Deep eutectic solvent enhances antibacterial activity of a modular lytic enzyme against Acinetobacter baumannii
Abstract
Supplementary Information:
The online version contains supplementary material available at 10.1038/s41598-024-80440-z.
Article type: Research Article
Keywords: Modular lytic enzyme, Endolysin, Deep eutectic solvent, Synergism, Antibacterial effect, Biofilm, Persisters, Antimicrobials, Antimicrobial resistance, Microbiology, Proteins
Affiliations: https://ror.org/011dv8m48grid.8585.00000 0001 2370 4076Laboratory of Extremophiles Biology, Department of Microbiology, Faculty of Biology, University of Gdansk, Wita Stwosza 59, Gdansk, 80-308 Poland; https://ror.org/02kzqn938grid.503422.20000 0001 2242 6780Univ. Lille, CNRS, Univ. Polytechnique Hauts-de-France, UMR 8520 IEMN – Institut d’Electronique de Microélectronique et de Nanotechnologie, Lille, 59000 France; https://ror.org/006x4sc24grid.6868.00000 0001 2187 838XInstitute of Nanotechnology and Materials Engineering, Faculty of Applied Physics and Mathematics, Gdańsk University of Technology, Narutowicza 11/12, Gdansk, 80-233 Poland; https://ror.org/011dv8m48grid.8585.00000 0001 2370 4076Department of General and Medical Biochemistry, Faculty of Biology, University of Gdansk, Wita Stwosza 59, Gdansk, 80-308 Poland; https://ror.org/011dv8m48grid.8585.00000 0001 2370 4076Bioimaging Laboratory, Faculty of Biology, University of Gdansk, Wita Stwosza 59, Gdansk, 80-308 Poland; https://ror.org/00cv9y106grid.5342.00000 0001 2069 7798Department of Biotechnology, Faculty of Bioscience Engineering, Ghent University, Valentin Vaerwyckweg 1, Ghent, 9000 Belgium; https://ror.org/011dv8m48grid.8585.00000 0001 2370 4076Collection of Plasmids and Microorganisms | KPD, Faculty of Biology, University of Gdansk, Wita Stwosza 59, Gdansk, 80-308 Poland
License: © The Author(s) 2025 CC BY 4.0 Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
Article links: DOI: 10.1038/s41598-024-80440-z | PubMed: 39814769 | PMC: PMC11735859
Relevance: Relevant: mentioned in keywords or abstract
Full text: PDF (3.4 MB)
Introduction
Amid the global crisis of antimicrobial resistance (AMR) driven by the increase in multidrug-resistant bacteria, the scientific community’s primary objective is to discover new and effective antibacterial agents1,2. Joint studies by the Institute for Health Metrics and Evaluation (IHME) and the University of Oxford revealed that in 2019 alone, approximately 13.66 million people died from microbe-induced sepsis. Over 65% of these deaths (8.88 million) were attributed to bacterial diseases, with 4.95 million associated with antimicrobial resistance1. The group of pathogens characterized by the highest resistance to currently available therapeutics is called ESKAPEE, an acronym for Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, Enterobacter spp., and Escherichia coli3,4.
A particularly challenging aspect of AMR is the formation of bacterial biofilm, a complex bacterial population that exhibits significantly greater resistance to antibacterial agents, including antibiotics, than planktonic bacteria5–7. While most antibacterial agents show the highest activity against cells in the outer layer of biofilms, the presence of extracellular polymeric substances (EPS) significantly reduces their efficacy in the inner layers7,8. The frequent presence of persister cells, a subpopulation that become dormant in the presence of antibacterial agents and regain physiological activity once exposure ceases9,10, in biofilms poses an additional challenge. This can lead to the rebuilding of biofilms, potentially causing recurrent infections11,12. Therefore, in pursuit of effective strategies counteracting AMR, it is imperative to explore novel approaches and assess their efficacy against planktonic cells and biofilms while expanding the study to include persister cells.
Recently, an alternative approach in antibacterial therapies focused on the application of enzybiotics (enzyme-based antibacterials) has emerged13,14. Among these enzymes, phage-originating endolysins, responsible for the degradation of the peptidoglycan layer of the bacterial cell wall, draw particular attention15,16. Most endolysins are effective against Gram-positive bacteria but have limited efficacy against Gram-negative bacteria17,18. Generally, endolysins active against Gram-negative bacteria have a globular structure with a single enzymatically active (catalytic) domain (EAD). In contrast, endolysins targeting Gram-positive bacteria usually have a multidomain, modular architecture18. The structure of modular endolysins or modular lytic enzymes (MLE) is determined by the presence of at least one EAD and cell wall-binding domain (CBD) connected by a linker18. The limited activity of endolysins against Gram-negative bacteria is a direct result of the presence of an outer membrane18. The outer membrane acts as a chemo-physical barrier that can be penetrated by molecules smaller than 600 Da, which enter through embedded porins. However, it presents an obstacle to larger molecules, such as endolysins17. A significant portion of the ESKAPEE group is represented by Gram-negative bacteria. Therefore, it is necessary to search for solutions to improve the efficacy of lysins against Gram-negative strains18. One promising solution is protein engineering aimed at obtaining modular lytic enzymes (MLEs) with desirable properties. Due to their modular structure (Fig. 1), endolysins of phage origin are well-suited for domain shuffling.
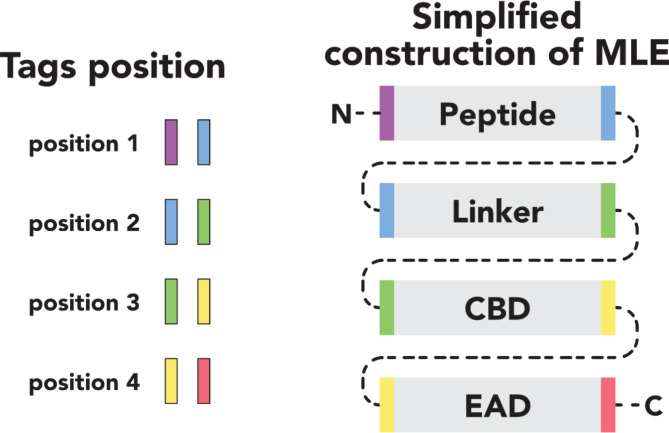
Moreover, the efficacy of endolysins against Gram-negative bacteria can be augmented by fusing them with an antimicrobial peptide (AMP). AMPs are typically small (10–60 amino acids), predominantly cationic peptides found abundantly in nature19. They are often considered a promising substitute for traditional antibiotics20,21. Among AMPs, a category referred to as outer membrane permeabilizing peptides (OMPs) exists22. Fusing an OMP with endolysin produces an MLE with potential activity against Gram-negative bacteria.
In 2020, the high-throughput VersaTile method for modular enzyme engineering was described18. This innovative DNA assembly method takes the advantage of the modular structure of endolysins and relies on their fusion with OMP. Using VersaTile, it is possible to replace and shuffle an infinite number of EADs, CBDs, linkers, and many OMPs18 allowing for quick generation of engineered lysin libraries and rapid screening to identify variants with high antibacterial activity18. In the pioneering publication on VersaTile18, a lead variant 1D10 consisting of OMP7–linker1–CBD6–EAD20 completely inhibited the growth of A. baumannii NCTC13423 and RUH134 in simulated human physiological conditions (50% human serum). Moreover, it slowed biofilm formation in a porcine skin wound infection model18. Obtaining MLE with high antibacterial activity against A. baumannii prompted us to use the VersaTile method to obtain variants containing EAD from the thermostable endolysin Ph2119 discovered in our laboratory23.
Another example of innovative compounds that has not yet been used in combination with endolysins, are natural deep eutectic solvents (NADESs)24. NADESs are an example of green chemistry materials, and in addition to water and lipids, constitute the third class of solvents, providing an alternative to conventional volatile and organic solvents25,26. These viscous liquids are formed as a result of the formation of an intricate hydrogen bond network between hydrogen bond donors (such as amines, sugars, carboxylic acids, and alcohols) and hydrogen bond acceptors (e.g., quaternary ammonium salts choline chloride or betaine)25,27. NADESs are formulated by grinding, sonication, or heating of parent substances and can be synthesized without using solvents or catalysts. They do not require any further purification27. To date, DESs have been used mainly in chemical transformations as substitutes for chemical solvents or in (bio)medicine as drug delivery systems or enhancers of antibiotic activity28–30. Depending on the selection of parent substances, NADESs may exhibit antibacterial properties, as shown in previous studies31,32. NADESs have been implemented to enhance the activity of protein-based drugs, such as tumor necrosis factor (TNF⍺) antibody33, glucagon-like peptide 1, (GLP-1)34 and insulin35. However, to our knowledge, there is currently no information on the synergy between NADESs and antimicrobial agents other than conventional small-molecule antibiotics. Moreover, the mechanism of action of NADES against biofilms and the effectiveness against persister cells have not yet been established.
Therefore, in response to the urgent need for new and effective alternatives to conventional antibiotics, it seems imperative to evaluate the antibacterial properties of MLE. Considering the above, the aim of this study was to investigate the antibacterial effect of modular lytic enzyme (MLE-15) containing the EAD of Ph2119 thermostable endolysin of Thermus scotoductus phage 2119, and reline as a representative of deep eutectic solvents alone and in combination against planktonic bacteria, biofilms and persister cells.
Materials and methods
Bacterial strains, media and culture conditions
All Gram-negative (Pseudomonas aeruginosa PAO1, Pseudomonas fluorescens DSM 50090, Klebsiella pneumoniae P3, Escherichia coli P4-7250, Acinetobacter baumannii CRAB, and Acinetobacter baumannii RUH134) and Gram-positive strains (Staphylococcus aureus Z8, Enterococcus faecalis Z9, and Bacillus subtilis subsp. spizizenii ATCC 6633) used in the study were supplied either by Leibniz-Institut DSMZ – Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH, Germany, the American Type Culture Collection (ATCC), or by the collection of bacterial cultures of the Department of Microbiology, University of Gdansk, Poland. The strains were stored in the form of glycerol stocks at − 80℃. For each analysis, an overnight culture was prepared in fresh Luria-Bertani (LB) broth or Mueller Hinton (MH) broth (Merck, Germany) and incubated at 37℃.
Preparation of reline
Reline was prepared at a molar ratio of 1:2 choline chloride (Sigma Aldrich, C1879-500G) and urea (Sigma Aldrich, U5378-1KG)36. DES was obtained by mixing and heating parent substances slowly, with occasional vortex stirring, until the content of the vessel was completely liquefied. After the substances melted, heating was stopped, and the obtained DES was let to cool down naturally to room temperature. The prepared DES was stored dry at room temperature until the experiments were performed.
Protein structure prediction
VersaTile method used to obtain the library of MLEs was in details described by Gerstmans et al.18. In preliminary screening by bacteria growth inhibitory assay18, out of 200 tested variants, MLE-15 completely inhibited bacterial growth of A. baumannii RUH134 and P. aeruginosa PAO1 (data not shown) and was assigned as a lead variant. This variant consisted of cecropin A (OMP), linker 2, the CBD of endolysin 201phi2-1, and the EAD Ph2119 (thermostable endolysin module) (Table 1; Fig. 2).
Table 1: Components of the modular lytic enzyme – MLE-15.
| Code | Position | Sequential position | Description | Origin | Accession number | Ref. |
|---|---|---|---|---|---|---|
| OMP7 | 1 | Gly2 – Lys37 | Cecropin A | Aedes aegypti (yellow fever mosquito) | AAF59831 | [ref. 37] |
| Link2 | 2 | Gly38-Ala51 | Flexible median linker | In silico design | – | [ref. 18] |
| CBD5 | 3 | Ile56- Gln140 | 201phi2-1gp229-CBD, CBD of endolysin 201phi2-1 | Pseudomonas phage 201phi2-1 | YP_001956952 | [ref. 38] |
| EAD29 | 4 | Arg143- Lys296 | Ph2119 | Thermus phage 2119 | HF20915.1 | [ref. 23] |
| The amino acid sequence of MLE-15 (304 aa) | ||||||
| MGGLKKLGKKLEGAGKRVFNAAEKALPVVAGAKALRKGAGAGAGAGAGAGAGAAGILKNGSKGDDVIRLQRKLIGLGYSVKDDGVFGDNTEKAVKAVQLRFNLKDDGIVGNNTWAVLDTPTTTRPALTDKDYQWAADYLQGSRILEPWNRWYRQKGVYRIRGTPPHYIVLHHTAGPVDQAPEVIRDFHEKGRGWPHIGYHYLVYQDGRVYKTLPNNAIPICVREFNPVSLCIAAVGDFSQGPAWPDNAPGWKALLELKDALVKAYPKAVLVLHKELTQTTCPGVLSWGMVAEKGGKKYHHHHHH | ||||||
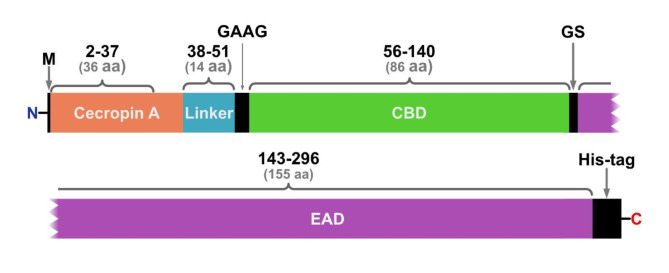
Using the amino acid sequences of the individual fragments, we aimed to construct a 3D model of MLE-15 with AlphaFold v2.3.239, ESMFold_advanced40, DeepFold41, Phyre242, OmegaFold43 and MODELLER44. Predictions were carried out on the Google Colab platform for most tools, with MODELLER executed locally and DeepFold on the Zhang Group platform. Structural homologs of MLE-15 used in MODELLER were found with Foldseek45. Side chain optimization was performed with GalaxyRefine46,47. Modeling accuracy was evaluated in Chimera 1.1648 using superposition to the existing model of the Ph2119 domain (PDB entry: 6SU5). To evaluate the cecropin A fragment, a structural model was predicted with ESMFold and OmegaFold. Foldseek was implemented for searching for similar peptides and calculating TM-scores. Protein visualization was also performed in Chimera 1.16.
Overproduction and purification of modular lytic enzyme
MLE-15 was overexpressed in the E. coli BL21(DE3) pRIL strain. For this purpose, 1 L of LB supplemented with kanamycin (100 µg/mL) and chloramphenicol (30 µg/mL) was used. An overnight culture of E. coli harboring the recombinant plasmid (pVTD-MLE-15) was diluted 100-fold and grown until OD600 of 0.4 − 0.5. Then, the protein overproduction was induced with 1 mM isopropyl-β-d-thiogalactopyranoside (IPTG) following the incubation for 4 h at 37 °C. Finally, bacterial cells were harvested by centrifugation (6,000 × g, 20 min, 4 °C), suspended in NPI-10 buffer (50 mM NaH2PO4, pH 8.0, 300 mM NaCl, 10 mM imidazole, 0.1% Triton X-100, and 10% (v/v) glycerol) and stored at − 80 °C.
Subsequently, purification using the ÄKTA Pure Protein Purification System (Cytiva) was performed. First, cells were disrupted by sonication (30 bursts of 10 s at an amplitude of 12 μm, 4℃). Then, the cell lysates were centrifuged (11 000 × g, 20 min, 4 °C) and heated (20 min, 65℃). After that, the supernatant with His-tagged MLE-15 was subjected to cobalt-based immobilized metal affinity chromatography with 1 mL HiTrap TALON crude columns (Cytiva), washed (NPI buffer with 20 mM imidazole), and eluted (NPI buffer without 0.1% Triton X-100 with 150 mM imidazole). Fractions with pure protein were dialyzed overnight against 20 mM HEPES buffer (pH 7.4) supplemented with 50% glycerol at 4℃. Finally, protein homogeneity was evaluated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), and the protein concentration was determined via Bradford assay using bovine serum albumin as a standard.
Nano differential scanning fluorimetry
MLE-15 sample at a protein concentration of 0.8 mg/mL (in 20 mM HEPES, pH 7.4, 10% glycerol, 25 mM NaCl buffer) was equilibrated to room temperature and centrifuged (10,000 × g, 1 min) to remove any residual air bubbles. Subsequently, the sample was loaded in four replicates into Prometheus standard capillaries and sealed with capillary sealing paste (NanoTemper Technologies) to prevent evaporation. The measurements were conducted at a temperature range from 20 to 110 °C with a 1 °C interval using the Prometheus NT.48 AGOHTU (NanoTemper Technologies) to determine the protein’s thermal stability. During the melting scans, the intrinsic protein fluorescence at 330 nm (F350) and 350 nm (F350) signal was recorded and further using the PR.ThermControl and PR.Stability Analysis software, respectively.
Determination of minimum inhibitory concentration (MIC)
MIC was determined according to Gerstmans et al.18 with modifications49. Briefly, an overnight culture of bacteria was diluted 100-fold in MH broth and grown to an OD600 = 0.5. After that, the cultures were diluted 5 000-fold in MH broth. Four hundred microliters of MLE-15 (200 µg/mL) or reline (50% v/v) prepared in MH broth were transferred onto first row of 96-well microtiter plates. To further wells two hundred microliters of MH was added and 2-fold dilutions of MLE-15 or reline were prepared. For experiments with MLE-15 and reline, reline was mixed with MH broth to achieve the sub-MIC concentration of 4.5% v/v. Solution was transferred onto 96-well microtiter plate. MLE-15 was added to the first row to final concentration of 200 µg/mL and 2-fold serial dilutions of MLE-15 were prepared as described above. Then, 20 µL of adjusted bacteria were added to each well (final count of approx. 2 × 105 cells). Sterile MH broth (200 µL) and medium inoculated with bacteria were used as negative (sterility) and positive (growth) controls, respectively. The samples were then incubated for 18 h at 37 °C, and the MIC values were determined as the lowest concentration that completely inhibited growth visible to the naked eye.
Antibacterial test
Antibacterial tests were performed followed the protocol of Szadkowska et al., 202350. All bacterial strains were grown in LB broth at 37 °C until the OD600 = 0.5. One milliliter of each culture was centrifuged (4,000 × g, 15 min, 20 °C) and washed with 20 mM HEPES buffer (pH 7.4) or saline (0.85% NaCl) for MLE-15 and reline, respectively. Subsequently, the harvested cells (10-fold dilution, approx. 107 cells) were resuspended in either 20 mM HEPES (pH 7.4) or saline supplemented with MLE-15 or reline (MIC value applied based on the results in point 2.6). 20 mM HEPES buffer (pH 7.4) was used as a negative control. The samples were incubated for 1.5 h at 37 °C. Then, the samples were centrifuged (4,000 × g, 15 min, 20 °C), washed with saline, suspended, serially diluted, plated onto Luria-Bertani agar (LA) plates, and incubated for 24 h at 37 °C. After incubation, the colonies were counted, and the results were expressed as log CFU/mL. The antibacterial tests with use of buffers with different tonicity (20 mM HEPES buffer pH 7.4 without or supplemented with 30 mM NaCl (hypotonic buffer), 150 mM NaCl (isotonic buffer), and 350 mM NaCl (hypertonic buffer) were performed according to Vázquez et al., 202451, but because of 600 µL reaction volume, tests were performed in Eppendorf tubes not in sterile 96-deep-well plates.
Biofilm eradication assay
An overnight culture of bacteria was refreshed in LB broth and adjusted to OD600 = 0.5. A 500 µL of adjusted culture was then transferred to a 24-well microtiter plate and 500 µL of LB broth was added and mixed with the culture. The inoculated plate was incubated at 37 °C for 24 h. The medium was removed, and the biofilm was gently washed twice with 1 mL of 20 mM HEPES buffer (pH 7.4) or saline for MLE-15 and reline, respectively. Next, biofilms were treated with 500 µL of MLE-15 or reline at the concentration of 2×MIC. The solutions of MLE-15 and reline were prepared in HEPES or saline respectively. Five hundred µL of HEPES buffer was used as a negative control. Samples were incubated at 37 °C for 3 h. Then, the supernatant was removed, and the biofilm was gently washed twice with saline. Next, the biofilm was harvested with a sterile swab and placed in 1 mL of saline. The samples were serially diluted, and appropriate dilutions were plated onto LB agar plates and incubated at 37 °C for 24 h. After incubation, the colonies were counted, and the results were expressed as log CFU/mL5.
Confocal laser scanning microscopy (CLSM)
Biofilms of A. baumannii RUH134 were prepared in the LabTek™ chamber slide system according to Kocot and Olszewska52. Initially, biofilms were prepared as described in Sect. 2.8. After treatment (MLE-15: 2×MIC, reline: 2×MIC, combination: MLE-15 1×MIC and reline sub-MIC) and subsequent washing, they were stained using LIVE/DEAD Bac Light Bacterial Viability Kit (Thermo Fisher Scientific; L7012) using 1.5 µL of SYTO 9 and 1.5 µL of propidium iodide (PI) diluted in 1 mL of saline. Biofilms were incubated at room temperature for 20 min without light. Next, the staining mixture was removed, and the chambers were detached from the slide. Then, a coverslip was placed on the gasket and mounted with low melting agarose. Images were acquired with a Leica STELLARIS 5 WLL confocal microscope (Wetzlar, Germany) with 495 nm (SYTO9) and 549 nm (PI) lines. Z-stacks (Z = 10 images collected) were acquired using 40× magnification at 1024 × 1024 pixels and zoom = 0.75 and 2. The images shown are maximum intensity projections, and orthogonal sections are displayed using LAS X software. Three-dimensional reconstructions were made using the FIJI (ImageJ™) 3D Viewer plugin displayed as volume. The sets of experimental combinations were visualized twice. Two z-stacks were taken and analyzed for each combination from different places in the chamber.
Persister cells induction and treatment
A. baumannii was grown in LB broth at 37 °C with agitation. The overnight culture was diluted to OD595 = 0.1 in fresh LB broth and supplemented with reline at concentrations of 1×MIC, 3×MIC, 5×MIC, MLE-15 at concentrations of 1×MIC, 3×MIC, 5×MIC, 10×MIC and antibiotic (meropenem) at a concentration of 10×MIC (5 µg/mL; based on preliminary screening tests). This was followed by incubation at 37 °C for 24 h. After 3, 6 and 24 h, the samples were serially diluted and plated onto LB agar. The plates were then incubated at 37 °C for 16 h. The colonies were counted, and the results are expressed as log CFU/mL53. The bacteria were plated on LB agar supplemented with meropenem (10×MIC) to exclude the possibility that the colonies originated from spontaneous meropenem-resistant mutants. The colonies that could not grow on meropenem supplemented LB agar were considered persister cells. After that, meropenem-tolerant persister cells were treated with reline or MLE-15 at a concentration of 1×MIC (12.5% v/v and 100 µg/mL, respectively), and incubated at 37 °C for 24 h. After 3, 6 and 24 h, the samples were diluted and plated onto LB agar. The plates were incubated at 37 °C for 16 h. The colonies were counted, and the results were expressed as log CFU/mL.
Statistical analysis
All the experiments were duplicated in three independent trials (n = 6). Bacterial counts and biofilm densities were expressed as log CFU/mL. Significant differences between the treated and control samples were determined via one-way analysis of variance (ANOVA). All the statistical analyses were performed using Statistica software ver. 13.3 (StatSoft Inc., Tulsa, OK, USA). Differences were considered significant at a level of probability p < 0.05.
Results and discussion
Protein model prediction
The composition of MLE-15 starts with cecropin A, which is connected via a linker to CBD and then to EAD. Bioinformatics was utilized to visualize the tertiary structure of MLE-15 and facilitate understanding of the protein’s overall morphology and the relative positions of its domains. The analysis began with the superposition of models of MLE-15 obtained through various prediction algorithms, including ESMFold, OmegaFold, MODELLER, Phyre2, DeepFold, and AlphaFold2 as well as the known crystallographic structure of EAD domain Ph2119 (PDB: 6SU5) (Fig. 3).
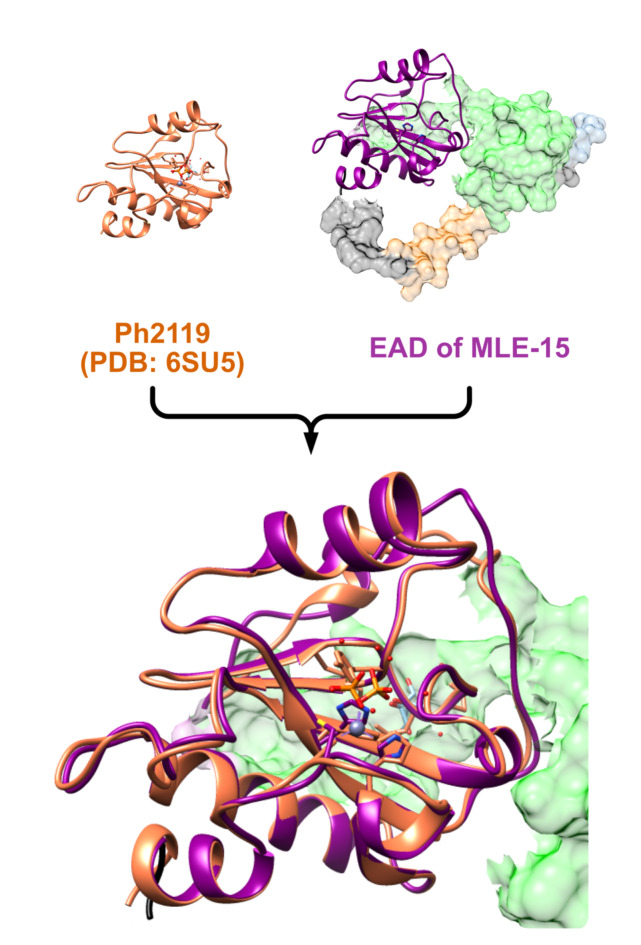
The model predicted by DeepFold was rejected first, because it did not adequately cover Ph2119 and improperly folded the protein, resulting in domains being scattered in a non-globular and highly improbable manner. The Phyre2 model was dismissed as its CBD domain did not align with the other models. While modeling with MODELLER, no template was found for Met1-Ala53 residues in the sequence of MLE-15. Hence, the model obtained with OmegaFold was used as a template by MODELLER instead of covering the missing template. However, this model was also rejected as it merely duplicated the OmegaFold model, rather than creating an original structure. The AlphaFold2 model, which suggested a loop structure between Met1 and Gly110, was discarded due to the expectation of an α-helical structure between Gly2 and Lys37, characteristic of cecropin A54. Subsequently, the investigation focused on cecropin A, the OMP component of MLE-15. ESMFold and OmegaFold algorithms predicted tertiary structure of the N-terminal part of the protein with high probability.
These two models were then analyzed using the Foldseek tool in search of similar structures in protein’s databases. No structures were identified using the ESMFold model, but the OmegaFold model produced several search hits. The TM-score, which measures topological similarity between protein structures, where 1.0 is the highest score, was the best for cecropin A2 (UniProt: Q963B0) with a score of 0.774. Other notable scores included cecropin B1 (UniProt: Q86PR5) at 0.73149 and cecropin C (UniProt: Q8MUF3) at 0.603777, all found in the SWISSPROT database. The only structure found in the RCSB Protein Data Bank was the H5 hemagglutinin mutant Y161A from the A/Viet Nam/1203/2004 H5N1 influenza virus, with similarities in chains B, D, and F. The highest TM-score was 0.67043 in chain D (PDB: 6E7H).
After comparing all models, the tertiary structure constructed using OmegaFold was chosen as the most accurate (Fig. 4). To our knowledge, this is the first protein model representing the structure of the modular enzyme consisting of four building blocks, obtained using the VersaTile methodology.
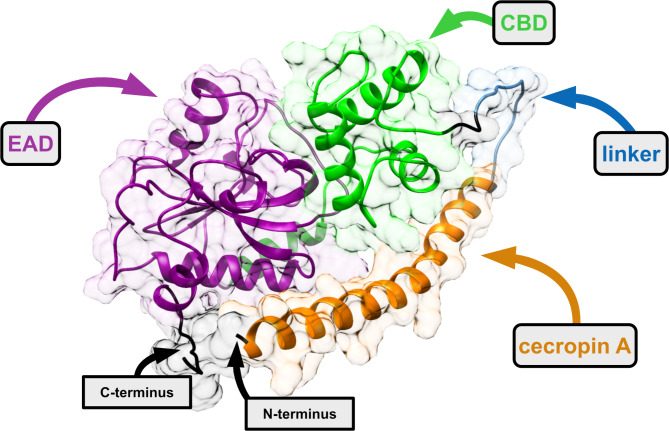
Overproduction, homogeneity and thermal stability of the MLE-15
MLE-15 was overproduced in the E. coli system (Fig. 5A) and purified via metal affinity chromatography through its C-terminal His-tag (Fig. 5B). Analysis of the subsequent purification steps visualized by SDS-PAGE showed that, after purification and dialysis against storage buffer (20 mM HEPES-NaOH, pH 7.4, 50% glycerol), MLE-15 migrated as a single band according to its predicted molecular mass of 33.43 kDa (Fig. 5B, lane 7). The final yield of recombinant protein purification was 10.5 mg/L of bacterial culture.
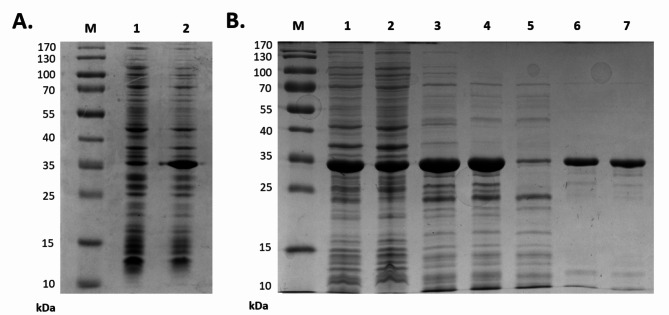
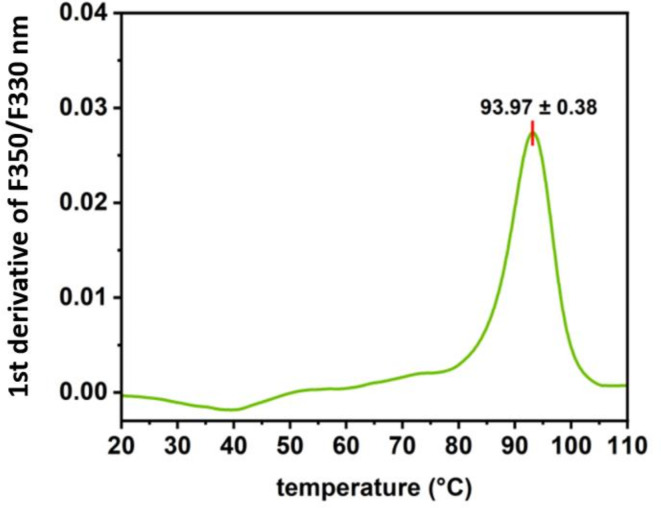
MLE-15 was present in a soluble protein fraction after heat treatment of the bacterial lysate, which proved the stability of the tested protein (Fig. 5, lane 4). To investigate the thermal stability of MLE-15, we implemented nanoDSF, which allowed us to determine the protein melting temperature at which 50% of the protein is in its unfolded form (Fig. 6). The melting temperature of MLE-15 was determined to be 93.97 ± 0.38 °C, which indicated that the enzyme is extremely stable at high temperatures, which is a desirable feature in the production of enzymes of potential use in the biotech industry.
Growth inhibitory effect of MLE-15, reline, and their combination
MIC values for MLE-15 and reline were determined against Gram-positive and Gram-negative bacteria (Table 2). The MIC values for MLE-15 were consistently 100 µg/mL across all tested strains. In contrast, the MIC values for reline varied by strain, ranging from 6.25 to 25% v/v. The lowest value of MIC for reline (6.25% v/v) was observed for (A) baumannii CRAB, the highest (25% v/v) for (B) subtilis, while the MIC for the remaining strains was 12.5% v/v. The MIC values of reline were also tested in comparison to MICs of its parent substances (Supplementary Table S1). All MIC values for reline were lower than the MICs of the constituent substances, therefore reline was selected for further testing. The effect of the combination of MLE-15 with reline was estimated based on fractional inhibitory concentrations (ƩFIC). The combination of MLE-15 with reline at its sub-MIC concentration (4.5% v/v) reduced the MIC of MLE-15 from 100 µg/mL to 3.15–50 µg/mL depending on the tested stain. However, for most strains, the ΣFIC value indicated an additive effect (0.5 < ΣFIC ≤ 1), with synergy observed only for (A) baumannii RUH134 and (B) subtilis (ΣFIC ≤ 0.5). The MIC values of MLE-15 were reduced from 100 µg/mL to 6.25 µg/mL for (A) baumannii RUH134 and 12.5 µg/mL for (B) subtilis. This data demonstrated that the presence of reline reduced the MIC of MLE-15 while maintaining its inhibitory effect on bacterial growth, suggesting that reline could potentially enhance the antibacterial properties of other substances.
Table 2: MIC values of MLE-15, reline, and MLE-15 in the presence of reline (sub-MIC, 4.5% v/v) against selected Gram-positive and Gram-negative bacterial strains. The effect of interaction was estimated based on fractional inhibitory concentrations (ƩFIC) as follows: ƩFIC ≤ 0.5 (synergistic), 0.5 < ƩFIC ≤ 1 (additive), 1 < ƩFIC ≤ 4 (indifferent) and ƩFIC > 4 (antagonistic).
| Strain | MLE-15 | Reline[% v/v] | MLE-15 + reline* | ||
|---|---|---|---|---|---|
| [µg/mL] | MLE-15 + reline | ƩFIC | Interaction | ||
| P. aeruginosa PAO1 | 100 | 12.5 | 50 | 0.88 | additive effect |
| P. fluorescens DSM 50,090 | 100 | 12.5 | 50 | 0.88 | additive effect |
| K. pneumoniae Z4 | 100 | 12.5 | 50 | 0.88 | additive effect |
| E. coli P4-7250 | 100 | 12.5 | 50 | 0.88 | additive effect |
| A. baumannii CRAB | 100 | 6.25 | 3.15 | 0.80 | additive effect |
| A. baumannii RUH134 | 100 | 12.5 | 6.25 | 0.45 | synergism |
| S. aureus Z7 | 100 | 12.5 | 12.5 | 0.51 | additive effect |
| E. faecalis Z9 | 100 | 12.5 | 50 | 0.88 | additive effect |
| B. subtilis ATCC6633 | 100 | 25 | 12.5 | 0.32 | synergism |
* sub-MIC 4.5% v/v.
Bactericidal effect of reline and MLE-15
Based on the observed MIC values, we assessed the antibacterial activity of MLE-15 and reline against Gram-positive and Gram-negative bacteria in planktonic and biofilm forms (Fig. 7). MLE-15 displayed activity against selected planktonic cells of Gram-negative bacteria, showing complete inhibition of the growth of A. baumannii RUH134, and a significant reduction in the number of cells of (A) baumannii CRAB (5.92 log units), and P. aeruginosa PAO1 (5.12 log units). On the other hand, reline caused significant reduction only in the number of planktonic cells of P. aeruginosa (2.26 log units) and (B) subtilis (1.84 log units). It should be noted, that the activity of MLE-15, similarly to other modular enzymes, such as 1D10 consisting of four building blocks (peptide, linker, CBD, and EAD), investigated by Gerstmans et al., (2020), may depend on the tonicity of the reaction buffer18,51. Our results show that in the absence of NaCl or in hypotonic buffer with 30 mM NaCl, MLE-15 caused eradication of A. baumannii RUH134 cells in antibacterial test (≥ 6.40 log units), while 3.19 log reduction of cells counts was observed in the presence of 150 mM NaCl (Supplementary Figure S1). The bactericidal effect of MLE-15 was abolished at 350 mM NaCl, what was also observed in case of 1D10 variant18. Therefore, it should be emphasized, that detailed reaction conditions need to be listed while studying activity of modular lytic enzymes. In relation to the biofilms, MLE-15 was ineffective against all strains tested (p > 0.05), while reline reduced the number of bacterial cells in all biofilms with a significant reduction of A. baumannii RUH134 (1.57 log units).
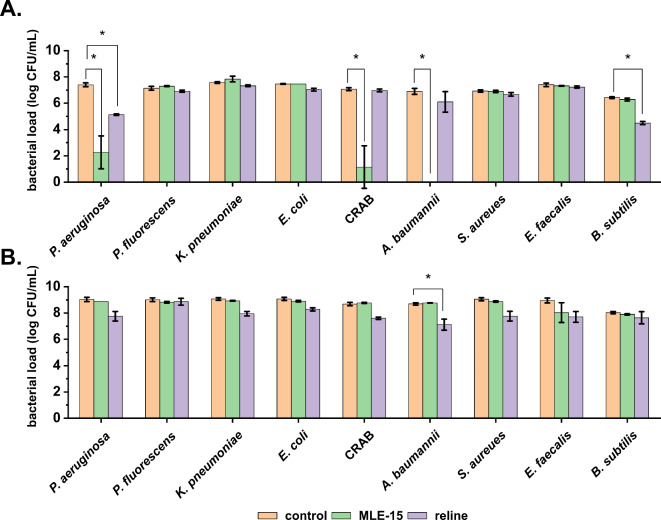
These findings are consistent with previous literature, where DESs were proven to affect biofilms while not completely eradicating them. For example, in the work of Nystedt et al.55, authors investigated three DESs comprising choline chloride and xylitol, choline chloride and glycerol, as well as betaine and sucrose against biofilms of S. aureus and P. aeruginosa. They found that these DESs reduced biofilm biomass by 27–67% for S. aureus and 34–49% for P. aeruginosa. In another study by Nava-Ocampo et al.56, the authors examined two DESs composed of betaine, urea, and water, as well as lactic acid, glucose, and water, against aerobic granular sludge imitating biofilm. The investigated DESs dissolved up to 70% of the extracellular matrix components, weakening the biofilm structure and potentially facilitating its eradication. Silva et al.32 used DESs based on fatty acids (capric acid, myristic acid, lauric acid, and stearic acid) against biofilms formed by S. aureus, E. coli, and C. albicans, demonstrating that a 10-minute exposure to a DES composed of capric acid and lauric acid resulted in over a 50% reduction in biomass of all the investigated biofilms.
However, merely reducing the number of cells in a biofilm without its complete eradication is insufficient in the fight against biofilms. This is due to possible regeneration of the biofilm, which is likely to cause recurrent infections. Despite this, our results and the previously cited studies indicate that DESs can disrupt biofilm structure. This disruption may facilitate the penetration of the antibacterial substances into the complex structure of the biofilm, resulting in more effective interaction with bacterial cells. Therefore, substances that can disrupt the biofilms’ structure (such as conventional DESs like reline) seem promising as co-drugs with other antimicrobials to enhance their efficacy against biofilms.
MLE-15 completely inhibited the growth of planktonic cells of A. baumannii RUH134 but did not show effectiveness in eradicating biofilms. On the other hand, reline was ineffective against planktonic cells but significantly reduced the bacterial load in biofilms of the same strain. Therefore, we investigated the effects of both antimicrobials against this strain in more detail. First, we assessed the impact of these substances against the matured biofilm (Fig. 8). The results confirmed that MLE-15 was ineffective against biofilm (p > 0.05), while reline significantly reduced the number of cells in the biofilm (0.54 log units).
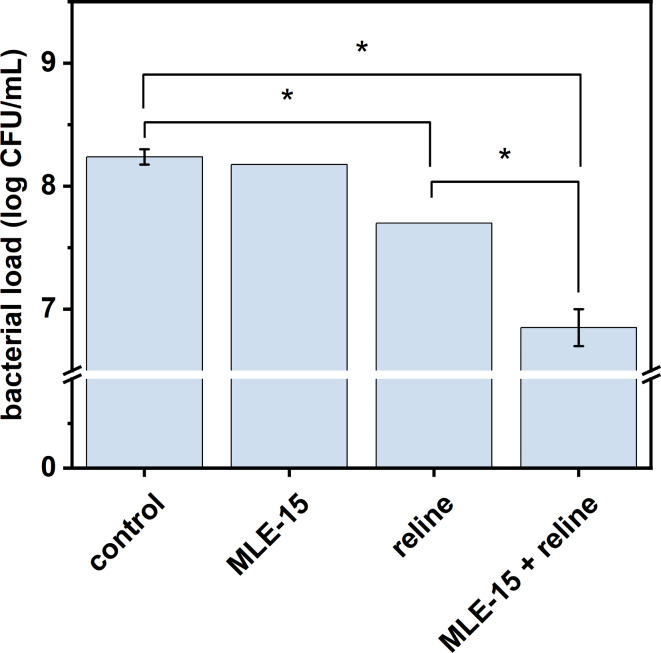
Considering the synergy when MLE-15 was combined with reline, the antibacterial effect of this combination was also examined. The experiment proved that the combination of MLE-15 and reline resulted in a greater reduction (1.39 log units) in the number of cells in the biofilm than when reline was used alone.
To sum up, we proved that MLE-15 at a concentration of 2×MIC was ineffective against A. baumannii RUH134 biofilm, while the presence of reline (sub-MIC) significantly reduced the presence of A. baumannii RUH134 in the biofilm at a concentration of 1×MIC. Thus, it confirmed again that the presence of reline enhanced the antibacterial activity of MLE-15.
A similar approach, where an external substance was used as a co-drug, was reported in a pioneering study on the assessment of the antibacterial properties of MLE, which was obtained using the VersaTile method18. Adding 0.2 mM EDTA – chelator of divalent cations – to lead variant 1D10 reduced its MIC value from 11.8 ± 5.2 to 4.4 ± 3.6 µg/mL in tests on A. baumannii. Moreover, it was shown that 0.2 mM EDTA slightly increased the antibacterial effect against A. baumannii strains in 90% human serum (reduction from 2.4 to 2.7 to 2.7–3.1 log units). The use of substances that enhance the antibacterial effect of lysins is known in the literature and has been reported as early as 201216,57,58. It is common to use EDTA or weak organic acids as outer membrane permeabilizers, allowing lysins to reach and degrade the peptidoglycan layer. For instance, Oliveira et al.57 used EDTA and weak organic acids (malic and citric) to evaluate the antibacterial properties of endolysin Lys68. Authors found that supplementation of Lys68 with 0.2 mM EDTA displayed an antibacterial effect against Pseudomonas strains, while the presence of 5 mM malic and 2 mM citric acid broadened the antibacterial effect of Lys68 to Salmonella Typhimurium. Moreover, it was demonstrated that combinations malic acid/Lys68 and citric acid/Lys68 reduced the number of Salmonella cells by 3–5 log units compared to control samples treated with acids alone57. Therefore, our research on combining the antibacterial action of MLE-15 and reline fits existing trends. This approach is also innovative due to the co-action of MLE with deep eutectic solvents represented here by reline, which has not been studied in this context so far. Moreover, our study indicates a possible mechanism of action combining MLE-15 with reline. We showed that MLE-15 had an antibacterial effect on planktonic cells of A. baumannii RUH134 at a concentration of 1×MIC, while using up to twice the concentration did not affect the biofilm. However, adding reline at a sub-MIC concentration allowed for a significant reduction of cells in the biofilm when combined with MLE-15.
Considering observations mentioned above and the lack of antibacterial effect of MLE-15 alone on the biofilms, one can assume that the activity of reline is involved in the dissolution of the extracellular matrix of the biofilm. Thus, we suspect that reline disrupted the EPS, facilitating the penetration of MLE-15 into the biofilm structure which allowed MLE-15 to act directly on bacterial cells. These observations were supported by microscopy images obtained while implementing LIVE/DEAD staining on the biofilms (Fig. 9). The untreated biofilm showed that A. baumannii RUH134 formed a densely-packed biofilm, mostly labeled with SYTO 9 (green cells), with few scattered, damaged cells labeled with PI (red cells), indicating high cell viability within the biofilm (Fig. 9A1). The biofilm treated with MLE-15 displayed a visual effect similar to the untreated control, which showed the lack of activity of MLE-15 against the biofilm of A. baumannii RUH134 (Fig. 9B1). On the contrary, biofilms treated with reline (Fig. 9C1) appeared yellowish-orange, while the combination of MLE-15 and reline (Fig. 9D1) resulted in red-stained spots. 3-dimensional analysis of the biofilms indicated that the control (Fig. 9A3, 9A5) and MLE-15-treated biofilms (Fig. 9B3, 9B5) were thicker than biofilms treated with reline (Fig. 9C3, 9C5) or the combination of MLE-15 with reline (Fig. 9D3, 9D5). Furthermore, structural analysis showed very few spaces devoid of cells in the control (Fig. 9A2, 9A4) and MLE-15-treated-biofilm (Fig. 9B2, 9B4), while reline-treated biofilm (Fig. 9C2, 9C4) and biofilm treated with a combination of both (Fig. 9D2, 9D4) resulted in the formation of holes and large areas without visible accumulated biomass. The observations mentioned above confirmed that MLE-15 alone affected neither the viability of A. baumannii RUH134 cells in the biofilm nor the structure of the biofilm. On the contrary, using reline alone caused a visible disintegration of the biofilm structure resulting in the cells stained yellowish-orange, indicating a specific effect on the physiology of bacterial cells. In turn, the use of reline combined with MLE-15 revealed damage to the biofilm structure by reline, which enabled the penetration of MLE-15 into the biofilm and a direct effect on the damage of the cytoplasmic membranes of bacterial cells which allowed the penetration of PI into the cells. Presumably, reline alone requires more prolonged action time to affect bacterial cells substantially. However, from the medical point-of-view, it seems more reasonable to use reline with another antibacterial substance, such as MLE-15, than to extend the time needed to inactivate bacterial pathogens.
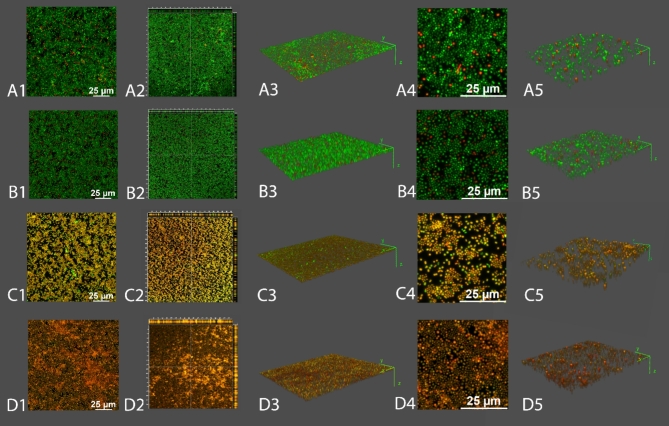
Previously-mentioned studies on assessing the impact of DESs on biofilms6,32,55 also indicated that DESs disrupted the biofilm structure. Our proposed combination of reline with MLE-15 seems to be important in the context of an effective fight against biofilms, which may give more satisfactory results than DESs alone.
Treatment and induction of emergence of persister cells
In addition to the biofilms, persister cells are the leading cause of recurring bacterial infections and pose a formidable challenge to modern medicine. Therefore, the next step in our studies was to investigate whether MLE-15 and reline cause the emergence of persisters of A. baumannii RUH134. Our study showed that MLE-15 (Fig. 10A) used at concentrations of 1×, 3×, 5× and 10×MIC as well as reline (Fig. 10B) used at concentration of 1×, 3× and 5×MIC did not cause to emergence of persister cells. Moreover, it was found that MLE-15, regardless of concentration, caused gradual cell death over time until complete eradication 72 h after exposure (Fig. 10A). In turn, reline at a concentration of 3× and 5×MIC caused complete eradication of A. baumannii RUH134 after only 7 h of treatment (Fig. 10B). These results are extremely valuable due to the eradication, not just the reduction of A. baumannii RUH134 cells and the lack of tolerance to these antimicrobials.
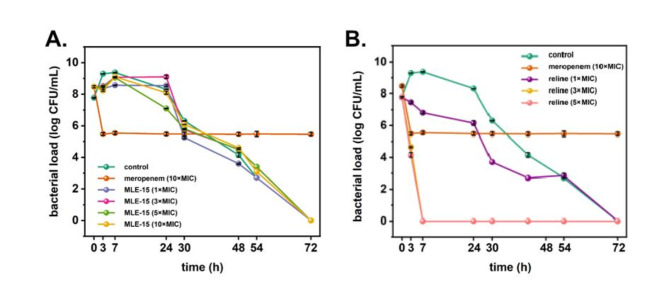
Finally, we assessed whether MLE-15, reline, and their combination could combat persister cells of A. baumannii RUH134 induced using an antibiotic – meropenem (10×MIC) (Fig. 11). Meropenem-induced persister cells were treated with MLE-15 (1×MIC), reline (1×MIC), and their combination (MLE-15–1×MIC and reline – sub-MIC). After the addition of MLE-15, the number of the cells remained stable at the level of approx. 5 log CFU/mL for a total of 24 h, which indicated the lack of the activity of MLE-15 against persisters. In turn, reline, and its combination with MLE-15 eradicated persister cells in just 3 h after treatment, and this effect was maintained in the following hours, which indicated strong activity against persister cells.
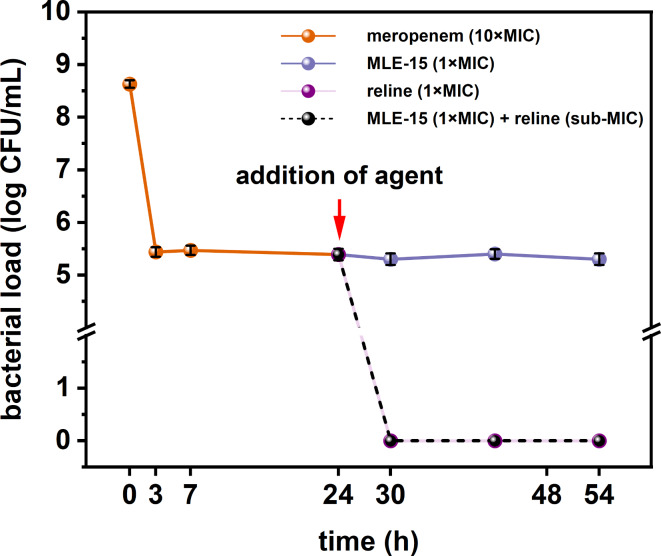
These observations are valuable due to the global AMR crisis. Antibiotic-tolerant persister cells are one of the reasons for the overuse of antibiotics. Therefore, our results show that reline and its combination with MLE-15 completely eradicated meropenem-induced persister cells and provided hope for effective antibacterial therapy in the near future.
Conclusion
In the current global crisis related to antimicrobial resistance, it is imperative to search for novel and innovative solutions to counteract bacterial infections effectively. In this study, we investigated the antibacterial activity of MLE-15, a modular lytic enzyme based on an enzymatically active domain from thermostable endolysin Ph2119, obtained with the VersaTile method. We also examined reline, a natural deep eutectic solvent and their combination. Our result showed that MLE-15 completely inhibited the growth of planktonic cells of extensively resistant strain A. baumannii RUH134, while reline significantly reduced the number of A. baumannii RUH134 cells in the biofilm. Moreover, both antimicrobials significantly reduced the number of cells in fully formed biofilm. This finding suggests that the activity of MLE-15, which was ineffective against biofilm when used alone, was enhanced in the presence of reline. It appears that reline disrupts extracellular matrix and destabilizes the biofilm structure. These findings were confirmed by microscopic analysis, which demonstrated that reline enhanced the antibacterial properties of MLE-15, indicating that combining MLE-15 with reline is a promising strategy for combating biofilms. Additionally, our study showed that MLE-15 and reline did not cause the emergence of persister cells, and reline alone or in combination with MLE-15 eradicated persister cells induced in the presence of meropenem. These findings suggest that reline, through its action on the outer layers and destabilizing function, acts as a permeability enhancer for MLE-15, enabling it to act directly on bacterial cells. The complete eradication of meropenem-persister cells of this strain highlights the potential of combining MLE-15 with reline in the fight against bacterial infections and antibiotic-resistant strains.
Endolysins, i.e. lytic enzymes of phage origin (e.g. MLE-15 created by VersaTile methodology) are currently facing great interest in the context of the therapy of bacterial infections (e.g. LysRODI against S. aureus59, Ph28 against S. epidermidis60, ORF28 gene product against E. faecalis61). There is also no shortage of advanced clinical or preclinical trials on the use of endolysins to combat bacterial infections62,63. Therefore, the issue of lytic enzymes in therapy is absolutely justified and necessary. However, most endolysins are tested against Gram-positive bacteria, because their effectiveness against Gram-negative bacteria is limited by the presence of an additional cover – the outer membrane. Consequently, studies like ours are a valuable contribution in research on combating Gram-negative bacteria.
In turn, DESs are known in biomedicine as treatment (e.g. wound healing64), antimicrobial (e.g. effect against Escherichia coli, Proteus mirabilis, Salmonella typhimurium, Pseudomonas aeruginosa and Staphylococcus aureus65) or tool for increasing the antibacterial properties of other therapeutics (e.g. increase the effects of clindamycin in in vivo tests in mice30). Nevertheless, our work is the first to indicate an increase in the antibacterial effect of lytic enzymes in the presence of DES (reline). Using a combination of two or more agents to obtain the desired antibacterial effect is a popular strategy in medicine today. Therefore, the research we propose meets current trends and perfectly combines both innovative antibacterial approaches – the use of lytic enzyme and DES. The combination of these two antimicrobials, which are currently being intensively studied, may constitute a real alternative to conventional antibiotics.
Supplementary Materials
References
- 1.Murray, C. J. et al. Feb., Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis, The Lancet, vol. 399, no. 10325, pp. 629–655, (2022). 10.1016/S0140-6736(21)02724-0
- 2.Huemer, M., Shambat, S. M., Brugger, S. D. & Zinkernagel, A. S. Antibiotic resistance and persistence—Implications for human health and treatment perspectives, EMBO Rep, vol. 21, no. 12, p. e51034, Dec. (2020). 10.15252/EMBR.202051034
- 3.Mulani, M. S., Kamble, E. E., Kumkar, S. N., Tawre, M. S. & Pardesi, K. R. Emerging strategies to combat ESKAPE pathogens in the era of antimicrobial resistance: A review, Front. Microbiol., 10, 403107. 10.3389/FMICB.2019.00539 (2019).
- 4.Yu, Z., Tang, J., Khare, T. & Kumar, V. The alarming antimicrobial resistance in ESKAPEE pathogens: can essential oils come to the rescue? Fitoterapia14010.1016/J.FITOTE.2019.104433 (Jan. 2020).
- 5.Szadkowska, M. et al. A Novel cryptic clostridial peptide that kills Bacteria by a cell membrane permeabilization mechanism. Microbiol. Spectr.10(5). 10.1128/SPECTRUM.01657-22 (2022).
- 6.Kocot, A. M., Wróblewska, B. & Cabo, M. L. Operational culture conditions determinate benzalkonium chloride resistance in L. monocytogenes-E. Coli dual species biofilms. Int. J. Food Microbiol.360, 109441. 10.1016/J.IJFOODMICRO.2021.109441 (Dec. 2021).
- 7.Sharma, D., Misba, L. & Khan, A. U. Antibiotics versus biofilm: an emerging battleground in microbial communities, Antimicrobial Resistance & Infection Control 2019 8:1, vol. 8, no. 1, pp. 1–10, May (2019). 10.1186/S13756-019-0533-3
- 8.Abebe, G. M. The role of bacterial biofilm in Antibiotic Resistance and Food Contamination. Int. J. Microbiol.202010.1155/2020/1705814 (2020).
- 9.Berkvens, A., Chauhan, P. & Bruggeman, F. J. Integrative biology of persister cell formation: molecular circuitry, phenotypic diversification and fitness effects. J. R Soc. Interface. 19(194). 10.1098/RSIF.2022.0129 (2022).
- 10.Balaban, N. Q. et al. Definitions and guidelines for research on antibiotic persistence. Nature Rev. Microbiol., 17(7), 441–448. 10.1038/s41579-019-0196-3 (2019).
- 11.Levin-Reisman, I. et al. Antibiotic tolerance facilitates the evolution of resistance. Science. 355(6327), 826–830. 10.1126/SCIENCE.AAJ2191 (2017).
- 12.Lebeaux, D., Ghigo, J. M. & Beloin, C. Biofilm-Related Infections: Bridging the Gap between Clinical Management and Fundamental Aspects of Recalcitrance toward Antibiotics, Microbiol. Mol. Biol. Rev.78(3), 510 (2014). 10.1128/MMBR.00013-14 (2014).
- 13.Weber, G., Buhren, B., Schrumpf, H., Wohlrab, J. & Gerber, P. Therapeutic enzymes: function and clinical implications., Adv Exp Med Biol, vol. 1148, pp. 233–252, Accessed: Jun. 13, 2024. [Online]. Available: http://link.springer.com/ (2019). 10.1007/978-981-13-7709-9
- 14.Danis-Wlodarczyk, K. M., Wozniak, D. J. & Abedon, S. T. Treating Bacterial Infections with Bacteriophage-Based Enzybiotics: In Vitro, In Vivo and Clinical Application, Antibiotics 2021, Vol. 10, Page 1497, vol. 10, no. 12, p. 1497, Dec. (2021). 10.3390/ANTIBIOTICS10121497
- 15.Schmelcher, M., Donovan, D. M. & Loessner, M. J. Bacteriophage endolysins as novel antimicrobials. Fut. Microbiol.7, (10), 1147–1171. 10.2217/FMB.12.97 (2012).
- 16.Plotka, M., Kapusta, M., Dorawa, S., Kaczorowska, A. K. & Kaczorowski, T. Ts2631 Endolysin from the Extremophilic Thermus scotoductus Bacteriophage vB_Tsc2631 as an Antimicrobial Agent against Gram-Negative Multidrug-Resistant Bacteria, Viruses 2019, Vol. 11, Page 657, vol. 11, no. 7, p. 657, Jul. (2019). 10.3390/V11070657
- 17.Jeong, T. H., Hong, H. W., Kim, M. S., Song, M. & Myung, H. Characterization of three different endolysins effective against gram-negative bacteria. Viruses, 15(3), 679, 10.3390/V15030679 (2023).
- 18.Gerstmans, H. et al. A VersaTile-driven platform for rapid hit-to-lead development of engineered lysins. Sci. Adv.6(23). 10.1126/SCIADV.AAZ1136 (2020).
- 19.Huan, Y., Kong, Q., Mou, H. & Yi, H. Antimicrobial peptides: classification, design, application and research progress in multiple fields. Front. Microbiol.11, 582779. 10.3389/FMICB.2020.582779 (2020).
- 20.Mazurkiewicz-Pisarek, A., Baran, J. & Ciach, T. Antimicrobial peptides: challenging Journey to the Pharmaceutical, Biomedical, and Cosmeceutical Use. Int. J. Mol. Sci.. 24(10), 9031. 10.3390/IJMS24109031 (2023).
- 21.Mahlapuu, M., Håkansson, J., Ringstad, L. & Björn, C. Antimicrobial Peptides: An Emerging Category of Therapeutic Agents, Front Cell Infect Microbiol. 6, 194. 10.3389/FCIMB.2016.00194 (2016).
- 22.Henriques, S. T., Melo, M. N. & Castanho, M. A. R. B. Cell-penetrating peptides and antimicrobial peptides: how different are they? Biochem. J.399(1), 1–7. 10.1042/BJ20061100 (2006)
- 23.Plotka, M. et al. Novel highly thermostable endolysin from Thermus scotoductus MAT2119 bacteriophage Ph2119 with amino acid sequence similarity to eukaryotic peptidoglycan recognition proteins. Appl. Environ. Microbiol.80 (3), 886–895. 10.1128/AEM.03074-13 (2014).
- 24.Siddiqui, R. et al. The increasing importance of novel deep eutectic solvents as potential effective antimicrobials and other medicinal properties. World J Microbiol Biotechnol, 39, (12), 1–10. 10.1007/S11274-023-03760-8 (2023).
- 25.Thapa, R. K., Kim, J. O. & Kim, J. Antimicrobial strategies for topical biofilm-based wound infections: past, present, and future. Journal of Pharmaceutical Investigation, 53(5), 627–641. 10.1007/S40005-023-00628-9 (2023).
- 26.Svigelj, R., Dossi, N., Grazioli, C. & Toniolo, R. Deep Eutectic solvents (DESs) and their application in Biosensor Development. Sens. 2021. 21, Page 4263, 21, (13), 4263. 10.3390/S21134263 (Jun. 2021).
- 27.Tiecco, M. et al. Advances in the development of novel green liquids: thymol/water, thymol/urea and thymol/phenylacetic acid as innovative hydrophobic natural deep eutectic solvents. J. Mol. Liq. 364, 120043. 10.1016/J.MOLLIQ.2022.120043 (Oct. 2022).
- 28.Pereira, C. V. et al. Unveil the Anticancer potential of Limomene Based Therapeutic Deep Eutectic solvents. Sci. Rep.9 (1). 10.1038/S41598-019-51472-7 (Dec. 2019).
- 29.Lu, C., Cao, J., Wang, N. & Su, E. Significantly improving the solubility of non-steroidal anti-inflammatory drugs in deep eutectic solvents for potential non-aqueous liquid administration. Medchemcomm7 (5), 955–959. 10.1039/C5MD00551E (May 2016).
- 30.Zakrewsky, M. et al. Jun., Choline and Geranate Deep Eutectic Solvent as a Broad-Spectrum Antiseptic Agent for Preventive and Therapeutic Applications, Adv Healthc Mater, vol. 5, no. 11, pp. 1282–1289, (2016). 10.1002/ADHM.201600086
- 31.Akbar, N. et al. Antimicrobial activity of Novel Deep Eutectic solvents. Sci. Pharm.91(1), 9. 10.3390/SCIPHARM91010009 (2023).
- 32.Silva, J. M., Silva, E., Reis, R. L. & Duarte, A. R. C. A closer look in the antimicrobial properties of deep eutectic solvents based on fatty acids. Sustain. Chem. Pharm.14, 100192. 10.1016/J.SCP.2019.100192 (Dec. 2019).
- 33.Angsantikul, P. et al. Ionic liquids and deep Eutectic solvents for enhanced delivery of antibodies in the gastrointestinal tract. Adv. Funct. Mater.31, 2002912. 10.1002/ADFM.202002912 (Oct. 2021).
- 34.Agatemor, C. et al. Choline-Geranate Deep Eutectic Solvent improves Stability and Half-Life of Glucagon-Like Peptide-1. Adv. Ther. (Weinh). 4 (3), 2000180. 10.1002/ADTP.202000180 (Mar. 2021).
- 35.Khamoushian, S. et al. Aug., Transdermal Delivery of Insulin Using Combination of Iontophoresis and Deep Eutectic Solvents as Chemical Penetration Enhancers: In Vitro and in Vivo Evaluations. J Pharm Sci. 112(8), 2249–2259. 10.1016/J.XPHS.2023.03.005 (2023).
- 36.Hammond, O. S., Bowron, D. T. & Edler, K. J. Liquid structure of the choline chloride-urea deep eutectic solvent (reline) from neutron diffraction and atomistic modelling. Green Chem.18 (9), 2736–2744. 10.1039/C5GC02914G (May 2016).
- 37.Silvestro, L., Weiser, J. N. & Axelsen, P. H. Antibacterial and antimembrane activities of cecropin A in Escherichia coli. Antimicrob. Agents Chemother.44(3), 602–607. 10.1128/AAC.44.3.602-607.2000 (2000).
- 38.Thomas, J. A. et al. Jul., Characterization of Pseudomonas chlororaphis myovirus 201ϕ2 – 1 via genomic sequencing, mass spectrometry, and electron microscopy, Virology, vol. 376, no. 2, pp. 330–338, (2008). 10.1016/J.VIROL.2008.04.004
- 39.Jumper, J. et al. Jul., Highly accurate protein structure prediction with AlphaFold, Nature 2021 596:7873, vol. 596, no. 7873, pp. 583–589, (2021). 10.1038/s41586-021-03819-2
- 40.Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model, Science , 379(6637), 1123–1130. 10.1126/SCIENCE.ADE2574 (2023).
- 41.Pearce, R., Li, Y., Omenn, G. S. & Zhang, Y. Fast and accurate ab initio protein structure prediction using deep learning potentials. PLoS Comput. Biol.18 (9), e1010539. 10.1371/JOURNAL.PCBI.1010539 (Sep. 2022).
- 42.Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis, Nature Protocols 2015 10:6, vol. 10, no. 6, pp. 845–858, May (2015). 10.1038/nprot.2015.053
- 43.Wu, R. et al. Jul., High-resolution de novo structure prediction from primary sequence, bioRxiv, p. 2022.07.21.500999, (2022). 10.1101/2022.07.21.500999
- 44.Šali, A. & Blundell, T. L. Comparative Protein Modelling by Satisfaction of Spatial Restraints, J Mol Biol, vol. 234, no. 3, pp. 779–815, Dec. (1993). 10.1006/JMBI.1993.1626
- 45.van Kempen, M. et al. May., Fast and accurate protein structure search with Foldseek, Nature Biotechnology 2023 42:2, vol. 42, no. 2, pp. 243–246, (2023). 10.1038/s41587-023-01773-0
- 46.Heo, L., Park, H. & Seok, C. GalaxyRefine: protein structure refinement driven by side-chain repacking. Nucleic Acids Res.41, W384–W388. 10.1093/NAR/GKT458 (Jul. 2013). no. W1.
- 47.Lee, G. R., Heo, L. & Seok, C. Effective protein model structure refinement by loop modeling and overall relaxation, Proteins: Structure, Function, and Bioinformatics, vol. 84, no. S1, pp. 293–301, Sep. (2016). 10.1002/PROT.24858
- 48.Pettersen, E. F. et al. Jan., UCSF ChimeraX: Structure visualization for researchers, educators, and developers, Protein Science, vol. 30, no. 1, pp. 70–82, (2021). 10.1002/PRO.3943
- 49.Minimum Inhibitory Concentration (MIC) and Minimum Bactericidal Concentration (MBC). Assays Using Broth Microdilution Method. Accessed: Jun. 13, 2024. [Online]. Available: https://www.protocols.io/view/minimum-inhibitory-concentration-mic-and-minimum-b-5qpvo3x6dv4o/v1
- 50.Szadkowska, M. et al. Molecular characterization of the PhiKo endolysin from Thermus thermophilus HB27 bacteriophage phiKo and its cryptic lytic peptide RAP-29. Front. Microbiol.14, 1303794. 10.3389/FMICB.2023.1303794 (2023).
- 51.Vázquez, R. et al. You get what you test for: the killing effect of phage lysins is highly dependent on buffer tonicity and ionic strength. Microb. Biotechnol.17 (7), e14513. 10.1111/1751-7915.14513 (Jul. 2024).
- 52.Kocot, A. M. & Olszewska, M. A. Interaction and inactivation of Listeria and Lactobacillus cells in single and mixed species biofilms exposed to different disinfectants, J Food Saf, vol. 39, no. 6, p. e12713, Dec. (2019). 10.1111/JFS.12713
- 53.Kuczyńska-Wiśnik, D., Stojowska-Swędrzyńska, K. & Laskowska, E. Intracellular Protective Functions and Therapeutical Potential of Trehalose, Molecules 2024, Vol. 29, Page 2088, vol. 29, no. 9, p. May 2024, (2088). 10.3390/MOLECULES29092088
- 54.Steiner, H. Secondary structure of the cecropins: antibacterial peptides from the moth Hyalophora cecropia. FEBS Lett.137 (2), 283–287. 10.1016/0014-5793(82)80368-2 (Jan. 1982).
- 55.Nystedt, H. L. et al. Neutral natural deep eutectic solvents as anti-biofilm agents, Biofilm, vol. 5, p. 100114, Dec. (2023). 10.1016/J.BIOFLM.2023.100114
- 56.Nava-Ocampo, M. F. et al. Natural deep eutectic solvents as biofilm structural breakers. Water Res.201, 117323. 10.1016/J.WATRES.2021.117323 (Aug. 2021).
- 57.Oliveira, H. et al. A Thermostable Salmonella Phage Endolysin, Lys68, with Broad Bactericidal properties against Gram-negative pathogens in Presence of weak acids. PLoS One. 9 (10), e108376. 10.1371/JOURNAL.PONE.0108376 (Oct. 2014).
- 58.Walmagh, M., Briers, Y., dos Santos, S. B., Azeredo, J. & Lavigne, R. Characterization of modular bacteriophage endolysins from Myoviridae Phages OBP, 201φ2 – 1 and PVP-SE1. PLoS One. 7 (5), e36991. 10.1371/JOURNAL.PONE.0036991 (May 2012).
- 59.Portilla, S., Fernández, L., Gutiérrez, D., Rodríguez, A. & García, P. Encapsulation of the Antistaphylococcal Endolysin LysRODI in pH-Sensitive liposomes. Antibiot. 2020. 9 (5), 242. 10.3390/ANTIBIOTICS9050242 (May 2020). Page 242.
- 60.Muharram, M. M., Abulhamd, A. T., Aldawsari, M. F., Alqarni, M. H. & Labrou, N. E. Development of Staphylococcus Enzybiotics: The Ph28 Gene of Staphylococcus epidermidis Phage PH15 Is a Two-Domain Endolysin, Antibiotics 2020, Vol. 9, Page 148, vol. 9, no. 4, p. 148, Mar. (2020). 10.3390/ANTIBIOTICS9040148
- 61.Zhang, H. et al. Jul., Bacteriophage φEf11 ORF28 Endolysin, a Multifunctional Lytic Enzyme with Properties Distinct from All Other Identified Enterococcus faecalis Phage Endolysins, Appl Environ Microbiol, vol. 85, no. 13, pp. 555–574, (2019). 10.1128/AEM.00555-19
- 62.Harhala, M. et al. Safety Studies of Pneumococcal Endolysins Cpl-1 and Pal, Viruses Vol. 10, Page 638, vol. 10, no. 11, p. 638, Nov. 2018, (2018). 10.3390/V10110638
- SY Jun. Preclinical safety evaluation of intravenously administered SAL200 containing the recombinant phage endolysin SAL-1 as a pharmaceutical ingredient. Antimicrob. Agents Chemother., 2014. [DOI | PubMed]
- JM Silva. Therapeutic role of Deep Eutectic solvents based on menthol and saturated fatty acids on Wound Healing. ACS Appl. Bio Mater., 2019. [DOI]
- 65.Radošević, K. et al. Antimicrobial, cytotoxic and antioxidative evaluation of natural deep eutectic solvents. Environ. Sci. Pollut. Res.25(14), 14188–14196. 10.1007/S11356-018-1669-Z (2018).
