University, Lund, Sweden; grid.7700.00000 0001 2190 4373Heidelberg University, Heidelberg, Germany; grid.6363.00000 0001 2218 4662New BIH Digital Health Center, Berlin Institute of Health (BIH) and Charité – Universitätsmedizin Berlin, Berlin, Germany; grid.466571.70000 0004 1756 6246CIBER Epidemiología y Salud Pública (CIBERESP), Madrid, Spain; Research Group on Statistics, Econometrics and Health (GRECS), UdG, Barcelona, Spain; Quantitative Genomics Laboratories (qGenomics), Barcelona, Spain; grid.507118.a0000 0001 0329 4954Icelandic Cancer Registry, Icelandic Cancer Society, Reykjavik, Iceland; grid.233520.50000 0004 1761 4404State Key Laboratory of Cancer Biology, and Xijing Hospital of Digestive Diseases, Fourth Military Medical University, Shaanxi, China; grid.5608.b0000 0004 1757 3470Department of Medicine (DIMED), Surgical Pathology Unit, University of Padua, Padua, Italy; grid.475435.4Rigshospitalet, Copenhagen, Denmark; grid.94365.3d0000 0001 2297 5165Center for Cancer Genomics, National Cancer Institute, National Institutes of Health, Bethesda, MD USA; grid.14848.310000 0001 2292 3357Department of Biochemistry and Molecular Medicine, University of Montreal, Montreal, QC Canada; grid.1011.10000 0004 0474 1797Australian Institute of Tropical Health and Medicine, James Cook University, Douglas, QLD Australia; Department of Neuro-Oncology, Istituto Neurologico Besta, Milano, Italy; grid.484025.fBioplatforms Australia, North Ryde, NSW Australia; grid.83440.3b0000000121901201Department of Pathology (Research), University College London Cancer Institute, London, UK; grid.415224.40000 0001 2150 066XDepartment of Surgical Oncology, Princess Margaret Cancer Centre, Toronto, ON Canada; grid.5645.2000000040459992XDepartment of Medical Oncology, Josephine Nefkens Institute and Cancer Genomics Centre, Erasmus Medical Center, Rotterdam, CN The Netherlands; grid.415184.d0000 0004 0614 0266The University of Queensland Thoracic Research Centre, The Prince Charles Hospital, Brisbane, QLD Australia; grid.5808.50000 0001 1503 7226CIBIO/InBIO – Research Center in Biodiversity and Genetic Resources, Universidade do Porto, Vairão, Portugal; grid.420746.30000 0001 1887 2462HCA Laboratories, London, UK; grid.10025.360000 0004 1936 8470University of Liverpool, Liverpool, UK; grid.22098.310000 0004 1937 0503The Azrieli Faculty of Medicine, Bar-Ilan University, Safed, Israel; grid.15276.370000 0004 1936 8091Department of Neurosurgery, University of Florida, Gainesville, FL USA; grid.26999.3d0000 0001 2151 536XDepartment of Pathology, Graduate School of Medicine, University of Tokyo, Tokyo, Japan; grid.7563.70000 0001 2174 1754University of Milano Bicocca, Monza, Italy; grid.21155.320000 0001 2034 1839BGI-Shenzhen, Shenzhen, China; grid.55325.340000 0004 0389 8485Department of Pathology, Oslo University Hospital Ulleval, Oslo, Norway; grid.38142.3c000000041936754XCenter for Biomedical Informatics, Harvard Medical School, Boston, MA USA; grid.5841.80000 0004 1937 0247Department Biochemistry and Molecular Biomedicine, University of Barcelona, Barcelona, Spain; grid.94365.3d0000 0001 2297 5165Office of Cancer Genomics, National Cancer Institute, National Institutes of Health, Bethesda, MD USA; grid.7497.d0000 0004 0492 0584Cancer Epigenomics, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.240145.60000 0001 2291 4776Department of Cancer Biology, The University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.240145.60000 0001 2291 4776Department of Surgical Oncology, The University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.47100.320000000419368710Department of Computer Science, Yale University, New Haven, CT USA; grid.47100.320000000419368710Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT USA; grid.47100.320000000419368710Program in Computational Biology and Bioinformatics, Yale University, New Haven, CT USA; grid.32224.350000 0004 0386 9924Center for Cancer Research, Massachusetts General Hospital, Boston, MA USA; grid.32224.350000 0004 0386 9924Department of Pathology, Massachusetts General Hospital, Boston, MA USA; grid.51462.340000 0001 2171 9952Department of Pathology, Memorial Sloan Kettering Cancer Center, New York, NY USA; grid.66875.3a0000 0004 0459 167XDivision of Gastroenterology and Hepatology, Mayo Clinic, Rochester, MN USA; grid.1013.30000 0004 1936 834XUniversity of Sydney, Sydney, NSW Australia; grid.4991.50000 0004 1936 8948University of Oxford, Oxford, UK; grid.5335.00000000121885934Department of Surgery, Academic Urology Group, University of Cambridge, Cambridge, UK; grid.8379.50000 0001 1958 8658Department of Medicine II, University of Würzburg, Wuerzburg, Germany; grid.26790.3a0000 0004 1936 8606Sylvester Comprehensive Cancer Center, University of Miami, Miami, FL USA; grid.20522.370000 0004 1767 9005Institut Hospital del Mar d’Investigacions Mèdiques (IMIM), Barcelona, Spain; grid.280664.e0000 0001 2110 5790Genome Integrity and Structural Biology Laboratory, National Institute of Environmental Health Sciences (NIEHS), Durham, NC USA; grid.425213.3St. Thomas’s Hospital, London, UK; Osaka International Cancer Center, Osaka, Japan; grid.411843.b0000 0004 0623 9987Department of Pathology, Skåne University Hospital, Lund University, Lund, Sweden; grid.422301.60000 0004 0606 0717Department of Medical Oncology, Beatson West of Scotland Cancer Centre, Glasgow, UK; grid.94365.3d0000 0001 2297 5165National Human Genome Research Institute, National Institutes of Health, Bethesda, MD USA; grid.1008.90000 0001 2179 088XCentre for Cancer Research, Victorian Comprehensive Cancer Centre, University of Melbourne, Melbourne, VIC Australia; grid.170205.10000 0004 1936 7822Department of Medicine, Section of Hematology/Oncology, University of Chicago, Chicago, IL USA; grid.452463.2German Center for Infection Research (DZIF), Partner Site Hamburg-Borstel-Lübeck-Riems, Hamburg, Germany; grid.7048.b0000 0001 1956 2722Bioinformatics Research Centre (BiRC), Aarhus University, Aarhus, Denmark; grid.410865.eDepartment of Biotechnology, Ministry of Science and Technology, Government of India, New Delhi, Delhi India; grid.410724.40000 0004 0620 9745National Cancer Centre Singapore, Singapore, Singapore; grid.253264.40000 0004 1936 9473Brandeis University, Waltham, MA USA; grid.17091.3e0000 0001 2288 9830Department of Urologic Sciences, University of British Columbia, Vancouver, BC Canada; grid.168010.e0000000419368956Department of Internal Medicine, Stanford University, Stanford, CA USA; grid.267308.80000 0000 9206 2401The University of Texas Health Science Center at Houston, Houston, TX USA; grid.7445.20000 0001 2113 8111Imperial College NHS Trust, Imperial College, London, INY UK; grid.7839.50000 0004 1936 9721Senckenberg Institute of Pathology, University of Frankfurt Medical School, Frankfurt, Germany; grid.266100.30000 0001 2107 4242Department of Medicine, Division of Biomedical Informatics, UC San Diego School of Medicine, San Diego, CA USA; grid.468222.8Center for Precision Health, School of Biomedical Informatics, The University of Texas Health Science Center, Houston, TX USA; Oxford Nanopore Technologies, New York, NY USA; grid.26999.3d0000 0001 2151 536XInstitute of Medical Science, University of Tokyo, Tokyo, Japan; grid.205975.c0000 0001 0740 6917Howard Hughes Medical Institute, University of California Santa Cruz, Santa Cruz, CA USA; grid.412857.d0000 0004 1763 1087Wakayama Medical University, Wakayama, Japan; grid.10698.360000000122483208Department of Internal Medicine, Division of Medical Oncology, Lineberger Comprehensive Cancer Center, University of North Carolina at Chapel Hill, Chapel Hill, NC USA; grid.267301.10000 0004 0386 9246University of Tennessee Health Science Center for Cancer Research, Memphis, TN USA; grid.412346.60000 0001 0237 2025Department of Histopathology, Salford Royal NHS Foundation Trust, Salford, UK; grid.5379.80000000121662407Faculty of Biology, Medicine and Health, University of Manchester, Manchester, UK; grid.11135.370000 0001 2256 9319BIOPIC, ICG and College of Life Sciences, Peking University, Beijing, China; grid.11135.370000 0001 2256 9319Peking-Tsinghua Center for Life Sciences, Peking University, Beijing, China; grid.239552.a0000 0001 0680 8770Children’s Hospital of Philadelphia, Philadelphia, PA USA; grid.240145.60000 0001 2291 4776Department of Bioinformatics and Computational Biology and Department of Systems Biology, The University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.4714.60000 0004 1937 0626Karolinska Institute, Stockholm, Sweden; grid.17063.330000 0001 2157 2938The Donnelly Centre, University of Toronto, Toronto, ON Canada; grid.256753.00000 0004 0470 5964Department of Medical Genetics, College of Medicine, Hallym University, Chuncheon, South Korea; grid.5612.00000 0001 2172 2676Department of Experimental and Health Sciences, Institute of Evolutionary Biology (UPF-CSIC), Universitat Pompeu Fabra, Barcelona, Spain; grid.411941.80000 0000 9194 7179Health Data Science Unit, University Clinics, Heidelberg, Germany; grid.32224.350000 0004 0386 9924Massachusetts General Hospital Center for Cancer Research, Charlestown, MA USA; grid.39158.360000 0001 2173 7691Hokkaido University, Sapporo, Japan; grid.272242.30000 0001 2168 5385Department of Pathology and Clinical Laboratory, National Cancer Center Hospital, Tokyo, Japan; grid.10698.360000000122483208Department of Genetics, University of North Carolina at Chapel Hill, Chapel Hill, NC USA; grid.418245.e0000 0000 9999 5706Computational Biology, Leibniz Institute on Aging – Fritz Lipmann Institute (FLI), Jena, Germany; grid.1008.90000 0001 2179 088XUniversity of Melbourne Centre for Cancer Research, Melbourne, VIC Australia; grid.266813.80000 0001 0666 4105University of Nebraska Medical Center, Omaha, NE USA; Syntekabio Inc, Daejeon, South Korea; grid.5650.60000000404654431Department of Pathology, Academic Medical Center, Amsterdam, AZ The Netherlands; grid.507779.b0000 0004 4910 5858China National GeneBank-Shenzhen, Shenzhen, China; grid.7497.d0000 0004 0492 0584Division of Molecular Genetics, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.24515.370000 0004 1937 1450Division of Life Science and Applied Genomics Center, Hong Kong University of Science and Technology, Clear Water Bay, Hong Kong, China; grid.59734.3c0000 0001 0670 2351Icahn School of Medicine at Mount Sinai, New York, NY USA; Geneplus-Shenzhen, Shenzhen, China; grid.43169.390000 0001 0599 1243School of Computer Science and Technology, Xi’an Jiaotong University, Xi’an, China; grid.431072.30000 0004 0572 4227AbbVie, North Chicago, IL USA; grid.6363.00000 0001 2218 4662Institute of Pathology, Charité – University Medicine Berlin, Berlin, Germany; grid.248762.d0000 0001 0702 3000Centre for Translational and Applied Genomics, British Columbia Cancer Agency, Vancouver, BC Canada; grid.418716.d0000 0001 0709 1919Edinburgh Royal Infirmary, Edinburgh, UK; grid.419491.00000 0001 1014 0849Berlin Institute for Medical Systems Biology, Max Delbrück Center for Molecular Medicine, Berlin, Germany; grid.5253.10000 0001 0328 4908Department of Pediatric Immunology, Hematology and Oncology, University Hospital, Heidelberg, Germany; grid.7497.d0000 0004 0492 0584German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.482664.aHeidelberg Institute for Stem Cell Technology and Experimental Medicine (HI-STEM), Heidelberg, Germany; grid.5386.8000000041936877XInstitute for Computational Biomedicine, Weill Cornell Medical College, New York, NY USA; grid.429884.b0000 0004 1791 0895New York Genome Center, New York, NY USA; grid.21107.350000 0001 2171 9311Department of Urology, James Buchanan Brady Urological Institute, Johns Hopkins University School of Medicine, Baltimore, MD USA; grid.26999.3d0000 0001 2151 536XDepartment of Preventive Medicine, Graduate School of Medicine, The University of Tokyo, Tokyo, Japan; grid.39382.330000 0001 2160 926XDepartment of Molecular and Cellular Biology, Baylor College of Medicine, Houston, TX USA; grid.39382.330000 0001 2160 926XDepartment of Pathology and Immunology, Baylor College of Medicine, Houston, TX USA; grid.413890.70000 0004 0420 5521Michael E. DeBakey Veterans Affairs Medical Center, Houston, TX USA; grid.5170.30000 0001 2181 8870Technical University of Denmark, Lyngby, Denmark; grid.49606.3d0000 0001 1364 9317Department of Pathology, College of Medicine, Hanyang University, Seoul, South Korea; grid.8756.c0000 0001 2193 314XAcademic Unit of Surgery, School of Medicine, College of Medical, Veterinary and Life Sciences, University of Glasgow, Glasgow Royal Infirmary, Glasgow, UK; grid.267370.70000 0004 0533 4667Department of Pathology, Asan Medical Center, College of Medicine, Ulsan University, Songpa-gu, Seoul South Korea; Science Writer, Garrett Park, MD USA; grid.419890.d0000 0004 0626 690XInternational Cancer Genome Consortium (ICGC)/ICGC Accelerating Research in Genomic Oncology (ARGO) Secretariat, Ontario Institute for Cancer Research, Toronto, ON Canada; grid.8954.00000 0001 0721 6013University of Ljubljana, Ljubljana, Slovenia; grid.170205.10000 0004 1936 7822Department of Public Health Sciences, University of Chicago, Chicago, IL USA; grid.240372.00000 0004 0400 4439Research Institute, NorthShore University HealthSystem, Evanston, IL USA; grid.5734.50000 0001 0726 5157Department for Biomedical Research, University of Bern, Bern, Switzerland; grid.411640.6Centre of Genomics and Policy, McGill University and Génome Québec Innovation Centre, Montreal, QC Canada; grid.10698.360000000122483208Carolina Center for Genome Sciences, University of North Carolina at Chapel Hill, Chapel Hill, NC USA; grid.510964.fHopp Children’s Cancer Center (KiTZ), Heidelberg, Germany; grid.7497.d0000 0004 0492 0584Pediatric Glioma Research Group, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.11485.390000 0004 0422 0975Cancer Research UK, London, UK; Indivumed GmbH, Hamburg, Germany; Genome Integration Data Center, Syntekabio, Inc, Daejeon, South Korea; grid.412004.30000 0004 0478 9977University Hospital Zurich, Zurich, Switzerland; grid.419765.80000 0001 2223 3006Clinical Bioinformatics, Swiss Institute of Bioinformatics, Geneva, Switzerland; grid.412004.30000 0004 0478 9977Institute for Pathology and Molecular Pathology, University Hospital Zurich, Zurich, Switzerland; grid.7400.30000 0004 1937 0650Institute of Molecular Life Sciences, University of Zurich, Zurich, Switzerland; grid.4305.20000 0004 1936 7988MRC Human Genetics Unit, MRC IGMM, University of Edinburgh, Edinburgh, UK; grid.50956.3f0000 0001 2152 9905Women’s Cancer Program at the Samuel Oschin Comprehensive Cancer Institute, Cedars-Sinai Medical Center, Los Angeles, CA USA; grid.4808.40000 0001 0657 4636Department of Biology, Bioinformatics Group, Division of Molecular Biology, Faculty of Science, University of Zagreb, Zagreb, Croatia; grid.412468.d0000 0004 0646 2097Department for Internal Medicine II, University Hospital Schleswig-Holstein, Kiel, Germany; grid.414733.60000 0001 2294 430XGenetics and Molecular Pathology, SA Pathology, Adelaide, SA Australia; grid.272242.30000 0001 2168 5385Department of Gastric Surgery, National Cancer Center Hospital, Tokyo, Japan; grid.272242.30000 0001 2168 5385Department of Bioinformatics, Division of Cancer Genomics, National Cancer Center Research Institute, Tokyo, Japan; grid.435025.50000 0004 0619 6198A.A. Kharkevich Institute of Information Transmission Problems, Moscow, Russia; grid.465331.6Oncology and Immunology, Dmitry Rogachev National Research Center of Pediatric Hematology, Moscow, Russia; grid.454320.40000 0004 0555 3608Skolkovo Institute of Science and Technology, Moscow, Russia; grid.253615.60000 0004 1936 9510Department of Surgery, The George Washington University, School of Medicine and Health Science, Washington, DC USA; grid.48336.3a0000 0004 1936 8075Endocrine Oncology Branch, Center for Cancer Research, National Cancer Institute, National Institutes of Health, Bethesda, MD USA; grid.1004.50000 0001 2158 5405Melanoma Institute Australia, Macquarie University, Sydney, NSW Australia; grid.116068.80000 0001 2341 2786MIT Computer Science and Artificial Intelligence Laboratory, Massachusetts Institute of Technology, Cambridge, MA USA; grid.413249.90000 0004 0385 0051Tissue Pathology and Diagnostic Oncology, Royal Prince Alfred Hospital, Sydney, NSW Australia; grid.9786.00000 0004 0470 0856Cholangiocarcinoma Screening and Care Program and Liver Fluke and Cholangiocarcinoma Research Centre, Faculty of Medicine, Khon Kaen University, Khon Kaen, Thailand; Controlled Department and Institution, New York, NY USA; grid.5386.8000000041936877XEnglander Institute for Precision Medicine, Weill Cornell Medicine, New York, NY USA; grid.410914.90000 0004 0628 9810National Cancer Center, Gyeonggi, South Korea; grid.255649.90000 0001 2171 7754Department of Biochemistry, College of Medicine, Ewha Womans University, Seoul, South Korea; grid.266100.30000 0001 2107 4242Health Sciences Department of Biomedical Informatics, University of California San Diego, La Jolla, CA USA; grid.410914.90000 0004 0628 9810Research Core Center, National Cancer Centre Korea, Goyang-si, South Korea; grid.264381.a0000 0001 2181 989XDepartment of Health Sciences and Technology, Sungkyunkwan University School of Medicine, Seoul, South Korea; Samsung Genome Institute, Seoul, South Korea; grid.417747.60000 0004 0460 3896Breast Oncology Program, Dana-Farber/Brigham and Women’s Cancer Center, Boston, MA USA; grid.51462.340000 0001 2171 9952Department of Surgery, Memorial Sloan Kettering Cancer Center, New York, NY USA; grid.62560.370000 0004 0378 8294Division of Breast Surgery, Brigham and Women’s Hospital, Boston, MA USA; grid.280664.e0000 0001 2110 5790Integrative Bioinformatics Support Group, National Institute of Environmental Health Sciences (NIEHS), Durham, NC USA; grid.7914.b0000 0004 1936 7443Department of Clinical Science, University of Bergen, Bergen, Norway; grid.412484.f0000 0001 0302 820XCenter For Medical Innovation, Seoul National University Hospital, Seoul, South Korea; grid.412484.f0000 0001 0302 820XDepartment of Internal Medicine, Seoul National University Hospital, Seoul, South Korea; grid.413454.30000 0001 1958 0162Institute of Computer Science, Polish Academy of Sciences, Warsawa, Poland; grid.7497.d0000 0004 0492 0584Functional and Structural Genomics, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.94365.3d0000 0001 2297 5165Laboratory of Translational Genomics, Division of Cancer Epidemiology and Genetics, National Cancer Institute, , National Institutes of Health, Bethesda, MD USA; grid.9647.c0000 0004 7669 9786Institute for Medical Informatics Statistics and Epidemiology, University of Leipzig, Leipzig, Germany; grid.240145.60000 0001 2291 4776Morgan Welch Inflammatory Breast Cancer Research Program and Clinic, The University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.7450.60000 0001 2364 4210Department of Hematology and Oncology, Georg-Augusts-University of Göttingen, Göttingen, Germany; grid.5718.b0000 0001 2187 5445Institute of Cell Biology (Cancer Research), University of Duisburg-Essen, Essen, Germany; grid.420545.20000 0004 0489 3985King’s College London and Guy’s and St. Thomas’ NHS Foundation Trust, London, UK; grid.251017.00000 0004 0406 2057Center for Epigenetics, Van Andel Research Institute, Grand Rapids, MI USA; grid.416100.20000 0001 0688 4634The University of Queensland Centre for Clinical Research, Royal Brisbane and Women’s Hospital, Herston, QLD Australia; grid.6190.e0000 0000 8580 3777Department of Pediatric Oncology and Hematology, University of Cologne, Cologne, Germany; grid.411327.20000 0001 2176 9917University of Düsseldorf, Düsseldorf, Germany; grid.418119.40000 0001 0684 291XDepartment of Pathology, Institut Jules Bordet, Brussels, Belgium; grid.8761.80000 0000 9919 9582Institute of Biomedicine, Sahlgrenska Academy at University of Gothenburg, Gothenburg, Sweden; grid.414235.50000 0004 0619 2154Children’s Medical Research Institute, Sydney, NSW Australia; ILSbio, LLC Biobank, Chestertown, MD USA; grid.2515.30000 0004 0378 8438Division of Genetics and Genomics, Boston Children’s Hospital, Harvard Medical School, Boston, MA USA; grid.49606.3d0000 0001 1364 9317Institute for Bioengineering and Biopharmaceutical Research (IBBR), Hanyang University, Seoul, South Korea; grid.205975.c0000 0001 0740 6917Department of Statistics, University of California Santa Cruz, Santa Cruz, CA USA; grid.482251.80000 0004 0633 7958National Genotyping Center, Institute of Biomedical Sciences, Academia Sinica, Taipei, Taiwan; grid.419538.20000 0000 9071 0620Department of Vertebrate Genomics/Otto Warburg Laboratory Gene Regulation and Systems Biology of Cancer, Max Planck Institute for Molecular Genetics, Berlin, Germany; grid.411640.6McGill University and Genome Quebec Innovation Centre, Montreal, QC Canada; grid.431797.fbiobyte solutions GmbH, Heidelberg, Germany; grid.137628.90000 0004 1936 8753Gynecologic Oncology, NYU Laura and Isaac Perlmutter Cancer Center, New York University, New York, NY USA; grid.4367.60000 0001 2355 7002Division of Oncology, Stem Cell Biology Section, Washington University School of Medicine, St. Louis, MO USA; grid.240145.60000 0001 2291 4776Department of Systems Biology, The University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.38142.3c000000041936754XHarvard University, Cambridge, MA USA; grid.48336.3a0000 0004 1936 8075Urologic Oncology Branch, Center for Cancer Research, National Cancer Institute, National Institutes of Health, Bethesda, MD USA; grid.5510.10000 0004 1936 8921University of Oslo, Oslo, Norway; grid.17063.330000 0001 2157 2938University of Toronto, Toronto, ON Canada; grid.11135.370000 0001 2256 9319Peking University, Beijing, China; grid.11135.370000 0001 2256 9319School of Life Sciences, Peking University, Beijing, China; grid.419407.f0000 0004 4665 8158Leidos Biomedical Research, Inc, McLean, VA USA; grid.5841.80000 0004 1937 0247Hematology, Hospital Clinic, Institut d’Investigacions Biomèdiques August Pi i Sunyer (IDIBAPS), University of Barcelona, Barcelona, Spain; grid.73113.370000 0004 0369 1660Second Military Medical University, Shanghai, China; Chinese Cancer Genome Consortium, Shenzhen, China; grid.414350.70000 0004 0447 1045Department of Medical Oncology, Beijing Hospital, Beijing, China; grid.412474.00000 0001 0027 0586Laboratory of Molecular Oncology, Key Laboratory of Carcinogenesis and Translational Research (Ministry of Education), Peking University Cancer Hospital and Institute, Beijing, China; grid.11914.3c0000 0001 0721 1626School of Medicine/School of Mathematics and Statistics, University of St. Andrews, St, Andrews, Fife UK; grid.64212.330000 0004 0463 2320Institute for Systems Biology, Seattle, WA USA; Department of Biochemistry and Molecular Biology, Faculty of Medicine, University Institute of Oncology-IUOPA, Oviedo, Spain; grid.476460.70000 0004 0639 0505Institut Bergonié, Bordeaux, France; grid.5335.00000000121885934Cancer Unit, MRC University of Cambridge, Cambridge, UK; grid.239546.f0000 0001 2153 6013Department of Pathology and Laboratory Medicine, Center for Personalized Medicine, Children’s Hospital Los Angeles, Los Angeles, CA USA; grid.1001.00000 0001 2180 7477John Curtin School of Medical Research, Canberra, ACT Australia; MVZ Department of Oncology, PraxisClinic am Johannisplatz, Leipzig, Germany; grid.5342.00000 0001 2069 7798Department of Information Technology, Ghent University, Ghent, Belgium; grid.5342.00000 0001 2069 7798Department of Plant Biotechnology and Bioinformatics, Ghent University, Ghent, Belgium; grid.240344.50000 0004 0392 3476Institute for Genomic Medicine, Nationwide Children’s Hospital, Columbus, OH USA; grid.5288.70000 0000 9758 5690Computational Biology Program, School of Medicine, Oregon Health and Science University, Portland, OR USA; grid.26009.3d0000 0004 1936 7961Department of Surgery, Duke University, Durham, NC USA; grid.425902.80000 0000 9601 989XInstitució Catalana de Recerca i Estudis Avançats (ICREA), Barcelona, Spain; grid.7080.f0000 0001 2296 0625Institut Català de Paleontologia Miquel Crusafont, Universitat Autònoma de Barcelona, Barcelona, Spain; grid.8756.c0000 0001 2193 314XUniversity of Glasgow, Glasgow, UK; grid.10403.360000000091771775Institut d’Investigacions Biomèdiques August Pi i Sunyer (IDIBAPS), Barcelona, Spain; grid.4367.60000 0001 2355 7002Division of Oncology, Washington University School of Medicine, St. Louis, MO USA; grid.7445.20000 0001 2113 8111Department of Surgery and Cancer, Imperial College, London, INY UK; grid.437060.60000 0004 0567 5138Applications Department, Oxford Nanopore Technologies, Oxford, UK; grid.266102.10000 0001 2297 6811Department of Obstetrics, Gynecology and Reproductive Services, University of California San Francisco, San Francisco, CA USA; grid.27860.3b0000 0004 1936 9684Department of Biochemistry and Molecular Medicine, University California at Davis, Sacramento, CA USA; grid.415224.40000 0001 2150 066XSTTARR Innovation Facility, Princess Margaret Cancer Centre, Toronto, ON Canada; grid.1029.a0000 0000 9939 5719Discipline of Surgery, Western Sydney University, Penrith, NSW Australia; grid.47100.320000000419368710Yale School of Medicine, Yale University, New Haven, CT USA; grid.10698.360000000122483208Department of Genetics, Lineberger Comprehensive Cancer Center, University of North Carolina at Chapel Hill, Chapel Hill, NC USA; grid.413103.40000 0001 2160 8953Departments of Neurology and Neurosurgery, Henry Ford Hospital, Detroit, MI USA; grid.5288.70000 0000 9758 5690Precision Oncology, OHSU Knight Cancer Institute, Oregon Health and Science University, Portland, OR USA; grid.13648.380000 0001 2180 3484Institute of Pathology, University Medical Center Hamburg-Eppendorf, Hamburg, Germany; grid.177174.30000 0001 2242 4849Department of Health Sciences, Faculty of Medical Sciences, Kyushu University, Fukuoka, Japan; grid.461593.c0000 0001 1939 6592Heidelberg Academy of Sciences and Humanities, Heidelberg, Germany; grid.1008.90000 0001 2179 088XDepartment of Clinical Pathology, University of Melbourne, Melbourne, VIC, Australia; grid.240614.50000 0001 2181 8635Department of Pathology, Roswell Park Cancer Institute, Buffalo, NY USA; grid.7737.40000 0004 0410 2071Department of Computer Science, University of Helsinki, Helsinki, Finland; grid.7737.40000 0004 0410 2071Institute of Biotechnology, University of Helsinki, Helsinki, Finland; grid.7737.40000 0004 0410 2071Organismal and Evolutionary Biology Research Programme, University of Helsinki, Helsinki, Finland; grid.4367.60000 0001 2355 7002Department of Obstetrics and Gynecology, Division of Gynecologic Oncology, Washington University School of Medicine, St. Louis, MO USA; grid.430183.d0000 0004 6354 3547Penrose St. Francis Health Services, Colorado Springs, CO USA; grid.410712.10000 0004 0473 882XInstitute of Pathology, Ulm University and University Hospital of Ulm, Ulm, Germany; grid.272242.30000 0001 2168 5385National Cancer Center, Tokyo, Japan; grid.418377.e0000 0004 0620 715XGenome Institute of Singapore, Singapore, Singapore; grid.47100.32000000041936871032Program in Computational Biology and Bioinformatics, Yale University, New Haven, CT USA; grid.453370.60000 0001 2161 6363German Cancer Aid, Bonn, Germany; grid.428397.30000 0004 0385 0924Programme in Cancer and Stem Cell Biology, Centre for Computational Biology, Duke-NUS Medical School, Singapore, Singapore; grid.10784.3a0000 0004 1937 0482The Chinese University of Hong Kong, Shatin, NT, Hong Kong China; grid.233520.50000 0004 1761 4404Fourth Military Medical University, Shaanxi, China; grid.5335.00000000121885934The University of Cambridge School of Clinical Medicine, Cambridge, UK; grid.240871.80000 0001 0224 711XSt. Jude Children’s Research Hospital, Memphis, TN USA; grid.415224.40000 0001 2150 066XUniversity Health Network, Princess Margaret Cancer Centre, Toronto, ON Canada; grid.205975.c0000 0001 0740 6917Center for Biomolecular Science and Engineering, University of California Santa Cruz, Santa Cruz, CA USA; grid.170205.10000 0004 1936 7822Department of Medicine, University of Chicago, Chicago, IL USA; grid.66875.3a0000 0004 0459 167XDepartment of Neurology, Mayo Clinic, Rochester, MN USA; grid.24029.3d0000 0004 0383 8386Cambridge Oesophagogastric Centre, Cambridge University Hospitals NHS Foundation Trust, Cambridge, UK; grid.253692.90000 0004 0445 5969Department of Computer Science, Carleton College, Northfield, MN USA; grid.8756.c0000 0001 2193 314XInstitute of Cancer Sciences, College of Medical Veterinary and Life Sciences, University of Glasgow, Glasgow, UK; grid.265892.20000000106344187Department of Epidemiology, University of Alabama at Birmingham, Birmingham, AL USA; grid.417691.c0000 0004 0408 3720HudsonAlpha Institute for Biotechnology, Huntsville, AL USA; grid.265892.20000000106344187O’Neal Comprehensive Cancer Center, University of Alabama at Birmingham, Birmingham, AL USA; grid.26091.3c0000 0004 1936 9959Department of Pathology, Keio University School of Medicine, Tokyo, Japan; grid.272242.30000 0001 2168 5385Department of Hepatobiliary and Pancreatic Oncology, National Cancer Center Hospital, Tokyo, Japan; grid.430406.50000 0004 6023 5303Sage Bionetworks, Seattle, WA USA; grid.410724.40000 0004 0620 9745Lymphoma Genomic Translational Research Laboratory, National Cancer Centre, Singapore, Singapore; grid.416008.b0000 0004 0603 4965Department of Clinical Pathology, Robert-Bosch-Hospital, Stuttgart, Germany; grid.17063.330000 0001 2157 2938Department of Cell and Systems Biology, University of Toronto, Toronto, ON Canada; grid.4714.60000 0004 1937 0626Department of Biosciences and Nutrition, Karolinska Institutet, Stockholm, Sweden; grid.410914.90000 0004 0628 9810Center for Liver Cancer, Research Institute and Hospital, National Cancer Center, Gyeonggi, South Korea; grid.264381.a0000 0001 2181 989XDivision of Hematology-Oncology, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, South Korea; grid.264381.a0000 0001 2181 989XSamsung Advanced Institute for Health Sciences and Technology, Sungkyunkwan University School of Medicine, Seoul, South Korea; grid.263136.30000 0004 0533 2389Cheonan Industry-Academic Collaboration Foundation, Sangmyung University, Cheonan, South Korea; grid.240324.30000 0001 2109 4251NYU Langone Medical Center, New York, NY USA; grid.239578.20000 0001 0675 4725Department of Hematology and Medical Oncology, Cleveland Clinic, Cleveland, OH USA; grid.266102.10000 0001 2297 6811Department of Radiation Oncology, University of California San Francisco, San Francisco, CA USA; grid.66875.3a0000 0004 0459 167XDepartment of Health Sciences Research, Mayo Clinic, Rochester, MN USA; grid.414316.50000 0004 0444 1241Helen F. Graham Cancer Center at Christiana Care Health Systems, Newark, DE USA; grid.5253.10000 0001 0328 4908Heidelberg University Hospital, Heidelberg, Germany; CSRA Incorporated, Fairfax, VA USA; grid.83440.3b0000000121901201Research Department of Pathology, University College London Cancer Institute, London, UK; grid.13097.3c0000 0001 2322 6764Department of Research Oncology, Guy’s Hospital, King’s Health Partners AHSC, King’s College London School of Medicine, London, UK; grid.1004.50000 0001 2158 5405Faculty of Medicine and Health Sciences, Macquarie University, Sydney, NSW Australia; grid.411158.80000 0004 0638 9213University Hospital of Minjoz, INSERM UMR 1098, Besançon, France; grid.7719.80000 0000 8700 1153Spanish National Cancer Research Centre, Madrid, Spain; grid.415180.90000 0004 0540 9980Center of Digestive Diseases and Liver Transplantation, Fundeni Clinical Institute, Bucharest, Romania; Cureline, Inc, South San Francisco, CA USA; grid.412946.c0000 0001 0372 6120St. Luke’s Cancer Centre, Royal Surrey County Hospital NHS Foundation Trust, Guildford, UK; grid.24029.3d0000 0004 0383 8386Cambridge Breast Unit, Addenbrooke’s Hospital, Cambridge University Hospital NHS Foundation Trust and NIHR Cambridge Biomedical Research Centre, Cambridge, UK; grid.416266.10000 0000 9009 9462East of Scotland Breast Service, Ninewells Hospital, Aberdeen, UK; grid.5841.80000 0004 1937 0247Department of Genetics, Microbiology and Statistics, University of Barcelona, IRSJD, IBUB, Barcelona, Spain; grid.30760.320000 0001 2111 8460Department of Obstetrics and Gynecology, Medical College of Wisconsin, Milwaukee, WI USA; grid.516089.30000 0004 9535 5639Hematology and Medical Oncology, Winship Cancer Institute of Emory University, Atlanta, GA USA; grid.16750.350000 0001 2097 5006Department of Computer Science, Princeton University, Princeton, NJ USA; grid.152326.10000 0001 2264 7217Vanderbilt Ingram Cancer Center, Vanderbilt University, Nashville, TN USA; grid.261331.40000 0001 2285 7943Ohio State University College of Medicine and Arthur G. James Comprehensive Cancer Center, Columbus, OH USA; grid.268441.d0000 0001 1033 6139Department of Surgery, Yokohama City University Graduate School of Medicine, Kanagawa, Japan; grid.7497.d0000 0004 0492 0584Division of Chromatin Networks, German Cancer Research Center (DKFZ) and BioQuant, Heidelberg, Germany; grid.10698.360000000122483208Research Computing Center, University of North Carolina at Chapel Hill, Chapel Hill, NC USA; grid.30064.310000 0001 2157 6568School of Molecular Biosciences and Center for Reproductive Biology, Washington State University, Pullman, WA USA; grid.5254.60000 0001 0674 042XFinsen Laboratory and Biotech Research and Innovation Centre (BRIC), University of Copenhagen, Copenhagen, Denmark; grid.17063.330000 0001 2157 2938Department of Laboratory Medicine and Pathobiology, University of Toronto, Toronto, ON Canada; grid.51462.340000 0001 2171 9952Department of Pathology, Human Oncology and Pathogenesis Program, Memorial Sloan Kettering Cancer Center, New York, NY USA; grid.411067.50000 0000 8584 9230University Hospital Giessen, Pediatric Hematology and Oncology, Giessen, Germany; grid.418189.d0000 0001 2175 1768Oncologie Sénologie, ICM Institut Régional du Cancer, Montpellier, France; grid.9764.c0000 0001 2153 9986Institute of Clinical Molecular Biology, Christian-Albrechts-University, Kiel, Germany; grid.8379.50000 0001 1958 8658Institute of Pathology, University of Wuerzburg, Wuerzburg, Germany; grid.418484.50000 0004 0380 7221Department of Urology, North Bristol NHS Trust, Bristol, UK; grid.419385.20000 0004 0620 9905SingHealth, Duke-NUS Institute of Precision Medicine, National Heart Centre Singapore, Singapore, Singapore; grid.17063.330000 0001 2157 2938Department of Computer Science, University of Toronto, Toronto, ON Canada; grid.5734.50000 0001 0726 5157Bern Center for Precision Medicine, University Hospital of Bern, University of Bern, Bern, Switzerland; grid.5386.8000000041936877XEnglander Institute for Precision Medicine, Weill Cornell Medicine and New York Presbyterian Hospital, New York, NY USA; grid.5386.8000000041936877XMeyer Cancer Center, Weill Cornell Medicine, New York, NY USA; grid.5386.8000000041936877XPathology and Laboratory, Weill Cornell Medical College, New York, NY USA; grid.411083.f0000 0001 0675 8654Vall d’Hebron Institute of Oncology: VHIO, Barcelona, Spain; grid.411475.20000 0004 1756 948XGeneral and Hepatobiliary-Biliary Surgery, Pancreas Institute, University and Hospital Trust of Verona, Verona, Italy; grid.22401.350000 0004 0502 9283National Centre for Biological Sciences, Tata Institute of Fundamental Research, Bangalore, India; grid.411377.70000 0001 0790 959XIndiana University, Bloomington, IN USA; grid.428965.40000 0004 7536 2436Department of Pathology, GZA-ZNA Hospitals, Antwerp, Belgium; grid.422639.80000 0004 0372 3861Analytical Biological Services, Inc, Wilmington, DE USA; grid.1013.30000 0004 1936 834XSydney Medical School, University of Sydney, Sydney, NSW Australia; grid.38142.3c000000041936754XcBio Center, Dana-Farber Cancer Institute, Harvard Medical School, Boston, MA USA; grid.38142.3c000000041936754XDepartment of Cell Biology, Harvard Medical School, Boston, MA USA; grid.410869.20000 0004 1766 7522Advanced Centre for Treatment Research and Education in Cancer, Tata Memorial Centre, Navi Mumbai, Maharashtra India; grid.266842.c0000 0000 8831 109XSchool of Environmental and Life Sciences, Faculty of Science, The University of Newcastle, Ourimbah, NSW Australia; grid.410718.b0000 0001 0262 7331Department of Dermatology, University Hospital of Essen, Essen, Germany; grid.7497.d0000 0004 0492 0584Bioinformatics and Omics Data Analytics, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.6363.00000 0001 2218 4662Department of Urology, Charité Universitätsmedizin Berlin, Berlin, Germany; grid.13648.380000 0001 2180 3484Martini-Clinic, Prostate Cancer Center, University Medical Center Hamburg-Eppendorf, Hamburg, Germany; grid.9764.c0000 0001 2153 9986Department of General Internal Medicine, University of Kiel, Kiel, Germany; grid.7497.d0000 0004 0492 0584German Cancer Consortium (DKTK), Partner site Berlin, Berlin, Germany; grid.239395.70000 0000 9011 8547Cancer Research Institute, Beth Israel Deaconess Medical Center, Boston, MA USA; grid.21925.3d0000 0004 1936 9000University of Pittsburgh, Pittsburgh, PA USA; grid.38142.3c000000041936754XDepartment of Ophthalmology and Ocular Genomics Institute, Massachusetts Eye and Ear, Harvard Medical School, Boston, MA USA; grid.240372.00000 0004 0400 4439Center for Psychiatric Genetics, NorthShore University HealthSystem, Evanston, IL USA; grid.251017.00000 0004 0406 2057Van Andel Research Institute, Grand Rapids, MI USA; grid.26999.3d0000 0001 2151 536XLaboratory of Molecular Medicine, Human Genome Center, Institute of Medical Science, University of Tokyo, Tokyo, Japan; grid.480536.c0000 0004 5373 4593Japan Agency for Medical Research and Development, Tokyo, Japan; grid.222754.40000 0001 0840 2678Korea University, Seoul, South Korea; grid.414467.40000 0001 0560 6544Murtha Cancer Center, Walter Reed National Military Medical Center, Bethesda, MD USA; grid.9764.c0000 0001 2153 9986Human Genetics, University of Kiel, Kiel, Germany; grid.38142.3c000000041936754XDepartment of Oncologic Pathology, Dana-Farber Cancer Institute, Harvard Medical School, Boston, MA USA; grid.5288.70000 0000 9758 5690Oregon Health and Science University, Portland, OR USA; grid.240145.60000 0001 2291 4776Center for RNA Interference and Noncoding RNA, The University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.240145.60000 0001 2291 4776Department of Experimental Therapeutics, The University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.240145.60000 0001 2291 4776Department of Gynecologic Oncology and Reproductive Medicine, The University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.15628.380000 0004 0393 1193University Hospitals Coventry and Warwickshire NHS Trust, Coventry, UK; grid.10417.330000 0004 0444 9382Department of Radiation Oncology, Radboud University Nijmegen Medical Centre, Nijmegen, GA The Netherlands; grid.170205.10000 0004 1936 7822Institute for Genomics and Systems Biology, University of Chicago, Chicago, IL USA; grid.459927.40000 0000 8785 9045Clinic for Hematology and Oncology, St.-Antonius-Hospital, Eschweiler, Germany; grid.51462.340000 0001 2171 9952Computational and Systems Biology Program, Memorial Sloan Kettering Cancer Center, New York, NY USA; grid.14013.370000 0004 0640 0021University of Iceland, Reykjavik, Iceland; grid.7497.d0000 0004 0492 0584Division of Computational Genomics and Systems Genetics, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.416266.10000 0000 9009 9462Dundee Cancer Centre, Ninewells Hospital, Dundee, UK; grid.410712.10000 0004 0473 882XDepartment for Internal Medicine III, University of Ulm and University Hospital of Ulm, Ulm, Germany; grid.418596.70000 0004 0639 6384Institut Curie, INSERM Unit 830, Paris, France; grid.268441.d0000 0001 1033 6139Department of Gastroenterology and Hepatology, Yokohama City University Graduate School of Medicine, Kanagawa, Japan; grid.10417.330000 0004 0444 9382Department of Laboratory Medicine, Radboud University Nijmegen Medical Centre, Nijmegen, GA The Netherlands; grid.7497.d0000 0004 0492 0584Division of Cancer Genome Research, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.163555.10000 0000 9486 5048Department of General Surgery, Singapore General Hospital, Singapore, Singapore; grid.4280.e0000 0001 2180 6431Cancer Science Institute of Singapore, National University of Singapore, Singapore, Singapore; grid.7737.40000 0004 0410 2071Department of Medical and Clinical Genetics, Genome-Scale Biology Research Program, University of Helsinki, Helsinki, Finland; grid.24029.3d0000 0004 0383 8386East Anglian Medical Genetics Service, Cambridge University Hospitals NHS Foundation Trust, Cambridge, UK; grid.21729.3f0000000419368729Irving Institute for Cancer Dynamics, Columbia University, New York, NY USA; grid.418812.60000 0004 0620 9243Institute of Molecular and Cell Biology, Singapore, Singapore; grid.410724.40000 0004 0620 9745Laboratory of Cancer Epigenome, Division of Medical Science, National Cancer Centre Singapore, Singapore, Singapore; Universite Lyon, INCa-Synergie, Centre Léon Bérard, Lyon, France; grid.66875.3a0000 0004 0459 167XDepartment of Urology, Mayo Clinic, Rochester, MN USA; grid.416177.20000 0004 0417 7890Royal National Orthopaedic Hospital – Stanmore, Stanmore, Middlesex UK; grid.6312.60000 0001 2097 6738Department of Biochemistry, Genetics and Immunology, University of Vigo, Vigo, Spain; Giovanni Paolo II / I.R.C.C.S. Cancer Institute, Bari, BA Italy; grid.7497.d0000 0004 0492 0584Neuroblastoma Genomics, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.414603.4Fondazione Policlinico Universitario Gemelli IRCCS, Rome, Italy, Rome, Italy; grid.5611.30000 0004 1763 1124University of Verona, Verona, Italy; grid.418135.a0000 0004 0641 3404Centre National de Génotypage, CEA – Institute de Génomique, Evry, France; grid.5012.60000 0001 0481 6099CAPHRI Research School, Maastricht University, Maastricht, ER The Netherlands; grid.418116.b0000 0001 0200 3174Department of Biopathology, Centre Léon Bérard, Lyon, France; grid.7849.20000 0001 2150 7757Université Claude Bernard Lyon 1, Villeurbanne, France; grid.419082.60000 0004 1754 9200Core Research for Evolutional Science and Technology (CREST), JST, Tokyo, Japan; grid.26999.3d0000 0001 2151 536XDepartment of Biological Sciences, Laboratory for Medical Science Mathematics, Graduate School of Science, University of Tokyo, Yokohama, Japan; grid.265073.50000 0001 1014 9130Department of Medical Science Mathematics, Medical Research Institute, Tokyo Medical and Dental University (TMDU), Tokyo, Japan; grid.10306.340000 0004 0606 5382Cancer Ageing and Somatic Mutation Programme, Wellcome Sanger Institute, Hinxton, UK; grid.412563.70000 0004 0376 6589University Hospitals Birmingham NHS Foundation Trust, Birmingham, UK; grid.4777.30000 0004 0374 7521Centre for Cancer Research and Cell Biology, Queen’s University, Belfast, UK; grid.240145.60000 0001 2291 4776Breast Medical Oncology, The University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.21107.350000 0001 2171 9311Department of Surgery, Johns Hopkins University School of Medicine, Baltimore, MD USA; grid.4714.60000 0004 1937 0626Department of Oncology-Pathology, Science for Life Laboratory, Karolinska Institute, Stockholm, Sweden; grid.5491.90000 0004 1936 9297School of Cancer Sciences, Faculty of Medicine, University of Southampton, Southampton, UK; grid.6988.f0000000110107715Department of Gene Technology, Tallinn University of Technology, Tallinn, Estonia; grid.42327.300000 0004 0473 9646Genetics and Genome Biology Program, SickKids Research Institute, The Hospital for Sick Children, Toronto, ON Canada; grid.189967.80000 0001 0941 6502Departments of Neurosurgery and Hematology and Medical Oncology, Winship Cancer Institute and School of Medicine, Emory University, Atlanta, GA USA; grid.5947.f0000 0001 1516 2393Department of Clinical and Molecular Medicine, Faculty of Medicine and Health Sciences, Norwegian University of Science and Technology, Trondheim, Norway; Argmix Consulting, North Vancouver, BC Canada; grid.5342.00000 0001 2069 7798Department of Information Technology, Ghent University, Interuniversitair Micro-Electronica Centrum (IMEC), Ghent, Belgium; grid.4991.50000 0004 1936 8948Nuffield Department of Surgical Sciences, John Radcliffe Hospital, University of Oxford, Oxford, UK; grid.9845.00000 0001 0775 3222Institute of Mathematics and Computer Science, University of Latvia, Riga, LV Latvia; grid.1013.30000 0004 1936 834XDiscipline of Pathology, Sydney Medical School, University of Sydney, Sydney, NSW Australia; grid.5335.00000000121885934Department of Applied Mathematics and Theoretical Physics, Centre for Mathematical Sciences, University of Cambridge, Cambridge, UK; grid.51462.340000 0001 2171 9952Department of Epidemiology and Biostatistics, Memorial Sloan Kettering Cancer Center, New York, NY USA; grid.21729.3f0000000419368729Department of Statistics, Columbia University, New York, NY USA; grid.8993.b0000 0004 1936 9457Department of Immunology, Genetics and Pathology, Science for Life Laboratory, Uppsala University, Uppsala, Sweden; grid.43169.390000 0001 0599 1243School of Electronic and Information Engineering, Xi’an Jiaotong University, Xi’an, China; grid.24029.3d0000 0004 0383 8386Department of Histopathology, Cambridge University Hospitals NHS Foundation Trust, Cambridge, UK; grid.4991.50000 0004 1936 8948Oxford NIHR Biomedical Research Centre, University of Oxford, Oxford, UK; grid.410427.40000 0001 2284 9329Georgia Regents University Cancer Center, Augusta, GA USA; grid.417286.e0000 0004 0422 2524Wythenshawe Hospital, Manchester, UK; grid.4367.60000 0001 2355 7002Department of Genetics, Washington University School of Medicine, St.Louis, MO USA; grid.423940.80000 0001 2188 0463Department of Biological Oceanography, Leibniz Institute of Baltic Sea Research, Rostock, Germany; grid.4991.50000 0004 1936 8948Wellcome Centre for Human Genetics, University of Oxford, Oxford, UK; grid.39382.330000 0001 2160 926XDepartment of Molecular and Human Genetics, Baylor College of Medicine, Houston, TX USA; grid.66875.3a0000 0004 0459 167XThoracic Oncology Laboratory, Mayo Clinic, Rochester, MN USA; grid.66875.3a0000 0004 0459 167XDepartment of Obstetrics and Gynecology, Division of Gynecologic Oncology, Mayo Clinic, Rochester, MN USA; grid.510975.f0000 0004 6004 7353International Institute for Molecular Oncology, Poznań, Poland; grid.22254.330000 0001 2205 0971Poznan University of Medical Sciences, Poznań, Poland; grid.7497.d0000 0004 0492 0584Genomics and Proteomics Core Facility High Throughput Sequencing Unit, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.410724.40000 0004 0620 9745NCCS-VARI Translational Research Laboratory, National Cancer Centre Singapore, Singapore, Singapore; grid.4367.60000 0001 2355 7002Edison Family Center for Genome Sciences and Systems Biology, Washington University, St. Louis, MO USA; grid.301713.70000 0004 0393 3981MRC-University of Glasgow Centre for Virus Research, Glasgow, UK; grid.5288.70000 0000 9758 5690Department of Medical Informatics and Clinical Epidemiology, Division of Bioinformatics and Computational Biology, OHSU Knight Cancer Institute, Oregon Health and Science University, Portland, OR USA; grid.33199.310000 0004 0368 7223School of Electronic Information and Communications, Huazhong University of Science and Technology, Wuhan, China; grid.21107.350000 0001 2171 9311Department of Applied Mathematics and Statistics, Johns Hopkins University, Baltimore, MD USA; grid.136593.b0000 0004 0373 3971Department of Cancer Genome Informatics, Graduate School of Medicine, Osaka University, Osaka, Japan; grid.7700.00000 0001 2190 4373Institute of Computer Science, Heidelberg University, Heidelberg, Germany; grid.1013.30000 0004 1936 834XSchool of Mathematics and Statistics, University of Sydney, Sydney, NSW Australia; grid.170205.10000 0004 1936 7822Ben May Department for Cancer Research, University of Chicago, Chicago, IL USA; grid.170205.10000 0004 1936 7822Department of Human Genetics, University of Chicago, Chicago, IL USA; grid.5386.8000000041936877XTri-Institutional PhD Program in Computational Biology and Medicine, Weill Cornell Medicine, New York, NY USA; grid.43169.390000 0001 0599 1243The First Affiliated Hospital, Xi’an Jiaotong University, Xi’an, China; grid.10784.3a0000 0004 1937 0482Department of Medicine and Therapeutics, The Chinese University of Hong Kong, Shatin, NT, Hong Kong China; grid.240145.60000 0001 2291 4776Department of Biostatistics, The University of Texas MD Anderson Cancer Center, Houston, TX USA; grid.428397.30000 0004 0385 0924Duke-NUS Medical School, Singapore, Singapore; grid.16821.3c0000 0004 0368 8293Department of Surgery, Ruijin Hospital, Shanghai Jiaotong University School of Medicine, Shanghai, China; grid.8756.c0000 0001 2193 314XSchool of Computing Science, University of Glasgow, Glasgow, UK; grid.55325.340000 0004 0389 8485Division of Orthopaedic Surgery, Oslo University Hospital, Oslo, Norway; grid.1002.30000 0004 1936 7857Eastern Clinical School, Monash University, Melbourne, VIC Australia; grid.414539.e0000 0001 0459 5396Epworth HealthCare, Richmond, VIC Australia; grid.38142.3c000000041936754XDepartment of Biostatistics and Computational Biology, Dana-Farber Cancer Institute and Harvard Medical School, Boston, MA USA; grid.261331.40000 0001 2285 7943Department of Biomedical Informatics, College of Medicine, The Ohio State University, Columbus, OH USA; grid.413944.f0000 0001 0447 4797The Ohio State University Comprehensive Cancer Center (OSUCCC – James), Columbus, OH USA; grid.267308.80000 0000 9206 2401The University of Texas School of Biomedical Informatics (SBMI) at Houston, Houston, TX USA; grid.10698.360000000122483208Department of Biostatistics, University of North Carolina at Chapel Hill, Chapel Hill, NC USA; grid.16753.360000 0001 2299 3507Department of Biochemistry and Molecular Genetics, Feinberg School of Medicine, Northwestern University, Chicago, IL USA; grid.1013.30000 0004 1936 834XFaculty of Medicine and Health, University of Sydney, Sydney, NSW Australia; grid.5645.2000000040459992XDepartment of Pathology, Erasmus Medical Center Rotterdam, Rotterdam, GD The Netherlands; grid.430814.a0000 0001 0674 1393Division of Molecular Carcinogenesis, The Netherlands Cancer Institute, Amsterdam, CX The Netherlands; grid.7400.30000 0004 1937 0650Institute of Molecular Life Sciences and Swiss Institute of Bioinformatics, University of Zurich, Zurich, Switzerland
License: © The Author(s) 2020, corrected publication 2022 CC BY 4.0 Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Article links: DOI: 10.1038/s41467-019-14052-x | PubMed: 32024819 | PMC: PMC7002770
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (16.5 MB)
Introduction
Approximately half of all solid tumours are characterized by low levels of molecular oxygen (hypoxia)1–4. Sub-regions of hypoxia can result from disrupted oxygen supply: irregular and disorganized tumour vasculature can reduce oxygen availability5. Hypoxia can also be caused by changes in oxygen demand: altered tumour metabolism6,7 can increase intra-cellular demand for oxygen, potentially extending hypoxia signalling to liquid tumours. The adaptation of tumour cells to this imbalance in oxygen supply and demand is associated with poor clinical prognosis in several cancer types, attributed at least in part to hypoxia-associated genomic instability and clonal selection8–16.
Previous work has provided insight into the molecular origins and consequences of tumour hypoxia and genomic instability. Dynamic cycling of hypoxia can select for cells with TP53 mutations and for those that are apoptosis-deficient17,18. Indeed mutations in TP53 occur at a higher frequency in hypoxic primary tumours of at least 9 types16. The abundance of proteins involved in homologous recombination (e.g. RAD51) and non-homologous end joining (e.g. Ku70) are reduced under hypoxia, and these changes can persist for 2 days after reoxygenation19–21. Genes central to efficient mismatch repair (e.g. MLH1 and MSH2) are also downregulated under hypoxia22,23. Further, co-presence of tumour hypoxia and high genomic instability14,15, specific cellular morphologies like intraductal and cribriform carcinoma24 or specific mutations like loss of PTEN16, synergistically predict for rapid relapse after definitive local therapy in some tumour types, particularly prostate cancer. These data underscore the relationship between hypoxia and DNA repair defects, and suggest the tumour microenvironment applies a selective pressure leading to the development of specific genomic profiles.
We previously evaluated the exomic and copy-number alteration (CNA) consequences of tumour hypoxia across 19 cancer types16. However, the influence of tumour hypoxia on pan-cancer driver alterations, mutational signatures, and subclonal architectures remains unclear. To fill this gap, we calculated tumour hypoxia scores for 1188 tumours with whole-genome sequencing (WGS) and RNA sequencing, spanning 27 cancer types. Genome sequencing data was aggregated by the Pan-Cancer Analysis of Whole Genomes (PCAWG) consortium and generated by the ICGC and TCGA projects. These sequencing data were re-analyzed with standardized, high-accuracy pipelines to align to the human genome (reference build hs37d5) and identify germline variants and somatic mutations, as described previously25. This sequencing data together with our high-quality hypoxia quantitation represents a powerful hypothesis-generating mechanism to suggest useful back-translational in vitro experiments and better define the hypoxia-associated mutator phenotype across cancers. We associated hypoxia with key driver alterations in coding and non-coding regions of the genome, and find hypoxia is associated with specific mutational signatures of unknown aetiology. We illustrate the joint impact of PTEN and the tumour microenvironment in influencing the evolutionary trajectory of tumours. Overall, these data highlight the genomic changes through which hypoxia drives aggressive cancers.
Results
The pan-cancer landscape of tumour hypoxia
We compiled a cohort of 1188 tumours from 27 cancer types via the PCAWG Consortium. All samples had matched tumour and reference normal WGS and tumour RNA sequencing data generated by the ICGC and TCGA projects. WGS25 and RNA-sequencing26 analyses were systematically carried out by centralized teams with consistent and high-accuracy bioinformatics pipelines. Normal reference samples had a mean WGS coverage of 39 reads per base-pair while coverage for tumour samples had a bimodal distribution with modes at 38 and 60 reads per base-pair25. All samples underwent an extensive and systematic quality assurance process25.
We used linear mixed-effect models to associate hypoxia with features of interest across cancers while adjusting for tumour purity, age, and sex27,28. Cancer type was further incorporated as a random effect in every model, allowing us to consider a different baseline value for the feature of interest for each cancer type. As a measure of effect size we report conditional R2 values, denoted as \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\), which reflect the variance explained by the fixed and random factors in each model29. We also report marginal R2 values, denoted as \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\), which reflect the variance explained only by the fixed factors29.
We scored tumour hypoxia in all 1188 tumours using a trio of mRNA-based hypoxia signatures from Buffa30, Winter31 and Ragnum32 (Fig. 1a, Supplementary Fig. 1a, b, Supplementary Table 1, Supplementary Data 1). Hypoxia scores from each of these independent signatures were strongly correlated (ρ = 0.71–0.88, all p < 2.2 × 10−16, AS89; Supplementary Fig. 1c) and consistently predicted squamous tumours of the lung (Lung-SCC), cervix (Cervix-SCC), and head (Head-SCC) as the most hypoxic (Supplementary Fig. 1d, e). Comparatively, chronic lymphocytic leukaemias (Lymph-CLL) and thyroid adenocarcinomas (Thy-AdenoCA) were the least hypoxic, consistent with previous16 reports (ρ = 0.94, p < 2.2 × 10−16, AS89; Fig. 1b, Supplementary Fig. 1f, h). Remarkably, subsets of patients from 23/27 cancer types have tumours with elevated hypoxia (hypoxia score > 0) and tumours consistently have elevated hypoxia compared to normal tissues (Supplementary Fig. 2a–c).
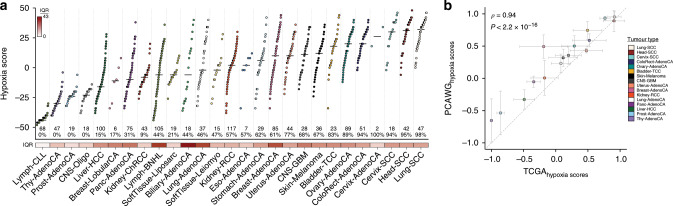
Considering the strong agreement between the Winter, Buffa and Ragnum hypoxia signatures (Fig. 1a, Supplementary Fig. 1c, d), we used the Buffa signature for subsequent analyses. The Buffa signature has been previously used for pan-cancer analyses and shows results consistent with those from other signatures16. We first assessed the degree of inter-tumoural heterogeneity in hypoxia that lies within individual cancer types rather than between them. Over 42% of the variance in hypoxia scores occurs within individual cancer types, highlighting the microenvironmental diversity between tumours arising in a single tissue. This variability in hypoxia score within cancer types was especially elevated in some tumour types, particularly biliary adenocarcinomas (interquartile range, IQR = 43.0; Biliary-AdenoCA), mature B-cell lymphomas (IQR = 36.0; Lymph-BNHL), lung adenocarcinomas (IQR = 34.0; Lung-AdenoCA) and breast adenocarcinomas (IQR = 32.0; Breast-AdenoCA). This was in contrast to chronic lymphocytic leukaemias (IQR = 2.0; Lymph-CLL) and prostate adenocarcinomas (IQR = 6.0; Prost-AdenoCA) where little inter-tumoural variability in hypoxia was observed. The variability in hypoxia score was not significantly associated with the median hypoxia score within cancer types (ρ = 0.20, p = 0.30, AS89; Supplementary Fig. 2d) or with sample size (ρ = 0.22, p = 0.27, AS89; Supplementary Fig. 2e). Overall, extensive heterogeneity exists in hypoxia levels within and across cancer types.
The genomic correlates of tumour hypoxia
To determine whether genomic instability arising from specific mutational classes is associated with hypoxia, we looked to identify hypoxia-associated pan-cancer mutational density and summary features33. As a positive control, we first considered the percentage of the genome with a copy-number aberration (PGA), an engineered feature that is a surrogate for genomic instability and is associated with hypoxia across several tumour types16 (Supplementary Fig. 2f). Indeed, in this diverse pan-cancer cohort, hypoxic tumours have elevated genomic instability while controlling for cancer type, tumour purity, age and sex27 (p = 2.41 × 10−8, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.022, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.57, linear mixed-effect model; Fig. 2a).
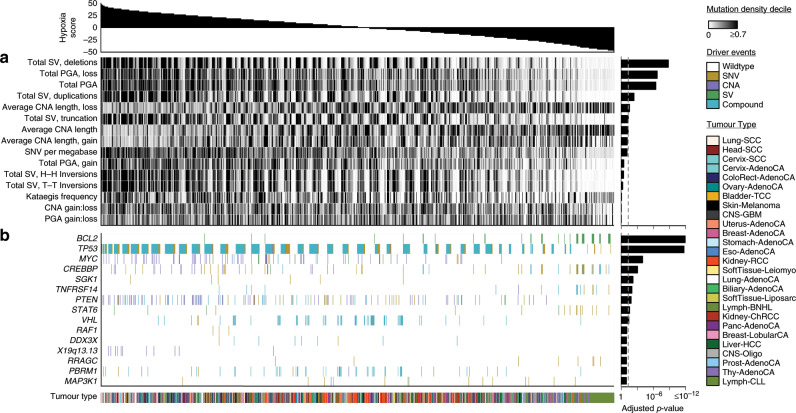
We then considered the association of hypoxia scores with 14 other metrics of the mutation density of CNAs, structural variants (SVs) and single nucleotide variants (SNVs) using linear mixed-effect models (Fig. 2a, Supplementary Fig. 2f, Supplementary Tables 2, 3). The strongest single correlate of tumour hypoxia was the total number of deletions, where patients with elevated hypoxia had more deletions (p = 1.11 × 10−10, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.023, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.59, linear mixed-effect model). Elevated numbers of other SVs such as duplications (p = 2.94 × 10−4, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.0084, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.60, linear mixed-effect model) and truncations (p = 3.29 × 10−3, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.0062, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.60, linear mixed-effect model) were also associated with high hypoxia, and we confirmed this within individual cancer types (Supplementary Fig. 3a). Other features associated with elevated hypoxia include smaller CNAs (p = 3.51 × 10−3, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.0065, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.59, linear mixed-effect model) and more SNVs/Mbp (p = 5.55 × 10−3, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.0054, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.60, linear mixed-effect model). Since mutational density features can be correlated, we wanted to further test if SNVs per megabase were independently associated with hypoxia after adjusting for the total number of deletions. We created a linear mixed-effect model associating hypoxia with the number of SNVs per megabase while adjusting for cancer type, age, sex, tumour purity and the number of deletions. We also created a second model which lacked our feature of interest, SNVs per megabase, and compared the two models using an ANOVA (see the “Methods” section). The p-value for this comparison was 0.011, suggesting that the number of SNVs per megabase are associated with hypoxia independent of the number of deletions (and other potential confounders included in the models). Overall, hypoxia is associated with increased numbers of most types of somatic mutations.
Considering the strong association of hypoxia with mutational density, we next looked to determine if these were only general effects or selectively affected specific genes or chromosome regions. We leveraged a catalogue of 653 driver mutations25, with CNA, SV and SNV data available for 1096 patients. In cases where a patient had multiple mutations in the same gene (e.g. a CNA and an SNV) we denoted these as compound events. We again used linear-mixed effect models to associate hypoxia with each driver feature across cancers (Fig. 2b). Adjusting for cancer type, tumour purity, age and sex, 10 driver events were associated with hypoxia across cancers (FDR < 0.10, linear-mixed effect models; Supplementary Fig. 2f, Supplementary Table 4). Tumours with mutations in BCL2 (FDR = 7.56 × 10−15, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.045, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.62, linear-mixed effect model) showed lower levels of hypoxia compared to those without. All alterations of BCL2 in this cohort were SVs, so it is important to note that this association could not be identified from previous exome-sequencing data. Similarly, mutations in the tumour suppressor TP53 were associated with elevated hypoxia across cancers (FDR = 1.97 × 10−12, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.043, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.59, linear-mixed effect model), consistent with previous descriptions of hypoxia-mediated selection of TP53-mutated cells17 and elevated hypoxia in breast cancers with TP53 mutations16. We also confirmed this association within individual cancer types (Supplementary Fig. 3b). Mutations of the oncogene MYC (FDR = 1.07 × 10−4, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.016, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.60, linear-mixed effect model) and tumour suppressor PTEN (FDR = 1.50 × 10−2, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.0098, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.59, linear mixed-effect model) were also associated with elevated hypoxia. Alterations in mitochondrial genes34 were not significantly associated with tumour hypoxia (Supplementary Fig. 3c). Thus, hypoxia is associated with both broad elevation of mutation density of most types of somatic variation, along with a consistent signature of alterations in oncogenes and tumour suppressors across cancers.
Hypoxia-associated mutational signatures
Previous work has used nonnegative matrix factorization to identify distinct mutational processes in cancer cells from endogenous and exogenous agents35. To identify hypoxia-associated mutational processes, we tested if hypoxia score was associated with the proportion of mutations attributed to each mutational signature using linear-mixed effect models. Of the 65 single base substitution (SBS) signatures tested, nine showed differential activity in hypoxic tumours compared to non-hypoxic ones, while controlling for cancer type, tumour purity, age and sex (FDR < 0.10, linear mixed-effect models; Fig. 3a, Supplementary Table 5). Of these, six were more active and three less active in tumours with elevated hypoxia. Since previous work has shown that DNA repair is impaired under hypoxia, it was not surprising to observe that a higher proportion of mutations were attributed to SBS3 (related to defective homologous recombination-based repair) in tumours with elevated hypoxia score (FDR = 1.98 × 10−3, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.016, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.60, linear-mixed effect model). Further, SBS6 (FDR = 1.98 × 10−3, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.0086, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.61, linear-mixed effect model) and SBS21 (FDR = 4.31 × 10−2, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.0051, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.61, linear-mixed effect model), both related to defective DNA mismatch repair, had a higher proportion of attributed mutations with increasing hypoxia. A lower proportion of mutations were also attributed to SBS1, previously related to the deamination of 5-methylcytosine, with increasing hypoxia (FDR = 8.33 × 10−8, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.033, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.61, linear-mixed effect model).
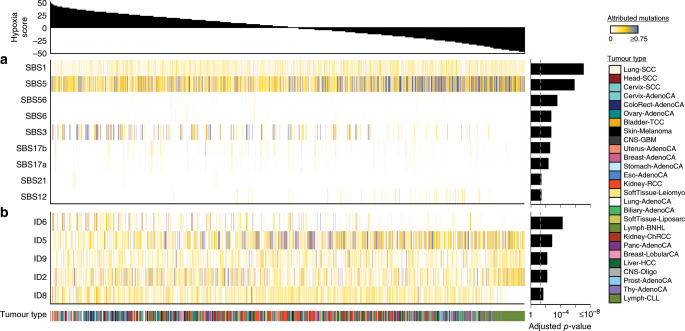
Intriguingly, hypoxia was also associated with a number of SBS signatures with unknown aetiology (Fig. 3b). The strongest of these was SBS5, where elevated hypoxia was associated with a significantly lower proportion of mutations attributed to the signature (FDR = 1.43 × 10−6, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.022, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.59, linear-mixed effect model). A significantly lower proportion of mutations were also attributed to SBS12 with increasing hypoxia score (FDR = 4.31 × 10−2, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.0066, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.60, linear-mixed effect model). In contrast, a higher proportion of mutations were attributed to SBS17a (FDR = 4.80 × 10−3, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.0072, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.61, linear mixed-effect model) and SBS17b (FDR = 2.83 × 10−3, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.079, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.61, linear mixed-effect model) with increasing hypoxia.
Analysis of small insertion and deletion (ID) signatures illustrated a similar story. Of the 17 ID signatures analyzed, the activity of 5 were associated with tumour hypoxia scores while controlling for cancer type, tumour purity, age and sex (FDR < 0.10, linear mixed-effect models; Fig. 3b, Supplementary Table 6). Of these, 3 were more active in tumours with elevated hypoxia while 2 were less active in them. The defective homologous recombination signature ID6 (FDR = 5.76 × 10−5, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.015, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.60, linear mixed-effect model) and defective DNA mismatch repair signature ID2 (FDR = 7.06 × 10−3, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.011, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.61, linear mixed-effect model) had a higher proportion of attributed mutations as hypoxia score increased. Several signatures with unknown aetiology were also significantly associated with hypoxia score, including ID5 (FDR = 1.54 × 10−3, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.016, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.60, linear mixed-effect model) and ID9 (FDR = 7.06 × 10−3, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.0068, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.60, linear mixed-effect model). These data suggest that oxygen levels play a direct or indirect role in the accumulation of specific mutations in cancer cells that are reflected by these signatures.
The subclonal hallmarks of tumour hypoxia
State-of-the-art methods for subclonal reconstruction rely on WGS data36, making the PCAWG dataset ideal for understanding the evolutionary pressures imposed by hypoxia. We and others have shown that some mutations consistently occur early during tumourigenesis while others occur later and that hypoxia is associated with CNAs occurring early in localized prostate cancer16,37,38. To explore if this interaction between the tumour microenvironment and mutational landscape exists more broadly in cancer, we assessed if hypoxia was related to the number of clonal or subclonal mutations across 1188 tumours from 27 cancer types38. Clonal mutations are common to all cells in a tumour while subclonal ones are only present in a subpopulation of cells. We found that elevated hypoxia was significantly associated with an increased number of clonal alterations across cancers (Bonferroni-adjusted p = 4.65 × 10−3, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.0074, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.60, linear mixed-effect model; Fig. 4a, Supplementary Table 7), particularly clonal SVs (p = 1.17 × 10−5, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.013, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.60, linear mixed-effect model). In contrast, tumour hypoxia was not significantly associated with the number of subclonal alterations (Bonferroni-adjusted p = 0.28, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.0039, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.60, linear mixed-effect model; Fig. 4a). Further, consistent with findings in prostate cancer16, hypoxia was not associated with the number of subclones detected (Bonferroni-adjusted p = 0.14, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.0051, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.60, linear mixed-effect model; Fig. 4a). These data suggest that hypoxia applies a selective pressure on tumours during their early evolution, prior to subclonal diversification.
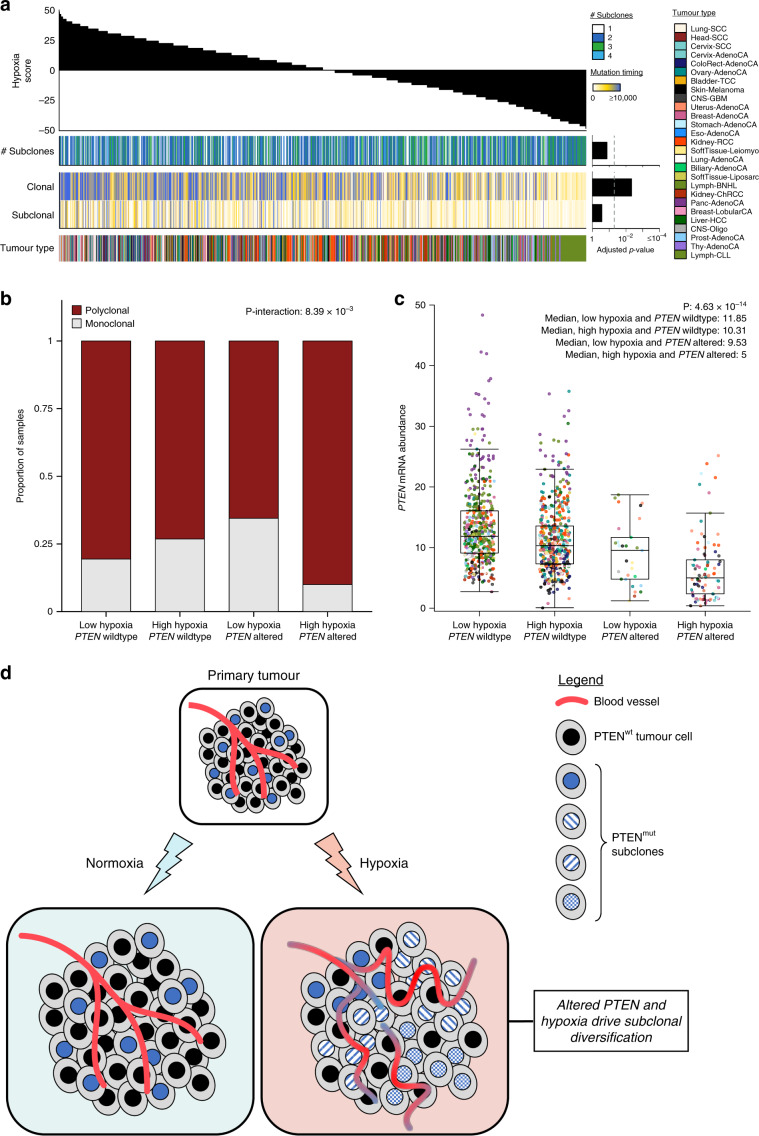
Next, we assessed if the mutational background of a tumour together with its oxygenation level was linked to its evolutionary trajectory. We previously demonstrated that patients with hypoxic polyclonal prostate tumours with loss of the tumour suppressor PTEN tend to have a poor prognosis16. Indeed, here we observed a significant interaction between tumour hypoxia and loss of PTEN in predicting subclonal architecture (pinteraction = 8.39 × 10−3, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.60, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.87, linear mixed-effect model; Fig. 4b). Specifically, tumours with both of these features tend to have a polyclonal architecture across cancers. The downstream impact of this interaction between the genome and the tumour microenvironment was observed in RNA data: tumours with both altered PTEN and elevated hypoxia had the lowest abundance of PTEN mRNA (p = 4.63 × 10−14, \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) = 0.054, \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) = 0.47, linear mixed-effect model; Fig. 4c). Thus, the evolutionary trajectory of a tumour may be driven by the presence of a mutation in a specific microenvironmental context (Fig. 4d).
Discussion
Hypoxia is a feature of many solid and liquid tumours and is associated with aggressive disease. We calculated hypoxia scores for 1188 tumours from 27 cancer types and showed the vast heterogeneity that exists in this microenvironmental feature within and across cancer types. This reinforces previous pushes for careful patient selection in prospective trials of hypoxia-targeting agents16.
For the first time, we characterized the pan-cancer whole-genome correlates of tumour hypoxia. We show the broad influence of the hypoxia-associated mutator phenotype: elevated hypoxia is associated with increased mutational load across all mutational classes (i.e. CNAs, SVs and SNVs). This supports previous in vitro work that demonstrated the contextual synthetic lethality of PARP inhibition in cells with defective DNA repair due to hypoxia39. Regarding this co-occurrence of genomic instability and hypoxia, our group16 and others40 have previously described this metabolic reprogramming as a series of distinct genomic alterations. This is supported by our finding that alterations in TP53, MYC and PTEN are more common in tumours with elevated hypoxia across cancers. Supporting these findings, previous in vitro work has shown that heterogeneous populations of cells where a small subpopulation have mutant TP53 can rapidly expand under cycling hypoxia to become the major subpopulation due to deficient apoptosis and selection17. We have also previously shown that tumours with TP53 mutations have elevated hypoxia within individual breast cancer subtypes, confirming that this association is not simply reflecting previously described molecular subtypes16.
Our study cannot conclusively say whether hypoxia exerts a selective pressure that enriches for specific genomic alterations or if these genomic changes directly result in hypoxia. Experimental studies of single genes support that both effects may contribute to the associations we describe17,22,41–43. While we have not specifically included in vitro experimental validation data in this report, we and others have previously validated associations first revealed by analysis of mRNA-based hypoxia signatures. For example, our group previously described microRNA-133a-3p as a hypoxia-associated miRNA in prostate cancer based on mRNA signature-based associations across multiple independent datasets16. We went on to validate that microRNA-133a-3p was indeed induced under hypoxia in multiple prostate cancer cell lines and confirmed its capacity to modulate cell proliferation and invasion. Similarly, Ye et al. applied hypoxia signatures to 10 independent datasets of cell lines and primary tumour fragments under hypoxia and normoxia44. These 10 datasets represented seven cancer types and within each dataset samples under hypoxia showed higher hypoxia scores compared to normoxic samples. Further, they generated predictions of drugs that would be more or less potent under hypoxia and validated four drug–hypoxia interactions in vitro. These data illustrate that hypoxia signatures applied to large cohorts of primary tumours can generate reliable hypotheses, many of which have been validated in controlled systems. However, some aerobic cancer cells may also mimic the biological state of hypoxia (i.e., pseudohypoxia) and this may affect signature-derived hypoxia estimates. Further, while pimonidazole (which was used to develop the Ragnum hypoxia signature32) reflects oxygen tensions below 10 mmHg (1.3% O2), it is difficult to directly relate hypoxia signature scores with oxygen tension45. Overall, hypoxia signalling can be distinct from microenvironmental hypoxia and this remains a critical caveat of this study.
Diving into the mutational processes related to hypoxia, we confirmed that several SBS and small indel signatures related to impaired DNA repair were associated with hypoxia. This raises the potential confounder that because hypoxic tumours have more mutations, we have more power to detect related mutational signatures. However, we demonstrated that hypoxia is indeed strongly associated with many mutational signatures with unknown aetiology, particularly SBS5, which is found in nearly all cancer types. Modelling these associations in vitro is particularly difficult and these data provide a high confidence measure of the mutational signatures that may be directly or indirectly driven by tumour oxygen levels. It is difficult to disentangle the timing of these events: whether a specific driver mutation gives rise to a specific mutational signature or if these are separate processes. Better mapping of the evolutionary timing of hypoxia will be particularly important in addressing this question and the advent of hypoxia signatures may facilitate future studies in this area.
We observed a significant association between elevated hypoxia and the number of clonal mutations. This supports the idea that hypoxia is an early event in cancer, as we have suggested previously16, and other models that link hypoxia to genomic instability and downstream clonal selection20,42. Previous work has also demonstrated that patients with allelic loss of PTEN and elevated hypoxia rapidly relapse after definitive treatment for localized prostate cancer16. Here, we showed that tumours with alterations in PTEN and elevated hypoxia are enriched for a polyclonal tumour architecture. This illustrates the joint influence of the tumour mutational landscape and microenvironment in guiding evolutionary trajectories across cancers. Further, these data suggest that increased subclonal diversification may be a novel route via which PTEN drives aggressive tumour phenotypes, in concert with tumour hypoxia, and this can be better defined with future back-translational in vitro experiments. The PCAWG dataset is the largest publicly available pan-cancer dataset to date and this limits our ability to validate our discoveries in independent datasets. Ultimately, it will be necessary to validate our findings in large, independent cohorts. Hypotheses generated here, particularly those around hypoxia and tumour evolution, will require long term, systematic in vitro modelling and will be the subject of future studies. Overall, this work shows that a hypoxic tumour microenvironment is associated with specific mutational processes and distinct somatic mutational profiles, and may direct the subclonal architecture of cancers.
Methods
Pan-cancer hypoxia scoring
Hypoxia scores were calculated for all 1188 tumours with mRNA abundance data (FPKM with upper-quartile normalization) using mRNA-abundance-based signatures of tumour hypoxia developed previously by Winter et al.31, Buffa et al. 30 and Ragnum et al. 32, as described previously14,16 (Supplementary Data 1). Briefly, patients with the top 50% of mRNA abundance values for each gene in a signature were given a score of +1. Patients with the bottom 50% of mRNA abundance values for that gene were given a score of −1. This was repeated for every gene in the signature to generate a hypoxia score for each patient, and this process was repeated for each of the three signatures used in the study. High scores suggest that the tumour was hypoxic and low scores are indicative of normoxia.
Hypoxia score comparison
To compare hypoxia scores generated by the different signatures, the median hypoxia score was calculated for each of the PCAWG cancer types based on each signature. The median hypoxia scores from each signature were then scaled from +1 to −1 using the plotrix package (v3.7). Scaled median hypoxia values for the PCAWG cancer types were also compared to scaled median hypoxia values from previously published16 TCGA data between comparable cancer groups. The groups compared are as follows (PCAWG cancer type and TCGA cancer type): Bladder-TCC and BLCA; Breast-AdenoCA and BRCA; Cervix-SCC and CESC; CNS-GBM and GBM; ColoRect-AdenoCA and COADREAD; Head-SCC and HNSC; Kidney-RCC and KIRC; Liver-HCC and LIHC; Lung-AdenoCA and LUAD; Lung-SCC and LUSC; Ovary-AdenoCA and OV; Panc-AdenoCA and PAAD; Prost-AdenoCA and PRAD; Skin-Melanoma and SKCM; Thy-AdenoCA and THCA; Uterus-Adeno and UCEC. Of the 27 cancer types in PCAWG, 11 (Cervix-AdenoCA, Stomach-AdenoCA, Eso-AdenoCA, Breast-LobularCA, SoftTissue-Leiomyo, Lymph-BNHL, SoftTissue-Liposarc, Biliary-AdenoCA, Kidney-ChRCC, CNS-Oligo and Lymph-CLL) did not have hypoxia data from comparable cancers in TCGA and were not used for the comparison16. For Spearman’s correlations, p-values were calculated using algorithm AS89.
Tumour vs. normal hypoxia comparison
Previously calculated pan-cancer tumour hypoxia scores were gathered for 7791 independent tumours from 19 cancer types based on the Buffa hypoxia signature16. Hypoxia scores were then calculated for all normal tissue samples related to the 19 cancer types with hypoxia scores (n = 640 independent normal tissue samples). Tumour hypoxia scores were compared to normal tissue hypoxia scores for each tissue type where at least 15 normal tissue samples were available (Mann–Whitney U-test). A total of 5649 independent tumours and 625 independent normal tissue samples were evaluated in the comparisons.
Linear mixed-effect models
We used linear mixed-effect models to associate hypoxia with features of interest (e.g., PGA, TP53 mutational status, etc.) across cancers using the lme4 package (v1.1-17). For each feature of interest, we compared a full model (i.e., a model with the feature of interest) to a null model (i.e. a model without the feature of interest) using an ANOVA to determine if hypoxia was significantly associated with the feature of interest across cancers. A generic example of this is shown below with Eqs. (1) and (2):
\[
{{{\mathrm{{full}}}}} = {{{\mathrm{{hypoxia}}}}}\sim {{{\mathrm{{feature}}}}} + {{{\mathrm{{purity}}}}} + {{{\mathrm{{age}}}}} + {{{\mathrm{{sex}}}}} + \left( {1\left| {{{{\mathrm{{cancer}}}}}} \right.} \right)
\]
\[
{{{\mathrm{{null}}}}} = {{{\mathrm{{hypoxia}}}}}\sim {{{\mathrm{{purity}}}}} + {{{\mathrm{{age}}}}} + {{{\mathrm{{sex}}}}} + \left( {1|{{{\mathrm{{cancer}}}}}} \right)
\]
All models were adjusted for tumour purity, patient age and sex27,28. Cancer type was incorporated as a random effect in every model. This allowed us to consider a different baseline value for the feature of interest for each cancer type. For each model a conditional R2 value is reported (\(R_{{{\mathrm{{{LMEM – C}}}}}}^2\)) which reflects the variance explained by the fixed and random factors29. We also report marginal R2 values for each model (\(R_{{{\mathrm{{{LMEM – M}}}}}}^2\)) which reflect the variance explained by the fixed factors only29. \(R_{{{\mathrm{{{LMEM – C}}}}}}^2\) and \(R_{{{\mathrm{{{LMEM – M}}}}}}^2\) values were calculated as described previously29.
All model diagnostics were done using the DHARMa package (0.2.0) which uses a simulation-based approach to create standardized residuals46. For each model, scaled residuals were generated using the simulateResiduals function. The full model was used as the input for fittedModel parameter and 1000 simulations were run. For correctly specified models, the scaled residuals were expected to be uniformly distributed and this was verified for each full model. We also compared the standardized residuals to the rank transformed predicted values to assess deviations from uniformity for each full model.
Mutational density analysis
Previously published data for 15 mutational density and summary features were downloaded for 1188 tumours33. We used linear mixed-effect models to associate each feature with hypoxia score across cancers and compared each full model with a null model. Cancer type was incorporated as a random effect in each model while tumour purity, age and sex were incorporated as fixed effects. Tumours belonging to cancer types with fewer than 15 samples were excluded from the analysis. A Bonferroni p-value adjustment was applied to the p-values from linear mixed-effect modelling since fewer than 20 tests were conducted. All models were adjusted for tumour purity based on previously published purity data33. The full model for evaluating PGA is shown below as an example as follows:
\[
{{{\mathrm{{full}}}}}_{{{\mathrm{{{PGA}}}}}} = {{{\mathrm{{hypoxia}}}}}\sim {{{\mathrm{{PGA + purity + age + sex}}}}} + \left( {1|{{{\mathrm{{cancer}}}}}} \right)
\]
To assess if SNVs per megabase were independently associated with hypoxia after adjusting for the total number of deletions we created two linear mixed-effect models. The full model associated hypoxia with SNVs per megabase while adjusting for cancer type, age, sex, tumour purity and the number of deletions (Eq. (4)). For comparison, a null model was created without our feature of interest, SNVs per megabase (Eq. (5)). The two models were compared using an ANOVA.
\[
\begin{array}{l}{{{\mathrm{{full}}}}} = {{{\mathrm{{hypoxia}}}}}\sim {{{\mathrm{{SNVs}}}}}\,{{{\mathrm{{per}}}}}\,{{{\mathrm{{megabase + purity + age + sex}}}}}\\ + \, {{{\mathrm{{total}}}}}\,{{{\mathrm{{deletions}}}}} + \left( {1|{{{\mathrm{{cancer}}}}}} \right)\end{array}
\]
\[
{{{\mathrm{{null}}}}} = {{{\mathrm{{hypoxia}}}}}\sim {{{\mathrm{{purity + age + sex +}}}}} {{{\mathrm{{total}}}}}\, {{{\mathrm{{deletions}}}}} + \left( {1|{{{\mathrm{{cancer}}}}}} \right)
\]
Driver mutations analysis
Data for driver mutations was first summarized at the gene level for 1096 tumours with previously published driver mutation data25. For each of the 653 driver features, we summarized if a patient had an SNV, CNA or SV. Some tumours had more than one type of event in a gene (e.g. a CNA and an SNV) and these events were classified as compound events. We then used linear mixed-effect models to associate the mutational status of each gene with hypoxia score and compared each full model with a null model. Cancer type was incorporated as a random effect in each model while tumour purity, age and sex were incorporated as fixed effects. The driver mutation analysis did not specifically consider the type of mutation in the gene and only considered if the gene had a mutation or was wildtype. Tumours belonging to cancer types with fewer than 15 samples were excluded from the analysis. An FDR adjustment was applied to the p-values from linear mixed-effect modelling. The full model for evaluating PTEN is shown below as an example as follows:
\[
{{{\mathrm{{full}}}}}_{{{\mathrm{{{PTEN}}}}}} = {{{\mathrm{{hypoxia}}}}}\sim {{{\mathrm{{PTEN + purity + age + sex}}}}} + \left( {1|{{{\mathrm{{cancer}}}}}} \right)
\]
Mutational signature analysis
Previously published data for mutations attributed to various specific signatures was downloaded for 1188 tumours35. For each tumour, we calculated the proportion of total mutations attributed to each mutational signature. The proportion of mutations attributed to each signature were calculated by dividing the number of mutations attributed to each signature by the total number of mutations in the tumour. We used linear mixed-effect models to associate the proportion of mutations attributed to each signature with hypoxia score and compared each full model with a null model. Cancer type was incorporated as a random effect in each model while purity, age and sex were incorporated as fixed effects. Tumours belonging to cancer types with fewer than 15 samples were excluded from the analysis. An FDR adjustment was applied to the p-values from linear mixed-effect modelling. The full model for SBS1 is shown below as an example as follows:
\[
{{{\mathrm{{full}}}}}_{{{\mathrm{{{SBS1}}}}}} = {{{\mathrm{{hypoxia}}}}}\sim {{{\mathrm{{SBS1 + purity + age + sex}}}}} + \left( {1|{{{\mathrm{{cancer}}}}}} \right)
\]
Subclonality analysis
Previously reported38 subclonal reconstruction data was used to summarize the number of clonal and subclonal mutations in all 1188 tumours. We used linear mixed-effect models to associate the number of these timed mutations with hypoxia score and compared each full model with a null model. Cancer type was incorporated as a random effect in each model while purity, age and sex were incorporated as fixed effects. Tumours belonging to cancer types with fewer than 15 samples were excluded from the analysis. A Bonferroni adjustment was applied to the p-values from linear mixed-effect modelling since fewer than 20 tests were conducted.
The number of subclones was calculated for all 1188 tumours based on the number of clusters of cells identified in each sample. A linear mixed-effects model was used to associate the number of subclones with hypoxia score and this model was compared to a null model. Cancer type was incorporated as a random effect while purity, age and sex were incorporated as fixed effects. Tumours belonging to cancer types with fewer than 15 samples were excluded from the analysis. The full model for associating the number of subclones with hypoxia score is shown below as follows:
\[
{{{\mathrm{{full}}}}}_{{{\mathrm{{{subclones}}}}}} = {{{\mathrm{{hypoxia}}}}}\sim {{{\mathrm{{subclones + purity + age + sex }}}}}+ \left( {1|{{{\mathrm{{cancer}}}}}} \right)
\]
Patients with only one identified cluster of cells were defined as monoclonal and patients with more than one identified cluster of cells were defined as polyclonal37. Hypoxia scores were median dichotomized to classify patients as hypoxic or normoxic. To test for an interaction between tumour hypoxia and PTEN mutational status in selecting for a particular subclonal architecture, we used linear mixed-effect models together with an ANOVA. An interaction model was first created where the relationship between the hypoxia scores and PTEN mutational status was modelled as an interaction (Eq. (9)). An additive model was also created where the relationship between hypoxia scores and PTEN mutational status was modelled in an additive manner (Eq. (10)):
\[
{{{\mathrm{{interaction}}}}} = {{{\mathrm{{clonality}}}}}\sim {{{\mathrm{{hypoxia}}}}} \ast {{{\mathrm{{PTEN + purity + age + sex}}}}} + \left( {1|{{{\mathrm{{cancer}}}}}} \right)
\]
\[
{{{\mathrm{{additive}}}}} = {{{\mathrm{{clonality}}}}}\sim {{{\mathrm{{hypoxia + PTEN + purity + age + sex}}}}} + \left( {1|{{{\mathrm{{cancer}}}}}} \right)
\]
The two models were compared using an ANOVA to test if hypoxia scores significantly interact with PTEN mutational status. Tumours belonging to cancer types with fewer than 15 samples were excluded from the analysis. The full model diagnostics were carried out using the DHARMa package, as described above.
All data analysis was performed in the R statistical environment (v3.4.3). Data visualization was performed using the BPG package47 (v5.9.1). Figures were compiled using Inkscape (v0.91).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Supplementary Materials
References
- CE Weber, PC Kuo. The tumor microenvironment. Surg. Oncol., 2012. [DOI | PubMed]
- MV Blagosklonny. Antiangiogenic therapy and tumor progression. Cancer Cell, 2004. [DOI | PubMed]
- P Vaupel, O Thews, M Hoeckel. Treatment resistance of solid tumors. Med. Oncol., 2001. [DOI | PubMed]
- N Dhani, A Fyles, D Hedley, M Milosevic. The clinical significance of hypoxia in human cancers. Semin. Nucl. Med., 2015. [DOI | PubMed]
- JM Brown, WR Wilson. Exploiting tumour hypoxia in cancer treatment. Nat. Rev. Cancer, 2004. [DOI | PubMed]
- VE Zannella. Reprogramming metabolism with metformin improves tumor oxygenation and radiotherapy response. Clin. Cancer Res., 2013. [DOI | PubMed]
- V Mucaj, JES Shay, MC Simon. Effects of hypoxia and HIFs on cancer metabolism. Int. J. Hematol., 2012. [DOI | PubMed]
- KMA Rouschop. The unfolded protein response protects human tumor cells during hypoxia through regulation of the autophagy genes MAP1LC3B and ATG5. J. Clin. Invest., 2010. [DOI | PubMed]
- P Lassen. HPV-associated p16-expression and response to hypoxic modification of radiotherapy in head and neck cancer. Radiother. Oncol., 2010. [DOI | PubMed]
- PJ Hoskin, AM Rojas, SM Bentzen, MI Saunders. Radiotherapy with concurrent carbogen and nicotinamide in bladder carcinoma. J. Clin. Oncol., 2010. [DOI | PubMed]
- M Milosevic. Tumor hypoxia predicts biochemical failure following radiotherapy for clinically localized prostate cancer. Clin. Cancer Res., 2012. [DOI | PubMed]
- B Kim. Prognostic assessment of hypoxia and metabolic markers in cervical cancer using automated digital image analysis of immunohistochemistry. J. Transl. Med., 2013. [DOI | PubMed]
- L Yang. Development and validation of a 28-gene hypoxia-related prognostic signature for localized prostate cancer. EBioMedicine, 2018. [DOI | PubMed]
- E Lalonde. Tumour genomic and microenvironmental heterogeneity for integrated prediction of 5-year biochemical recurrence of prostate cancer: a retrospective cohort study. Lancet Oncol., 2014. [DOI | PubMed]
- E Lalonde. Translating a prognostic DNA Genomic Classifier into the clinic: retrospective validation in 563 localized prostate tumors. Eur. Urol., 2017. [DOI | PubMed]
- V Bhandari. Molecular landmarks of tumor hypoxia across cancer types. Nat. Genet., 2019. [DOI | PubMed]
- TG Graeber. Hypoxia-mediated selection of cells with diminished apoptotic potential in solid tumours. Nature, 1996. [DOI | PubMed]
- AE Greijer, E van der Wall. The role of hypoxia inducible factor 1 (HIF-1) in hypoxia induced apoptosis. J. Clin. Pathol., 2004. [DOI | PubMed]
- RS Bindra. Down-regulation of Rad51 and decreased homologous recombination in hypoxic cancer cells. Mol. Cell. Biol., 2004. [DOI | PubMed]
- RG Bristow, RP Hill. Hypoxia and metabolism: hypoxia, DNA repair and genetic instability. Nat. Rev. Cancer, 2008. [DOI | PubMed]
- AX Meng. Hypoxia down-regulates DNA double strand break repair gene expression in prostate cancer cells. Radiother. Oncol., 2005. [DOI | PubMed]
- VT Mihaylova. Decreased expression of the DNA mismatch repair gene Mlh1 under hypoxic stress in mammalian cells. Mol. Cell. Biol., 2003. [DOI | PubMed]
- M Koshiji. HIF-1α induces genetic instability by transcriptionally downregulating MutSα expression. Mol. Cell, 2005. [DOI | PubMed]
- MLK Chua. A prostate cancer “ Nimbosus ”: genomic instability and SChLAP1 dysregulation underpin aggression of intraductal and cribriform subpathologies. Eur. Urol., 2017. [DOI | PubMed]
- 25.The ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature10.1038/s41586-020-1969-6 (2020).
- 26.PCAWG Transcriptome Core Group. et al. Genomic basis for RNA alterations in cancer. Nature10.1038/s41586-020-1970-0 (2020).
- CH Li, S Haider, Y-J Shiah, K Thai, PC Boutros. Sex differences in cancer driver genes and biomarkers. Cancer Res., 2018. [DOI | PubMed]
- 28.Li, C. H. et al. Sex differences in oncogenic mutational processes. Nat. Commun. In Press (2020).
- S Nakagawa, H Schielzeth. A general and simple method for obtaining R2 from generalized linear mixed-effects models. Methods Ecol. Evol., 2013. [DOI]
- FM Buffa, AL Harris, CM West, CJ Miller. Large meta-analysis of multiple cancers reveals a common, compact and highly prognostic hypoxia metagene. Br. J. Cancer, 2010. [DOI | PubMed]
- SC Winter. Relation of a hypoxia metagene derived from head and neck cancer to prognosis of multiple cancers. Cancer Res., 2007. [DOI | PubMed]
- HB Ragnum. The tumour hypoxia marker pimonidazole reflects a transcriptional programme associated with aggressive prostate cancer. Br. J. Cancer, 2015. [DOI | PubMed]
- 33.Yousif, F. et al. The origins and consequences of localized and global somatic hypermutation. Preprint at https://www.biorxiv.org/content/10.1101/287839v2 (2018).
- 34.Yuan, Y. et al. Comprehensive molecular characterization of mitochondrial genomes in human cancers. Nat. Genet. 10.1038/s41588-019-0557-x (2020).
- 35.Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature10.1038/s41586-020-1943-3 (2020).
- AG Deshwar. PhyloWGS: reconstructing subclonal composition and evolution from whole-genome sequencing of tumors. Genome Biol., 2015. [DOI | PubMed]
- SMG Espiritu. The evolutionary landscape of localized prostate cancers drives clinical aggression. Cell, 2018. [DOI | PubMed]
- 38.Gerstung, M. et al. The evolutionary history of 2,658 cancers. Nature10.1038/s41586-019-1907-7 (2020).
- N Chan. Contextual synthetic lethality of cancer cell kill based on the tumor microenvironment. Cancer Res., 2010. [DOI | PubMed]
- S Haider. Genomic alterations underlie a pan-cancer metabolic shift associated with tumour hypoxia. Genome Biol., 2016. [DOI | PubMed]
- AD Nguyen, JG McDonald, RK Bruick, RA DeBose-Boyd. Hypoxia stimulates degradation of 3-hydroxy-3-methylglutaryl-coenzyme A reductase through accumulation of lanosterol and hypoxia-inducible factor-mediated induction of insigs. J. Biol. Chem., 2007. [DOI | PubMed]
- KR Luoto, R Kumareswaran, RG Bristow. Tumor hypoxia as a driving force in genetic instability. Genome Integr., 2013. [DOI | PubMed]
- Y Chen, R Cairns, I Papandreou, A Koong, NC Denko. Oxygen consumption can regulate the growth of tumors, a new perspective on the Warburg effect. PLoS One, 2009. [DOI | PubMed]
- Y Ye. Characterization of hypoxia-associated molecular features to aid hypoxia-targeted therapy. Nat. Metab., 2019. [DOI | PubMed]
- C Rosenberger, S Rosen, A Paliege, SN Heyman. Pimonidazole adduct immunohistochemistry in the rat kidney: detection of tissue hypoxia. Methods Mol. Biol., 2009. [DOI | PubMed]
- PK Dunn, GK Smyth. Randomized quantile residuals. J. Comput. Graph. Stat., 1996
- C P’ng. BPG: seamless, automated and interactive visualization of scientific data. BMC Bioinforma., 2019. [DOI]
