Elucidating the transcriptomic response of adult-derived mHypoA-2/12 mouse hypothalamic neuron cell line to cannabidiol (CBD) exposure
Abstract
Supplementary Information:
The online version contains supplementary material available at 10.1007/s13353-025-00970-8.
Article type: Research Article
Keywords: CBD, Hypothalamic neurons, Apoptosis, Extracellular matrix, RNA-seq
Affiliations: https://ror.org/012dxyr07grid.410701.30000 0001 2150 7124Faculty of Veterinary Medicine, University of Agriculture in Kraków, Redzina 1 C, 30-248, Krakow, Poland; https://ror.org/05f2age66grid.419741.e0000 0001 1197 1855Department of Animal Molecular Biology, National Research Institute of Animal Production, Krakowska 1, 32-083 Balice, Poland; https://ror.org/012dxyr07grid.410701.30000 0001 2150 7124Department of Animal Physiology and Endocrinology, University of Agriculture in Kraków, Mickiewicza 24/28, 30‑059 Krakow, Poland
License: © The Author(s) 2025 CC BY 4.0 Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
Article links: DOI: 10.1007/s13353-025-00970-8 | PubMed: 40335839 | PMC: PMC12605594
Relevance: Core topic: mentioned in title and keywords
Full text: PDF (2.7 MB)
Introduction
Cannabidiol (CBD) is one of the major phytocannabinoids found in hemp (Cannabis sativa), alongside the more well-known tetrahydrocannabinol (THC) (Meissner and Cascella ref. 2022). Unlike THC, CBD does not have psychotomimetic effects, making it an attractive subject of scientific research and medical interest (Sainz-Cort et al. ref. 2021). CBD has a broad spectrum of potential therapeutic benefits, especially concerning the nervous system (Singh et al. ref. 2023). CBD has neuroprotective potential and anti-inflammatory, analgesic, and anxiolytic properties (Patricio et al. ref. 2020). It is being investigated for use in the treatment of various neurological conditions such as epilepsy, multiple sclerosis, Parkinson’s disease, anxiety, depression, and sleep disorders (de Fátima dos Santos Sampaio et al. ref. 2024). Notably, CBD has gained recognition as an effective treatment for some forms of refractory epilepsy, confirmed by numerous clinical trials and accepted by regulatory agencies such as the US Food and Drug Administration (FDA) (Abu-Sawwa and Stehling ref. 2020) which approved the CBD-based medication Epidiolex for treatment of specific types of epilepsy. CBD actions in the brain are largely complex and not fully understood. Major targeted brain structures include the hippocampus, prefrontal cortex, mesolimbic system, amygdala, caudate, substantia nigra, and hypothalamus (Crippa et al. ref. 2004, ref. 2010; Castillo et al. ref. 2010; Batalla et al. ref. 2020; Lawn et al. ref. 2020).
CBD can potentially act on hypothalamic cells and functions through various molecular mechanisms involving several receptors present in hypothalamic neurons. It has been shown that CBD affects type 1 endocannabinoid (CB1) receptors that are widely distributed throughout the brain, including the hypothalamic suprachiasmatic nucleus (SCN), arcuate nucleus, and paraventricular nucleus (PVN) (Cruz-Martínez et al. ref. 2018). CBD has a low affinity to CB1 and acts as its negative allosteric modulator (Tham et al. ref. 2019). CBD may also modulate CB1 activity indirectly, for example, by affecting the levels of endocannabinoids such as anandamide (Scherma et al. ref. 2019). By inhibiting FAAH (fatty acid amide hydrolase) enzyme, CBD leads to an increase in the level of anandamide that acts on CB1 and CB2 receptors (Leweke et al. ref. 2012). CBD can also serve as an agonist of the 5-HT1 A receptor, which is involved in regulating mood, providing antidepressant, anti-anxiety, and antipsychotic effects (Resstel et al. ref. 2009). Additionally, CBD activates TRPV1 receptors, which are involved in pain processes, thermoregulation, and modulation of neurotransmission (Anand et al. ref. 2020). Other CBD actions in the hypothalamus can be mediated through the activation of PPARγ receptors (Khosropoor et al. ref. 2023). Available data suggests that activation of these receptors in the hypothalamus can affect energy metabolism and appetite regulation (Li et al. ref. 2018). Interestingly, CBD may also act on adenosine receptors, which play a role in regulating sleep (Huang et al. ref. 2014), inflammation (Blackburn et al. ref. 2009), and neuroprotection (Sagredo et al. ref. 2007; Liou et al. ref. 2008; Gonca and Darıcı ref. 2015; Zhang et al. ref. 2020). Finally, CBD may bind with high affinity to the GPR55 receptor (Lauckner et al. ref. 2008; Patricio et al. ref. 2022) and act as an antagonistic ligand, which may affect pain perception (Armin et al. ref. 2021) and inflammation (Ono et al. ref. 2023).
Each of these CBD actions may contribute to the broad spectrum of biological effects in the hypothalamus, highlighting the complexity and multifaceted nature of this compound’s effects on the brain. Specifically, some recent research showed that CBD impacts the synthesis and release of neurotransmitters in the rat Hypo-E22 cells and isolated hypothalamus, including dopamine, norepinephrine, and serotonin (di Giacomo et al. ref. 2020). In the rat ventromedial hypothalamus, CBD has been shown to cause panicolytic-like effects and reduce fear-induced responses, which may be important for managing anxiety-related behaviors (Khan et al. ref. 2020). Furthermore, CBD influences the hypothalamic–pituitary–adrenal (HPA) axis by enhancing its responsivity and normalizing anxiety-like behaviors, particularly in females, suggesting sex-specific effects on stress responses (Jenkins et al. ref. 2025).
Given the fundamental role of gene expression in the hypothalamus’s regulatory functions, this study undertook the analysis of changes arising in hypothalamic cells’ transcriptome following various CBD treatments. For this purpose, we used a hypothalamic cellular model, which allowed us to simplify the complexity of hypothalamic structures and complex interactions with other brain structures, but also abolished the need for animal sacrifice at this stage of experiments. Various hypothalamic cell models, derived from both embryonic (HypoE) and adult primary (HypoA) cultures, are currently available (Dalvi et al. ref. 2011, ref. 2021). These cell lines, immortalized by using a retroviral vector encoding the temperature-sensitive SV40 large T antigen, maintain fundamental characteristics of hypothalamic neurons, including neuronal morphology and expression of hypothalamus-specific markers. To investigate the influence of CBD on transcriptomic profiles of hypothalamic neurons, we employ adult-derived mHypoA-2/12 mouse cell lines. These cell lines have been demonstrated to be valuable models expressing key feeding-related neuropeptides, including NPY and AgRP, along with receptors such as NPY Y1R, NPY Y5R, melanocortin 3 receptor (MC3R), insulin receptor (IR), leptin receptor (Lep-R), ghrelin receptor (GHSR), and glucagon-like peptide receptor type 1 and 2 (GLP-1R, GLP-2R) (Dalvi et al. ref. 2011, ref. 2021).
In the course of the study, the neural cells were treated with different CBD doses and vehicle as a control for 6 and 24 h (h). The experiment setup allowed us to evaluate both dosage and time-dependent effect of CBD on hypothalamic cells, particularly on their viability, apoptosis and transcriptome profile, including specific genes expression. The transcriptome-wide method we employed enabled the formulation of a simplified, general hypothesis that CBD impacts the cells’ transcriptome. Our detailed analysis focused on identifying the altered pathways and associated biological processes. The results showed coordinated gene expression changes across different CBD treatments, revealing some of the mechanisms of CBD action in hypothalamic neural cells.
Material and methods
Cell culture and treatments
The adult mouse hypothalamic cell lines mHypoA-2/12 (Clu177) were purchased from Cedarlane (Canada) and cultured in 6 well plates (Thermo Fisher Scientific, USA) with Dulbecco’s Modified Eagle’s Medium with GlutaMAX (DMEM, 4500 mg/L, Gibco—Thermo Fisher Scientific, USA) supplemented with 10% fetal bovine serum (FBS, Gibco, Thermo Fisher Scientific, USA) and 1% Gibco™ Antibiotic–Antimycotic (10,000 units/mL of penicillin, 10,000 μg/mL of streptomycin, and 25 μg/mL of Gibco Amphotericin B, Thermo Fisher Scientific, USA) maintained at 37 °C and 5% CO2 until 75% confluency. The following day, the cells were cultured in DMEM with GlutaMAX, supplemented with CBD (0.325, 0.75, 1.5, and 3 µM) (MERCK, USA) either for 6 h (n = 3 for each concentration) or for 24 h (n = 3 for each concentration). CBD was suspended in DMSO and diluted to a final concentration in DMEM with GlutaMAX (final DMSO concentration of 0.1% in assay media). The vehicle control was 0.1% DMSO in the DMEM with GlutaMAX.
Cellular tests
Cell viability assay
Cell viability assays were performed on separate cell cultures treated at the same time as ones for RNA-Seq using the RealTime-Glo™ MT Cell Viability Assay according to the manufacturer’s protocol (Promega, USA). Cells were seeded at a density of 4000 cells/well into white-walled 96-well opaque assay plates. After the plated cells were allowed to attach overnight, they were treated with CBD concentrations ranging from 0.325 to 3 µM. The MT Cell Viability Substrate and NanoLuc Enzyme were equilibrated to 37 °C, 2 × RealTime-Glo reagent was prepared, and an equal volume was added to each well. For time zero measurements, cells were incubated with reagent for 20 min at 37 °C, and luminescence intensity was determined on a TECAN Infinite M200 PRO microplate reader. Luminescence was measured at 6 and 24 h after the addition of CBD.
Caspase activity
Caspase 3/7 activity was determined according to the manufacturer’s protocol. Briefly, mHypoA-2/12 cells (15 × 103 cells per well) were grown in a 96-well white plate and exposed to CBD for 6 h or 24 h. Cells with the addition of 5 μM staurosporine (0.1% final DMSO; Merck, USA) for 6 h were used as a positive control. For the assay, Caspase-Glo 3/7 reagent was added to all the wells in a 1:1 ratio and after shaking at room temperature for 30 min, the lysates were analyzed with the luminometer (Infinite M200 PRO, TECAN, Switzerland).
Transcriptome analysis
After the treatment time elapsed, cells were trypsinized, transferred to centrifuge tubes, and centrifuged at 500 RCF for 5 min to form pellets. Following removal of the supernatant, the cell pellets were snap-frozen at − 80 °C and stored until RNA isolation. Total RNA was isolated using the standard TRI Reagent™ Solution (ThermoFisher Scientific, USA) procedure. The quality of the RNA isolates was evaluated using the TapeStation 4150 System (Agilent, USA), and RNA quantification was performed using the Qubit RNA BR assay (ThermoFisher Scientific, USA).
Library preparation was carried out with 50 ng of total RNA using the QuantSeq 3′ mRNA-Seq Library Prep Kit FWD (Lexogen, Austria). This kit generates one fragment per transcript at the 3′ end, reducing the number of raw reads generated per sample to approximately 3 million (M). The quality of the indexed libraries was checked using the TapeStation 4150 System (Agilent, USA) and quantified using the Qubit dsDNA BR kit (ThermoFisher Scientific, USA). Finally, the library pools were sequenced commercially by the OMRF Clinical Genomics Center (CGC) in a single-end 150 bp run on a NovaSeq 6000 System (Illumina, USA), yielding at least 6–7 M reads per sample. These raw sequences and read counts were deposited in the Gene Expression Omnibus (GEO) and Sequence Read Archive (SRA) databases of the National Center for Biotechnology Information (NCBI) under accession number GSE270378.
Bioinformatic analysis
The raw sequencing reads received were quality-checked using FastQC (v0.11.9) software. Filtering and trimming were done using Flexbar software (3.5.0) (Dodt et al. ref. 2012). During filtering, low-quality read ends, adapter sequences, and reads that were too short after trimming were removed. High-quality cleaned reads were mapped to the Mouse GRC39 genome assembly using the STAR aligner software (2.7.5c) (Dobin et al. ref. 2013). Mapped reads were counted using MM109 (Ensembl) annotation and Htseq-count (1.99.2) (Anders et al. ref. 2015) software. Normalization of red counts and differential expression (DE) analysis were performed using DESeq2 (v3.16) (Love et al. ref. 2014) software. An FDR < 0.1 was required to consider a gene as differentially expressed. The DE genes were analyzed for their functional implications by enrichment analysis in a specific GO BP (gene ontology biological processes), KEEG (Kyoto Encyclopedia of Genes and Genomes), and GO CC (cellular components) categories using iDEP2.0 server (Ge et al. ref. 2018). Additional functional enrichment analysis was performed with WebGestalt (WEB-based GEne SeT AnaLysis Toolkit) (Liao et al. ref. 2019). Venn diagrams were prepared using online tools (https://bioinformatics.psb.ugent.be/webtools/Venn/) (Heberle et al. ref. 2015).
qPCR validation of 3′mRNA-Seq
Due to the insufficient amount or total RNA from initial treatments and purification, qPCR validation of 3′mRNA-Seq results was conducted based on replicated experiments on the same cell line. The cells were treated as in the main experiment with all CBD doses and vehicle as control. Only 24-h treatment was replicated. For validation, four genes that were differently expressed in at least three comparisons (treatments) in a major experiment (in 24-h treatment) were selected. The genes included Mmp13, Naca, Ifitm2, and Gm8960. The Hprt gene was used as an endogenous control (Zamani et al. ref. 2020). Primer sequences are provided in Supplementary File 1.
Genes were tested using RT-qPCR to verify the reliability of the analysis. For this purpose, cDNA was synthesized using 500 ng of RNA and the High-Capacity RNA-to-cDNA kit (Thermo Fisher Scientific), following the manufacturer’s protocol. RT-qPCR was performed with the AmpliQ 5 × HOT EvaGreen® qPCR Mix Plus (ROX) kit (Novazym, Poznan, Poland) and primers for mRNA sequences spanning two adjacent exons. Each sample was run in triplicate using Quant Studio 7 Flex (Thermo Fisher Scientific). The relative expression levels of each gene were calculated using the ΔΔCt method. Standardization was performed based on Hprt as the internal control.
Relative expression concordance between RNA-Seq and qPCR methods was assessed based on the correlation coefficient analysis on mean expression values within groups. Calculations were performed using JASP 0.11.1 software. The Spearman correlation coefficient was selected for this analysis, following data distribution evaluation using the Shapiro–Wilk test.
Results
Effect of CBD exposure on cells viability and apoptosis
To assess the impact of CBD on mHypoA-2/12 cell viability, the cells were treated with increasing concentrations of CBD (0.25, 0.75, 1.5, and 3 µM) for 6 or 24 h. Notably, within 24 h of exposure, CBD significantly enhanced cell survival across all concentrations (Fig. 1; p < 0.002 vs. vehicle control). As illustrated in Fig. 1, there is a notable increase in cell viability after 6 h of CBD treatment, particularly with concentrations of 0.75 and 1.5 µM (Fig. 1; p < 0.0004).
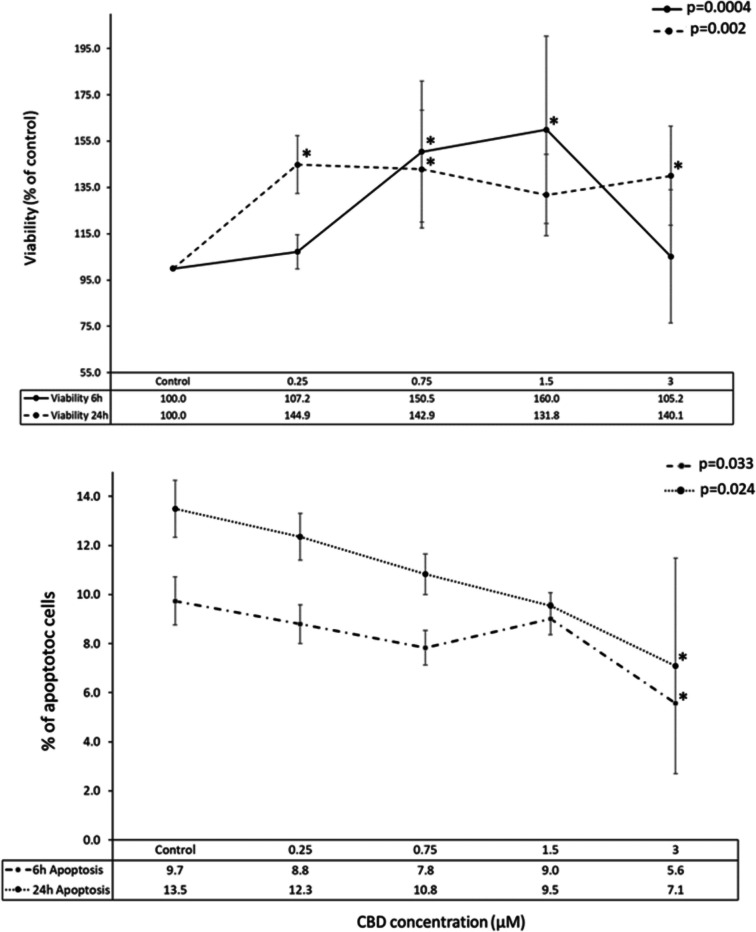
To investigate whether CBD-induced cell death occurred via apoptosis, we measured caspase 3/7 activity. Interestingly, at the highest CBD concentration (3 µM), caspase 3/7 activity significantly decreased after both 6 and 24 h (Fig. 1; p < 0.033 and p < 0.024, respectively). However, no significant changes in caspase 3/7 activity were observed in the remaining experimental conditions.
Sequencing statistics and general genes expression profile differentiation
In total, for 3′mRNA-Seq experiment, more than 210 M sequencing reads were generated for 30 samples—around 7 M per sample on average. Of the raw reads, on average 98.9% passed initial filtering and 54% were uniquely mapped against the newest available reference genome sequence. More than 75% of the mapped reads were located within annotation features (genes) (Supplementary File 2). Gene-related read counts were further used for differential expression analysis between various experimental conditions and transcriptome profile differentiation analysis.
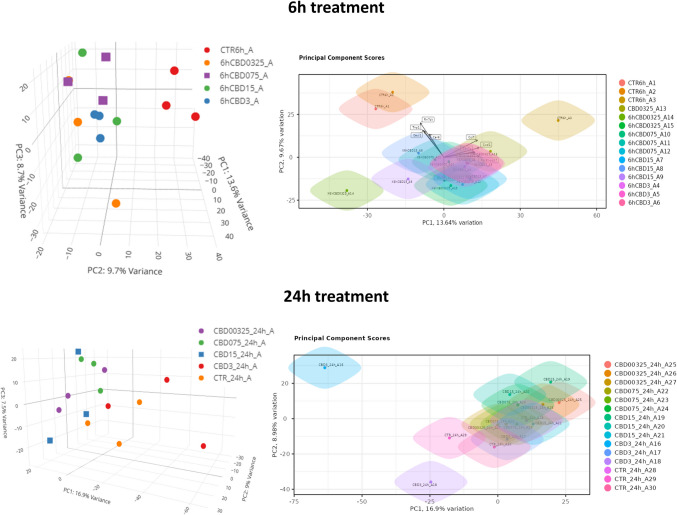
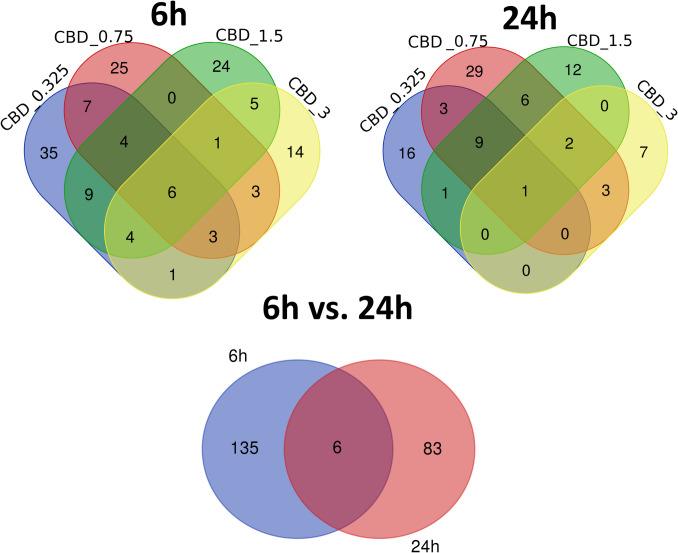
In 6-h treatment, CBD supplementation at various concentrations induced only a minor change in neural cells transcriptomes, but genially, expression profile of treated cells differed form control cells (vehicle). This was visible in principal components analysis (PCA) in which control cells clustered separately from a mixed group of cells treated with different concentrations of CBD. At 24-h treatment, the separation of control cells from treatments was not so clearly visible, and they co-localized in the PCA graph with 3 µM CBD-treated cells (Fig. 2).
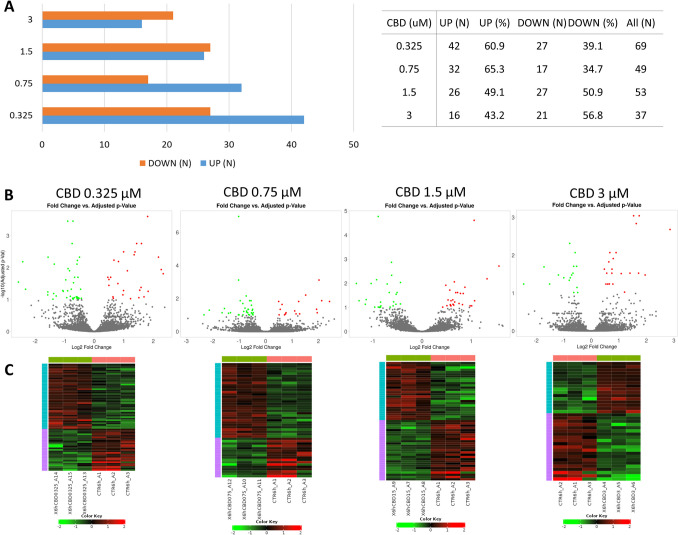
In 6-h treatment, the highest amount of genes was altered by the lowest applied CBD concentration (0.325 µM; n = 369) and in contrast, the lowest number of genes was altered by the highest CBD dose (3 µM; n = 37) (Supplementary File 3). At lower CBD concentrations, the genes were mainly downregulated (60.9 and 65.3% for 0.325 and 0.7 µM CBD, respectively), while the 3 μM CBD dose was associated with predominant genes upregulation (56.8%) (Fig. 3).
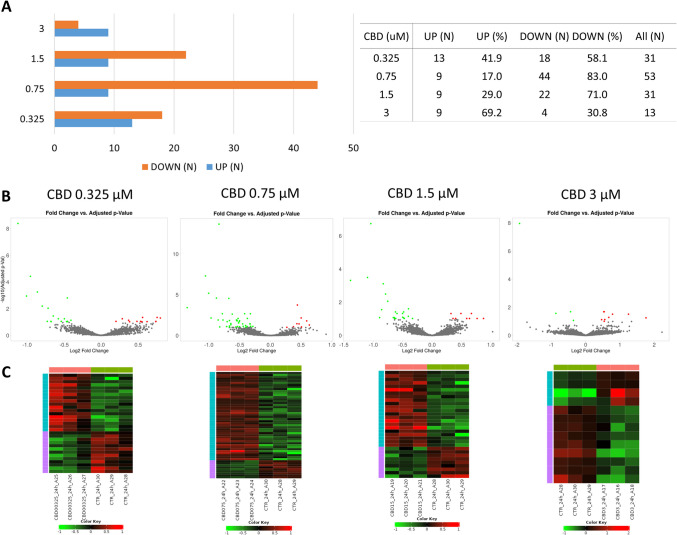
Following 24 h of treatment, again the highest amount of genes was altered by lower CBD concentrations (0.0325 and 0.075 µM; n = 331 and 53, respectively) (Supplementary File 3). The lowest amount of altered genes was observed for cells treated with 3 µM of CBD (n = 313). In the most treatments, (except 3 µM CBD), CBD caused mainly downregulation of genes expression (Fig. 4).
Of the 141 different genes whose expression was altered by CBD in 6-h treatment, six were affected by all applied CBD concentrations, and other 11 by at least three different CBD doses. The cells treated with 3 μM CBD solution had the most distinct profile of genes expression changes showed by the lowest amount of genes that were commonly altered by other treatments (Fig. 5). After 24-h incubation with CBD, of the 89 altered genes, only one was common for all treatments, however nine others were common for three lower CBD doses. All those nine genes were downregulated (Fig. 5). What is more, six common genes were altered in CBD-treated cells at 6 and 24 h (Supplementary File 4).
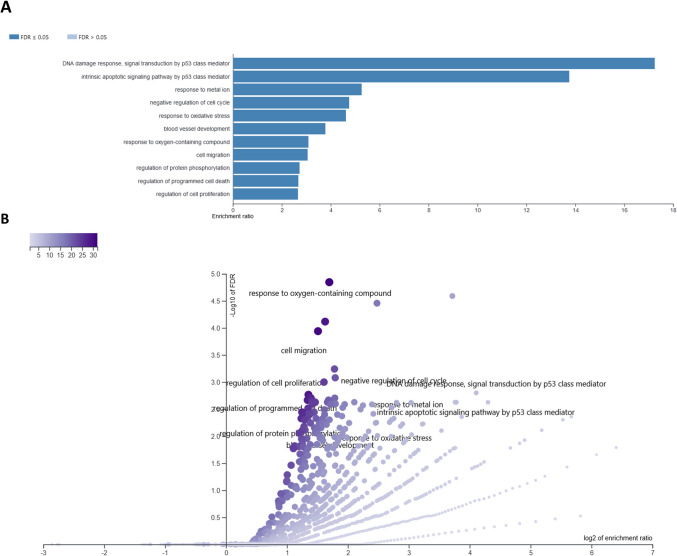
Functional annotation of genes altered by CBD in neural cells
After 6-h treatment of neural cells with different concentrations of CBD, in total 141 different genes were affected, from 37 (3 μM CBD) to 69 (0.325 μM) per treatment. The overrepresentation tests for those genes in GO BP and KEEG categories are presented in Supplementary File 5. Among top 50 overrepresented (FDR < 0.1) BP and KEEG pathways for lower CBD concentrations, only processes associated with upregulated genes have been found, however, in case of 3 μM concentration, both up- and downregulated genes were enriched. A comparative analysis of GO BP among various CBD treatments showed, that processes such as cell population proliferation, immune system process, cellular response to stress, cellular response to decreased oxygen levels, cellular response to hypoxia, DNA damage response signal transduction by p53 class mediator, positive regulation of programmed cell death, apoptotic process, and positive regulation of metabolic process were overrepresented by genes altered by at least three different CBD treatments. A similar analysis performed for KEEG pathways showed that pathways such as pathways in cancer, P53 signaling, and osteoclast differentiation were found to be overrepresented for at least three different CBD treatments (Supplementary File 5; Table 1). Comparative analysis of the enriched (FDR < 0.1) cellular components (Supplementary File 6) showed that categories related to, e.g., PCNA-p21 complex, PUMA-BCL-xl complex, nucleolus, extracellular region, extracellular matrix, collagen-containing extracellular matrix, external encapsulating structure, and extracellular space were associated with genes altered by more than one CBD treatment (Supplementary File 6). Combined analysis of all 141 genes altered by CBD in 6-h treatment revealed that (following redundancy removal by affinity propagation) the enriched BP (FDR < 0.1) included among others: response to oxygen-containing compound, cell migration, regulation of cell proliferation, intrinsic apoptotic signaling pathway by p53 class mediator, regulation of programmed cell death, response to metal ion, negative regulation of cell cycle, response to oxidative stress, and others. Detailed information on this analysis can be found in Supplementary File 7 and Fig. 6. In depth analysis of six genes that were altered by all CBD concentrations at 6-h treatment, and other 11 that were common for at least three different CBD doses (Supplementary File 4) showed that the genes were enriched (FDR < 0.1) in BP such as: wound healing involved in inflammatory response, inflammatory response to wounding and cellular response to stress. They also included two genes directly involved in extracellular matrix functioning, namely Mmp3 and Timp1 genes.
Table 1: Comparative analysis of biological processes and KEEG pathways overrepresented by different CBD treatments (concentrations) at 6-h treatment. Processes overrepresented by at least two treatments are presented along with the engaged genes
| 6-h treatment | ||
|---|---|---|
| Common for CBD treatment | Category | Genes |
| Biological processes | ||
| CBD_0.325 CBD_0.75 CBD_1.5 CBD_3 | Cell population proliferation | ENSMUSG00000052837, ENSMUSG00000022528, ENSMUSG00000005125, ENSMUSG00000005413, ENSMUSG00000023034, ENSMUSG00000023067, ENSMUSG00000027803, ENSMUSG00000032402, ENSMUSG00000015837, ENSMUSG00000024190, ENSMUSG00000020184, ENSMUSG00000040435, ENSMUSG00000022528, ENSMUSG00000052684, ENSMUSG00000029377, ENSMUSG00000052837, ENSMUSG00000041313, ENSMUSG00000056708, ENSMUSG00000038418, ENSMUSG00000030717, ENSMUSG00000032501 |
| Immune system process | ENSMUSG00000090272, ENSMUSG00000005125, ENSMUSG00000005413, ENSMUSG00000015837, ENSMUSG00000022528, ENSMUSG00000024190, ENSMUSG00000029561, ENSMUSG00000032402, ENSMUSG00000034459, ENSMUSG00000035692, ENSMUSG00000041827, ENSMUSG00000023067, ENSMUSG00000052837, ENSMUSG00000052684, ENSMUSG00000027111, ENSMUSG00000090272, ENSMUSG00000021250, ENSMUSG00000039883, ENSMUSG00000041313, ENSMUSG00000029377, ENSMUSG00000038418, ENSMUSG00000073489, ENSMUSG00000032501, ENSMUSG00000042406, | |
| Regulation of cell population proliferation | ENSMUSG00000052837, ENSMUSG00000022528, ENSMUSG00000005125, ENSMUSG00000005413, ENSMUSG00000023034, ENSMUSG00000023067, ENSMUSG00000027803, ENSMUSG00000032402, ENSMUSG00000024190, ENSMUSG00000020184, ENSMUSG00000040435, ENSMUSG00000022528, ENSMUSG00000052684, ENSMUSG00000029377, ENSMUSG00000052837, ENSMUSG00000041313, ENSMUSG00000056708, ENSMUSG00000038418, ENSMUSG00000030717, ENSMUSG00000032501, | |
| Cellular response to stress | ENSMUSG00000002083, ENSMUSG00000024190, ENSMUSG00000040435, ENSMUSG00000023067, ENSMUSG00000005125, ENSMUSG00000005413, ENSMUSG00000015837, ENSMUSG00000020184, ENSMUSG00000021453, ENSMUSG00000021701, ENSMUSG00000032402, ENSMUSG00000052684, ENSMUSG00000009630, ENSMUSG00000023067, ENSMUSG00000038418, ENSMUSG00000021250, ENSMUSG00000028494, ENSMUSG00000056708, ENSMUSG00000024843, ENSMUSG00000040435, ENSMUSG00000042406, ENSMUSG00000030717, ENSMUSG00000032501, ENSMUSG00000073489, | |
| CBD_0.325 CBD_0.75 CBD_1.5 | Cellular response to decreased oxygen levels | ENSMUSG00000002083, ENSMUSG00000005125, ENSMUSG00000005413, ENSMUSG00000020184 ENSMUSG00000052684, ENSMUSG00000038418, ENSMUSG00000021250, |
| Cellular response to hypoxia | ENSMUSG00000002083, ENSMUSG00000005125, ENSMUSG00000005413, ENSMUSG00000020184, ENSMUSG00000052684, ENSMUSG00000038418, ENSMUSG00000021250 | |
| DNA damage response signal transduction by p53 class mediator | ENSMUSG00000020184, ENSMUSG00000023067, ENSMUSG00000023067, ENSMUSG00000005125 | |
| CBD_0.325 CBD_0.75 CBD_3 | Positive regulation of programmed cell death | ENSMUSG00000023034, ENSMUSG00000002083, ENSMUSG00000021453, ENSMUSG00000005413, ENSMUSG00000023067, ENSMUSG00000024190, ENSMUSG00000040435, ENSMUSG00000052684, ENSMUSG00000030717, ENSMUSG00000042406, |
| Negative regulation of cell population proliferation | ENSMUSG00000022528, ENSMUSG00000005413, ENSMUSG00000023067, ENSMUSG00000032402, ENSMUSG00000005125, ENSMUSG00000024190, ENSMUSG00000040435, ENSMUSG00000052684, ENSMUSG00000030717, ENSMUSG00000032501, | |
| Response to stress | ENSMUSG00000002083, ENSMUSG00000005413, ENSMUSG00000024190, ENSMUSG00000029561, ENSMUSG00000034459, ENSMUSG00000040435, ENSMUSG00000041827, ENSMUSG00000090272, ENSMUSG00000023067, ENSMUSG00000005125, ENSMUSG00000015837, ENSMUSG00000020184, ENSMUSG00000021453, ENSMUSG00000032402, ENSMUSG00000035692, ENSMUSG00000021701, ENSMUSG00000052684, ENSMUSG00000009630, ENSMUSG00000023067, ENSMUSG00000038418, ENSMUSG00000021250, ENSMUSG00000028494, ENSMUSG00000056708, ENSMUSG00000024843, ENSMUSG00000005413, ENSMUSG00000073489, ENSMUSG00000042406, ENSMUSG00000030717, ENSMUSG00000032501 | |
| Apoptotic process | ENSMUSG00000002083, ENSMUSG00000020184, ENSMUSG00000023034, ENSMUSG00000005413, ENSMUSG00000015837, ENSMUSG00000021453, ENSMUSG00000032402, ENSMUSG00000040435, ENSMUSG00000021701, ENSMUSG00000023067, ENSMUSG00000024190, ENSMUSG00000052684, ENSMUSG00000009630, ENSMUSG00000042406, ENSMUSG00000003541, ENSMUSG00000030717, ENSMUSG00000073489 | |
| Cell death | ENSMUSG00000002083, ENSMUSG00000020184, ENSMUSG00000023034, ENSMUSG00000005413, ENSMUSG00000015837, ENSMUSG00000021453, ENSMUSG00000032402, ENSMUSG00000040435, ENSMUSG00000021701, ENSMUSG00000023067, ENSMUSG00000024190, ENSMUSG00000052684, ENSMUSG00000009630, ENSMUSG00000042406, ENSMUSG00000003541, ENSMUSG00000030717, ENSMUSG00000073489 | |
| Programmed cell death | ENSMUSG00000002083, ENSMUSG00000020184, ENSMUSG00000023034, ENSMUSG00000005413, ENSMUSG00000015837, ENSMUSG00000021453, ENSMUSG00000032402, ENSMUSG00000040435, ENSMUSG00000021701, ENSMUSG00000023067, ENSMUSG00000024190, ENSMUSG00000052684, ENSMUSG00000009630, ENSMUSG00000042406, ENSMUSG00000003541, ENSMUSG00000030717, ENSMUSG00000073489 | |
| Signal transduction by p53 class mediator | ENSMUSG00000023067, ENSMUSG00000005125, ENSMUSG00000020184, ENSMUSG00000002083 ENSMUSG00000030717, ENSMUSG00000073489 | |
| CBD_0.325 CBD_1.5 CBD_3 | Positive regulation of metabolic process | ENSMUSG00000002083, ENSMUSG00000027803, ENSMUSG00000040435, ENSMUSG00000090272, ENSMUSG00000022528, ENSMUSG00000023034, ENSMUSG00000032402, ENSMUSG00000000078, ENSMUSG00000015837, ENSMUSG00000020184, ENSMUSG00000021701, ENSMUSG00000023067, ENSMUSG00000035692, ENSMUSG00000052837, ENSMUSG00000005413, ENSMUSG00000029377, ENSMUSG00000038418, ENSMUSG00000021250, ENSMUSG00000027111, ENSMUSG00000056708, ENSMUSG00000001627, ENSMUSG00000032501, ENSMUSG00000038508, ENSMUSG00000073489, ENSMUSG00000042406, ENSMUSG00000030717, |
| KEEG | ||
| CBD_0.325 CBD_0.75 CBD_1.5 | Colorectal cancer | ENSMUSG00000023067, ENSMUSG00000002083, ENSMUSG00000032402, ENSMUSG00000021453, ENSMUSG00000052684, ENSMUSG00000029377, ENSMUSG00000021250 |
| Human papillomavirus infection | ENSMUSG00000023067, ENSMUSG00000022528, ENSMUSG00000020184, ENSMUSG00000041827, ENSMUSG00000029561, ENSMUSG00000009630, ENSMUSG00000027111, ENSMUSG00000026478, | |
| Mitophagy-animal | ENSMUSG00000015837, ENSMUSG00000008348, ENSMUSG00000052684, ENSMUSG00000015837 | |
| Pathways in cancer | ENSMUSG00000023067, ENSMUSG00000022528, ENSMUSG00000005413, ENSMUSG00000002083, ENSMUSG00000032402, ENSMUSG00000020184, ENSMUSG00000021453, ENSMUSG00000052684, ENSMUSG00000021250, ENSMUSG00000027111, ENSMUSG00000026478 | |
| P53 signaling pathway | ENSMUSG00000023067, ENSMUSG00000002083, ENSMUSG00000020184, ENSMUSG00000021453, | |
| Platinum drug resistance | ENSMUSG00000023067, ENSMUSG00000002083, ENSMUSG00000020184 | |
| Epstein-Barr virus infection | ENSMUSG00000023067, ENSMUSG00000022528, ENSMUSG00000020184, ENSMUSG00000021453, ENSMUSG00000052684 | |
| Breast cancer | ENSMUSG00000023067, ENSMUSG00000022528, ENSMUSG00000021453 ENSMUSG00000052684, ENSMUSG00000021250, | |
| Osteoclast differentiation | ENSMUSG00000052837, ENSMUSG00000015837, ENSMUSG00000003545, ENSMUSG00000052684, ENSMUSG00000021250, ENSMUSG00000052837, | |
| CBD_0.325 CBD_1.5 | Endocrine resistance | ENSMUSG00000023067, ENSMUSG00000020184, ENSMUSG00000021250 |
| Maturity onset diabetes of the young | ENSMUSG00000022528 | |
| Melanoma | ENSMUSG00000023067, ENSMUSG00000020184, ENSMUSG00000021453 | |
| Prostate cancer | ENSMUSG00000023067, ENSMUSG00000020184, | |
| Small cell lung cancer | ENSMUSG00000023067, ENSMUSG00000021453, ENSMUSG00000027111, ENSMUSG00000026478, | |
| Cellular senescence | ENSMUSG00000023067, ENSMUSG00000032402, ENSMUSG00000020184, ENSMUSG00000015837, ENSMUSG00000021453 | |
| Bladder cancer | ENSMUSG00000023067, ENSMUSG00000020184, | |
| HIF-1 signaling pathway | ENSMUSG00000023067, ENSMUSG00000005413, | |
| MicroRNAs in cancer | ENSMUSG00000023067, ENSMUSG00000005413, ENSMUSG00000020184, ENSMUSG00000041313, ENSMUSG00000023067 | |
| Glioma | ENSMUSG00000023067, ENSMUSG00000020184, ENSMUSG00000021453 | |
| Chronic myeloid leukemia | ENSMUSG00000023067, ENSMUSG00000032402, ENSMUSG00000020184, ENSMUSG00000021453, | |
| Fluid shear stress and atherosclerosis | ENSMUSG00000005413, ENSMUSG00000015837, ENSMUSG00000024190, ENSMUSG00000021250, ENSMUSG00000005413 | |
| CBD_0.325 CBD_3 | Transcriptional misregulation in cancer | ENSMUSG00000023067, ENSMUSG00000020184, ENSMUSG00000021453, ENSMUSG00000055435, ENSMUSG00000043613 |
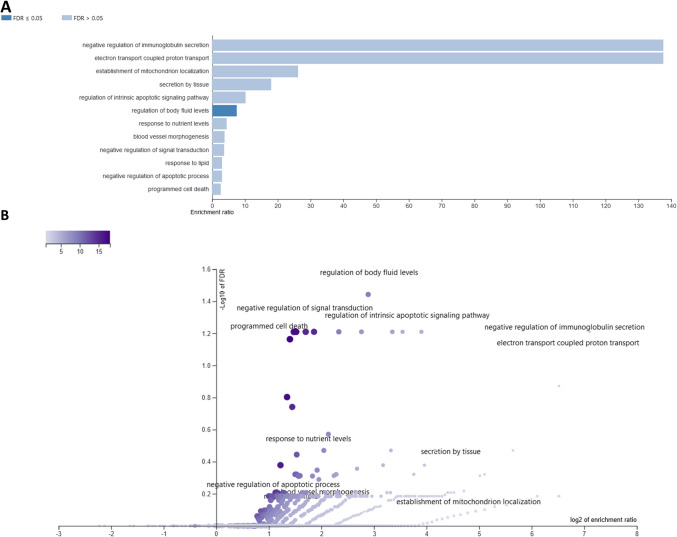
After 24 h CBD treatment, 89 different genes were altered with respect to vehicle (control), from 13 (3 μM) to 53 (0.75 μM) per treatment. Among the top 50 enriched BP (Supplementary File 8) comparative analysis revealed: cell motility and migration, acute-phase response, monocyte chemotaxis, negative regulation of leukocyte migration, and amine transport-related biological processes, as overrepresented by at least two different CBD treatments. Among the common for different treatments overrepresented (FDR < 0.1) KEEG pathways, there were, e.g., Parkinson disease, thermogenesis, RNA degradation, vascular smooth muscle contraction, platelet activation, oxytocin signaling pathway, and calcium signaling pathway (Supplementary File 8; Table 2). Only genes altered by two extreme CBD doses enriched any cellular components (CC) categories. Genes affected by 0.325 μM CBD treatment enriched CC associated with, e.g., mitochondrial respiratory chain complex I, NADH dehydrogenase complex (upregulated genes) and extracellular space, cell–cell junction, intercellular canaliculus (downregulated genes) and others (Supplementary File 6). 3 μM CBD dose, however, affected genes that were enriched in CC such as elongator holoenzyme complex, organellar ribosome, stress fiber and mitochondrial ribosome (Supplementary File 6). Combined analysis of all 89 CBD-affected genes in 24-h treatment revealed that following affinity propagation redundancy removal several processes remained enriched (FDR < 0.1), namely, regulation of body fluid levels, negative regulation of signal transduction, regulation of intrinsic apoptotic signaling pathway, negative regulation of immunoglobulin secretion, and programmed cell death (Supplementary File 9; Fig. 7). Nine genes that were downregulated in three of four CBD treatments (24 h) were enriched (FDR < 0.1) in BP associated with: cell chemotaxis and migration, regulation of amine transport, regulation of amino acid transport, regulation of monocyte chemotaxis and myeloid leukocyte migration. Single gene that was commonly affected by all CBD doses in 24-h treatment was Mylk that encode Myosin light chain kinase.
Table 2: Comparative analysis of biological processes and KEEG pathways overrepresented by different CBD treatments (concentrations) at 24-h treatment. Processes overrepresented by at least two treatments are presented along with the engaged genes
| 24-h treatment | ||
|---|---|---|
| CBD concentration | Category | Genes |
| Biological processes | ||
| CBD_0.325 CBD_1.5 CBD_3 | Regulation of cell migration | ENSMUSG00000032531; ENSMUSG00000061353; ENSMUSG00000022836; ENSMUSG00000024190; ENSMUSG00000032714; ENSMUSG00000026822; ENSMUSG00000026981; ENSMUSG00000054836; ENSMUSG00000004951; ENSMUSG00000022836; |
| Localization of cell | ENSMUSG00000032531; ENSMUSG00000061353; ENSMUSG00000040026; ENSMUSG00000022836; ENSMUSG00000024190; ENSMUSG00000026620; ENSMUSG00000032714; ENSMUSG00000061315; ENSMUSG00000026822; ENSMUSG00000026981; ENSMUSG00000054836; ENSMUSG00000022836; | |
| Cell motility | ENSMUSG00000032531; ENSMUSG00000061353; ENSMUSG00000040026; ENSMUSG00000022836; ENSMUSG00000024190; ENSMUSG00000026620; ENSMUSG00000032714; ENSMUSG00000061315; ENSMUSG00000026822; ENSMUSG00000026981; ENSMUSG00000054836 | |
| Regulation of cell motility | ENSMUSG00000032531; ENSMUSG00000061353; ENSMUSG00000022836; ENSMUSG00000024190; ENSMUSG00000032714; ENSMUSG00000026822; ENSMUSG00000026981; ENSMUSG00000054836; ENSMUSG00000022836 | |
| Regulation of cellular component movement | ENSMUSG00000032531; ENSMUSG00000061353; ENSMUSG00000022836; ENSMUSG00000024190; ENSMUSG00000032714; ENSMUSG00000038848; ENSMUSG00000026822; ENSMUSG00000026981 ENSMUSG00000054836; ENSMUSG00000022836 | |
| Cell migration | ENSMUSG00000032531; ENSMUSG00000061353; ENSMUSG00000040026; ENSMUSG00000022836; ENSMUSG00000024190; ENSMUSG00000026620; ENSMUSG00000032714; ENSMUSG00000061315; ENSMUSG00000026822; ENSMUSG00000026981; ENSMUSG00000054836 | |
| Regulation of locomotion | ENSMUSG00000032531; ENSMUSG00000061353; ENSMUSG00000022836; ENSMUSG00000024190; ENSMUSG00000032714; ENSMUSG00000038848; ENSMUSG00000026822; ENSMUSG00000026981; ENSMUSG00000054836; ENSMUSG00000022836; | |
| CBD_0.325 CBD_1.5 | Acute-phase response | ENSMUSG00000040026; ENSMUSG00000026981; |
| Movement of cell or subcellular component | ENSMUSG00000032531; ENSMUSG00000061353; ENSMUSG00000040026; ENSMUSG00000022836; ENSMUSG00000024190; ENSMUSG00000026620; ENSMUSG00000032714; ENSMUSG00000038848; ENSMUSG00000061315; ENSMUSG00000026822; ENSMUSG00000026981; ENSMUSG00000032536; ENSMUSG00000054836 | |
| Monocyte chemotaxis | ENSMUSG00000024190; ENSMUSG00000061353; | |
| Negative regulation of leukocyte migration | ENSMUSG00000024190; ENSMUSG00000061353; | |
| Locomotion | ENSMUSG00000032531; ENSMUSG00000061353; ENSMUSG00000040026; ENSMUSG00000022836; ENSMUSG00000024190; ENSMUSG00000026620; ENSMUSG00000032714; ENSMUSG00000038848; ENSMUSG00000061315; ENSMUSG00000026822; ENSMUSG00000026981; ENSMUSG00000054836; | |
| Negative regulation of amine transport | ENSMUSG00000038530; ENSMUSG00000026981; | |
| Negative regulation of anion transport | ENSMUSG00000038530; ENSMUSG00000026981; | |
| Regulation of amino acid transport | ENSMUSG00000038530; ENSMUSG00000026981; | |
| Regulation of amine transport | ENSMUSG00000038530; ENSMUSG00000061353; ENSMUSG00000026981 | |
| Negative regulation of amino acid transport | ENSMUSG00000038530; ENSMUSG00000026981; | |
| Negative regulation of organic acid transport | ENSMUSG00000038530; ENSMUSG00000026981; | |
| Amine transport | ENSMUSG00000038530; ENSMUSG00000061353; ENSMUSG00000026981 | |
| Regulation of monocyte chemotaxis | ENSMUSG00000024190; ENSMUSG00000061353; | |
| Gastrulation | ENSMUSG00000024190; ENSMUSG00000026981; ENSMUSG00000022330 | |
| CBD_1.5 CBD_3 | Regulation of leukocyte differentiation | ENSMUSG00000027523; ENSMUSG00000052713; ENSMUSG00000048546; ENSMUSG00000004951; |
| KEEG | ||
| CBD_0.325 CBD_1.5 | Parkinson disease | ENSMUSG00000064363; ENSMUSG00000064367; ENSMUSG00000027523 |
| Thermogenesis | ENSMUSG00000064363; ENSMUSG00000064367; ENSMUSG00000027523 | |
| CBD_0.75 CBD_3 | RNA degradation | ENSMUSG00000022283; ENSMUSG00000048546; ENSMUSG00000048546 |
| CBD_1.5 CBD_3 | Vascular smooth muscle contraction | ENSMUSG00000027523; ENSMUSG00000022836 |
| Gastric acid secretion | ENSMUSG00000027523; ENSMUSG00000022836; | |
| Platelet activation | ENSMUSG00000027523; ENSMUSG00000022836; | |
| Oxytocin signaling pathway | ENSMUSG00000027523; ENSMUSG00000022836; | |
| Salivary secretion | ENSMUSG00000027523; ENSMUSG00000027447 | |
| Amoebiasis | ENSMUSG00000027523; ENSMUSG00000004951; | |
| Calcium signaling pathway | ENSMUSG00000027523; ENSMUSG00000022836; | |
Only six common genes have been affected by CBD at 6-h and 24-h treatments. They did not enrich any BP category but included genes associated with, e.g., cell chemotaxis, cell migration, acute-phase response, response to interleukin-1 (Dusp1, Il1rn, Saa3), and degradation of extracellular matrix proteins (Mmp13).
qPCR validation results
For the four validated genes, the Spearman rank correlation coefficient ranged from moderate to high, with values from 0.3 for Gm8960 to 0.9 for Ifitm2 (average of 0.625). However, none of these values was statistically significant, likely due to the low number of observations within the dataset. For all genes combined, the correlation coefficient was moderate (0.538) and statistically significant (p = 0.016) (Supplementary File 1). This suggests the reliability of the obtained results, especially when a laboratory-reproduced experiment was used as a validation control.
Discussion
Cannabidiol (CBD) exhibits diverse biological effects in the brain, including the hypothalamus. Research on CBD’s impact on hypothalamic cells revealed its potential to modulate anxiety, stress responses, and other neurobiological functions. For example, in the case of anxiolytic effects, CBD microinjected into the ventromedial hypothalamus (VMH) in rats reduced panic attack-like behaviors and unconditioned fear-induced antinociception, potentially through CB1 receptor signaling (Khan et al. ref. 2020). CBD was also found to enhance neurogenesis and exhibit neuroprotective effects. Repeated CBD administration in chronically stressed mice increased hippocampal neurogenesis and reduced anxiety-like behaviors (Campos et al. ref. 2013). Additionally, CBD influenced histone modifications in the hypothalamus, which can affect gene expression. A recent study revealed that systemic administration of CBD modified the levels of certain histones (e.g., H3 K9ac) in the hypothalamus, indicating potential epigenetic effects of CBD (Pastrana-Trejo et al. ref. 2021). An important action of CBD in the hypothalamus is the modulation of the stress-induced activation of the hypothalamic–pituitary–adrenal (HPA) axis (Viudez-Martínez et al. ref. 2018). These complex responses to CBD in the hypothalamus require a molecular basis that can be observed only in isolated and simplified models, minimizing the complexity and multicellular interactions of brain structures.
In this study, we focused on the effects of CBD on proliferation, viability, and gene expression in a cellular model comprising hypothalamic neurons. We found that CBD stimulated cell viability, especially at longer incubation times (24 h) and at lower or intermediate CBD concentrations. Our results showed a noticeable decrease in apoptotic processes intensity with increasing CBD concentrations, although statistical significance was only observed at the highest CBD concentration. These effects were accompanied by moderate changes in cellular transcriptome profiles, with a greater number of genes affected by higher CBD concentrations and shorter incubation times (indicating a stronger initial response). The transcriptomic response varied between the two time points studied, with more genes upregulated at 6 h and a higher number of downregulated genes following prolonged CBD incubation.
All these observations require in depth analysis and comparison with currently available literature data.
CBD affects hypothalamic neurons’ viability
The observation that CBD increases cell viability in in vitro cultured cells is atypical for healthy unchallenged cells, where CBD’s effect is predominantly associated with viability reduction (Pagano et al. ref. 2020). The results obtained here contradict the findings from previously published data on hypothalamic and other neural cells (di Giacomo et al. ref. 2020; Drummond-Main et al. ref. 2023; Jantas et al. ref. 2024). In most of these studies, CBD alone reduced cell viability primarily at higher concentrations, with no effect observed at lower CBD doses. However, this negative effect on cell viability was reversed when cells were challenged, e.g., with H2O2 to induce oxidative stress, where CBD treatment resulted in increased cell viability (di Giacomo et al. ref. 2020; Jantas et al. ref. 2024). These results indicate that CBD may exert effects that promote neuronal viability, as observed in this study on hypothalamic neurons. It remains unclear why this effect was observed here in unchallenged cells. It is possible that the CBD doses used are within a well-tolerated concertation range for mHypoA-2/12 mouse cell line, thus only the positive or neutral effect of CBD on their proliferation was observed. Alternatively, hypothalamic neural cells may react differently to CBD, similar to some other cell types such as PNT2 cells (Śledziński et al. ref. 2023) or certain specific cancer cell lines (Omer et al. ref. 2024), where CBD stimulates viability under basic culture conditions.
CBD affects cell apoptosis
In this study, a trend toward reduction in apoptotic process intensity (as indicated by 3/7 caspase activity assay) was observed in mHypoA-2/12 cells. This findings, similarly to the viability trend, is unusual for healthy, untreated cells. Nevertheless, reducing apoptosis levels after CBD administration is not uncommon in cells exposed to various stressors. One study demonstrated that CBD protects dopaminergic neurons from oxidative stress and apoptosis by reducing reactive oxygen species (ROS) production and maintaining mitochondrial integrity, independently of CB1 and CB2 cannabinoid receptors (Mendivil-Perez et al. ref. 2023). Furthermore, a review on CBD’s mechanisms in regulating apoptosis and autophagy suggests that these effects may vary based on biological context, cell type, and CBD concentration (Fu et al. ref. 2023). Another study showed that CBD could mitigate perfluorooctane sulfonic acid (PFOS)-induced cardiomyocyte apoptosis by preserving mitochondrial dynamics and metabolic energy homeostasis (Wang et al. ref. 2023). These documented protective and anti-apoptotic effects of CBD are well-supported in the literature, reinforcing the findings of this study.
Transcriptomic evidence on apoptosis and viability regulation
The observed improvement in cell viability along with decreased apoptosis reflects neuroprotective potential and metabolic activity of CBD in hypothalamic neurons. To elucidate the underlying mechanisms at the transcriptome level, we conducted a detailed analysis of biological processes overrepresented by genes differentially expressed following various CBD treatments. Among the numerous processes affected by these treatments, those related to response to oxygen-containing compounds, cellular response to hypoxia, regulation of cell proliferation and cell cycle, regulation of programmed cell death, intrinsic apoptotic signaling pathway by p53 class mediator, DNA damage response, immune system processes, and regulation of metabolic processes were prominently altered in the 6-h treatment (Supplementary File 5; Supplementary File 7). In the 24-h treatment, processes involving regulation of cell migration, immune response, intrinsic apoptosis, and regulation of apoptosis were the most abundantly represented (Supplementary File 6; Supplementary File 9).
The analysis of these processes and the involved genes suggests that one of the important CBD actions in hypothalamic cells is the direct or indirect modulation of cellular apoptosis, particularly intrinsic apoptosis, potentially through the p53 pathway activity modulation (Supplementary File 5; Supplementary File 7). Intrinsic apoptosis involves mitochondrial proteins released in response to various stressors such as ultraviolet radiation, osmotic stress, growth factor withdrawal, chemotherapeutic agents, and natural compounds (Jan and Chaudhry ref. 2019). Internal stimuli like high cytosolic Ca2+ concentrations, hypoxia, oxidative stress, and DNA damage also initiate mitochondrial pathway-mediated apoptosis (Carlsson et al. ref. 2022). CBD’s activities in many of these intracellular processes have been previously documented (Ryan et al. ref. 2009; Russo et al. ref. 2019; Gross et al. ref. 2021; Pereira et al. ref. 2021). For instance, CBD has been shown to induce mitochondrial damage and cytochrome C release in cell lines derived from acute lymphoblastic leukemia of T lineage (T-ALL), and disrupt calcium homeostasis in those cells (Olivas-Aguirre et al. ref. 2019). Additionally, CBD has been demonstrated to induce DNA damage in human-derived cell lines under conditions relevant to consumer exposure (Russo et al. ref. 2019). While examining the key genes that were altered in our dataset and had major contributions to the regulation of apoptotic pathways, we found that the most important hub genes in 6-h treatment were Bbc3 (PUMA; which was downregulated by two lower CBD doses), and Mdm2 (that was upregulated by lowest and intermediate CBD doses). The Bbc3 (Bcl-2-binding component 3), also known as the p53 upregulated modulator of apoptosis (PUMA) is a pro-apoptotic protein, a member of the Bcl-2 protein family (Nakano and Vousden ref. 2001). Upon activation by p53, PUMA interacts with antiapoptotic Bcl-2 family members, thereby releasing Bax and/or Bak to signal apoptosis to the mitochondria (Han et al. ref. 2001). Subsequent mitochondrial dysfunction triggers the caspase cascade, culminating in cell death. In contrast, mouse double minute 2 homolog (Mdm2), also known as E3 ubiquitin-protein ligase, is a critical negative regulator of the p53 tumor suppressor (Mendoza et al. ref. 2014). Mdm2 functions as both an E3 ubiquitin ligase, recognizing the N-terminal trans-activation domain (TAD) of p53, and an inhibitor of p53 transcriptional activation, thereby suppressing p53-mediated apoptosis (de Rozieres et al. ref. 2000). Mdm2 has also been shown to promote proliferation and inhibit apoptosis in pituitary adenoma cells by directly interacting with p53 (Wang et al. ref. 2020). Therefore, the interplay between Bbc3 (PUMA) and Mdm2 seems to be pivotal for CBD-mediated regulation of apoptosis via the p53 pathway. In the anticipated mechanism modulated by CBD, PUMA expression is inhibited directly or indirectly by CBD, thereby suppressing apoptosis, while the upregulation of Mdm2 aims to negatively regulate p53, further mitigating excessive apoptotic activity. These changes in gene expression align with the observed effects of CBD on viability and apoptosis in hypothalamus cells in this study.
The other genes identified in this study as differentially expressed, which may significantly impact apoptosis processes, proliferation regulation, and immune responses, include among others Cdkn1a (p21, Cyclin-dependent kinase inhibitor 1 A), Ndrg1 (N-myc downstream–regulated gene), and Smad3, which contributes to the maintenance of genomic integrity, cell cycle regulation, and apoptosis. The regulation of Cdkn1a and Ndrg1 by p53, and their roles in cellular stress and apoptosis, underscore their critical functions in preserving cellular integrity (Kreis et al. ref. 2019; Schonkeren et al. ref. 2019). Cdkn1a is a direct transcriptional target of p53. Upon cellular stress signals such as DNA damage, p53 binds to the promoter region of Cdkn1a, thereby inducing its expression. This leads to the inhibition of cyclin-dependent kinases (CDKs), resulting in cell cycle arrest at the G1 phase. Such cell cycle arrest allows for DNA damage repair before cell division proceeds, thereby preventing the propagation of mutations (Kreis et al. ref. 2019). Smad3, a member of the SMAD family of proteins involved in TGF-β signaling, interacts with both p53 and Cdkn1a. In some contexts, Smad3 can directly regulate p21 expression, contributing to cell cycle regulation and apoptosis. Additionally, Smad3 signaling can intersect with p53 pathways to modulate cellular responses to stress and DNA damage. Smad3 involvement in TGF-β signaling adds another layer of complexity to the regulation of cell cycle, apoptosis, and stress response. Its interaction with both p53 and Cdkn1a underscores the intricate crosstalk between different signaling pathways in coordinating cellular responses to various stimuli (Wang et al. ref. 2016a).
CBD’s ability to downregulate the expression of genes Cdkn1a, Ndrg1, Smad3, and Bbc2 can have both beneficial and potentially adverse effects. It may offer advantages such as anti-inflammatory and antitumor properties, as well as neuroprotective effects. However, there are risks, including potential disruption of cellular homeostasis, impaired cellular defense mechanisms, and unintended effects on other cellular processes. Further research would be needed to fully understand the consequences of modulating all mentioned genes and to evaluate the therapeutic potential of such interventions, particularly in the context of neurodegenerative diseases.
At 24 h of treatment, the set of genes associated with apoptosis significantly differed, suggesting the alteration of downstream elements of the intrinsic apoptosis mechanism with prolonged CBD treatments. Among the apoptosis-related genes, that were altered predominantly at 24-h treatment, there were for example Hspb1 (Heat shock protein beta-1), which product is known to inhibit apoptosis by stabilizing the actin cytoskeleton and preventing cytochrome c release from mitochondria (Park et al. ref. 2002); Tmbim6 (Transmembrane BAX inhibitor motif containing 6), encoding an anti-apoptotic protein that inhibits mitochondrial calcium uptake and oxidative stress-induced cell death (Kim et al. ref. 2021); Cyld (Cylindromatosis), a tumor suppressor gene that negatively regulates NF-κB signaling, promoting apoptosis by removing K63-linked polyubiquitin chains from target proteins (Fernández-Majada et al. ref. 2016); and Dusp1 (Dual specificity phosphatase 1 also known as MAP kinase phosphatase-1), involved in the negative regulation of MAPK signaling, which can lead to the inhibition of apoptosis (Wang et al. ref. 2016b). These and other apoptosis-related genes altered by CBD in this study suggest that CBD’s effect on intrinsic apoptosis persists over time, though later stages may involve different molecular mechanisms compared to short-term treatments.
The mechanism by which CBD affects cell viability, particularly at the transcriptome level, remains unclear. However, our findings suggest that CBD can stimulate the expression of genes associated with mitochondrial respiratory chain complex I, providing new insights into this process. In our study, the two most significantly affected genes were mt-Nd4 and mt-Nd5. These genes encode NADH-ubiquinone oxidoreductase chain proteins, which are subunits of NADH dehydrogenase (ubiquinone). This enzyme, located in the inner mitochondrial membrane, is the largest of the five complexes in the electron transport chain. NADH dehydrogenase and NAD (nicotinamide adenine dinucleotide) play a crucial role in regulating cell viability through its involvement in energy metabolism and protection against oxidative stress (Čermáková et al. ref. 2021). Our findings align with previous studies on the brains of CBD-treated rats, where increased mitochondrial calcium accumulation enhanced the activity of calcium-sensitive dehydrogenases. This, in turn, promoted NADH availability and enhanced oxidative phosphorylation (Valvassori et al. ref. 2013). Our results further support the broader impact of CBD on mitochondrial metabolism across various cell types (Olivas-Aguirre et al. ref. 2019).
Modulation of stress-like neurons’ responses by CBD
CBD induces cellular responses with transcriptomic signatures resembling known stress responses, primarily evident at the 6-h treatment (initial response). Among the cellular processes overrepresented by DEGs, we identified ones associated with general stress response, cellular response to external stimuli, cellular response to chemical stimuli, response to inorganic substances, interleukin-27-mediated signaling pathway, response to hypoxia, DNA damage response signal transduction mediated by p53 resulting in cell cycle arrest, response to endoplasmic reticulum stress, response to oxidative stress, and others. While the cellular reactions to external and chemical stimuli are obvious, other processes require further analysis and explanation.
An intriguing CBD-associated cellular response is the interleukin-27-mediated signaling pathway, enriched with genes altered by the lowest CBD concentration. Interleukin-27 (IL-27) is recognized as a pivotal regulator of T cell activation and differentiation, influencing T cell responses in autoimmune conditions within the central nervous system (Iwasaki et al. ref. 2015). Recent evidence also suggests its neuroprotective properties in the retina and brain (Nortey et al. ref. 2022). IL-27, secreted by and interacting with infiltrating microglia, macrophages, astrocytes, and neurons, enhances neuronal survival by modulating pro- and anti-inflammatory cytokines, neuroinflammatory pathways, oxidative stress, apoptosis, autophagy, and epigenetic changes (Nortey et al. ref. 2022). The pathway-associated genes such as Oasl1 and Oasl2 were upregulated by lower CBD doses in this research. Oasl proteins (2′−5′-oligoadenylate synthase) may indirectly affect neurodegeneration by regulating inflammatory processes and immune responses in the brain (Ghosh et al. ref. 2019). However, their specific roles and regulatory mechanisms in the context of CBD action, and their implications for CBD consumption, warrant further investigation.
CBD’s effect on extracellular matrix
The genes altered by CBD, primarily in short-term treatments, enrich cellular components such as the extracellular region, extracellular matrix (ECM), and collagen-containing extracellular matrix. The interaction between CBD and the ECM has been previously documented, mainly in different types of fibroblastic cells. It has been shown that CBD, at lower concentrations, increases the production of metalloproteinases (MMPs), while the highest concentrations decrease both the production of MMPs and MMP-2 protein activity (Rawal et al. ref. 2012). A previous transcriptome analysis revealed that CBD pre-treatment enriched genes and functional associations between proteins mainly related to ECM organization and cell interactions in the mouse brain (Prieto et al. ref. 2023). In our previous transcriptomic study on human dermal fibroblasts, we found that CBD affected several genes connected with ECM formation (especially its collagen constituent), which can have serious implications for the fibrosis process (in press). In this study, several genes connected to ECM functioning were altered, the most significant being most likely Mmp3 (downregulated by all 6 h CBD treatments), Mmp13 (downregulated by all 24-h treatments and one 6-h treatment), Timp1 (downregulated by all 6-h treatments), and Col11a1 (downregulated by a single 6-h treatment).
Mmp3 (matrix metalloproteinase-3) is a matrix metalloproteinase, a proteolytic enzyme capable of degrading many ECM components, including proteoglycans, collagens (types III, IV, V, IX, and X), laminin, and fibronectin. Mmp3 plays a key role in tissue remodeling, wound healing, and pathological processes such as inflammation and cancer development (Kandhwal et al. ref. 2022). Its activity is tightly regulated because excessive ECM degradation can lead to diseases such as arthritis and cancer (Mehner et al. ref. 2015). Research suggests that CBD can modulate the activity of MMPs (Gęgotek et al. ref. 2021), which leads to reduced inflammation and protection of the ECM from excessive degradation. Similarly, Mmp13 (matrix metalloproteinase-13) degrades type II collagen as well as other collagen types. It is particularly important in the remodeling of cartilage and bones (Hu and Ecker ref. 2021). One study suggests that CBD derivate can reduce the expression Mmp13 (Jin et al. ref. 2023) and now we further confirm this in this study.
Timp1 (tissue inhibitor of metalloproteinases-1) inhibits the activity of metalloproteinases such as Mmp3 and Mmp13, protecting the ECM from excessive degradation (Brew et al. ref. 2000). Previous research has shown that CBD can increase the expression of Timp1, supporting the protection of the ECM against degradation and potentially benefiting the treatment of inflammation and cancer (Solinas et al. ref. 2012, ref. 2013).
Finally, Col11a1 (collagen type XI alpha 1 chain) is a component of type XI collagen, which supports the structural integrity of cartilage (Shi et al. ref. 2022). In this study, we found that its transcript abundance can be modulated by CBD.
In summary, all mentioned ECM-related genes participate in the process of connective tissue remodeling, where their coordinated action enables the dynamic maintenance and reconstruction of the ECM. Mmp3 and Mmp13 are responsible for the degradation of ECM components, while Timp1 regulates their activity, protecting the ECM against excessive degradation. Col11a1, on the other hand, is important for maintaining the structure of the ECM. This complex interaction is crucial for tissue health and regeneration, and the modulation of this process by CBD can have various implications, particularly related to pathologies such as degenerative joint diseases and cancer.
Transcriptomic insights into the regulation of hypothalamic functions by CBD
In the analysis of genes altered by CBD after 24 h of treatment, we identified those enriched in the dopamine and serotonin biosynthetic processes, as well as in the dopamine receptor signaling pathway. Dopamine is crucial in the hypothalamus, influencing neuroendocrine and autonomic functions. It regulates the HPA axis, with D1 and D2 receptors activating the HPA axis in response to severe stress in rats (Belda and Armario ref. 2009). Given these roles, we focused on genes involved in these pathways and found that Aldh2, protein of which assists in dopamine and serotonin synthesis, was upregulated by the lowest CBD dose after 24 h of treatment. What is more, a Rgs4 gene engaged in the regulation of the dopamine receptor signaling pathway was downregulated in most of the 24 h CBD treatments. Aldh2 (aldehyde dehydrogenase 2 family member) belongs to the aldehyde dehydrogenase family of proteins and is essential in the major oxidative pathway of alcohol metabolism (Adams and Rans ref. 2013; Chen et al. ref. 2014). This links CBD’s effect on neural cells to its presumed positive properties in therapies for alcohol use disorders (De Ternay et al. ref. 2019), though further studies are needed to explore this aspect. Regarding the Rgs4 gene (regulator of G protein signaling 4), its downregulation observed in most 24-h CBD treatments may affect dopamine D2 and D3 receptors, which are major targets for current antipsychotic drugs (Min et al. ref. 2012). Studies using cDNA microarrays have consistently shown decreased RGS4 expression in the prefrontal cortex of subjects with schizophrenia, suggesting implications for CBD in anxiety and mental disorder therapies (McGuire et al. ref. 2018).
Conclusions
The study evaluated the effects of CBD on adult-derived hypothalamic neurons, focusing on cell proliferation, survival, and gene expression. Our results demonstrate that CBD significantly enhances neuronal viability, particularly with extended incubation times of 24 h and at lower to intermediate concentrations. CBD showed a trend toward reducing apoptosis, with statistical significance achieved at the highest concentration. CBD also caused moderate changes in the cellular transcriptome, with more genes affected by higher concentrations and shorter incubation times, indicating a stronger initial response. The gene expression profile varied between time points, with an increase in upregulated genes observed at 6 h and a predominance of downregulated genes with prolonged incubation. Detailed analysis of gene-associated processes showed that CBD altered several cellular responses, resulting in intrinsic apoptosis suppression, immune response modulation, extracellular matrix reorganization, and other cellular mechanism alterations. We found that CBD primarily affects intrinsic apoptosis through p53 modulation, likely by influencing the expression of Bbc3, Mdm2, Cdkn1a, and Smad3 genes. CBD also impacted several genes related to ECM organization, including important MMPs (Mmp-3, Mmp-13) and their inhibitors (such as Timp1), as well as collagen components. Additionally, CBD-influenced genes associated with serotonin and dopamine biosynthesis, including the Aldh2 gene, link CBD’s action in hypothalamic neurons to its proposed benefits in treating alcohol use disorders. These findings confirm the hypothalamus as an important target for CBD actions, highlighting its diverse impact on the central nervous system. Future research will investigate the initiation and underlying mechanisms of these action in more detail.
Supplementary Materials
- Supplementary file1 Primers used for qPCR RNA-Seq validation and obtained correlation coefficients (XLSX 10 KB) (XLSX)
- Supplementary file2 Sequencing reads and mapping statistics (XLSX 15 KB) (XLSX)
- Supplementary file3 Differential expression analysis for cells treated with different concentrations of CBD for 6 and 24 h (including genes regulation tables) (XLSX 17489 KB) (XLSX)
- Supplementary file4 Comparative analysis for genes affected by different CBD concentrations and differ treatment times (including altered genes annotations) (XLSX 24 KB) (XLSX)
- Supplementary file5 Altered genes (by all CBD concentrations separately at 6 h treatment) overrepresentation tests in GO biological processes and KEEG pathways categories (including comparative reanalysis of processes/pathways) (XLSX 444 KB) (XLSX)
- Supplementary file6 Altered genes (by all CBD concentrations separately at 6 h and 24 h treatments) overrepresentation tests in GO cellular components categories (XLSX 22 KB) (XLSX)
- Supplementary file7 Altered genes (by all CBD concentrations jointly at 6 h treatment) overrepresentation tests in GO biological processes (HTML 493 KB) (HTML)
- Supplementary file8 Altered genes (by all CBD concentrations separately at 24 h treatment) overrepresentation tests in GO biological processes and KEEG pathways categories (including comparative reanalysis of processes/pathways) (XLSX 438 KB) (XLSX)
- Supplementary file9 Altered genes (by all CBD concentrations jointly at 24 h treatment) overrepresentation tests in GO biological processes (HTML 323 KB) (HTML)
References
- R Abu-Sawwa, C Stehling. Epidiolex (cannabidiol) primer: frequently asked questions for patients and caregivers. J Pediatr Pharmacol Ther JPPT off J PPAG, 2020. [DOI]
- KE Adams, TS Rans. Adverse reactions to alcohol and alcoholic beverages. Ann Allergy, Asthma Immunol off Publ Am Coll Allergy, Asthma, Immunol, 2013. [DOI]
- U Anand, B Jones, Y Korchev. CBD Effects on TRPV1 signaling pathways in cultured DRG neurons. J Pain Res, 2020. [DOI | PubMed]
- S Anders, PT Pyl, W Huber. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics, 2015. [DOI | PubMed]
- S Armin, S Muenster, M Abood, K Benamar. GPR55 in the brain and chronic neuropathic pain. Behav Brain Res, 2021. [DOI | PubMed]
- A Batalla, J Bos, A Postma, MG Bossong. The impact of cannabidiol on human brain function: a systematic review. Front Pharmacol, 2020. [DOI | PubMed]
- X Belda, A Armario. Dopamine D1 and D2 dopamine receptors regulate immobilization stress-induced activation of the hypothalamus-pituitary-adrenal axis. Psychopharmacology, 2009. [DOI | PubMed]
- Blackburn MR, Vance CO, Morschl E, Wilson CN (2009) Adenosine receptors and inflammation. Handb Exp Pharmacol 215–269. 10.1007/978-3-540-89615-9_8
- K Brew, D Dinakarpandian, H Nagase. Tissue inhibitors of metalloproteinases: evolution, structure and function. Biochim Biophys Acta, 2000. [DOI | PubMed]
- AC Campos, Z Ortega, J Palazuelos. The anxiolytic effect of cannabidiol on chronically stressed mice depends on hippocampal neurogenesis: involvement of the endocannabinoid system. Int J Neuropsychopharmacol, 2013. [DOI | PubMed]
- MJ Carlsson, AS Vollmer, P Demuth. p53 triggers mitochondrial apoptosis following DNA damage-dependent replication stress by the hepatotoxin methyleugenol. Cell Death Dis, 2022. [DOI | PubMed]
- A Castillo, MR Tolón, J Fernández-Ruiz. The neuroprotective effect of cannabidiol in an in vitro model of newborn hypoxic-ischemic brain damage in mice is mediated by CB(2) and adenosine receptors. Neurobiol Dis, 2010. [DOI | PubMed]
- P Čermáková, A Maďarová, P Baráth. Differences in mitochondrial NADH dehydrogenase activities in trypanosomatids. Parasitology, 2021. [DOI | PubMed]
- C-H Chen, JCB Ferreira, ER Gross, D Mochly-Rosen. Targeting aldehyde dehydrogenase 2: new therapeutic opportunities. Physiol Rev, 2014. [DOI | PubMed]
- JAS Crippa, GN Derenusson, TB Ferrari. Neural basis of anxiolytic effects of cannabidiol (CBD) in generalized social anxiety disorder: a preliminary report. J Psychopharmacol, 2010. [DOI | PubMed]
- AM Cruz-Martínez, JG Tejas-Juárez, JM Mancilla-Díaz. CB1 receptors in the paraventricular nucleus of the hypothalamus modulate the release of 5-HT and GABA to stimulate food intake in rats. Eur Neuropsychopharmacol J Eur Coll Neuropsychopharmacol, 2018. [DOI]
- PS Dalvi, A Nazarians-Armavil, S Tung, DD Belsham. Immortalized neurons for the study of hypothalamic function. Am J Physiol Regul Integr Comp Physiol, 2011. [DOI | PubMed]
- P Dalvi, N Loganathan, EK Mcilwraith. Chapter 2 – Hypothalamic cell models. Ulloa-Aguirre A, Tao Y-XBT-CE in H and D, 2021
- JAS de Crippa, AW Zuardi, GEJ Garrido. Effects of cannabidiol (CBD) on regional cerebral blood flow. Neuropsychopharmacol Off Publ Am Coll Neuropsychopharmacol, 2004. [DOI]
- S de Rozieres, R Maya, M Oren, G Lozano. The loss of mdm2 induces p53 mediated apoptosis. Oncogene, 2000. [DOI | PubMed]
- de Fátima dos Santos Sampaio M, de Paiva YB, Sampaio TB et al (2024) Therapeutic applicability of cannabidiol and other phytocannabinoids in epilepsy, multiple sclerosis and Parkinson’s disease and in comorbidity with psychiatric disorders. Basic Clin Pharmacol Toxicol 134:574–601. 10.1111/bcpt.13997
- J De Ternay, M Naassila, M Nourredine. Therapeutic prospects of cannabidiol for alcohol use disorder and alcohol-related damages on the liver and the brain. Front Pharmacol, 2019. [DOI | PubMed]
- A Dobin, CA Davis, F Schlesinger. STAR: ultrafast universal RNA-seq aligner. Bioinformatics, 2013. [DOI | PubMed]
- M Dodt, JT Roehr, R Ahmed, C Dieterich. FLEXBAR-flexible barcode and adapter processing for next-generation sequencing platforms. Biology (Basel), 2012. [DOI | PubMed]
- CD Drummond-Main, Y Ahn, M Kesler. Cannabidiol impairs brain mitochondrial metabolism and neuronal integrity. Cannabis Cannabinoid Res, 2023. [DOI | PubMed]
- V Fernández-Majada, P-S Welz, MA Ermolaeva. The tumour suppressor CYLD regulates the p53 DNA damage response. Nat Commun, 2016. [DOI | PubMed]
- Z Fu, P-Y Zhao, X-P Yang. Cannabidiol regulates apoptosis and autophagy in inflammation and cancer: a review. Front Pharmacol, 2023. [DOI | PubMed]
- SX Ge, EW Son, R Yao. iDEP: an integrated web application for differential expression and pathway analysis of RNA-Seq data. BMC Bioinformatics, 2018. [DOI | PubMed]
- A Gęgotek, S Atalay, A Wroński. Cannabidiol decreases metalloproteinase activity and normalizes angiogenesis factor expression in UVB-irradiated keratinocytes from psoriatic patients. Oxid Med Cell Longev, 2021. [DOI | PubMed]
- A Ghosh, L Shao, P Sampath. Oligoadenylate-synthetase-family protein OASL Inhibits activity of the DNA sensor cGAS during DNA virus infection to limit interferon production. Immunity, 2019. [DOI | PubMed]
- di Giacomo V, Chiavaroli A, Orlando G et al (2020) Neuroprotective and neuromodulatory effects induced by cannabidiol and cannabigerol in rat Hypo-E22 cells and isolated hypothalamus. Antioxidants (Basel, Switzerland) 9. 10.3390/antiox9010071
- E Gonca, F Darıcı. The effect of cannabidiol on ischemia/reperfusion-induced ventricular arrhythmias: the role of adenosine A1 receptors. J Cardiovasc Pharmacol Ther, 2015. [DOI | PubMed]
- C Gross, DA Ramirez, S McGrath, DL Gustafson. Cannabidiol induces apoptosis and perturbs mitochondrial function in human and canine glioma cells. Front Pharmacol, 2021. [DOI | PubMed]
- J Han, C Flemington, AB Houghton. Expression of bbc3, a pro-apoptotic BH3-only gene, is regulated by diverse cell death and survival signals. Proc Natl Acad Sci U S A, 2001. [DOI | PubMed]
- H Heberle, GV Meirelles, FR da Silva. InteractiVenn: a web-based tool for the analysis of sets through Venn diagrams. BMC Bioinformatics, 2015. [DOI | PubMed]
- Hu Q, Ecker M (2021) Overview of MMP-13 as a promising target for the treatment of osteoarthritis. Int J Mol Sci 22. 10.3390/ijms22041742
- Huang Z-L, Zhang Z, Qu W-M (2014) Chapter fourteen – roles of adenosine and its receptors in sleep–wake regulation. In: Mori ABT-IR of N (ed) Adenosine receptors in neurology and psychiatry. Academic Press, pp 349–371
- Y Iwasaki, K Fujio, T Okamura, K Yamamoto. Interleukin-27 in T cell immunity. Int J Mol Sci, 2015. [DOI | PubMed]
- R Jan, G-E-S Chaudhry. Understanding apoptosis and apoptotic pathways targeted cancer therapeutics. Adv Pharm Bull, 2019. [DOI | PubMed]
- Jantas D, Leśkiewicz M, Regulska M et al (2024) Protective effects of cannabidiol (CBD) against qxidative stress, but not excitotoxic-related neuronal cell damage-an in vitro study. Biomolecules 14. 10.3390/biom14050564
- BW Jenkins, HA Spina, K Nicholson. Cannabidiol (CBD) potentiates physiological and behavioral markers of hypothalamic-pituitary-adrenal (HPA) axis responsivity in female and male mice. Psychopharmacology, 2025. [DOI | PubMed]
- Z Jin, Y Zhan, L Zheng. Cannabidiol-loaded poly lactic-co-glycolic acid nanoparticles with improved bioavailability as a potential for osteoarthritis therapeutic. ChemBioChem, 2023. [DOI | PubMed]
- M Kandhwal, T Behl, S Singh. Role of matrix metalloproteinase in wound healing. Am J Transl Res, 2022. [PubMed]
- AU Khan, LL Falconi-Sobrinho, T Dos Anjos-Garcia. Cannabidiol-induced panicolytic-like effects and fear-induced antinociception impairment: the role of the CB(1) receptor in the ventromedial hypothalamus. Psychopharmacology, 2020. [DOI | PubMed]
- S Khosropoor, MS Alavi, L Etemad, A Roohbakhsh. Cannabidiol goes nuclear: the role of PPARγ. Phytomedicine, 2023. [DOI | PubMed]
- H-K Kim, G-H Lee, KR Bhattarai. TMBIM6 (transmembrane BAX inhibitor motif containing 6) enhances autophagy through regulation of lysosomal calcium. Autophagy, 2021. [DOI | PubMed]
- N-N Kreis, F Louwen, J Yuan. The multifaceted p21 (Cip1/Waf1/CDKN1A) in cell differentiation, migration and cancer therapy. Cancers, 2019. [DOI | PubMed]
- JE Lauckner, JB Jensen, H-Y Chen. GPR55 is a cannabinoid receptor that increases intracellular calcium and inhibits M current. Proc Natl Acad Sci U S A, 2008. [DOI | PubMed]
- W Lawn, J Hill, C Hindocha. The acute effects of cannabidiol on the neural correlates of reward anticipation and feedback in healthy volunteers. J Psychopharmacol, 2020. [DOI | PubMed]
- FM Leweke, D Piomelli, F Pahlisch. Cannabidiol enhances anandamide signaling and alleviates psychotic symptoms of schizophrenia. Transl Psychiatry, 2012. [DOI | PubMed]
- Q Li, Q Yu, L Lin. Hypothalamic peroxisome proliferator-activated receptor gamma regulates ghrelin production and food intake. Neuropeptides, 2018. [DOI | PubMed]
- Y Liao, J Wang, EJ Jaehnig. WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res, 2019. [DOI | PubMed]
- GI Liou, JA Auchampach, CJ Hillard. Mediation of cannabidiol anti-inflammation in the retina by equilibrative nucleoside transporter and A2A adenosine receptor. Invest Ophthalmol vis Sci, 2008. [DOI | PubMed]
- MI Love, W Huber, S Anders. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol, 2014. [DOI | PubMed]
- P McGuire, P Robson, WJ Cubala. Cannabidiol (CBD) as an adjunctive therapy in schizophrenia: a multicenter randomized controlled trial. Am J Psychiatry, 2018. [DOI | PubMed]
- C Mehner, E Miller, A Nassar. Tumor cell expression of MMP3 as a prognostic factor for poor survival in pancreatic, pulmonary, and mammary carcinoma. Genes Cancer, 2015. [DOI | PubMed]
- Meissner H, Cascella M (2022) Cannabidiol (CBD). Treasure Island (FL) StatPearls Publishing, 2025
- M Mendivil-Perez, AA Felizardo-Otalvaro, M Jimenez-Del-Rio, C Velez-Pardo. Cannabidiol protects dopaminergic-like neurons against paraquat- and maneb-induced cell death through safeguarding DJ-1CYS(106) and caspase 3 independently of cannabinoid receptors: relevance in Parkinson’s disease. ACS Chem Neurosci, 2023. [DOI | PubMed]
- M Mendoza, G Mandani, J Momand. The MDM2 gene family. Biomol Concepts, 2014. [DOI | PubMed]
- C Min, S-Y Cheong, S-J Cheong. RGS4 exerts inhibitory activities on the signaling of dopamine D2 receptor and D3 receptor through the N-terminal region. Pharmacol Res, 2012. [DOI | PubMed]
- K Nakano, KH Vousden. PUMA, a novel proapoptotic gene, is induced by p53. Mol Cell, 2001. [DOI | PubMed]
- AN Nortey, KN Garces, AS Hackam. Exploring the role of interleukin-27 as a regulator of neuronal survival in central nervous system diseases. Neural Regen Res, 2022. [DOI | PubMed]
- M Olivas-Aguirre, L Torres-López, JS Valle-Reyes. Cannabidiol directly targets mitochondria and disturbs calcium homeostasis in acute lymphoblastic leukemia. Cell Death Dis, 2019. [DOI | PubMed]
- S Omer, S Pathak, M Mansour. Effects of cannabidiol, ∆9-tetrahydrocannabinol, and WIN 55–212–22 on the viability of canine and human non-Hodgkin lymphoma cell lines. Biomolecules, 2024. [DOI | PubMed]
- T Ono, T Yamashita, R Kano. GPR55 contributes to neutrophil recruitment and mechanical pain induction after spinal cord compression in mice. Brain Behav Immun, 2023. [DOI | PubMed]
- S Pagano, M Coniglio, C Valenti. Biological effects of Cannabidiol on normal human healthy cell populations: systematic review of the literature. Biomed Pharmacother, 2020. [DOI | PubMed]
- H-S Park, S-G Cho, CK Kim. Heat shock protein hsp72 is a negative regulator of apoptosis signal-regulating kinase 1. Mol Cell Biol, 2002. [DOI | PubMed]
- JC Pastrana-Trejo, F Duarte-Aké, R Us-Camas. Effects on the post-translational modification of H3K4Me3, H3K9ac, H3K9Me2, H3K27Me3, and H3K36Me2 levels in cerebral cortex, hypothalamus and pons of rats after a systemic administration of cannabidiol: a preliminary study. Cent Nerv Syst Agents Med Chem, 2021. [DOI | PubMed]
- F Patricio, AA Morales-Andrade, A Patricio-Martínez, ID Limón. Cannabidiol as a therapeutic target: evidence of its neuroprotective and neuromodulatory function in Parkinson’s disease. Front Pharmacol, 2020. [DOI | PubMed]
- F Patricio, E Morales Dávila, A Patricio-Martínez. Intrapallidal injection of cannabidiol or a selective GPR55 antagonist decreases motor asymmetry and improves fine motor skills in hemiparkinsonian rats. Front Pharmacol, 2022. [DOI | PubMed]
- SR Pereira, B Hackett, DN O’Driscoll. Cannabidiol modulation of oxidative stress and signalling. Neuronal Signal, 2021. [DOI | PubMed]
- Prieto JP, Fort R, Eastman G et al (2023) Cannabidiol prevents the locomotor sensitization induced by cocaine and caffeine and upregulates genes of extracellular matrix and anti-inflammatory pathways in the nucleus accumbens: a transcriptome-wide analysis. bioRxiv 2023.09.28.560030. 10.1101/2023.09.28.560030
- SY Rawal, MK Dabbous, DA Tipton. Effect of cannabidiol on human gingival fibroblast extracellular matrix metabolism: MMP production and activity, and production of fibronectin and transforming growth factor β. J Periodontal Res, 2012. [DOI | PubMed]
- LBM Resstel, RF Tavares, SFS Lisboa. 5-HT1A receptors are involved in the cannabidiol-induced attenuation of behavioural and cardiovascular responses to acute restraint stress in rats. Br J Pharmacol, 2009. [DOI | PubMed]
- C Russo, F Ferk, M Mišík. Low doses of widely consumed cannabinoids (cannabidiol and cannabidivarin) cause DNA damage and chromosomal aberrations in human-derived cells. Arch Toxicol, 2019. [DOI | PubMed]
- D Ryan, AJ Drysdale, C Lafourcade. Cannabidiol targets mitochondria to regulate intracellular Ca2+ levels. J Neurosci off J Soc Neurosci, 2009. [DOI]
- O Sagredo, JA Ramos, A Decio. Cannabidiol reduced the striatal atrophy caused 3-nitropropionic acid in vivo by mechanisms independent of the activation of cannabinoid, vanilloid TRPV1 and adenosine A2A receptors. Eur J Neurosci, 2007. [DOI | PubMed]
- A Sainz-Cort, D Jimenez-Garrido, E Muñoz-Marron. Opposite roles for cannabidiol and δ-9-tetrahydrocannabinol in psychotomimetic effects of cannabis extracts: a naturalistic controlled study. J Clin Psychopharmacol, 2021. [DOI | PubMed]
- M Scherma, P Masia, V Satta. Brain activity of anandamide: a rewarding bliss?. Acta Pharmacol Sin, 2019. [DOI | PubMed]
- SL Schonkeren, M Massen, R van der Horst. Nervous NDRGs: the N-myc downstream-regulated gene family in the central and peripheral nervous system. Neurogenetics, 2019. [DOI | PubMed]
- W Shi, Z Chen, H Liu. COL11A1 as an novel biomarker for breast cancer with machine learning and immunohistochemistry validation. Front Immunol, 2022. [DOI | PubMed]
- K Singh, B Bhushan, DK Chanchal. Emerging therapeutic potential of cannabidiol (CBD) in neurological disorders: a comprehensive review. Behav Neurol, 2023. [DOI | PubMed]
- P Śledziński, A Nowak-Terpiłowska, P Rzymski. In vitro evidence of selective pro-apoptotic action of the pure cannabidiol and cannabidiol-rich extract. Molecules, 2023. [DOI | PubMed]
- M Solinas, P Massi, AR Cantelmo. Cannabidiol inhibits angiogenesis by multiple mechanisms. Br J Pharmacol, 2012. [DOI | PubMed]
- M Solinas, P Massi, V Cinquina. Cannabidiol, a non-psychoactive cannabinoid compound, inhibits proliferation and invasion in U87-MG and T98G glioma cells through a multitarget effect. PLoS ONE, 2013. [DOI | PubMed]
- M Tham, O Yilmaz, M Alaverdashvili. Allosteric and orthosteric pharmacology of cannabidiol and cannabidiol-dimethylheptyl at the type 1 and type 2 cannabinoid receptors. Br J Pharmacol, 2019. [DOI | PubMed]
- SS Valvassori, DV Bavaresco, G Scaini. Acute and chronic administration of cannabidiol increases mitochondrial complex and creatine kinase activity in the rat brain. Rev Bras Psiquiatr, 2013. [DOI | PubMed]
- A Viudez-Martínez, MS García-Gutiérrez, J Manzanares. Cannabidiol regulates the expression of hypothalamus-pituitary-adrenal axis-related genes in response to acute restraint stress. J Psychopharmacol, 2018. [DOI | PubMed]
- G Wang, Y Yu, C Sun. STAT3 selectively interacts with Smad3 to antagonize TGF-β. Oncogene, 2016. [DOI | PubMed]
- J Wang, J-Y Zhou, D Kho. Role for DUSP1 (dual-specificity protein phosphatase 1) in the regulation of autophagy. Autophagy, 2016. [DOI | PubMed]
- Y Wang, J Zhao, C Zhang. MDM2 promotes the proliferation and inhibits the apoptosis of pituitary adenoma cells by directly interacting with p53. Endokrynol Pol, 2020. [DOI | PubMed]
- X Wang, T Xu, D Luo. Cannabidiol alleviates perfluorooctanesulfonic acid-induced cardiomyocyte apoptosis by maintaining mitochondrial dynamic balance and energy metabolic homeostasis. J Agric Food Chem, 2023. [DOI | PubMed]
- A Zamani, KL Powell, A May, BD Semple. Validation of reference genes for gene expression analysis following experimental traumatic brain injury in a pediatric mouse model. Brain Res Bull, 2020. [DOI | PubMed]
- Y Zhang, H Cao, X Qiu. Neuroprotective effects of adenosine A1 receptor signaling on cognitive impairment induced by chronic intermittent hypoxia in mice. Front Cell Neurosci, 2020. [DOI | PubMed]
