Pan-Cancer Analysis Reveals Ribonuclease K (RNASEK) as a Potential Prognostic Biomarker in Pancreatic Cancer and a Diagnostic Indicator Across Multiple Human Cancers
Abstract
Background: Cancer is marked by rapid abnormal cell growth, leading to high mortality. The human Ribonuclease K (RNASEK) gene is involved in various cellular processes, such as viral infection, immune response, and maintaining cellular homeostasis. RNASEK, found in metazoans, contributes to tumor development, but the lack of systemic pan-cancer investigation into the diagnostic and prognostic function of RNASEK, epigenetic regulation, and interaction with the immune cell infiltration remains unclear. This study investigated RNASEK as a potential pan-cancer biomarker.
Methodology: Public databases such as Tumor Immune Estimation Resource (TIMER), Gene Expression Profiling Interactive Analysis (GEPIA), and University of Alabama at Birmingham Cancer Data Analysis Portal (UALCAN) assessed RNASEK expression patterns. Gene Expression Omnibus (GEO) datasets validated these expressions. UALCAN examined RNASEK’s expression differences, DNA methylation, and clinical features. TIMER analyzed SNHG8 expression about immune cell infiltration, while prognosis was evaluated through GEPIA, UALCAN, and Kaplan-Meier (KM) Plotter. cBioPortal reviewed the genetic alterations of RNASEK.
Results: Our study revealed a significant upregulation of RNASEK (P < 0.05) in six cancers: bladder (BLCA), cholangiocarcinoma (CHOL), esophageal (ESCA), head/neck squamous cell (HNSC), liver (LIHC), and thyroid (THCA). This was accompanied by notable hypomethylation in BLCA, HNSC, LIHC, and Uterine Corpus Endometrial Carcinoma (UCEC), associated with increased RNASEK expression. Significant differences (P < 0.05) were noted between stage 1 and stage 3 in ESCA, HNSC, and THCAas well as significant differences (P < 0.05) in HNSC between African-American and Asian populations. Additionally, age-related expression differences were significant (P < 0.05) in HNSC across young (21-40 years), middle-aged (41-60 years), and older (61-80 years) groups. A weak positive correlation (P < 0.05) existed between RNASEK expression and various immune cell infiltrations such as B cells, CD8+ T-cells, CD4+ T-cells, macrophages, neutrophils, and dendritic cells in patients with BLCA, ESCA, HNSC, and LIHC, while THCA presented moderate negative correlations with CD4+ T-cells and neutrophils. Moreover, High RNASEK expression indicated a good prognosis in pancreatic adenocarcinoma (PAAD) (hazard ratio (HR) 0.49, P = 0.0007). RNASEK was altered in less than 1% (95 samples out of 10,967 samples) across various tumor types. The highest alteration rates were identified as significant deletions in miscellaneous neuroepithelial tumors, one case out of 31 cases (3.23%), amplifications in sarcoma, four cases out of 255 cases (1.96%), and mutations in endometrial cancer, which is two cases out of 586 (0.34%).
Conclusions: In conclusion, this study’s pan-cancer analysis revealed that RNASEK could be a potential diagnostic biomarker in six cancer types, including BLCA, CHOL, ESCA, HNSC, LIHC, and THCA, and as a prognostic biomarker in PAAD.
Article type: Research Article
Keywords: diagnostic biomarker, paad, pan-cancer, prognostic biomarker, rnasek
License: Copyright © 2025, Hassan Elsheikh et al. CC BY 4.0 This is an open access article distributed under the terms of the Creative Commons Attribution License CC-BY 4.0., which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Article links: DOI: 10.7759/cureus.84574 | PubMed: 40546550 | PMC: PMC12180778
Relevance: Relevant: mentioned in keywords or abstract
Full text: PDF (6.1 MB)
Introduction
The World Health Organization (WHO) estimates that there will be 19.3 million new cancer diagnoses and approximately 10 million cancer deaths globally in 2020. The incidence and mortality of cancer are increasing rapidly annually, and major obstacles affect the quality of life in every country worldwide, with no cure for cancer [ref. 1]. Considering the complexity of tumorigenesis and the successful application of cancer biomarkers [ref. 2,ref. 3], it is important to be curious about any gene of interest and to explore its prognostic value and underlying molecular mechanisms in cancers [ref. 4]. Cancer research has developed exponentially during the past decade because of the revolutionary high-throughput sequencing technology and the data it generates [ref. 5].
RNAs are key for gene expression and cell functions. Some RNases break down RNA molecules in various biological processes [ref. 6]. Several human RNases are involved in pathogenic conditions, such as inflammatory disorders and autoimmune diseases [ref. 7], or the inhibition of tumor growth and metastasis [ref. 8], Additionally, RNA degradation by ribonucleases is a key process that affects cell growth, apoptosis, and angiogenesis and plays a critical role in the development of human cancers [ref. 9].
The ribonuclease K (RNASEK) is located on chromosome 17 and belongs to a highly conserved protein family in metazoans [ref. 6, ref. 10], whose normal function appears to center on intracellular transport and endocytosis [ref. 11]. It is a small protein with 95-101 amino acid polypeptide chains. The involvement of RNASEK in regulating essential cellular processes, especially those connected to endocytosis, viral entry, and elements of immune signaling, is demonstrated. RNASEK localizes to the endoplasmic reticulum and cell surface, and it contributes to clathrin-mediated endocytosis and viral uptake through interactions with elements like the vacuolar-type H⁺-ATPase proton pump [ref. 11,ref. 12].
In addition, previous studies have supported the role of RNASEK in carcinogenesis and metastasis [ref. 12]. Moreover, the RNASEK gene is transcribed via expressed sequence tag (EST) analysis in the vast majority of normal and cancerous human tissues; 75% of these sequences are present in cancer tissues, and 25% of them are present in normal tissues [ref. 6]. This makes RNASEK a potential diagnostic and/or prognostic biomarker in various types of cancers [ref. 5]. Adamopoulos et al. reported eight novel alternatively spliced variants of the human RNASEK gene identified in 55 human cancer cell lines via 3′ RACE and next-generation sequencing (NGS) methodologies [ref. 5]. These eight novel RNASEK transcripts are widely expressed in different types of human cell lines, including brain tumors, lung adenocarcinomas, leukemia, lymphomas, melanoma, normal pancreas, normal embryonic kidney, ovarian cancer, cervical cancer, prostate cancer, renal cell carcinoma, bladder cancer (BCLA), hepatocellular carcinoma, gastric adenocarcinoma, colon cancer, brain tumors, lung adenocarcinoma, normal pancreas, normal embryonic kidney, and neck and head squamous cell carcinoma [ref. 5]. However, despite the progress made in considering the role of RNASEK in human cancers, the possible participation remains unclear and needs to be further elucidated. While previous studies have suggested the role of RANASK in tumor progression and carcinogenesis, comprehensive pan-cancer research on this gene has not been conducted. Therefore, we conducted a pan-cancer analysis to evaluate RNASEK expression levels across various tumor types, aiming to assess its potential role as a diagnostic and prognostic biomarker.
Materials and methods
Gene expression profile of RNASEK across cancers
TIMER, also known as the Tumor Immune Estimation Resource (https://cistrome.shinyapps.io/timer/, accessed in June 2024), is a database that enables the analysis of differential gene expression between tumor and normal tissues [ref. 13]. Additionally, we used the GEPIA database (Gene Expression Profiling Interactive Analysis) (http://gepia.cancer-pku.cn/, accessed in June 2024), a web-based tool used for exploring RNASEK expression patterns across different cancers and differences among all stages, via TCGA datasets (The Cancer Genome Atlas Program) [ref. 14]. UALCAN (University of Alabama at Birmingham Cancer Data Analysis Portal) (UALCAN (uab.edu, accessed in June 2024) is a user-friendly and powerful OMICS tool for cancer transcriptome analysis that includes RNA-seq expression data from the TCGA dataset [ref. 15]. Furthermore, we used the UALCAN database to identify the relationships between RNASEK expression and different pathological and clinical parameters, such as cancer stage, race, and age. The expression pattern of RNASEK was evaluated across various cancers using TIMER, GEPIA, and UALCAN databases.
Methylation analysis of RNASEK
The UALCAN database is a website that allows users to evaluate the epigenetic regulation of gene expression by promoter methylation. Thus, we searched UALCAN (UALCAN (uab.edu, accessed in June 2024) to explore RNASEK promoter DNA methylation levels in various cancers and to determine the differences between tumors and normal tissues [ref. 15].
Immune cell infiltration analysis
TIMER (https://cistrome.shinyapps.io/timer/, accessed in June 2024) [ref. 13] is an analysis network of tumor immune cell infiltration. It was used to analyze the relationship between RNASEK expression and tumor-infiltrating immune cells in tumors across various cancer types. We subsequently evaluated significant (P < 0.05) correlations between the abundance of immune cells, such as B cells, CD8+ T-cells, CD4+ T-cells, neutrophils, macrophages, and dendritic cells, and RNASEK expression.
Overall survival of RNASEK
We investigated the overall survival of RNASEK across various human cancers via three databases. The first database is GEPIA (http://gepia.cancer-pku.cn/, accessed in June 2024). GEPIA is a website that can provide customizable functions such as patient survival analysis. The second database we used was UALCAN (UALCAN (uab.edu), which is an interactive website that can be used to explore patient survival rates among different cancers [ref. 15]. Finally, we used Kaplan-Meier (KM) plotter (https://kmplot.com/analysis/), an online survival analysis tool used to assess the relationship between the expression of every gene, including mRNAs, miRNAs, proteins, DNA, and the survival outcomes of patients in 3,500+ samples from 21 tumor types [ref. 16].
Genetic alteration analysis of RNASEK
To perform a pan-cancer alteration frequency analysis of RNASEK, we used the cBioPortal database (cBioPortal for Cancer Genomics, accessed in June 2024) [ref. 17]. It is a web platform that allows for visualization, exploration, and analysis of cancer data. cBioPortal allows us to uncover the genetic alterations of RNASEK in 10,967 tumor samples from 32 studies, specifically the TCGA Pan-Cancer Atlas studies.
Validation analysis
We performed differential expression analysis via the GEO2R tool (https://www.ncbi.nlm.nih.gov/geo/, accessed in June 2024) [ref. 18], an interactive web tool that allows researchers to compare two or more groups of samples to identify differentially expressed genes. Differential expression gene profiles were visualized via volcano plots generated via the bioinformatics.com.cn platform (http://www.bioinformatics.com.cn/srplot) [ref. 19], which is an online platform that is mainly used for data analysis and visualization. Statistical analyses were performed based on the criteria of |Log2FC| > 1 and adj P-value < 0.05 for the identification of the differentially expressed genes.
Results
Gene expression profile of RNASEK across various types of cancers
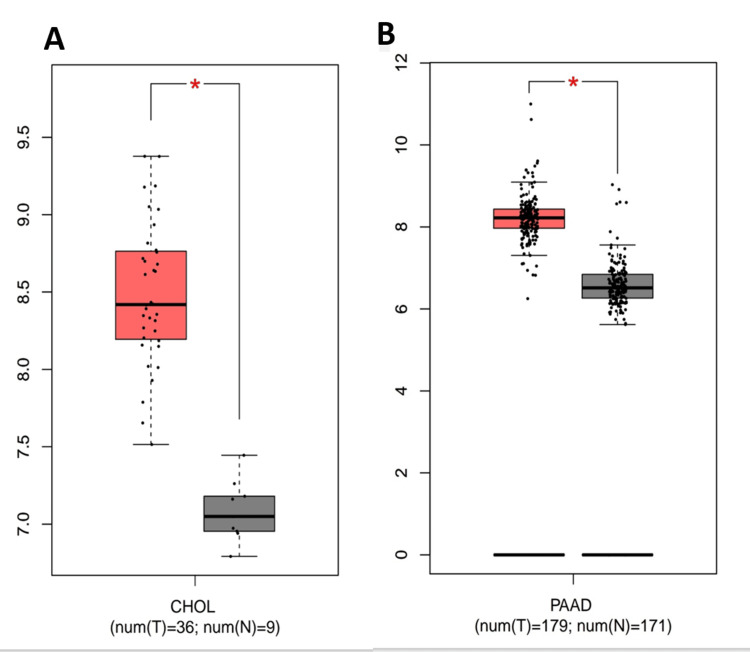
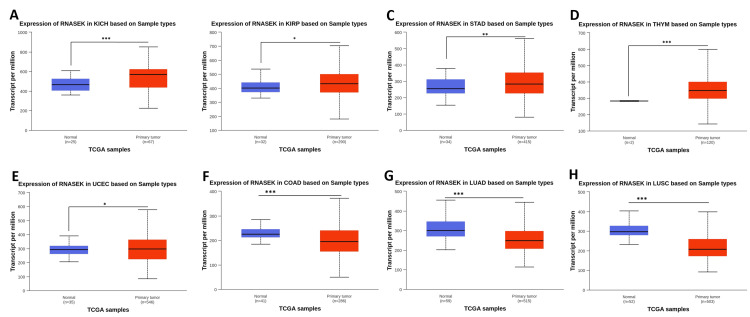
We investigated the differential expression of RNASEK between normal and tumor tissues via the TIMER database (result shown in Figure 1A). RNASEK was significantly upregulated in seven types of cancer (P < 0.05), including BLCA, CHOL, ESCA, HNSC, LIHC, KICH, and THCA. However, highly significant (P < 0.05) upregulation of skin cutaneous melanoma (SKCM) was observed between the tumor and metastasis groups. Furthermore, there is considerable downregulation in four types of cancer, including colon adenocarcinoma (COAD), kidney renal clear cell carcinoma (KIRC), lung adenocarcinoma (LUAD), and lung squamous cell carcinoma (LUSC). The results from the GEPIA database revealed that RNASEK was significantly upregulated (P < 0.05) in cholangiocarcinoma (CHOL) and pancreatic adenocarcinoma (PAAD) (Appendix A). Furthermore, we investigated RNASEK expression via the UALCAN database. The results revealed that RNASEK was significantly (P < 0.05) upregulated in 11 types of cancers, including BLCA, CHOL, ESCA, HNSC, LIHC, and THCA (Figures 1B–1G), KICH, KIRP, STAD, THYM, and UCEC (Appendix B). Significant downregulation was detected in three cancers: COAD, LUAD, and LUSC (Appendix B). By cross-referencing the results confirmed by two databases, TIMER and UALCAN, we concluded that RNASEK was commonly upregulated in six types of cancers, including BLCA, CHOL, ESCA, HNSC, LIHC, and THCA (Figures 1A–1G).
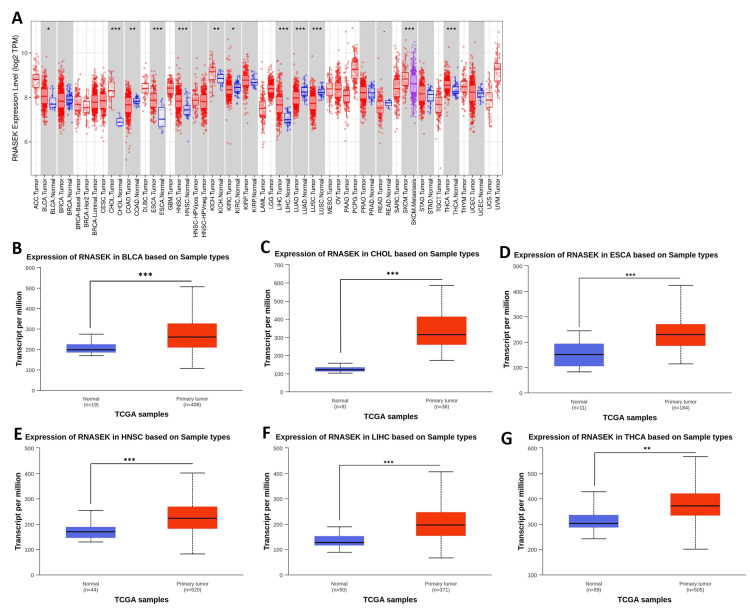
RNASEK expression and DNA methylation
We examined the DNA methylation levels of RNASEK in various tumors using the UALCAN database. The results revealed that the methylation levels of RNASEK in BLCA, HNSC, LIHC, and UCEC tissues were significantly (P < 0.05) lower than those in normal tissues, which may explain the high RNASEK expression in these tumors (Figures 2A–2D).
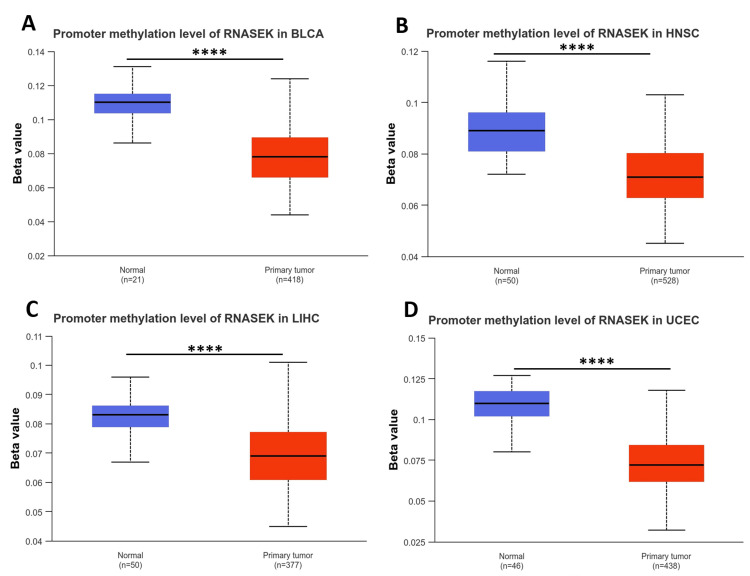
Clinicopathological parameters (stage, race, and age) associated with RNASEK expression
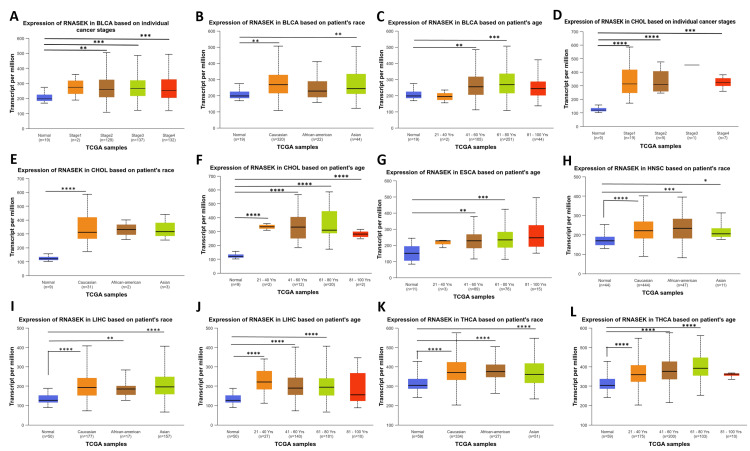
We studied RNASEK expression concerning pathological and clinical features, including stage, race, and age, in six types of cancers (BLCA, CHOL, ESCA, HNSC, LIHC, and THCA) via the UALCAN database. Our study observed RNASEK expression variations across cancer stages in ESCA, with significant differences (P < 0.001) noted between stages 1 and 2, as well as 1 and 3 (Figure 3A). In HNSC, stage 1 showed marked differences (P < 0.05) when compared to stages 3 and 4, and stage 2 differed from stage 4 (Figure 3B). LIHC patients exhibited significant differences (P < 0.05) between stages 2 and 3 against stage 4 (Figure 3C). In THCA, stage 1 was significantly different (P < 0.05) from stage 3 (Figure 3D). As shown in Figure 3E, racial disparities were evident in ESCA between African-Americans and Asian individuals (P < 0.05). Age-wise, we examined the impact of age on RNASEK expression; individuals were categorized into five groups: normal, young (21-40 years), middle-aged (41-60 years), older (61-80 years), and elderly (81-100 years). RNASEK expression significantly varied in HNSC across age groups, which revealed differences (P < 0.05) in the young-aged compared to middle-aged and older people (Figure 3F). The comparison between tumor tissues and stages of cancers, race, and age in six cancers under study was provided in Appendix C.
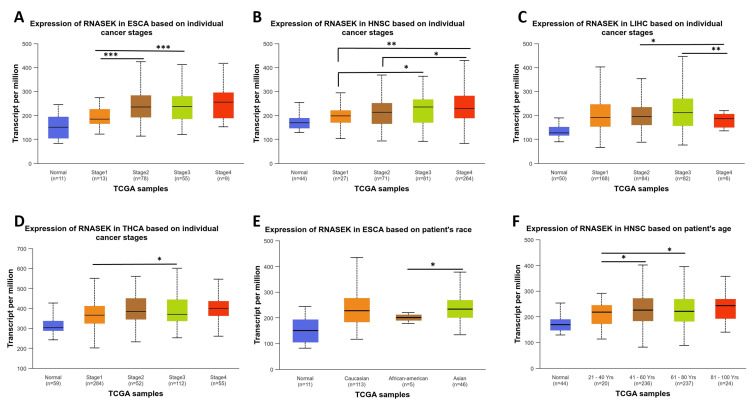
Correlation between RNASEK expression and immune cell infiltration
We investigated the correlation between RNASEK expression and immune cells (B cells, CD8+ T-cells, CD4+ T-cells, macrophages, neutrophils, and dendritic cells) in six cancers, including BLCA, CHOL, ESCA, HNSC, LIHC, and THCA. As shown in Figure 4, our study found no significant correlation in CHOL, while identifying a significant (P < 0.05) weak positive correlation between RNASEK expression and macrophages in BLCA (correlation = 0.11, P = 0.036), ESCA (correlation = 0.236, P = 0.0014), HNSC (correlation = 0.233, P = 2.20E-07), and LIHC (correlation = 0.181, P = 0.0008). Additionally, we observed weak positive correlations with B cells in HNSC (correlation = 0.1, P = 0.048), and LIHC (correlation = 0.175, P = 0.0011), and CD8+ T-cells in HNSC (correlation = 0.108, P = 0.018), LIHC (correlation = 0.202, P = 0.0001), and THCA (correlation = 0.183, P = 4.71E-05). CD4+ T-cells had a weak positive correlation with ESCA (correlation = 0.207, P = 0.005). Neutrophils showed weak positive correlations in BLCA (correlation = 0.136, P = 0.009) and LIHC (correlation = 0.124, P = 0.021). Dendritic cells have a weak positive correlation with BLCA (correlation = 0.229, P = 9.93E-06), HNSC (correlation = 0.127, P = 0.0053), and LIHC (correlation = 0.166, P = 0.0021). Notably, THCA exhibited moderate negative correlations with several immune cells, including macrophages (correlation = -0.444, P = 4.90E-25), B cells (correlation = -0.379, P = 7.01E-18), CD4+ T-cells (correlation = -0.537, P = 7.30E-38), neutrophils (correlation = -0.439, P = 2.27E-24), and dendritic cells (correlation = -0.366, P = 8.23E-17).
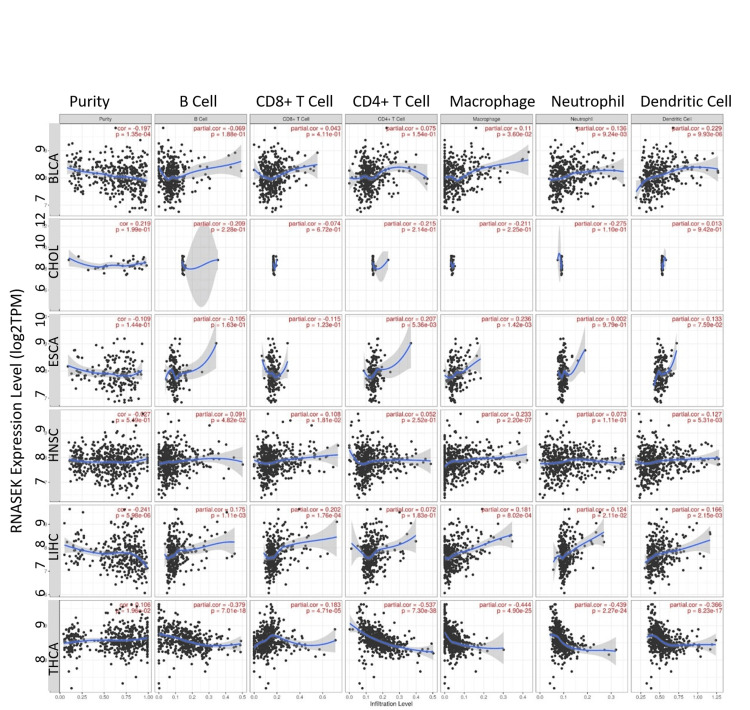
Overall survival analysis of RNASEK across all human cancers
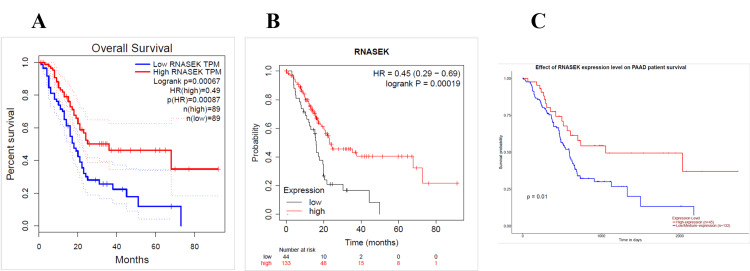
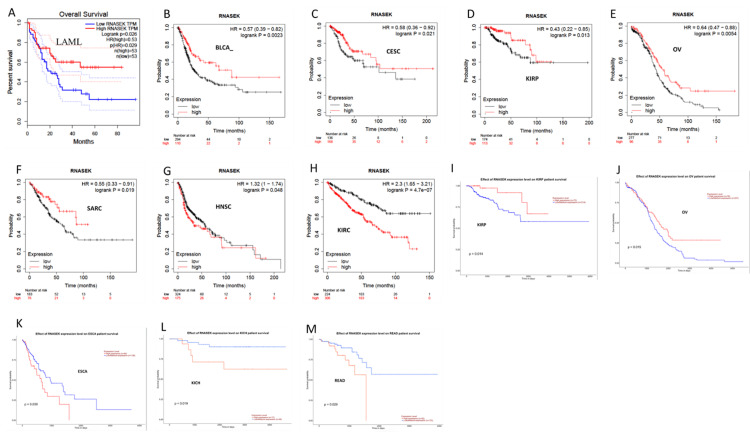
We identified the relation between RNASEK expression and overall survival by using the GEPIA, UALCAN, and Kaplan-Meier (KM) databases. The overall survival data obtained via GEPIA were statistically analyzed, and the P-value was < 0.05; the low and high cutoffs were 50%, and the confidence interval (CI) was 95%. We found that high expression of RNASEK was associated with good prognosis in PAAD patients (hazard ratio (HR): 0.49, P = 0.0007) (Figure 5A) and acute myeloid leukemia (LAML) patients (HR: 0.53, P = 0.029) (Appendix D). According to the KM database, our results revealed a correlation between high RNASEK expression and good prognosis in five types of cancers: BLCA (HR: 0.57, P = 0.002), cervical squamous cell carcinoma (CESC) (HR: 0.58, P = 0.021), KIRP (HR: 0.43, P = 0.013), ovarian cancer (OV) (HR: 0.64, P = 0.005), sarcoma (SARC) (HR: 0.55, P = 0.019) (Appendix D) and PAAD (HR: 0.45, P = 0.0001) (Figure 5B). In addition, high RNASEK expression was correlated with poor prognosis in HNSC (HR: 1.32 (1-1.74), P = 0.048) and KIRC (HR: 2.3 (1.65- 3.21), P = 0.048) (Appendix D). Furthermore, overall survival analysis via UALCAN revealed that high expression of RNASEK was associated with good prognosis in three cancers, namely, kidney renal papillary cell carcinoma (KIRP) (P = 0.014), OV (P = 0.015) (Appendix D), and PAAD (P = 0.01) (Figure 5C), whereas poor prognosis was detected in ESCA (P = 0.039), KICH (P = 0.019), and READ (P = 0.029) (Appendix D). By cross-referencing the results obtained from three databases (GEPIA, KM, and UALCAN), we found that PAAD was associated with good prognosis and high expression of RNASEK (Figures 5A–5C).
Genetic alteration analysis of RNASEK
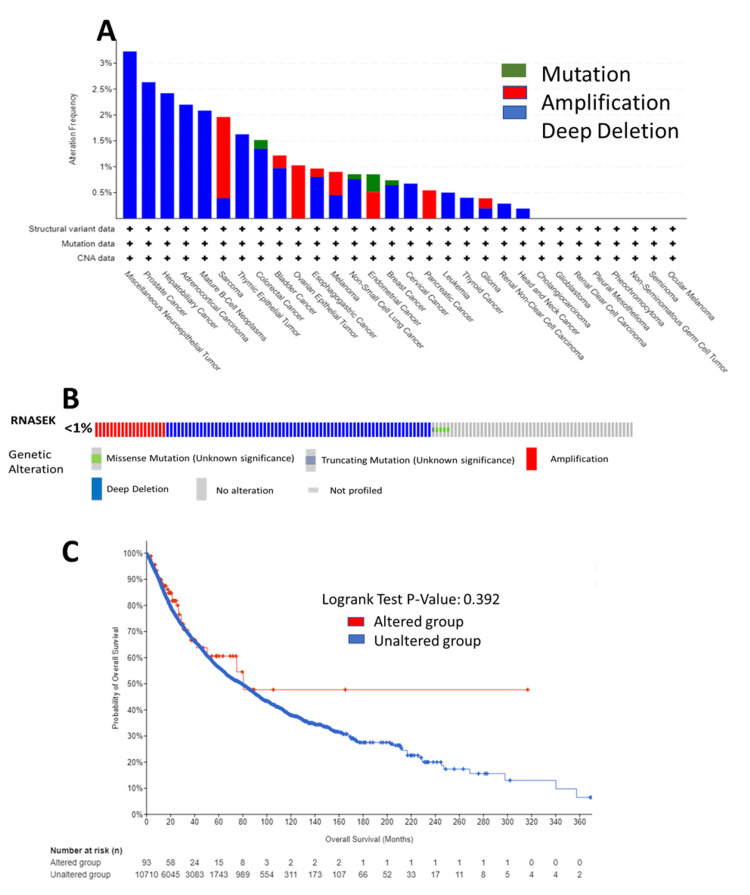
We analyzed RNASEK genetic alterations in the TCGA Pan-Cancer Atlas across 32 studies (10,967 samples) via the cBioPortal database; RNASEK was altered in 95 samples (<1%). As shown in Figure 6A, the RNASEK alterations included deep deletions, followed by amplification and mutation. In the form of deep deletions, RNASEK was altered in nineteen types of cancer, with the highest rate observed in miscellaneous neuroepithelial tumors, where one case out of 31 (3.23%) was affected. Amplification was identified in eight types of cancer, including SARC, BLCA, OV epithelial tumor, esophagogastric cancer (ESCA), melanoma, endometrial cancer, PAAD, and glioma, with the highest rate detected in SARC, at four cases out of 255 (1.96%). The frequency of mutations was found in four types of cancer: endometrial cancer, with two cases out of 586 (0.34%); colorectal cancer, with one case out of 594 cases (0.17%); non-small cell lung cancer, with one case out of 1,053 cases (0.09%); and breast cancer, with one case out of 1,084 cases.
Validation
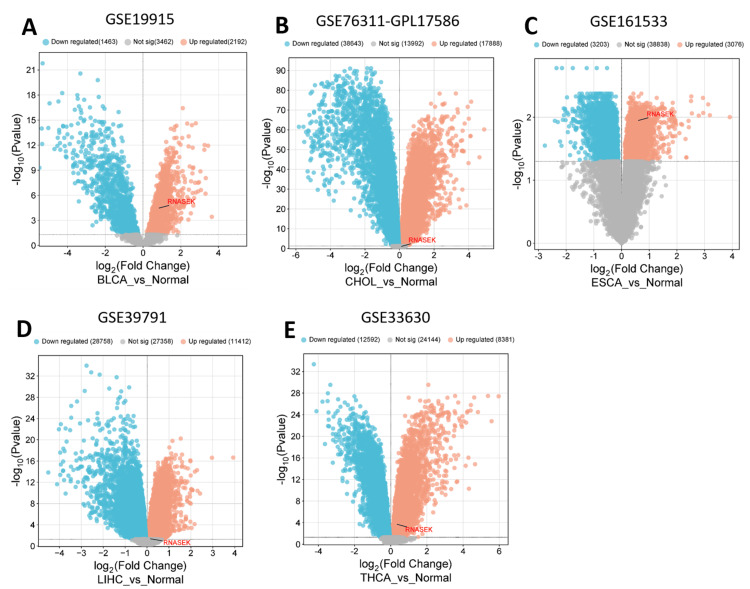
We validated the expression of RNASEK in five cancers, including BLCA, CHOL, LIHC, ESCA, and THCA, via public datasets obtained from the Gene Expression Omnibus (GEO). We employed the GEO2R tool to indicate the differential expression of RNASEK (|Log2FC| > 0.5, adjusted P-value < 0.05), and we utilized the http://www.bioinformatics.com.cn/srplot platform to visualize the volcano plots for differentially expressed genes. As shown in Figure 7, RNASEK expression was significantly upregulated in five cancers: BLCA (GSE19915-GPL3883, tumor = 76 samples, normal = 8 samples), CHOL (GSE76311-GPL17586, tumor = 92 samples, non-tumor = 93 samples), ESCA (GSE161533, tumor = 40 samples, normal = 16 samples), LIHC (GSE139791, tumor = 72 samples, non-tumor = 72 samples), and THCA (GSE133630, tumor = 49 samples, non-tumor = 45 samples).
Discussion
Researchers worldwide consider pan-cancer biomarker analysis as a useful and cost-effective screening tool to better understand the role of blood or other body fluids as biomarkers [ref. 20,ref. 21]. This study investigated how RNASEK functions as a diagnostic and prognostic biomarker in a human pan-cancer dataset. Moreover, the expression of the RNASEK gene and its potential implications in various types of cancer have been explored. We used a variety of databases from TCGA, such as TIMER, GEPIA, UALCAN, cBioportal, and KM, for a comprehensive examination of RNASEK in 33 types of tumors, covering gene expression, prognosis, gene alterations, immune infiltration, DNA methylation, and validation.
Our findings demonstrated significant RNASEK upregulation across six cancer types: BLCA, CHOL, ESCA, HNSC, LIHC, and THCA, which aligns with previous research showing threefold higher RNASEK expression in various cancer tissues like heart, brain, placenta, lung, liver, skeletal muscle, kidney, and pancreas compared to normal tissues [ref. 6]. RNASEK is expressed as one main transcript in nearly all human tissues and developmental stages, as well as in many carcinomas, which may explain its involvement in the carcinogenesis process [ref. 22]. Identifying eight novel alternatively spliced RNASEK variants with widespread expression in human cancer cell lines further emphasizes its biological significance in cancer development [ref. 5]. Furthermore, the importance of RNASEK in cancer is supported by the general role of ribonucleases in RNA degradation, which influences critical cellular processes including growth, apoptosis, and angiogenesis [ref. 9]. Additionally, other human RNases have been implicated as pathogenic factors in inflammatory disorders and autoimmune diseases [ref. 7] and the suppression of tumor growth and metastasis [ref. 8]. However, the specific role of human RNASEK in cancer progression remains poorly understood, and further investigations are needed.
Epigenetic modifications, such as DNA methylation can significantly influence gene expression without altering the underlying DNA sequence, making them key targets for cancer research, and recognized as crucial factors in cancer development and progression, regulate cell proliferation, and potential biomarkers for early detection, diagnosis, and treatment of various tumors [ref. 23]. Abnormal patterns of DNA methylation have been linked to the occurrence and proliferation of cancer [ref. 24]. Our study found that RNASEK expression is elevated in several cancer types (BLCA, HNSC, LIHC, UCEC) due to reduced DNA methylation. This suggests that RNASEK methylation could serve as a potential biomarker for prognosis in these patients.
Our analysis of RNASEK expression across cancer stages revealed significant differences between stage 1 compared to stage 3 in ESCA, HNSC, and THCA, and significant differences were observed between stages 2 and 3 when compared to stage 4 in LIHC patients. These differences among stages suggest RNASEK’s fundamental role in these cancers’ pathogenesis and potential involvement in advanced disease progression. These findings present an interesting contrast with previous research on prostate cancer, where RNASEK downregulation was associated with cancer development, and its overexpression correlated with reduced tumor aggressiveness and improved survival [ref. 25].
Our analysis of RNASEK expression across racial groups demonstrated significant differences among HNSC, LIHC, and THCA cancers between normal control tissue and all three major racial groups under study (Caucasian, African American, and Asian populations). ESCA showed variations between African American individuals compared to Asian individuals. This finding could suggest that RNASEK upregulation mechanisms in these cancers may be conserved across different genetic backgrounds. For BLCA and ESCA cancers, significant differences were found between normal tissue and both Caucasian and Asian groups, but not African American populations, potentially indicating protective factors or distinct environmental influences in African American populations for these specific cancers. In CHOL, significant differences were only observed between normal and Caucasian groups, suggesting possible genetic or environmental factors specific to Caucasian populations that influence RNASEK expression in this cancer type. These varied racial patterns in RNASEK expression demonstrate the complex interaction between genetic predisposition, environmental factors, and cancer development, emphasizing the importance of considering racial differences in cancer research and treatment strategies.
Regarding age, differences in RNASEK expression were also found to be an important factor in our research. By categorizing individuals into five age groups (normal, young, middle-aged, older, and elderly), we identified different patterns among various forms of cancer in comparison to normal controls. In CHOL, LIHC, and THCA, all age categories older than 20 showed significant differences. Variations are also observed in the young-aged compared to middle-aged and older people in HNSC. This broad age range of RNASEK upregulation suggests that its role in these cancers may not be strictly age-dependent. However, further investigation into the mechanisms underlying the role of RNASEK in cancer biology as a diagnostic or therapeutic intervention.
Tumor immune infiltrating cells are closely associated with tumor progression, immune checkpoint inhibition function, and patient prognosis [ref. 26–ref. 28]. Our results revealed a consistent weak positive correlation between RNASEK expression and macrophage abundance in BLCA, ESCA, HNSC, and LIHC patients. This relationship suggests that RNASEK may play a role in macrophage recruitment or survival within the tumor microenvironment. Thus, tumor-associated macrophages play complex roles in cancer pathophysiology. The results for LIHC were particularly noteworthy, as RNASEK expression showed weak positive correlations also with B cells, CD8+ T-cells, neutrophils, and dendritic cells. This association with multiple immune cell types suggests that in LIHC, RNASEK might be involved in tumor progression. The findings in THCA, RNASEK expression showed a weak positive correlation with CD8+ T-cells, while it exhibited significant weak negative correlations with the infiltration of B cells, CD4+ T-cells, macrophages, neutrophils, and dendritic cells. This contrasting pattern suggests a complex and potentially unique role for RNASEK in the immune landscape of thyroid cancer. Our study provides novel insights into the potential role of RNASEK in cancer immunity. However, further research is necessary to fully elucidate its immunological functions and mechanisms of action in human cancers.
In various types of cancer, the mRNA expression of RNASEK showed notable prognostic connections in pan-cancer patients. Elevated RNASEK levels were linked to good prognosis in LAML, BLCA, CESC, KIRP, OV, and PAAD, but poor prognosis in ESCA, HNSC, KICH, KIRP, and READ patients. On the other hand, in late-stage ovarian cancer, increased expression of RNASEK has been associated with resistance to chemotherapy and worse survival rates. The differing observations indicate that the involvement of RNASEK in cancer is intricate and reliant on context, varying based on cancer type and stage of progression [ref. 29]. In summary, PAAD was selected as a cancer with great potential prognostic value. However, the impact of RNASEK on survival varied across different cancers, suggesting its complex role in tumorigenesis. Further investigation is required to elucidate the specific mechanisms underlying RNASEK’s prognostic value in these cancers.
Furthermore, our genomic analysis identified frequent genetic alterations in RNASEK particularly deep deletions in miscellaneous neuroepithelial tumors, amplifications in SARC, and finally mutations in endometrial cancers. This finding suggests that RNASEK might play a role in the progression of tumors in these cancers. Another study stated the potential importance of Genetic mutations in tumor development and progression [ref. 30].
Limitations
Our study provides valuable insights into the role of RNASEK across different cancers using publicly available databases. However, some limitations should be considered. First, all the analyses were based on bioinformatics tools, further experiments to clarify the precise mechanisms by which RNASEK contributes to cancer development and progression are needed in the future. Second, the original data of these databases were mainly derived from databases, the method of collecting and processing data may not be consistent from database to database, which might cause systematic bias. Finally, the sample size and diversity of patient populations in public databases may not fully represent the full scope of cancer globally, affecting the ability to apply findings to diverse ethnicities and regions. The study primarily focuses on mRNA expression levels of RNASEK. Further examination of protein expression, post-translational modifications, and other functional aspects of the protein is advantageous.
Conclusions
In conclusion, this study’s pan-cancer analysis revealed that RNASEK could be a potential diagnostic biomarker in six types of cancers, including BLCA, CHOL, ESCA, HNSC, LIHC, and THCA. Further, we confirmed RNASEK upregulation in BLCA, CHOL, LIHC, ESCA, and THCA by the GEO datasets. Additionally, it shows promise as a prognostic biomarker for PAAD. Our findings highlighted the significant reduction in the methylation levels of RNASEK in BLCA, HNSC, LIHC, and UCEC tissues, which may explain the high RNASEK expression in these tumors. Our study showed marked RNASEK expression variations across cancer stages in ESCA, HNSC, LIHC, and THCA patients. Racial differences were evident in ESCA between African-American and Asian populations. Age-wise, RNASEK expression significantly varied in HNSC across age groups. Furthermore, there is a weak positive correlation between RNASEK expression and macrophages in BLCA, ESCA, HNSC, and LIHC. Weak positive correlations were also noted with B cells and CD8+ T-cells in HNSC, LIHC, and THCA. CD4+ T-cells showed a weak positive correlation with ESCA. Additionally, neutrophils and dendritic cells displayed weak positive correlations in BLCA, HNSC, and LIHC. Conversely, THCA demonstrated moderate negative correlations with various immune cells, including macrophages, B cells, CD4+ T-cells, neutrophils, and dendritic cells. Moreover, PAAD was linked with good prognosis and high expression of RNASEK. RNASEK alterations were rare across tumor types; they exhibited deep deletions in miscellaneous neuroepithelial tumors, amplification in SARC, and mutations were identified in endometrial cancer.
References
- H Sung, J Ferlay, RL Siegel. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin, 2021. [PubMed]
- D Daassi, KM Mahoney, GJ Freeman. The importance of exosomal PDL1 in tumour immune evasion. Nat Rev Immunol, 2020. [PubMed]
- T Soussi, KG Wiman. TP53: an oncogene in disguise. Cell Death Differ, 2015. [PubMed]
- Y Gu, S Tang, Z Wang, L Cai, H Lian, Y Shen, Y Zhou. A pan-cancer analysis of the prognostic and immunological role of β-actin (ACTB) in human cancers. Bioengineered, 2021. [PubMed]
- PG Adamopoulos, CK Kontos, A Scorilas, DC Sideris. Identification of novel alternative transcripts of the human Ribonuclease κ (RNASEK) gene using 3′ RACE and high-throughput sequencing approaches. Genomics, 2020. [PubMed]
- MA Economopoulou, EG Fragoulis, DC Sideris. Molecular cloning and characterization of the human RNase kappa, an ortholog of Cc RNase. Nucleic Acids Res, 2007. [PubMed]
- B Schwartz, O Shoseyov, VO Melnikova. ACTIBIND, a T2 RNase, competes with angiogenin and inhibits human melanoma growth, angiogenesis, and metastasis. Cancer Res, 2007. [PubMed]
- N Ohta, S Okazaki, S Fukase. Serum concentrations of eosinophil cationic protein and eosinophils of patients with Kimura’s disease. Allergol Int, 2007. [PubMed]
- VA Shlyakhovenko. Ribonucleases in tumor growth. Exp Oncol, 2009. [PubMed]
- TN Rampias, EG Fragoulis, DC Sideris. Genomic structure and expression analysis of the RNase kappa family ortholog gene in the insect Ceratitis capitata. FEBS J, 2008. [PubMed]
- JM Perreira, AM Aker, G Savidis. Rnasek is a V-ATPase-associated factor required for endocytosis and the replication of rhinovirus, influenza A virus, and dengue virus. Cell Rep, 2015. [PubMed]
- BA Hackett, A Yasunaga, D Panda, MA Tartell, KC Hopkins, SE Hensley, S Cherry. RNASEK is required for internalization of diverse acid-dependent viruses. Proc Natl Acad Sci U S A, 2015. [PubMed]
- T Li, J Fan, B Wang. TIMER: A web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res, 2017
- Z Tang, C Li, B Kang, G Gao, C Li, Z Zhang. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res, 2017
- DS Chandrashekar, B Bashel, SA Balasubramanya, CJ Creighton, I Ponce-Rodriguez, BV Chakravarthi, S Varambally. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia, 2017. [PubMed]
- B Győrffy. Survival analysis across the entire transcriptome identifies biomarkers with the highest prognostic power in breast cancer. Comput Struct Biotechnol J, 2021. [PubMed]
- E Cerami, J Gao, U Dogrusoz. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov, 2012. [PubMed]
- T Barrett, SE Wilhite, P Ledoux. NCBI GEO: archive for functional genomics data sets–update. Nucleic Acids Res, 2013
- D Tang, M Chen, X Huang. SRplot: a free online platform for data visualization and graphing. PLoS One, 2023
- Network Cancer Genome Atlas Research, JN Weinstein, EA Collisson. The cancer genome atlas pan-cancer analysis project. Nat Gen, 2013
- T Ching, K Peplowska, S Huang. Pan-cancer analyses reveal long intergenic non-coding RNAs relevant to tumor diagnosis, subtyping and prognosis. EBioMedicine, 2016. [PubMed]
- MN Kiritsi, EG Fragoulis, DC Sideris. Essential cysteine residues for human RNase κ catalytic activity. FEBS J, 2012. [PubMed]
- Y Pan, G Liu, F Zhou, B Su, Y Li. DNA methylation profiles in cancer diagnosis and therapeutics. Clin Exp Med, 2018. [PubMed]
- TB Nguyen, DN Do, T Nguyen-Thanh, VB Tatipamula, HT Nguyen. Identification of five hub genes as key prognostic biomarkers in liver cancer via integrated bioinformatics analysis. Biology (Basel), 2021
- AP Burlaka, SV Virko, VM Grygorenko, OА Samoylenko, AV Verbinenko, VO Shlyakhovenko. The changes of NO level and RNase activity in tumor tissue accompanying the progression of prostate cancer. Exp Oncol, 2022. [PubMed]
- PA Ascierto, KD Lewis, AM Di Giacomo. Prognostic impact of baseline tumour immune infiltrate on disease-free survival in patients with completely resected, BRAF(v600) mutation-positive melanoma receiving adjuvant vemurafenib. Ann Oncol, 2020. [PubMed]
- X Liu, J Xu, B Zhang. The reciprocal regulation between host tissue and immune cells in pancreatic ductal adenocarcinoma: new insights and therapeutic implications. Mol Cancer, 2019. [PubMed]
- SS Wang, W Liu, D Ly, H Xu, L Qu, L Zhang. Tumor-infiltrating B cells: their role and application in anti-tumor immunity in lung cancer. Cell Mol Immunol, 2019. [PubMed]
- Y-S Kim, JD Hwan, S Bae. Identification of differentially expressed genes using an annealing control primer system in stage III serous ovarian carcinoma. BMC Cancer, 2010. [PubMed]
- H Teimouri, AB Kolomeisky. Temporal order of mutations influences cancer initiation dynamics. Phys Biol, 2021
