Involvement of the hemP-hemA-smlt0796-smlt0797 Operon in Hemin Acquisition by Stenotrophomonas maltophilia
Abstract
The hemin acquisition system of Stenotrophomonas maltophilia was elucidated in this study. To identify the TonB-dependent outer membrane receptor for hemin in S. maltophilia, the hemin acquisition systems of Pseudomonas aeruginosa were referenced. PhuR, HasA, and HxuA are three known TonB-dependent outer membrane receptors involved in hemin acquisition by P. aeruginosa. Thus, HemA (Smlt0795) and Smlt2937, the orthologs of PhuR and HasA/HxuA in S. maltophilia, were first considered. KJΔEnt, a stenobactin-null strain, was used as the parental strain for the hemin utilization assay. Deletion of hemA, but not Smlt2937, of KJΔEnt impaired hemin acquisition under iron-depleted conditions, indicating that HemA is the TonB-dependent receptor for hemin uptake. The hemA gene is a member of the hemP-hemA-smlt0796-smlt0797 operon, whose expression was upregulated in a fur mutant and under iron-depleted conditions. The contribution of the hemP-hemA-smlt0796-smlt0797 operon to hemin acquisition was investigated by in-frame deletion mutant construction and hemin utilization assays. Inactivation of hemP, smlt0796, and smlt0797 of KJΔEnt insignificantly affected hemin acquisition under iron-depleted conditions. However, hemP deletion in a fur mutant increased hemin acquisition under iron-depleted conditions. Collectively, we revealed that (i) HemA likely functions as the outer membrane receptor for hemin uptake; (ii) HemP, a predicted transcriptional factor, apparently functions as a repressor of the expression of the hemA transcript; and (iii) in a fur mutant, HemP has a negative impact on hemin acquisition under iron-depleted conditions.
IMPORTANCEStenotrophomonas maltophilia is an emerging multidrug-resistant opportunistic pathogen, increasing the difficulty of treatment of this infection. Iron is a critical element for bacterial viability. Heme is the most abundant iron source in the human host; thus, heme is the major iron source for a pathogen in the infection niche. Blocking iron acquisition from heme can be an alternative strategy to control S. maltophilia infection. Although several hemin acquisition systems have been reported in various pathogens, very little is known about the hemin acquisition systems of S. maltophilia. By in-frame deletion mutant construction and hemin utilization assays, we demonstrated that HemA (Smlt0795) is the TonB-dependent outer membrane receptor for hemin uptake and that HemP (Smlt0794), a predicted transcriptional factor, had a negative impact on hemin acquisition in a fur mutant. The negative regulatory role of HemP in hemin acquisition is first reported.
Article type: Research Article
Keywords: HemA, HemP, hemin, TonB-dependent receptor
Affiliations: a Department of Pathology and Laboratory Medicine, Shin Kong Wu Ho-Su Memorial Hospital, Taipei, Taiwan; b School of Medical Laboratory Science and Biotechnology, Taipei Medical University, Taipei, Taiwan; c School of Medicine, College of Medicine, Fu-Jen Catholic University, New Taipei City, Taiwan; d Department of Biotechnology and Laboratory Science in Medicine, National Yang Ming Chiao Tung University, Taipei, Taiwan; e Department of Medical Laboratory Science and Biotechnology, Asia University, Taichung, Taiwan; f Department of Pathology and Laboratory Medicine, Taipei Veterans General Hospitalgrid.278247.c, Taipei, Taiwan; g School of Medical Laboratory Science and Biotechnology, College of Medical Science and Technology, Taipei Medical University, Taipei, Taiwan; h Division of Infectious Diseases, Department of Medicine, Taipei Veterans General Hospitalgrid.278247.c, Taipei, Taiwan; i School of Medicine, National Yang Ming Chiao Tung University, Taipei, Taiwan
License: Copyright © 2022 Shih et al. CC BY 4.0 This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International license.
Article links: DOI: 10.1128/spectrum.00321-22 | PubMed: 35658602 | PMC: PMC9241770
Relevance: Core topic: mentioned in title and keywords
Full text: PDF (294 KB)
INTRODUCTION
Iron is an essential nutrient for microorganisms. During infection, a coordinated human host cell response limits the availability of iron to the microbes, a process referred to as nutritional immunity (ref. 1). To escape the stress of nutritional immunity that is imposed by the host, pathogens have evolved several strategies to obtain iron during infection. Heme accounts for the majority of the iron pool in vertebrates and is the largest source of iron for bacterial pathogens within the host (ref. 2). Free heme is not readily available in the host, as the majority is located in hemoglobin and sequestered within erythrocytes (ref. 3). To facilitate the use of heme, certain bacteria secrete hemolysin or hemoglobin protease to degrade hemoglobin, which releases hemin (ref. 4). Gram-negative bacteria are generally equipped with TonB-dependent outer membrane proteins (OMPs) as specific receptors for the direct uptake of hemin from host cells. In addition, certain bacteria can synthesize and secrete hemophores, which are small proteins with a high affinity for heme, and take up the hemophore-heme complex via specific TonB-dependent receptors (ref. 5). TonB-dependent receptors, a family of β-barrel proteins, are involved in the uptake and transport of ferric iron-associated complexes. The transport process requires energy and a complex of three inner membrane proteins, TonB-ExbB-ExbD, to transduce this energy to the outer membrane. Periplasmic hemin-binding proteins shuttle hemin from the periplasm into the cytoplasm via inner membrane ABC-type transporters such as HmuU of Ensifer meliloti (ref. 6). In the cytoplasm, hemin is bound to cytoplasmic hemin-binding proteins and is subsequently degraded by heme oxygenase (ref. 7).
There are three known ways for Pseudomonas aeruginosa to utilize hemin as an iron source for growth under iron-depleted conditions: the Phu (Pseudomonas heme uptake) system, the Has (heme assimilation) system, and the Hxu system (ref. 8, ref. 9). The Phu system is the major mechanism by which P. aeruginosa can directly take up hemin from the external environment via the PhuR receptor when cytoplasmic iron is depleted (ref. 8). The Has system is a hemophore-dependent mechanism. When P. aeruginosa grows under iron-depleted conditions, hemophores are endogenously synthesized and are secreted into the extracellular environment. Hemophores specifically capture hemins, and the hemophore-hemin complex is taken up via the HasR receptor (ref. 8). The Hxu system was recently identified and is capable of the direct uptake of hemin as an iron source. However, in P. aeruginosa, it has been verified that the Hxu system plays a major role in signaling the presence of hemin in the extracellular environment and a minor role in hemin acquisition (ref. 9).
The ferric uptake regulator (Fur) is a transcriptional regulator that participates in the regulation of iron homeostasis in several pathogens. Under iron-replete conditions, Fur utilizes Fe2+ as a corepressor, and the Fur-Fe2+ complex binds to a specific sequence, the Fur box, consequently repressing the transcription of iron-responsive genes and operons (ref. 10). When the intracellular ferrous iron level is too low to bind with Fur, free-form Fur dissociates from the Fur box, allowing the RNA polymerase to bind and initiate the transcription of genes related to the acquisition and utilization of iron sources (ref. 11).
Stenotrophomonas maltophilia is a Gram-negative, free-living bacterium commonly found in soil, water, and plants. This bacterium is increasingly recognized as an opportunistic pathogen in immunocompromised and cystic fibrosis patients (ref. 12). S. maltophilia infections are difficult to treat due to the intrinsic and acquired resistance of the bacterium to several antibiotics such as aminoglycosides, β-lactams, and macrolides (ref. 13). The bacterium can colonize many different host environments, which may be driven by its ability to scavenge iron during infection. The putative iron acquisition systems of S. maltophilia have been pointed out by Kalidasan et al. using an in silico approach, mainly siderophore- and/or heme-mediated iron acquisition systems (ref. 14). The siderophore system of S. maltophilia has been well characterized. In response to iron-depleted stress, stenobactin is synthesized in the cytoplasm by enzymes encoded by the entCEBB′FA gene cluster and exported into the extracellular environment for ferric iron acquisition (ref. 15, ref. 16). However, the understanding of the strategies used by S. maltophilia to acquire heme-containing iron is still limited. In this study, we present the first characterization of hemP (Smlt0794) and hemA (Smlt0795) in hemin acquisition by S. maltophilia. HemA is the TonB-dependent outer membrane receptor of hemin. We also show a unique property of HemP as a transcriptional factor in repressing hemP-hemA-smlt0796-smlt0797 operon expression, which is distinct from the functional homologs, HemP/HmuP, in other heme-utilizing bacteria (ref. 17ref. –ref. 20).
RESULTS
S. maltophilia can utilize hemin as an iron source to support growth under iron-depleted conditions.
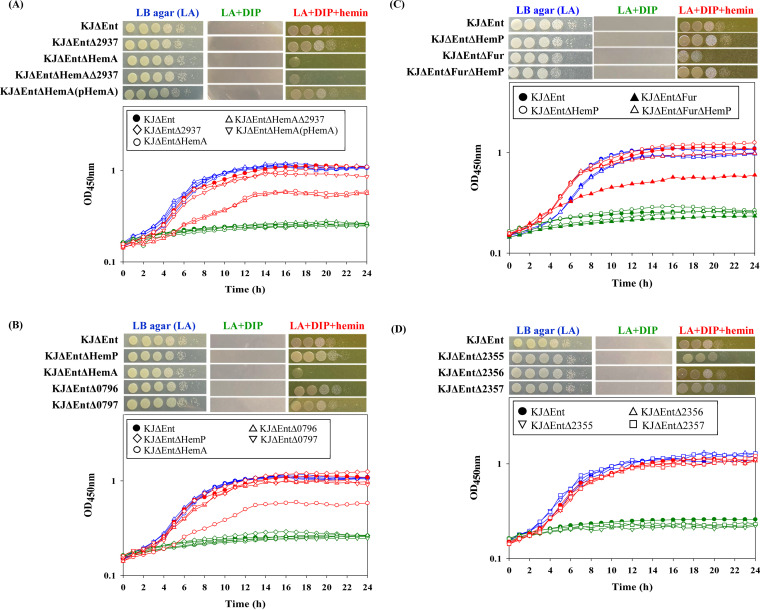
We used iron utilization assays to assess hemin utilization of S. maltophilia KJ under iron-depleted conditions. To avoid the bias caused by stenobactin, KJΔEnt, a stenobactin-null mutant carrying entC and entF deletions (ref. 21), was used. KJΔEnt was unable to grow in 50 μg/mL 2,2′-dipyridyl (DIP)-containing Luria-Bertani (LB) agar but grew when 150 μM hemin was included in the medium (Fig. 1A), indicating that S. maltophilia KJ is capable of utilizing hemin as an iron source for growth under iron-depleted conditions.
Smlt0795 (HemA), rather than Smlt2937, is the TonB-dependent OMP for hemin uptake.
A homolog search of hemophores in the S. maltophilia K279a genome was conducted using the HasA proteins of P. aeruginosa (GenBank accession no. AAG06795), Serratia marcescens (accession no. CAA57068), and Acinetobacter baumannii (accession no. SKV34882) as the queries, but no HasA homolog was identified. Thus, S. maltophilia seems not to synthesize a hemophore for hemin uptake. PhuR (PA4710), HasR (PA3408), and HxuA (PA1302) are three well-known TonB-dependent OMP receptors involved in heme-associated uptake in P. aeruginosa (ref. 9). To identify OMPs for hemin uptake in S. maltophilia, we used the PhuR, HasR, and HxuA proteins of P. aeruginosa as queries for a candidate search. The search revealed that the most similar homolog of PhuR in the S. maltophilia genome was Smlt0795 (37% identity and 52% similarity). Smlt0795 was designated hemA here based on the following results. In addition, when HasR and HxuA were used as queries, the same candidate, Smlt2937, was revealed. Smlt2937 shared 26% identity and 40% similarity with HasR and 27% identity and 41% similarity with HxuA.
The known roles of PhuR, HasR, and HxuA in hemin uptake (ref. 8, ref. 9), together with sequence comparisons, suggested the involvement of hemA and Smlt2937 in hemin uptake. To test this hypothesis, we constructed hemA and Smlt2937 single- and double-deletion mutants from the stenobactin-null strain KJΔEnt (ref. 21), yielding KJΔEntΔHemA, KJΔEntΔ2937, and KJΔEntΔHemAΔ2937. The viability of each mutant and the parental strain (KJΔEnt) was examined in iron-replete medium and iron-depleted medium with and without hemin. All strains tested had comparable growth in LB agar, but growth was stunted in LB agar containing 50 μg/mL DIP. The growth of KJΔEnt was restored when 150 μM hemin was added to DIP-containing LB agar. KJΔEntΔHemA displayed compromised viability in hemin-supplemented medium compared to that of KJΔEnt, but KJΔEntΔ2937 did not (Fig. 1A). Complementation of KJΔEntΔHemA with plasmid pHemA restored growth (Fig. 1A). Furthermore, the deletion of Smlt2397 from the chromosome of KJΔEntΔΗemA did not further compromise viability in hemin-supplemented medium (Fig. 1A), supporting the insignificance of Smlt2397 for hemin uptake. To further quantitatively confirm the role of hemA in hemin acquisition, a growth assay in liquid medium was carried out. The involvement of hemA in hemin acquisition was also supported (Fig. 1A). Consequently, HemA, rather than Smlt2937, is the major TonB-dependent OMP for hemin uptake in S. maltophilia KJ, which is consistent with the previous prediction proposed by Kalidasan et al. (ref. 14).
Role of the hemP-hemA-smlt0796-smlt0797 operon in hemin acquisition.
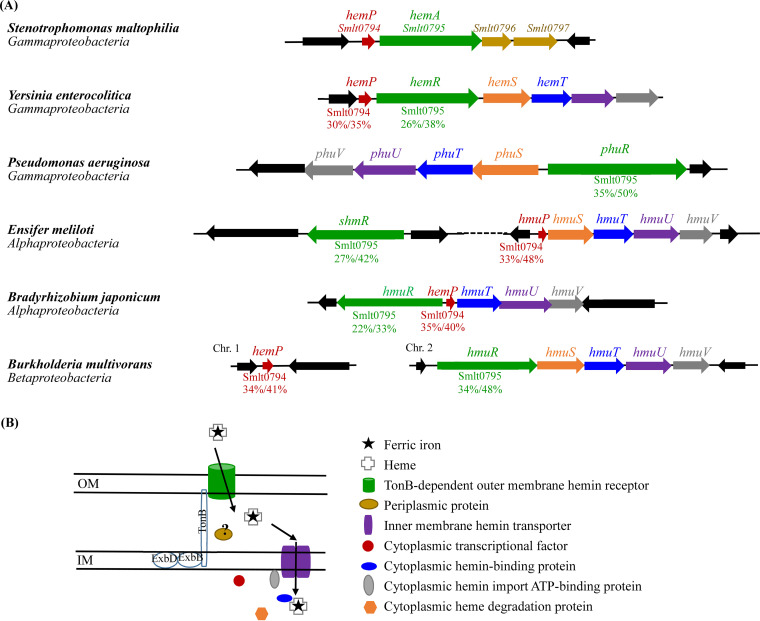
The TonB-dependent OMP genes are usually located near other genes related to iron uptake. The genomic organization surrounding hemA was assessed in the S. maltophilia K279a genome. A four-gene cluster, Smlt0794 to Smlt0797 (Fig. 2A), interested us. The Smlt0794 gene encodes a 66-amino-acid (aa) cytoplasmic protein that shows protein identities of 35%, 33%, 35%, and 34% with HemP of Yersinia enterocolitica, HmuP of Ensifer meliloti, HmuP of Bradyrhizobium japonicum, and HemP of Burkholderia multivorans, respectively (Fig. 3). The HemP and HmuP proteins are transcriptional activators of the genes encoding outer membrane hemin receptors in these heme uptake systems (ref. 17ref. –ref. 20). Thus, we designated the Smlt0794 gene hemP here. HemA is a TonB-dependent outer membrane protein responsible for the uptake of hemin based on our above-mentioned results (Fig. 1A). Smlt0796 and Smlt0797 are predicted to be periplasmic proteins by the CELLO v.2.5 subcellular localization predictor (http://cello.life.nctu.edu.tw/). We also performed signal peptide prediction for Smlt0796 and Smlt0797 with SignalP 6.0 (https://services.healthtech.dtu.dk/service.php?SignalP-6.0). Smlt0797 had a 20-aa predicted signal peptide, but no predicted signal peptide was revealed for Smlt0796.
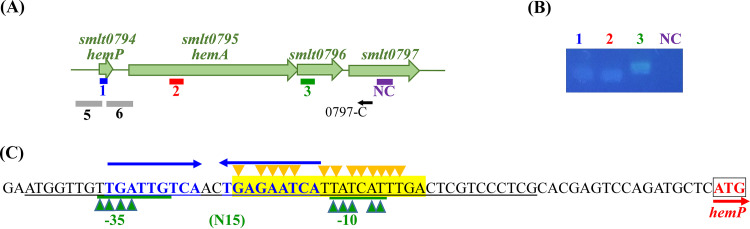
Reverse transcriptase PCR (RT-PCR) was performed to verify whether the four genes formed an operon. Given that the iron uptake system is generally inactive under iron-replete conditions, we prepared DNA-free RNA from the logarithmic growth phase of KJΔFur, a fur isogenic in-frame deletion mutant (ref. 16), grown in LB broth. The results indicated that smlt0794, hemA, smlt0796, and smlt0797 form an operon (Fig. 2B).
To study the function of the hemP-hemA-smlt0796-smlt0797 operon in hemin utilization, the isogenic in-frame single-deletion mutants were individually constructed in KJΔEnt, yielding KJΔEntΔHemP, KJΔEntΔHemA, KJΔEntΔ0796, and KJΔEntΔ0797. KJΔEntΔHemP, KJΔEntΔ0796, and KJΔEntΔ0797 exhibited growth comparable to that of the parental strain in iron-depleted medium supplemented with hemin (Fig. 1B). The results of growth in liquid medium were consistent with this conclusion (Fig. 1B). Thus, only hemA, but not hemP, smlt0796, and smlt0797, is required for hemin uptake in S. maltophilia.
Comparison of hemin acquisition systems of microorganisms.
Given that HemP homologs have been reported in Y. enterocolitica, E. meliloti, B. japonicum, and B. multivorans and that P. aeruginosa and S. maltophilia are often coisolated from cystic fibrosis patients, the hemin acquisition systems in the five microorganisms were compared to the hemP-hemA-smlt0796-smlt0797 operon of S. maltophilia.
The conservation of hemP homologs in gammaproteobacteria is not widespread, although hemP or hmuP homologs are highly conserved in alpha- and betaproteobacteria (ref. 14) (Fig. 3A). Among gammaproteobacteria, no hemP homolog was identified in the genome of P. aeruginosa PAO1, while Y. enterocolitica and S. maltophilia contained hemP homologs (Fig. 3A). Usually, hemin utilization-associated genes are adjacently located, such as hemP-hemRSTUV in Y. enterocolitica, phuR-phuSTUV in P. aeruginosa, hmuP-hmuSTUV in E. meliloti, hmuR-hemPTUV in B. japonicum, and hmuR-hmuSTUV in B. multivorans. However, no hmuSTUV or phuSTUV homologs were found in the hemPA cluster of S. maltophilia (Fig. 3A). Furthermore, Smlt0796 and Smlt0797 did not display significant identities to the known components of other hemin utilization systems (Fig. 3A).
Protein alignments of the HemP and HemA homologs were performed. Although the overall amino acid sequence similarity of the five HemP family proteins was low, a conserved KLILXK motif was found in the C terminus of S. maltophilia HemP, similar to most of those reported for HemP/HmuP proteins (ref. 18) (Fig. 4A). In addition, the C-terminal histidine and tyrosine residues, which have been shown to participate in heme binding (ref. 18), were conserved in S. maltophilia HemP (Fig. 4A). Furthermore, Fig. 4B shows the alignment of the HemA homologs, whereby the majority of the TonB-dependent outer membrane receptors share two motifs, the FRAP and NPNL domains (ref. 22), which are partially conserved in S. maltophilia HemA (Fig. 4B).

Regulation of hemP-hemA-smlt0796-smlt0797 operon expression.
Since a 189-bp intergenic region was present between hemP and hemA, we hypothesized whether a promoter upstream of hemA exists. The promoter-xylE transcriptional fusion constructs pHemPxylE and pHemAxylE were prepared to test this notion (Fig. 2A and Table 1). A promoter activity assay was carried out in XOLNG minimal medium (ref. 23) with and without FeSO4 to assess the putative promoter activities under iron-replete and iron-depleted conditions. Meanwhile, we also investigated the impact of the growth phase on promoter activities; thus, the promoter activities were analyzed in logarithmic phase (8 h) and stationary phase (18 h). In logarithmic phase, KJ(pHemPxylE) displayed weak expression of the xylE gene in iron-replete medium and showed an ~10-fold increase in iron-depleted medium. However, KJ(pHemAxylE) expressed no significant catechol-2,3-dioxygenase (C23O) activity regardless of the presence of iron (Fig. 5A), indicating that the hemP-hemA-smlt0796-smlt0797 operon is driven by the promoter upstream of hemP, which is active under iron-depleted conditions. By comparing the differences between the logarithmic and stationary phases, we noticed that the bacterial growth phase had an insignificant impact on PhemP promoter activity regardless of the iron levels (Fig. 5A).
TABLE 1: Bacterial strains and plasmids used in this study
| Strain or plasmid | Property(ies) and/or genotype | Reference or source |
|---|---|---|
| Strains | ||
| S. maltophilia | ||
| KJ | Clinical S. maltophilia isolate | ref. 30 |
| KJΔEnt | S. maltophilia KJ mutant of entF and entA; ΔentF ΔentA | ref. 21 |
| KJΔEntΔ2937 | S. maltophilia KJ mutant of entF, entA, and smlt2937; ΔentF ΔentA Δsmlt2937 | This study |
| KJΔEntΔHemA | S. maltophilia KJ mutant of entF, entA, and hemA; ΔentF ΔentA ΔhemA | This study |
| KJΔEntΔHemAΔ2937 | S. maltophilia KJ mutant of entF, entA, hemA, and smlt2937; ΔentF ΔentA ΔhemA Δsmlt2937 | This study |
| KJΔEntΔHemP | S. maltophilia KJ mutant of entF, entA, and hemP; ΔentF ΔentA ΔhemP | This study |
| KJΔEntΔ0796 | S. maltophilia KJ mutant of entF, entA, and smlt0796; ΔentF ΔentA Δsmlt0796 | This study |
| KJΔEntΔ0797 | S. maltophilia KJ mutant of entF, entA, and smlt0797; ΔentF ΔentA Δsmlt0797 | This study |
| KJΔEntΔ0796Δ0797 | S. maltophilia KJ mutant of entF, entA, smlt0796, and smlt0797; ΔentF ΔentA ΔhemA Δsmlt0796 Δsmlt0797 | This study |
| KJΔEntΔFur | S. maltophilia KJ mutant of entF, entA, and fur; ΔentF ΔentA Δfur | This study |
| KJΔEntΔFurΔHemP | S. maltophilia KJ mutant of entF, entA, fur, and hemP; ΔentF ΔentA Δfur ΔhemP | This study |
| KJΔEntΔ2355 | S. maltophilia KJ mutant of entF, entA, and smlt2355; ΔentF ΔentA Δsmlt2355 | This study |
| KJΔEntΔ2356 | S. maltophilia KJ mutant of entF, entA, and smlt2356; ΔentF ΔentA Δsmlt2356 | This study |
| KJΔEntΔ2357 | S. maltophilia KJ mutant of entF, entA, and smlt2357; ΔentF ΔentA Δsmlt2357 | This study |
| E. coli | ||
| DH5α | F− ϕ80dlacZΔM15 Δ(lacZYA-argF)U169 deoR recA1 endA1 hsdR17(rK− mK+) phoA supE44λ thi-1 gyrA96 relA1 | Invitrogen |
| S17-1 | λpir+ mating strain | ref. 35 |
| Plasmids | ||
| pEX18Tc | sacB oriT; Tcr | ref. 36 |
| pRK415 | Mobilizable broad-host-range plasmid cloning vector, RK2 origin; Tcr | ref. 37 |
| pRKXylE | pRK415-derived vector for construction of the promoter-xylE transcriptional fusion; the orientation of the xylE gene in this plasmid is opposite that of PlacZ of pRK415; Tcr | ref. 34 |
| pHemA | pRK415 with an intact hemA gene; Tcr | This study |
| pHemPxylE | pRK415 with a 559-bp DNA fragment upstream from the hemP start codon and a PhemP::xylE transcriptional fusion | This study |
| pHemAxylE | pRK415 with a 220-bp DNA fragment upstream from the hemA start codon and a PhemA::xylE transcriptional fusion | This study |
| pΔ2937 | pEX18Tc with an internally deleted smlt2937 gene; Tcr | This study |
| pΔHemA | pEX18Tc with an internally deleted hemA gene; Tcr | This study |
| pΔHemP | pEX18Tc with an internally deleted hemP gene; Tcr | This study |
| pΔ0796 | pEX18Tc with an internally deleted smlt0796 gene; Tcr | This study |
| pΔ0797 | pEX18Tc with an internally deleted smlt0797 gene; Tcr | This study |
| pΔ2355 | pEX18Tc with an internally deleted Smlt2355 gene; Tcr | This study |
| pΔ2356 | pEX18Tc with an internally deleted Smlt2356 gene; Tcr | This study |
| pΔ2357 | pEX18Tc with an internally deleted Smlt2937 gene; Tcr | This study |
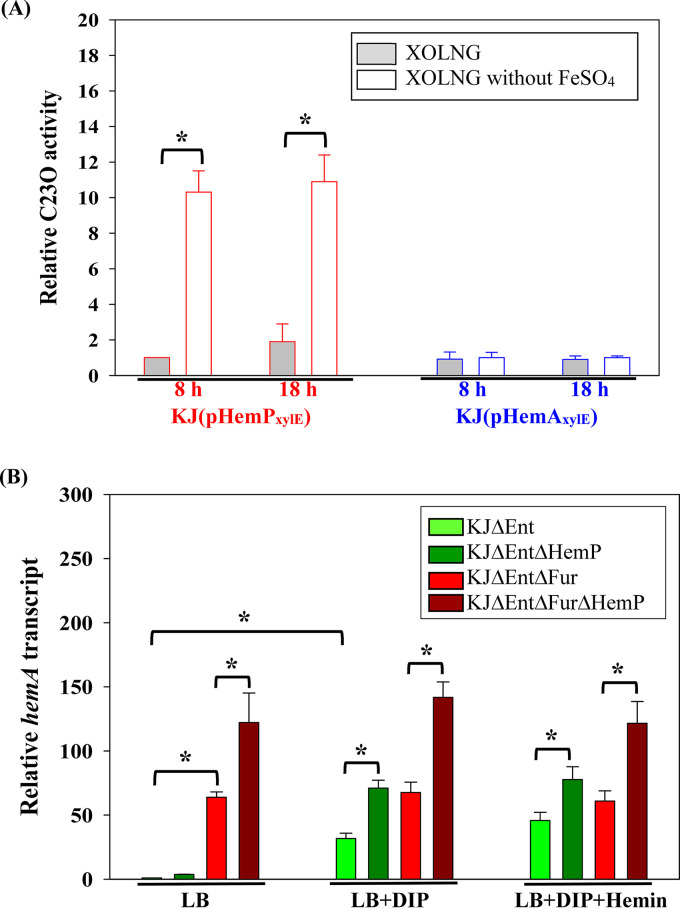
Next, we investigated the role of Fur in regulating the expression of the hemP-hemA-smlt0796-smlt0797 operon using the XOLNG minimal medium system; nevertheless, the fur deletion mutant (KJΔEntΔFur) grew poorly in XOLNG medium. We alternatively used LB medium with and without DIP to represent the iron-depleted and iron-replete conditions, respectively. Under iron-replete conditions, the level of the hemA transcript of KJΔFur had a 63.9-fold ± 4.1-fold increase compared to that of KJ cells (Fig. 5B), indicating that the hemP-hemA-smlt0796-smlt0797 operon is negatively regulated by Fur. The putative “Fur box” of S. maltophilia (ref. 24) was found upstream of the hemP-hemA-smlt0796-smlt0797 operon (Fig. 2C). In response to DIP treatment, the hemA transcript level increased in KJ cells but not in KJΔFur cells (Fig. 5B).
To assess the role of HemP in hemP-hemA-smlt0796-smlt0797 operon expression, we performed a pairwise comparison of hemA transcripts of KJ and KJΔHemP as well as KJΔFur and KJΔFurΔHemP. The assayed conditions included iron-replete medium (LB medium), iron-depleted medium (LB medium plus DIP), and iron-depleted medium supplemented with hemin (LB medium plus DIP and hemin). The inactivation of hemP resulted in an ~2-fold increase in the hemA transcript level in a fur mutant background and/or under DIP-treated conditions (Fig. 5B).
To further assess the roles of Fur and HemP in hemin acquisition under iron-depleted conditions, the viabilities of KJΔEnt, KJΔEntΔHemP, KJΔEntΔFur, and KJΔEntΔFurΔHemP in DIP- and hemin-supplemented media were evaluated. The viability of KJΔEntΔHemP was slightly, but not significantly, better than that of KJΔEnt in DIP- and hemin-supplemented media (Fig. 1C). However, compared to KJΔEnt, KJΔEntΔFur displayed compromised viability in DIP- and hemin-supplemented media, and this compromise was attenuated when the ΔhemP mutation was introduced into the chromosome of KJΔEntΔFur (Fig. 1C). Thus, HemP appears to exert a negative impact on hemin utilization in a fur mutant but not in the parental strain background.
Significance of the hemP-hemA-smlt0796-smlt0797 operon in hemin acquisition when siderophore-mediated iron uptake is active.
To understand the significance of hemP-hemA-smlt0796-smlt0797 when siderophore-mediated iron uptake is active, we constructed a set of mutants in the wild-type KJ background, yielding KJΔHemP, KJΔHemA, KJΔ0796, and KJΔ0797, and a hemin utilization assay was carried out. KJΔHemA, but not KJΔHemP, KJΔ0796, and KJΔ0797, displayed compromised growth in iron-depleted medium supplemented with hemin (see Fig. S1 in the supplemental material), similar to the phenotype observed in strains of a stenobactin-null background (Fig. 1B). This observation implies that the affinity of stenobactin for ferric iron is not high enough to sequester ferric iron from hemin, signifying the importance of HemA in iron acquisition when hemin is the sole iron source available.
The putative HmuV, HmuU, and HmuT homologs (Smlt2357, Smlt2356, and Smlt2355, respectively) are not involved in hemin acquisition.
Once hemin is taken up by HemA and transported into the periplasmic space, a hemin cytoplasmic membrane permease is required for the transport of hemin into the cytosol. Kalidasan et al. conducted a genome-wide analysis of putative iron acquisition systems of S. maltophilia (ref. 14). Based on their analysis, Smlt2357, Smlt2356, and Smlt2355 are homologs of hmuV, hmuU, and hmuT, respectively, which are known inner membrane transporters of hemin in other bacteria (ref. 14). Thus, the involvement of Smlt2357, Smlt2356, and Smlt2355 in hemin acquisition was investigated. The Δsmlt2357, Δsmlt2356, and Δsmlt2355 alleles were introduced into the chromosome of KJΔEnt to generate the mutants KJΔEntΔ2357, KJΔEntΔ2356, and KJΔEntΔ2355. All three mutants displayed viability comparable to that of the parental strain in iron-depleted medium supplemented with hemin (Fig. 1D), suggesting that Smlt2357, Smlt2356, and Smlt2355 are not individually critical for hemin acquisition in S. maltophilia.
DISCUSSION
In most of the known hemin acquisition systems, the genes associated with the transport of hemin across both the outer and inner membranes are generally clustered, such as those of Y. enterocolitica, P. aeruginosa, B. japonicum, and B. multivorans (ref. 17ref. –ref. 20) (Fig. 3A). However, in S. maltophilia, hemA, which encodes the TonB-dependent outer membrane receptor, is clustered with hemP, but these genes do not cluster with the genes associated with the transport of hemin across the inner membrane.
Unlike most gammaproteobacteria, but like alpha- and betaproteobacteria, S. maltophilia, a member of the Gammaproteobacteria, contains the hemP gene. HemP/HmuP is a well-known transcriptional activator essential for the expression of hemin uptake TonB-dependent outer membrane receptors in alpha- and betaproteobacteria (ref. 17ref. –ref. 20). However, in this study, our results suggested that HemP of S. maltophilia acts as a transcriptional repressor, having a negative effect on the hemA transcript level. To our knowledge, the negative regulatory role of S. maltophilia HemP represents a novel property that has not been observed in other HemP superfamily proteins.
Based on the established hemin acquisition systems in other Gram-negative bacteria, hemin is bound to the TonB-dependent OMP and then taken to the periplasmic space. Iron complexes are transported from the periplasm to the cytoplasm through ABC transporters. The ABC transport system generally consists of an inner membrane hemin permease, a cytoplasmic hemin-binding protein, a cytoplasmic hemin import ATP-binding protein, and a cytoplasmic heme degradation protein (Fig. 3B) (ref. 25). Interestingly, unlike FecB and FepB in the ferric citrate and ferrienterobactin acquisition systems of Escherichia coli (ref. 26, ref. 27), no periplasmic hemin-binding proteins were reported in these known hemin acquisition systems (Fig. 3B). These hemin acquisition-associated genes are frequently located in the same operon as or near outer membrane receptor genes. We therefore predicted that Smlt0796 and Smlt0797 are members of the ABC transport system for hemin acquisition in S. maltophilia. However, our results did not support this but rather supported that Smlt0796 and Smlt0797 are not individually crucial for hemin acquisition in S. maltophilia (Fig. 1B). Similar observations were also reported for the hemPRST cluster of Y. enterocolitica (Fig. 3A), in which hemP, hemR, and hemS participate in hemin acquisition but hemT is not necessary for hemin uptake (ref. 17). To obtain insight into their putative functions, we assessed the homology of Smlt0796 and Smlt0797 with known proteins. Nevertheless, no positive results were obtained. Thus, the exact functions of Smlt0796 and Smlt0797 are still unclear at present.
Interestingly, under iron-depleted and hemin-supplemented conditions, HemP functioned as a transcriptional factor negatively regulating the expression of the hemA transcript in either a parental strain (KJΔEnt) or a fur mutant (KJΔEntΔFur) (Fig. 5B). However, with respect to hemin utilization under iron-depleted conditions, the negative impact of HemP on hemin utilization was observed in a fur mutant (KJΔEntΔFur) but not in a parental strain (KJΔEnt) (Fig. 1C). By further inspecting the results shown in Fig. 1C and Fig. 5B, we noticed that the hemA transcript level of KJΔEnt in DIP- and hemin-supplemented LB medium had a 45.8-fold ± 6.3-fold increase compared to that in LB medium (Fig. 5B), and KJΔEnt was able to utilize hemin as the sole iron source to support growth under iron-depleted conditions (Fig. 1C). Thus, the hemA upregulation level in KJΔEnt should be enough to support hemin utilization. However, even though the hemA transcript level of KJΔEntΔFur was higher than that of KJΔEnt (Fig. 5B), the viability of KJΔEntΔFur was worse than that of KJΔEnt in DIP- and hemin-supplemented medium (Fig. 1C). HemA is prerequisite for hemin utilization; nevertheless, functional periplasmic and cytoplasmic hemin transport system components are required for hemin utilization. Based on this rationale, the underlying mechanism responsible for the compromised hemin utilization of KJΔEntΔFur may be the defect in periplasmic or cytoplasmic hemin transport components rather than HemA. Furthermore, our results also supported that the Δfur-mediated reduction in hemin utilization under iron-depleted conditions can be HemP dependent since KJΔEntΔFurΔHemP displayed viability comparable to that of KJΔEnt in DIP- and hemin-supplemented media (Fig. 1C).
HemP (or HmuP) functions as a transcriptional activator and is essential for the expression of hemin uptake outer membrane receptors, which has been reported in Y. enterocolitica, B. japonicum, E. meliloti, and B. multivorans (ref. 17ref. –ref. 20). Distinct from these systems, S. maltophilia HemP negatively regulated the expression of hemA in a fur mutant and/or under iron-depleted conditions (Fig. 5B). This discovery led us to speculate whether the HemP-binding region overlaps or is near the Fur box; thus, Fur-Fur box binding sequesters the opportunity for HemP binding. It has been proposed that HemP/HmuP specifically binds to a conserved DNA motif present upstream of the genes encoding the heme outer membrane receptor. The conserved DNA motif is named as a HmuP-responsive element in E. meliloti, an alphaproteobacterium (ref. 28), and is assigned as a HemP-binding region in B. multivorans, a betaproteobacterium (ref. 20). The HmuP-responsive element is a direct repeat motif that is widely distributed among alpha- and betaproteobacteria but is absent in gammaproteobacteria (ref. 28). In B. multivorans, the HemP-binding region of an inverted repeat frame was revealed by Sato et al. (ref. 20). Thus, we inspected the promoter-containing region of the hemP-hemA-smlt0796-smlt0797 operon and found an inverted repeat sequence (underlined), TGATTGTCAACTGAGAATCA, although it was not similar to the HemP-binding region of B. multivorans, GGCTGCGCGCCCGGGCGCAGCC (ref. 20). This is highly likely to be the HemP-binding region for HemP in S. maltophilia (Fig. 2C) based on the rationale that the binding motif of HemP/HmuP has a direct repeat or an inverted repeat feature. The putative HemP-binding region partially overlaps the Fur box, providing an explanation for the repressor role of HemP in hemA expression in a fur mutant and/or under iron-depleted conditions. Although hemin is an important iron source for invading bacteria, excess hemin is also very toxic to bacteria, most likely due to the generation of hydroxyl radicals by the heme-mediated Fenton reaction (ref. 29). The two-stage negative regulatory circuit for hemin acquisition by Fur and HemP emphasizes the importance of the optimal expression of the hemin acquisition system.
MATERIALS AND METHODS
Bacterial strains, media, plasmids, and primers.
The bacterial strains and plasmids used in this study are listed in Table 1. S. maltophilia KJ has been used as a wild-type strain in our previous series of studies since 2008 (ref. 30), and an array of isogenic deletion mutants of S. maltophilia KJ is available in our stock. The complete genome sequence of S. maltophilia KJ is not available at present. Nevertheless, based on our previous experience, the genome of the KJ strain is highly similar to that of the K279a strain (ref. 31) regarding gene organizations and DNA sequences. Thus, we chose S. maltophilia K279a as a reference strain for bioinformatics analysis and PCR primer design in this study.
Luria-Bertani (LB) medium and XOLNG medium were used as nutrient and minimal media, respectively. XOLNG medium was prepared as described previously (ref. 23). XOLNG medium without FeSO4·7H2O was used as iron-depleted minimal medium. Cells were grown at 37°C with shaking in broth. Hemin (150 μM), DIP (50 μg/mL), and tetracycline (10 μg/mL) were added when required.
Mutant construction.
S. maltophilia chromosomal in-frame deletion mutants were generated by allelic replacement using the suicide vector pEX18Tc, as previously described (ref. 32). Briefly, two DNA fragments containing ~500 bp of the N terminus and C terminus of the intended deletion region were amplified from S. maltophilia KJ by PCR using the designed primer pairs. The primer pairs used are listed in Table S1 in the supplemental material. The PCR amplicons were subsequently cloned into pEX18Tc, generating recombinant plasmids pΔ2397, pΔHemA, pΔHemP, pΔ0796, pΔ0797, pΔ2355, pΔ2356, and pΔ2357 (Table 1). To delete each target gene, the pEX18Tc-derived recombinant plasmids were transferred to S. maltophilia KJ by conjugation. Transconjugant selection, double-crossover mutant selection, and confirmation were carried out as described previously (ref. 32). The correctness of mutants was confirmed by PCR (Fig. S2) and DNA sequencing. Double and triple mutants were constructed from the single mutant sequentially using the same procedure.
Hemin utilization assay.
A hemin utilization assay was designed to investigate whether the assayed strain under iron-depleted conditions can utilize exogenous hemin as the iron source for growth. During LB agar preparation, 50 μg/mL DIP was added to create iron-depleted conditions (ref. 21). If needed, 150 μM hemin (Sigma-Aldrich) was supplemented as an iron source. Logarithmic-phase bacterial cells tested at 2 × 105 CFU/μL were 10-fold serially diluted. Five microliters of the bacterial suspension was spotted onto LB agar containing 50 μg/mL DIP and 150 μM hemin. After a 24-h incubation at 37°C, bacterial viability was imaged.
Construction of complementation plasmid pHemA.
The hemA complementation plasmid pHemA was constructed by amplifying hemA from S. maltophilia KJ with primers HemA-F and HemA-R and cloning into pRK415 under the control of the vector lacZ promoter. The lacZ promoter of pRK415 was constitutively active in S. maltophilia.
Reverse transcriptase PCR.
DNA-free RNA preparation, reverse transcriptase PCRs (RT-PCRs), and data analysis were performed as described previously (ref. 33). S. maltophilia KJΔFur cultured overnight was inoculated into fresh LB medium at an initial optical density at 450 nm (OD450) of 0.15 and grown for 5 h. DNA-free RNA isolated from KJΔFur cells was reverse transcribed using the primer 0797-C (Table S1). The 0797-C-derived cDNA was used as a template for PCR using primers sets HemPQ95-F/R, HemAQ93-F/R, 0796Q120-F/R, and 0797Q108-F/R (Table S1). The 0797Q108-F/R primer set targeted the region downstream of the 0797-C primer and within the C terminus of the smlt0797 gene, which was used as a negative control to rule out DNA contamination.
Construction of promoter-xylE transcriptional fusions.
The DNA segments containing the upstream 559-bp region of hemP and the upstream 220-bp region of hemA were amplified by PCR using the primer sets HemAN-F/R and HemPN-F/HemPN-R and cloned into plasmid pRKXylE (ref. 34) with compatible restriction enzyme sites, yielding pHemPxylE and pHemAxylE, respectively. The correctness of the orientation of the cloned fragments was confirmed by DNA sequencing.
C23O activity determination.
The activities of catechol-2,3-dioxygenase encoded by the xylE gene were measured as described previously (ref. 34). Each assay was run in duplicate at least three times, and the data given are averages.
Bioinformatics assay.
A protein homolog search was carried out using the BLASTp tool from the NCBI (https://blast.ncbi.nlm.nih.gov/Blast.cgi) against the S. maltophilia K279a genome (GenBank accession no. AM743169.1).
Statistical analysis.
Student’s t test was used for comparison of means between the groups. The Bonferroni correction method was applied to adjust the P values.
References
- Nutritional immunity: transition metals at the pathogen-host interface.. Nat Rev Microbiol, 2012. [DOI | PubMed]
- Heme synthesis and acquisition in bacterial pathogens.. J Mol Biol, 2016. [DOI | PubMed]
- Bacterial heme sources: the role of heme, hemoprotein receptors and hemophores.. Curr Opin Microbiol, 2000. [DOI | PubMed]
- In vitro cytotoxic effect of alpha-hemolytic Escherichia coli on human blood granulocytes.. Infect Immun, 1984. [DOI | PubMed]
- Iron acquisition from heme and hemoglobin by a Serratia marcescens extracellular protein.. Proc Natl Acad Sci USA, 1994. [DOI | PubMed]
- The hmuUV genes of Sinorhizobium meliloti 2011 encode the permease and ATPase components of an ABC transport system for the utilization of both haem and the hydroxamate siderophores, ferrrichrome and ferrioxamine B.. Mol Microbiol, 2008. [DOI | PubMed]
- Bacterial heme oxygenases.. Antioxid Redox Signal, 2004. [DOI | PubMed]
- Genetics and regulation of two distinct haem-uptake systems, phu and has, in Pseudomonas aeruginosa.. Microbiology (Reading), 2000. [DOI | PubMed]
- Pseudomonas aeruginosa possesses three distinct systems for sensing and using the host molecule haem.. Environ Microbiol, 2019. [DOI | PubMed]
- Transcriptional regulation by ferric uptake regulator (Fur) in pathogenic bacteria.. Front Cell Infect Microbiol, 2013. [DOI | PubMed]
- Iron acquisition strategies of bacterial pathogens.. Microbiol Spectr, 2016. [DOI]
- Stenotrophomonas maltophilia as an emerging ubiquitous pathogen: looking beyond contemporary antibiotic therapy.. Front Microbiol, 2017. [DOI | PubMed]
- Mechanisms of antimicrobial resistance in Stenotrophomonas maltophilia: a review of current knowledge.. Expert Rev Anti Infect Ther, 2020. [DOI | PubMed]
- Putative iron acquisition systems in Stenotrophomonas maltophilia.. Molecules, 2018. [DOI | PubMed]
- Stenotrophomonas maltophilia produces an EntC-dependent catecholate siderophore that is distinct from enterobactin.. Microbiology (Reading), 2017. [DOI | PubMed]
- AmpR of Stenotrophomonas maltophilia is involved in stenobactin synthesis and enhanced β-lactam resistance in an iron-depleted condition.. J Antimicrob Chemother, 2020. [DOI | PubMed]
- Hemin uptake system of Yersinia enterocolitica: similarities with other TonB-dependent systems in gram-negative bacteria.. EMBO J, 1992. [DOI | PubMed]
- A new small regulatory protein, HmuP, modulates haemin acquisition in Sinorhizobium meliloti.. Microbiology (Reading), 2010. [DOI | PubMed]
- HmuP is a coactivator of Irr-dependent expression of heme utilization genes in Bradyrhizobium japonicum.. J Bacteriol, 2012. [DOI | PubMed]
- The small protein HemP is a transcriptional activator for the hemin uptake operon in Burkholderia multivorans ATCC 17616.. Appl Environ Microbiol, 2017. [DOI | PubMed]
- The involvement of PacIRA system of Stenotrophomonas maltophilia in the uptake of Pseudomonas aeruginosa pyochelin and intraspecies competition for iron acquisition.. J Microbiol Immunol Infect, 2022. [DOI | PubMed]
- Use of heme-protein complexes by the Yersinia enterocolitica HemR receptor: histidine residues are essential for receptor function.. J Bacteriol, 1999. [DOI | PubMed]
- AmpI functions as an iron exporter to alleviate β-lactam-mediated reactive oxygen species stress in Stenotrophomonas maltophilia.. Antimicrob Agents Chemother, 2019. [DOI | PubMed]
- Iron is a signal for Stenotrophomonas maltophilia biofilm formation, oxidative stress response, OMPs expression, and virulence.. Front Microbiol, 2015. [DOI | PubMed]
- Role and regulation of heme iron acquisition in gram-negative pathogens.. Front Cell Infect Microbiol, 2013. [DOI | PubMed]
- Nucleotide sequences of the fecBCDE genes and locations of the proteins suggest a periplasmic-binding-protein-dependent transport mechanism for iron(III) dicitrate in Escherichia coli.. J Bacteriol, 1989. [DOI | PubMed]
- Escherichia coli periplasmic protein FepB binds ferrienterobactin.. Microbiology (Reading), 1995. [DOI | PubMed]
- Highly conserved nucleotide motif present in the 5′-UTR of the heme-receptor gene shmR are required for HmuP-dependent expression of shmR in Ensifer meliloti.. Biometals, 2019. [DOI | PubMed]
- Overcoming the heme paradox: heme toxicity and tolerance in bacterial pathogens.. Infect Immun, 2010. [DOI | PubMed]
- Induction of L1 and L2 β-lactamases of Stenotrophomonas maltophilia.. Antimicrob Agents Chemother, 2008. [DOI | PubMed]
- The complete genome, comparative and functional analysis of Stenotrophomonas maltophilia reveals an organism heavily shielded by drug resistance determinants.. Genome Biol, 2008. [DOI | PubMed]
- AmpDI is involved in expression of the chromosomal L1 and L2 beta-lactamases of Stenotrophomonas maltophilia.. Antimicrob Agents Chemother, 2009. [DOI | PubMed]
- Contribution of resistance-nodulation-division efflux pump operon smeU1-V-W-U2-X to multidrug resistance of Stenotrophomonas maltophilia.. Antimicrob Agents Chemother, 2011. [DOI | PubMed]
- A linkage between SmeIJK efflux pump, cell envelope integrity, and σE-mediated envelope stress response in Stenotrophomonas maltophilia.. PLoS One, 2014. [DOI | PubMed]
- Plasmid vectors for the genetic analysis and manipulation of rhizobia and other Gram-negative bacteria.. Methods Enzymol, 1986. [DOI | PubMed]
- A broad-host-range Flp-FRT recombination system for site-specific excision of chromosomally-located DNA sequences: application for isolation of unmarked Pseudomonas aeruginosa mutants.. Gene, 1998. [DOI | PubMed]
- Improved broad host-range plasmids for DNA cloning in gram-negative bacteria.. Gene, 1988. [DOI | PubMed]
