Retrospective evaluation of whole exome and genome mutation calls in 746 cancer samples
Abstract
The Cancer Genome Atlas (TCGA) and International Cancer Genome Consortium (ICGC) curated consensus somatic mutation calls using whole exome sequencing (WES) and whole genome sequencing (WGS), respectively. Here, as part of the ICGC/TCGA Pan-Cancer Analysis of Whole Genomes (PCAWG) Consortium, which aggregated whole genome sequencing data from 2,658 cancers across 38 tumour types, we compare WES and WGS side-by-side from 746 TCGA samples, finding that ~80% of mutations overlap in covered exonic regions. We estimate that low variant allele fraction (VAF < 15%) and clonal heterogeneity contribute up to 68% of private WGS mutations and 71% of private WES mutations. We observe that ~30% of private WGS mutations trace to mutations identified by a single variant caller in WES consensus efforts. WGS captures both ~50% more variation in exonic regions and un-observed mutations in loci with variable GC-content. Together, our analysis highlights technological divergences between two reproducible somatic variant detection efforts.
Article type: Research Article
Keywords: Comparative genomics, Communication and replication, Cancer genomics, Genetic databases
Affiliations: grid.4367.60000 0001 2355 7002The McDonnell Genome Institute at Washington University, St. Louis, MO 63108 USA; grid.4367.60000 0001 2355 7002Division of Oncology, Department of Medicine, Washington University School of Medicine, St. Louis, MO 63108 USA; grid.4367.60000 0001 2355 7002Alvin J. Siteman Cancer Center, Washington University School of Medicine, St. Louis, MO 63108 USA; grid.47100.320000000419368710Yale School of Medicine, Yale University, New Haven, CT 06520 USA; grid.47100.320000000419368710Program in Computational Biology and Bioinformatics, Yale University, New Haven, CT 06520 USA; grid.419890.d0000 0004 0626 690XComputational Biology Program, Ontario Institute for Cancer Research, Toronto, ON M5G 0A3 Canada; grid.42327.300000 0004 0473 9646The Hospital for Sick Children, Toronto, ON M5G 1X8 Canada; grid.240145.60000 0001 2291 4776Department of Bioinformatics and Computational Biology, The University of Texas MD Anderson Cancer Center, Houston, TX 77030 USA; grid.5288.70000 0000 9758 5690Molecular and Medical Genetics, OHSU Knight Cancer Institute, Oregon Health and Science University, Portland, OR 97239 USA; grid.5288.70000 0000 9758 5690Biomedical Engineering, Oregon Health and Science University, Portland, OR 97239 USA; grid.39382.330000 0001 2160 926XHuman Genome Sequencing Center, Baylor College of Medicine, Houston, TX 77030 USA; Castle Biosciences Inc, Friendswood, TX 77546 USA; grid.4367.60000 0001 2355 7002Department of Medicine and Department of Genetics, Washington University School of Medicine, St. Louis, MO 63110 USA; grid.66859.34Broad Institute of MIT and Harvard, Cambridge, MA 02142 USA; grid.38142.3c000000041936754XHarvard Medical School, Boston, MA 02115 USA; grid.32224.350000 0004 0386 9924Center for Cancer Research, Massachusetts General Hospital, Boston, MA 02114 USA; grid.32224.350000 0004 0386 9924Department of Pathology, Massachusetts General Hospital, Boston, MA 02114 USA; grid.205975.c0000 0001 0740 6917UC Santa Cruz Genomics Institute, University of California Santa Cruz, Santa Cruz, CA 95064 USA; grid.205975.c0000 0001 0740 6917Howard Hughes Medical Institute, University of California Santa Cruz, Santa Cruz, CA 95064 USA; grid.32224.350000 0004 0386 9924Massachusetts General Hospital Center for Cancer Research, Charlestown, MA 02114 USA; grid.94365.3d0000 0001 2297 5165National Human Genome Research Institute, National Institutes of Health, Bethesda, MD 20894 USA; grid.51462.340000 0001 2171 9952Marie-Josée and Henry R. Kravis Center for Molecular Oncology, Memorial Sloan Kettering Cancer Center, New York, NY 10065 USA; grid.419890.d0000 0004 0626 690XOntario Institute for Cancer Research, Toronto, ON M5G 0A3 Canada; grid.434706.20000 0004 0410 5424Canada’s Michael Smith Genome Sciences Centre, BC Cancer, Vancouver, BC V5Z 4S6 Canada; grid.5288.70000 0000 9758 5690Computational Biology Program, School of Medicine, Oregon Health and Science University, Portland, OR 97239 USA; DNAnexus Inc, Mountain View, CA 94040 USA; grid.240145.60000 0001 2291 4776Department of Systems Biology, UT MD Anderson Cancer Center, Houston, TX 77030 USA; grid.5288.70000 0000 9758 5690Precision Oncology, OHSU Knight Cancer Institute, Oregon Health and Science University, Portland, OR 97239 USA; grid.9227.e0000000119573309Computer Network Information Center, Chinese Academy of Sciences, Beijing, China; grid.64212.330000 0004 0463 2320Institute for Systems Biology, Seattle, WA 98109 USA; grid.205975.c0000 0001 0740 6917Department of Biomolecular Engineering, University of California Santa Cruz, Santa Cruz, CA 95064 USA; grid.240145.60000 0001 2291 4776Department of Bioinformatics and Computational Biology and Department of Systems Biology, The University of Texas MD Anderson Cancer Center, Houston, TX 77030 USA; grid.39382.330000 0001 2160 926XDepartment of Molecular and Human Genetics, Baylor College of Medicine, Houston, TX 77030 USA; grid.205975.c0000 0001 0740 6917Biomolecular Engineering Department, University of California Santa Cruz, Santa Cruz, CA 95064 USA; grid.43169.390000 0001 0599 1243School of Electronic and Information Engineering, Xi’an Jiaotong University, Xi’an, China; grid.43169.390000 0001 0599 1243The First Affiliated Hospital, Xi’an Jiaotong University, Xi’an, China; grid.484013.aCenter for Digital Health, Berlin Institute of Health and Charitè-Universitätsmedizin Berlin, 10178 Berlin, Germany; grid.7497.d0000 0004 0492 0584Heidelberg Center for Personalized Oncology (DKFZ-HIPO), German Cancer Research Center (DKFZ), 69120 Heidelberg, Germany; grid.17063.330000 0001 2157 2938Department of Medical Biophysics, University of Toronto, Toronto, ON M5S Canada; grid.19006.3e0000 0000 9632 6718Department of Human Genetics, University of California Los Angeles, Los Angeles, CA 90095 USA; grid.17063.330000 0001 2157 2938Department of Pharmacology, University of Toronto, Toronto, ON M5S Canada; grid.7497.d0000 0004 0492 0584Division of Theoretical Bioinformatics, German Cancer Research Center (DKFZ), 69120 Heidelberg, Germany; grid.7700.00000 0001 2190 4373Institute of Pharmacy and Molecular Biotechnology and BioQuant, Heidelberg University, 69117 Heidelberg, Germany; grid.10306.340000 0004 0606 5382Wellcome Sanger Institute, Wellcome Genome Campus, Hinxton, CB10 1SA UK; grid.240145.60000 0001 2291 4776University of Texas MD Anderson Cancer Center, Houston, TX 77030 USA; grid.265892.20000000106344187Department of Genetics, Informatics Institute, University of Alabama at Birmingham, Birmingham, AL 77030 USA; grid.5612.00000 0001 2172 2676Universitat Pompeu Fabra (UPF), 08002 Barcelona, Spain; grid.473715.3Centre for Genomic Regulation (CRG), The Barcelona Institute of Science and Technology, Barcelona, Spain; grid.7700.00000 0001 2190 4373Heidelberg University, 69117 Heidelberg, Germany; grid.484013.aNew BIH Digital Health Center, Berlin Institute of Health (BIH) and Charité-Universitätsmedizin Berlin, 08036 Berlin, Germany; grid.21155.320000 0001 2034 1839BGI-Shenzhen, Shenzhen, China; grid.10097.3f0000 0004 0387 1602Barcelona Supercomputing Center (BSC), 08034 Barcelona, Spain; grid.5841.80000 0004 1937 0247Department Biochemistry and Molecular Biomedicine, University of Barcelona, 08007 Barcelona, Spain; grid.47100.320000000419368710Department of Computer Science, Yale University, New Haven, CT 06520 USA; grid.47100.320000000419368710Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06520 USA; grid.225360.00000 0000 9709 7726European Molecular Biology Laboratory, European Bioinformatics Institute (EMBL-EBI), Cambridge, CB10 1SD UK; grid.4709.a0000 0004 0495 846XGenome Biology Unit, European Molecular Biology Laboratory (EMBL), 22607 Heidelberg, Germany; grid.473715.3CNAG-CRG, Centre for Genomic Regulation (CRG), Barcelona Institute of Science and Technology (BIST), 08036 Barcelona, Spain; grid.412541.70000 0001 0684 7796Vancouver Prostate Centre, Vancouver, BC V6H 3Z6 Canada; grid.17091.3e0000 0001 2288 9830Department of Urologic Sciences, University of British Columbia, Vancouver, BC V6T 1Z4 Canada; grid.24515.370000 0004 1937 1450Division of Life Science and Applied Genomics Center, Hong Kong University of Science and Technology, Clear Water Bay, Hong Kong, China; Geneplus-Shenzhen, Shenzhen, China; grid.43169.390000 0001 0599 1243School of Computer Science and Technology, Xi’an Jiaotong University, Xi’an, China; grid.5253.10000 0001 0328 4908National Center for Tumor Diseases (NCT) Heidelberg, 69120 Heidelberg, Germany; grid.7497.d0000 0004 0492 0584German Cancer Consortium (DKTK), 69120 Heidelberg, Germany; Genome Integration Data Center, Syntekabio, Inc, Daejeon, South Korea; grid.255649.90000 0001 2171 7754Department of Biochemistry, College of Medicine, Ewha Womans University, Seoul, South Korea; grid.431797.fBiobyte solutions GmbH, 69126 Heidelberg, Germany; grid.32224.350000 0004 0386 9924Massachusetts General Hospital, Boston, MA 02114 USA; grid.154185.c0000 0004 0512 597XDepartment of Molecular Medicine (MOMA), Aarhus University Hospital, 8200 Aarhus N, Denmark; grid.10392.390000 0001 2190 1447Institute of Medical Genetics and Applied Genomics, University of Tübingen, 72074 Tübingen, Germany; grid.7048.b0000 0001 1956 2722Bioinformatics Research Centre (BiRC), Aarhus University, 8000 Aarhus, Denmark; grid.419890.d0000 0004 0626 690XGenome Informatics Program, Ontario Institute for Cancer Research, Toronto, ON M5G 0A3 Canada; grid.266102.10000 0001 2297 6811Department of Radiation Oncology, University of California San Francisco, San Francisco, CA 94110 USA; grid.61971.380000 0004 1936 7494Simon Fraser University, Burnaby, BC V5A 1S6 Canada; grid.411377.70000 0001 0790 959XIndiana University, Bloomington, IN 47405 USA; grid.7497.d0000 0004 0492 0584Bioinformatics and Omics Data Analytics, German Cancer Research Center (DKFZ), 69120 Heidelberg, Germany; grid.17063.330000 0001 2157 2938Department of Computer Science, University of Toronto, Toronto, ON M5S Canada; grid.425902.80000 0000 9601 989XInstitució Catalana de Recerca i Estudis Avançats (ICREA), Barcelona, Spain; grid.65499.370000 0001 2106 9910Department of Medical Oncology, Dana-Farber Cancer Institute, Boston, MA 02215 USA; grid.5254.60000 0001 0674 042XFinsen Laboratory and Biotech Research and Innovation Centre (BRIC), University of Copenhagen, 1165 Copenhagen, Denmark; grid.6363.00000 0001 2218 4662Department of Urology, Charité Universitätsmedizin Berlin, 10117 Berlin, Germany; grid.4367.60000 0001 2355 7002Department of Mathematics, Washington University in St. Louis, St. Louis, MO 63130 USA; grid.4367.60000 0001 2355 7002Department of Genetics, Washington University School of Medicine, St.Louis, MO 63110 USA; grid.423940.80000 0001 2188 0463Department of Biological Oceanography, Leibniz Institute of Baltic Sea Research, 18119 Rostock, Germany; grid.51462.340000 0001 2171 9952Department of Epidemiology and Biostatistics, Memorial Sloan Kettering Cancer Center, New York, NY 10065 USA; grid.7737.40000 0004 0410 2071Applied Tumor Genomics Research Program, Research Programs Unit, University of Helsinki, 00100 Helsinki, Finland; grid.51462.340000 0001 2171 9952Memorial Sloan Kettering Cancer Center, New York, NY 10065 USA; grid.26999.3d0000 0001 2151 536XGenome Science Division, Research Center for Advanced Science and Technology, University of Tokyo, Tokyo, 113-8654 Japan; grid.170205.10000 0004 1936 7822Department of Surgery, University of Chicago, Chicago, IL 60637 USA; grid.414067.00000 0004 0647 8419Department of Surgery, Division of Hepatobiliary and Pancreatic Surgery, School of Medicine, Keimyung University Dongsan Medical Center, Daegu, South Korea; grid.256155.00000 0004 0647 2973Department of Oncology, Gil Medical Center, Gachon University, Incheon, South Korea; grid.257022.00000 0000 8711 3200Hiroshima University, Hiroshima, 739-8511 Japan; grid.415310.20000 0001 2191 4301King Faisal Specialist Hospital and Research Centre, Al Maather, Riyadh, Saudi Arabia; grid.7719.80000 0000 8700 1153Bioinformatics Unit, Spanish National Cancer Research Centre (CNIO), 28029 Madrid, Spain; grid.13648.380000 0001 2180 3484Bioinformatics Core Facility, University Medical Center Hamburg, 20251 Hamburg, Germany; grid.418481.00000 0001 0665 103XHeinrich Pette Institute, Leibniz Institute for Experimental Virology, 20251 Hamburg, Germany; grid.419890.d0000 0004 0626 690XOntario Tumour Bank, Ontario Institute for Cancer Research, Toronto, ON M5G 0A3 Canada; grid.240145.60000 0001 2291 4776Department of Pathology, The University of Texas MD Anderson Cancer Center, Houston, TX 77030 USA; grid.48336.3a0000 0004 1936 8075Laboratory of Pathology, Center for Cancer Research, National Cancer Institute, Bethesda, MD 20814 USA; grid.266100.30000 0001 2107 4242Department of Cellular and Molecular Medicine and Department of Bioengineering, University of California San Diego, La Jolla, CA 92093 USA; grid.266100.30000 0001 2107 4242UC San Diego Moores Cancer Center, San Diego, CA 92037 USA; grid.1008.90000 0001 2179 088XSir Peter MacCallum Department of Oncology, Peter MacCallum Cancer Centre, University of Melbourne, Melbourne, VIC 3010 Australia; grid.11794.3a0000000109410645Centre for Research in Molecular Medicine and Chronic Diseases (CiMUS), Universidade de Santiago de Compostela, 15705 Santiago de Compostela, Spain; grid.11794.3a0000000109410645Department of Zoology, Genetics and Physical Anthropology, (CiMUS), Universidade de Santiago de Compostela, 15705 Santiago de Compostela, Spain; grid.6312.60000 0001 2097 6738The Biomedical Research Centre (CINBIO), Universidade de Vigo, 36310 Vigo, Spain; grid.416177.20000 0004 0417 7890Royal National Orthopaedic Hospital-Bolsover, London, 36310 UK; grid.240145.60000 0001 2291 4776Department of Genomic Medicine, The University of Texas MD Anderson Cancer Center, Houston, TX 77030 USA; grid.39382.330000 0001 2160 926XQuantitative and Computational Biosciences Graduate Program, Baylor College of Medicine, Houston, TX 77030 USA; grid.249880.f0000 0004 0374 0039The Jackson Laboratory for Genomic Medicine, Farmington, CT 06032 USA; grid.9764.c0000 0001 2153 9986Institute of Human Genetics, Christian-Albrechts-University, 24118 Kiel, Germany; grid.410712.1Institute of Human Genetics, Ulm University and Ulm University Medical Center, 89081 Ulm, Germany; grid.1003.20000 0000 9320 7537Queensland Centre for Medical Genomics, Institute for Molecular Bioscience, University of Queensland, St. Lucia, Brisbane, QLD 4072 Australia; grid.412346.60000 0001 0237 2025Salford Royal NHS Foundation Trust, Salford, M6 8HD UK; grid.411475.20000 0004 1756 948XDepartment of Surgery, Pancreas Institute, University and Hospital Trust of Verona, 37129 Verona, Italy; grid.248762.d0000 0001 0702 3000Department of Molecular Oncology, BC Cancer Research Centre, Vancouver, BC V5Z 1L3 Canada; grid.83440.3b0000000121901201University College London, London, WC1E 6BT UK; grid.272242.30000 0001 2168 5385Division of Cancer Genomics, National Cancer Center Research Institute, National Cancer Center, Tokyo, 104-0045 Japan; DLR Project Management Agency, Bonn, Germany; grid.410818.40000 0001 0720 6587Tokyo Women’s Medical University, Tokyo, 162-8666 Japan; grid.51462.340000 0001 2171 9952Center for Molecular Oncology, Memorial Sloan Kettering Cancer Center, New York, NY 10065 USA; grid.148313.c0000 0004 0428 3079Los Alamos National Laboratory, Los Alamos, NM 87545 USA; grid.417184.f0000 0001 0661 1177Department of Pathology, University Health Network, Toronto General Hospital, Toronto, ON M5G 2C4 Canada; grid.240404.60000 0001 0440 1889Nottingham University Hospitals NHS Trust, Nottingham, NG5 1PB UK; grid.7497.d0000 0004 0492 0584Epigenomics and Cancer Risk Factors, German Cancer Research Center (DKFZ), 69120 Heidelberg, Germany; grid.17063.330000 0001 2157 2938Department of Molecular Genetics, University of Toronto, Toronto, ON M5S Canada; grid.494618.6Vector Institute, Toronto, ON M5G 1M1 Canada; grid.9764.c0000 0001 2153 9986Hematopathology Section, Institute of Pathology, Christian-Albrechts-University, Kiel, 24118 Germany; grid.10698.360000000122483208Department of Pathology and Laboratory Medicine, School of Medicine, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA; grid.55325.340000 0004 0389 8485Department of Cancer Genetics, Institute for Cancer Research, Oslo University Hospital, The Norwegian Radium Hospital, 0450 Oslo, Norway; grid.5841.80000 0004 1937 0247Pathology, Hospital Clinic, Institut d’Investigacions Biomèdiques August Pi i Sunyer (IDIBAPS), University of Barcelona, 08007 Barcelona, Spain; grid.5335.00000000121885934Department of Veterinary Medicine, Transmissible Cancer Group, University of Cambridge, Cambridge, CB2 1TN UK; grid.8756.c0000 0001 2193 314XWolfson Wohl Cancer Research Centre, Institute of Cancer Sciences, University of Glasgow, Glasgow, G12 8QQ UK; grid.10698.360000000122483208Lineberger Comprehensive Cancer Center, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA; Dana-Farber/Boston Children’s Cancer and Blood Disorders Center, Boston, MA 02115 USA; grid.38142.3c000000041936754XDepartment of Pediatrics, Harvard Medical School, Boston, MA 02115 USA; Leeds Institute of Medical Research at St. James’s, University of Leeds, St. James’s University Hospital, Leeds, LS2 9JT UK; grid.411475.20000 0004 1756 948XDepartment of Pathology and Diagnostics, University and Hospital Trust of Verona, 37129 Verona, Italy; grid.412744.00000 0004 0380 2017Department of Surgery, Princess Alexandra Hospital, Brisbane, QLD 4102 Australia; grid.1003.20000 0000 9320 7537Surgical Oncology Group, Diamantina Institute, University of Queensland, Brisbane, QLD 4072 Australia; grid.67105.350000 0001 2164 3847Department of Population and Quantitative Health Sciences, Case Western Reserve University School of Medicine, Cleveland, OH 44106 USA; grid.443867.a0000 0000 9149 4843Research Health Analytics and Informatics, University Hospitals Cleveland Medical Center, Cleveland, OH 44106 USA; grid.413144.70000 0001 0489 6543Gloucester Royal Hospital, Gloucester, GL1 3NN UK; grid.419890.d0000 0004 0626 690XDiagnostic Development, Ontario Institute for Cancer Research, Toronto, ON M5G 0A3 Canada; grid.22072.350000 0004 1936 7697Arnie Charbonneau Cancer Institute, University of Calgary, Calgary, AB T2N 1N4 Canada; grid.22072.350000 0004 1936 7697Departments of Surgery and Oncology, University of Calgary, Calgary, AB T2N 1N4 Canada; grid.55325.340000 0004 0389 8485Department of Pathology, Oslo University Hospital, The Norwegian Radium Hospital, Oslo, Norway; grid.419890.d0000 0004 0626 690XPanCuRx Translational Research Initiative, Ontario Institute for Cancer Research, Toronto, ON M5G 0A3 Canada; grid.280502.d0000 0000 8741 3625Department of Oncology, Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins University School of Medicine, Baltimore, MD 21231 USA; grid.430506.4University Hospital Southampton NHS Foundation Trust, Southampton, SO16 6YD UK; grid.439344.d0000 0004 0641 6760Royal Stoke University Hospital, Stoke-on-Trent, ST4 6QG UK; grid.419890.d0000 0004 0626 690XGenome Sequence Informatics, Ontario Institute for Cancer Research, Toronto, ON M5G 0A3 Canada; grid.459583.60000 0004 4652 6825Human Longevity Inc, San Diego, CA M5G 0A3 USA; grid.1018.80000 0001 2342 0938Olivia Newton-John Cancer Research Institute, La Trobe University, Heidelberg, VIC 3086 Australia; grid.440163.40000 0001 0352 8618Genome Canada, Ottawa, ON K2P 1P1 Canada; grid.272799.00000 0000 8687 5377Buck Institute for Research on Aging, Novato, CA 94945 USA; grid.189509.c0000000100241216Duke University Medical Center, Durham, NC 27710 USA; grid.10423.340000 0000 9529 9877Department of Human Genetics, Hannover Medical School, 30625 Hannover, Germany; grid.50956.3f0000 0001 2152 9905Center for Bioinformatics and Functional Genomics, Cedars-Sinai Medical Center, Los Angeles, CA 90048 USA; grid.50956.3f0000 0001 2152 9905Department of Biomedical Sciences, Cedars-Sinai Medical Center, Los Angeles, CA 90048 USA; grid.9619.70000 0004 1937 0538The Hebrew University Faculty of Medicine, Jerusalem, 9112102 Israel; grid.4868.20000 0001 2171 1133Barts Cancer Institute, Barts and the London School of Medicine and Dentistry, Queen Mary University of London, London, E1 4NS UK; grid.9647.c0000 0004 7669 9786Department of Computer Science, Bioinformatics Group, University of Leipzig, 04109 Leipzig, Germany; grid.9647.c0000 0004 7669 9786Interdisciplinary Center for Bioinformatics, University of Leipzig, 04109 Leipzig, Germany; grid.9647.c0000 0004 7669 9786Transcriptome Bioinformatics, LIFE Research Center for Civilization Diseases, University of Leipzig, 04109 Leipzig, Germany; grid.42505.360000 0001 2156 6853USC Norris Comprehensive Cancer Center, University of Southern California, Los Angeles, CA 90007 USA; grid.411475.20000 0004 1756 948XDepartment of Diagnostics and Public Health, University and Hospital Trust of Verona, Verona, 37129 Italy; grid.7048.b0000 0001 1956 2722Department of Mathematics, Aarhus University, 8000 Aarhus, Denmark; Instituto Carlos Slim de la Salud, Mexico City, Mexico; grid.1005.40000 0004 4902 0432Cancer Division, Garvan Institute of Medical Research, Kinghorn Cancer Centre, University of New South Wales (UNSW Sydney), Sydney, NSW 2052 Australia; grid.1005.40000 0004 4902 0432South Western Sydney Clinical School, Faculty of Medicine, University of New South Wales (UNSW Sydney), Liverpool, NSW 2052 Australia; grid.411714.60000 0000 9825 7840West of Scotland Pancreatic Unit, Glasgow Royal Infirmary, Glasgow, G4 0SF UK; grid.189509.c0000000100241216The Preston Robert Tisch Brain Tumor Center, Duke University Medical Center, Durham, NC 27710 USA; grid.410872.80000 0004 1774 5690National Institute of Biomedical Genomics, Kalyani, West Bengal 741251 India; grid.5510.10000 0004 1936 8921Institute of Clinical Medicine and Institute of Oral Biology, University of Oslo, 0315 Oslo, Norway; grid.10698.360000000122483208University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA; grid.411475.20000 0004 1756 948XARC-Net Centre for Applied Research on Cancer, University and Hospital Trust of Verona, Verona, 0315 Italy; grid.18886.3f0000 0001 1271 4623The Institute of Cancer Research, London, SM2 5NG UK; grid.428397.30000 0004 0385 0924Centre for Computational Biology, Duke-NUS Medical School, Singapore, 169857 Singapore; grid.428397.30000 0004 0385 0924Programme in Cancer and Stem Cell Biology, Duke-NUS Medical School, Singapore, 169857 Singapore; grid.4514.40000 0001 0930 2361Division of Oncology and Pathology, Department of Clinical Sciences Lund, Lund University, Lund, Sweden; grid.411327.20000 0001 2176 9917Department of Pediatric Oncology, Hematology and Clinical Immunology, Heinrich-Heine-University, 40225 Düsseldorf, Germany; Laboratory for Medical Science Mathematics, RIKEN Center for Integrative Medical Sciences, Yokohama, Japan; RIKEN Center for Integrative Medical Sciences, Yokohama, Japan; Department of Internal Medicine/Hematology, Friedrich-Ebert-Hospital, Neumünster, Germany; grid.47100.320000000419368710Departments of Dermatology and Pathology, Yale University, New Haven, CT 06520 USA; grid.4991.50000 0004 1936 8948Radcliffe Department of Medicine, University of Oxford, Oxford, OX1 2JD UK; grid.14709.3b0000 0004 1936 8649Canadian Center for Computational Genomics, McGill University, Montreal, QC H3A 0G4 Canada; grid.14709.3b0000 0004 1936 8649Department of Human Genetics, McGill University, Montreal, QC H3A 0G4 Canada; grid.412330.70000 0004 0628 2985Faculty of Medicine and Health Technology, Tampere University and Tays Cancer Center, Tampere University Hospital, 33520 Tampere, Finland; grid.415967.80000 0000 9965 1030Haematology, Leeds Teaching Hospitals NHS Trust, Leeds, LS1 3EX UK; grid.418116.b0000 0001 0200 3174Translational Research and Innovation, Centre Léon Bérard, Lyon, France; grid.249335.aFox Chase Cancer Center, Philadelphia, PA 19111 USA; grid.17703.320000000405980095International Agency for Research on Cancer, World Health Organization, Lyon, France; grid.421605.40000 0004 0447 4123Earlham Institute, Norwich, NR4 7UZ UK; grid.8273.e0000 0001 1092 7967Norwich Medical School, University of East Anglia, Norwich, NR4 7TJ UK; grid.5590.90000000122931605Department of Molecular Biology, Faculty of Science, Radboud Institute for Molecular Life Sciences, Radboud University, Nijmegen, HB 6525 XZ The Netherlands; CRUK Manchester Institute and Centre, Manchester, M20 4GJ UK; grid.17063.330000 0001 2157 2938Department of Radiation Oncology, University of Toronto, Toronto, ON M5S Canada; grid.5379.80000000121662407Division of Cancer Sciences, Manchester Cancer Research Centre, University of Manchester, Manchester, M13 9PL UK; grid.415224.40000 0001 2150 066XRadiation Medicine Program, Princess Margaret Cancer Centre, Toronto, ON M5G 2C1 Canada; grid.38142.3c000000041936754XDepartment of Pathology, Brigham and Women’s Hospital, Harvard Medical School, Boston, MA 02115 USA; grid.21107.350000 0001 2171 9311Department of Surgery, Division of Thoracic Surgery, The Johns Hopkins University School of Medicine, Baltimore, MD 21205 USA; grid.430814.aDivision of Molecular Pathology, The Netherlands Cancer Institute, Oncode Institute, Amsterdam, CX 1066 The Netherlands; grid.7497.d0000 0004 0492 0584Division of Applied Bioinformatics, German Cancer Research Center (DKFZ), 69120 Heidelberg, Germany; German Cancer Genome Consortium (DKTK), Heidelberg, Germany; grid.5170.30000 0001 2181 8870Center for Biological Sequence Analysis, Department of Bio and Health Informatics, Technical University of Denmark, 2800 Lyngby, Denmark; grid.5254.60000 0001 0674 042XNovo Nordisk Foundation Center for Protein Research, University of Copenhagen, Copenhagen, 1165 Denmark; grid.1003.20000 0000 9320 7537Institute for Molecular Bioscience, University of Queensland, St. Lucia, Brisbane, QLD 4072 Australia; grid.5586.e0000 0004 0639 2885Federal Ministry of Education and Research, Berlin, Germany; grid.1013.30000 0004 1936 834XMelanoma Institute Australia, University of Sydney, Sydney, NSW 2006 Australia; grid.16149.3b0000 0004 0551 4246Pediatric Hematology and Oncology, University Hospital Muenster, 48149 Muenster, Germany; grid.21107.350000 0001 2171 9311Department of Pathology, Johns Hopkins University School of Medicine, Baltimore, MD 21205 USA; grid.280502.d0000 0000 8741 3625McKusick-Nathans Institute of Genetic Medicine, Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins University School of Medicine, Baltimore, MD 21231 USA; grid.418158.10000 0004 0534 4718Foundation Medicine, Inc, Cambridge, MA 02141 USA; grid.168010.e0000000419368956Department of Biomedical Data Science, Stanford University School of Medicine, Stanford, CA 94305 USA; grid.168010.e0000000419368956Department of Genetics, Stanford University School of Medicine, Stanford, CA 94305 USA; grid.266102.10000 0001 2297 6811Bakar Computational Health Sciences Institute and Department of Pediatrics, University of California, San Francisco, CA 94110 USA; grid.5510.10000 0004 1936 8921Institute of Clinical Medicine, Faculty of Medicine, University of Oslo, Oslo, 0315 Norway; grid.94365.3d0000 0001 2297 5165National Cancer Institute, National Institutes of Health, Bethesda, MD 20892 USA; grid.5072.00000 0001 0304 893XRoyal Marsden NHS Foundation Trust, London and Sutton, Sutton, SM2 5PT UK; grid.5335.00000000121885934Department of Oncology, University of Cambridge, Cambridge, CB2 1TN UK; grid.5335.00000000121885934Li Ka Shing Centre, Cancer Research UK Cambridge Institute, University of Cambridge, Cambridge, CB2 1TN UK; grid.14925.3b0000 0001 2284 9388Institut Gustave Roussy, 94800 Villejuif, France; grid.5335.00000000121885934Department of Haematology, University of Cambridge, Cambridge, CB2 1TN UK; grid.5841.80000 0004 1937 0247Anatomia Patológica, Hospital Clinic, Institut d’Investigacions Biomèdiques August Pi i Sunyer (IDIBAPS), University of Barcelona, 08007 Barcelona, Spain; grid.451322.30000 0004 1770 9462Spanish Ministry of Science and Innovation, Madrid, Spain; grid.412590.b0000 0000 9081 2336University of Michigan Comprehensive Cancer Center, Ann Arbor, MI 48109 USA; grid.5734.50000 0001 0726 5157Department for BioMedical Research, University of Bern, 3012 Bern, Switzerland; grid.5734.50000 0001 0726 5157Department of Medical Oncology, Inselspital, University Hospital and University of Bern, 3012 Bern, Switzerland; grid.5734.50000 0001 0726 5157Graduate School for Cellular and Biomedical Sciences, University of Bern, 3012 Bern, Switzerland; grid.8982.b0000 0004 1762 5736University of Pavia, 27100 Pavia, Italy; grid.265892.20000000106344187University of Alabama at Birmingham, Birmingham, AL 35294 USA; grid.417184.f0000 0001 0661 1177UHN Program in BioSpecimen Sciences, Toronto General Hospital, Toronto, ON M5G 2C4 Canada; grid.59734.3c0000 0001 0670 2351Department of Urology, Icahn School of Medicine at Mount Sinai, New York, NY 10029 USA; grid.1009.80000 0004 1936 826XCentre for Law and Genetics, University of Tasmania, Sandy Bay Campus, Hobart, TAS 7005 Australia; grid.7700.00000 0001 2190 4373Faculty of Biosciences, Heidelberg University, 69117 Heidelberg, Germany; grid.28046.380000 0001 2182 2255Department of Biochemistry, Microbiology and Immunology, Faculty of Medicine, University of Ottawa, Ottawa, ON K1N 6N5 Canada; grid.66875.3a0000 0004 0459 167XDivision of Anatomic Pathology, Mayo Clinic, Rochester, MN 55905 USA; grid.94365.3d0000 0001 2297 5165Division of Cancer Epidemiology and Genetics, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892 USA; grid.417154.20000 0000 9781 7439Illawarra Shoalhaven Local Health District L3 Illawarra Cancer Care Centre, Wollongong Hospital, Wollongong, NSW 2500 Australia; BioForA, French National Institute for Agriculture, Food, and Environment (INRAE), ONF, Orléans, France; grid.21107.350000 0001 2171 9311Department of Biostatistics, Bloomberg School of Public Health, Johns Hopkins University, Baltimore, MD 21218 USA; grid.266100.30000 0001 2107 4242University of California San Diego, San Diego, CA 92093 USA; grid.66875.3a0000 0004 0459 167XDivision of Experimental Pathology, Mayo Clinic, Rochester, MN 55905 USA; grid.1013.30000 0004 1936 834XCentre for Cancer Research, The Westmead Institute for Medical Research, University of Sydney, Sydney, NSW 2006 Australia; grid.413252.30000 0001 0180 6477Department of Gynaecological Oncology, Westmead Hospital, Sydney, NSW 2145 Australia; PDXen Biosystems Inc, Seoul, South Korea; grid.37172.300000 0001 2292 0500Korea Advanced Institute of Science and Technology, Daejeon, South Korea; grid.36303.350000 0000 9148 4899Electronics and Telecommunications Research Institute, Daejeon, South Korea; grid.455095.80000 0001 2189 059XInstitut National du Cancer (INCA), Boulogne-Billancourt, France; grid.410724.40000 0004 0620 9745Division of Medical Oncology, National Cancer Centre, Singapore, 169610 Singapore; grid.411475.20000 0004 1756 948XMedical Oncology, University and Hospital Trust of Verona, Verona, 37129 Italy; grid.412468.d0000 0004 0646 2097Department of Pediatrics, University Hospital Schleswig-Holstein, 23562 Kiel, Germany; grid.231844.80000 0004 0474 0428Hepatobiliary/Pancreatic Surgical Oncology Program, University Health Network, Toronto, ON M5G 1L7 Canada; grid.9654.e0000 0004 0372 3343School of Biological Sciences, University of Auckland, Auckland, 1010 New Zealand; grid.1008.90000 0001 2179 088XDepartment of Surgery, University of Melbourne, Parkville, VIC 3010 Australia; grid.416107.50000 0004 0614 0346The Murdoch Children’s Research Institute, Royal Children’s Hospital, Parkville, VIC 3052 Australia; grid.1042.7Walter and Eliza Hall Institute, Parkville, VIC 3052 Australia; grid.416166.20000 0004 0473 9881Lunenfeld-Tanenbaum Research Institute, Mount Sinai Hospital, Toronto, ON M5G 1X5 Canada; grid.8273.e0000 0001 1092 7967University of East Anglia, Norwich, NR4 7TJ UK; grid.240367.4Norfolk and Norwich University Hospital NHS Trust, Norwich, NR4 7UY UK; grid.433802.e0000 0004 0465 4247Victorian Institute of Forensic Medicine, Southbank, VIC 3006 Australia; grid.38142.3c000000041936754XDepartment of Biomedical Informatics, Harvard Medical School, Boston, MA 02115 USA; grid.5335.00000000121885934Department of Chemistry, Centre for Molecular Science Informatics, University of Cambridge, Cambridge, CB2 1TN UK; grid.38142.3c000000041936754XLudwig Center at Harvard Medical School, Boston, MA 02115 USA; grid.1008.90000 0001 2179 088XPeter MacCallum Cancer Centre, University of Melbourne, Melbourne, VIC 3010 Australia; grid.32224.350000 0004 0386 9924Physics Division, Optimization and Systems Biology Lab, Massachusetts General Hospital, Boston, MA 02114 USA; grid.39382.330000 0001 2160 926XDepartment of Medicine, Baylor College of Medicine, Houston, TX 77030 USA; grid.6190.e0000 0000 8580 3777University of Cologne, 50923 Cologne, Germany; grid.450294.e0000 0004 0641 0756International Genomics Consortium, Phoenix, AZ 85004 USA; grid.419890.d0000 0004 0626 690XGenomics Research Program, Ontario Institute for Cancer Research, Toronto, ON M5G 0A3 Canada; grid.439436.fBarking Havering and Redbridge University Hospitals NHS Trust, Romford, UK; grid.1013.30000 0004 1936 834XChildren’s Hospital at Westmead, University of Sydney, Sydney, NSW 2006 Australia; grid.411475.20000 0004 1756 948XDepartment of Medicine, Section of Endocrinology, University and Hospital Trust of Verona, 37129 Verona, Italy; grid.51462.340000 0001 2171 9952Computational Biology Center, Memorial Sloan Kettering Cancer Center, New York, NY 10065 USA; grid.5801.c0000 0001 2156 2780Department of Biology, , ETH Zurich, Zürich, Switzerland; grid.5801.c0000 0001 2156 2780Department of Computer Science, ETH Zurich, Zurich, Switzerland; grid.419765.80000 0001 2223 3006SIB Swiss Institute of Bioinformatics, Lausanne, Switzerland; grid.5386.8000000041936877XWeill Cornell Medical College, New York, NY 10021 USA; grid.5335.00000000121885934Academic Department of Medical Genetics, University of Cambridge, Addenbrooke’s Hospital, Cambridge, CB2 1TN UK; grid.5335.00000000121885934MRC Cancer Unit, University of Cambridge, Cambridge, CB2 1TN UK; grid.10698.360000000122483208Departments of Pediatrics and Genetics, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA; grid.492568.4Seven Bridges Genomics, Charlestown, MA 02129 USA; Annai Systems, Inc, Carlsbad, CA 92013 USA; grid.5608.b0000 0004 1757 3470Department of Pathology, General Hospital of Treviso, Department of Medicine, University of Padua, 35122 Treviso, Italy; grid.9851.50000 0001 2165 4204Department of Computational Biology, University of Lausanne, 1015 Lausanne, Switzerland; grid.8591.50000 0001 2322 4988Department of Genetic Medicine and Development, University of Geneva Medical School, 1205 CH Geneva, Switzerland; grid.8591.50000 0001 2322 4988Swiss Institute of Bioinformatics, University of Geneva, 1205 CH Geneva, Switzerland; grid.451388.30000 0004 1795 1830The Francis Crick Institute, London, NW1 1AT UK; grid.5596.f0000 0001 0668 7884University of Leuven, 3000 Leuven, Belgium; grid.418377.e0000 0004 0620 715XComputational and Systems Biology, Genome Institute of Singapore, Singapore, 138672 Singapore; grid.4280.e0000 0001 2180 6431School of Computing, National University of Singapore, Singapore, 119077 Singapore; grid.4991.50000 0004 1936 8948Big Data Institute, Li Ka Shing Centre, University of Oxford, Oxford, OX1 2JD UK; grid.17063.330000 0001 2157 2938The Edward S. Rogers Sr. Department of Electrical and Computer Engineering, University of Toronto, Toronto, ON M5S Canada; grid.418119.40000 0001 0684 291XBreast Cancer Translational Research Laboratory JC Heuson, Institut Jules Bordet, 1000 Brussels, Belgium; grid.5596.f0000 0001 0668 7884Department of Oncology, Laboratory for Translational Breast Cancer Research, KU Leuven, 3000 Leuven, Belgium; grid.473715.3Institute for Research in Biomedicine (IRB Barcelona), The Barcelona Institute of Science and Technology, 08036 Barcelona, Spain; grid.5612.00000 0001 2172 2676Research Program on Biomedical Informatics, Universitat Pompeu Fabra, 08002 Barcelona, Spain; grid.415224.40000 0001 2150 066XDivision of Medical Oncology, Princess Margaret Cancer Centre, Toronto, ON M5G 2C1 Canada; grid.5386.8000000041936877XDepartment of Physiology and Biophysics, Weill Cornell Medicine, New York, NY 10065 USA; grid.5386.8000000041936877XInstitute for Computational Biomedicine, Weill Cornell Medicine, New York, NY 10065 USA; grid.415596.a0000 0004 0440 3018Department of Pathology, UPMC Shadyside, Pittsburgh, PA 15232 USA; Independent Consultant, Wellesley, USA; grid.8993.b0000 0004 1936 9457Department of Cell and Molecular Biology, Science for Life Laboratory, Uppsala University, 752 36 Uppsala, Sweden; grid.256896.6Hefei University of Technology, Anhui, China; grid.5284.b0000 0001 0790 3681Translational Cancer Research Unit, GZA Hospitals St.-Augustinus, Center for Oncological Research, Faculty of Medicine and Health Sciences, University of Antwerp, Antwerp, 2000 Belgium; grid.25879.310000 0004 1936 8972University of Pennsylvania, Philadelphia, PA 19104 USA; grid.52788.300000 0004 0427 7672The Wellcome Trust, London, NW1 2BE UK; grid.415490.d0000 0001 2177 007XDepartment of Pathology, Queen Elizabeth University Hospital, Glasgow, G51 4TF UK; grid.1049.c0000 0001 2294 1395Department of Genetics and Computational Biology, QIMR Berghofer Medical Research Institute, Brisbane, QLD 4006 Australia; grid.5335.00000000121885934Department of Oncology, Centre for Cancer Genetic Epidemiology, University of Cambridge, Cambridge, CB2 1TN UK; grid.5335.00000000121885934Department of Public Health and Primary Care, Centre for Cancer Genetic Epidemiology, University of Cambridge, Cambridge, CB2 1TN UK; grid.453281.90000 0004 4652 6665Prostate Cancer Canada, Toronto, ON M5C 1M1 Canada; grid.5335.00000000121885934University of Cambridge, Cambridge, CB2 1TN UK; grid.4514.40000 0001 0930 2361Department of Laboratory Medicine, Translational Cancer Research, Lund University Cancer Center at Medicon Village, Lund University, Lund, Sweden; grid.413448.e0000 0000 9314 1427CIBER Epidemiología y Salud Pública (CIBERESP), Madrid, Spain; Research Group on Statistics, Econometrics and Health (GRECS), UdG, Barcelona, Spain; Quantitative Genomics Laboratories (qGenomics), Barcelona, Spain; grid.507118.a0000 0001 0329 4954Icelandic Cancer Registry, Icelandic Cancer Society, Reykjavik, Iceland; grid.233520.50000 0004 1761 4404State Key Laboratory of Cancer Biology, and Xijing Hospital of Digestive Diseases, Fourth Military Medical University, Shaanxi, China; grid.5608.b0000 0004 1757 3470Department of Medicine (DIMED), Surgical Pathology Unit, University of Padua, 35122 Padua, Italy; grid.65499.370000 0001 2106 9910Department of Cancer Biology, Dana-Farber Cancer Institute, Boston, MA 02215 USA; grid.475435.4Rigshospitalet, Copenhagen, Denmark; grid.94365.3d0000 0001 2297 5165Center for Cancer Genomics, National Cancer Institute, National Institutes of Health, Bethesda, MD 20814 USA; grid.14848.310000 0001 2292 3357Department of Biochemistry and Molecular Medicine, University of Montreal, Montreal, QC H3T 1J4 Canada; grid.1011.10000 0004 0474 1797Australian Institute of Tropical Health and Medicine, James Cook University, Douglas, QLD 4811 Australia; Department of Neuro-Oncology, Istituto Neurologico Besta, Milano, Italy; grid.484025.fBioplatforms Australia, North Ryde, NSW Australia; grid.83440.3b0000000121901201Department of Pathology (Research), University College London Cancer Institute, London, WC1E 6DD UK; grid.415224.40000 0001 2150 066XDepartment of Surgical Oncology, Princess Margaret Cancer Centre, Toronto, ON M5G 2C1 Canada; grid.5645.2000000040459992XDepartment of Medical Oncology, Josephine Nefkens Institute and Cancer Genomics Centre, Erasmus Medical Center, 3015 GD Rotterdam, The Netherlands; grid.415184.d0000 0004 0614 0266The University of Queensland Thoracic Research Centre, The Prince Charles Hospital, Brisbane, QLD 4032 Australia; grid.5808.50000 0001 1503 7226CIBIO/InBIO-Research Center in Biodiversity and Genetic Resources, Universidade do Porto, 4099-002 Vairão, Portugal; grid.420746.30000 0001 1887 2462HCA Laboratories, London, UK; grid.10025.360000 0004 1936 8470University of Liverpool, Liverpool, L69 3BX UK; grid.22098.310000 0004 1937 0503The Azrieli Faculty of Medicine, Bar-Ilan University, Safed, 5290002 Israel; grid.15276.370000 0004 1936 8091Department of Neurosurgery, University of Florida, Gainesville, FL 32611 USA; grid.26999.3d0000 0001 2151 536XDepartment of Pathology, Graduate School of Medicine, University of Tokyo, Tokyo, 113-8654 Japan; grid.28665.3f0000 0001 2287 1366National Genotyping Center, Institute of Biomedical Sciences, Academia Sinica, Taipei, Taiwan; grid.7563.70000 0001 2174 1754University of Milano Bicocca, Monza, 20126 Italy; grid.55325.340000 0004 0389 8485Department of Pathology, Oslo University Hospital Ulleval, 0450 Oslo, Norway; grid.38142.3c000000041936754XCenter for Biomedical Informatics, Harvard Medical School, Boston, MA 02115 USA; grid.94365.3d0000 0001 2297 5165Office of Cancer Genomics, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892 USA; grid.7497.d0000 0004 0492 0584Cancer Epigenomics, German Cancer Research Center (DKFZ), 69120 Heidelberg, Germany; grid.240145.60000 0001 2291 4776Department of Cancer Biology, The University of Texas MD Anderson Cancer Center, Houston, TX 77030 USA; grid.240145.60000 0001 2291 4776Department of Surgical Oncology, The University of Texas MD Anderson Cancer Center, Houston, TX 77030 USA; grid.51462.340000 0001 2171 9952Department of Pathology, Memorial Sloan Kettering Cancer Center, New York, NY 10065 USA; grid.66875.3a0000 0004 0459 167XDivision of Gastroenterology and Hepatology, Mayo Clinic, Rochester, MN 55905 USA; grid.1013.30000 0004 1936 834XUniversity of Sydney, Sydney, NSW 2006 Australia; grid.4991.50000 0004 1936 8948University of Oxford, Oxford, OX1 2JD UK; grid.24029.3d0000 0004 0383 8386Cambridge University Hospitals NHS Foundation Trust, Cambridge, CB2 0QQ UK; grid.5335.00000000121885934Department of Surgery, Academic Urology Group, University of Cambridge, Cambridge, CB2 0QQ UK; grid.8379.50000 0001 1958 8658Department of Medicine II, University of Würzburg, 97070 Wuerzburg, Germany; grid.26790.3a0000 0004 1936 8606Sylvester Comprehensive Cancer Center, University of Miami, Miami, FL 33146 USA; grid.20522.370000 0004 1767 9005Institut Hospital del Mar d’Investigacions Mèdiques (IMIM), Barcelona, Spain; grid.280664.e0000 0001 2110 5790Genome Integrity and Structural Biology Laboratory, National Institute of Environmental Health Sciences (NIEHS), Durham, NC 27709 USA; grid.425213.3St. Thomas’s Hospital, London, SE1 7EH UK; Osaka International Cancer Center, Osaka, Japan; Department of Pathology, Skåne University Hospital, Lund University, Lund, Sweden; grid.422301.60000 0004 0606 0717Department of Medical Oncology, Beatson West of Scotland Cancer Centre, Glasgow, G12 0YN UK; grid.1008.90000 0001 2179 088XCentre for Cancer Research, Victorian Comprehensive Cancer Centre, University of Melbourne, Melbourne, VIC 3010 Australia; grid.170205.10000 0004 1936 7822Department of Medicine, Section of Hematology/Oncology, University of Chicago, Chicago, IL 60637 USA; grid.452463.2German Center for Infection Research (DZIF), Partner Site Hamburg-Borstel-Lübeck-Riems, Hamburg, Germany; grid.454780.a0000 0001 0683 2228Department of Biotechnology, Ministry of Science and Technology, Government of India, New Delhi, Delhi 110016 India; grid.410724.40000 0004 0620 9745National Cancer Centre Singapore, Singapore, 169610 Singapore; grid.253264.40000 0004 1936 9473Brandeis University, Waltham, MA 02453 USA; grid.168010.e0000000419368956Department of Internal Medicine, Stanford University, Stanford, CA 94305 USA; grid.267308.80000 0000 9206 2401The University of Texas Health Science Center at Houston, Houston, TX 77030 USA; grid.7445.20000 0001 2113 8111Imperial College NHS Trust, Imperial College, London, INY SW7 2BU UK; grid.7839.50000 0004 1936 9721Senckenberg Institute of Pathology, University of Frankfurt Medical School, 60323 Frankfurt, Germany; grid.266100.30000 0001 2107 4242Department of Medicine, Division of Biomedical Informatics, UC San Diego School of Medicine, San Diego, CA 92093 USA; grid.468222.8Center for Precision Health, School of Biomedical Informatics, The University of Texas Health Science Center, Houston, TX 77030 USA; Oxford Nanopore Technologies, New York, NY OX4 4DQ USA; grid.26999.3d0000 0001 2151 536XInstitute of Medical Science, University of Tokyo, Tokyo, 113-8654 Japan; grid.412857.d0000 0004 1763 1087Wakayama Medical University, Wakayama, 641-8510 Japan; grid.10698.360000000122483208Department of Internal Medicine, Division of Medical Oncology, Lineberger Comprehensive Cancer Center, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA; grid.267301.10000 0004 0386 9246University of Tennessee Health Science Center for Cancer Research, Memphis, TN 38163 USA; grid.412346.60000 0001 0237 2025Department of Histopathology, Salford Royal NHS Foundation Trust, Salford, M6 8HD UK; grid.5379.80000000121662407Faculty of Biology, Medicine and Health, University of Manchester, Manchester, M13 9PL UK; grid.11135.370000 0001 2256 9319Peking University, 100871 Beijing, China; grid.239552.a0000 0001 0680 8770Children’s Hospital of Philadelphia, Philadelphia, PA 19104 USA; grid.4714.60000 0004 1937 0626Karolinska Institute, 171 77 Stockholm, Sweden; grid.17063.330000 0001 2157 2938The Donnelly Centre, University of Toronto, Toronto, ON M5S Canada; grid.256753.00000 0004 0470 5964Department of Medical Genetics, College of Medicine, Hallym University, Chuncheon, South Korea; grid.5612.00000 0001 2172 2676Department of Experimental and Health Sciences, Institute of Evolutionary Biology (UPF-CSIC), Universitat Pompeu Fabra, 08002 Barcelona, Spain; grid.411941.80000 0000 9194 7179Health Data Science Unit, University Clinics, Heidelberg, Germany; grid.39158.360000 0001 2173 7691Hokkaido University, Sapporo, 060-0808 Japan; grid.272242.30000 0001 2168 5385Department of Pathology and Clinical Laboratory, National Cancer Center Hospital, Tokyo, 104-0045 Japan; grid.10698.360000000122483208Department of Genetics, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA; grid.418245.e0000 0000 9999 5706Computational Biology, Leibniz Institute on Aging-Fritz Lipmann Institute (FLI), 07745 Jena, Germany; grid.1008.90000 0001 2179 088XUniversity of Melbourne Centre for Cancer Research, Melbourne, VIC 3010 Australia; grid.266813.80000 0001 0666 4105University of Nebraska Medical Center, Omaha, NE 68198 USA; Syntekabio Inc, Daejeon, South Korea; grid.5650.60000000404654431Department of Pathology, Academic Medical Center, Amsterdam, 1105 AZ The Netherlands; China National GeneBank-Shenzhen, Shenzhen, China; grid.7497.d0000 0004 0492 0584Division of Molecular Genetics, German Cancer Research Center (DKFZ), 69120 Heidelberg, Germany; grid.59734.3c0000 0001 0670 2351Icahn School of Medicine at Mount Sinai, New York, NY 10029 USA; grid.431072.30000 0004 0572 4227AbbVie, North Chicago, IL 60064 USA; grid.6363.00000 0001 2218 4662Institute of Pathology, Charité – University Medicine Berlin, Berlin, Germany; grid.248762.d0000 0001 0702 3000Centre for Translational and Applied Genomics, British Columbia Cancer Agency, Vancouver, BC V8R 6V5 Canada; grid.418716.d0000 0001 0709 1919Edinburgh Royal Infirmary, Edinburgh, EH16 4SA UK; grid.419491.00000 0001 1014 0849Berlin Institute for Medical Systems Biology, Max Delbrück Center for Molecular Medicine, 13125 Berlin, Germany; grid.5253.10000 0001 0328 4908Department of Pediatric Immunology, Hematology and Oncology, University Hospital, Heidelberg, 69120 Heidelberg, Germany; grid.7497.d0000 0004 0492 0584German Cancer Research Center (DKFZ), 69120 Heidelberg, Germany; grid.482664.aHeidelberg Institute for Stem Cell Technology and Experimental Medicine (HI-STEM), 69120 Heidelberg, Germany; grid.5386.8000000041936877XInstitute for Computational Biomedicine, Weill Cornell Medical College, New York, NY 10021 USA; grid.429884.b0000 0004 1791 0895New York Genome Center, New York, NY 10013 USA; grid.21107.350000 0001 2171 9311Department of Urology, James Buchanan Brady Urological Institute, Johns Hopkins University School of Medicine, Baltimore, MD 21205 USA; grid.26999.3d0000 0001 2151 536XDepartment of Preventive Medicine, Graduate School of Medicine, The University of Tokyo, Tokyo, 113-8654 Japan; grid.39382.330000 0001 2160 926XDepartment of Molecular and Cellular Biology, Baylor College of Medicine, Houston, TX 77030 USA; grid.39382.330000 0001 2160 926XDepartment of Pathology and Immunology, Baylor College of Medicine, Houston, TX 77030 USA; grid.413890.70000 0004 0420 5521Michael E. DeBakey Veterans Affairs Medical Center, Houston, TX 77030 USA; grid.5170.30000 0001 2181 8870Technical University of Denmark, 2800 Lyngby, Denmark; grid.49606.3d0000 0001 1364 9317Department of Pathology, College of Medicine, Hanyang University, Seoul, South Korea; Academic Unit of Surgery, School of Medicine, College of Medical, Veterinary and Life Sciences, University of Glasgow, Glasgow Royal Infirmary, Glasgow, G12 8QQ UK; grid.267370.70000 0004 0533 4667Department of Pathology, Asan Medical Center, College of Medicine, Ulsan University, Songpa-gu, Seoul, South Korea; Science Writer, Garrett Park, MD USA; grid.419890.d0000 0004 0626 690XInternational Cancer Genome Consortium (ICGC)/ICGC Accelerating Research in Genomic Oncology (ARGO) Secretariat, Ontario Institute for Cancer Research, Toronto, ON M5G 0A3 Canada; grid.8954.00000 0001 0721 6013University of Ljubljana, 1000 Ljubljana, Slovenia; grid.170205.10000 0004 1936 7822Department of Public Health Sciences, University of Chicago, Chicago, IL 60637 USA; grid.240372.00000 0004 0400 4439Research Institute, NorthShore University HealthSystem, Evanston, IL 60201 USA; grid.5734.50000 0001 0726 5157Department for Biomedical Research, University of Bern, Bern, 3012 Switzerland; grid.411640.6Centre of Genomics and Policy, McGill University and Génome Québec Innovation Centre, Montreal, QC H3A 0G4 Canada; grid.10698.360000000122483208Carolina Center for Genome Sciences, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA; Hopp Children’s Cancer Center (KiTZ), Heidelberg, Germany; grid.7497.d0000 0004 0492 0584Pediatric Glioma Research Group, German Cancer Research Center (DKFZ), 69120 Heidelberg, Germany; grid.11485.390000 0004 0422 0975Cancer Research UK, London, NG34 7SY UK; Indivumed GmbH, Hamburg, Germany; grid.412004.30000 0004 0478 9977University Hospital Zurich, 8091 Zurich, Switzerland; grid.419765.80000 0001 2223 3006Clinical Bioinformatics, Swiss Institute of Bioinformatics, 1015 Geneva, Switzerland; grid.412004.30000 0004 0478 9977Institute for Pathology and Molecular Pathology, University Hospital Zurich, 8091 Zurich, Switzerland; grid.7400.30000 0004 1937 0650Institute of Molecular Life Sciences, University of Zurich, 8091 Zurich, Switzerland; grid.4305.20000 0004 1936 7988MRC Human Genetics Unit, MRC IGMM, University of Edinburgh, Edinburgh, EH8 9YL UK; grid.50956.3f0000 0001 2152 9905Women’s Cancer Program at the Samuel Oschin Comprehensive Cancer Institute, Cedars-Sinai Medical Center, Los Angeles, CA 90048 USA; grid.4808.40000 0001 0657 4636Department of Biology, Bioinformatics Group, Division of Molecular Biology, Faculty of Science, University of Zagreb, 10000 Zagreb, Croatia; grid.412468.d0000 0004 0646 2097Department for Internal Medicine II, University Hospital Schleswig-Holstein, 23562 Kiel, Germany; grid.414733.60000 0001 2294 430XGenetics and Molecular Pathology, SA Pathology, Adelaide, SA 5011 Australia; grid.272242.30000 0001 2168 5385Department of Gastric Surgery, National Cancer Center Hospital, Tokyo, Japan; grid.272242.30000 0001 2168 5385Department of Bioinformatics, Division of Cancer Genomics, National Cancer Center Research Institute, Tokyo, Japan; grid.435025.50000 0004 0619 6198A.A. Kharkevich Institute of Information Transmission Problems, Moscow, Russia; Oncology and Immunology, Dmitry Rogachev National Research Center of Pediatric Hematology, Moscow, Russia; grid.454320.40000 0004 0555 3608Skolkovo Institute of Science and Technology, Moscow, Russia; grid.253615.60000 0004 1936 9510Department of Surgery, The George Washington University, School of Medicine and Health Science, Washington, DC 20052 USA; grid.94365.3d0000 0001 2297 5165Endocrine Oncology Branch, Center for Cancer Research, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892 USA; grid.1004.50000 0001 2158 5405Melanoma Institute Australia, Macquarie University, Sydney, NSW 2109 Australia; grid.116068.80000 0001 2341 2786MIT Computer Science and Artificial Intelligence Laboratory, Massachusetts Institute of Technology, Cambridge, MA 02139 USA; grid.413249.90000 0004 0385 0051Tissue Pathology and Diagnostic Oncology, Royal Prince Alfred Hospital, Sydney, NSW 2050 Australia; grid.9786.00000 0004 0470 0856Cholangiocarcinoma Screening and Care Program and Liver Fluke and Cholangiocarcinoma Research Centre, Faculty of Medicine, Khon Kaen University, Khon Kaen, 40002 Thailand; Controlled Department and Institution, New York, NY USA; grid.5386.8000000041936877XEnglander Institute for Precision Medicine, Weill Cornell Medicine, New York, NY 10065 USA; grid.410914.90000 0004 0628 9810National Cancer Center, Gyeonggi, South Korea; grid.266100.30000 0001 2107 4242Health Sciences Department of Biomedical Informatics, University of California San Diego, La Jolla, CA 92093 USA; grid.410914.90000 0004 0628 9810Research Core Center, National Cancer Centre Korea, Goyang-si, South Korea; grid.264381.a0000 0001 2181 989XDepartment of Health Sciences and Technology, Sungkyunkwan University School of Medicine, Seoul, South Korea; Samsung Genome Institute, Seoul, South Korea; grid.417747.60000 0004 0460 3896Breast Oncology Program, Dana-Farber/Brigham and Women’s Cancer Center, Boston, MA 02215 USA; grid.51462.340000 0001 2171 9952Department of Surgery, Memorial Sloan Kettering Cancer Center, New York, NY 10065 USA; grid.62560.370000 0004 0378 8294Division of Breast Surgery, Brigham and Women’s Hospital, Boston, MA 02115 USA; grid.280664.e0000 0001 2110 5790Integrative Bioinformatics Support Group, National Institute of Environmental Health Sciences (NIEHS), Durham, NC 27709 USA; grid.7914.b0000 0004 1936 7443Department of Clinical Science, University of Bergen, 5007 Bergen, Norway; grid.412484.f0000 0001 0302 820XCenter For Medical Innovation, Seoul National University Hospital, Seoul, South Korea; grid.412484.f0000 0001 0302 820XDepartment of Internal Medicine, Seoul National University Hospital, Seoul, South Korea; grid.413454.30000 0001 1958 0162Institute of Computer Science, Polish Academy of Sciences, Warsawa, Poland; grid.7497.d0000 0004 0492 0584Functional and Structural Genomics, German Cancer Research Center (DKFZ), 69120 Heidelberg, Germany; grid.94365.3d0000 0001 2297 5165Laboratory of Translational Genomics, Division of Cancer Epidemiology and Genetics, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892 USA; grid.9647.c0000 0004 7669 9786Institute for Medical Informatics Statistics and Epidemiology, University of Leipzig, 04109 Leipzig, Germany; grid.240145.60000 0001 2291 4776Morgan Welch Inflammatory Breast Cancer Research Program and Clinic, The University of Texas MD Anderson Cancer Center, Houston, TX 77030 USA; grid.7450.60000 0001 2364 4210Department of Hematology and Oncology, Georg-Augusts-University of Göttingen, 37073 Göttingen, Germany; grid.5718.b0000 0001 2187 5445Institute of Cell Biology (Cancer Research), University of Duisburg-Essen, 47057 Essen, Germany; grid.420545.2King’s College London and Guy’s and St. Thomas’ NHS Foundation Trust, London, WC2R 2LS UK; grid.251017.00000 0004 0406 2057Center for Epigenetics, Van Andel Research Institute, Grand Rapids, MI 49503 USA; grid.416100.20000 0001 0688 4634The University of Queensland Centre for Clinical Research, Royal Brisbane and Women’s Hospital, Herston, QLD 4029 Australia; grid.6190.e0000 0000 8580 3777Department of Pediatric Oncology and Hematology, University of Cologne, 50923 Cologne, Germany; grid.411327.20000 0001 2176 9917University of Düsseldorf, 40225 Düsseldorf, Germany; grid.418119.40000 0001 0684 291XDepartment of Pathology, Institut Jules Bordet, Brussels, 1000 Belgium; grid.8761.80000 0000 9919 9582Institute of Biomedicine, Sahlgrenska Academy at University of Gothenburg, 413 90 Gothenburg, Sweden; grid.414235.50000 0004 0619 2154Children’s Medical Research Institute, Sydney, NSW 2145 Australia; ILSbio, LLC Biobank, Chestertown, MD USA; Division of Genetics and Genomics, Boston Children’s Hospital, Harvard Medical School, Boston, MA 02115 USA; grid.49606.3d0000 0001 1364 9317Institute for Bioengineering and Biopharmaceutical Research (IBBR), Hanyang University, Seoul, South Korea; grid.205975.c0000 0001 0740 6917Department of Statistics, University of California Santa Cruz, Santa Cruz, CA 95064 USA; grid.419538.20000 0000 9071 0620Department of Vertebrate Genomics/Otto Warburg Laboratory Gene Regulation and Systems Biology of Cancer, Max Planck Institute for Molecular Genetics, 14195 Berlin, Germany; grid.411640.6McGill University and Genome Quebec Innovation Centre, Montreal, QC H3A 0G4 Canada; grid.137628.90000 0004 1936 8753Gynecologic Oncology, NYU Laura and Isaac Perlmutter Cancer Center, New York University, New York, NY 10003 USA; grid.4367.60000 0001 2355 7002Division of Oncology, Stem Cell Biology Section, Washington University School of Medicine, St. Louis, MO 63110 USA; grid.38142.3c000000041936754XHarvard University, Cambridge, MA 02138 USA; grid.94365.3d0000 0001 2297 5165Urologic Oncology Branch, Center for Cancer Research, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892 USA; grid.5510.10000 0004 1936 8921University of Oslo, 0315 Oslo, Norway; grid.17063.330000 0001 2157 2938University of Toronto, Toronto, ON M5S Canada; grid.11135.370000 0001 2256 9319School of Life Sciences, Peking University, Beijing, China; grid.419407.f0000 0004 4665 8158Leidos Biomedical Research, Inc, McLean, VA USA; grid.5841.80000 0004 1937 0247Hematology, Hospital Clinic, Institut d’Investigacions Biomèdiques August Pi i Sunyer (IDIBAPS), University of Barcelona, 08007 Barcelona, Spain; grid.73113.370000 0004 0369 1660Second Military Medical University, Shanghai, China; Chinese Cancer Genome Consortium, Shenzhen, China; grid.414350.70000 0004 0447 1045Department of Medical Oncology, Beijing Hospital, Beijing, China; grid.412474.00000 0001 0027 0586Laboratory of Molecular Oncology, Key Laboratory of Carcinogenesis and Translational Research (Ministry of Education), Peking University Cancer Hospital and Institute, Beijing, China; grid.11914.3c0000 0001 0721 1626School of Medicine/School of Mathematics and Statistics, University of St. Andrews, St. Andrews, Fife KY16 9AJ UK; Department of Biochemistry and Molecular Biology, Faculty of Medicine, University Institute of Oncology-IUOPA, Oviedo, Spain; grid.476460.70000 0004 0639 0505Institut Bergonié, Bordeaux, France; grid.5335.00000000121885934Cancer Unit, MRC University of Cambridge, Cambridge, CB2 1TN UK; grid.239546.f0000 0001 2153 6013Department of Pathology and Laboratory Medicine, Center for Personalized Medicine, Children’s Hospital Los Angeles, Los Angeles, CA 90027 USA; grid.1001.00000 0001 2180 7477John Curtin School of Medical Research, Canberra, ACT 2601 Australia; MVZ Department of Oncology, PraxisClinic am Johannisplatz, Leipzig, Germany; grid.5342.00000 0001 2069 7798Department of Information Technology, Ghent University, Ghent, Belgium; grid.5342.00000 0001 2069 7798Department of Plant Biotechnology and Bioinformatics, Ghent University, Ghent, Belgium; grid.240344.50000 0004 0392 3476Institute for Genomic Medicine, Nationwide Children’s Hospital, Columbus, OH 43205 USA; grid.26009.3d0000 0004 1936 7961Department of Surgery, Duke University, Durham, NC 27708 USA; grid.7080.fInstitut Català de Paleontologia Miquel Crusafont, Universitat Autònoma de Barcelona, Barcelona, Spain; grid.8756.c0000 0001 2193 314XUniversity of Glasgow, Glasgow, G12 8QQ UK; grid.10403.36Institut d’Investigacions Biomèdiques August Pi i Sunyer (IDIBAPS), Barcelona, Spain; grid.7445.20000 0001 2113 8111Department of Surgery and Cancer, Imperial College, London, INY SW7 2BU UK; grid.437060.60000 0004 0567 5138Applications Department, Oxford Nanopore Technologies, Oxford, OX4 4DQ UK; grid.266102.10000 0001 2297 6811Department of Obstetrics, Gynecology and Reproductive Services, University of California San Francisco, San Francisco, CA 94110 USA; grid.27860.3b0000 0004 1936 9684Department of Biochemistry and Molecular Medicine, University California at Davis, Sacramento, CA 95616 USA; grid.415224.40000 0001 2150 066XSTTARR Innovation Facility, Princess Margaret Cancer Centre, Toronto, ON M5G 2C1 Canada; grid.1029.a0000 0000 9939 5719Discipline of Surgery, Western Sydney University, Penrith, NSW 2150 Australia; grid.10698.360000000122483208Department of Genetics, Lineberger Comprehensive Cancer Center, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA; grid.413103.40000 0001 2160 8953Departments of Neurology and Neurosurgery, Henry Ford Hospital, Detroit, MI 48202 USA; grid.13648.380000 0001 2180 3484Institute of Pathology, University Medical Center Hamburg-Eppendorf, 20251 Hamburg, Germany; grid.177174.30000 0001 2242 4849Department of Health Sciences, Faculty of Medical Sciences, Kyushu University, Fukuoka, Japan; grid.461593.c0000 0001 1939 6592Heidelberg Academy of Sciences and Humanities, Heidelberg, Germany; grid.1008.90000 0001 2179 088XDepartment of Clinical Pathology, University of Melbourne, Melbourne, VIC 3010 Australia; grid.240614.50000 0001 2181 8635Department of Pathology, Roswell Park Cancer Institute, Buffalo, NY 14203 USA; grid.7737.40000 0004 0410 2071Department of Computer Science, University of Helsinki, 00100 Helsinki, Finland; grid.7737.40000 0004 0410 2071Institute of Biotechnology, University of Helsinki, 00100 Helsinki, Finland; grid.7737.40000 0004 0410 2071Organismal and Evolutionary Biology Research Programme, University of Helsinki, 00100 Helsinki, Finland; grid.4367.60000 0001 2355 7002Department of Obstetrics and Gynecology, Division of Gynecologic Oncology, Washington University School of Medicine, St. Louis, MO 63110 USA; grid.430183.dPenrose St. Francis Health Services, Colorado Springs, CO 80923 USA; grid.410712.1Institute of Pathology, Ulm University and University Hospital of Ulm, Ulm, 89081 Germany; grid.272242.30000 0001 2168 5385National Cancer Center, Tokyo, Japan; grid.418377.e0000 0004 0620 715XGenome Institute of Singapore, Singapore, 138672 Singapore; grid.453370.60000 0001 2161 6363German Cancer Aid, Bonn, Germany; grid.428397.30000 0004 0385 0924Programme in Cancer and Stem Cell Biology, Centre for Computational Biology, Duke-NUS Medical School, 169857 Singapore, Singapore; grid.10784.3a0000 0004 1937 0482The Chinese University of Hong Kong, Shatin, NT, Hong Kong, China; grid.233520.50000 0004 1761 4404Fourth Military Medical University, Shaanxi, China; grid.5335.00000000121885934The University of Cambridge School of Clinical Medicine, Cambridge, CB2 1TN UK; grid.240871.80000 0001 0224 711XSt. Jude Children’s Research Hospital, Memphis, TN 38105 USA; grid.415224.40000 0001 2150 066XUniversity Health Network, Princess Margaret Cancer Centre, Toronto, ON M5G 1L7 Canada; grid.205975.c0000 0001 0740 6917Center for Biomolecular Science and Engineering, University of California Santa Cruz, Santa Cruz, CA 95064 USA; grid.170205.10000 0004 1936 7822Department of Medicine, University of Chicago, Chicago, IL 60637 USA; grid.66875.3a0000 0004 0459 167XDepartment of Neurology, Mayo Clinic, Rochester, MN 55905 USA; grid.24029.3d0000 0004 0383 8386Cambridge Oesophagogastric Centre, Cambridge University Hospitals NHS Foundation Trust, Cambridge, CB2 0QQ UK; grid.253692.90000 0004 0445 5969Department of Computer Science, Carleton College, Northfield, MN 55057 USA; grid.8756.c0000 0001 2193 314XInstitute of Cancer Sciences, College of Medical Veterinary and Life Sciences, University of Glasgow, Glasgow, G12 8QQ UK; grid.265892.20000000106344187Department of Epidemiology, University of Alabama at Birmingham, Birmingham, AL 35294 USA; grid.417691.c0000 0004 0408 3720HudsonAlpha Institute for Biotechnology, Huntsville, AL 35806 USA; grid.265892.20000000106344187O’Neal Comprehensive Cancer Center, University of Alabama at Birmingham, Birmingham, AL 35294 USA; grid.26091.3c0000 0004 1936 9959Department of Pathology, Keio University School of Medicine, Tokyo, Japan; grid.272242.30000 0001 2168 5385Department of Hepatobiliary and Pancreatic Oncology, National Cancer Center Hospital, Tokyo, Japan; grid.430406.50000 0004 6023 5303Sage Bionetworks, Seattle, WA USA; grid.410724.40000 0004 0620 9745Lymphoma Genomic Translational Research Laboratory, National Cancer Centre, Singapore, 98121 Singapore; grid.416008.b0000 0004 0603 4965Department of Clinical Pathology, Robert-Bosch-Hospital, 70376 Stuttgart, Germany; grid.17063.330000 0001 2157 2938Department of Cell and Systems Biology, University of Toronto, Toronto, ON M5S Canada; grid.4714.60000 0004 1937 0626Department of Biosciences and Nutrition, Karolinska Institutet, 171 77 Stockholm, Sweden; grid.410914.90000 0004 0628 9810Center for Liver Cancer, Research Institute and Hospital, National Cancer Center, Gyeonggi, South Korea; grid.264381.a0000 0001 2181 989XDivision of Hematology-Oncology, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, South Korea; grid.264381.a0000 0001 2181 989XSamsung Advanced Institute for Health Sciences and Technology, Sungkyunkwan University School of Medicine, Seoul, South Korea; grid.263136.30000 0004 0533 2389Cheonan Industry-Academic Collaboration Foundation, Sangmyung University, Cheonan, South Korea; grid.240324.30000 0001 2109 4251NYU Langone Medical Center, New York, NY 10016 USA; grid.239578.20000 0001 0675 4725Department of Hematology and Medical Oncology, Cleveland Clinic, Cleveland, OH 44195 USA; grid.66875.3a0000 0004 0459 167XDepartment of Health Sciences Research, Mayo Clinic, Rochester, MN 55905 USA; grid.414316.50000 0004 0444 1241Helen F. Graham Cancer Center at Christiana Care Health Systems, Newark, DE 19713 USA; grid.5253.10000 0001 0328 4908Heidelberg University Hospital, 69120 Heidelberg, Germany; CSRA Incorporated, Fairfax, VA USA; grid.83440.3b0000000121901201Research Department of Pathology, University College London Cancer Institute, London, WC1E 6DD UK; grid.13097.3c0000 0001 2322 6764Department of Research Oncology, Guy’s Hospital, King’s Health Partners AHSC, King’s College London School of Medicine, London, WC2R 2LS UK; grid.1004.50000 0001 2158 5405Faculty of Medicine and Health Sciences, Macquarie University, Sydney, NSW 2109 Australia; grid.411158.80000 0004 0638 9213University Hospital of Minjoz, INSERM UMR 1098, Besançon, France; grid.7719.80000 0000 8700 1153Spanish National Cancer Research Centre, Madrid, 28029 Spain; grid.415180.90000 0004 0540 9980Center of Digestive Diseases and Liver Transplantation, Fundeni Clinical Institute, Bucharest, Romania; Cureline, Inc, South San Francisco, CA USA; grid.412946.c0000 0001 0372 6120St. Luke’s Cancer Centre, Royal Surrey County Hospital NHS Foundation Trust, Guildford, UK; grid.24029.3d0000 0004 0383 8386Cambridge Breast Unit, Addenbrooke’s Hospital, Cambridge University Hospital NHS Foundation Trust and NIHR Cambridge Biomedical Research Centre, Cambridge, CB2 0QQ UK; grid.416266.10000 0000 9009 9462East of Scotland Breast Service, Ninewells Hospital, Aberdeen, UK; grid.5841.80000 0004 1937 0247Department of Genetics, Microbiology and Statistics, University of Barcelona, IRSJD, IBUB, Barcelona, Spain; grid.30760.320000 0001 2111 8460Department of Obstetrics and Gynecology, Medical College of Wisconsin, Milwaukee, WI 53226 USA; grid.189967.80000 0001 0941 6502Hematology and Medical Oncology, Winship Cancer Institute of Emory University, Atlanta, GA 30322 USA; grid.16750.350000 0001 2097 5006Department of Computer Science, Princeton University, Princeton, NJ 08544 USA; grid.152326.10000 0001 2264 7217Vanderbilt Ingram Cancer Center, Vanderbilt University, Nashville, TN 37235 USA; grid.261331.40000 0001 2285 7943Ohio State University College of Medicine and Arthur G. James Comprehensive Cancer Center, Columbus, OH 43210 USA; grid.268441.d0000 0001 1033 6139Department of Surgery, Yokohama City University Graduate School of Medicine, Kanagawa, 236-0027 Japan; grid.10698.360000000122483208Research Computing Center, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA; grid.30064.310000 0001 2157 6568School of Molecular Biosciences and Center for Reproductive Biology, Washington State University, Pullman, WA 99163 USA; grid.17063.330000 0001 2157 2938Department of Laboratory Medicine and Pathobiology, University of Toronto, Toronto, ON M5S Canada; grid.51462.340000 0001 2171 9952Department of Pathology, Human Oncology and Pathogenesis Program, Memorial Sloan Kettering Cancer Center, New York, NY 10065 USA; grid.411067.50000 0000 8584 9230University Hospital Giessen, Pediatric Hematology and Oncology, Giessen, Germany; Oncologie Sénologie, ICM Institut Régional du Cancer, Montpellier, France; grid.9764.c0000 0001 2153 9986Institute of Clinical Molecular Biology, Christian-Albrechts-University, Kiel, Germany; grid.8379.50000 0001 1958 8658Institute of Pathology, University of Wuerzburg, 97070 Wuerzburg, Germany; grid.418484.50000 0004 0380 7221Department of Urology, North Bristol NHS Trust, Bristol, UK; grid.419385.20000 0004 0620 9905SingHealth, Duke-NUS Institute of Precision Medicine, National Heart Centre Singapore, Singapore, 169609 Singapore; grid.5734.50000 0001 0726 5157Bern Center for Precision Medicine, University Hospital of Bern, University of Bern, Bern, 3012 Switzerland; grid.413734.60000 0000 8499 1112Englander Institute for Precision Medicine, Weill Cornell Medicine and New York Presbyterian Hospital, New York, NY 10065 USA; grid.5386.8000000041936877XMeyer Cancer Center, Weill Cornell Medicine, New York, NY 10065 USA; grid.5386.8000000041936877XPathology and Laboratory, Weill Cornell Medical College, New York, NY 10065 USA; Vall d’Hebron Institute of Oncology: VHIO, Barcelona, Spain; grid.411475.20000 0004 1756 948XGeneral and Hepatobiliary-Biliary Surgery, Pancreas Institute, University and Hospital Trust of Verona, Verona, Italy; grid.22401.350000 0004 0502 9283National Centre for Biological Sciences, Tata Institute of Fundamental Research, Bangalore, 560065 India; Department of Pathology, GZA-ZNA Hospitals, Antwerp, Belgium; grid.422639.8Analytical Biological Services, Inc, Wilmington, DE 19801 USA; grid.1013.30000 0004 1936 834XSydney Medical School, University of Sydney, Sydney, NSW 2050 Australia; grid.38142.3c000000041936754XcBio Center, Dana-Farber Cancer Institute, Harvard Medical School, Boston, MA 02115 USA; grid.38142.3c000000041936754XDepartment of Cell Biology, Harvard Medical School, Boston, MA 02115 USA; grid.410871.b0000 0004 1769 5793Advanced Centre for Treatment Research and Education in Cancer, Tata Memorial Centre, Navi Mumbai, Maharashtra 400012 India; grid.266842.c0000 0000 8831 109XSchool of Environmental and Life Sciences, Faculty of Science, The University of Newcastle, Ourimbah, NSW 2308 Australia; grid.410718.b0000 0001 0262 7331Department of Dermatology, University Hospital of Essen, 45147 Essen, Germany; grid.13648.380000 0001 2180 3484Martini-Clinic, Prostate Cancer Center, University Medical Center Hamburg-Eppendorf, 20251 Hamburg, Germany; grid.9764.c0000 0001 2153 9986Department of General Internal Medicine, University of Kiel, 24118 Kiel, Germany; German Cancer Consortium (DKTK), Partner site Berlin, Berlin, Germany; grid.239395.70000 0000 9011 8547Cancer Research Institute, Beth Israel Deaconess Medical Center, Boston, MA 02215 USA; grid.21925.3d0000 0004 1936 9000University of Pittsburgh, Pittsburgh, PA 15260 USA; grid.38142.3c000000041936754XDepartment of Ophthalmology and Ocular Genomics Institute, Massachusetts Eye and Ear, Harvard Medical School, Boston, MA 02115 USA; grid.240372.00000 0004 0400 4439Center for Psychiatric Genetics, NorthShore University HealthSystem, Evanston, IL 60031 USA; grid.251017.00000 0004 0406 2057Van Andel Research Institute, Grand Rapids, MI 49503 USA; grid.26999.3d0000 0001 2151 536XLaboratory of Molecular Medicine, Human Genome Center, Institute of Medical Science, University of Tokyo, Tokyo, Japan; grid.480536.c0000 0004 5373 4593Japan Agency for Medical Research and Development, Tokyo, Japan; grid.222754.40000 0001 0840 2678Korea University, Seoul, South Korea; grid.414467.40000 0001 0560 6544Murtha Cancer Center, Walter Reed National Military Medical Center, Bethesda, MD 20814 USA; grid.9764.c0000 0001 2153 9986Human Genetics, University of Kiel, 24118 Kiel, Germany; Department of Oncologic Pathology, Dana-Farber Cancer Institute, Harvard Medical School, Boston, MA 02115 USA; grid.5288.70000 0000 9758 5690Oregon Health and Science University, Portland, OR 97239 USA; grid.240145.60000 0001 2291 4776Center for RNA Interference and Noncoding RNA, The University of Texas MD Anderson Cancer Center, Houston, TX 77030 USA; grid.240145.60000 0001 2291 4776Department of Experimental Therapeutics, The University of Texas MD Anderson Cancer Center, Houston, TX 77030 USA; grid.240145.60000 0001 2291 4776Department of Gynecologic Oncology and Reproductive Medicine, The University of Texas MD Anderson Cancer Center, Houston, TX 77030 USA; grid.15628.38University Hospitals Coventry and Warwickshire NHS Trust, Coventry, UK; grid.10417.330000 0004 0444 9382Department of Radiation Oncology, Radboud University Nijmegen Medical Centre, Nijmegen, 6525 GA The Netherlands; grid.170205.10000 0004 1936 7822Institute for Genomics and Systems Biology, University of Chicago, Chicago, IL USA; grid.459927.40000 0000 8785 9045Clinic for Hematology and Oncology, St.-Antonius-Hospital, 60637 Eschweiler, Germany; grid.51462.340000 0001 2171 9952Computational and Systems Biology Program, Memorial Sloan Kettering Cancer Center, New York, NY 10065 USA; grid.14013.370000 0004 0640 0021University of Iceland, Reykjavik, Iceland; grid.7497.d0000 0004 0492 0584Division of Computational Genomics and Systems Genetics, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.416266.10000 0000 9009 9462Dundee Cancer Centre, Ninewells Hospital, Dundee, UK; grid.410712.1Department for Internal Medicine III, University of Ulm and University Hospital of Ulm, Ulm, Germany; grid.418596.70000 0004 0639 6384Institut Curie, INSERM Unit 830, Paris, France; grid.268441.d0000 0001 1033 6139Department of Gastroenterology and Hepatology, Yokohama City University Graduate School of Medicine, Kanagawa, Japan; grid.10417.330000 0004 0444 9382Department of Laboratory Medicine, Radboud University Nijmegen Medical Centre, Nijmegen, 6525 GA The Netherlands; grid.7497.d0000 0004 0492 0584Division of Cancer Genome Research, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.163555.10000 0000 9486 5048Department of General Surgery, Singapore General Hospital, Singapore, Singapore; grid.4280.e0000 0001 2180 6431Cancer Science Institute of Singapore, National University of Singapore, Singapore, Singapore; grid.7737.40000 0004 0410 2071Department of Medical and Clinical Genetics, Genome-Scale Biology Research Program, University of Helsinki, Helsinki, Finland; grid.24029.3d0000 0004 0383 8386East Anglian Medical Genetics Service, Cambridge University Hospitals NHS Foundation Trust, Cambridge, UK; grid.21729.3f0000000419368729Irving Institute for Cancer Dynamics, Columbia University, New York, NY 10027 USA; grid.418812.60000 0004 0620 9243Institute of Molecular and Cell Biology, Singapore, Singapore; grid.410724.40000 0004 0620 9745Laboratory of Cancer Epigenome, Division of Medical Science, National Cancer Centre Singapore, Singapore, Singapore; Universite Lyon, INCa-Synergie, Centre Léon Bérard, Lyon, France; grid.66875.3a0000 0004 0459 167XDepartment of Urology, Mayo Clinic, Rochester, MN 55905 USA; grid.416177.20000 0004 0417 7890Royal National Orthopaedic Hospital-Stanmore, Stanmore, Middlesex UK; Giovanni Paolo II / I.R.C.C.S. Cancer Institute, Bari, BA Italy; grid.7497.d0000 0004 0492 0584Neuroblastoma Genomics, German Cancer Research Center (DKFZ), Heidelberg, Germany; grid.414603.4Fondazione Policlinico Universitario Gemelli IRCCS, Rome, Italy, Rome, Italy; grid.5611.30000 0004 1763 1124University of Verona, Verona, Italy; grid.418135.a0000 0004 0641 3404Centre National de Génotypage, CEA-Institute de Génomique, Evry, France; grid.5012.60000 0001 0481 6099CAPHRI Research School, Maastricht University, Maastricht, ER The Netherlands; grid.418116.b0000 0001 0200 3174Department of Biopathology, Centre Léon Bérard, Lyon, France; grid.7849.20000 0001 2150 7757Université Claude Bernard Lyon 1, Villeurbanne, France; grid.419082.60000 0004 1754 9200Core Research for Evolutional Science and Technology (CREST), JST, Tokyo, Japan; grid.26999.3d0000 0001 2151 536XDepartment of Biological Sciences, Laboratory for Medical Science Mathematics, Graduate School of Science, University of Tokyo, Yokohama, Japan; grid.265073.50000 0001 1014 9130Department of Medical Science Mathematics, Medical Research Institute, Tokyo Medical and Dental University (TMDU), Tokyo, Japan; grid.412563.70000 0004 0376 6589University Hospitals Birmingham NHS Foundation Trust, Birmingham, UK; grid.4777.30000 0004 0374 7521Centre for Cancer Research and Cell Biology, Queen’s University, Belfast, UK; grid.240145.60000 0001 2291 4776Breast Medical Oncology, The University of Texas MD Anderson Cancer Center, Houston, TX 77030 USA; grid.21107.350000 0001 2171 9311Department of Surgery, Johns Hopkins University School of Medicine, Baltimore, MD 21205 USA; grid.4714.60000 0004 1937 0626Department of Oncology-Pathology, Science for Life Laboratory, Karolinska Institute, 171 77 Stockholm, Sweden; grid.5491.90000 0004 1936 9297School of Cancer Sciences, Faculty of Medicine, University of Southampton, Southampton, UK; grid.6988.f0000000110107715Department of Gene Technology, Tallinn University of Technology, Tallinn, Estonia; grid.42327.300000 0004 0473 9646Genetics and Genome Biology Program, SickKids Research Institute, The Hospital for Sick Children, Toronto, ON M5G 1X8 Canada; grid.189967.80000 0001 0941 6502Departments of Neurosurgery and Hematology and Medical Oncology, Winship Cancer Institute and School of Medicine, Emory University, Atlanta, GA 30322 USA; grid.5947.f0000 0001 1516 2393Department of Clinical and Molecular Medicine, Faculty of Medicine and Health Sciences, Norwegian University of Science and Technology, Trondheim, Norway; Argmix Consulting, North Vancouver, BC Canada; grid.5342.00000 0001 2069 7798Department of Information Technology, Ghent University, Interuniversitair Micro-Electronica Centrum (IMEC), Ghent, Belgium; grid.4991.50000 0004 1936 8948Nuffield Department of Surgical Sciences, John Radcliffe Hospital, University of Oxford, Oxford, OX1 2JD UK; grid.9845.00000 0001 0775 3222Institute of Mathematics and Computer Science, University of Latvia, Riga, LV 1586 Latvia; grid.1013.30000 0004 1936 834XDiscipline of Pathology, Sydney Medical School, University of Sydney, Sydney, NSW 2006 Australia; grid.5335.00000000121885934Department of Applied Mathematics and Theoretical Physics, Centre for Mathematical Sciences, University of Cambridge, Cambridge, 10027 UK; grid.21729.3f0000000419368729Department of Statistics, Columbia University, New York, NY USA; grid.8993.b0000 0004 1936 9457Department of Immunology, Genetics and Pathology, Science for Life Laboratory, Uppsala University, 752 36 Uppsala, Sweden; grid.24029.3d0000 0004 0383 8386Department of Histopathology, Cambridge University Hospitals NHS Foundation Trust, Cambridge, CB2 0QQ UK; grid.4991.50000 0004 1936 8948Oxford NIHR Biomedical Research Centre, University of Oxford, Oxford, OX1 2JD UK; grid.410427.40000 0001 2284 9329Georgia Regents University Cancer Center, Augusta, GA 30912 USA; grid.417286.e0000 0004 0422 2524Wythenshawe Hospital, Manchester, M23 9LT UK; grid.4991.50000 0004 1936 8948Wellcome Centre for Human Genetics, University of Oxford, Oxford, OX1 2JD UK; grid.66875.3a0000 0004 0459 167XThoracic Oncology Laboratory, Mayo Clinic, Rochester, MN 55905 USA; grid.66875.3a0000 0004 0459 167XDepartment of Obstetrics and Gynecology, Division of Gynecologic Oncology, Mayo Clinic, Rochester, MN 55905 USA; International Institute for Molecular Oncology, Poznań, 60-203 Poland; grid.22254.330000 0001 2205 0971Poznan University of Medical Sciences, 61-701 Poznań, Poland; grid.7497.d0000 0004 0492 0584Genomics and Proteomics Core Facility High Throughput Sequencing Unit, German Cancer Research Center (DKFZ), 69120 Heidelberg, Germany; grid.410724.40000 0004 0620 9745NCCS-VARI Translational Research Laboratory, National Cancer Centre Singapore, Singapore, 169610 Singapore; grid.4367.60000 0001 2355 7002Edison Family Center for Genome Sciences and Systems Biology, Washington University, St. Louis, MO 63130 USA; grid.301713.70000 0004 0393 3981MRC-University of Glasgow Centre for Virus Research, Glasgow, G61 1QH UK; grid.5288.70000 0000 9758 5690Department of Medical Informatics and Clinical Epidemiology, Division of Bioinformatics and Computational Biology, OHSU Knight Cancer Institute, Oregon Health and Science University, Portland, OR 97239 USA; grid.33199.310000 0004 0368 7223School of Electronic Information and Communications, Huazhong University of Science and Technology, 430074 Wuhan, China; grid.21107.350000 0001 2171 9311Department of Applied Mathematics and Statistics, Johns Hopkins University, Baltimore, MD 21218 USA; grid.136593.b0000 0004 0373 3971Department of Cancer Genome Informatics, Graduate School of Medicine, Osaka University, Osaka, 565-0871 Japan; grid.1013.30000 0004 1936 834XSchool of Mathematics and Statistics, University of Sydney, Sydney, NSW 2006 Australia; grid.170205.10000 0004 1936 7822Ben May Department for Cancer Research and Department of Human Genetics, University of Chicago, Chicago, IL 60637 USA; grid.5386.8000000041936877XTri-Institutional PhD Program in Computational Biology and Medicine, Weill Cornell Medicine, New York, NY 10065 USA; grid.10784.3a0000 0004 1937 0482Department of Medicine and Therapeutics, The Chinese University of Hong Kong, Shatin, NT, Hong Kong, China; grid.240145.60000 0001 2291 4776Department of Biostatistics, The University of Texas MD Anderson Cancer Center, Houston, TX 77030 USA; grid.428397.30000 0004 0385 0924Duke-NUS Medical School, Singapore, 169857 Singapore; grid.16821.3c0000 0004 0368 8293Department of Surgery, Ruijin Hospital, Shanghai Jiaotong University School of Medicine, Shanghai, China; grid.8756.c0000 0001 2193 314XSchool of Computing Science, University of Glasgow, Glasgow, G12 8QQ UK; grid.55325.340000 0004 0389 8485Division of Orthopaedic Surgery, Oslo University Hospital, 0450 Oslo, Norway; grid.1002.30000 0004 1936 7857Eastern Clinical School, Monash University, Melbourne, VIC 3800 Australia; grid.414539.e0000 0001 0459 5396Epworth HealthCare, Richmond, VIC 3121 Australia; grid.65499.370000 0001 2106 9910Department of Biostatistics and Computational Biology, Dana-Farber Cancer Institute and Harvard Medical School, Boston, MA 02215 USA; grid.261331.40000 0001 2285 7943Department of Biomedical Informatics, College of Medicine, The Ohio State University, Columbus, OH 43202 USA; grid.261331.40000 0001 2285 7943The Ohio State University Comprehensive Cancer Center (OSUCCC–James), Columbus, OH 43202 USA; grid.11135.370000 0001 2256 9319BIOPIC, ICG and College of Life Sciences, Peking University, 100871 Beijing, China; grid.267308.80000 0000 9206 2401The University of Texas School of Biomedical Informatics (SBMI) at Houston, Houston, TX 77030 USA; grid.10698.360000000122483208Department of Biostatistics, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599 USA; grid.16753.360000 0001 2299 3507Department of Biochemistry and Molecular Genetics, Feinberg School of Medicine, Northwestern University, Chicago, IL 60208 USA; grid.1013.30000 0004 1936 834XFaculty of Medicine and Health, University of Sydney, Sydney, NSW 2006 Australia; grid.5645.2000000040459992XDepartment of Pathology, Erasmus Medical Center Rotterdam, 1066 CX Rotterdam, GD The Netherlands; grid.430814.aDivision of Molecular Carcinogenesis, The Netherlands Cancer Institute, 1066 CX Amsterdam, The Netherlands; grid.7400.30000 0004 1937 0650Institute of Molecular Life Sciences and Swiss Institute of Bioinformatics, University of Zurich, 8006 Zurich, Switzerland
License: © The Author(s) 2020 CC BY 4.0 Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Article links: DOI: 10.1038/s41467-020-18151-y | PubMed: 32958763 | PMC: PMC7505971
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (776 KB)
Introduction
Complementary efforts of The Cancer Genome Atlas (TCGA) and the International Cancer Genome Consortium (ICGC) have recently produced two of the highest quality and most elaborate and reproducible somatic variant call sets from exome and whole genome-level data in cancer genomics, respectively. The motivation for these efforts stems from the notion that “scientific crowd sourcing” and combining mutation callers can provide very strong results.
These two efforts produced variant calls from 10 different callers, namely Radia1, Varscan2, MuSE3, MuTect4, Pindel5,6, Indelocator7, SomaticSniper8 for WES and MuSE, Broad-Pipeline (anchored by MuTect), Sanger-pipeline, German Cancer Research Center pipeline (DKFZ), and SMuFin9, for WGS. Briefly, the PCAWG Consortium aggregated whole genome sequencing data from 2658 cancers across 38 tumor types generated by the ICGC and TCGA projects. These sequencing data were re-analyzed with standardized, high-accuracy pipelines to align to the human genome (reference build hs37d5) and identify germline variants and somatically acquired mutations10. Of the 885 TCGA samples in ICGC, 746 were included in the latest exome call set produced by both the Multi-Center Mutation Calling in Multiple Cancers (MC3) effort and the Pan-Cancer Analysis of Whole Genomes (PCAWG) Consortium set. These 746 samples represent a critical benchmark for high-level analysis of similarities and differences between exome and genome somatic variant detection methods.
Reproducibility of mutations identified by both whole exome capture sequencing and whole genome sequencing (WGS) techniques remains an important issue, not only for the scientific use of large, established data sets, but for data designs of future research projects. Previous work analyzing exome capture effects on sequence read quality has shown that GC-content bias is the major source of variation in coverage11. A performance comparison across exome-captured platforms demonstrated that for most technologies, both high and low GC-content result in reduced coverage in read depth12. Belkadi et al. compared mutation calls between WGS and WES, observing that ~3% of coding variants with high quality were only detected in WGS, and WGS also had a more uniform distribution of coverage depth, genotype quality, and minor read ratio13. Furthermore, due to the relatively high error rate per read in next-generation sequencing14, the detectability of mutations with low variant allele fractions (VAFs) is limited by background noise. Despite these studies’ nuanced preference towards WGS, others contend that WES will remain a better choice until costs of WGS fall15. The decision to sequence exomes or whole genomes is further confounded as more recent publications in oncology select either WGS16–20 or WES21–24. Recognizing the unresolved nature of this issue, Schwarze et al. have called for more comprehensive studies comparing the WES and WGS studies, especially as this issue has important ramifications for the clinic25.
Our analysis provides confidence that mutation calls within the captured exonic regions of these two data sets are largely consistent. We highlight common sample, cohort, and caller-specific challenges in cancer variant detection from the TCGA and ICGC efforts. We show that variants that are most confidently called in one database i.e., called by multiple callers, are very likely to be called in the other. We assess how reproducibility impacts higher-level mutation signature analysis and illustrate the need for caution in assessing performance that can only be identified by the overlap of these two data sets. Finally, we explore the capacity of WGS to detect recurrent non-coding mutations captured by whole exome sequencing.
Results
Data and workflow
We used publicly available data from the MC3 and PCAWG repositories, consisting of ~3.6 M and ~47 M variants, respectively (Fig. 1a). 746 samples were sequenced by both WES and WGS, comprising various aliquots and portions of the same tumor (Supplementary Data 1, Fig. 1b). Effects of these differences are discussed below for preliminary results, but we ultimately used the entire set of 746 samples in the variant overlap analysis, since the effects of tumor partitioning did not play a significant role (Supplementary Fig. 1). By restricting the public data sets to overlapping samples, we reduced the total corpus to ~220 K (6.1%) and ~23 M (49.6%) mutations for exome and whole genome, respectively. It is notable that there is an enrichment of variants in hypermutated samples from COAD, HNSC, LUAD, and STAD in the PCAWG set used in this study (Supplementary Fig. 2). To begin building a comparable set of mutations between these two studies, we further restricted the whole genome data set to exon regions provided by the MC3 analysis working group. This reduced the WGS data set to 1.6% of its original size, within range of total exome material estimations26 (Fig. 1a). The next step involved removing poorly-covered variants potentially caused by technical anomalies by limiting mutations to those captured in coverage files (distributed as.wig files). A reciprocal coverage strategy was used, meaning PCAWG mutations were restricted to covered genomic regions in MC3 and vice versa, thereby maintaining a complementary set of callable genomic regions. Removal of mutations in uncovered regions reduced the remaining PCAWG data set by approximately one-half, from 387,166 to 183,424 mutations. We also identified 4241 MC3 and 2219 PCAWG mutations that were present in the respective MAF but were not marked as covered in the coverage files provided by a single group. This suggests that different tools consider different minimum coverage strategies. These mutations reflect 2.0% and 1.2%, respectively, of the total mutation discrepancy and were removed because some callers had limited capacity to identify mutations in poorly-covered regions (see “Methods” section). Finally, filter flags provided by MC3 were used to assess somatic mutation filtering strategies. At this stage, we performed filter optimization to comprehensively evaluate all possible combinations of MC3 filters (Supplementary Fig. 3a). Ultimately, we decided to only remove OxoG labeled artifacts and duplicated events produced by these filters (see “Methods” section, Supplementary Data 2). Since each stage of this filtering workflow resulted in many alternative decisions and outcomes, we built MAFit, a web-based graphical user interface that allows users to easily customize comparisons of merged mutations (https://mbailey.shinyapps.io/MAFit/). A MC3 filter assessment also shows that many variants with filter flags in MC3 are present in the PCAWG variant call set, suggesting a need for improved filtering strategies (Supplementary Fig. 3b).
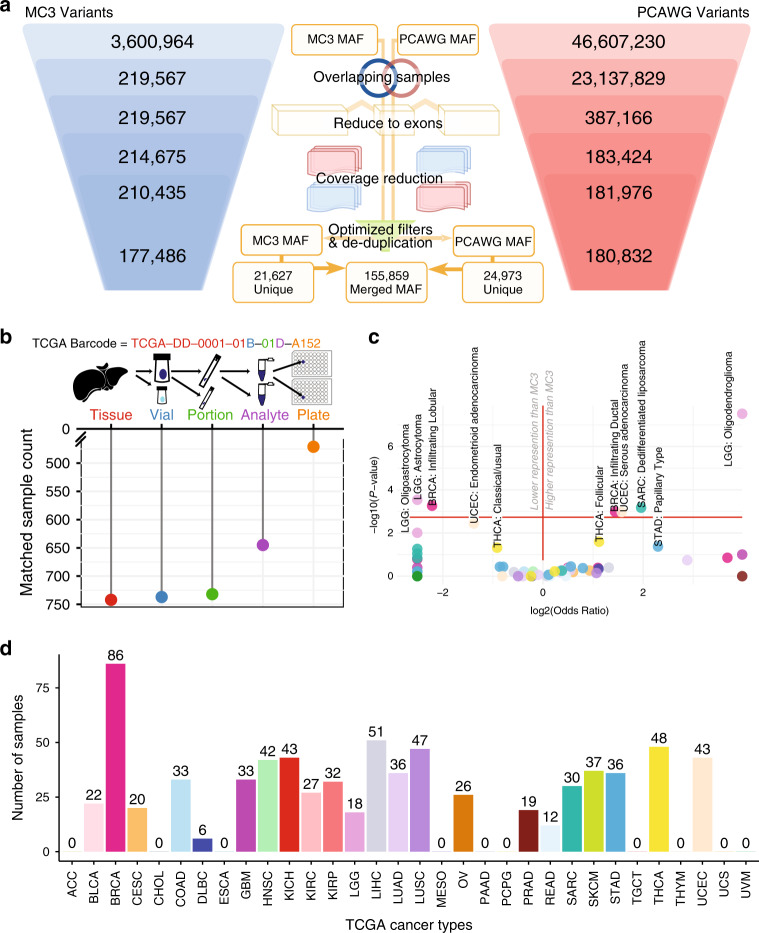
TCGA samples comprise a sizable fraction of the PCAWG sample pool (~30%, Supplementary Data 1) Additional WGS sequencing from TCGA allowed for mutation validation27 and insights into non-coding mutations, such as in TET2. However, this selection process could have potentially influenced our basic comparison of exome-sequenced samples and genome-sequenced samples in two fundamental ways. First, vagaries of tumor extraction and tissue storage protocols may have resulted in many different portions of a tumor being stored, introducing the possibility that different subclones of the same tumor could be present. These could have very different genetic makeups. This information was captured in different substrings of the TCGA identification barcode (see “Methods” section). From the 746 TCGA barcodes, we found that 64% (477) could be traced to the same well of a microtiter plate (Fig. 1b). After correcting for cancer type, we modeled both the impact of matching barcode identifiers between MC3 and PCAWG and variant concordance, finding that differing barcodes did not have an appreciable impact. This result was seen for all samples, even when excluding the hypermutator (Fig. 1). Second, each AWG was able to independently select samples for WGS, which, while not affecting mutation calling, does raise potential biases when comparing PCAWG results to TCGA exome cohort data. An enrichment analysis was performed to identify which tumor subtypes may have been preferentially selected for different cancer types. We found that four tumor subtypes were enriched in the PCAWG effort from TCGA samples: infiltrating ductal breast cancer, endometrial serous adenocarcinoma, differentiated liposarcoma, and low grade oligodendroglioma (FDR < 0.05, Fig. 1c, Supplementary Data 3, and see “Methods” section). Final tumor sample counts for each cancer type are shown in Fig. 1d.
Landscape of mutational overlap between WGS and WES calls
Limiting our analysis to coding regions with sufficient coverage yielded a total of 202,459 variants (155,859 matched, 21,627 unique MC3 variants, and 24,973 unique PCAWG mutations), with 76.7% in concordance between MC3 and PCAWG and 10.7% and 12.3% being unique in MC3 and PCAWG, respectively (Fig. 2a). Concordance can be further separated into SNPs and indels, with 79% and 57% overlapping, respectively (Supplementary Fig. 4). Variant overlap was further investigated to reveal its association with mutation caller, sample, and cancer type (Fig. 2a–c). Consensus variant calling showed 91,705 (45.3%) concordant variants were captured in the intersection of Sanger, MuSE, DKFZ, and Broad callers from PCAWG, as well as Varscan, SomaticSniper, Radia, MuTect, and MuSE callers from MC3. Notably, an additional 7.7% were identified by all SNV mutation detection algorithms, except SomaticSniper. The reduced sensitivity of SomaticSniper is related to its algorithmic consideration of tumor contamination in the matched normal (e.g., skin) for liquid tumors8. After optimizing for filtering strategies, we performed a sample level comparison and found that 70% of samples had greater than 80% mutation concordance across the two cohorts. An additional 20% of samples had greater than 80% mutation recoverability in one or the other technique (Fig. 2b). Skin Cutaneous Melanoma performed the best among all cancer types and had the highest variant-matching rates for both MC3 and PCAWG (Fig. 2c). Generally, when considering all MC3 and all PCAWG mutations separately, we observed that PCAWG variant matching rates were generally higher, especially for Kidney Chromophobe (KICH), Brain Lower Grade Glioma (LGG), Ovarian Serous Cystadenocarcinoma (OV), Rectum Adenocarcinoma (READ), and Thyroid Carcinoma (THCA). The differences in OV are likely driven by whole genome amplified library preparation. Generally, the median fractions for matching MC3 variants were lower than those of matching PCAWG variants. This result was unexpected because MC3 provided fewer unique variants overall, suggesting that a large fraction of PCAWG unique variants reside in a few samples. Furthermore, after accounting for hypermutators, we identified a correlation between non-silent mutations per megabase and mean consensus percentages at the cancer level in both PCAWG consensus percentages (Mann–Whitney p-value = 1.97 × 10−3) and MC3 consensus percentages (Mann–Whitney p-value = 6.59 × 10−4, see “Methods” section, Supplementary Fig. 1c, d). Despite strong rank statistics, neither set exhibited strong correlation values for MC3 variants or PCAWG variants, R2 statistics = 0.31 and 0.17 respectively with the majority of cancer types exceeding 80% mean concordance. Thus, one may expect to observe slightly higher variant fidelity in samples with more mutations.
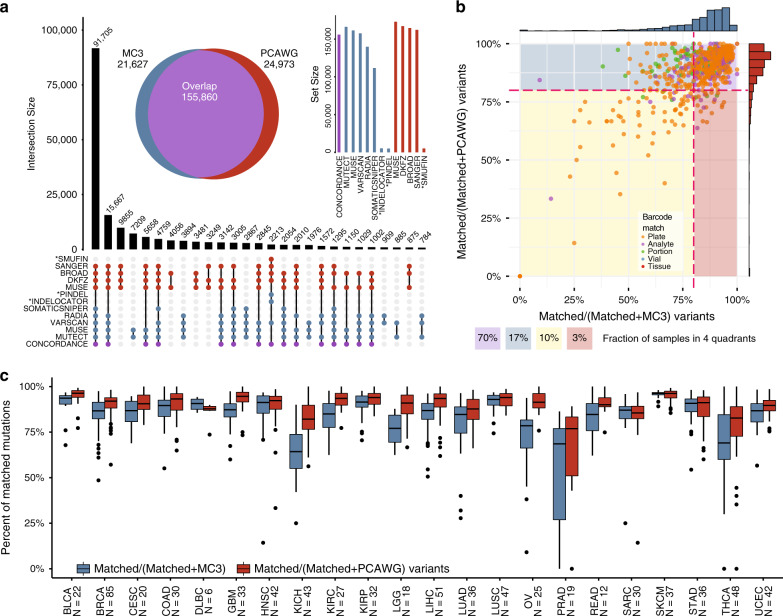
Variant allele fraction affects call-rates
After achieving a comparable data set and merging MC3 and PCAWG variants, we found that low VAF is the prevailing attribute of unique mutations. VAF is a fundamental factor in somatic variant detection, as well as sub-clonal structure prediction, and is used to predict subclonal tumor growth rates and metastatic potential. To explore the contribution of VAF, we sought to distinguish the contribution of subclonal structure and statistical chance when exploring private mutations in a single call set. We articulate our findings in six broad categories: modeling sequence noise, departure from idealized behavior, sub-clonal modeling, annotation differences, variant-caller effects, and analysis correlations.
Association of variant allele fraction with recoverability
We have observed that variants with low VAF are less likely to be reported in both call-sets. This finding relates to the lower sensitivity of somatic variant callers for variants with low VAF. To illustrate this principle, we estimated the expected overlap rate between MC3 and PCAWG at different VAFs. The sensitivity of MuSE across a range of VAFs and read depths in synthetic data was reported in Fan et al., 20163. We used these reported benchmarking characteristics of MuSE to estimate the expected overlap rate between the MuSE call-sets of MC3 and PCAWG across a range of VAFs (see “Methods” section). These expectations, which involve lower overlap rates at lower VAFs, generally tracked observed data but tended to overestimate observed overlap rates, especially for predicting the recovery fraction of MC3 variants in PCAWG. (Fig. 3a) The discrepancies between expectations and observations may relate to simplifying assumptions that made this modeling possible (see “Methods” section).
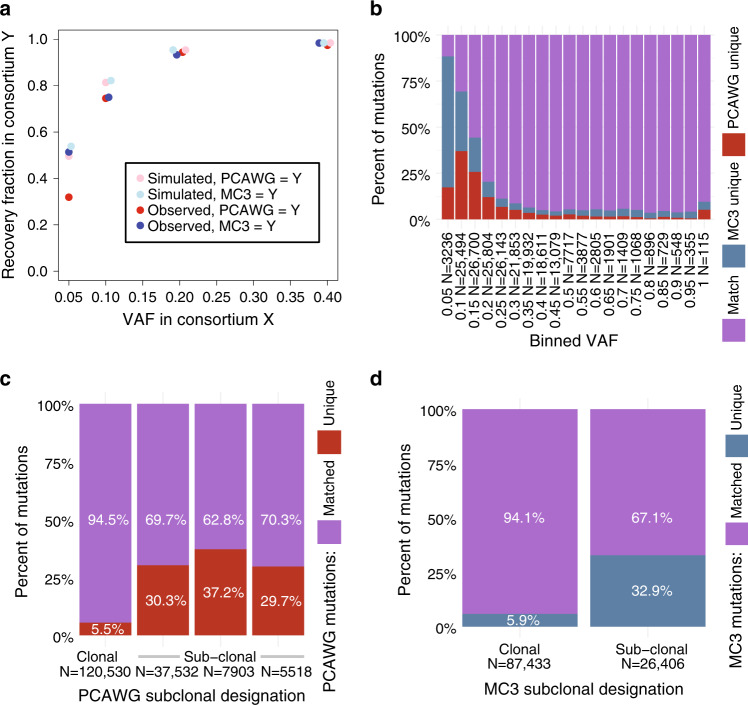
More generally, we observed that VAF had a greater association with variant recovery rates than predicted by the binomial model (Fig. 3b). A random forest regression model trained on five statistics characteristics of VAF distribution per PCAWG sample and another five for that of the corresponding MC3 call-set predicted the fraction of variants per sample unique to PCAWG with 0.85 (0.86—when restricting to variants called by MuSE) Spearman correlation of test-set observations and a 0.68 (0.78) coefficient of determination (R2).
The strong association of VAF with recovery rates by call-set, despite modest explanatory power of the binomial, indicates important departures from idealized behavior. These departures could include explanations such as: PCR amplification violates the assumption of independence of reads, imputed read depths are systematically inflated, or some low-VAF variants represent sequencing artifacts. We conclude that non-ideal effects of VAF predict the majority of sample-level variance in fraction of co-called variants.
Exploring subclonality
One possible explanation for some variants being private to one call-set is that the sequencing aliquots for the two sequencing projects came from subclonally-distinct microregions of the same tumor. To investigate this possibility, we tested whether the MC3 and PCAWG call-sets differed from each other systematically at the subclonal level (Fig. 3c, d). We hypothesized that tumors with a more complex subclonal structure (i.e., greater number of subclones) would have larger systematic differences in the VAF of shared variants between the MC3 and PCAWG call-sets. We found a small but highly significant effect: each additional subclone increased the average absolute difference in VAF of the shared variants between MC3 and PCAWG by 0.003, with a p-value of 1.3 × 10−11 (linear regression); this effect reversed after controlling for tumor purity, indicating that the observed trend does not provide evidence of this interesting concept in re-sequencing (see “Methods” section for details). We do not have evidence that systematic VAF differences between call-sets of the same underlying sample associate with tumor heterogeneity. Real time effects of VAF differences between these two data sets can be observed using the online MAFit tool (Fig. 4).
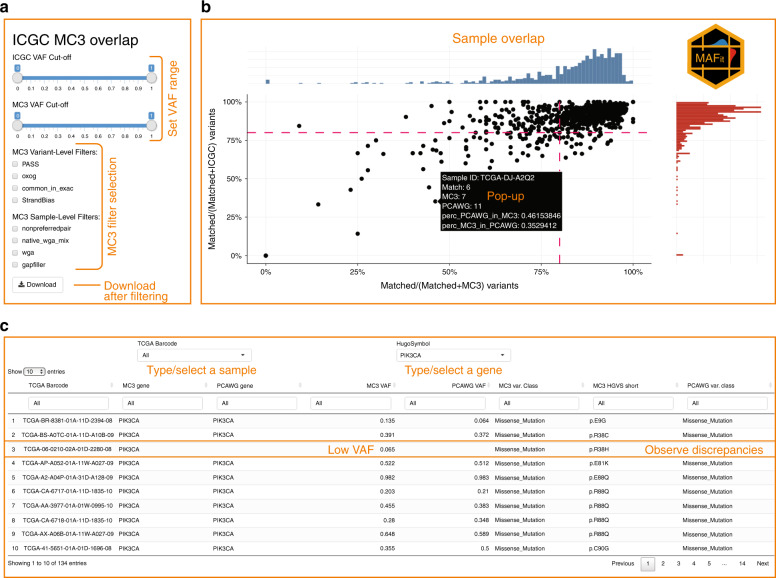
Annotation differs by call-set
Genome annotation is critical for biological interpretation and downstream analysis of sequencing data. In order to avoid issues that arise from annotation differences, we only considered genomic locations in our intersection strategy. In doing so, we observed 2153 annotation differences where MC3 and PCAWG had different genes annotated for the same mutation. After restricting the mutation type to missense mutations and indels, 789 annotations differences remained. Most of these had the same mutation types annotated by both call-sets (690 SNPs, 15 insertions, 50 deletions), but some discrepancies remained. Notably, 413 out of 789 mismatch variants are labeled coding in MC3 but non-coding in PCAWG (Supplementary Data 4). We also observed four mutations that were annotated as cancer gene mutations by MC3, but as non-cancer gene mutations by PCAWG, and another four mutations that were annotated as cancer gene mutations by PCAWG, but as non-cancer gene mutations by MC3. One such example subsumed two mutations on chromosomal location 3p21.1 (genomic locations chr3:52442525 and chr3:52442604) that were annotated as missense mutations of BAP1 by MC3, but as 5’Flank SNPs of PHF7 by PCAWG. While identical pipelines resolve such differences, we stress the potential for misinterpretations when combining these publicly-available datasets.
Effects of software
Another important issue we assess is the degree to which differences in bioinformatics pipelines impact concordance. We extracted calls from MuSE and MuTect, both of which were executed on each dataset, and examined 6 subsets of results: MuSE-only-calls and all calls save MuSE-calls (the complement), MuTect-only-calls and their complement, and MuSE + MuTect calls and their complement. MuSE and Mutect each generate around 95% of the total calls, of which each respective subset shows close to 80% concordance between WES and WGS (Supplementary Fig. 5). These call sets themselves overlap almost completely, with their combination (MuSE + MuTect) giving a marginally higher concordance. Conversely, the data-specific caller combinations (referred to above as the complements) each furnish small call sets which vary considerably between WES and WGS (concordance as low as 15%). Because of the vast difference in the sizes of the MuSE/MuTect and the complementary call sets, there is little difference in the original analysis versus analyses restricted to variant callers common to both platforms. Differences in software pipelines do not appear to be significant confounding factors in concordance here.
Effects on higher-level analysis
We also sought to assess how higher-level analyses might be impacted using mutation signature analysis as a representative. We ran SignatureAnalyzer28 to ascertain signatures between matched WGS and WES samples for each case. A total of 563 of 739 cases (76%) showed the same dominant signature between WES and WGS and the multi-element signature vectors for each case are very highly correlated with one another, the average Pearson coefficient being almost 90%, with a cohort significance of <2 × 10−6 (Fisher’s Test, “Methods” section, Supplementary Fig. 6). These observations suggest that signature analysis is relatively insensitive to data type when concordance is high, as it is here.
Landscape of private WES and WGS mutations
After identifying many possible sources of variation among private variants, we sought to characterize the fraction of variation explained by previously identified factors (Supplementary Fig. 7, see “Methods” section). As displayed, subclonal and low VAF variants make up the largest fractions of explained variants for private MC3 and PCAWG variants. Notably, for private MC3 calls, indels (not called by MuSE or MuTect) are the next highest source of variation explained. GC-content and poor performing cancers such as THCA, KICH, and PRAD make up a smaller portion of the total number of private mutations.
Variants present in only one public call-set
We sought to classify cancer driver mutations uniquely identified by MC3. After removing two outlier samples having excesses of unique mutations (TCGA-CA-6717-01A-11D-1835-10, TCGA-BR-6452-01A-12D-1800-08), we observed 424 mutations in cancer genes28 (median read depth = 97, median alternative allele count = 9) The four most frequently mutated genes were: KMT2C (22-mutations), PIK3CA (12), SPTA1 (9), and NCOR1 (9). Interestingly, the majority of unique PIK3CA mutations not identified by PCAWG were at 2 locations: E542K/G (5), and E545K (4). Whether this phenomenon reflects technical bias of WGS or is a product of subclonality warrants further investigation.
The MC3 effort produced two mutation files: one controlled access somatic mutation file that represents nearly all mutations found by all callers, and a second was modified by the scientific community for public use. There are two critical differences in these files involving the reporting of mutations in exonic regions and mutations reported by a single variant caller. Since we limited our analysis strictly to exonic regions, we observed that 92% of the 9138 PCAWG private mutations found in the MC3 controlled access file were only identified by a single variant caller (Supplementary Fig. 8). As expected, the highest unique variant caller overlap was observed in MuTect and MuSE, two tools that were used by both MC3 and PCAWG. This observation accounts for 30% of PCAWG private variants.
We investigated how many variants unique to the MC3 somatic public access call-set could be found in the PCAWG germline call-set for the same patients. We identified a total of six such variants (each in a different sample), five of which were flagged in the MC3 public call-set with one filter or another. Overall, this indicates that variants that have been incorrectly designated as germline or somatic are an extremely uncommon source of variation between the two projects.
Variants in GC-extreme intervals
Since it is well-known that the efficiency of exome capture is adversely affected by very high or very low GC-content29,30, we sought to test whether GC-content was associated with call rates in MC3 and PCAWG. We used a plug-in for VEP31 to annotate all matched and private SNVs with CADD32 in order to annotate each variant with the percentage of the neighboring 100 bases that are a G or C. First, we assessed how the distribution of read depth across GC-content changes between MC3 and PCAWG (Fig. 5b). PCAWG was found to have a fairly uniform read depth across GC-content bins, while MC3 read depth was concentrated in regions of moderate GC-content (Fig. 5c). The low read depth in MC3 at regions of extreme GC-content was in turn associated with lower variant recovery rates in these regions but did not grossly affect the number of variants recovered by MC3 because regions of extreme GC-content are relatively rare in the genome overall and in exome-capture regions in particular.
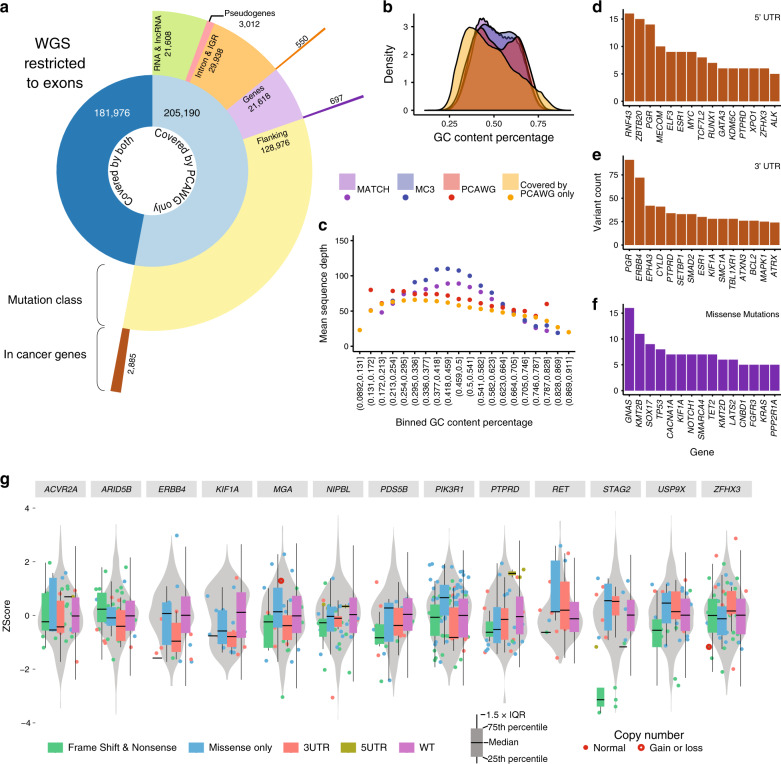
An in-depth analysis of these regions revealed that 76 mutations in known driver genes, identified in the combined TCGA data by Bailey et al. 2018, were missed in GC poor (GC fraction < 0.3) or GC rich (GC > 0.7) regions28. Three such instances revealed VHL mutations in KIRC that were overlooked in GC rich regions of this gene (two of these three recur). In addition, these 3 samples are not reported to carry a VHL mutation in the MC3 public data set. Other such instances include 7 SOX17 mutations, LATS2 (6), and CACNA1A (6). These findings emphasize the advantages of uniform coverage using WGS.
The bases flanking a mutation (tri-nucleotide context) affect mutation rate, which should be approximately equal between MC3 and PCAWG, and also the rate of introduction of sequencing artifacts. Large differences in the call-rates of MC3 and PCAWG and particular nucleotide sequences could indicate a sequencing artifact unique to one or the other call-set, which might arise from different procedures for computationally filtering or biochemically preventing sequencing oxidation products. Therefore, we sought to test whether the trinucleotide context of variants correlated with relative call-rates in MC3 and PCAWG. Before applying the MC3 OxoG filters, we found a huge predominance of CA variants unique to MC3, with the trinucleotide contexts most specific to one database or another being 7-9 times more specific than the least specific trinucleotide contexts. After applying the MC3 OxoG filters, nucleotide contexts differed by less than four-fold in their specificities. The residual differential specificity by trinucleotide context after filtering can either indicate differences in sequencing artifact abundance and filtration by project, or could merely be a consequence of the fact that nucleotide context is also correlated with VAF and the distance from transcription start sites, which may independently affect MC3 and PCAWG relative call-rates.
We extended the nucleotide context and performed mutation spectrum analysis, comparing all MC3 and all PCAWG mutations found after restricting the two data sets to exonic regions as described above (Step 3 of Fig. 1a). We then calculated transition and transversion frequencies in each cancer type. After removing hypermutated samples and OxoG artifacts, we used a chi-squared test to determine the similarities and differences between cancer types in the full exome space compared versus the captured exome space. Strikingly, we did not identify significant differences in mutation spectrum in the majority of cancers. We did observe significant differences (FDR < 0.05) in the mutation spectrum for COAD, KICH, LUAD, and OV (Supplementary Data 5). These observations included strong discrepancies for AG and CG transition differences in KICH and OV, respectively. AT and CA transversions contributed mostly to COAD and LUAD differences (Supplementary Fig. 9). While these differences may reflect sequencing artifacts, such as whole genome amplified DNA in OV or low sample size, we believe the data can still provide more information pertaining to additional cancer genes and oncogenic mechanisms.
Non-Coding/Flanking intersections with low coverage
With the growing use of WGS in many labs, we sought to identify which mutations are gained by extending to this form of data. One major observation from our pipeline highlighted that some variants in exome regions were not well covered by WES (Fig. 1a Step 3). Using this mutation set we investigated the most recurrent members as derived by WGS but not by MC3 in exonic regions as defined by gencode.v19 (Fig. 5a). We observed 697 mutations in cancer genes28 uniquely called by WGS (Supplementary Data 6). We defined flanking mutations as all non-translated mutations near exons, i.e., 5’UTR, 3’UTR, 5’Flanking, and 3’Flanking regions, as they make up the majority of mutations not present in the MC3 public MAF. Recurrent mutation analysis identified the most frequently mutated genes in 5’UTR (Fig. 5d), 3’UTR (Fig. 5e), and missense mutations (Fig. 5f). We found the most frequently mutated 3’UTR in cancer genes was PGR (91 mutations allowing for multiple mutations per sample), followed by ERBB4 (72), EPHA3 (42), CYLD (41), and PTPRD (34). To extend this analysis, we used RNAseq data collected by TCGA to determine mutation type specific cis-expression patterns, which clearly shows correlation of UTR mutations on RNA abundance (Fig. 5g).
Finally, similar to previous studies33,34, we investigated the potential effect of non-coding mutations when determining significantly mutated genes (SMG). Using MuSiC35 with the no-skip-non-coding option, we rescued non-coding mutations annotated by PCAWG and included them in the significantly mutated gene (SMG) analysis. We only performed SMG analysis on cancer types having greater than 35 samples (BRCA-Breast-AdenoCa, HNSC-Head-SCC, KICH-Kidney-ChRCC, LIHC-Liver-HCC, LUAD-Lung-AdenoCA, LUSC-Lung-SCC, SKCM-Skin-Melanoma, STAD-Stomach-AdenoCA, THCA-Thy-AdenoCA, and UCEC-Uterus-AdenoCA). We initially identified potential driver-gene candidates (FUT9, MMP16, SNHG14, and SFTPB, Fig. 6) not previously reported in Pan-Cancer whole genome analysis, but further investigation did not support these candidates with the exception of SFTPB.
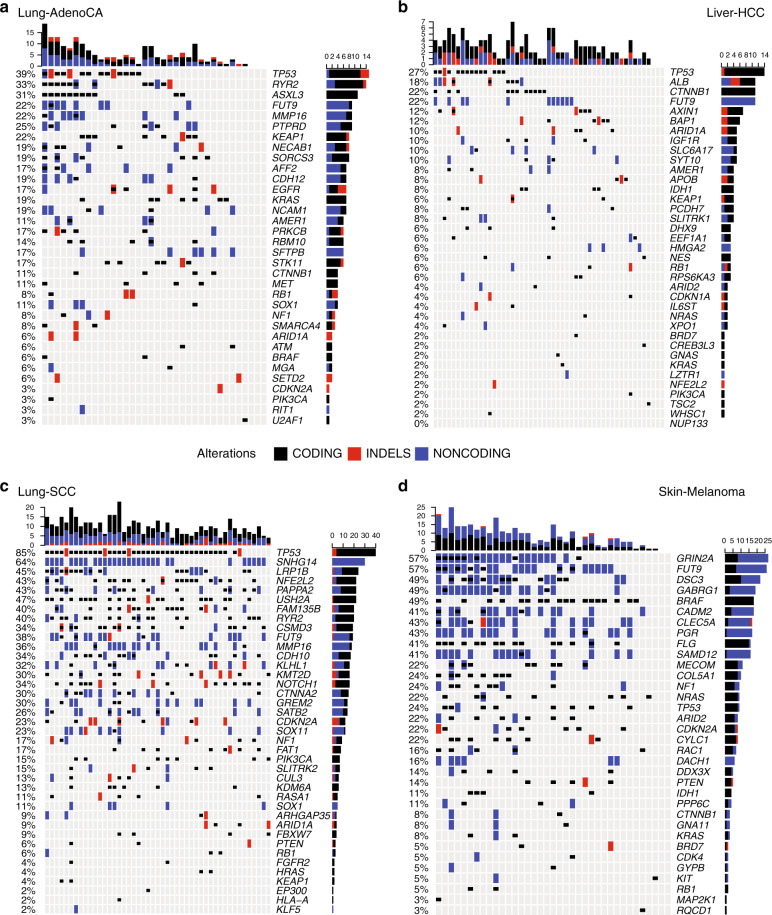
SFTPB (FDR 1.56e−07) in LUAD was recently reported to be significantly mutated using a larger set of these same data34. As reported, this gene is involved in a lineage-defining surfactant protein. While six mutations contributed to its SMG status, only 1 3’UTR mutation was reported for LUAD in the MC3 controlled data set. Furthermore, this single indel was only found by one variant caller (Varscan). We confirmed the impact of SFTPB UTR mutations by performing a genome-wide association analysis of expression differences and found that samples with SFTPB mutations showed lower RNA abundance in PCDHA7, a gene known to be involved in cells’ self-recognition and non-self-discrimination (chi-squared p-value 3.6 × 10-8). While other promising candidates exist, such as FUT9, a fucosyltransferase involved in organ bud progression during embryogenesis and has been implicated in cancer initiation36, we found no additional evidence for supporting its driver status.
Discussion
The research community is increasingly leveraging technology advances to integrate data at larger scales. We performed a comparative evaluation of ~750 samples with joint exome and whole genome sequencing mutation calls provided by two consensus mutation calling efforts, MC3 and PCAWG. This joint data set is encouraging, suggesting that ~80% of the predicted somatic mutations were captured by both efforts. Furthermore, a combined 90% of samples have greater than 80% variant concordance when considering covered exonic mutations from individual cohorts separately. Analysis of this data set also revealed three major contributors to variant discrepancies: (1) low variant allele fraction, (2) variant filtering decisions, and (3) technological limitations. Software differences were not an appreciable confounder.
Distinct advantages and disadvantages accompany somatic mutation calling when utilizing captured WES or WGS. We found that ~70% of the discrepancies between whole genome and whole exome sequencing are influenced by low variant allele fraction. This information holds many implications in identifying subclonal heterogeneity in the tumor of interest. Other discrepant calls originate from the decisions made on how to filter and distribute publicly available mutation calls. Higher-order mutation signature analysis does not appear to be inordinately affected by these differences. We show that reported germline variants were negligible, but nearly 30% of the private PCAWG mutations were not reported by MC3 because only a single variant detection algorithm identified them. We want to emphasize that, while somatic variant detection in cancer is commonplace, there are still many issues to reconcile.
Finally, we found additional mutations only observable in exonic regions using either WES or WGS. For example, WES uniquely identified 424 mutations in cancer genes with median VAF of ~10%. We also highlight ~700 WGS mutations from cancer genes, of which ~10% are attributable to regions of high and low CG-content; thus, highlighting the advantages of more uniform coverage from WGS.
Only about 2% of the genome is protein coding. For the last dozen years, cancer genomics has provided a comprehensive molecular characterization of many different tumor types, thanks in large part to The Cancer Genome Atlas and other publicly funded efforts. The community is just starting to explore how exomics, transcriptomics, proteomics, and methylomics can be woven together across this 2% of the genome. We anticipate a general transition from WES to WGS, but this analysis is meanwhile reassuring that few clerical mutations were overlooked in WES and that WGS is capable of recapitulating previous genomic findings.
Methods
Human research participants
The Cancer Genome Atlas (TCGA) collected both tumor and non-tumor biospecimens from human samples with informed consent under authorization of local institutional review boards (https://cancergenome.nih.gov/abouttcga/policies/informedconsent).
Sample overlap
TCGA barcodes carry metadata that reflect tumor portions and different aliquots. As noted in Fig. 1b, TCGA barcode differ slightly in the comparison between MC3 and WGS aliquots. A brief description of the breakdown of the TCGA barcode is outlined below.
Example: TCGA-DD-0001-01B-01D-A152
- – TCGA-Project
- – DD-Tissue source site: the tissue location of tumor that matches clinical metadata.
- – 0001-Participant code
- – 01-Sample type: i.e., solid tumor (01), primary blood derived tumor (03), solid tissue normal (11), blood derived normal (10)
- – B-Vial: the order in a sequence of samples, i.e., A = first in sequence, B = second in sequence
- – 01-Portion: sequential order of the 100–120 mg of samples
- – D-Analyte: molecular analyte type for analysis, i.e., D for DNA and W for WGA.
- – A152-Plate: sequential location of a 96-well plate
A lookup table outlining these fields is located at the GDC: https://gdc.cancer.gov/resources-tcga-users/tcga-code-tables. In order to determine the role of aliquot differences in assessing mutation concordance, we re-analyzed the clonality and overall mutation overlap after stratifying for exact barcode differences. We observed that the effect of matching barcodes on match variant frequency has little effect.
Assessing cancer subtype selection preference
Analysis working groups for TCGA projects were primarily subdivided according to cancer types. Scientific experts gathered in consortia from around the country to participate in characterizing many tumors using high throughput data generated on many substrates, e.g., WES, RNAseq, etc. At the conclusion of these projects, groups were asked to hand-select a subset of samples to perform validation sequencing (WGS, the samples used in this analysis). The selection criteria differed from group-to-group and sometimes resulted in an overabundance of one subtype over another. To determine cancer subtype selection bias, we performed an enrichment analysis. Using clinical data we calculated (for each cancer type) the subtype fraction in the WES cancer cohort and measured whether the fraction was similar to WGS set of samples using a Fisher’s exact test.
Defining exonic regions
We used the same definition as Ellrott et al. to reduce whole genome and exome calls to define genomic coordinates27.
Coverage calculations
Fixed-step (step = 1) Wiggle coverage files (.wig) for both MC3 and PCAWG were provided by the Broad Institute. The wig format is a binary readout of sufficient sequencing coverage for genomic data. Here, sufficient coverage is defined as bases with 8 or more reads at a given location. These wig files were processed and reduced to exonic regions using the wig2bed function from BEDOPTS37.
After the preliminary screen of coverage-reduced MAF files, we observed that matching mutations (identified by PCAWG and MC3) were removed from one technology and not the other after the coverage reduction step. To account for this issue, we performed a self-coverage reduction step to that identified 6460 mutations. We describe some properties of those mutations here. The median tumor depth reported by MC3 from these variants is 12 reads (+/− 3 median absolute difference). The median tumor depth reported by PCAWG in this region is 39 reads (+/− 35.6 median absolute difference), suggesting wide variance of tumor read depths that were removed. However, the mode tumor depth of the PCAWG variants was 13, justifying this removal of variants with low read support. Finally, we determined how many of these poorly-covered variants originated from cancer driver genes. We observed 126 mutations from the MC3 file, and 156 cancer mutations were eliminated at this stage in the comparison.
Overlapping mutations
After reducing the variants to be within exome sequencing target region, within same exon definitions, and having enough sequencing depth, the remaining variants from ICGC and PCAWG were stored in a SQLite database to enable fast lookup. We then executed a full join between the two sources of variants by matching the donor ID, sample ID, and the genomic range of each variant. The full join output was further cleaned up to remove duplicated filters due to naming variations and duplicated variants due to DNPs.
- – Matching IDs
- – Matching chromosomes
- – End position greater than or equal to start position
- – Start position is less than or equal to end position.
Deduplication of variants
After merging the PCAWG and MC3 data, we observed different strategies were taken by MC3 and PCAWG to capture neighboring variants, i.e., complex indels, di-nucleotide (DNP) and tri-nucleotide (TNP) polymorphisms. To address complex indel events (SNVs in indel regions), the MC3 working group absorbed the variants made by SNV callers into the assignment made by Pindel. Conversely, PCAWG merged DNP and TNP events into a single event. These strategies resulted in many duplication events from MC3 and PCAWG: 1731 and 62, respectively. These events encompassed 3457 and 119 events, respectively. To address these differences, we merged PCAWG variants into MC3s complex indel events, and MC3 variants into single DNP or TNP events.
Filtering optimization
After reducing the starting pool of possible mutations from 746 samples to covered exons, we sought to identify the optimal set of MC3 filters that provide the largest number of samples with greater than 80% concordance from the two technologies with the simplest schema. This was performed comprehensively using all possible combinations of filters, often with more than one filter per variant, with the MC3 cohort (131,071 filter combinations). Filter flags include: “common_in_exac”, “gapfiller”, “native_wga_mix”, “nonpreferredpair”, “oxog”, “StrandBias”, and “wga”. We pre-defined the exclusion of variants in MC3 flagged as OxoG along with the inclusion of all PASS variants. The comprehensive filter analysis resulted in two major clusters of variant recoverability (Supplementary Fig. 3). Here, we observed the computational trade-off of identifying more matched variants at the cost of more unique MC3 calls. Below, we highlight five strategies considered for analysis (Supplementary Data 2).
- Only consider variants labeled PASS by the MC3 filter column.
- Only remove variants labeled OxoG by MC3.
- Prioritize G1 (samples in the most recoverable quadrant, MC3 and PCAWG samples with greater than or equal to 80% from both efforts.)
- Prioritize total number of matched variants.
- Maximize total number of samples in the most recoverable quadrant (Fig. 2b) while maximizing the difference between unique MC3 variants and matched variants thus generating fewer unique calls.
After considering complexity, we chose to move forward with strategy 2 for the entirety of this study due to its simplicity and relative similarity to other filtering schemes. We recognize that by selecting a single filtering strategy, we are limiting the data slightly and likely introducing some false positive variant calls. However, this strategy allowed us to maintain larger sample sizes and to capture ~15,000 more matched variants than the PASS only strategy at the cost of ~3500 unique mutations calls for MC3.
Assessing mutations per megabase and cancer type concordance
Mutations per megabase data were collected from the broader TCGA dataset and reduced following the same methods outlined previously28. Briefly, this systematically removed hypermutators from the dataset. This resulted in a set of 625 samples from the MC3/PCAWG dataset studied here and 8852 TCGA samples. Both Pearson and Mann-Whitney correlations statistics were performed to assess the association of non-silent mutations per megabase and concordance statistics.
Simulation of sequencing noise and recoverability
Fan et al. benchmarked the sensitivity of MuSE at recovering somatic variants across 24 combinations of VAF and read depth3. When simulating the recovery of PCAWG variants in MC3 we assumed that the VAF observed in PCAWG was the true VAF. We matched the observed VAF of each variant to the closest VAF reported in Fan et al.
For our analysis, the best value to use as the read depth when predicting the MC3 recovery rate of PCAWG variants would be the MC3 read depth at the same site and sample as the PCAWG variant. However, it was not practical to obtain MC3 read depths at sites without MC3 variants, so instead we simulated an MC3 read depth for each PCAWG variant by randomly sampling from the read depths of observed MC3 variants from the same sample as the PCAWG sample. We then matched these simulated read depths for each variant to the closest read depths reported in Fan et al.
For the binned VAFs and read depths for each PCAWG variant obtained as above, we pulled the corresponding sensitivities of MuSE from the Fan et al. paper and simulated MC3 variants with probability equal to these sensitivities.
Integrating clonality
Both consortia considered clonality in their comprehensive characterization of the somatic mutations. Locations of these files are provided in the data availability section. Here, we provide a brief summary of the strategies used to compile these resources. First, the PanCancer Atlas working-groups used MC3 mutations to predict subclonal structures using ABSOLUTE38. This tool uses copy number, recurrent karyotype, and mutation data to calculate copy number purity and cluster identification. Furthermore, the PanCancer Atlas working group only made the distinction of clonal and subclonal mutations and did not attempt to further assign sub-clonal mutations to other likely heterogeneous clusters. PCAWG, on the other hand, used a consensus calling approach incorporating 11 different clustering tools. Here, we evaluated cluster-ID which represents those mutations that are clonal (ID = 1), with other clusters representing mutations that are subclonal (ID = 2 through 4). For this analysis, we restricted our data to SNVs to be consistent with calls made by the PanCancer Atlas calls of MC3 mutations.
Fraction of private variation explained
In Supplementary Fig. 7 we provide a breakdown of different sources of variant described in our analysis using publicly available data. For MC3 all private variants were classified as into 3 variant types (Indel, MissensePlus, and Other). Specifically, indels are comprised of: “Frame_Shift_Del”, “In_Frame_Ins”, “Frame_Shift_Ins”, and “In_Frame_Del”. MissensePlus variants are comprised of: “Missense_Mutation”, “Nonsense_Mutation”, “Nonstop_Mutation”, “Splice_Site”. And Other variants are comprised of: “RNA”, “3’UTR”, “5’UTR”, “5’Flank”, “Silent”, “3’Flank”, “Intron”, “Translation_Start_Site”.
On the other hand, PCAWG variants were also categorized into Indels, MissensePlus, and Other. Specifically, indels are comprised of: “Frame_Shift_Del”, “Frame_Shift_Ins”, “De_novo_Start_InFrame”, “Start_Codon_Ins”, “Stop_Codon_Ins”, “In_Frame_Del”, “In_Frame_Ins”, “Stop_Codon_Del”, and “Start_Codon_Del”. MissensePlus variants are comprised of: “Missense_Mutation”, “Nonsense_Mutation”, “Nonstop_Mutation”, “Splice_Site”. And other variants are comprised of: “5’UTR”, “RNA”, “5’Flank”, “Silent”, “3’UTR”, “Intron”, “IGR”, “lincRNA”, “De_novo_Start_OutOfFrame”, and “Start_Codon_SNP”.
In addition to the three variant type categories, six additional sources of variation were added to private variants: Subclonal, VAF5, VAF10, MMcomplement, THCA KICH or PRAD, and GCcontents. As mentioned, subclonal variants are tagged if labeled as identified by the TCGA or ICGC consortia. VAF5 tags all variants with less that 5% VAF. VAF10 tags all variants with VAF greater than or equal to 5% and less that 10%. MMcompelement tags all variants not detected by MuSE or MuTect. And finally, GCcontent was calculated as the fraction of G or C nucleotides in a 100 bp window surround a variant. This was calculated using a CADD plug-in to VEP. The GCcontent flag was assigned to a variant if GC fraction was less and 0.3 or greater than 0.7.
MAFit: online comparison and visualization tool
MAFit (maf Interaction Tool) is a shinyapp39 tool to visualize and extract mutations from the intersection of PCAWG and MC3 call sets. It is an interactive and graphical web-based interface built using R Shiny. The interface consists of three panels: a main scatter plot display, a side box of control widgets, and a mutation table in the bottom pane. Any alteration of a control widget will update the scatter plot and the mutation table, enabling very rapid browsing. There is also a button to download the current data set displayed in the scatter plot.
The main panel displays a scatter plot with marginal histograms of mutations grouped by sample. The axes are percentage of matched mutations versus matched plus call-set-unique mutations. Mouse-hovering on a data point initiates a pop-up window showing specific information for this sample, such as TCGA barcode, number of matched mutations, and numbers of mutations unique to each call-set.
The side panel contains two sets of control widgets which can be used to select data based on different criteria. The top control set consists of two sliders to set the VAF cut-offs for each call-set. Both ends of the slider can be adjusted so that users can obtain a desired interval of the VAF. The bottom control set consists of check-boxes of both variant-level and sample-level MC3 filters. If only variant-level filters are checked, all PCAWG-only mutations will be retained; if at least one sample-level filter is checked, mutations from samples that do not have any checked filters flagged (variant-level or sample-level) will be filtered out. Both variant-level and sample-level filters result in the union of mutations with any checked filter will be shown.
The bottom panel displays a table of mutations based on the selection criteria from the side panel. Columns include information on each mutation, such as TCGA barcode, gene name, VAF, variant class, Human Genome Variant Society (HGVS) nomenclature, etc. Users can sort the table by each column. There are two drop-down selection boxes where users can view mutations from a specific TCGA barcode or Hugo symbol. There is also a search bar, which results in mutations that contain the input in any columns.
Controlled access files
Having worked with both the TCGA and ICGC consortia, we were privy to the controlled access data (all MC3 somatic variant calls and PCAWG germline calls). These data sets allowed for the further interrogation of unique variants called by both groups.
We performed a mutation variant intersection of MC3 controlled access mutations with unique PCAWG variants in the captured exonic regions. These data can be downloaded using necessary credentials from https://gdc.cancer.gov/about-data/publications/mc3-2017.
We intersected the MC3 public MAF with the PCAWG germline call-set, donor-by-donor. Six MC3 somatic variants were identified as germline variants in PCAWG for the same donor. Of these, five were flagged in MC3 as OxoG or non-preferred-pair artifacts, and only one was marked PASS in MC3. This PASS variant had a depth in the matched MC3 normal of well over 100 with no alternate reads. The minimal intersection between the MC3 somatic call-set and the donor-matched PCAWG germline call-set is evidence that germline contamination in MC3 calls is low.
Assessment of impact on mutation signature analysis
We ran SignatureAnalyzer40 on the corpus of WES and WGS samples from our TCGA cases. This tool reports a vector of J = 7 normalized weights corresponding to mutational signatures. Once weights were computed, we used COSMIC signatures as a reference in order to synchronize labels of the fractional weights between WES and WGS data for each case to enable proper comparison. We excluded 5 cases in which signatures were not computed for WGS data. Using the synchronized results, we then assessed both the number of cases for which the tool identified the same dominant signature between WES and WGS data and also evaluated the correlation between WES and WGS vectors for each case using least-squares regression. Statistical significance of each correlation was calculated using a 2-tailed t-test. Statistical power of each correlation was limited by the paucity of signature weights because the underlying t-statistic is proportional to the square root of J – 2. However, because these cases, and therefore their hypothesis tests, are independent, the cohort constitutes multiple tests of the same hypothesis regarding signatures derived from WES and WGS data. Therefore, we combined individual P-values into an overall cohort P-value using Fisher’s log-transform. Namely, the transform of negative 2 times the natural log of the product of the K = 739 individual P-values is, itself, chi-square distributed with 2 K degrees of freedom. Using this transform, we found an overall cohort P-value of <2 × 10−6.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Supplementary Materials
References
- 1.Radenbaugh, A. J. et al. RADIA: RNA and DNA integrated analysis for somatic mutation detection. PLoS ONE9, e111516 (2014).
- DC Koboldt. VarScan 2: Somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res., 2012. [DOI | PubMed]
- Y Fan. MuSE: accounting for tumor heterogeneity using a sample-specific error model improves sensitivity and specificity in mutation calling from sequencing data. Genome Biol., 2016. [DOI | PubMed]
- K Cibulskis. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol., 2013. [DOI | PubMed]
- K Ye, MH Schulz, Q Long, R Apweiler, Z Ning. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics, 2009. [DOI | PubMed]
- K Ye. Systematic discovery of complex insertions and deletions in human cancers. Nat. Med., 2016. [DOI | PubMed]
- MA Chapman. Initial genome sequencing and analysis of multiple myeloma. Nature, 2011. [DOI | PubMed]
- DE Larson. Somaticsniper: Identification of somatic point mutations in whole genome sequencing data. Bioinformatics, 2012. [DOI | PubMed]
- V Moncunill. Comprehensive characterization of complex structural variations in cancer by directly comparing genome sequence reads. Nat. Biotechnol., 2014. [DOI | PubMed]
- 10.The ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Network. Pan-cancer analysis of whole genomes. Nature578, 82–93 (2019).
- D Sims, I Sudbery, NE Ilott, A Heger, CP Ponting. Sequencing depth and coverage: key considerations in genomic analyses. Nat. Rev. Genet., 2014. [DOI | PubMed]
- 12.Chilamakuri, C. S. R. et al. Performance comparison of four exome capture systems for deep sequencing. BMC Genomics15, 449 (2014).
- A Belkadi. Whole-genome sequencing is more powerful than whole-exome sequencing for detecting exome variants. Proc. Natl Acad. Sci. USA, 2015. [DOI | PubMed]
- DH Spencer. Performance of common analysis methods for detecting low-frequency single nucleotide variants in targeted next-generation sequence data. J. Mol. Diagnostics, 2014. [DOI]
- A Alfares. Whole-genome sequencing offers additional but limited clinical utility compared with reanalysis of whole-exome sequencing. Genet. Med., 2018. [DOI | PubMed]
- J Phallen. Early noninvasive detection of response to targeted therapy in non–small cell lung cancer. Cancer Res., 2019. [DOI | PubMed]
- D Głodzik. Mutational mechanisms of amplifications revealed by analysis of clustered rearrangements in breast cancers. Ann. Oncol., 2018. [DOI | PubMed]
- S Ren. Whole-genome and transcriptome sequencing of prostate cancer identify new genetic alterations driving disease progression [Figure presented]. Eur. Urol., 2018. [DOI | PubMed]
- S Zhang. Genome evolution analysis of recurrent testicular malignant mesothelioma by whole-genome sequencing. Cell. Physiol. Biochem., 2018. [DOI | PubMed]
- S Nik-Zainal. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature, 2016. [DOI | PubMed]
- A Mendoza-Alvarez. Whole-exome sequencing identifies somatic mutations associated with mortality in metastatic clear cell kidney carcinoma. Front. Genet., 2019. [DOI | PubMed]
- P Yan. Whole exome sequencing of ulcerative colitis-associated colorectal cancer based on novel somatic mutations identified in Chinese patients. Inflamm. Bowel Dis., 2019. [DOI | PubMed]
- 23.Mogensen, M. B. et al. Genomic alterations accompanying tumour evolution in colorectal cancer: tracking the differences between primary tumours and synchronous liver metastases by whole-exome sequencing. BMC Cancer18, 752 (2018).
- M Sponziello. Whole exome sequencing identifies a germline MET mutation in two siblings with hereditary wild-type RET medullary thyroid cancer. Hum. Mutat., 2018. [DOI | PubMed]
- K Schwarze, J Buchanan, JC Taylor, S Wordsworth. Are whole-exome and whole-genome sequencing approaches cost-effective? A systematic review of the literature. Genet. Med., 2018. [DOI | PubMed]
- M Choi. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc. Natl Acad. Sci. USA, 2009. [DOI | PubMed]
- K Ellrott. Scalable open science approach for mutation calling of tumor exomes using multiple genomic pipelines. Cell Syst., 2018. [DOI | PubMed]
- MH Bailey. Comprehensive characterization of cancer driver genes and mutations. Cell, 2018. [DOI | PubMed]
- 29.Meienberg, J. et al. New insights into the performance of human whole-exome capture platforms. Nucleic Acids Res.43, e76 (2015).
- MJ Clark. Performance comparison of exome DNA sequencing technologies. Nat. Biotechnol., 2011. [DOI | PubMed]
- W McLaren. Deriving the consequences of genomic variants with the Ensembl API and SNP effect predictor. Bioinformatics, 2010. [DOI | PubMed]
- M Kircher. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet., 2014. [DOI | PubMed]
- 33.Rheinbay, E. et al. Analyses of non-coding somatic drivers in 2,658 cancer whole genomes. Nature578, 102–111 (2019).
- M Imielinski, G Guo, M Meyerson. Insertions and deletions target lineage-defining genes in human cancers. Cell, 2017. [DOI | PubMed]
- ND Dees. MuSiC: Identifying mutational significance in cancer genomes. Genome Res., 2012. [DOI | PubMed]
- N Auslander. An integrated computational and experimental study uncovers FUT 9 as a metabolic driver of colorectal cancer. Mol. Syst. Biol., 2017. [DOI | PubMed]
- S Neph. BEDOPS: High-performance genomic feature operations. Bioinformatics, 2012. [DOI | PubMed]
- SL Carter. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol., 2012. [DOI | PubMed]
- 39.Chang, W., Cheng, J., Allaire, J., Xie, Y. & McPherson, J. shiny: Web Application Framework for R. R Packag (2016).
- 40.Kasar, S. et al. Whole-genome sequencing reveals activation-induced cytidine deaminase signatures during indolent chronic lymphocytic leukaemia evolution. Nat. Commun. 6, 1–12 (2015).
- JR Conway, A Lex, N Gehlenborg. UpSetR: An R package for the visualization of intersecting sets and their properties. Bioinformatics, 2017. [DOI | PubMed]
- Z Gu, R Eils, M Schlesner. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics, 2016. [DOI | PubMed]
