Oncogenic role of TWF2 in human tumors: A pan-cancer analysis
Abstract
To develop effective medicines, researchers must first understand the common and distinct mechanisms that drive oncogenic processes in human cancers. TWF1 and TWF2 belong to the actin-depolymerizing factor homology family. TWF1 has been identified as an important gene in lung, breast, and pancreatic cancer in recent investigations. TWF2’s role in cancer remains largely unknown, no comprehensive pan-cancer studies have been conducted. We utilized the The Cancer Genome Atlas and Gene Expression Omnibus datasets to investigate the role of TWF2 in different types of cancers. TWF2 transcription in cancers and the number of TWF2 mutations were examined as part of our study. We also examined the possible functional pathways involved in TWF2-mediated oncogenicity. Our pan-cancer analysis provided a complete overview of the oncogenic effects of TWF2 in a wide range of human malignancies.
Article type: Research Article
Keywords: TWF2, cancer, survival, prognosis, immune infiltration
Affiliations: Department of Thoracic Oncology, Tianjin Medical University Cancer Institute and Hospital, National Clinical Research Center for Cancer, Key Laboratory of Cancer Prevention and Therapy of Tianjin, Tianjin, China; Tianjin’s Clinical Research Center for Cancer, Tianjin, 300060, China; Department of Oncology, Jining No. 1 People’s Hospital, Jining, 272000, Shandong, China; Department of Thoracic Oncology, Tianjin Medical University Cancer Institute and Hospital, National Clinical Research Center for Cancer, Key Laboratory of Cancer Prevention and Therapy of Tianjin, Tianjin’s Clinical Research Center for Cancer, Tianjin, 300060, China
License: © 2022 Wenjie Liu et al., published by De Gruyter CC BY 4.0 This work is licensed under the Creative Commons Attribution 4.0 International License.
Article links: DOI: 10.1515/med-2022-0547 | PubMed: 36128444 | PMC: PMC9449685
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (4.5 MB)
Introduction
To gain a deeper understanding of the complex process of cancer formation, it is necessary to recognize and characterize new pan-cancer genes. The Cancer Genome Atlas (TCGA) and the Gene Expression Omnibus (GEO) contain a substantial amount of cancer-related functional genomic datasets from various cancer types that can be utilized for pan-cancer analysis [ref. 1,ref. 2,ref. 3] (Table 1).
Table 1: Abbreviation list
| Abbreviation | Definition |
|---|---|
| ACC | Adrenocortical carcinoma |
| BLCA | Bladder urothelial carcinoma |
| BRCA | Breast invasive carcinoma |
| CESC | Cervical squamous cell carcinoma and endocervical adenocarcinoma |
| CHOL | Cholangiocarcinoma |
| COAD | Colon adenocarcinoma |
| READ | Rectum adenocarcinoma |
| DLBC | Lymphoid neoplasm diffuse large B-cell lymphoma |
| ESCA | Esophageal carcinoma |
| GBM | Glioblastoma multiforme |
| HNSC | Head and neck squamous cell carcinoma |
| KICH | Kidney chromophobe |
| KIRC | Kidney renal clear cell carcinoma |
| KIRP | Kidney renal papillary cell carcinoma |
| LAML | Acute myeloid leukemia |
| LGG | Brain lower grade glioma |
| LIHC | Liver hepatocellular carcinoma |
| LUAD | Lung adenocarcinoma |
| LUSC | Lung squamous cell carcinoma |
| MESO | Mesothelioma |
| OV | Ovarian serous cystadenocarcinoma |
| PAAD | Pancreatic adenocarcinoma |
| PCPG | Pheochromocytoma and paraganglioma |
| PRAD | Prostate adenocarcinoma |
| READ | Rectum adenocarcinoma |
| SARC | Sarcoma |
| SKCM | Skin cutaneous melanoma |
| STAD | Stomach adenocarcinoma |
| TGCT | Testicular germ cell tumors |
| THCA | Thyroid carcinoma |
| THYM | Thymoma |
| UCEC | Uterine corpus endometrial carcinoma |
| UCS | Uterine carcinosarcoma |
| UVM | Uveal melanoma |
TWF (Twinfilin), a protein that regulates actin dynamics, is an evolutionarily conserved protein with two Actin-Depolymerizing Factor Homology domains [ref. 4]. TWF proteins bind to actin monomers and heterodimeric capping proteins [ref. 5,ref. 6]. TWF1 (Twinfilin Actin Binding Protein 1) and TWF2 (Twinfilin Actin Binding Protein 2) showed different tissue distributions in mammals, and initial studies have indicated that TWF1 was the major isoform in the developing embryo and non-muscle tissues in most adult mouse, whereas TWF2 was mainly expressed in heart, skeletal muscle, and spleen [ref. 7]. Recent research studies have suggested that TWF1 was highly expressed in various solid tumors and may be regarded as an important gene in lung, pancreas, and breast cancers [ref. 8,ref. 9,ref. 10,ref. 11]. It has previously been demonstrated that the transcription of TWF1 in LUAD tissues is linked to a poorer TNM stage, more lymph node metastases, a larger tumor size, and late clinical staging, among other factors [ref. 9]. Homologous to the actin depolymerizing factor (ADF) as a member of the ADF homology family, TWF2 is a protein with two ADF-homology domains. A6RP, A6r, or PTK9L may alternatively be referred to as TWF2. TWF2’s role in cancer development, on the other hand, remained a mystery.
The transcription profile of TWF2 was investigated in a pan-cancer analysis using data from TCGA and GEO databases. When comparing TWF2 transcription profiles across different types of cancers, the survival status, genetic alterations, and essential biological pathways were all considered. The results of this comprehensive analysis suggest that TWF2 may play a role in the pathogenesis and prognosis of a wide spectrum of malignancies.
Materials and methods
Gene mapping analysis
TWF2 genome location information was acquired from the UCSC (http://genome.ucsc.edu/) genome browser [ref. 12]. The National Center for Biotechnology Information (https://ncbi.nlm.nih.gov/) conducted a conserved functional domain analysis of TWF2 in diverse species.
HPA-gene transcription analysis
The transcription levels of TWF2 under physiological conditions in different cell and tissue types were analyzed using the HPA (https://www.proteinatlas.org) (Human Protein Atlas) database. The internal normalization pipeline was used when combining the HPA and Genotype-Tissue Expression (GTEx) transcriptomics datasets. This consensus dataset consists of normalized expression (nTPM) levels for 55 tissue types.
Gene transcription analysis
In our study, Tumour Immune Estimation Resource 2 (TIMER2) was used to analyze the transcription profile of TWF2 in tumors and adjacent normal tissues. For tumors that lack or contain just a small amount of healthy tissue, we utilized the Gene Expression Profiling Interactive Analysis 2 (GEPIA2) tool to generate box plots from GTEx databases using the GTEx databases, with a p-value threshold of 0.01, with a fold change of log2FC cutoff of one, and “Match TCGA healthy and GTEx data.” All TCGA tumors were analyzed using the HEPIA2 program for TWF2 transcription analysis. Violin and box plots were created using translated expression data log2 [transcripts per million + 1].
The UALCAN (http://ualcan.path.uab.edu) databases were utilized to evaluate tumor omics data and to undertake protein transcription analysis from CPTAC (Clinical Proteomic Tumor Analysis Consortium) databases, which we found to be very useful. A two-tailed p value of less than 0.05 was considered statistically significant [ref. 13]. *p < 0.05; **p < 0.01; ***p < 0.001.
Survival prognosis analysis
All TCGA tumors were utilized to construct TWF2 survival maps and survival plots to assess Disease-Free Survival (DFS) and Overall Survival (OS). To divide the transcription cohorts into low- and high-expression cohorts, a cut-off (50 percent) was utilized [ref. 14]. Log-rank tests were used to analyze the validity of our hypotheses. The Kaplan–Meier plotter was utilized to pool many GEO databases that analyze the Post Progression Survival (PPS), Progression-Free Survival (PFS), First Progression (FP), and OS (http://kmplot.com/analysis/). High and low transcription groups were identified using the “auto-selection of the best cutoff” function and calculated the HR, CI, and log-rank p-values for our study.
Genetic alteration analysis
We collected mutant site information, mutation type, mutation frequency, and Copy Number Alteration (CNA) from all TCGA malignancies using the cBioPortal (https://www.cbioportal.org/) databases.
Correlation of TWF2 and MSI/TMB
Analysis of TWF2 transcription and Microsatellite Instability (MSI) or Tumour Mutational Burden (TMB) in TCGA tumors was performed using the web of “http://sangerbox.com/Tool” [ref. 15]. Spearman’s rank correlation tests were used to obtain the p value and partial correlation value.
Immune infiltration analysis
TWF2 transcription and immune infiltrates were examined in all TCGA tumors using the TIMER2 program. We focused on malignancy-related fibroblasts, neutrophils, T-cell regulatory cells, and endothelial cells. Estimates were made using TIMER, CIBERSORT-ABS, CIBERSORT, QUANTISEQ, EPIC, MCPCOUNTER, XCELL, and TIDE algorithms. The purity-adjusted Spearman’s rank correlation test was used to calculate the p-value and partial correlation (cor) values.
TWF2-related gene enrichment analysis
The interactome network was further analyzed using the STRING website (https://cn.string-db.org/) to determine TWF2 binding proteins by available experiments. We chose the significance of network edges (“evidence”), the active interaction sources (“experiments”), the maximum number of participants to display (“no more than 50 interactors”), and the minimal interaction score [“Medium confidence (0.400)”] to reduce the bias.
In order to extract the 100 highest TWF2-related genes from TCGA cancer and healthy tissue datasets, the program GEPIA2 was employed. In the subsequent stage, we determined the relationship between TWF2 and the target genes that we previously identified using Pearson correlation analysis. All statistical significance values (p-value and correlation coefficient) were computed and presented in the appropriate plot panels for each of the variables. p-Value and partially correlated from the purity-adjusted Spearman’s rank correlation test are displayed as heatmaps of the transcription patterns for the selected genes. GO|KEGG (Gene Ontology | Kyoto Encyclopedia of Genes and Genomes) enrichment and pathway analysis were done by integrating and filtering two sets of data. The BP, CC, and MF, together with KEGG pathway analysis, were visualized using the R packages “clusterProfiler” and “ggplot2” R project software (https://www.r-project.org/) (version 3.6.3) in this investigation [ref. 16].
Results
Gene transcription analysis
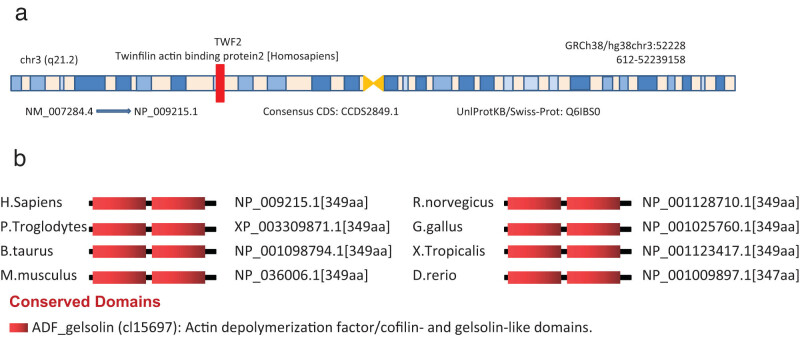
Human TWF2 (NM 007284.4 mRNA or NP 009215.1 protein, Figure A1a) was the focus of this study because of its potential oncogenesis. Figure A1b shows that the ADF gelsolin (cl15697) domain is present in all TWF2 proteins from diverse species, including those from humans and other primate species, such as pan troglodytes, homo sapiens, and bos taurus.
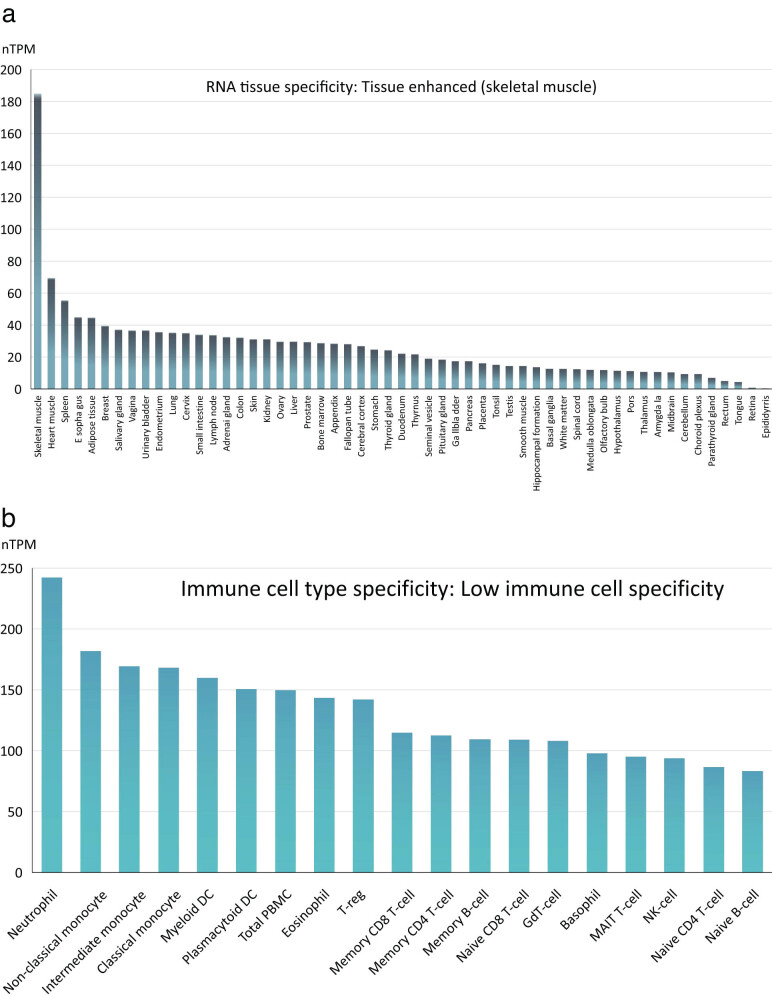
The transcription patterns of TWF2 in various cell lines and non-tumor tissues were investigated in greater depth. We constructed our model based on the GTEx and HPA datasets, as shown in Figure A2a. This comparison revealed that TWF2 was substantially expressed, mainly in the skeletal muscle, granulocytes, and monocytes examined in this study. Analysis of the HPA datasets revealed that neutrophils had the highest TWF2 expression, followed by non-classical monocyte (Figure A2b).
Using TIMER2, we then examined the transcription of TWF2 in tumors and adjacent normal tissues from TCGA datasets. This revealed a significant difference in the transcription between the two tissues. According to Figure 1a, the transcription level for the transcription factor TWF2 in the tumor tissues of UCEC, LIHC, KIRC, BRCA, CHOL, KIRP, and THCA with p < 0.001, READ, BLCA, and ESCA, with p < 0.01, that PCPG, with a p < 0.05, is higher than that of the corresponding control tissues. This was not the case with LUSC and PRAD, which had significantly reduced levels of TWF2 compared to control tissues (p < 0.001). TWF2 transcription variations between tumor and non-tumor tissues were further examined using the GTEx dataset in cases where TCGA data were unavailable. Both TGCT and DLBC had a higher level of transcription in tumor tissues than expected (Figure 1b, p < 0.05). According to the research, TWF2 is overexpressed in the vast majority of human malignancies. A correlation between higher TWF2 transcription and advanced tumor pathological staging was found using the GEPIA2 program in KICH and PAAD (Figure 1c, all p < 0.05).
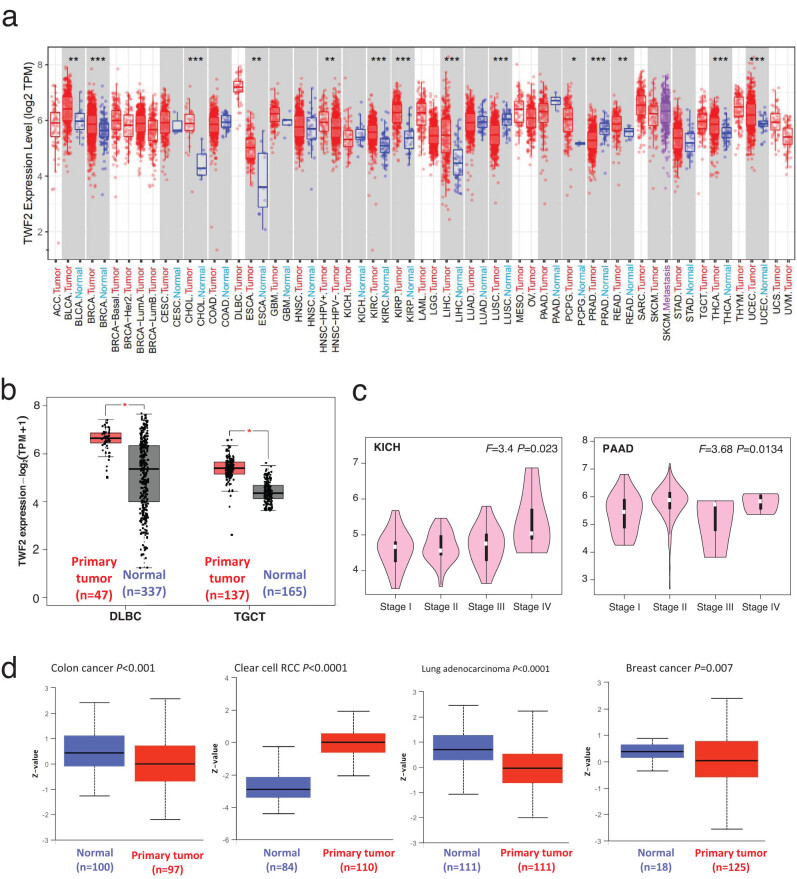
The large-scale proteome program from the CPTAC (National Cancer Institute’s Clinical Proteomic Tumor Analysis Consortium) enabled us to analyze TWF2 at the protein level, in addition to its transcription. When TWF2 expression in clear cell RCC was compared to that in healthy tissues, we determined that it was much higher than that in healthy tissues, whereas it was much lower in colon cancer, LUAD, and breast cancer tissues (Figure 1d; p < 0.01).
Survival analysis
The next step was to examine whether the presence of TWF2 was associated with a better prognosis or longer OS. To divide the cancer cases into high- and low-transcription groups, the TWF2 transcription level was measured. Then, using data from TCGA and GEO datasets, we investigated the relationship between TWF2 transcription and the prognosis of various tumor patients. TWF2 transcription was associated with poorer OS in several forms of cancer, including LAML (p = 0.03), LGG (p < 0.0001), and LIHC (p = 0.01) (Figure 2a). In the DFS study, TWF2 overexpression was associated with a poorer prognosis in HNSC (p = 0.02) and KIRC (p < 0.01) (Figure 2b).
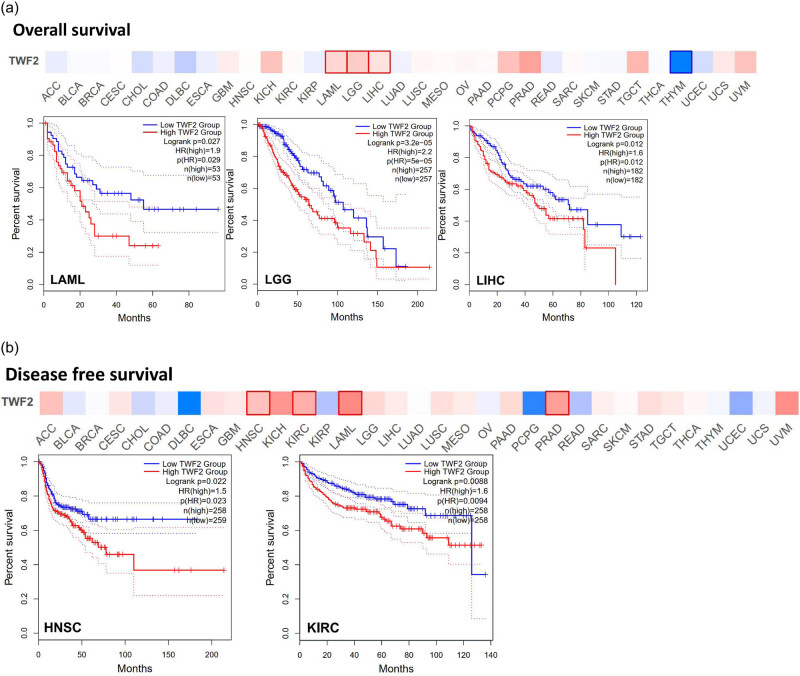
When we utilized the Kaplan–Meier plotter datasets to analyze the survival data, we discovered a correlation between high TWF2 transcription levels and poorer OS and PPS for gastrointestinal cancer. In contrast, we discovered a statistically significant difference between high TWF2 transcription levels and improved OS for lung and ovarian cancers (Figure 3).
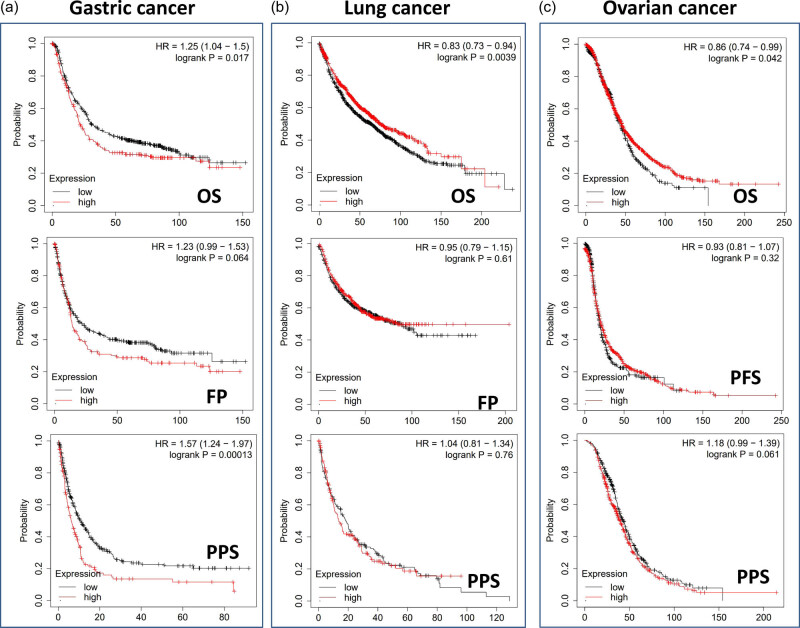
Genetic alterations analysis
Mutations in the human genome are the root cause of nearly all human malignancies. Therefore, we decided to investigate TWF2 genetic changes in human tumor tissues. According to our analysis, the frequency of TWF2 alteration (>4%) is highest in DLBC with “deep deletion” as the predominant kind of alteration. The “amplification” kind of Copy Number Alteration was most common in UCEC, occurring at a frequency of ∼2%. In addition, we noted that “deep deletion” of TWF2 in tumors frequently occurs in almost all cancer types (Figure 4a). Figure 4b further depicts the types, sites, and case numbers of the TWF2 genetic alteration. We found that missense mutation of TWF2 was the main type of gene alteration and P342 alteration, which was detected in three cases of SKCM (Figure 4b).
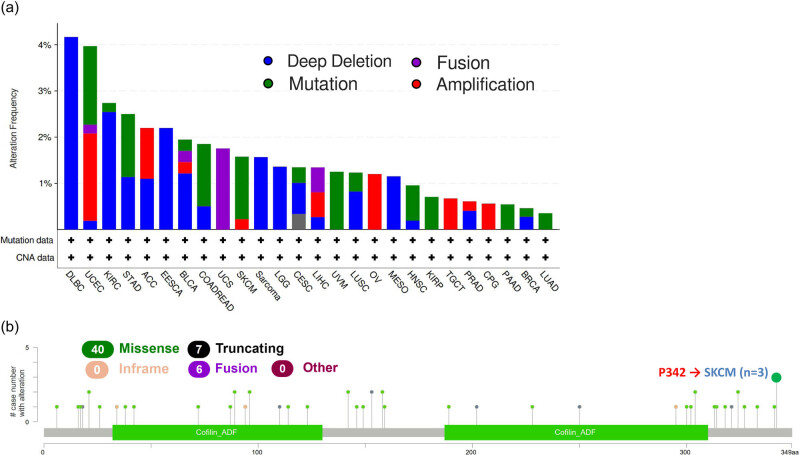
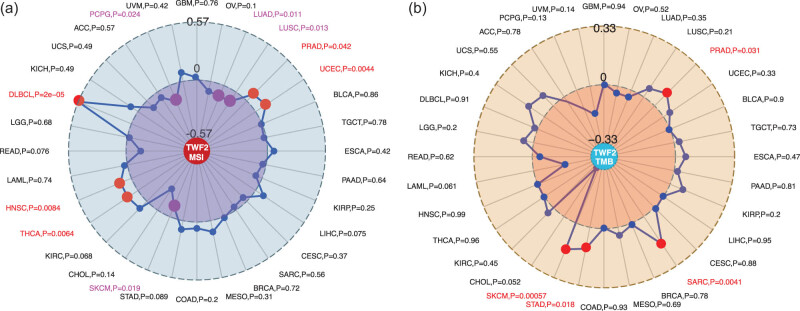
Using TCGA tumor samples, we examined the correlation between the transcription of TWF2, MSI, and TMB. According to our findings, TWF2 transcription was positively correlated with MSI in PRAD, UCEC, THCA, HNSC, and DLBC (all p < 0.05) but was negatively correlated with LUAD, LUSC, SKCM, and PCPG (all p < 0.05) (Figure 5a). Additionally, TWF2 transcription in TMB was associated with STAD, SARC, PRAD, and SKCM (all p < 0.05) (Figure 5b).
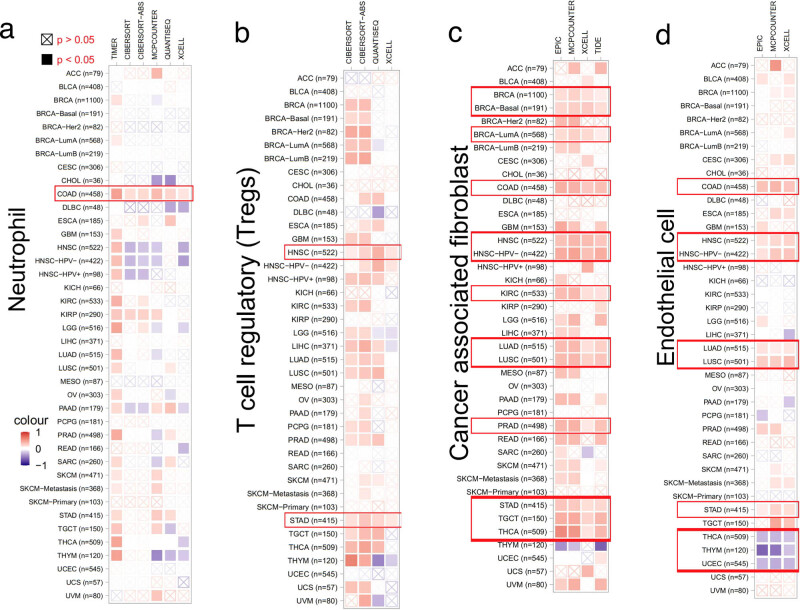
Immune infiltration analysis
We anticipated that changing TWF2 transcript level or genetic changes in TWF2 would influence the tumor-infiltrating immune cell reaction because of the established role of the actin cytoskeleton in cell migration pathways and the involvement of TWF2 in the regulation of actin cytoskeleton structure [ref. 17,ref. 18,ref. 19]. As the results shown in Figure 6, we utilized the TIMER2 program to investigate the link between the infiltration of distinct endothelial and immune cells and TWF2 transcription in different tumor types from TCGA. Intriguingly, we found a positive association between TWF2 transcription and the predicted neutrophil infiltration value in COAD (Figure 6a). There is a positive association between TWF2 transcription and T-cell regulatory in HNSC and STAD (Figure 6b); cancer-associated fibroblasts for BRCA, BRCA-Basal, BRCA-LumA, COAD, HNSC, HNSC-HPV (−), KIRC, LUAD, LUSC, PRAD, STAD, TGCT, and THCA (Figure 6c); and endothelial cell infiltration for COAD, HNSC, HNSC-HPV (−), LUAD, LUSC, and STAD. TWF2 transcription was negatively correlated with endothelial cells in THCA, THYM, and UCEC (Figure 6d).
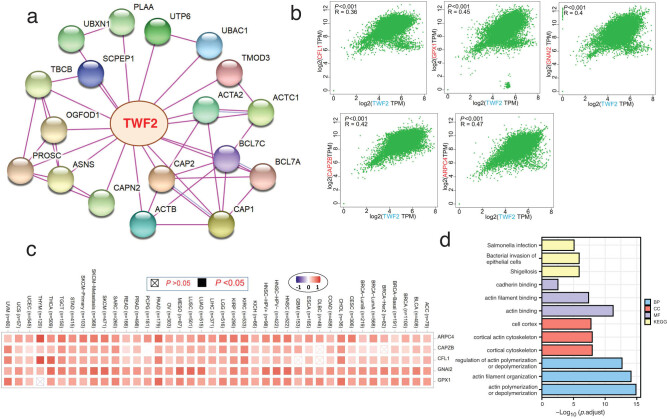
TWF2 partner enrichment analysis
The penultimate phase in our inquiry into the molecular mechanism of the TWF2 gene in cancer and development was to filter out all recognized TWF2-interacting proteins and TWF2 expression-correlated genes, which we did as the last step. STRING allowed us to identify a total of 50 interacted TWF2 proteins that had been previously identified. Figure 7a depicts the interaction network of 18 proteins. TWF2 was shown to be associated with the transcription of the top 100 genes that had been combined from TCGA tumor transcription data. TWF2 transcription was found to be positively linked with the transcription of CFL1 (R = 0.36), GPX1 (R = 0.45), GNAI2 (R = 0.4), CAPZB (R = 0.42), and ARPC4 (R = 0.47) genes (all p < 0.001) (Figure 7b). TWF2 was found to have a high positive connection with the five genes listed above in the majority of tumor types, according to heatmap data (Figure 7c). We integrated the two databases and performed GO and KEGG enrichment studies on the combined results. A search for GO|KEGG pathways found that “shigellosis,” “actin binding,” “cortical cytoskeleton,” and “acting polymerization or depolymerization” were among the top hits, indicating that the influence of TWF2 on tumor etiology maybe mediated through these pathways (Figure 7d).
Discussion
It is unclear whether TWF2 is involved in the oncogenesis of specific tumor types or whether it is involved in more general pathways that contribute to tumor pathogenesis. Therefore, we conducted a TWF2 pan-cancer analysis in this study. So far, there have been few studies related to TWF2 in the field of cancer research. When CHAF1B was knocked down, the protein and mRNA levels of TWF2 were considerably reduced in the human hepatocellular carcinoma cell line HUH-7, thus decreasing the invasion and migration of the tumor [ref. 20].
We utilized TCGA and GEO datasets and various bioinformatics techniques to investigate the oncogenic role of TWF2 in this study. The data from the “HomoloGene” research revealed the conservation of the TWF2 protein structure across species. TWF2 transcription was shown to be higher in cancerous tissues than in normal tissues in a study comparing the two types of tissues. According to survival analysis, TWF2 transcription was found to be associated with poor prognosis in various forms of cancer. Genomic changes in TWF2 have been observed in tumor samples spanning a variety of cancer types, including deletions, amplifications, and mutations. TWF2 transcription, MSI, and tumor mutational burden have been found to be positively associated with various cancers.
TWF2 transcription in tumor samples from patients was higher in UCEC, THCA, READ, PCPG, LIHC, KIRP, KIRC, ESCA, CHOL, BRCA, and BLCA tumors than in the control samples. This was not the case in our study of samples from patients with LUSC and PRAD, which had lower levels of TWF2 expression. Depending on the type of tumor, the TWF2 transcription level can vary significantly. This could be due to its distinct functions and mechanisms in different tumors. Both TWF1 and TWF2 had a significantly higher expression in UCEC, LIHC, BRCA, CHOL, KIRP, THCA, and ESCA tissues [ref. 11]. TWF2 overexpression was also reported to be related to poorer prognosis in patients with cancers such as LAML, LGG, LIHC, HNSC, and KIRC, which express a high level of TWF2. Thus, TWF2 may be one of an indicator for cancer patients’ prognosis, which is supported by these findings.
In contrast, our previous research of TWF1 [ref. 11] showed a quite different mutation profile. The frequency of TWF1 alteration (>5%) is the highest in uterine tumors with “mutation” as the primary type. ACC had the highest incidence of “amplification” type of CNA, with a frequency of ∼4%. TWF1 “amplification” kind in tumors frequently occurs in almost all tumors. This may be caused by the structural differences and tissue distribution differences of TWF1 and TWF2 gene subtypes, and the genetic alteration differences may explain the functional differences of TWF1 and TWF2 proteins and the differences in the expression and regulation of cell signaling pathways [ref. 7].
In this study, TWF2 transcription, MSI, and TMB were linked. Our data showed that TWF2 transcription is positively correlated with the number of endothelial and tumor-associated fibroblast cells that have been deconvolved using several immunodeconvolution methods. TWF2 was positively associated with cancer-associated fibroblasts in STAD and TGCT and with endothelial cells in LUAD, LUSC, and STAD. However, the relationship between TWF1 and corresponding immune cells in the above tumors was largely opposite [ref. 11]. This may mean that there are differences in the roles of TWF1 and TWF2 in the immune microenvironment in these tumors.
Despite the fact that our research generated helpful results, we recognize that it has several limitations. First, with further studying, the relationship between TWF2 and tumor may become clearer and closer. Relevant results are constantly updated, so the results presented in this study may not be comprehensive. Second, a large number of experiments need to be validated and further explored.
TWF2 transcription was found to be statistically associated with clinical prognosis, immune cell infiltration, MSI, and tumor mutation burden in a range of human malignancies, helping to clarify the role of TWF2 in carcinogenesis from various perspectives.
References
- K Tomczak, P Czerwińska, M Wiznerowicz. The cancer genome atlas (TCGA): An immeasurable source of knowledge. Contemp Oncol = Współczesna onkologia, 2015. [PubMed]
- E Clough, T Barrett. The gene expression omnibus database. Methods Mol Biol, 2016. [PubMed]
- A Blum, P Wang, JC Zenklusen. SnapShot: TCGA-analyzed tumors. Cell, 2018. [PubMed]
- S Palmgren, M Vartiainen, P Lappalainen. Twinfilin, a molecular mailman for actin monomers. J Cell Sci, 2002. [PubMed]
- S Palmgren, PJ Ojala, MA Wear, JA Cooper, P Lappalainen. Interactions with PIP2, ADP-actin monomers, and capping protein regulate the activity and localization of yeast twinfilin. J Cell Biol, 2001. [PubMed]
- PJ Ojala, VO Paavilainen, MK Vartiainen, R Tuma, AG Weeds, P Lappalainen. The two ADF-H domains of twinfilin play functionally distinct roles in interactions with actin monomers. Mol Biol Cell, 2002. [PubMed]
- MK Vartiainen, EM Sarkkinen, T Matilainen, M Salminen, P Lappalainen. Mammals have two twinfilin isoforms whose subcellular localizations and tissue distributions are differentially regulated. J Biol Chem, 2003. [PubMed]
- R Samaeekia, V Adorno-Cruz, J Bockhorn, YF Chang, S Huang, A Prat. miR-206 inhibits stemness and metastasis of breast cancer by targeting MKL1/IL11 pathway. Clin Cancer Res An Off J Am Assoc Cancer Research, 2017
- Z Kaishang, P Xue, Z Shaozhong, F Yingying, Z Yan, S Chanjun. Elevated expression of Twinfilin-1 is correlated with inferior prognosis of lung adenocarcinoma. Life Sci, 2018. [PubMed]
- LL Sun, M Cheng, XD Xu. MicroRNA-30c inhibits pancreatic cancer cell proliferation by targeting twinfilin 1 and indicates a poor prognosis. World J Gastroenterol, 2019. [PubMed]
- G Huo, Y Wang, J Chen, Y Song, C Zhang, H Guo. A pan-cancer analysis of the oncogenic role of twinfilin actin binding protein 1 in human tumors. Front Oncol, 2021. [PubMed]
- WJ Kent, CW Sugnet, TS Furey, KM Roskin, TH Pringle, AM Zahler. The human genome browser at UCSC. Genome Res, 2002. [PubMed]
- F Chen, DS Chandrashekar, S Varambally, CJ Creighton. Pan-cancer molecular subtypes revealed by mass-spectrometry-based proteomic characterization of more than 500 human cancers. Nat Commun, 2019. [PubMed]
- Z Tang, B Kang, C Li, T Chen, Z Zhang. GEPIA2: An enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res, 2019. [PubMed]
- R Bonneville, MA Krook, EA Kautto, J Miya, MR Wing, HZ Chen. Landscape of microsatellite instability across 39 cancer types. JCO Precis Oncol, 2017
- G Yu, LG Wang, Y Han, QY He. clusterProfiler: An R package for comparing biological themes among gene clusters. Omics A J Integr Biol, 2012
- SS Wang, W Liu, D Ly, H Xu, L Qu, L Zhang. Tumor-infiltrating B cells: Their role and application in anti-tumor immunity in lung cancer. Cell Mol Immunol, 2019. [PubMed]
- R Sales de Sá, M Miranda Galvis, B Mariz, AA Leite, L Schultz, OP Almeida. Increased tumor immune microenvironment CD3 + and CD20 + lymphocytes predict a better prognosis in oral tongue squamous cell carcinoma. Front Cell Dev Biol, 2020. [PubMed]
- MV Dieci, F Miglietta, V Guarneri. Immune infiltrates in breast cancer: Recent updates and clinical implications. Cells, 2021. [PubMed]
- X Peng, H Fu, J Yin, Q Zhao. CHAF1B knockdown blocks migration in a hepatocellular carcinoma model. Oncol Rep, 2018. [PubMed]
