Dual agonistic and antagonistic roles of ZC3H18 provide for co-activation of distinct nuclear RNA decay pathways
Abstract
The RNA exosome is a versatile ribonuclease. In the nucleoplasm of mammalian cells, it is assisted by its adaptors the nuclear exosome targeting (NEXT) complex and the poly(A) exosome targeting (PAXT) connection. Via its association with the ARS2 and ZC3H18 proteins, NEXT/exosome is recruited to capped and short unadenylated transcripts. Conversely, PAXT/exosome is considered to target longer and adenylated substrates via their poly(A) tails. Here, mutational analysis of the core PAXT component ZFC3H1 uncovers a separate branch of the PAXT pathway, which targets short adenylated RNAs and relies on a direct ARS2-ZFC3H1 interaction. We further demonstrate that similar acidic-rich short linear motifs of ZFC3H1 and ZC3H18 compete for a common ARS2 epitope. Consequently, while promoting NEXT function, ZC3H18 antagonizes PAXT activity. We suggest that this organization of RNA decay complexes provides co-activation of NEXT and PAXT at loci with abundant production of short exosome substrates.
Article type: Research Article
Keywords: nuclear RNA decay, activation, inhibition, PAXT, NEXT, ARS2, ZC3H18
Affiliations: Department of Molecular Biology and Genetics, Universitetsbyen 81, Aarhus University, Aarhus, Denmark; Department of Structural Cell Biology, Max Planck Institute of Biochemistry, Am Klopferspitz 18, Martinsried/Munich, Germany; Department of Biochemistry and Molecular Biology, University of Southern Denmark, Campusvej 55, Odense M, Denmark
License: © 2023 The Author(s) CC BY 4.0 This is an open access article under the CC BY license (http://creativecommons.org/licenses/by/4.0/).
Article links: DOI: 10.1016/j.celrep.2023.113325 | PubMed: 37889751 | PMC: PMC10720265
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (20.2 MB)
Introduction
Mammalian genomic DNA is pervasively transcribed by RNA polymerase II (RNAPII), generating a large volume of unstable noncoding RNA (ncRNA) along with the canonical production of mRNA and stable ncRNA.ref. bib1 Relevantly, a large share of RNAPII transcription initiation events is estimated to be non-productive and subjected to premature transcription termination with the released transcript being rapidly removed by RNA decay.ref. bib2,ref. bib3,ref. bib4 Such termination can occur outside of, or early within, conventional transcription units (TUs) and be induced by the presence of transcription start site (TSS)-proximal polyadenylation (pA) sites,ref. bib5,ref. bib6,ref. bib7 recruiting the cleavage and pA complex and yielding short pA+ RNA.ref. bib8 Features triggering early transcription termination can also be more elusiveref. bib9,ref. bib10,ref. bib11 and activate the integrator-ref. bib12 or ZC3H4-WDR82 (restrictor) complexes,ref. bib13,ref. bib14,ref. bib15,ref. bib16 giving rise to pA− RNA. Given their essential roles in integral quality control processes that regulate genomic output and maintain transcriptome homeostasis, mechanisms connecting early transcription termination and transcript turnover remain important study objects.ref. bib17,ref. bib18
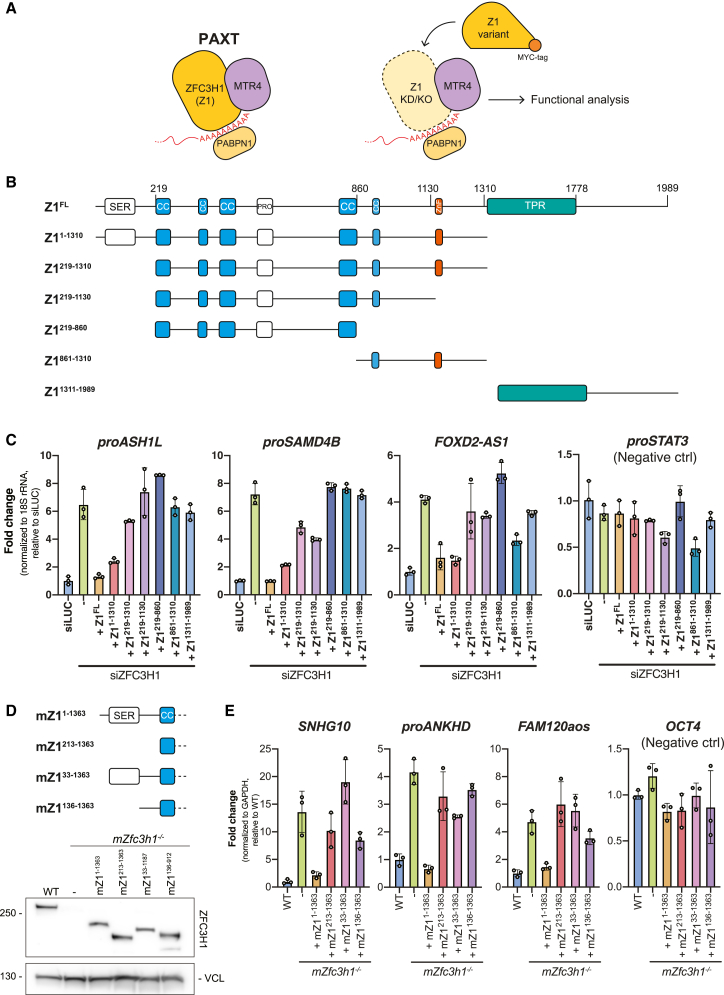
Nuclear RNA decay is primarily handled by the 3′-5′ exonucleolytic activity of the RNA exosome complex.ref. bib17,ref. bib19 To gain access to its RNAPII-derived substrates, the exosome-associated RNA helicase MTR4ref. bib20,ref. bib21,ref. bib22 can contact at least two distinct nucleoplasmic adaptors; the nuclear exosome targeting (NEXT) complexref. bib23 and the poly(A) exosome targeting (PAXT) connection,ref. bib24,ref. bib25 targeting pA− and pA+ transcripts, respectively.ref. bib3,ref. bib26 The NEXT complex, formed by a dimer of MTR4-ZCCHC8-RBM7 heterotrimers,ref. bib27,ref. bib28 can be recruited to short, TSS-proximal, pA− transcripts by connecting to the RNA-bound cap-binding complex (CBC) via the ARS2 and ZC3H18 proteins.ref. bib3,ref. bib29,ref. bib30 The PAXT connection, on the other hand, consists of a core MTR4-ZFC3H1 heterodimer that associates with the nuclear poly(A) binding protein (PABPN1) in an RNA-dependent manner (Figure 1A, left). In addition, less-well-described interactions with the RNA binding proteins ZC3H3, RBM26, or RBM27 are suggested to occur. Together, this facilitates the decay of a wide range of pA+ RNAs.ref. bib24,ref. bib25,ref. bib31,ref. bib32
The 5′ end of every RNAPII-produced transcript, regardless of its stability, is m7G cap modified and nascently bound by the CBC during the early stages of transcription.ref. bib33 The CBC in turn associates with ARS2, forming the trimeric CBCA complex,ref. bib29,ref. bib34,ref. bib35 which acts as a central hub for the competitive exchanges of RNA sorting factors, also termed “classifiers,”ref. bib36 that ultimately direct the RNA toward a productive or a destructive fate.ref. bib18 For example, CBCA can connect to factors, such as ALYREF, PHAX, or FLASH, to promote the cellular transport of mRNA, snRNA, or replication-dependent histone (RDH) RNA, respectively.ref. bib18,ref. bib36 Alternatively, CBCA can direct its bound transcripts toward RNA decay.ref. bib29 A key factor facilitating this is ZC3H18, which was shown to connect the CBCA and NEXT complexes,ref. bib29,ref. bib37 while directly competing with productive CBCA interactors such as PHAX.ref. bib38,ref. bib39 While the CBCA-ZC3H18-NEXT association was validated through protein domain mapping,ref. bib37 an analogous connection to PAXT was merely suggested based on the presence of ZC3H18 in PAXT interactome data but has lacked further experimental validation.ref. bib24 Indeed, PAXT activity was originally proposed to rely on prolonged RNA nuclear residence times as a means of purging nuclei of pA+ transcripts inefficiently managed by export factors, such as ALYREF.ref. bib40,ref. bib41,ref. bib42,ref. bib43 Hence, bona fide PAXT substrates were considered longer and more processed than their NEXT-sensitive counterparts.ref. bib24 More recent genome-wide analyses have, however, revealed that PAXT also targets short and unspliced RNAs which, apart from their pA+ 3′-end status, are biochemically reminiscent of NEXT substrates.ref. bib3,ref. bib26 Whether these TSS-proximal pA+ transcripts rely on the same recruitment mechanism as longer and processed PAXT substrates, and which role, if any, ZC3H18 plays in the process, remains unclear.
Here, we find that the N-terminal domain of ZFC3H1 is instrumental for PAXT targeting of short pA+ transcripts. This is achieved by its direct binding to ARS2 via a conserved acidic-rich short linear motif (SLiM), consistent with observations reported in ZFC3H1 homologs.ref. bib44,ref. bib45 Surprisingly, however, we find that ZC3H18, which promotes NEXT-mediated RNA decay, exhibits an inhibitory effect on the ARS2-dependent PAXT decay pathway. This inhibition is explained by a competition between ZFC3H1 and ZC3H18, both of which utilize a similar SLiM to bind a common surface on ARS2. We suggest that this intricate setup provides the possibility for increased NEXT activity to lift the ZC3H18 inhibition of PAXT, thereby providing for co-activation of the two pathways in situations where the demand for RNA turnover is high.
Results
The ZFC3H1 N terminus harbors important information for its function in RNA decay
ZFC3H1 is the central hub of PAXT, mediating direct contact with MTR4 and further associating with accessory PAXT componentsref. bib25,ref. bib46 (Figure 1A, left). To dissect the relative contributions of ZFC3H1 domains in RNA decay, we examined ZFC3H1 through the domain-predicted generation of six C-terminally MYC-tagged ZFC3H1 variants (Z1x) (Figure 1B). These truncated proteins, as well as their full-length (FL) counterpart, were individually and stably expressed in HeLa cells depleted of endogenous ZFC3H1 using UTR-specific siRNA (siZFC3H1) (Figure S1A). Possible functional complementation was then assessed by qRT-PCR using primers toward known PAXT substrates (Figure 1A, right), which revealed that only the Z1FL and Z11-1310 variants consistently repressed the upregulation of RNA levels induced by siZFC3H1 (Figure 1C). This activity was particularly significant for the Z11-1310 variant due to its relatively low expression (Figure S1A). In contrast, higher expression of the N-terminally truncated Z1219-1310 variant yielded only marginal activity, which immediately highlighted the N terminus (1–218 aa) of ZFC3H1 as being important for targeting of the tested substrates (Figure 1C). Due to the somewhat variable expression levels of the ZFC3H1 variants in HeLa cells (Figure S1A), we validated these results by introducing homologous mouse (mZ1x) constructs into our previously established ZFC3H1 knockout mouse embryonic stem cells (mESCs)ref. bib47 (Figure S1B). With a caveat that we could not achieve expression of mZ1FL or mZ11364-1992, we consistently observed that only the mZ11-1363 variant, containing the intact N terminus, was able to repress the upregulation of PAXT targets triggered by the Zfc3h1−/− conditions (Figure S1C). Using the mESC system, we further narrowed down the functional region by interrogating activities of two additional N-terminal truncations of mZ11-1363 (mZ133-1363 and mZ1136-1363, Figure 1D), which revealed that the first 33 aa of ZFC3H1 are important for its function (Figure 1E). From this, we conclude that the extreme N terminus of ZFC3H1 plays an important role in the decay of select PAXT-sensitive RNAs.
A conserved short linear motif connects ZFC3H1 to the CBC via ARS2
To further investigate the N terminus of ZFC3H1, we first addressed the conservation of this region by multiple sequence alignment analysis. This revealed tandem copies of a highly conserved and acidic SLiM within the first 33 aa of ZFC3H1 (Figure S2A). Interestingly, an identical motif was previously highlighted in the S. pombe ZFC3H1 homolog, spRed1, where it was linked to the binding of the spArs2 protein.ref. bib44 Moreover, while our study was under completion, such ARS2 binding was confirmed by in vitro assays using human proteins.ref. bib45 Inspired by this, we introduced point mutations into the SLiM of Z1FL (Figure 2A, upper left) and assessed their impact by expressing the mutant protein Z1FL(MUT) in HeLa cells depleted of endogenous ZFC3H1 (Figure 2A, upper right; Figure S2B). Like the N-terminal deletions, the Z1FL(MUT) construct was unable to repress RNA levels of selected PAXT substrates in siZFC3H1-treated cells (Figure 2A, lower).
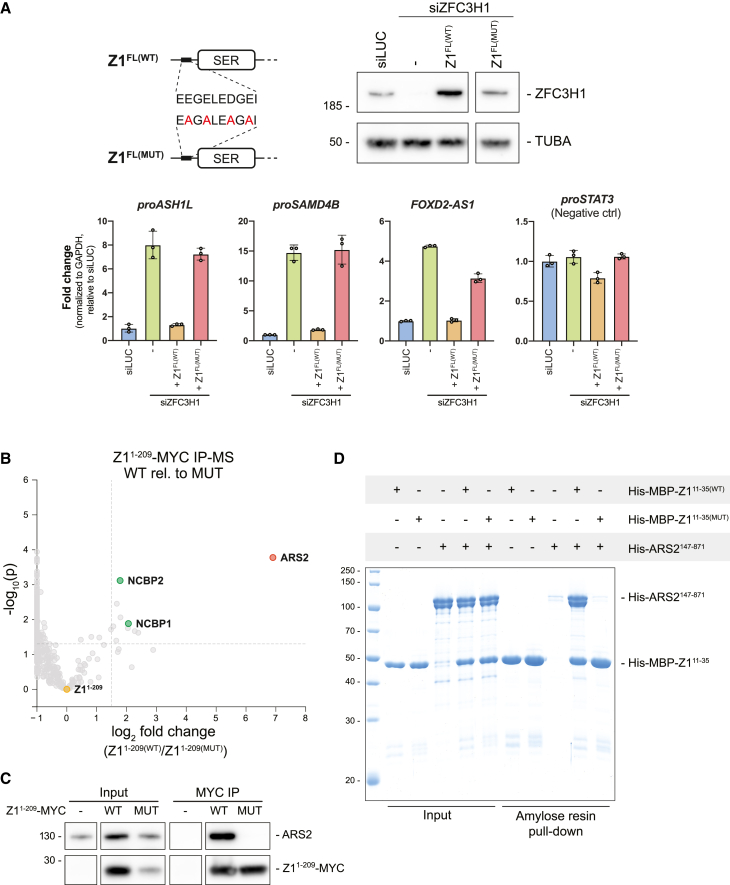
To interrogate the effect of these point mutations on the interactome of the ZFC3H1 N terminus, we generated wild-type (WT) (Z11-209(WT)) and mutant (Z11-209(MUT)) MYC-tagged variants and subjected them to immunoprecipitation (IP) followed by mass spectrometry (IP-MS). Plotting the relative enrichment of interacting proteins in Z11-209(WT) vs. Z11-209(MUT) IPs, we observed a significant enrichment of ARS2 and CBC components, NCBP1 and NCBP2 (Figure 2B). This interaction of ARS2 with the acidic-rich N terminus of ZFC3H1 was confirmed by IP-WB analysis (Figures 2C and S2C) and by in vitro pull-downs, demonstrating that the recombinant Z111-35(WT), but not Z111-35(MUT), directly interacts with ARS2147-871 (Figure 2D). To finally demonstrate that the conserved SLiM of ZFC3H1 is not only sufficient but also necessary for ARS2 binding, we performed GFP-ARS2 IPs from cells expressing either Z1FL(WT) or Z1FL(MUT). As expected, the Z1FL(MUT) protein did not co-immunoprecipitate (coIP) with ARS2 (Figure S2D). Consistent with previous studies, we therefore conclude that the conserved acidic SLiM of ZFC3H1 binds ARS2 directly and further that this interaction is functionally relevant.
The ZFC3H1-ARS2 connection facilitates the decay of short, adenylated transcripts
What might then be the mechanistic consequence of the ZFC3H1-ARS2 interaction, given that ZFC3H1/MTR4 was assumed to gain substrate access via the RNA pA tail and its associated PABPN1ref. bib24? Providing a clue, we recently reported that artificial tethering of ARS2 to a pA+ reporter RNA made PAXT-mediated decay more efficient.ref. bib48 Hence, we set out to determine the exact role of ARS2 in PAXT-mediated RNA decay by revisiting our previously published RNA sequencing (RNA-seq) data from siARS2-treated HeLa cells.ref. bib30 Utilizing a set of PAXT-sensitive targets, defined as upregulated upon depletion of ZFC3H1 and ZC3H3,ref. bib3 a total of 1,116 was split into 296 ARS2-dependent and 820 ARS2-independent substrates (Figure 3A). Stratifying these RNAs by their corresponding TU lengths revealed that ARS2-dependent PAXT targets were generally shorter (Figures 3A and S3A, left) and with fewer exons (Figure S3B, left) than their ARS2-independent counterparts. In this way, ARS2 depletion appeared to primarily affect shorter substrates as also seen for the NEXT pathway (Figures S3A and S3B, right). We note that this trend was, to a milder extent, also observed in the control set of PAXT-insensitive RNAs (Figure 3A, right), consistent with the idea that ARS2 impacts the metabolism of a wider variety of short RNAs.
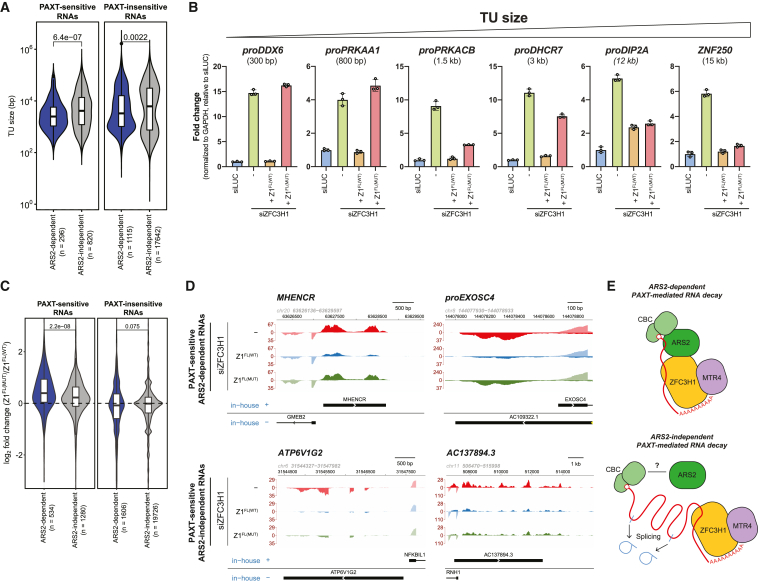
As ARS2 is a central RNA sorting factor,ref. bib36 its depletion will affect several aspects of RNA metabolism. We therefore investigated this size-based substrate trend by analyzing RNA isolated from siZFC3H1 HeLa cells complemented with either FL ZFC3H1 (Z1FL(WT)) or its mutated counterpart incapable of binding ARS2 (Z1FL(MUT)). As the qRT-PCR amplicons utilized in our earlier analysis of the Z1FL(MUT) construct (Figure 2A) were designed to detect short PAXT substrates, we here included amplicons detecting PAXT substrates of increasing lengths. This strategy revealed that, while the Z1FL(MUT) variant was unable to target short PAXT-sensitive transcripts, it was fully capable of repressing such longer RNAs (Figure 3B). We further analyzed the functionality of the Z1FL(MUT) construct in the decay of ARS2-independent substrates at a global scale by conducting triplicate RNA-seq experimentation of total RNA purified from the Z1FL(WT)– and Z1FL(MUT)-expressing siZFC3H1-treated cells (Figure S3C). As predicted, ARS2-dependent PAXT substrates were significantly upregulated upon Z1FL(MUT) relative to Z1FL(WT) expression, while a less pronounced effect was observed for ARS2-independent RNAs (Figures 3C and 3D). The latter might, at least partly, result from the lower expression levels of the Z1FL(MUT) protein (Figure 2A, upper right).
Because the CBCA complex could, in principle, suffice for recruitment of ZFC3H1 to ARS2-dependent substrates, we investigated the extent to which PABPN1 would also be required for the decay of these RNAs. To this end, we utilized previous siPABPN1 RNA-seq data,ref. bib24 which revealed that PABPN1 depletion results in the upregulation of both ARS2-dependent and -independent PAXT substrates (Figure S3D).
We conclude that at least two independent branches of PAXT-mediated RNA decay exist. One takes advantage of the CBCA connection to enhance access to shorter substrates, resembling polyadenylated versions of those targeted by the NEXT pathway (Figure 3E, top). The other branch targets longer and more spliced RNAs, possibly relying more on the pA tail for factor recruitment (Figure 3E, bottom).
ZC3H18 antagonizes the ARS2-dependent PAXT pathway
The physical link between the CBCA and NEXT complexes was reported to be bridged by the ZC3H18 protein (Figure 4A), the depletion of which results in the accumulation of NEXT substrates.ref. bib15,ref. bib29,ref. bib37 A comparable connection between ZC3H18 and ZFC3H1 was also speculated to link the CBCA to PAXTref. bib24; however, this idea was never further elaborated. Given these biochemical interactions and our finding that NEXT- and ARS2-dependent PAXT substrates are of largely similar lengths, we were prompted to investigate how ZC3H18 might impact ARS2-dependent PAXT activity.
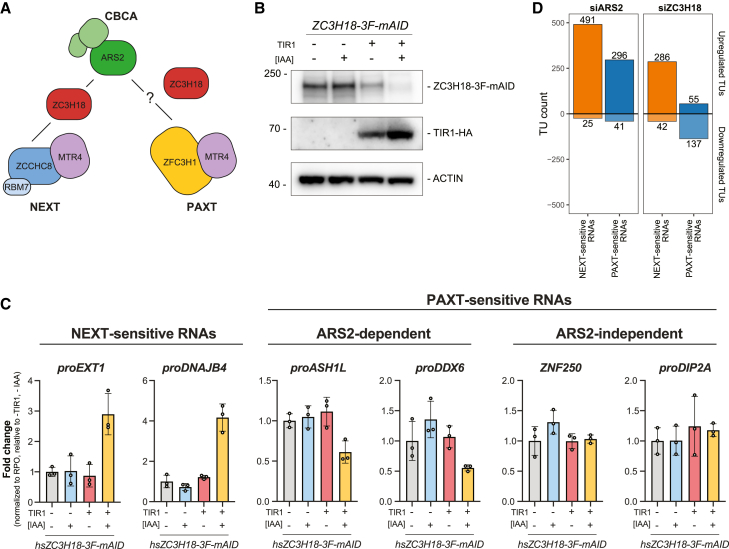
Our previous approach to study ZC3H18 function utilized siRNA-mediated factor depletion, which led to significant, albeit somewhat moderate increases of NEXT-sensitive RNAs in siZC3H18 conditions.ref. bib29,ref. bib30,ref. bib37 We therefore turned to a conditional degron-based technique to rapidly deplete endogenous ZC3H18 via a fused mini-auxin-inducible degron and 3×FLAG (3F-mAID) tag.ref. bib49 After 16 h of auxin (IAA) treatment, ZC3H18-3F-mAID protein levels were efficiently depleted (Figure 4B). Subsequent qRT-PCR assessment of selected NEXT-sensitive RNAs revealed their significant, and expected, upregulation (Figure 4C, left). In contrast, and to our surprise, levels of ARS2-dependent PAXT substrates were reduced upon ZC3H18 depletion (Figure 4C, middle). Thus, ZC3H18 may normally antagonize decay of these transcripts. Because levels of ARS2-independent PAXT substrates yielded no significant effect upon ZC3H18 depletion (Figure 4C, right), the inhibitory function of ZC3H18 might be enacted via ARS2. To validate these findings, we generated analogous Zc3h18-3F-mAID mESCs and included previously established Zcchc8-3F-mAID and Zfc3h1-3F-mAID linesref. bib50 to perform a comparative rapid factor depletion time course (Figure S4A). Consistent with our HeLa cell data, NEXT-sensitive RNAs (FigureS4B, top: proNUP85, proSMG5) were increasingly upregulated over the 8-h time course following depletion of ZC3H18 and ZCCHC8, while PAXT (ZFC3H1)-sensitive RNAs (Figure S4B, bottom: FAM120aos, proANKHD1) were reduced by ZC3H18 depletion.
To corroborate these diverse functions of ZC3H18 in NEXT- and PAXT-mediated RNA decay genome-wide, we revisited siZC3H18 RNA-seq dataref. bib30 and assessed changes in NEXT- or PAXT-sensitive transcripts in siARS2 vs. siZC3H18 conditions. In general, NEXT-sensitive RNAs were upregulated in both siARS2 and siZC3H18 conditions (Figure 4D, orange bar plots). PAXT-sensitive RNAs, on the other hand, were only preferentially upregulated in siARS2 conditions but predominantly downregulated upon ZC3H18 depletion (Figure 4D, blue bar plots). Thus, while ZC3H18 promotes NEXT activity it rather appears to inhibit the ARS2-dependent PAXT pathway.
ZFC3H1 and ZC3H18 compete for ARS2 binding
As ZC3H18 only antagonizes the ARS2-dependent branch of the PAXT pathway, we tested whether ZC3H18 might negatively affect ARS2-ZFC3H1 association by performing IPs of MYC-ARS2 stably integrated in HeLa following control of ZC3H18 levels by siRNA depletion. Consistent with its role in bridging the CBCA and NEXT complexes, ZC3H18 depletion led to reduced ARS2-ZCCHC8 association (Figures 5A and S5A). Conversely, levels of ZFC3H1 were increased in ARS2 IPs when ZC3H18 was depleted. The observed downregulation of PAXT-sensitive RNAs in this condition (Figure 4C) was therefore likely to be due to the reinforced interaction between ARS2 and ZFC3H1, which in turn enhances ARS2-dependent PAXT activity.
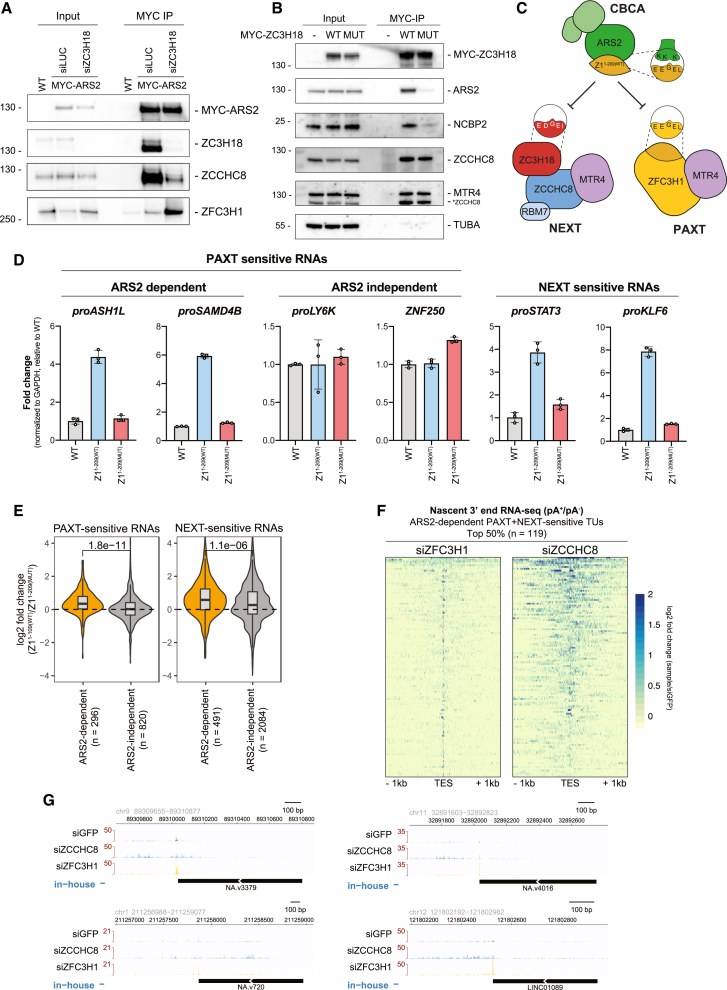
In line with the possibility that ZC3H18 and ZFC3H1 might compete for binding to the same region of ARS2, sequence analysis of ZC3H18 revealed three copies of a ZFC3H1-like acidic SLiM in a highly conserved region of the protein (Figure S5B). To examine whether this motif indeed contributes to the ZC3H18-ARS2 interaction, we introduced diagnostic point mutations into this putative ZC3H18 acidic SLiM (ZC3H18MUT) analogous to the mutations of Z1FL(MUT) (Figures 2A and S5B). We then stably integrated MYC-tagged ZC3H18WT and ZC3H18MUT variants into HeLa cells and performed MYC IPs. Both ZC3H18WT and ZC3H18MUT constructs were expressed at comparable levels and co-immunoprecipitated ZCCHC8 and MTR4 with similar efficiencies (Figure 5B), demonstrating that the introduced changes did not affect NEXT binding. However, in contrast, ARS2 and NCBP2 levels were both significantly reduced in the ZC3H18MUT IP, confirming that ZC3H18 and ZFC3H1 utilize similar motifs to interact with ARS2. Moreover, like the ZFC3H1 acidic SLiM, the ZC3H18 SLiM was also sufficient to bind ARS2 in vitro in a WT-specific manner (Figure S5Cref. bib15). We note that a previous study reported a coiled-coil ZC3H18 domain outside of this conserved region to contribute to ARS2 binding,ref. bib37 the relevance of which now remains to be further investigated. Finally, we performed in vitro pull-downs of WT SLiM containing ZFC3H1 and ZC3H18 peptides in combination with either a WT ARS2147-871 peptide or a ZnF domain mutant (K719A, K722A, K734A), previously shown to disrupt binding between ARS2 and effector proteins.ref. bib15,ref. bib45,ref. bib48,ref. bib51 Here, we reassuringly confirmed that ZFC3H1 and ZC3H18 SLiM peptides both bound to WT ARS2, while these interactions were lost with the ZnF mutant (Figure S5D). We conclude that both ZC3H18 and ZFC3H1 can directly interact with the ZnF/effector domain of ARS2 via their acidic SLiM motifs.
Given the competition between ZFC3H1 and ZC3H18 for ARS2 binding, one might expect that decreased ZFC3H1 levels would consequently affect NEXT activity. However, ZFC3H1 depletion did not significantly affect NEXT-sensitive transcripts,ref. bib3,ref. bib24 possibly because ZFC3H1 protein is only approximately half as abundant as ZC3H18 in HeLa cells.ref. bib52 We therefore turned to overexpression of the Z11-209(WT) variant containing the ARS2-binding motif. As Z11-209(WT) cannot form the full PAXT connection, we assumed that its overexpression might inhibit both NEXT- and PAXT-mediated RNA decay pathways by saturating the corresponding binding site on ARS2 (Figure 5C). Hence, we performed MYC-ARS2 IPs following DOX-induced overexpression of the Z11-209(WT) variant, which significantly reduced the association of ZC3H18 and by extension ZCCHC8 with ARS2 (Figure S5E). Furthermore, this also reduced the association of endogenous ZFC3H1 with MYC-ARS2, thereby reducing both NEXT and PAXT proteins on ARS2. To validate the functional consequences of this, we analyzed RNA from equivalent samples while also utilizing the Z11-209(MUT), deficient for ARS2 binding (Figures 2C and 2D), as a negative control. As expected, both NEXT-sensitive and ARS2-dependent PAXT-sensitive RNAs were upregulated upon overexpression of Z11-209(WT), while Z11-209(MUT) overexpression showed no significant substrate changes (Figure 5D). In addition, ARS2-independent PAXT substrates remained unaffected, supporting the idea that the observed phenotype was due to the blocking of the common ZFC3H1/ZC3H18 interaction surface on ARS2. To obtain a global view, we further performed RNA-seq analysis of the same samples (Figure S3C), which confirmed a Z11-209(WT) overexpression-specific upregulation of both NEXT- and PAXT-sensitive ARS2-dependent RNAs, while no significant changes were observed for ARS2-independent substrates (Figure 5E).
At first glance, it seemed counterintuitive for two RNA decay pathways to compete for ARS2-bound RNAs. However, because PAXT and NEXT target distinct substrate species (pA+ vs. pA− RNAs, respectively), this could possibly be rationalized by individual TUs expressing transcripts sensitive to both pathways.ref. bib3 To provide a visual demonstration of this phenomenon, we stratified TUs sensitive to both PAXT and NEXT, adding a further criteria of ARS2 sensitivity (n = 238). Transcript production from this TU selection was then analyzed using our previously published 3′-end 4-thiouridine RNA-seq data from HeLa cells depleted of ZFC3H1 or ZCCHC8.ref. bib3 To capture both non-adenylated and adenylated transcript 3′ ends (pA−/+), we utilized libraries generated from RNAs following their in vitro pA.ref. bib53 3′-end RNA-seq signals from siZFC3H1 and siZCCHC8 relative to control samples falling within a 2-kb region surrounding annotated transcript end sites (TESs) were then defined, and the top 50% ARS2-dependent PAXT- and NEXT-sensitive TUs were visualized (Figure 5F). This representation, as well as selected genome browser views (Figure 5G), highlighted the ample presence of dual PAXT- and NEXT-sensitive TUs, which were largely driven by ZFC3H1-sensitive 3′ ends at annotated TESs embedded in more heterogeneous ZCCHC8-sensitive 3′ ends.
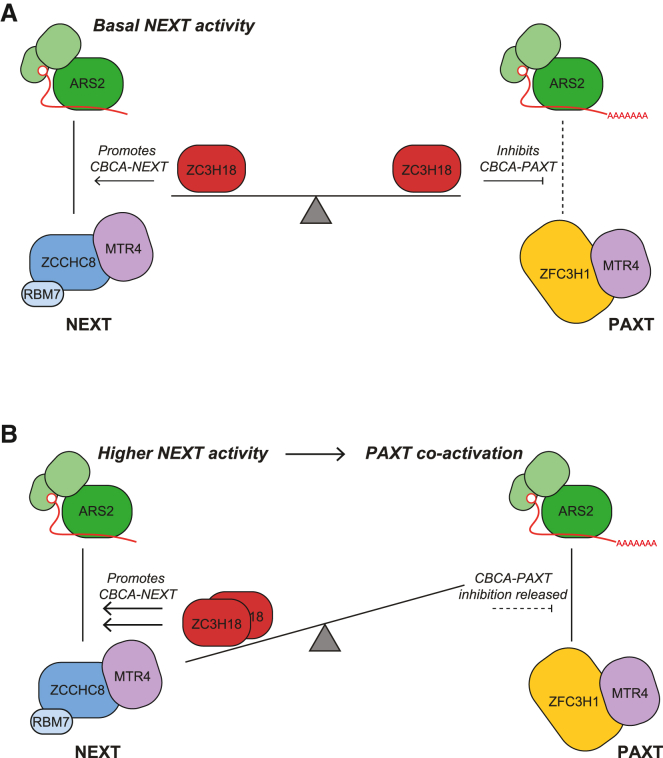
Having ZC3H18 inhibit the ARS2-dependent PAXT pathway would be detrimental for the efficient removal of transcripts produced from such dually sensitive loci. We therefore suggest that demanding engagement of ZC3H18 with NEXT will locally prevent its PAXT-inhibitory function, allowing for co-activation of the PAXT-ARS2 pathway to degrade pA+ transcripts generated from the same TU (Figure 6, see discussion).
Discussion
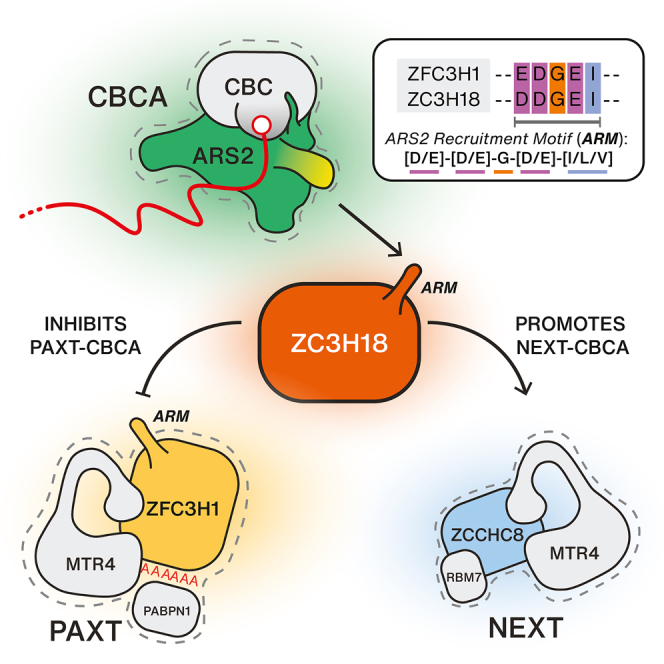
The PAXT connection targets a variety of nuclear pA+ RNAs. At one end, competition between PAXT and nuclear export factorsref. bib42,ref. bib43,ref. bib54 has led to a proposed nuclear timer model in which PAXT assembles on pA+ RNA with a prolonged nuclear residence time due to inefficient export.ref. bib18,ref. bib40,ref. bib41 Now, we show that PAXT targeting can be enhanced by binding of the core PAXT protein ZFC3H1 to the CBC-associated protein ARS2, providing a more direct response to the production of early terminated and short pA+ RNAs. Remarkably, this distinct branch of the PAXT pathway is antagonized by the NEXT-associated ZC3H18 protein via an unprecedented crosstalk between the two RNA decay factors ZFC3H1 and ZC3H18 on CBCA-bound RNAs. While the opposing roles of ZC3H18 in PAXT- and NEXT-mediated RNA decay may at first appear contradictory, we propose that it provides for possible co-activation of both pathways in demanding situations with larger volumes of early terminated RNAs.
Although previous studies of PAXT-mediated RNA decay have acknowledged the existence of a link between CBCA and PAXT, its importance was not subjected to critical scrutiny due to a primary focus on PAXT recruitment via the RNA pA+ tail.ref. bib24,ref. bib25,ref. bib46 However, an artificial PAXT-sensitive RNA reporter system showed a clear requirement for ARS2, implying a role of CBCA in PAXT-mediated RNA decay, at least in some situations.ref. bib48 Moreover, the S. pombe CBCA homolog was reported to form a submodule of the Mtl1-Red1 core complex, homologous to the MTR4-ZFC3H1 core of the PAXT connection,ref. bib55 linked via an interaction between spArs2 and spRed1.ref. bib44,ref. bib45 Indeed, we confirm here that a group of PAXT-sensitive RNAs display a dependency on ARS2 for their degradation. While we find no categorical difference in biochemical features between ARS2-dependent and -independent PAXT substrates, the former tends to be shorter, less spliced, and more “nascent-like.” This is fitting with the concept of ARS2 being crucial for fate decisions of RNAs produced within a promoter-proximal range of a few kilobases,ref. bib15,ref. bib30 which is exemplified by its role in the biogenesis of short functional snRNAs and RDH RNAs,ref. bib29,ref. bib30,ref. bib34,ref. bib56,ref. bib57 the termination of short, unstable RNAs,ref. bib15,ref. bib30 and the decay of unstable promoter-proximal transcripts via NEXTref. bib3,ref. bib15,ref. bib24,ref. bib30 or PAXT.ref. bib3 Any role of ARS2 in the metabolism of longer and more processed transcripts remains elusive. We therefore suggest that these depend on features/factors that become more influential once an RNA “moves out” of the ARS2-dependent phase and becomes more of a subject to post-transcriptional processing steps and the assembly into export-competent RNPs. Non-optimal transcripts in this category that fail in any of these productive steps may still fall prey to PAXT-mediated decayref. bib18 but are not dependent on ARS2 (Figure 3E).
As a biochemical account of the ability of ARS2 to be involved in promotor-proximal RNA metabolism, the CBCA provides a central hub capable of competitive exchanges of multiple RNA sorting factors. Binary competition between ARS2 interactors has been widely reported such as ZC3H18 and PHAX,ref. bib38 ZC3H18 and NCBP3,ref. bib39 and PHAX and NCBP3.ref. bib51 The basis for this competition was shown to be mediated by a common binding surface on ARS2, the lysine-rich surface of the ZnF domain,ref. bib15,ref. bib45,ref. bib48,ref. bib51 which we coin as the “effector” domain. In agreement with recent studies,ref. bib44,ref. bib45 we demonstrate here that a highly conserved acidic-rich SLiM in ZFC3H1 is responsible for the direct binding of ARS2. Refined searches of this SLiM consensus ([DE]-[DE]-G-[DE]-[ILV]) have revealed its presence in several other ARS2-interacting proteins involved in nuclear RNA metabolism pathwaysref. bib15,ref. bib44,ref. bib45 and the introduction of diagnostic mutations disrupts their interactions with ARS2. We therefore coin this SLiM motif consensus as an “ARS2 recruitment motif” or ARM. Together, this implies that ZFC3H1 impacts early CBCA-mediated nuclear RNA fate decisions via competition for ARS2 binding with other nuclear RNA sorting factors. Previous data support the notion that these factors do not show preference for specific RNA biotypes during nascent stages of transcription but rather that they dynamically sample and exchange on the CBCA until additional cues are sensed to form a signature that ultimately settles the fate of the transcript.ref. bib18,ref. bib38 The close proximity of a pA+ tail to the CBCA, along with the association of PABPN1, may be such a cue to tip the scales of competition in favor of the CBCA-PAXT connection, explaining why ARS2-dependent PAXT substrates tend to be short. On the other hand, the presence of introns might result in a preferential recruitment of productive factors to the nascent RNP, thus leaning spliced PAXT substrates toward the ARS2-independent group.
While factor exchange on the ARS2 effector domain was considered a productive vs. destructive RNP concept, we report here the surprising observation that two ARS2-interacting factors, ZFC3H1 and ZC3H18, targeting transcripts for PAXT- and NEXT-mediated degradation, respectively, also compete between themselves. Moreover, we demonstrate that this competition has unprecedented functional consequences for nuclear RNA decay. Based on our findings, we propose a model where basal NEXT activity, under standard conditions, leaves nuclear amounts of ZC3H18 and NEXT components in excess. This, in turn, allows for ZC3H18 molecules, unoccupied by NEXT, to dampen the CBCA-PAXT connection via competitive ARS2 binding (Figure 6A), which is likely possible due to the relatively low nuclear levels of ZFC3H1 compared with ZC3H18 and ZCCHC8.ref. bib52 This inhibition may then be lifted in physiological conditions, or in certain sub-compartments of the nucleus, that demand intense NEXT activity, for example, at loci, yielding a higher load of early terminated transcripts (Figure 6B). Because a large number of TUs produce both NEXT- and PAXT-sensitive transcripts,ref. bib3 an increased production of NEXT substrates is naturally predicted to coincide with a parallel increase in short PAXT-sensitive RNAs.
Our appreciation of the richness of RNA transactions taking place during the early phases of RNAPII transcription has grown dramatically in recent years, and we are now beginning to understand the biochemical underpinnings of the competition/crosstalk occurring between differently involved pathways. While there are most certainly additional biochemical connections to be delineated, future efforts will also start to address how these are orchestrated inside the cell nucleus to achieve sufficient processing and transport of the needed short RNA biotypes while preventing spurious transcripts to overwhelm such production.
Limitations of the study
Some aspects of this study were based on siRNA-mediated depletion of factors in human cells over 2–3 days. These long-term depletion experiments could be associated with both secondary and compensatory effects. To remedy this, we have complemented our analyses, where possible, using rapid depletion systems or providing orthogonal data in mESCs to support our conclusions.
As ARS2 interacts with a multitude of RNA sorting factors in a mutually exclusive manner, any perturbations of acidic SLiM/ARM-containing proteins through depletion/overexpression may disrupt the balance of ARS2 complexes. In this study, we only address the impact on ARS2-dependent RNA decay via ZFC3H1 and ZC3H18 through their perturbations. Any effects on reported ARS2-dependent RNA processingref. bib34,ref. bib57 or transcription termination activitiesref. bib15,ref. bib16 were outside the scope of this manuscript.
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| Mouse monoclonal anti-ACTIN | Sigma-Aldrich | Cat# A2228; RRID: RRID:AB_476697 |
| Mouse monoclonal anti-FLAG M2 | Sigma-Aldrich | Cat# F1804: RRID:AB_262044 |
| Rat monoclonal anti-HA | Sigma-Aldrich | Cat# 11867423001; RRID:AB_390918 |
| Rabbit polyclonal anti-MTR4 (SKIV2L2) | Abcam | Cat# ab70551; RRID:AB_1270701 |
| Rabbit monoclonal anti-MYC | Cell Signaling | Cat# 2278; RRID:AB_490778 |
| Rabbit polyclonal anti-ALPHA-TUBULIN | Rockland | Cat# 600-401-880; RRID:AB_2612816 |
| Mouse monoclonal anti-VINCULIN (VCL) | Sigma-Aldrich | Cat# V9131; RRID:AB_477629 |
| Rabbit polyclonal anti-ZC3H18 | Sigma-Aldrich | Cat# HPA040847; RRID:AB_10794865 |
| Mouse polyclonal anti-ZCCHC8 | Abcam | Cat# ab68739; RRID:AB_1271512 |
| Rabbit polyclonal anti-ZCCHC8 | Bethyl | Cat# A301-806A; RRID:AB_1233063 |
| Rabbit polyclonal anti-ZCCHC8 | Novus Biologicals | Cat# NB10094995; RRID: AB_1262274 |
| Rabbit polyclonal anti-ZFC3H1 | Sigma-Aldrich | Cat# HPA007151; RRID:AB_1846133 |
| Rabbit polyclonal anti-ARS2 | GeneTex | Cat# GTX119872; RRID: AB_10720168 |
| Mouse monoclonal anti-GFP (B-2) | Santa Cruz | Cat# sc-9996; RRID:AB_627695 |
| Goat polyclonal anti-rabbit immunoglobulins/HRP | Agilent | Cat# P0448; RRID:AB_2617138 |
| Goat polyclonal anti-mouse immunoglobulins/HRP | Agilent | Cat# P0447; RRID:AB_2617137 |
| Rabbit polyclonal anti-rat immunoglobulins/HRP | Agilent | Cat# P0450; RRID:AB_2630354 |
| Bacterial and virus strains | ||
| DH5-alpha competent E.coli strain | N/A | N/A |
| BL21 Star (DE3) pRARE E.coli strain | EMBL Heidelberg Core Facility | N/A |
| Chemicals, peptides, and recombinant proteins | ||
| GSK3 inhibitor (CHIR99021) | Sigma-Aldrich | Cat# SML1046 |
| MEK1/2 inhibitor (PD0325901) | Sigma-Aldrich | Cat# PZ0162 |
| Indole-3-acetic acid sodium salt (IAA) | Sigma-Aldrich | Cat# I5148-10G |
| TRIzol | Thermo Fisher | Cat# 15596026 |
| Critical commercial assays | ||
| siLentFect | BioRad | Cat# 1703362 |
| Lipofectamine 3000 Transfection Reagent | Thermo Fisher | Cat# L300001 |
| Viafect Transfection Reagent | Promega | Cat# E4981 |
| TURBO DNase kit | Thermo Fisher | Cat# AM2238 |
| SuperScript III Reverse Transcriptase | Thermo Fisher | Cat# 1808044 |
| Platinum SYBR Green qPCR SuperMix | Thermo Fisher | Cat# 11733046 |
| Benzonase nuclease | Millipore | Cat# 70746 |
| Benzonase nuclease | Merck | Cat# 71206 |
| Bioanalyzer RNA 6000 Nano Kit | Agilent | Cat# NC1783726 |
| Ribolock RNase Inhibitor | Thermo Fisher | Cat# EO0381 |
| cOmplete, Mini, EDTA-free Protease Inhibitor Cocktail | Roche | Cat# 04693159001 |
| NEBuilder HiFi DNA Assembly cloning kit | NEB | Cat# E5520S |
| GeneJET PCR Purification Kit | Thermo Fisher | Cat# K0701 |
| Purelink Genomic DNA mini purification Kit | Thermo Fisher | Cat# K182001 |
| SuperSignal West Femto Maximum Sensitivity Substrate | Thermo Fisher | Cat# 34096 |
| Protein G Dynabeads | Thermo Fisher | Cat# 10009D |
| Pierce Anti-c-Myc Magnetic Beads | Thermo Fisher | Cat #88842 |
| ChromoTek GFP-Trap Magnetic Particles M-270 | Proteintech | Cat #gtdk |
| NuPAGE LDS Sample Buffer (4x) | Thermo Fisher | Cat# NP0007 |
| NuPAGE Sample Reducing Agent (10x) | Thermo Fisher | Cat# NP0004 |
| Amylose Resin | NEB | Cat# E8021 |
| Superdex 75 Increase 10/300 GL | Cytiva | Cat# 29148721 |
| HiTrap Heparin HP | Cytiva | Cat# 17040601 |
| HIS-Select Nickel Affinity Gel | Millipore | Cat# P6611 |
| MagReSyn HILIC beads | Resyn Biosciences | Cat# MR-HLC002 |
| Deposited data | ||
| RNA-seq data | This study | GEO: GSE212557 |
| IP-MS data | This study | PRIDE: PXD045842 |
| Experimental models: Cell lines | ||
| HeLa Kyoto | Anthony A. Hyman | N/A |
| HeLa Kyoto ZFC3H1FL(WT)-MYC | This study | N/A |
| HeLa Kyoto ZFC3H1FL(MUT)-MYC | This study | N/A |
| HeLa Kyoto ZFC3H11−1310-MYC | This study | N/A |
| HeLa Kyoto ZFC3H1219−1310-MYC | This study | N/A |
| HeLa Kyoto ZFC3H1219−1130-MYC | This study | N/A |
| HeLa Kyoto ZFC3H1219−860-MYC | This study | N/A |
| HeLa Kyoto ZFC3H1861−1310-MYC | This study | N/A |
| HeLa Kyoto ZFC3H11311−1989-MYC | This study | N/A |
| HeLa Kyoto ZFC3H11−209(WT)-MYC | This study | N/A |
| HeLa Kyoto ZFC3H11−209(MUT)-MYC | This study | N/A |
| HeLa Kyoto MYC-ZC3H18WT | This study | N/A |
| HeLa Kyoto MYC-ZC3H18MUT | This study | N/A |
| HeLa Kyoto MYC-ARS2 | This study | N/A |
| HeLa Kyoto ZC3H18-3F-mAID | This study | N/A |
| HeLa Kyoto ZC3H18-3F-mAID OsTIR1-HA | This study | N/A |
| Mouse ES-E14TG2A | ATCC | Cat# CRL-1821; RRID:CVCL_9108 |
| Mouse ES-E14TG2A Zfc3h1−/− #1 | Garland et al.ref. bib47 | N/A |
| Mouse ES-E14TG2A Zfc3h1−/− MYC-mZ11−1363 | This study | N/A |
| Mouse ES-E14TG2A Zfc3h1−/− MYC-mZ1213−1363 | This study | N/A |
| Mouse ES-E14TG2A Zfc3h1−/− MYC-mZ1213−1187 | This study | N/A |
| Mouse ES-E14TG2A Zfc3h1−/− MYC-mZ1213−612 | This study | N/A |
| Mouse ES-E14TG2A Zfc3h1−/− MYC-mZ1913−1363 | This study | N/A |
| Mouse ES-E14TG2A Zfc3h1−/− MYC-mZ133−1363 | This study | N/A |
| Mouse ES-E14TG2A Zfc3h1−/− MYC-mZ1136−1363 | This study | N/A |
| Mouse ES-E14TG2A OsTIR1-HA | Garland et al.ref. bib50 | N/A |
| Mouse ES-E14TG2A OsTIR1-HA Zcchc8-3F-mAID | Garland et al.ref. bib50 | N/A |
| Mouse ES-E14TG2A OsTIR1-HA Zfc3h1-3F-mAID | Garland et al.ref. bib50 | N/A |
| Mouse ES-E14TG2A OsTIR1-HA Zc3h18-3F-mAID | This study | N/A |
| Oligonucleotides | ||
| siRNAs oligonucleotides | See Table S1 | N/A |
| sgRNA oligonucleotides | See Table S2 | N/A |
| qRT-PCR oligonucleotides | See Table S3 | N/A |
| Recombinant DNA | ||
| pB/TO[ZFC3H1FL(WT)-MYC] HYG | This study | N/A |
| pB/TO[ZFC3H11FL(MUT)-MYC] HYG | This study | N/A |
| pB/TO[ZFC3H11−1310-MYC] HYG | This study | N/A |
| pB/TO[ZFC3H1219−1310-MYC] HYG | This study | N/A |
| pB/TO[ZFC3H1219−1130-MYC] HYG | This study | N/A |
| pB/TO[ZFC3H1219−860-MYC] HYG | This study | N/A |
| pB/TO[ZFC3H1861−1310-MYC] HYG | This study | N/A |
| pB/TO[ZFC3H11311−1989-MYC] HYG | This study | N/A |
| pB/TO[ZFC3H11−209(WT)-MYC] HYG | This study | N/A |
| pB/TO[ZFC3H11−209(MUT)-MYC] HYG | This study | N/A |
| pB/TO[MYC-ZC3H18(WT)] HYG | This study | N/A |
| pB/TO[MYC-ZC3H18(MUT)] HYG | This study | N/A |
| pB[MYC-mZFC3H11−1363] BLAST | This study | N/A |
| pB[MYC-mZFC3H1213−1363] BLAST | This study | N/A |
| pB[MYC-mZFC3H1213−1187] BLAST | This study | N/A |
| pB[MYC-mZFC3H1213−612] BLAST | This study | N/A |
| pB[MYC-mZFC3H1913−1363] BLAST | This study | N/A |
| pB[MYC-mZFC3H133−1363] BLAST | This study | N/A |
| pB[MYC-mZFC3H1136−1363] BLAST | This study | N/A |
| pB[OsTIR1-HA] ZEOCIN | Garland et al.ref. bib50 | N/A |
| pGCT[hZC3H18-3F-mAID] HYG | This study | N/A |
| pGCT[hZC3H18-3F-mAID] NEO | This study | N/A |
| pGCT[mZC3H18-3F-mAID] HYG | This study | N/A |
| pGCT[mZC3H18-3F-mAID] NEO | This study | N/A |
| pB[MYC-ARS2] BLAST | This study | N/A |
| pCDNA5-GFP-ARS2 | Melko et al.ref. bib48 | N/A |
| pcDNA5-ZFC3H1-3F | Meola et al.ref. bib24 | N/A |
| Software and algorithms | ||
| R | N/A | N/A |
| RStudio | N/A | N/A |
| CHOPCHOP (v3) | Labun et al.ref. bib58 | https://chopchop.cbu.uib.no/ |
| HISAT2 (v2.2.1) | Kim et al.ref. bib59 | https://github.com/DaehwanKimLab/hisat2 |
| DEseq2 (v1.32.0) | Love et al.ref. bib60 | https://bioconductor.org/packages/release/bioc/html/DESeq2 |
| seqNdisplayR | N/A | https://github.com/THJlab/seqNdisplayR/ |
| ImageJ (v1.51) | Schneider, Rasband, and Eliceiriref. bib61 | https://ImageJ.nih.gov/ij/ |
| AriaMx (v1.71) | Agilent | https://www.agilent.com/ |
| Graphpad Prism (9.0.0) | Graphpad | https://www.graphpad.com/scientific-software/prism/ |
| Aminode | Chang et al.ref. bib62 | http://www.aminode.org/search |
Resource availability
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the lead contact, Torben Heick Jensen (thj@mbg.au.dk).
Materials availability
Resources and reagents used or generated in this study will be available upon request to the lead contact.
Experimental model and subject participant details
HeLa cell culture and transfections
HeLa Kyoto cells and descended cell lines were cultured in DMEM, GlutaMAX (Gibco) supplemented with 1× Pen-Strep (Gibco) and 10% FBS (Gibco) at 37°C, 5% CO2. Cells were passaged every 3–4 days by aspirating medium, dissociating cells with 0.05% Trypsin-EDTA (Gibco) briefly at 37°C, resuspending in culture medium and plating an appropriate amount of cell suspension. Cell lines were seeded in 6-well plates with culture medium without Pen-Strep prior to transfection with plasmid DNA using Lipofectamine 3000 (Invitrogen) or with 20 nm siRNA using siLentFect (Bio-Rad) and RPMI 1640 medium (Gibco), according to the manufacturers’ instructions. A list of siRNAs used in this study are included in Table S1. Cells were incubated for 72 h following siRNA transfection to achieve target protein depletions. For antibiotic selection, Hygromycin B (Invitrogen) was used at 200 μg/mL, Zeocin (Invitrogen) was used at 100 μg/mL. For mAID-tagged cell lines, 750 μM Indole-3-acetic acid sodium salt (IAA, Sigma) was supplemented to the medium and cells were incubated for the indicated timepoints before harvest. For transient expression of ZFC3H1 and ARS2, pcDNA5-ZFC3H1-3×FLAGref. bib24 and pcDNA5-GFP-ARS2ref. bib48 plasmids were transfected using Lipofectamine 3000 (Invitrogen), according to the manufacturer’s instructions.
mES cell culture and transfections
E14TG2a mES cells (male genotype, XY) and descended cell lines were cultured on 0.2% gelatin coated plates in 2i/LIF containing medium (1:1 mix of DMEM/F12 (Gibco) and Neurobasal (Gibco) supplemented with 1× Pen-Strep (Gibco), 2 μM Glutamax, 50 μM b-mercaptoethanol (Gibco), 0.1 mM Non-Essential Amino Acids (Gibco), 1 mM Sodium Pyruvate (Gibco), 0.5× N2 Supplement (Gibco), 0.5× B27 Supplement (Gibco), 3 μM GSK3-inhibitor (CHIR99021), 1 μM MEK-inhibitor (PD0325901) and Leukemia Inhibitory Factor (LIF, produced in house)) at 37°C, 5% CO2. Cells were passaged every 2–3 days by aspirating medium, dissociated cells with 0.05% Trypsin-EDTA (Gibco) briefly at 37°C before the addition of an equal volume of 1× Trypsin Inhibitor (Sigma) and gentle disruption by pipetting. Cells were pelleted by centrifugation to remove trypsin before resuspending in 2i/LIF medium and plating ∼8×104 cells/ml. Cell lines were transfected with single plasmids using Viafect (Promega) or multiple plasmids using Lipofectamine 3000 (Invitrogen) in 6 well plates according to the manufacturer’s instructions. For antibiotic selection, Blasticidin (BSD) was used at 10 μg/mL, Hygromycin B was used at 100 μg/mL, Genetecin was used at 250 μg/mL. For depletions in mAID-tagged cell lines, 750 μM Indole-3-acetic acid sodium salt (IAA, Sigma) was supplemented to the medium and cells were incubated for the indicated timepoints before harvest. A full list of cell lines used or generated in this study is found in the key resources table.
Method details
CRISPR/Cas9 knock-out/in cells
The generation of Zfc3h1−/− KO mES cell lines was described in.ref. bib47 CRISPR/Cas9 mediated genomic knock-ins of C-terminal 3xFLAG(3F) mini-AID (mAID) tags were carried out using pGolden (pGCT) homology dependent repair (HDR) donor vectors.ref. bib26,ref. bib50 The generation of Zcchc8-3F-mAID and Zfc3h1-3F-mAID ES cells was described in,ref. bib50 and Zc3h18-3F-mAID ES cell lines were generated using the same approach. Single guide RNAs (sgRNAs) were designed using the CHOPCHOP tool (v3)ref. bib58 and cloned into the pSLCas(BB)2A-PURO vector (pX459 Addgene plasmid ID: #48139) as previously describedref. bib64 (Table S3). OsTIR1-expressing ES cells were co-transfected using Lipofectamine 3000 (Thermo) with 2 donor plasmids harboring distinct selection markers (HYG/NEO) along with a sgRNA/Cas9 vector in a 1:1:1 ratio. Colonies were maintained under HYG/NEO double selection for the donor plasmid markers to increase the likelihood of homozygous knock-in cells. Single cell clones, that survived the selection process, were expanded before screening by western blotting analysis and confirmed by sanger sequencing of the targeted locus.
cDNA cloning and exogenous expression of ZFC3H1
Human ZFC3H1 and ZC3H18 cDNA constructs were cloned using full-length cDNA plasmids (pcDNA5-ZFC3H1-3×FLAG,ref. bib24 pcDNA5-ZC3H18-3×FLAGref. bib29) as a template. Fragments were amplified by Phusion PCR (NEB), using standard conditions, and cloned into DOX-inducible Tet-On (TO) piggyBAC (pB) vectors harboring a C- or N-terminal MYC-tag and Hygromycin B (HYG) selection marker using NEBuilder HiFi DNA assembly (NEB). HeLa Kyoto cells were transfected with pB/TO-Z1x-MYC or pB/TO-ZC3H18x-MYC vectors along with a pB transposase expressing vector (pBase) and a reverse tetracycline transactivator expressing pB vector containing Zeocin selection marker (pB-rtTA). Cell pools were selected using Hygromycin B and Zeocin until negative control cells no longer survived (7–14 days). For induction of expression constructs, cells were incubated for 1–2 days in culture medium supplemented with 1 μg/mL Doxycycline (Sigma-Aldrich) before harvest. Expression was validated by WB analysis using antibodies against MYC, ZFC3H1 or ZC3H18.
mZFC3H1 cDNA constructs were cloned, using a full-length cDNA plasmid as a template (pCR-XL-TOPO[mZFC3H1], Transomic Technologies), into a piggyBAC (pB) vector containing an N-terminal MYC tag and BSD selection marker using NEBbuilder HiFi DNA assembly (NEB). Zfc3h1−/− mES cells were transfected with pB-MYC-mZFC3H1x-BSD vectors along with a piggyBAC transposase expressing vector (pBase) in a 1:1 ratio using Viafect (Promega). Cell pools were selected with BSD for ∼ 7–10 days or until negative control cells no longer survived. Expression of constructs were validated by western blotting analysis using MYC antibodies. All generated constructs used and generated in this study are listed in the key resources table.
RNA isolation and qRT-PCR analysis
Total RNA was isolated using TRIzol reagent (Invitrogen) according to the manufacturer’s instructions. Extracted RNA was treated with TURBO DNase (Invitrogen) according to the manufacturer’s instructions followed by cDNA synthesis from 2 μg RNA using SuperScript III reverse transcriptase (Invitrogen) and a mixture of 80 pmol random primers (Invitrogen) and 20 pmol oligo d(T)20VN (Merck). RNA reactions were performed in the presence of Ribolock RNase inhibitor (Thermo). qPCR was performed using Platinum SYBR Green (Invitrogen) and AriaMX Real-Time PCR machine (Agilent Technologies) or ViiA7 Real-Time PCR machine (Applied Biosystems). Primers used for qPCR are listed in Table S3.
RNA-sequencing (RNA-seq)
Three biological replicates were prepared for each sample. RNA integrity was assessed using a BioAnalyzer 2000 (Agilent) using RNA Nano chips. Samples were ribodepleted and strand specific libraries were prepared by BGI Tech Solutions (Europe) and sequenced on the DNBseq platform (BGI) (100 bp, paired end).
Western blotting analysis
Whole cell protein lysates were prepared using RSB100 buffer (10 mM Tris-HCl pH 7.5, 100 mM NaCl, 2.5 mM MgCl2, 0.5% NP-40, 0.5% Triton X-100) freshly supplemented with protease inhibitors (Roche). Samples were denatured by the addition of NuPAGE Loading Buffer (Invitrogen) and NuPAGE Sample Reducing Agent (Invitrogen) before boiling at 95°C for 5 min. SDS-PAGE was carried out on either NuPAGE 4%–12% Bis-Tris or 3%–8% Tris-Acetate (Invitrogen) gels. Western blotting analysis was carried out according to standard protocols with the primary antibodies and HRP- conjugated secondary antibodies listed in the key resources table. Bands were visualized by SuperSignal West Femto ECL substrate (Thermo Scientific) and captured using an Amersham ImageQuant 800 imaging system (GE Healthcare). Images were processed using ImageJ (v1.53).ref. bib61
IP experiments
Cells from a confluent 10 cm dish were standardly extracted in HT150 extraction buffer (20 mM HEPES pH 7.4, 150/200 mM NaCl, 0.5% v/v Triton X-100) freshly supplemented with protease inhibitors (Roche). Lysates were sheared by sonication for 3 × 5 s and clarified by centrifugation at 20,000 rcf for 10 min. Cleared lysates were incubated with Pierce anti-c-MYC magnetic beads (Thermo Scientific) or ChromoTek GFP-Trap Magnetic Particles M-270 (Proteintech) overnight at 4°C. Beads were washed 3x with HT150 extraction buffer and transferred to a fresh tube for the final wash. Proteins were eluted by incubation at 75°C in NuPAGE Loading Buffer (Invitrogen) for 15 min. Eluates were mixed with NuPAGE Sample Reducing Agent (Invitrogen) and denatured for a further 5 min at 95°C before proceeding with western blotting analysis.
Protein expression and purification
Recombinant human ARS2147−871, ARS2147−817(ZnFmut), ZFC3H111−35(WT), ZFC3H111−35(MUT), ZC3H18190−211(WT) and ZC3H18190−211(MUT) were expressed in BL21 (DE3) E. coli cells. ARS2 isoform 4 constructs were cloned from synthetic gene codon optimized for E. coli (Invitrogen) as N-terminal 10×His tagged fusion protein cleavable with 3C protease. ZFC3H1 variants were cloned from a synthetic gene codon optimized for S. frugiperda (Invitrogen) as N-terminal 6×His-MBP fusion constructs. After E. coli cells were grown to OD600nm of 1.2 at 37°C in terrific broth (TB) medium, recombinant protein expression was induced with 0.5 mM IPTG and incubated at 18°C for 16 h. Cultures were harvested by centrifugation at 2000g for 15 min, and subsequently bacteria were lysed by sonication in 20 mM HEPES/NaOH pH 7.5, 500 mM NaCl, 20 mM imidazole, 5 mM β-mercaptoethanol, 0.5 mM AEBSF, and 15 U/ml benzonase (Merck). All proteins were initially purified using nickel-based affinity chromatography (IMAC, HIS-Select resin (Sigma-Aldrich)), washed with the wash buffer (lysis buffer supplemented with 1M NaCl, 10 mM MgSO4, 50 mM KCl, and 2 mM ATP) and eluted with 20 mM HEPES/NaOH pH 7.5, 100 mM NaCl, 300 mM imidazole, 2 mM DTT. ARS2 variants were subsequently loaded onto 1 mL HiTrap Heparin HP (Cytiva) and gradually eluted with buffer containing 1M NaCl. In the final purification step, ARS2, ZFC3H1 and ZC3H18 eluates were subjected to size exclusion chromatography on a Superdex 75 Increase 10/300 GL column (Cytiva) equilibrated in 20 mM HEPES/NaOH pH 7.5, 100 mM NaCl, 2 mM DTT.
In vitro pull-down assays
For in vitro pull-down assays, ARS2, ZFC3H1 and ZC3H18 variants were mixed at a final concentration of 10 or 30 μM for each protein in pull-down buffer containing 20 mM HEPES/NaOH pH 7.5, 100 mM NaCl, 2 mM DTT, 0.01% Tween 20 at a total volume of 20 μL and incubated on ice for 30 min. The protein mixtures were then incubated with amylose resin (NEB) for 1 h at 4°C. Beads were washed 3x times with pull-down buffer and bound proteins were eluted with SDS loading buffer. Input samples and eluates were analyzed by SDS-PAGE (12.5% gel) and visualized by Coomassie staining.
Mass spectrometry
Three biological samples were prepared for each sample. After affinity enrichment, eluted proteins were precipitated on MagResyn HILIC beads in 70% acetonitrile, washed in 95% acetonitrile and 70% ethanol to remove detergentsref. bib65 and digested with trypsin and Lys-C. The resulting peptides were desalted on StageTips and subjected to LCMS analysis on an EASY nano-LC 1200 system (80 min gradient) coupled to an Orbitrap Exploris 480 mass spectrometer (Thermo Scientific). MS data were acquired by data-dependent acquisition and subjected to MaxQuant database searchesref. bib66 using a human Uniprot reviewed sequence database (2021), ‘match between runs’, and ‘label-free Quantification’.
Data visualization
Genome browser views based on BigWig files were generated using the R package seqNdisplayR (https://rdrr.io/github/THJlab/seqNdisplayR/). RT-qPCR data was imported from the AriaMx Software (v1.71, Agilent) or QuantStudio Real-time PCR Software (v1.3, Applied Biosystems) and plotted using Graphpad Prism (v9.0.0). Data for multiple sequence alignments was extracted from the Aminode webtoolref. bib62 aligned using Clustal Omegaref. bib67 and displayed using Snapgene (v6.1.1). Volcano plots visualizing the analyzed MS data were generated using custom JavaScript/CSS webtool. Heatmaps were created using the R package Pheatmap (v1.0.12). Displayed data, nascent pA+/− 3′ends,ref. bib3 were centered around the annotated TES and log2 fold change was calculated within 2 nt bins. Displayed TUs represent top 50% of NEXT+PAXT ARS2-sensitive subset extracted after ranking according to mean log2fold change of siZCCHC8 relative to ctrl within the displayed region.
Quantification and statistical analysis
Quantification of western blots
Quantification of western blots following immunoprecipitation was performed on non-saturated exposed images using ImageJ (v1.51). Values of co-precipitated proteins were normalized to the bait protein values and plotted relative to control conditions using Graphpad Prism (v9.0.0).
RNA-seq data analysis
For all RNAseq data reads were mapped to GRCh38 with HISAT2 (v 2.2.1; ref. bib59) default settings and the following approach: Genome and splice site information for ‘H. sapiens, UCSC hg38 and Refseq gene annotations’ were obtained from the HISAT2 download page (ftp://ftp.ccb.jhu.edu/pub/infphilo/hisat2/data/). Data for canonical chromosomes (i.e., named chr1-22,X and Y but omitting all other sequence contigs) was selected to create a custom index for read mapping. After read mapping only proper read pairs with both reads mapping to the above genome annotation were used for further analysis. Exon read counts overlapping our HeLa in-house annotationsref. bib9 were collected using the featureCounts tools from the subread software suite (v 2.0.1; ref. bib68) with settings ‘-p -C -s 2 -t exon’. Differentially expressed transcripts were obtained from those raw read counts using R package DESeq2 (v 1.32.0) at a false discovery rate (FDR) cutoff of 0.1. Exon counts for differentially up- and down-regulated transcripts were compared using custom scripts in R. For classification of transcripts into NEXT and PAXT-dependent groups, we used differentially upregulated genes from siRBM7, siZCCHC8, siZFC3H1 -treated HeLa cellsref. bib24 (GSE84172) and similar data from siZC3H3 -treated cellsref. bib25 (GSE131255). Transcripts significantly upregulated (log2FC > 0 and padj <.1) in siRBM7, siZCCHC8, siZFC3H1 and siZC3H3 were selected and used to defined the set of NEXT targets (sig. upregulated in siRBM7 and siZCCHC8, but not sig. upregulated in siZC3H3 and siZFC3H1), PAXT targets (sig. upregulated in siZC3H3 and siZFC3H1, but not sig. upregulated in siRBM7 and siZCCHC8) and NEXT+PAXT targets (upregulated in all 4 knock-downs). ARS2-dependent transcripts were similarly defined as being significantly upregulated (FDR <.1) upon siARS2 knock-downref. bib30 (GSE99344).
Mass spectrometry analysis
Processed LFQ intensities of identified proteins were normalized to the LFQ intensity of the bait protein. Subsequently, Student’s t-test was performed to obtain differential protein enrichment in Z11−209(WT) IP relative to Z11−209(MUT) IP.
References
- S. Djebali, C.A. Davis, A. Merkel, A. Dobin, T. Lassmann, A. Mortazavi, A. Tanzer, J. Lagarde, W. Lin, F. Schlesinger. Landscape of transcription in human cells. Nature, 2012. [DOI | PubMed]
- B. Steurer, R.C. Janssens, B. Geverts, M.E. Geijer, F. Wienholz, A.F. Theil, J. Chang, S. Dealy, J. Pothof, W.A. van Cappellen. Live-cell analysis of endogenous GFP-RPB1 uncovers rapid turnover of initiating and promoter-paused RNA Polymerase II. Proc National Acad Sci, 2018. [DOI]
- G. Wu, M. Schmid, L. Rib, P. Polak, N. Meola, A. Sandelin, T.H. Jensen. A Two-Layered Targeting Mechanism Underlies Nuclear RNA Sorting by the Human Exosome. Cell Rep., 2020. [DOI | PubMed]
- J.T. Zimmer, N.A. Rosa-Mercado, D. Canzio, J.A. Steitz, M.D. Simon. STL-seq reveals pause-release and termination kinetics for promoter-proximal paused RNA polymerase II transcripts. Mol. Cell, 2021. [DOI | PubMed]
- A.E. Almada, X. Wu, A.J. Kriz, C.B. Burge, P.A. Sharp. Promoter directionality is controlled by U1 snRNP and polyadenylation signals. Nature, 2013. [DOI | PubMed]
- E. Ntini, A.I. Järvelin, J. Bornholdt, Y. Chen, M. Boyd, M. Jørgensen, R. Andersson, I. Hoof, A. Schein, P.R. Andersen. Polyadenylation site–induced decay of upstream transcripts enforces promoter directionality. Nat. Struct. Mol. Biol., 2013. [DOI | PubMed]
- Y. Chen, A.A. Pai, J. Herudek, M. Lubas, N. Meola, A.I. Järvelin, R. Andersson, V. Pelechano, L.M. Steinmetz, T.H. Jensen, A. Sandelin. Principles for RNA metabolism and alternative transcription initiation within closely spaced promoters. Nat. Genet., 2016. [DOI | PubMed]
- N.J. Proudfoot. Transcriptional termination in mammals: Stopping the RNA polymerase II juggernaut. Science, 2016. [DOI | PubMed]
- S. Lykke-Andersen, K. Žumer, E.Š. Molska, J.O. Rouvière, G. Wu, C. Demel, B. Schwalb, M. Schmid, P. Cramer, T.H. Jensen. Integrator is a genome-wide attenuator of non-productive transcription. Mol. Cell, 2021. [DOI | PubMed]
- N.D. Elrod, T. Henriques, K.-L. Huang, D.C. Tatomer, J.E. Wilusz, E.J. Wagner, K. Adelman. The Integrator Complex Attenuates Promoter-Proximal Transcription at Protein-Coding Genes. Mol. Cell, 2019. [DOI | PubMed]
- F. Beckedorff, E. Blumenthal, L.F. daSilva, Y. Aoi, P.R. Cingaram, J. Yue, A. Zhang, S. Dokaneheifard, M.G. Valencia, G. Gaidosh. The Human Integrator Complex Facilitates Transcriptional Elongation by Endonucleolytic Cleavage of Nascent Transcripts. Cell Rep., 2020. [DOI]
- M.S. Mendoza-Figueroa, D.C. Tatomer, J.E. Wilusz. The Integrator Complex in Transcription and Development. Trends Biochem. Sci., 2020. [DOI | PubMed]
- L.M.I. Austenaa, V. Piccolo, M. Russo, E. Prosperini, S. Polletti, D. Polizzese, S. Ghisletti, I. Barozzi, G.R. Diaferia, G. Natoli. A first exon termination checkpoint preferentially suppresses extragenic transcription. Nat. Struct. Mol. Biol., 2021. [DOI | PubMed]
- C. Estell, L. Davidson, P.C. Steketee, A. Monier, S. West. ZC3H4 restricts non-coding transcription in human cells. Elife, 2021. [DOI]
- J.O. Rouvière, A. Salerno-Kochan, S. Lykke-Andersen, W. Garland, Y. Dou, O. Rathore, E.Š. Molska, G. Wu, M. Schmid, A. Bugai. ARS2 instructs early transcription termination-coupled RNA decay by recruiting ZC3H4 to nascent transcripts. Mol. Cell, 2023. [DOI | PubMed]
- C. Estell, L. Davidson, J.D. Eaton, H. Kimura, V.A.M. Gold, S. West. A restrictor complex of ZC3H4, WDR82, and ARS2 integrates with PNUTS to control unproductive transcription. Mol. Cell, 2023. [DOI | PubMed]
- M. Schmid, T.H. Jensen. Controlling nuclear RNA levels. Nat. Rev. Genet., 2018. [DOI | PubMed]
- W. Garland, T.H. Jensen. 2020. [DOI]
- P. Mitchell, E. Petfalski, A. Shevchenko, M. Mann, D. Tollervey. The Exosome: A Conserved Eukaryotic RNA Processing Complex Containing Multiple 3′→5′ Exoribonucleases. Cell, 1997. [DOI | PubMed]
- B. Schuch, M. Feigenbutz, D.L. Makino, S. Falk, C. Basquin, P. Mitchell, E. Conti. The exosome-binding factors Rrp6 and Rrp47 form a composite surface for recruiting the Mtr4 helicase. Embo J, 2014. [DOI | PubMed]
- S. Falk, F. Bonneau, J. Ebert, A. Kögel, E. Conti. Mpp6 Incorporation in the Nuclear Exosome Contributes to RNA Channeling through the Mtr4 Helicase. Cell Rep., 2017. [DOI | PubMed]
- E.V. Wasmuth, J.C. Zinder, D. Zattas, M. Das, C.D. Lima. Structure and reconstitution of yeast Mpp6-nuclear exosome complexes reveals that Mpp6 stimulates RNA decay and recruits the Mtr4 helicase. Elife, 2017. [DOI]
- M. Lubas, M.S. Christensen, M.S. Kristiansen, M. Domanski, L.G. Falkenby, S. Lykke-Andersen, J.S. Andersen, A. Dziembowski, T.H. Jensen. Interaction Profiling Identifies the Human Nuclear Exosome Targeting Complex. Mol. Cell, 2011. [DOI | PubMed]
- N. Meola, M. Domanski, E. Karadoulama, Y. Chen, C. Gentil, D. Pultz, K. Vitting-Seerup, S. Lykke-Andersen, J.S. Andersen, A. Sandelin, T.H. Jensen. Identification of a Nuclear Exosome Decay Pathway for Processed Transcripts. Mol. Cell, 2016. [DOI | PubMed]
- T. Silla, M. Schmid, Y. Dou, W. Garland, M. Milek, K. Imami, D. Johnsen, P. Polak, J.S. Andersen, M. Selbach. The human ZC3H3 and RBM26/27 proteins are critical for PAXT-mediated nuclear RNA decay. Nucleic Acids Res., 2020. [DOI | PubMed]
- M. Gockert, M. Schmid, L. Jakobsen, M. Jens, J.S. Andersen, T.H. Jensen. Rapid factor depletion highlights intricacies of nucleoplasmic RNA degradation. Nucleic Acids Res., 2022. [DOI | PubMed]
- P. Gerlach, W. Garland, M. Lingaraju, A. Salerno-Kochan, F. Bonneau, J. Basquin, T.H. Jensen, E. Conti. Structure and regulation of the nuclear exosome targeting complex guides RNA substrates to the exosome. Mol. Cell, 2022. [DOI | PubMed]
- M.R. Puno, C.D. Lima. Structural basis for RNA surveillance by the human nuclear exosome targeting (NEXT) complex. Cell, 2022. [DOI | PubMed]
- P.R. Andersen, M. Domanski, M.S. Kristiansen, H. Storvall, E. Ntini, C. Verheggen, A. Schein, J. Bunkenborg, I. Poser, M. Hallais. The human cap-binding complex is functionally connected to the nuclear RNA exosome. Nat. Struct. Mol. Biol., 2013. [DOI | PubMed]
- C. Iasillo, M. Schmid, Y. Yahia, M.A. Maqbool, N. Descostes, E. Karadoulama, E. Bertrand, J.-C. Andrau, T.H. Jensen. ARS2 is a general suppressor of pervasive transcription. Nucleic Acids Res., 2017. [DOI | PubMed]
- K. Ogami, P. Richard, Y. Chen, M. Hoque, W. Li, J.J. Moresco, J.R. Yates, B. Tian, J.L. Manley. An Mtr4/ZFC3H1 complex facilitates turnover of unstable nuclear RNAs to prevent their cytoplasmic transport and global translational repression. Gene Dev., 2017. [DOI | PubMed]
- S.M. Bresson, N.K. Conrad. The Human Nuclear Poly(A)-Binding Protein Promotes RNA Hyperadenylation and Decay. PLoS Genet., 2013. [DOI]
- T. Gonatopoulos-Pournatzis, V.H. Cowling. Cap-binding complex (CBC). Biochem. J., 2014. [DOI | PubMed]
- M. Hallais, F. Pontvianne, P.R. Andersen, M. Clerici, D. Lener, N.E.H. Benbahouche, T. Gostan, F. Vandermoere, M.-C. Robert, S. Cusack. CBC–ARS2 stimulates 3′-end maturation of multiple RNA families and favors cap-proximal processing. Nat. Struct. Mol. Biol., 2013. [DOI | PubMed]
- W.M. Schulze, S. Cusack. Structural basis for mutually exclusive co-transcriptional nuclear cap-binding complexes with either NELF-E or ARS2. Nat. Commun., 2017. [DOI | PubMed]
- S. Lykke-Andersen, J.O. Rouvière, T.H. Jensen. ARS2/SRRT: at the nexus of RNA polymerase II transcription, transcript maturation and quality control. Biochem Soc T, 2021. [DOI]
- K. Winczura, M. Schmid, C. Iasillo, K.R. Molloy, L.M. Harder, J.S. Andersen, J. LaCava, T.H. Jensen. Characterizing ZC3H18, a Multi-domain Protein at the Interface of RNA Production and Destruction Decisions. Cell Rep., 2018. [DOI | PubMed]
- S. Giacometti, N.E.H. Benbahouche, M. Domanski, M.-C. Robert, N. Meola, M. Lubas, J. Bukenborg, J.S. Andersen, W.M. Schulze, C. Verheggen. Mutually Exclusive CBC-Containing Complexes Contribute to RNA Fate. Cell Rep., 2017. [DOI | PubMed]
- Y. Dou, I. Barbosa, H. Jiang, C. Iasillo, K.R. Molloy, W.M. Schulze, S. Cusack, M. Schmid, H. Le Hir, J. LaCava, T.H. Jensen. NCBP3 positively impacts mRNA biogenesis. Nucleic Acids Res., 2020. [DOI | PubMed]
- D. Libri. Nuclear Poly(A)-Binding Proteins and Nuclear Degradation: Take the mRNA and Run?. Mol. Cell, 2010. [DOI | PubMed]
- N. Meola, T.H. Jensen. Targeting the nuclear RNA exosome: Poly(A) binding proteins enter the stage. RNA Biol., 2017. [DOI | PubMed]
- T. Silla, E. Karadoulama, D. Mąkosa, M. Lubas, T.H. Jensen. The RNA Exosome Adaptor ZFC3H1 Functionally Competes with Nuclear Export Activity to Retain Target Transcripts. Cell Rep., 2018. [DOI | PubMed]
- J. Fan, B. Kuai, G. Wu, X. Wu, B. Chi, L. Wang, K. Wang, Z. Shi, H. Zhang, S. Chen. Exosome cofactor hMTR4 competes with export adaptor ALYREF to ensure balanced nuclear RNA pools for degradation and export. Embo J, 2017. [DOI | PubMed]
- N. Dobrev, Y.L. Ahmed, A. Sivadas, K. Soni, T. Fischer, I. Sinning. The zinc-finger protein Red1 orchestrates MTREC submodules and binds the Mtl1 helicase arch domain. Nat. Commun., 2021. [DOI | PubMed]
- A.-E. Foucher, L. Touat-Todeschini, A.B. Juarez-Martinez, A. Rakitch, H. Laroussi, C. Karczewski, S. Acajjaoui, M. Soler-López, S. Cusack, C.D. Mackereth. Structural analysis of Red1 as a conserved scaffold of the RNA-targeting MTREC/PAXT complex. Nat. Commun., 2022. [DOI | PubMed]
- Y. Wang, J. Fan, J. Wang, Y. Zhu, L. Xu, D. Tong, H. Cheng. ZFC3H1 prevents RNA trafficking into nuclear speckles through condensation. Nucleic Acids Res., 2021. [DOI | PubMed]
- W. Garland, I. Comet, M. Wu, A. Radzisheuskaya, L. Rib, K. Vitting-Seerup, M. Lloret-Llinares, A. Sandelin, K. Helin, T.H. Jensen. A Functional Link between Nuclear RNA Decay and Transcriptional Control Mediated by the Polycomb Repressive Complex 2. Cell Rep., 2019. [DOI | PubMed]
- M. Melko, K. Winczura, J.O. Rouvière, M. Oborská-Oplová, P.K. Andersen, T. Heick Jensen. Mapping domains of ARS2 critical for its RNA decay capacity. Nucleic Acids Res., 2020. [DOI | PubMed]
- T. Natsume, M.T. Kanemaki. Conditional Degrons for Controlling Protein Expression at the Protein Level. Annu. Rev. Genet., 2017. [DOI | PubMed]
- W. Garland, I. Müller, M. Wu, M. Schmid, K. Imamura, L. Rib, A. Sandelin, K. Helin, T.H. Jensen. Chromatin modifier HUSH co-operates with RNA decay factor NEXT to restrict transposable element expression. Mol. Cell, 2022. [DOI | PubMed]
- W.M. Schulze, F. Stein, M. Rettel, M. Nanao, S. Cusack. Structural analysis of human ARS2 as a platform for co-transcriptional RNA sorting. Nat. Commun., 2018. [DOI | PubMed]
- M.Y. Hein, N.C. Hubner, I. Poser, J. Cox, N. Nagaraj, Y. Toyoda, I.A. Gak, I. Weisswange, J. Mansfeld, F. Buchholz. A Human Interactome in Three Quantitative Dimensions Organized by Stoichiometries and Abundances. Cell, 2015. [DOI | PubMed]
- G. Wu, M. Schmid, T.H. Jensen. 3′ End sequencing of pA+ and pA− RNAs. Methods Enzymol., 2021. [DOI | PubMed]
- J. Fan, B. Kuai, K. Wang, L. Wang, Y. Wang, X. Wu, B. Chi, G. Li, H. Cheng. mRNAs are sorted for export or degradation before passing through nuclear speckles. Nucleic Acids Res., 2018. [DOI | PubMed]
- Y. Zhou, J. Zhu, G. Schermann, C. Ohle, K. Bendrin, R. Sugioka-Sugiyama, T. Sugiyama, T. Fischer. The fission yeast MTREC complex targets CUTs and unspliced pre-mRNAs to the nuclear exosome. Nat. Commun., 2015. [DOI | PubMed]
- J.J. Gruber, S.H. Olejniczak, J. Yong, G. La Rocca, G. Dreyfuss, C.B. Thompson. Ars2 Promotes Proper Replication-Dependent Histone mRNA 3′ End Formation. Mol. Cell, 2012. [DOI | PubMed]
- C. O’Sullivan, J. Christie, M. Pienaar, J. Gambling, P.E.B. Nickerson, S.C. Alford, R.L. Chow, P.L. Howard. Mutagenesis of ARS2 Domains To Assess Possible Roles in Cell Cycle Progression and MicroRNA and Replication-Dependent Histone mRNA Biogenesis. Mol. Cell Biol., 2015. [DOI | PubMed]
- K. Labun, T.G. Montague, M. Krause, Y.N. Torres Cleuren, H. Tjeldnes, E. Valen. CHOPCHOP v3: expanding the CRISPR web toolbox beyond genome editing. Nucleic Acids Res., 2019. [DOI | PubMed]
- D. Kim, B. Langmead, S.L. Salzberg. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods, 2015. [DOI | PubMed]
- M.I. Love, W. Huber, S. Anders. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol, 2014. [DOI | PubMed]
- C.A. Schneider, W.S. Rasband, K.W. Eliceiri. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods, 2012. [DOI | PubMed]
- K.T. Chang, J. Guo, A. di Ronza, M. Sardiello. Aminode: Identification of Evolutionary Constraints in the Human Proteome. Sci. Rep., 2018. [DOI | PubMed]
- Y. Perez-Riverol, J. Bai, C. Bandla, D. García-Seisdedos, S. Hewapathirana, S. Kamatchinathan, D.J. Kundu, A. Prakash, A. Frericks-Zipper, M. Eisenacher. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res., 2022. [DOI | PubMed]
- F.A. Ran, P.D. Hsu, C.-Y. Lin, J.S. Gootenberg, S. Konermann, A.E. Trevino, D.A. Scott, A. Inoue, S. Matoba, Y. Zhang, F. Zhang. Double Nicking by RNA-Guided CRISPR Cas9 for Enhanced Genome Editing Specificity. Cell, 2013. [DOI | PubMed]
- T.S. Batth, M.A.X. Tollenaere, P. Rüther, A. Gonzalez-Franquesa, B.S. Prabhakar, S. Bekker-Jensen, A.S. Deshmukh, J.V. Olsen. Protein Aggregation Capture on Microparticles Enables Multipurpose Proteomics Sample Preparation. Mol. Cell. Proteomics, 2019. [DOI | PubMed]
- J. Cox, M. Mann. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol., 2008. [DOI | PubMed]
- F. Sievers, A. Wilm, D. Dineen, T.J. Gibson, K. Karplus, W. Li, R. Lopez, H. McWilliam, M. Remmert, J. Söding. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol., 2011. [DOI | PubMed]
- Y. Liao, G.K. Smyth, W. Shi. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics, 2014. [DOI | PubMed]
