Chemical Diversity of Three Endophytic Talaromyces Strains and Their Potential for Biocontrol against the Cocoa Pathogenic Fungus
Abstract
Biological control using beneficial endophytic fungi is a sustainable alternative to agrochemicals for managing plant diseases. This study assessed three endophytic Talaromyces strains, isolated from different Brazilian host plants, for antagonistic activity toward Moniliophthora perniciosa, the causative agent of cocoa witches’ broom disease. All strains inhibited the pathogen’s mycelial growth in vitro, with Talaromyces pinophilus J6 exhibiting the strongest effect. Discriminant metabolites of the T. pinophilus J6 were unveiled by liquid chromatography‐tandem high‐resolution mass spectrometry‐based metabolomics combined with principal component analysis and a dereplication approach. These results underscore the promise of endophytic Talaromyces spp. as novel biocontrol agents and will contribute to ongoing research on sustainable agriculture. Moreover, chemical comprehension of endophytic fungi from the same genus that come from different host plants opens new insights into chemical ecology implicated in the endophyte‐plant interactions.
Article type: Research Article
Keywords: antifungal assay, biocontrol, metabolomics, witches’ broom disease of cacao
Affiliations: Department of Organic Chemistry, Institute of Chemistry Universidade Federal da Bahia Salvador Brazil; Departament of Biological Sciences Universidade Estadual de Santa Cruz Ilhéus Brazil; Institute For Research in Pharmaceuticals and Medications Universidade Federal da Paraíba João Pessoa Brazil
License: © 2026 The Author(s). Chemistry & Biodiversity published by Wiley‐VHCA AG. CC BY 4.0 This is an open access article under the terms of the http://creativecommons.org/licenses/by/4.0/ License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Article links: DOI: 10.1002/cbdv.202503152 | PubMed: 41518658 | PMC: PMC12790416
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (1.7 MB)
Introduction
Theobroma cacao L. (Sterculiaceae) is an economically important crop in several tropical countries, and its commercially valuable beans constitute about 10% of the cacao fruit’s fresh weight [ref. 1]. Brazil is the seventh largest cocoa producer in the world and also the seventh largest exporter of the product and its derivatives. Between January and May 2022, 14 038 tons of chocolate, 20 232 tons of by‐products, and 273 tons of cocoa beans were exported by Brazil [ref. 2]. Fungal diseases are significant threats to economically important crops, leading to substantial losses due to their rapid spread and ability to adapt to different environmental conditions [ref. 3]. Moniliophthora perniciosa is a Basidiomycete that causes the witches’ broom disease of cacao, being the cause of the rapid decline in cacao production in Brazil [ref. 4]. In 1989, witches’ broom disease became established in Bahia, a state in northeastern Brazil, and, together with other factors, it has contributed to the worsening impoverishment of the region, which continues to struggle to recover from its effects [ref. 5].
The use of chemical fungicides for the management of witches’ broom disease has disadvantages such as the development of resistance by the plant pathogens, environmental contamination, and human health disturbances [ref. 6]. Alternative methods of controlling fungal crop diseases have been widely developed, including the use of disease‐resistant cultivars, adequate water and soil management, fertilization, crop rotations, and biological control agents, aiming to maintain or increase agricultural production with a reduced application of chemical agents [ref. 7]. Among the promising alternative control methods is the use of antagonistic microorganisms, such as beneficial endophytic fungi, capable of protecting the plant from the action of pathogens [ref. 8].
Endophytic microorganisms live inside plant tissues for at least part of their life cycles and are generally asymptomatic [ref. 9]. Within complex relationships, endophytic strains receive nutrients and protection, while plants have advantages, such as greater resistance in environments with intense stress caused by biotic (such as insects, herbivores, nematodes, and phytopathogenic microorganisms) or abiotic factors (such as pH, temperature, drought, and saline stresses) [ref. 10, ref. 11]. Endophytic fungi inhabit a similar ecological niche to that occupied by phytopathogens, thus being able to protect their hosts and control pathogens through competition, production of antagonistic substances, direct parasitism, or even inducing resistance or tolerance [ref. 12]. Consequently, endophytes have attracted increasing attention as biological control agents and as inducers of plant defense responses [ref. 13, ref. 14], which are key points for sustainable agriculture intensification. Their ability to synthesize a wide variety of specialized metabolites, such as alkaloids, terpenoids, polyketides, and peptides, underlies their potent antifungal, antibacterial, and insecticidal activities. Moreover, endophytic fungi enhance plant immunity through multiple mechanisms, including competitive exclusion, antimicrobial compound production, activation of defense pathways, phosphate solubilization, siderophore production, and modulation of phytohormones [ref. 15].
The endophyte microbiota composition significantly differs between distinct geographic areas [ref. 16]. Specific environmental characteristics act as “ecological filters” in the selection of the endophytic strains that are better adapted to local conditions. Additionally, some endophytes show host specificity [ref. 17] or are compatible with only specific host genotypes [ref. 18]. Within this context, the diversity of biomes and endemic plants found in Brazil represents a potential source of new beneficial microbial resources. Conducting studies to explore the biocontrol potential of endophyte biodiversity is a promising and powerful alternative to agrochemicals, offering control over a variety of plant fungal diseases. Fungi belonging to the Talaromyces genus have emerged as sustainable and effective agents for plant disease management. These organisms produce a broad range of bioactive metabolites exhibiting strong antifungal, antibacterial, and nematicidal activities, and secrete hydrolytic enzymes (such as chitinases, glucanases, and proteases) capable of degrading pathogen cell walls [ref. 19]. Moreover, their metabolites can induce systemic resistance and promote plant growth, enhancing overall crop vigor without reliance on synthetic stimulants. For instance, evidence from agricultural systems has demonstrated the efficacy of Talaromyces purpurogenus Q2 in enhancing the resistance of bitter gourd to fungal pathogens by activating the lignin biosynthesis pathway and simultaneously promoting plant growth [ref. 20].
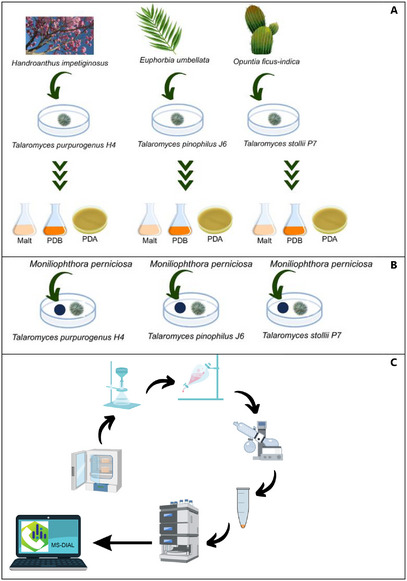
In this study, we investigated the antagonistic potential of three endophytic Talaromyces strains, each isolated from a distinct Brazilian host plant, against M. perniciosa, the causal agent of witches’ broom disease in cocoa (Scheme 1). Firstly, we qualitatively evaluated the interaction between the endophytes Talaromyces spp. and the pathogen by in vitro antagonism bioassays. To elucidate the chemical basis underlying their activity, we employed liquid chromatography‐high‐resolution mass spectrometry (LC‐HRMS)‐based metabolomics combined with principal component analysis and a dereplication approach to profile and compare their specialized metabolites. By linking chemical diversity to biocontrol performance, our results not only identify promising fungal candidates for sustainable cocoa disease management but also advances the understanding of the chemical ecology in endophyte–plant–pathogen interactions.
Results and Discussion
Endophytic Strains Identification
Three endophytic fungal strains—designated H4, J6, and P7 ‐ were isolated from the aerial parts of Handroanthus impetiginosus, Euphorbia umbellata, and Opuntia ficus‐indica, respectively. All colonies exhibited a consistent phenotypic progression, characterized by an initial yellow pigmentation that gradually shifted to green within seven days (Figure S1). Such reproducibility across isolates may reflect underlying metabolic or developmental similarities worthy of future investigation. Notably, this pattern was observed in the isolates H4, J6, and P7, which were obtained from taxonomically and ecologically distinct host plants, prompting further interest in molecular identification, comparative metabolomic profiling, and evaluation of their biological potential.
Genetic characterization of the endophytes H4, J6, and P7 elucidated the identity of all these strains as belonging to the Talaromyces genus (see phylogenetic trees in Figure S2A–C). The endophyte H4 clustered with sequences of Talaromyces spp. (83% similarity to reference sequences from this genus), while the strains J6 and P7 shared 99% and 100% identity, respectively, with the type strains Talaromyces pinophilus CBS 631.66 (JX091381) and Talaromyces stollii CBS 408.93 (JX315633). Talaromyces species have been frequently reported as endophytes in various plant hosts, reflecting their ecological versatility and adaptability to distinct environmental niches [ref. 21]. The observation that isolates H4, J6, and P7 originated from taxonomically unrelated host species raises intriguing questions regarding host specificity, ecological plasticity, and the conservation of biosynthetic pathways that may facilitate adaptation to different plant environments within the genus Talaromyces. Comparative metabolomic studies could provide deeper insights into these ecological and evolutionary dynamics.
The scientific interest in metabolites of fungi belonging to the Talaromyces genus (Ascomycota) has shown a constant increment since they are regarded as important natural product resources for producing various small molecules with diverse chemical structures and biological activities [ref. 22]. In our study, the three evaluated host plants naturally occur in distinct environmental conditions, which likely influence their specialized metabolism and the composition of their associated endophytic microbiota. Based on this, we hypothesize that the geographic origin of each host plant may also impact the metabolic profiles of the isolated endophytes (from the same genus), potentially shaping their biosynthetic capabilities and biological activities.
Inhibition of the Crop Pathogenic Fungus by the Evaluated Endophytic Fungi
The antagonistic potentials of Talaromyces sp. H4, T. pinophilus J6, and T. stollii P7 were separately evaluated against M. perniciosa, the causal agent of witches’ broom disease in cacao, using co‐culture assays for 7 and 14 days (Figure S3).
When compared with the control assay (M. perniciosa, Figure 1A), all Talaromyces spp. displayed inhibitory activity against M. perniciosa mycelial growth (Figure 1B–D), as the pathogen exhibited reduced growth in co‐culture with the endophytes relative to cultivation alone. T. pinophilus J6 (Figure 1C) displayed the strongest inhibitory effect, with M. perniciosa showing practically no mycelial development.
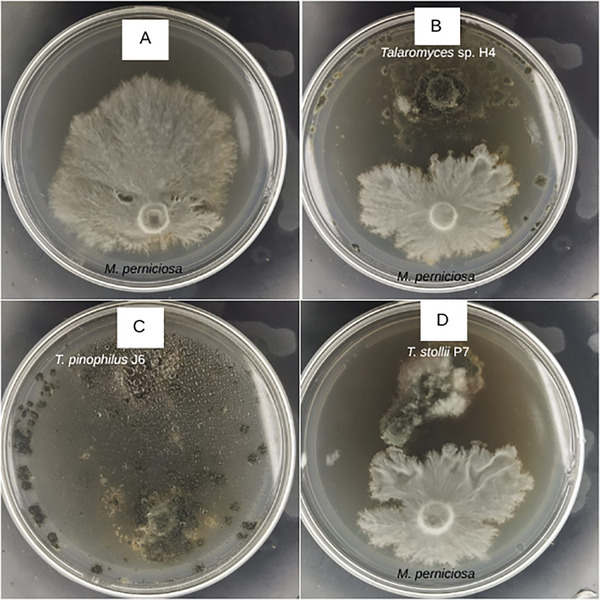
Inhibition outcomes were visually evaluated (for qualitative measurement) and categorized according to specialized literature as follows: (i) inhibition at a distance, when the opposing mycelia ceased growth before making contact; (ii) partial replacement, when the endophyte overgrew part of the pathogen colony without reaching its opposite edge; and (iii) complete replacement, when the endophyte entirely overgrew the pathogen colony. Accordingly, the interactions between M. perniciosa and Talaromyces sp. H4 (Figure 1B), T. pinophilus J6 (Figure 1C), and T. stollii P7 (Figure 1D) were classified as (i), (iii), and (i), respectively.
Microbial biological control agents have emerged as a viable alternative to chemical fungicides, playing a pivotal role in modern sustainable agriculture. Among these, endophytic fungi have garnered attention for their capacity to produce bioactive metabolites that suppress phytopathogens, promote plant growth, and enhance stress tolerance. Our preliminary inhibition findings reinforce the importance of chemically characterizing the endophytic Talaromyces spp., as such analyses could elucidate the distinct bioactivities observed against M. perniciosa.
Untargeted Metabolomics Analysis
Chemical investigations aimed at differentiating the metabolic profiles of the evaluated endophytes began with culturing the three Talaromyces strains on solid potato dextrose agar (PDA), liquid potato dextrose broth (PDB), and malt broth media. Liquid media were incubated under agitation and static conditions.
Base peak chromatograms (BPCs) of the ethyl acetate extracts from the controls (culture media without fungi, Figures S4) and the Talaromyces strains J6, H4, and P7 (Figures S5–S7, respectively) provide a visual overview of the chemical complexity of the samples and support the subsequent feature extraction and multivariate analyses. The BPCs exhibited stable baselines and well‐resolved peaks, confirming the high quality of the LC–HRMS acquisition. The positive ionization mode yielded superior signal intensity and a greater number of detected features; therefore, it was selected for the metabolomic analyses. The comparative analysis of BPCs revealed that cultures grown in Malt, PDA, and PDB media produced distinct chemical profiles across all evaluated fungal strains. Notably, agitation conditions induced clear differences in the chemical patterns of cultures in Malt medium compared with static conditions. In contrast, for cultures grown in PDB medium, the chemical profiles were relatively similar under both static and agitated conditions.
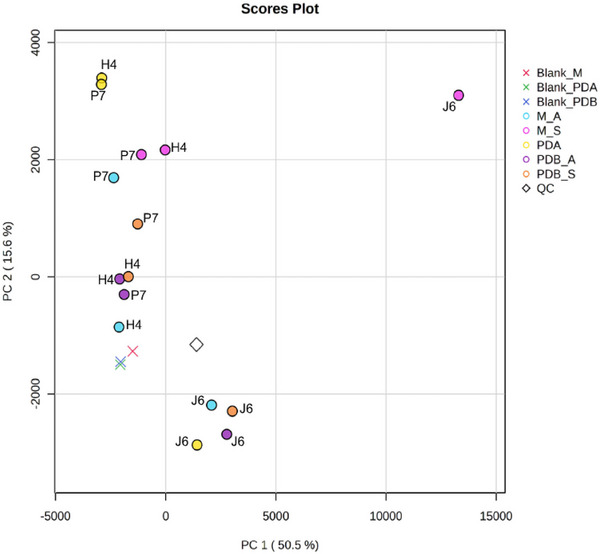
Based on retention time and exact mass (m/z), the spot ID for each feature was analyzed using MS‐DIAL. Subsequently, the metabolic profiles of the ethyl acetate extracts from all Talaromyces spp. were explored through principal component analysis (PCA, Figure 2) and Hierarchical Clustering Analysis shown as a dendrogram (Figure S8). The PCA showed that 66.1% of the total variation in the data was represented by the first two principal components. Clustering analysis based on metabolomic similarity revealed a clear separation of T. pinophilus J6 from Talaromyces sp. H4 and T. stollii P7, regardless of the culture conditions. Thus, with negative scores for PC2, T. pinophilus J6 grown in PDA, PDB (under agitation or static conditions), and Malt (under agitation) were joined. Interestingly, the extract from T. pinophilus J6 grown in Malt medium incubated under static conditions was placed separately from all samples. In other words, among all assayed Talaromyces spp., the T. pinophilus J6 was chemically distinct (confirmed by Hierarchical Clustering Dendrogram), which may be correlated with its superior inhibition potential against M. perniciosa (Figure 1C).
Chemical Characterization of Talaromyces Strains
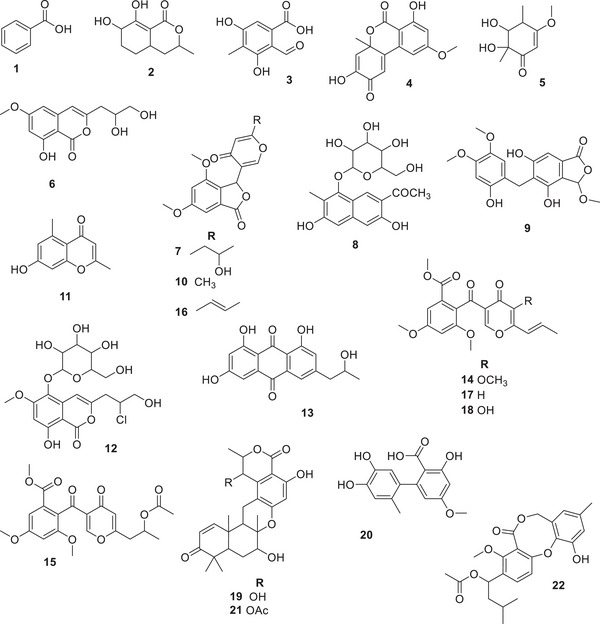
At the sequence, the three Talaromyces spp. were chemically characterized according to the tandem MS (MS/MS) fragmentation patterns, molecular formula, and in‐house database search, which includes over 700 specialized metabolites previously reported to Talaromyces sp. Twenty‐two metabolites were putatively identified (Table 1), and their chemical structures are depicted in Figure 3.
TABLE 1: Specialized metabolites putatively annotated in the ethyl acetate extracts of endophytic Talaromyces strains H4, J6, and P7, based on ultra‐high‐performance liquid chromatography‐electrospray ionization‐Quadrupole time‐of‐flight‐ mass spectrometry (UHPLC‐ESI‐QTOF‐MS) analysis and dereplication using a curated in‐house database. The table includes retention times (RT), observed m/z values, calculated errors (ppm), compound names, molecular formulas, and the Talaromyces strains in which each metabolite was previously reported.
| No. | Retention time (min) | m/z, Adduct | m/z error (ppm) | Putative identification (Molecular formula) | Reported Talaromyces source (Reference) |
|---|---|---|---|---|---|
| 1 | 17.52 | 140.0698, [M+NH4]+ | −5.72 | benzoic acid [a] | T. pinophilus AF‐02 |
| (C7H6O2) | [ref. 23] | ||||
| 2 | 19.67 | 216.1217, [M+NH4]+ | −6.02 | (3S,4aR,7S)‐7,8‐dihydroxy‐3‐ methyl‐3,4,10,5,6,7‐hexahydro‐1H‐isochromen‐1‐one [b] | Talaromyces sp. |
| (C10H14O4) | [ref. 24] | ||||
| 3 | 25.02 | 179.0329, [M+H‐H2O]+ | −5.59 | 2‐formyl‐3,5‐dihydroxy‐4‐methylbenzoic acid [c] | Talaromyces sp. T1BF |
| (C9H8O5) | [ref. 25] | ||||
| 4 | 27.19 | 311.0514, [M+Na]+ | −3.86 | dehydroaltenusin [c] | T. flavus |
| (C15H12O6) | [ref. 26] | ||||
| 5 | 27.85 | 209.0794, [M+Na]+ | 4.79 | talaketide [b] | Talaromyces sp. CY‐3 |
| (C9H14O4) | [ref. 27] | ||||
| 6 | 28.31 | 289.0695, [M+Na]+ | 4.15 | diaportinol [c] | T. minnesotensis BTBU20220184 |
| (C13H14O6) | [ref. 28] | ||||
| 7 | 32.85 | 347.1110, [M+H]+ | −4.33 | hydroxydihydrovermistatin [d] | T. thailandiasis |
| (C18H18O7) | [ref. 29] | ||||
| 8 | 33.26 | 377.1213, [M+H‐H2O]+ | −4.77 | talaminaphtholglycoside [d] | T. minnesotensis BTBU20220184 |
| (C19H22O9) | [ref. 28] | ||||
| 9 | 33.57 | 345.0952, [M+H‐H2O]+ | −3.48 | talaromycolide A [d] | T. pinophilus AF‐02. |
| (C18H18O8) | [ref. 23] | ||||
| 10 | 34.21 | 303.0849, [M+H]+ | −4.62 | penisimplicissin [c] | T. thailandiasis |
| (C16H14O6) | [ref. 29] | ||||
| 11 | 34.23 | 191.0692, [M+H]+ | −5.76 | 7‐hydroxy‐2,5‐dimethylchromon [c] | T. flavus |
| (C11H10O3) | [ref. 30] | ||||
| 12 | 35.94 | 485.0818, [M+Na]+ | −0.62 | talamiisocoumaringlycoside B [b] | T. minnesotensis BTBU20220184 |
| (C19H23ClO11) | [ref. 28] | ||||
| 13 | 36.27 | 337.0669, [M+Na]+ | −3.26 | isorhodoptilometrin [c] | Talaromyces sp. |
| (C17H14O6) | [ref. 31] | ||||
| 14 | 37.54 | 411.1027, [M+Na]+ | −5.59 | 3‐O‐methylfunicone [c] | T. pinophilus |
| (C20H20O8) | [ref. 32] | ||||
| 15 | 38.54 | 419.1314, [M+H]+ | −5.49 | actofunicone [b] | T. flavus FKI‐0076 |
| (C21H22O9) | [ref. 25] | ||||
| 16 | 40.82 | 351.0824, [M+Na]+ | −4.27 | vermistatin [c] | T. thailandiasis |
| (C18H16O6) | [ref. 29] | ||||
| 17 | 43.02 | 359.1112, [M+H]+ | −3.62 | deoxyfunicone [d] | T. flavus FKI‐0076 |
| (C19H18O7) | [ref. 25] | ||||
| 18 | 43.36 | 357.0960, [M+H‐H2O]+ | −2.53 | funicone [c] | T. flaus IFM52668. |
| (C19H18O8) | [ref. 25] | ||||
| 19 | 45.15 | 465.1867, [M+Na]+ | −3.66 | chrodrimanin A [a] | T. thailandensis PSUSPSF059 |
| (C25H30O7) | [ref. 33] | ||||
| 20 | 45.52 | 313.0671, [M+Na]+ | −3.84 | biphenyl derivative [b] | T. flavus |
| (C15H14O6) | [ref. 30] | ||||
| 21 | 46.41 | 507.1968, [M+Na]+ | −4.14 | chrodrimanin B [a] | T. thailandensis PSUSPSF059 |
| (C27H32O8) | [ref. 33] | ||||
| 22 | 52.11 | 437.1553, [M+Na]+ | −4.12 | vermixocin B [d] | Talaromyces sp. LF458 |
| (C23H26O7) | [ref. 34] |
[a] mainly detected in both Talaromyces sp. H4 and T. stollii P7 cultures.
[b] detected in all Talaromyces spp. cultures.
[c] mainly detected T. pinophilus J6 cultures.
[d] exclusively detected T. pinophilus J6 cultures.
Besides the 22 annotated metabolites, the MS‐DIAL analysis detected an additional 22 features (after removing blank features) that could not be confidently annotated using our in‐house database. The features that were detected but not annotated (Table S1) are potential candidates for future biochemical and biocontrol studies.
Certain compounds were predominantly detected in both Talaromyces sp. H4 and T. stollii P7 cultures (designated with a superscript letter ‘a’ in Table 1), while others were mainly or exclusively associated with T. pinophilus J6 cultures (designated with superscript letters ‘c’ and ‘d’ in Table 1). Figure S9 shows the relative distribution of each 22 annotated metabolites across the evaluated Talaromyces spp. Some specialized metabolites are conserved across the Talaromyces genus (designated with a superscript letter ‘b’ in Table 1), indicating shared biosynthetic pathways.
The use of an in‐house library for dereplication significantly enhanced the confidence in metabolite identification. Benzoic acid (1, m/z 140.0698), chrodrimanin A and B (19 and 21, m/z 465.1867 and 507.1968, respectively) were mainly detected in both Talaromyces sp. H4 and T. stollii P7 cultures. On the other hand, some metabolites were mainly detected in T. pinophilus J6 cultures: 2‐formyl‐3,5‐dihydroxy‐4‐methylbenzoic acid (3, m/z 179.0329), dehydroaltenusin (4, m/z 311.0514), diaportinol (6, m/z 289.0695), penisimplicissin (10, m/z 303.0849), 7‐hydroxy‐2,5‐dimethylchromon (11, m/z 191.0692), isorhodoptilometrin (13, m/z 337.0669), 3‐O‐methylfunicone (14, m/z 411.1027), vermistatin (16, m/z 351.0824), and funcione (18, m/z 357.0960). Other ones have exclusively been detected in the T. pinophilus J6 cultures: hydroxydihydrovermistatin (7, m/z 347.1110), talaminaphtholglycoside (8, m/z 377.1213), talaromycolide A (9, m/z 345.0964), deoxyfunicone (17, m/z 359.1112), and vermixocin B (22, m/z 437.1553).
The dereplication approach employed by us revealed a diverse array of specialized metabolites (Table 1). The presence of compounds with varied chemical scaffolds highlights their potential not only as biocontrol agents against plant pathogens but also as valuable resources for pharmaceutical and industrial biotechnological applications. Additionally, the chemical characterization of the Talaromyces strains evaluated herein showed some degree of specificity, which corroborates the chemical distribution pattern seen in PCA and HCA analysis. Talaromyces strains isolated from distinct host plants produce interesting repertoires of specialized metabolites. These chemical profiles appear to play a critical role in determining their antagonistic potential against M. perniciosa, the causal agent of witches’ broom disease. In particular, T. pinophilus J6 exhibited enhanced biocontrol activity, which correlates with the presence of bioactive compounds not detected in the other Talaromyces isolates.
The superior inhibitory activity displayed by T. pinophilus J6 against M. perniciosa (in comparison with the other two Talaromyces strains, Figure 1) could be due to the presence of chemicals with known antifungal activities. Funicone‐like compounds (14, 15, 17, and 18) and phthalide derivatives, such as talaromycolide A (9) and vermistatin (16), have been reported as antimicrobial leads [ref. 23, ref. 25, ref. 29] and were mainly or exclusively detected in T. pinophilus J6 cultures.
Funicone‐like compounds have frequently been isolated from the Talaromyces genus and display a range of biological activities, such as fungicides, antivirals, and antitumorals [ref. 35]. Funicones and structurally related compounds represent a homogeneous group of fungal polyketides that were initially characterized as determinants of the antagonistic abilities by the producers against other microorganisms, but were later found to possess remarkable biological properties that have promoted their consideration as drug prospects [ref. 36]. Previously, 3‐O‐methylfunicone was tested on broad bean (Vicia faba) leaves against the pea aphid Acyrthosiphon pisum and displayed an interesting mortality [ref. 32]. It completely inhibited the growth of Rhizoctonia solani and other species of phytopathogenic fungi at a concentration of 0.1 mg/mL [ref. 37]. Funicone‐like compounds have been found exclusively or almost exclusively in Talaromyces strains and can be considered as candidates for the assessment of phylogenetic relations [ref. 38].
Conclusions
In summary, our results implied the potential of the endophytic Talaromyces spp. for further exploring their molecular diversity of natural products under more diverse culture conditions and discovering new bioactive compounds. Moreover, the chemical comparison of Talaromyces strains isolated from distinct host plants enriches our understanding of endophyte–host interactions and their influence on specialized metabolism. We demonstrated that endophytes from the same genus, isolated from distinct host plants, exhibited different chemical metabolism profiles, which were implicated in inhibitory activity against the phytopathogenic fungus at varying levels. Altogether, this study contributes valuable insights into the chemical ecology of endophytic fungi and lays the groundwork for future exploration of their bioactive compounds in agricultural and pharmaceutical contexts.
Experimental
Endophytic Fungi Isolation and Identification
The endophytic fungi coded as H4, J6, and P7 were isolated, respectively, from aerial parts of Handroanthus impetiginosus (Mart. ex DC.) Mattos, Euphorbia umbellata (Pax) Bruyns, and Opuntia ficus‐indica L. Mill, as previously reported by our research group [ref. 39, ref. 40].
Host plants were collected from their characteristic Brazilian biomes: H. impetiginosus from the Cerrado biome in Minas Gerais State (S 21° 18′ 49.15″, W 45° 57′ 28.53″), and E. umbellata from the Atlantic Forest‐Cerrado transition area in Bahia State (S 14°54’46.81’, W 40°48’02.33’). In contrast, O. ficus‐indica was collected from the Caatinga biome, also in Bahia State (S 12° 59′ 21.06 W 38° 30′ 56.30). All samples were collected during the dry season. Only healthy, asymptomatic leaves were selected from plants in the vegetative phase, without visible flowers or fruits. E. umbellata and O. ficus‐indica were identified by Dr. Nádia Roque (Institute of Biology, Federal University of Bahia). A voucher specimen for each species was deposited in the Alexandre Leal Costa Herbarium (Federal University of Bahia) under the code ALBC 136527. H. impetiginosus was identified by Dr. Lúcia G. Lohmann (Department of Botany, Institute of Biosciences, University of São Paulo), and its voucher specimen was deposited in the Herbarium of the Federal University of Alfenas under the code 2535. The study was registered in the Brazilian System for the Management of Genetic Heritage and Associated Traditional Knowledge (SisGen) under the codes AE18457, AEFEF8D, and A30225F.
All strains were submitted to DNA sequencing and phylogenetic analysis for their identification. For this, the 7‐days cultures of the endophytic fungi were submitted to genomic DNA extraction and purification according to a previously reported protocol [ref. 41] by using glass microspheres (425–600 µm diameter, Sigma‐Aldrich).
ITS1–5.8S–ITS2 region from H4 was amplified using the primer pair ITS1–ITS4 [ref. 42]. The beta‐tubulin (benA) gene sequences were generated from J6 and P7 strains—Bt2a and Bt2b were the primers used. After purification (GFX PCR DNA and Gel Band Purification Kit; GE Healthcare, Little Chalfont, UK), the DNA fragments sequencing from each strain were carried out in an ABI 3500XL Series (Applied Biosystems, Foster City, USA) automatic sequencing. The DNA sequences were analyzed by Clustal X [ref. 43] with the MEGA 11.0 program [ref. 44]. Phylogenetic trees (Figure S1) were constructed using the Kimura model [ref. 45] and Neighbor‐Joining [ref. 46] parameter modeling (1000 bootstrap replications). The sequences data obtained from the endophytic fungi H4, J6, and P7 were registered at the GenBank database (https://www.ncbi.nlm.nih.gov/genbank/) with accession numbers MK749843, PQ963936, and PV755238, respectively.
Fungal Antagonism Assays
The endophytic fungal isolates H4, J6, and P7 were evaluated for antagonistic activity against the phytopathogen M. perniciosa. Dual‐culture assays were conducted in 9 cm Petri dishes containing PDA medium. Agar plugs (6 mm in diameter) from actively growing cultures (7 days old) of each endophytic strain and the pathogen were placed on opposite sides of the plate at the same time. The plates were incubated at 28°C under light conditions for 14 days, in triplicate. The inhibitions were evaluated after 7 and 14 days of the beginning of cultures.
The interaction patterns were categorized following established criteria [ref. 47]: (i) inhibition at a distance: the opposing mycelia stopped growing before contact; (ii) partial replacement: the endophyte overgrew part of the pathogen colony without reaching the opposite edge; (iii) complete replacement: the endophyte fully overgrew the pathogen colony.
Endophytic Fungi Cultures and Metabolites Extraction
The three endophytic fungi were grown in different culture media and conditions: (i) Petri dishes containing 20 mL of PDA, (ii) 250 mL‐Erlenmeyer flasks containing 100 mL of PDB, and (iii) 250 mL‐Erlenmeyer flasks containing 100 mL of malt extract medium (2.0% malt extract, 2.0% glucose, and 0.1% peptone). The liquid cultures were established with 10 plugs (6 mm diameter) of fungus, incubated under agitation (120 rpm), or under static conditions. All cultures were carried out in triplicate and incubated simultaneously for 15 days at 28°C. The crude extracts were obtained by extraction with ethyl acetate. The ethyl acetate extracts were concentrated using a rotary evaporator under reduced pressure at a temperature of 40°C to avoid thermal degradation of sensitive metabolites. After concentration, the residues were dried under a gentle stream of nitrogen to remove any remaining solvent before LC‐HRMS analysis. The blanks were achieved under the same conditions and consisted of the culture media without fungi.
Ultra‐HPLC‐HRMS Fungal Extracts Analysis
All crude extracts from Talaromyces cultures grown on Malt (shaken and static), PDB (shaken and static), and PDA were analyzed by ultra‐HPLC‐HRMS (UHPLC‐HRMS).
One microliter of each sample (0.5 mg/mL) was injected separately into the LC‐40D X3 (Shimadzu), equipped with the CBM‐40, DGU‐40S, LC 40D X3, SIL‐40C X3, and CTD‐40S modules, and coupled to an LCMS‐9050 mass spectrometer (Shimadzu) with an electrospray ionization (ESI) source and a Quadrupole Time‐of‐Flight (Q‐TOF) analyzer. The LC experiments were carried out using a C18 column (Kromasil‐250 mm × 4.6 mm x 5.0 µm). A linear gradient elution system was from 5% to 100% methanol (Supelco HPLC grade) in water for 60 min, under a flow rate of 0.6 mL/min and column temperature set to 40°C. The mass spectrometer’s analysis parameters included a capillary voltage of 4.0 kV, a nebulizer gas (N2) flow rate of 3.0 L/min, a drying gas flow rate of 10 mL/min, and an interface temperature of 300°C.
The LC‐HRMS analyses were initially conducted in both positive and negative ionization modes. The positive ion mode provided higher signal intensity, greater feature richness, and better reproducibility for the ethyl acetate extracts of the Talaromyces strains. Consequently, we selected the positive ion mode (m/z 100–1200) for the metabolomic analyses presented in this study.
Fragmentation was performed by data‐independent acquisition (DIA), with an event time of 0.034 s, a loop time of 1.188 s, and a Q1 resolution of 34.4. Argon was the gas used in the collision cell, which operated at a collision energy of 30‐50. Product ions of m/z 50‐1200 were detected. The samples were randomly analyzed. The results from the sample replicates, quality control (QC) samples, and a blank analyzed at the beginning, middle, and end of the sequence were evaluated to ensure analytical consistency and instrument performance.
Mass Data Processing, Dereplication, and Metabolomics
To generate the metabolite annotation, the raw data from UHPLC‐MS obtained in positive ion mode were directly uploaded and deconvoluted using MS‐DIAL software (ver. 4.9).
MS‐DIAL is open‐source software that can be easily deployed in any laboratory environment with little or no programming required, primarily for analyzing and processing metabolomics data [ref. 48].
Data collection is performed with 0.01 Da MS1 tolerance and 0.05 Da MS2. Peak detection was applied with 5E5 amplitude for minimum peak height (threshold) and 0.1 Da mass slice width. Deconvolution parameters were set as follows: sigma window value of 0.5 and MS/MS abundance cutoff of 10 amplitudes. Alignment parameters setting included as reference file the QC sample (the most complex file), a retention time tolerance of 0.05 min. Finally, features based on blank information were removed.
All BPC data were inspected to verify chromatographic consistency and reproducibility across replicates before proceeding to feature extraction. After the processing and features generation, a dereplication methodology is used on an in‐house database.
Metabolite annotation was performed using a curated in‐house database comprising chemicals previously reported for the genus Talaromyces. This database includes molecular formulae, accurate masses, and MS/MS fragmentation patterns compiled from research articles and reviews (see references in Supporting Information). In total, our database contains 773 reported features. MS/MS data were acquired using DIA on a high‐resolution mass spectrometer in both positive and negative ion modes. The recorded fragmentation spectra were matched against the in‐house database using accurate mass (±6 ppm) and diagnostic fragment ions. Only tentative metabolite identifications were reported in this study.
The potential candidates for each metabolite were obtained by consulting the in‐house database. The dereplication was performed manually using our curated in‐house library of Talaromyces metabolites. Each detected feature was compared with reference data from the library based on exact mass and MS/MS fragmentation patterns. Although custom library integration could improve reproducibility, our in‐house spectral library was not MSP‐compliant for MS‐DIAL, which required manual dereplication.
To visualize sample grouping, an unsupervised multivariate data analysis was performed using MetaboAnalyst 6.0 software (https://www.metaboanalyst.ca). PCA analysis was used to explore the dataset with the capability to verify the linear/polynomial association among variable matrices by reducing the predictive model dimensions, allowing for simple discrimination among samples and the discrimination causing metabolite attributes.
Author Contributions
Cecília L. S. Pereira: formal analysis, methodology. Gabrielle S. Paraguai: formal analysis and methodology – antagonism bioassays. Thiago A. M. Brito: formal analysis – mass data acquisition and analysis. Josean F. Tavares: funding acquisition, writing, review, and editing. Sônia C. O. Melo: formal analysis and methodology – antagonism bioassays. Eliane O. Silva: writing – original draft, project administration, supervision.
Conflicts of Interest
The authors declare no conflicts of interest.
Supplementary Materials
References
- Cacao Pod Husks (Theobroma cacao L.): Composition and Hot‐water‐soluble Pectins,”. Industrial Crops and Products, 2011
- Brazil: 7th Largest Cocoa Producer in the World (. 2022
- Mechanisms Underlying the Protective Effects of Beneficial Fungi Against Plant Diseases,”. Biological Control, 2018
- A Molecular Diagnostic for Moniliophthora perniciosa, the Causal Agent of Witches’ Broom Disease of Cacao, That Differentiates It From Its Sister Taxon Moniliophthora roreri ,”. Crop Protection, 2022
- 5 F. Lamberti , Economic History of Cocoa in Southern Bahia. Its Role on Economy, Society and Culture (Fundação Getúlio Vargas, 2017).
- The Most Important Fungal Diseases of Cereals ‐Problems and Possible Solutions,”. Agronomy, 2021
- Evolving Challenges and Strategies for Fungal Control in the Food Supply Chain,”. Fungal Biology Reviews, 2021. [PubMed]
- Endophytic Fungi: A Tool for Plant Growth Promotion and Sustainable Agriculture,”. Mycology, 2022. [PubMed]
- Exploitation of Endophytes for Sustainable Agricultural Intensification,”. Molecular Plant Pathology, 2017. [PubMed]
- Chemical Ecology of Fungi,”. Natural Product Reports, 2015. [PubMed]
- 11 A. Basit , S. T. Shah , I. Ullah , I. Ullah , and H. I. Mohamed , “Microbial Bioactive Compounds Produced by Endophytes (Bacteria and Fungi) and Their Uses in Plant Health,” in Mohamed in Plant Growth‐Promoting Microbes for Sustainable Biotic and Abiotic Stress Management, eds. H. I. Mohamed , H. E.‐D. S. El‐Beltagi , and K. A. Abd‐Elsalam (Springer International Publishing, 2021), 285–318.
- Endophytic Trichoderma Strains Isolated From Forest Species of the Cerrado‐Caatinga Ecotone Are Potential Biocontrol Agents Against Crop Pathogenic Fungi,”. PLoS ONE, 2022. [PubMed]
- Endophytic Fungi: Biological Control and Induced Resistance to Phytopathogens and Abiotic Stresses,”. Pathogens, 2021. [PubMed]
- Microorganisms in Plant Growth and Development: Roles in Abiotic Stress Tolerance and Secondary Metabolites Secretion,”. Microorganisms, 2022. [PubMed]
- Endophytic Fungi as Biocontrol Agents: A Metabolite‐driven Approach to Crop Protection and Sustainable Agriculture,”. Physiological and Molecular Plant Pathology, 2025
- A Multiscale Study of Fungal Endophyte Communities of the Foliar Endosphere of Native Rubber Trees in Eastern Amazon,”. Scientific Reports, 2018. [PubMed]
- Endophytes: A Rich Source of Functional Metabolites,”. Natural Product Reports, 2001. [PubMed]
- Biology and Chemistry of Endophytes,”. Natural Product Reports, 2006. [PubMed]
- 19 A. Abbas , S. Ali , M. Mubeen , et al., “Talaromyces spp. Are Promising Biocontrol Agents for Sustainable Agriculture,” in Microbial Biocontrol Techniques (Springer, 2024), 245–280, 10.1007/978-981-97-8739-5_13.
- The Control Efficacy and Mechanism of Talaromyces purpurogenus on Fusarium wilt of Bitter Gourd,”. Biological Control, 2022
- Isolation and Characterization of a Newly Discovered Plant Growth‐Promoting Endophytic Fungal Strain From the Genus Talaromyces ,”. Scientific Reports, 2024. [PubMed]
- Research Advances in the Structures and Biological Activities of Secondary Metabolites From Talaromyces,”. Frontiers in Microbiology, 2022. [DOI | PubMed]
- Talaromycolides A–C, Novel Phenyl‐Substituted Phthalides Isolated From the Green Chinese Onion‐Rerived Fungus Talaromyces Pinophilus AF‐02,”. Journal of Agricultural and Food Chemistry, 2015. [PubMed]
- Tubulin Inhibitors From an Endophytic Fungus Isolated From Cedrus deodara ,”. Journal of Natural Products, 2013. [PubMed]
- The Bioactive Secondary Metabolites From Talaromyces Species,”. Nat Prod Bioprospect, 2016. [PubMed]
- Secondary Metabolites of the Endophytic Fungi Talaromyces flavus Cultivated in Pumpkin Medium and Their Bioactivity,”. Chemistry of Natural Compounds, 2023
- Secondary Metabolites With α‐glucosidase Inhibitory Activity From Mangrove Endophytic Fungus Talaromyces sp. CY‐3,”. Marine Drugs, 2021. [PubMed]
- New Secondary Metabolites From Marine‐derived Fungus Talaromyces minnesotensis BTBU20220184,”. Marine Drugs, 2024. [PubMed]
- Merodrimanes and Other Constituents From Talaromyces thailandiasis ,”. Journal of Natural Products, 2007. [PubMed]
- The Metabolites of Talaromyces flavus: Part 1. Metabolites of the Organic Extracts,”. Canadian Journal of Chemistry, 1990
- New Dipyrroloquinones From a Plant‐derived Endophytic Fungus Talaromyces sp,”. Molecules, 2023. [PubMed]
- Secondary Metabolites From the Endophytic Fungus Talaromyces pinophilus ,”. Natural Product Research, 2017. [PubMed]
- Talarostatin, a Vermistatin Derivative From the Soil‐derived Fungus Talaromyces thailandensis PSU‐SPSF059,”. Natural Product Research, 2024. [PubMed]
- Acetylcholinesterase Inhibitors From a Marine Fungus Talaromyces sp. Strain LF458,”. Marine Biotechnology, 2015. [PubMed]
- Chemical Synthesis of Funicone Analogs,”. Tetrahedron, 2012
- New Insights Into Chemical and Biological Properties of Funicone‐Like Compounds,”. Toxins, 2022. [PubMed]
- 3‐O‐Methylfunicone, a Fungitoxic Metabolite Produced by the Fungus Penicillium pinophilum ,”. Phytochemistry, 1999
- The Outstanding Chemodiversity of Marine‐derived Talaromyces ,”. Biomolecules, 2023. [PubMed]
- Computer‐Guided Trypanocidal Activity of Natural Lactones Produced by Endophytic Fungus of Euphorbia umbellata ,”. Chemistry and Biodiversity, 2021. [DOI | PubMed]
- Natural Trypanocidal Product Produced by Endophytic Fungi Through Co‐Culturing,”. Folia Microbiologica, 2020. [DOI | PubMed]
- A Rapid and Efficient Method of Fungal Genomic DNA Extraction, Suitable for PCR Based Molecular Methods,”. Plant Pathology & Quarantine, 2015
- 42 T. J. White , T. Bruns , S. Lee , and J. W. Taylor , “Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics,” in PCR Protocol: A Guide to Methods and Applications, eds. M. A. Innis , D. H. Gelfand , J. J. Sninsky , and T. J. White . (Academic Press, 1990), 315–322.
- The CLUSTAL_X Windows Interface: Flexible Strategies for Multiple Sequence Alignment Aided by Quality Analysis Tools,”. Nucleic Acids Research, 1997. [PubMed]
- MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0,”. Molecular Biology and Evolution, 2007. [PubMed]
- A Simple Method for Estimating Evolutionary Rates of Base Substitutions Through Comparative Studies of Nucleotide Sequences,”. Journal of Molecular Evolution, 1980. [PubMed]
- The Neighbor‐Joining Method: A New Method for Reconstructing Phylogenetic Trees,”. Molecular Biology and Evolution, 1987. [PubMed]
- Antifungal Volatile Organic Compounds From the Endophyte Nodulisporium sp. Strain GS4d2II1a: A Qualitative Change in the Intraspecific and Interspecific Interactions With Pythium aphanidermatum ,”. Microbial Ecology, 2016. [PubMed]
- MS‐DIAL: Data‐independent MS/MS Deconvolution for Comprehensive Metabolome Analysis,”. Nature Methods, 2015. [PubMed]
