Forty sites of TRP channel regulation
Abstract
Transient receptor potential (TRP) channels are polymodal molecular sensors that integrate chemical, thermal, mechanical and electrical stimuli and convert them into ionic currents that regulate senses of taste, smell, vision, hearing, touch and contribute to perception of temperature and pain. TRP channels are implicated in the pathogenesis of numerous human diseases, including cancers, and represent one of the most ardently pursued drug targets. Recent advances in structural biology, particularly associated with the cryo-EM “resolution revolution”, yielded numerous TRP channel structures in complex with ligands that might have therapeutic potential. In this review, we describe the recent progress in TRP channel structural biology, focusing on the description of identified binding sites for small molecules, their relationship to membrane lipids, and interaction of TRP channels with other proteins. The characterized binding sites and interfaces create a diversity of druggable targets and provide a roadmap to aid in the design of new molecules for tuning TRP channel function in disease conditions.
Article type: Research Article
Keywords: TRP channels, Cryo-EM, X-ray crystallography, Gating, Activation, Inhibition, Block, Ligand, Agonist, Antagonist
License: CC BY 4.0 This is an open access article under the CC BY license (https://creativecommons.org/licenses/by/4.0/).
Article links: DOI: 10.1016/j.cbpa.2024.102550 | PubMed: 39615427 | PMC: PMC11788071
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (2.1 MB)
Introduction
The superfamily of cation-selective TRP channels includes seven subfamilies: TRPV (vanilloid, TRPV1–6), TRPC (canonical, TRPC1–7), TRPM (melastatin, TRPM1–8), TRPN (NOMPC-like), TRPA (ankyrin, TRPA1), TRPP (polycystin, TRPP1–3) and TRPML (mucolipin, TRPML1–3). We will discuss structural pharmacology of all subfamilies, except TRPN, for which the available structural information is limited.
Structural architecture of TRP channel subfamilies
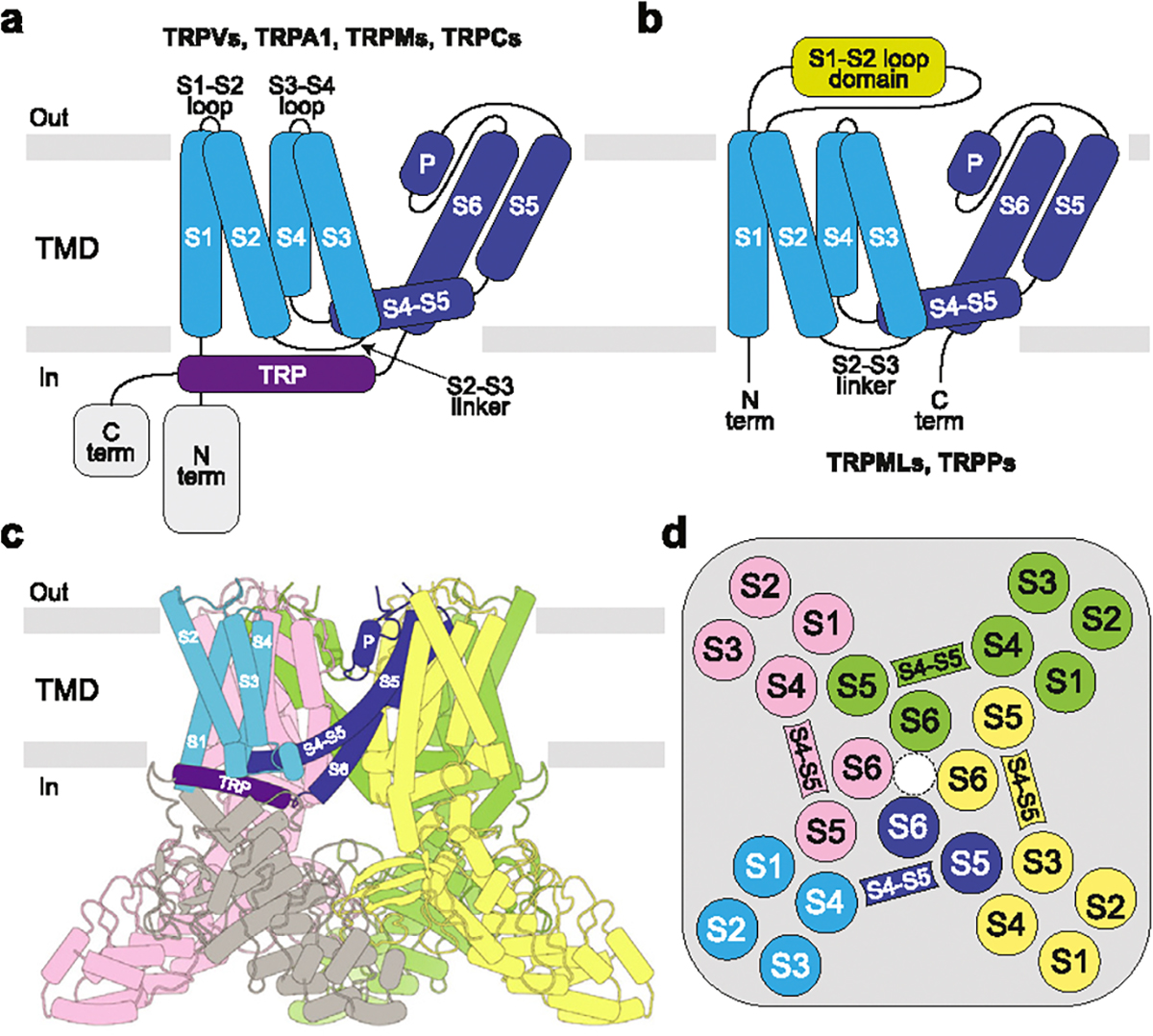
TRP channels are tetramers, with individual subunits composed of the intracellular N- and C-terminal domains and the transmembrane domain (TMD) in the middle (Figure 1). TMD is the most conserved region across the TRP channel superfamily, which includes six transmembrane (TM) helices (S1–S6) and a reentrant pore loop (P-loop) between S5 and S6, followed by the family signature domain–TRP helix. The first four TM helices comprise the S1–S4 or voltage sensing-like domain (VSLD). Four S1–S4 domains surround the ion channel pore in the center of the TMD, contributed by the pore domains (S5, P-loop and S6) of all four subunits. The N- and C-terminal domains are subfamily-specific and entirely absent in TRPP and TRPML channels, which also lack the TRP-helix but instead have extended S1–S2 loops that form polycystin or mucolipin domains, respectively.
Ligand binding sites
Structural studies characterized the interaction of ligands with specific regions of the TRP channel proteins which can be classified as forty distinct ligand binding sites, based on their location with respect to the conserved structural elements (Figure 2, Supplementary Table 1). These sites are located either within the individual domains (N- or C-terminal or TMD) or at the domain interfaces. The most common binding sites, which have been reported for several TRP channel subfamilies, are vanilloid (#1), S1–S4 base (#13), portal (#10–11), S4–S5 linker (#19–20) and S2–S3 calcium (#22) sites. All five have dual functionality and can bind both activators and inhibitors.
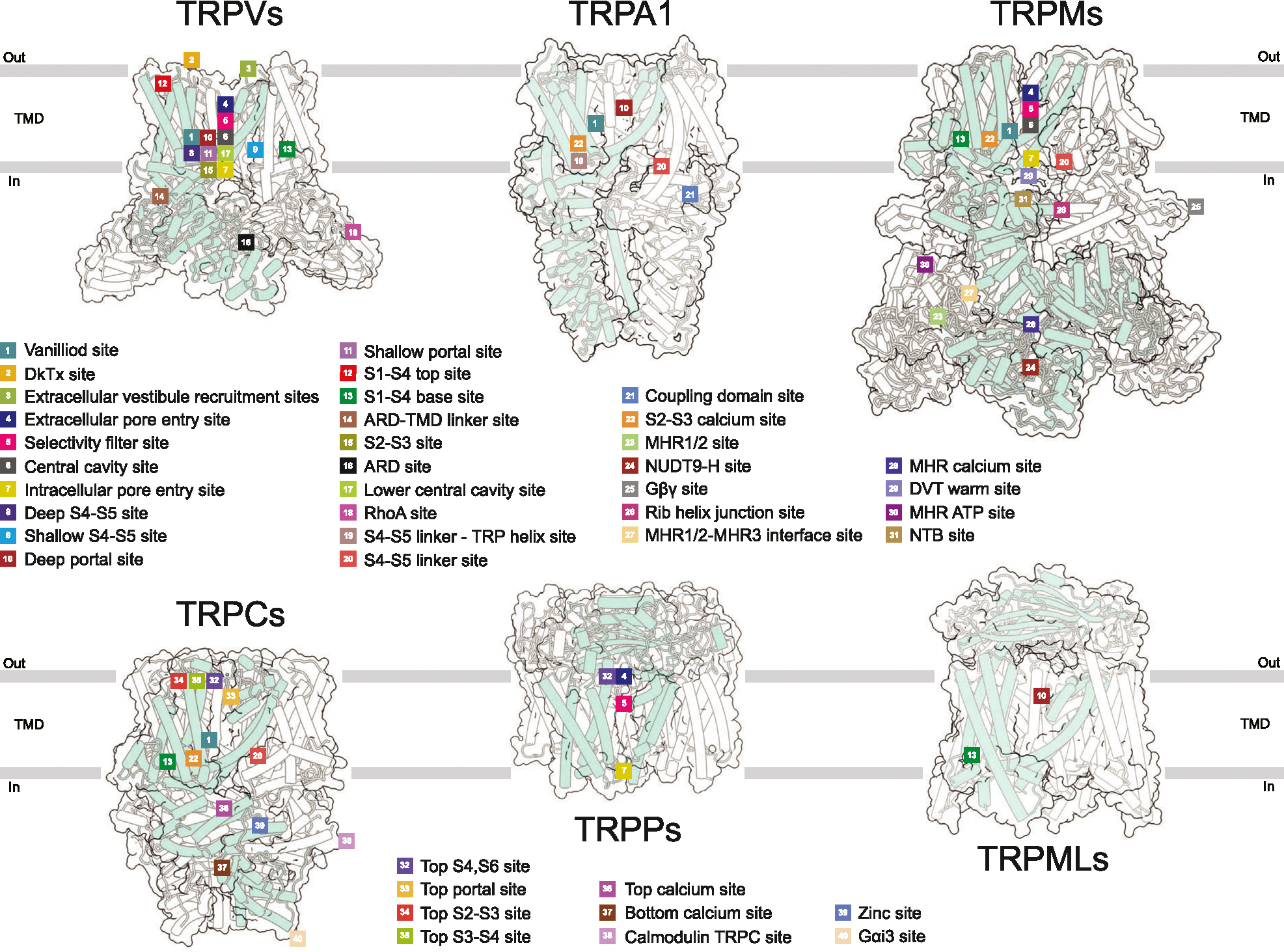
The vanilloid site (#1) is located at the interface between the S1–S4 and pore domains and contributed by the S4–S5 linker, S3, S4 and TRP helices of one subunit and S5 and S6 helices of the adjacent subunit [ref. 1]. Both agonists (capsaicin and resiniferatoxin) and competitive antagonists (capsazepine, SB-366791, and SAF312) can occupy this site in TRPV1. The agonists engage Arg557 into a salt bridge with Glu570, pulling S5 towards S1–S4 domain and leading to channel opening. In contrast, the antagonists disfavor the Arg557-Glu570 interaction and stabilize the closed state [ref. 2,ref. 3]. The vanilloid site also binds phosphoinositol lipids, lysophosphatidic acid, and cholesterol in TRPV1; agonist 2-APB and cholesterol in TRPV2; agonist THCV and inhibitor Trpvicin in TRPV3; cholesteryl hemisuccinate (CHS) and channel blockers PCHPDs in TRPV6; inhibitor BTDM in TRPC6; antagonist NDNA in TRPM5; activator AITC in TRPM8; inhibitors CCT128930, NS8593, VER155008, and activator naltriben in TRPM7; and non-covalent agonist GNE551 in TRPA1 (Supplementary Table 1).
The S1–S4 base (#13) site represents a cavity formed by the intracellular ends of all four helices of the S1–S4 bundle and the TRP helix [ref. 1]. In the absence of added ligands (apo condition), this site in TRPV subfamily representatives is typically occupied by a lipid. The S1–S4 base site also binds activator 2-APB and inhibitor osthole in TRPV3; agonists 4α-PDD and Agonist1, and antagonists A1, A2, GSK1016790A, and HC-067047 in TRPV4; inhibitor ZINC17988990 in TRPV5; inhibitor 2-APB in TRPV6; agonists Cryosim-3, icilin, and WS-12 in TRPM8; inhibitors GFB-8438, GFB-8749, and GFB-9289 in TRPC4; activator riluzole in TRPC5; antagonist AM-1473 and SAR7334 in TRPC6; and phospholipids PI(3,5)P2 and PI(4,5)P2 in TRPML1 (Supplementary Table 1).
The portal site (#10–11) is the closest to the pore site located at the intersubunit interface, contributed by S5 of one subunit as well as P-loop and S6 of another subunit [ref. 1]. The portal site accommodates cholesterol in TRPV1; activator CBD in TRPV2; inhibitor dyclonine in TRPV3; inhibitor THCV in TRPV6; inhibitor GDC-0334 in TRPA1; inhibitors HC-070 and Pico145 (HC-608) in TRPC5; as well as agonists ML-SA1 and temsirolimus, and inhibitor ML-SI3 in TRPML1 (Supplementary Table 1). The S4–S5 linker site (#19–20) is contributed by S1, S4–S5 linker and TRP helix as well as the linker domain in TRPC and TRPM channels or interfacial helix in TRPA1. The S4–S5 site accommodates the antagonist 3–60 in TRPA1; CHS in TRPC3–6 and TRPM8; and PI(4,5)P2 in TRPM3 and TRPM8 (Supplementary Table 1).
The S2–S3 calcium site (#22) represents a common site for Ca2+ ion in TRPA1, TRPC3-TRPC6, TRPM2, TRPM4–5, and TRPM8 (Supplementary Table 1). It is comprised of four residues (D, E, N, or Q in different combinations) that belong to intracellular regions of S2 and S3. Ca2+ ion bound to the S2–S3 site can either have the activating (TRPC3,6 and TRPM4,5) or regulatory (TRPA1, TRPM2,8) roles, either enhancing ligand binding or desensitization [ref. 4,ref. 5]. Ca2+ is displaced from the S2–S3 site in TRPC6 by the inhibitors AM-1473 [ref. 6] and SAR7334 [ref. 5]. On the other hand, Ca2+ helps to coordinate the activator riluzole in TRPC5 [ref. 7] and cooling agonist icillin in TRPM8 [ref. 8].
While different ligands — both activators and inhibitors — can bind to a single site, the same ligand can bind to different sites in one ion channel simultaneously. For example, 2-APB binds to three distinct sites in TRPV3 (#12, #13, #14), but it also binds to TRPV2 (#1, #9) and TRPV6 (#13) [ref. 1,ref. 9,ref. 10]. Naltriben binds to two different sites in TRPM7, resulting in either channel activation (NTB site, #31) or inhibition (vanilloid site, #1) [ref. 11]. Strikingly, recent structural studies of TRPM4 showed that decavanadate (DVT) binding is temperature-dependent, occurring either at the MHR1/2-MHR3 (#27) or warm DVT (#29) sites [ref. 12] (Supplementary Table 1). In addition, different sites can be occupied by different ligands, which activate or inhibit TRP channels cooperatively or synergistically [ref. 13–ref. 17]. For instance, the activation of TRPV2 by 2-APB and CBD applied simultaneously is much stronger than the cumulative activation produced by these agonists applied individually [ref. 16].
Lipid binding sites
In the apo state, many ligand-binding sites are occupied by annular lipids, which in the high-resolution cryo-EM structures of TRP channels are represented by non-protein densities around the TMD. Some lipids are required for protein stability (structural lipids) [ref. 18,ref. 19], while others modulate channel function [ref. 20], playing activatory or inhibitory roles (Figure 3). The two classes of lipid molecules that are commonly recognized in cryo-EM structures due to their distinct features are cholesterol and phospholipids.
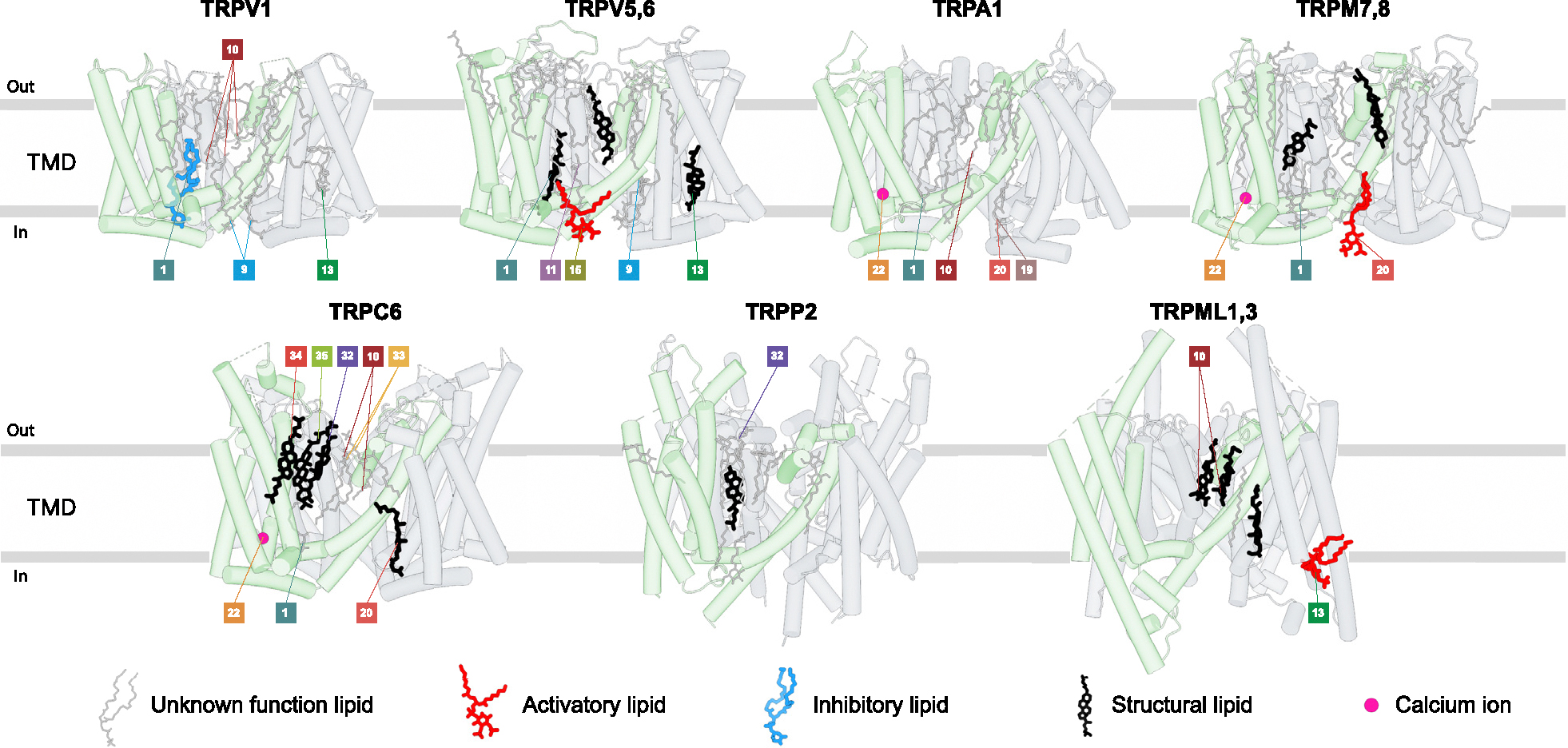
Densities for cholesterol or its more soluble acidic ester CHS, often used in protein purifications, have a clearly recognizable flat shape of sterol with holes and bumps corresponding to cyclohexane rings and methyl groups, respectively. Among the vanilloid-subfamily TRP channels, temperature-insensitive calcium-selective TRPV5–6 appear to require cholesterol or CHS for their structural integrity, as every structure of these channels show structural lipids at the ligand-binding sites #1 (vanilloid), #13 (S1–S4 base), and #33 (top portal) (Figure 3). In temperature-sensitive TRPV channels, on the other hand, only one site (#1, vanilloid) was reported to bind endogenous cholesterol in TRPV2 [ref. 10] and two sites (deep portal [ref. 19] and adjacent to vanilloid [ref. 13] sites) in TRPV1. Cholesterol was also reported to sensitize TRPV3 [ref. 21] and regulate function, expression, and the cellular response to tension of TRPV4 [ref. 22]. However, the corresponding sites of cholesterol binding to TRPV3 and TRPV4 await structural determination. While CHS is frequently used as a substitute for endogenous cholesterol, it is important to keep in mind that the hydrophilic hemisuccinate group in CHS versus the hydrophobic hydrocarbon group in cholesterol may cause differences in effects of these lipids on certain TRP channels.
Densities for phospholipids have a distinct head-and-two-tails appearance. The head density of the majority of phospholipids (excluding inositol phospholipids) has an elongated cylindrical shape, making it difficult to distinguish between different types of phospholipids. Accordingly, while many putative phospholipids consistently appear in different TRP channel structures, assigning their structural or functional roles is often difficult due to unclear chemical identity of the lipid and uncertain binding site specificity. In rare cases, the functional role of phospholipids becomes apparent when their site occupancy changes during gating. An example is a phospholipid in TRPV3, which occupies the vanilloid site (#1) in the closed state but must leave this site for the channel to open in response to heat [ref. 23,ref. 24] or agonist binding [ref. 9]. For inositol phospholipids, the head density resembles a donut, with additional bumps representing the number of phosphate groups attached to the inositol ring. Phosphatidylinositol lipids are known to regulate many TRP channels, serving as essential cofactors for their activity [ref. 20]. Thus, TRPV1 has a residential phosphatidylinositol lipid that is crucial for its stability and defines the temperature activation threshold and affinity to ligands [ref. 19]. PI(4,5)P2 was shown to activate all TRPM subfamily members except TRPM1 [ref. 20], with structures available for TRPM8 and TRPM3 [ref. 8,ref. 17,ref. 25,ref. 26]. On the other hand, the intracellular TRPML1 channel is inhibited by PI(4,5)P2 and activated by PI(3,5)P2, both binding to the same site [ref. 14,ref. 27]. PI(4,5)P2 was reported necessary for TRPV1–4, TRPA1, and TRPC4–5 activity but also as their inhibitor [ref. 20,ref. 28]. Structures of these channels bound to PI(4,5) P2 should untangle their complex interaction.
While several ligand-binding sites host lipids in the apo condition (Figure 3), many lipid binding locations have not been identified as sites for ligands. We anticipate future discoveries of ligands targeting the corresponding protein-lipid interfaces and acting by outcompeting endogenous lipids.
TRP channel protein binding partners
In addition to small molecules, the function of ion channels in general, and TRP channels in particular, is often regulated by other proteins, which represent either transmembrane auxiliary subunits or soluble binding partners. Cryo-EM structures have been recently solved for complexes of TRPV5/6 with CaM [ref. 29–ref. 32], TRPV4 with RhoA [ref. 33–ref. 35], TRPC4 with CaM [ref. 36], TRPM3 with Gβγ [ref. 25], and TRPC5 with Gαi3 [ref. 37] (Figure 4).
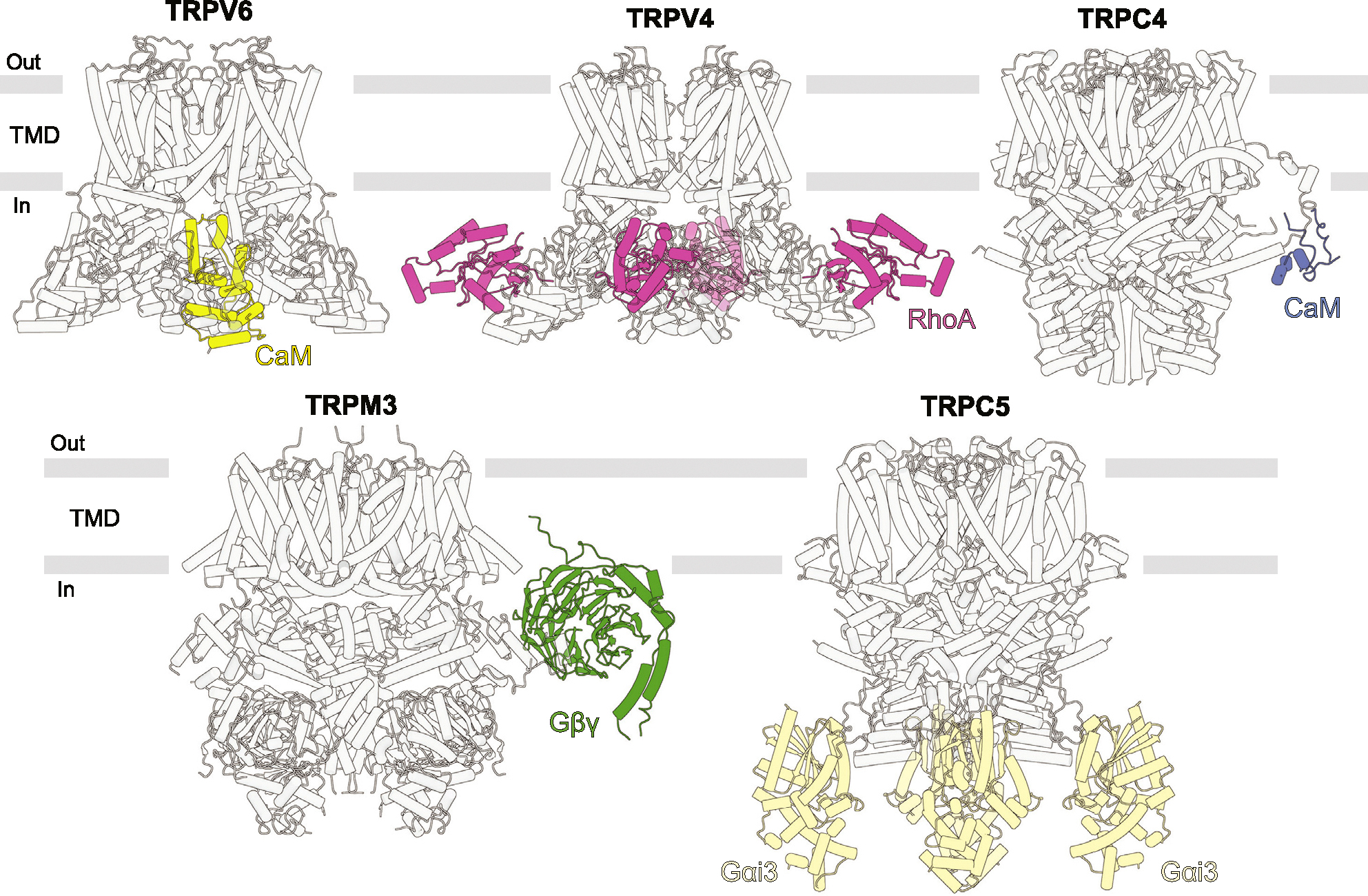
The TRPV5/6-CaM structures reveal the mechanism of CaM-mediated inactivation as a pore block by K115 of CaM, which forms a unique cation-π interaction with a cubic cage of four tryptophan indole rings at the ion channel intracellular entrance. The structures of TRPV4-RhoA were captured in different conformations of TRPV4 but showed no difference in RhoA binding, providing limited information about the mechanism of RhoA-mediated TRPV4 inhibition. RhoA, which is known to undergo a posttranslational modification by prenylation, can be anchored to the plasma membrane through its prenylated C-terminus. This interaction may enable TRPV4 to sense alterations in cellular shape and morphology resulting from osmotic shock or mechanical forces [ref. 33]. Like TRPV5–6, TRPC4 undergoes inactivation by CaM [ref. 36]. In the TRPC4-CaM structure, CaM binds to the rib helix of TRPC4. The authors proposed that this binding stabilizes a previously disordered region, which is directly connected to the TRP helix, restricts its mobility and allosterically locks the channel in the closed state.
The structure of TRPM3-Gβγ was solved in the closed state. Understanding the mechanism of TRPM3 inhibition by Gβγ will require solving the open-state structure of TRPM3. Similarly, Gαi3 is known to potentiate TRPC5 [ref. 37] but the TRPC5-Gαi3 structure is solved in the closed state. Likewise, solving the structure of TRPC5-Gαi3 in the open state will be necessary to understand the mechanism of TRPC5 potentiation by Gαi3. Similar to the TRPV4-RhoA interaction, the action of Gαi3 and Gβγ might be dependent on posttranslational modifications of these G proteins by lipidation [ref. 25,ref. 37]. Lipidation may indirectly affect TRP channels by increasing the local concentration of G proteins near membrane and causing clustering of TRP channels.
In particular cellular conditions, interaction of TRP channels with protein binding partners may depend on additional factors. One such factor is the functional state of the binding partner. For example, GTPase RhoA in its inactive, GDP-bound form can inhibit TRPV4 [ref. 38], while activated, GTP-bound Gαi3 can potentiate TRPC5 [ref. 37]. Another important factor is the concentration of second messengers. Ca2+ binding to both CaM and TRPC5 is required for inactivation of TRP channels by CaM and for the TRPC5-Gαi3 complex formation [ref. 29–ref. 32,ref. 36,ref. 37,ref. 39]. PIP2 has been reported as a stabilizing factor for the TRPC5-Gαi3 and TRPM3-Gβγ interactions [ref. 25,ref. 37]. Many interactions of TRP channels with other proteins, like TRPV3 with Ano1 [ref. 40] or TMEM79 [ref. 41], and their regulation by different cofactors remain elusive and will require additional structural studies to understand the corresponding molecular mechanisms.
Conformational changes caused by ligand binding
Numerous TRP channel structures solved in complex with ligands have revealed conformational changes and molecular mechanisms associated with channel regulation and gating. Nevertheless, a substantial number of TRP channel structures in complex with activators or inhibitors have been solved in apo-like states, where protein conformational changes occur only locally, in close proximity to the bound ligands. Typically, such structures are less informative and require functional experiments combined with mutagenesis or MD simulations to gain insights into the mechanism of ligand action. When ligands do induce global conformational changes, they typically resolve one or two new states of the channel conformational ensemble distinct from the closed resting (apo) state: the ligand-bound open state with the conducting pore and/or ligand-bound inhibited or inactivated/desensitized state with the closed pore (Figure 5a and b).
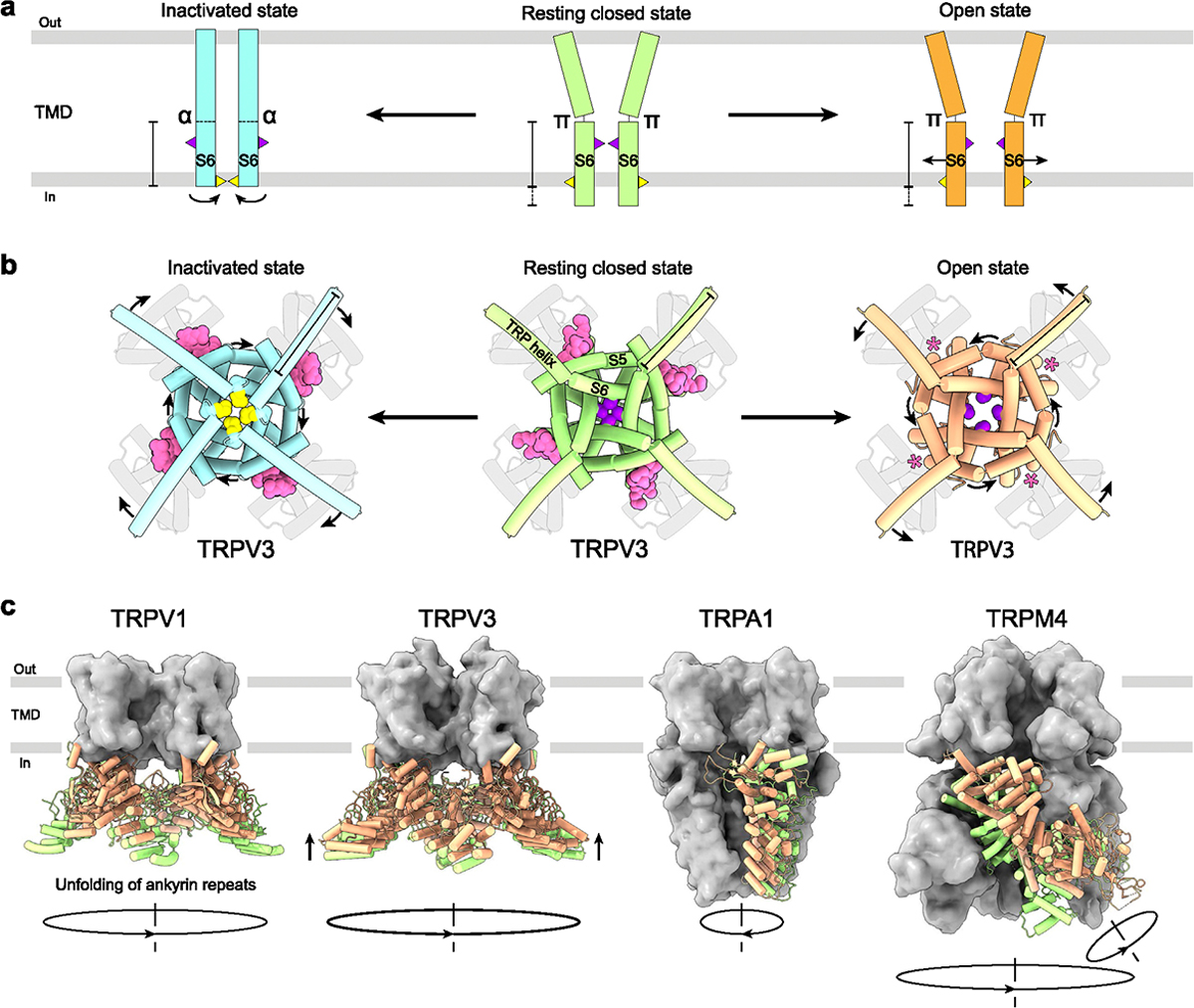
The most pronounced conformational changes associated with TRP channel gating typically happen in the S4eS5 linker, S5, the N-terminal part of the TRP helix, and most importantly S6, which forms the channel gate. When channel pore dilates during opening, S6 moves away from the pore center (Figure 5a). Meanwhile, the N-terminal part of the TRP helix tilts to compensate for the movement of S6 (Figure 5b). This tilt of TRP helix and movement of S6 helix during the closed-to-open transition in vanilloid (V), melatonin (M), and mucolipin (ML) subfamilies results in counterclockwise rotation of the pore-forming helices S5 and S6 around the pore axis when viewed intracellularly (Figure 5b). Importantly, since no open-state structures have been solved for TRPC and TRPP subfamilies, whether these channels follow the same pattern of conformational changes during opening remains unclear.
Inactivation, on the other hand, is often linked to secondary structure changes in S6 itself. Thus, in TRPV3, inactivation is accompanied by a ~100° rotation of the C-terminal portion of S6 converting it from a π-bulge-containing to entirely α-helical (Figure 5a). The rotation in S6 exposes a completely different set of residues to the channel pore, with methionines M677 and isoleucines I674 forming the narrowest part of the human TRPV3 pore in the inactivated and closed/open states, respectively [ref. 9]. In addition, inactivation is accompanied by a two helical turns shortening of S6 and two helical turns elongation of the TRP helix (Figure 5a and b). Similar transitions were observed in TRPV1, TRPV4, and TRPM2 in response to binding of agonists or inhibitors [ref. 19,ref. 34,ref. 42,ref. 43]. Despite this similarity, conformational changes that happen upon inactivation in the majority of other TRP channels do not follow the same pattern [ref. 4,ref. 5,ref. 15,ref. 29,ref. 30,ref. 32,ref. 33,ref. 35,ref. 44–ref. 49]. For example, the closed state of TRPV6 is characterized by α-helical S6, while the open and inactivated states have S6 with the π-bulge in the middle [ref. 50]. In addition, structures reported in intermediate [ref. 19,ref. 24,ref. 51,ref. 52] and non-canonical pentameric [ref. 53] conformations indicate that the 3-state model of TRP channel gating (Figure 5a and b) is an obvious oversimplification, signifying the need of developing more complete and sophisticated gating models.
The global structural rearrangements during activation are unique for each TRP channel subfamily and characterized by distinct architectural features (Figure 2). In TRPV1 channel, pore opening is accompanied by unfolding of the N-terminal ankyrin repeat domains (ARDs). During TRPV3 activation, the intracellular skirt domain moves towards the TMD and rotates 8° clockwise around the fourfold rotational symmetry axis when viewed intracellularly, while four C-termini unwrap and form α-helices [ref. 9]. Opening of the TRPM4 channel involves a 52°-rotation of the N-terminal melastatin homology region (MHR) domains [ref. 12] (Figure 5c). In contrast to TRPV, TRPM, and TRPML channels, activation of TRPA1 is accompanied by movement of S5–S6, TRP helices, and N-terminal domains in the opposite direction, clockwise when viewed intracellularly [ref. 54] (Figure 5c). The molecular mechanisms guiding these global conformational rearrangements, and a possible role of currently unresolved structural domains await future structural studies.
Conclusion
In this review, we provided an update on the current status of TRP channel structural pharmacology and classified 40 discovered binding sites that regulate TRP channel function. We highlight the importance of lipids and ions in modulating channel function and describe commonly observed conformational changes induced by ligand binding. Given modern advances in structural biology, we anticipate discovery of many more new sites, which will aid in drug design for treatment of TRP channel-related diseases.
Supplementary Materials
References
- MV Yelshanskaya, AI Sobolevsky. Ligand-binding sites in vanilloid-subtype TRP channels.. Front Pharmacol, 2022. [PubMed]
- Y Gao, E Cao, D Julius, Y Cheng. TRPV1 structures in nanodiscs reveal mechanisms of ligand and lipid action.. Nature, 2016. [PubMed]
- A Neuberger, M Oda, YA Nikolaev, KD Nadezhdin, EO Gracheva, SN Bagriantsev, AI Sobolevsky. Human TRPV1 structure and inhibition by the analgesic SB-366791.. Nat Commun, 2023. [PubMed]
- MM Diver, Y Cheng, D Julius. Structural insights into TRPM8 inhibition and desensitization.. Science, 2019. [PubMed]
- W Guo, Q Tang, M Wei, Y Kang, J-X Wu, L Chen. Structural mechanism of human TRPC3 and TRPC6 channel regulation by their intracellular calcium-binding sites.. Neuron, 2022. [PubMed]
- Y Bai, X Yu, H Chen, D Horne, R White, X Wu, P Lee, Y Gu, S Ghimire-Rijal, DCH Lin. Structural basis for pharmacological modulation of the TRPC6 channel.. Elife, 2020. [PubMed]
- Y Yang, M Wei, L Chen. Structural identification of riluzole-binding site on human TRPC5.. Cell Discov, 2022. [PubMed]
- Y Yin, SC Le, AL Hsu, MJ Borgnia, H Yang, S-Y Lee. Structural basis of cooling agent and lipid sensing by the cold-activated TRPM8 channel.. Science, 2019. [PubMed]
- KD Nadezhdin, A Neuberger, LS Khosrof, IA Talyzina, J Khau, MV Yelshanskaya, AI Sobolevsky. TRPV3 activation by different agonists accompanied by lipid dissociation from the vanilloid site.. Sci Adv, 2024. [PubMed]
- N Su, W Zhen, H Zhang, L Xu, Y Jin, X Chen, C Zhao, Q Wang, X Wang, S Li. Structural mechanisms of TRPV2 modulation by endogenous and exogenous ligands.. Nat Chem Biol, 2023. [PubMed]
- KD Nadezhdin, L Correia, C Narangoda, DS Patel, A Neuberger, T Gudermann, MG Kurnikova, V Chubanov, AI Sobolevsky. Structural mechanisms of TRPM7 activation and inhibition.. Nat Commun, 2023. [PubMed]
- J Hu, SJ Park, T Walter, IJ Orozco, G O’Dea, X Ye, J Du, W Lü. Physiological temperature drives TRPM4 ligand recognition and gating.. Nature, 2024. [PubMed]
- J Fan, H Ke, J Lei, J Wang, M Tominaga, X Lei. Structural basis of TRPV1 inhibition by SAF312 and cholesterol.. Nat Commun, 2024. [PubMed]
- M Fine, P Schmiege, X Li. Structural basis forPtdInsP2-mediated human TRPML1 regulation.. Nat Commun, 2018. [PubMed]
- Y Yin, CG Park, F Zhang, GF J, S Feng, Y Suo, W Im, SY Lee. Mechanisms of sensory adaptation and inhibition of the cold and menthol receptor TRPM8.. Sci Adv, 2024. [PubMed]
- RA Pumroy, AD Protopopova, TC Fricke, IU Lange, FM Haug, PT Nguyen, PN Gallo, BB Sousa, GJL Bernardes, V Yarov-Yarovoy. Structural insights into TRPV2 activation by small molecules.. Nat Commun, 2022. [PubMed]
- Y Yin, F Zhang, S Feng, KJ Butay, MJ Borgnia, W Im, S-Y Lee. Activation mechanism of the mouse cold-sensing TRPM8 channel by cooling agonist and PIP2.. Science, 2022. [PubMed]
- K Saotome, AK Singh, MV Yelshanskaya, AI Sobolevsky. Crystal structure of the epithelial calcium channel TRPV6.. Nature, 2016. [PubMed]
- WR Arnold, A Mancino, FR Moss, A Frost, D Julius, Y Cheng. Structural basis of TRPV1 modulation by endogenous bioactive lipids.. Nat Struct Mol Biol, 2024. [PubMed]
- T Rohacs. Phosphoinositide regulation of TRP channels: a functional overview in the structural era.. Annu Rev Physiol, 2024. [PubMed]
- AS Klein, A Tannert, M Schaefer. Cholesterol sensitises the transient receptor potential channel TRPV3 to lower temperatures and activator concentrations.. Cell Calcium, 2014. [PubMed]
- M Lakk, GF Hoffmann, A Gorusupudi, E Enyong, A Lin, PS Bernstein, T Toft-Bertelsen, N MacAulay, MH Elliott, D Krizaj. Membrane cholesterol regulates TRPV4 function, cytoskeletal expression, and the cellular response to tension.. J Lipid Res, 2021. [PubMed]
- AK Singh, LL McGoldrick, L Demirkhanyan, M Leslie, E Zakharian, AI Sobolevsky. Structural basis of temperature sensation by the TRP channel TRPV3.. Nat Struct Mol Biol, 2019. [PubMed]
- KD Nadezhdin, A Neuberger, YA Trofimov, NA Krylov, V Sinica, N Kupko, V Vlachova, E Zakharian, RG Efremov, AI Sobolevsky. Structural mechanism of heat-induced opening of a temperature-sensitive TRP channel.. Nat Struct Mol Biol, 2021. [PubMed]
- C Zhao, R MacKinnon. Structural and functional analyses of a GPCR-inhibited ion channel TRPM3.. Neuron, 2023. [PubMed]
- C Zhao, Y Xie, L Xu, F Ye, X Xu, W Yang, F Yang, J Guo. Structures of a mammalian TRPM8 in closed state.. Nat Commun, 2022. [PubMed]
- N Gan, Y Han, W Zeng, Y Wang, J Xue, Y Jiang. Structural mechanism of allosteric activation of TRPML1 by PI(3,5)P2 and rapamycin.. Proc Natl Acad Sci USA, 2022. [PubMed]
- T Rohacs. Phosphoinositide regulation of TRP channels.. Handb Exp Pharmacol, 2014. [PubMed]
- TET Hughes, RA Pumroy, AT Yazici, MA Kasimova, EC Fluck, KW Huynh, A Samanta, SK Molugu, ZH Zhou, V Carnevale. Structural insights on TRPV5 gating by endogenous modulators.. Nat Commun, 2018. [PubMed]
- AK Singh, LL McGoldrick, EC Twomey, AI Sobolevsky. Mechanism of calmodulin inactivation of the calcium-selective TRP channel TRPV6.. Sci Adv, 2018. [PubMed]
- S Dang, MK Van Goor, D Asarnow, Y Wang, D Julius, Y Cheng, J Van Der Wijst. Structural insight into TRPV5 channel function and modulation.. Proc Natl Acad Sci USA, 2019. [PubMed]
- EC Fluck, AT Yazici, T Rohacs, VY Moiseenkova-Bell. Structural basis of TRPV5 regulation by physiological and pathophysiological modulators.. Cell Rep, 2022. [PubMed]
- DH Kwon, F Zhang, BA McCray, S Feng, M Kumar, JM Sullivan, Im W, CJ Sumner, S-Y Lee. TRPV4-Rho GTPase complex structures reveal mechanisms of gating and disease.. Nat Commun, 2023. [PubMed]
- KD Nadezhdin, IA Talyzina, A Parthasarathy, A Neuberger, DX Zhang. Sobolevsky AI: Structure of human TRPV4 in complex with GTPase RhoA.. Nat Commun, 2023. [PubMed]
- J Fan, C Guo, D Liao, H Ke, J Lei, W Xie, Y Tang, M Tominaga, Z Huang, X Lei. Structural pharmacology of TRPV4 antagonists.. Adv Sci, 2024
- D Vinayagam, D Quentin, J Yu-Strzelczyk, O Sitsel, F Merino, M Stabrin, O Hofnagel, M Yu, MW Ledeboer, G Nagel. Structural basis of TRPC4 regulation by calmodulin and pharmacological agents.. Elife, 2020. [PubMed]
- J Won, J Kim, H Jeong, J Kim, S Feng, B Jeong, M Kwak, J Ko, W Im, I So. Molecular architecture of the Gαi-bound TRPC5 ion channel.. Nat Commun, 2023. [PubMed]
- BA McCray, E Diehl, JM Sullivan, WH Aisenberg, NW Zaccor, AR Lau, DJ Rich, B Goretzki, UA Hellmich, TE Lloyd. Neuropathy-causing TRPV4 mutations disrupt TRPV4-RhoA interactions and impair neurite extension.. Nat Commun, 2021. [PubMed]
- FM Bokhovchuk, N Bate, NV Kovalevskaya, BT Goult, C Spronk, GW Vuister. The structural basis of calcium-dependent inactivation of the transient receptor potential vanilloid 5 channel.. Biochemistry, 2018. [PubMed]
- Y Yamanoi, J Lei, Y Takayama, S Hosogi, Y Marunaka, M Tominaga. TRPV3-ANO1 interaction positively regulates wound healing in keratinocytes.. Commun Biol, 2023. [PubMed]
- J Lei, RU Yoshimoto, T Matsui, M Amagai, MA Kido, M Tominaga. Involvement of skin TRPV3 in temperature detection regulated by TMEM79 in mice.. Nat Commun, 2023. [PubMed]
- KD Nadezhdin, A Neuberger, YA Nikolaev, LA Murphy, EO Gracheva, SN Bagriantsev, AI Sobolevsky. Extracellular cap domain is an essential component of the TRPV1 gating mechanism.. Nat Commun, 2021
- Y Yin, M Wu, AL Hsu, WF Borschel, MJ Borgnia, GC Lander, S-Y Lee. Visualizing structural transitions of ligand-dependent gating of the TRPM2 channel.. Nat Commun, 2019
- X Zhou, M Li, D Su, Q Jia, H Li, X Li, J Yang. Cryo-EM structures of the human endolysosomal TRPML3 channel in three distinct states.. Nat Struct Mol Biol, 2017. [PubMed]
- W Zhen, Z Zhao, S Chang, X Chen, Y Wan, F Yang. Structural basis of ligand activation and inhibition in a mammalian TRPV4 ion channel.. Cell Discov, 2023. [PubMed]
- TE Hughes, JS Del Rosario, A Kapoor, AT Yazici, Y Yudin, EC Fluck, M Filizola, T Rohacs, VY Moiseenkova-Bell. Structure-based characterization of novel TRPV5 inhibitors.. Elife, 2019. [PubMed]
- A Neuberger, KD Nadezhdin, AI Sobolevsky. Structural mechanisms of TRPV6 inhibition by ruthenium red and econazole.. Nat Commun, 2021. [PubMed]
- A Neuberger, YA Trofimov, MV Yelshanskaya, KD Nadezhdin, NA Krylov, RG Efremov, AI Sobolevsky. Structural mechanism of human oncochannel TRPV6 inhibition by the natural phytoestrogen genistein.. Nat Commun, 2023. [PubMed]
- A Neuberger, YA Trofimov, MV Yelshanskaya, J Khau, KD Nadezhdin, LS Khosrof, NA Krylov, RG Efremov, AI Sobolevsky. Molecular pathway and structural mechanism of human oncochannel TRPV6 inhibition by the phytocannabinoid tetrahydrocannabivarin.. Nat Commun, 2023. [PubMed]
- MV Yelshanskaya, KD Nadezhdin, MG Kurnikova, AI Sobolevsky. Structure and function of the calcium-selective TRP channel TRPV6.. J Physiol, 2021. [PubMed]
- DH Kwon, F Zhang, JG Fedor, Y Suo, S-Y Lee. Vanilloid-dependent TRPV1 opening trajectory from cryoEM ensemble analysis.. Nat Commun, 2022. [PubMed]
- DH Kwon, F Zhang, Y Suo, J Bouvette, MJ Borgnia, S-Y Lee. Heat-dependent opening of TRPV1 in the presence of capsaicin.. Nat Struct Mol Biol, 2021. [PubMed]
- S Lansky, JM Betancourt, J Zhang, Y Jiang, ED Kim, N Paknejad, CM Nimigean, P Yuan, S Scheuring. A pentameric TRPV3 channel with a dilated pore.. Nature, 2023. [PubMed]
- J Zhao, JV Lin King, CE Paulsen, Y Cheng, D Julius. Irritant-evoked activation and calcium modulation of the TRPA1 receptor.. Nature, 2020. [PubMed]
