Prognostic value of TMTC1 in pan-cancer analysis
Abstract
Background:
Transmembrane and tetratricopeptide repeat containing 1 (TMTC1) is a recently discovered enzyme involved in the O-mannosylation of cadherins and protocadherins. It has been implicated in various types of cancer, but the overall prognostic significance of TMTC1 in pan-cancer and its potential as an immunotherapeutic target remain unclear.
Methods:
We applied various bioinformatics methods to investigate the potential oncogenic roles of TMTC1 using public databases. This analysis involved examining the expression, prognosis, genetic alterations, immune infiltration, immunotherapy response, drug sensitivity, and regulatory mechanisms of the TMTC1 gene in diverse cancer types.
Results:
In this study, we observed that TMTC1 expression is reduced in 19 types of cancer (ACC, BLCA, BRCA, CESC, COAD, ESCA, GBM, KICH, KIRC, KIRP, LAML, LUAD, LUSC, PRAD, READ, STAD, THCA, UCEC, and UCS) compared to normal tissues. Conversely, TMTC1 expression is elevated in OV and PAAD relative to normal tissues. Moreover, our analysis revealed that high expression of TMTC1 was associated with worse overall survival (OS) outcomes in patients with ACC, BLCA, COAD, GBM, KIRP, OV, STAD, and UCEC, but better OS outcomes in patients with CESC, KIRC, LUSC, and PAAD. Notably, patients with TMTC1 mutations or deep deletions demonstrated longer OS, while those with TMTC1 amplification showed shorter OS. There was a significant correlation between the expression level of TMTC1 and the infiltration of cancer-associated fibroblasts (CAFs) and endothelial cells. Using data from six real-world immunotherapy cohorts of BLCA, SKCM and RCC, we discovered that high TMTC1 expression was associated with better OS or progression-free survival (PFS). Lastly, through TMTC1-related gene enrichment analysis, some biological processes and pathways were found to be significantly enriched, such as vascular endothelial growth factor receptor signaling pathway and ECM-receptor interaction.
Conclusions:
Our study demonstrates the prognostic significance of TMTC1 in pan-cancer and highlights its potential as an immunotherapeutic target.
Article type: Research Article
Keywords: Pan-cancer, Prognosis, Immunotherapy
Affiliations: The International Peace Maternity and Child Health Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai, China; Shanghai Key Labouratory of Embryo Original Diseases, 200030, Shanghai, China; Institute of Birth Defects and Rare Diseases, School of Medicine, Shanghai Jiao Tong University, 200030, Shanghai, China; Department of Obstrics and Gynecology, The First People’s Hospital of Jiande, Hangzhou, China
License: © 2024 The Authors. Published by Elsevier Ltd. CC BY 4.0 This is an open access article under the CC BY license (http://creativecommons.org/licenses/by/4.0/).
Article links: DOI: 10.1016/j.heliyon.2024.e38308 | PubMed: 39397950 | PMC: PMC11471174
Relevance: Relevant: mentioned in keywords or abstract
Full text: PDF (19.5 MB)
Introduction
Cancer, characterized by the abnormal growth and spread of cells, remains a major global health concern, causing substantial morbidity and mortality [ref. 1,ref. 2]. To develop effective therapeutic strategies, it is imperative to gain a deep understanding of the intricate molecular mechanisms responsible for the initiation and progression of cancer. Recent years have witnessed remarkable advancements in high-throughput sequencing technologies, revolutionizing the exploration of molecular alterations across different types of cancer. Consequently, the identification and characterization of cancer-related genes and their involvement in tumor formation and development have emerged as indispensable undertakings in the field of cancer research.
The TMTC (transmembrane (TM) and tetratricopeptide (TPR) repeat-containing) proteins family consists of four members in humans, namely TMTC1-4, and serves as resident proteins of the endoplasmic reticulum (ER) [ref. 3]. The TMTCs proteins have an N-terminal composed of 11 transmembrane regions and multiple intermittent loops, while their C-terminal is located in the ER lumen and consists of TPR repeats [ref. [3], ref. [4], ref. [5]]. The TMTCs proteins play essential roles in ER function. For example, TMTC1 and TMTC2 can interact with the ER calcium uptake pump SERCA2B via their C-terminal TPR domains, participating in the regulation of ER calcium homeostasis [ref. 5]. TMTC3 interacts with PDIA3 and regulates proteasome activity and the expression of XBP-1, a stress response protein, thereby participating in the ER stress response [ref. 4]. Similarly, TMTC4 also interacts with the ER calcium pump SERCA2b, altering ER calcium dynamics, which leads to overactivation of the downstream unfolded protein response and cell death [ref. 6]. Recent studies have revealed that TMTCs function as glycosyltransferases, playing a crucial role in the O-mannosylation process of cadherins [ref. 7] and integrin β [ref. 8]. Consequently, they actively contribute to the progression of cancer [ref. 3,ref. 7,ref. 9,ref. 10].
TMTC1, as an important member of the TMTC family, plays a crucial role in various physiological pathways. Studies have revealed its involvement in protein glycosylation [ref. 3], endoplasmic reticulum calcium homeostasis [ref. 5], cellular adhesion [ref. 8], cell differentiation [ref. 11], and inflammation [ref. 12]. Furthermore, accumulating evidence suggests that dysregulation of TMTC1 is associated with several human diseases, including cancer. For instance, TMTC1 mutations have been linked to schizophrenia [ref. 13]. In the context of cancer, TMTC1 is upregulated in ovarian cancer and promotes migration and invasion of ovarian cancer cells through modifications in O-mannosylation and the activity of integrins β1 and β4 [ref. 8]. High TMTC1 mRNA expression is also correlated with decreased survival in gastric cancer patients [ref. 14]. Additionally, TMTC1 has been identified as being involved in multiple other cancers, such as breast cancer [ref. 15,ref. 16]、renal cancer [ref. 16]、glioblastoma [ref. 17] and nasopharyngeal carcinoma [ref. 18]. Although previous studies have suggested the potential role of TMTC1 in cancer, a comprehensive analysis of its expression and prognostic significance across different types of cancer is currently lacking. Hence, it is necessary to systematically examine TMTC1 expression in multiple cancer types in order to unravel its involvement in carcinogenesis and evaluate its potential as a prognostic biomarker.
Our research is the first pan-cancer analysis of TMTC1 by utilizing TCGA, GTEx, HPA, and GEO databases. Our analysis comprehensively examined several critical aspects including gene expression, prognosis, genetic alterations, immune infiltration, immunotherapy response, drug sensitivity, and regulatory mechanisms of TMTC1 in pan-cancer. By elucidating the role of TMTC1 in different cancer types, this study may pave the way for targeted therapeutic interventions that could improve patient outcomes and potentially provide novel avenues for cancer treatment.
Materials and methods
Data acquisition
Gene expression data, clinical phenotype data, and genetic alterations (mutations and copy number variations) data of thirty-three cancer types form TCGA were acquired from the UCSC Xena database (https://xenabrowser.net/datapages/) [ref. 19]. The thirty-three cancers of interest in this study, with their full names and abbreviations, are presented in Table 1. Gene expression profile of human normal tissues were retrieved from GTEx (https://commonfund.nih.gov/GTEx) [ref. 20]. Two immunotherapy cohorts of SKCM were obtained from the European Nucleotide Archive (https://www.ebi.ac.uk/ena/browser/view/PRJEB23709) and GEO (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE91061), respectively [ref. 21,ref. 22]. Two immunotherapy cohorts of BLCA [ref. 23,ref. 24] were obtained from available data package (http://research-pub.gene.com/IMvigor210CoreBiologies) and GEO (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE176307). Two immunotherapy cohorts of RCC were obtained from available data package (http://research-pub.gene.com/IMvigor210CoreBiologies) and Motzer’s articles [ref. 25]. An immunotherapy cohort of KIRC cohort were extracted from the supplementary files of Braun’s articles [ref. 26]. The IC50 values and transcriptomic data for cell lines treated with 198 drugs were downloaded from the GDSC2 database (https://www.cancerrxgene.org).
Table 1: List of cancer types.
| Study Abbreviation | Study Name |
|---|---|
| ACC | Adrenocortical carcinoma |
| BLCA | Bladder Urothelial Carcinoma |
| BRCA | Breast invasive carcinoma |
| CESC | Cervical squamous cell carcinoma and endocervical adenocarcinoma |
| CHOL | Cholangiocarcinoma |
| COAD | Colon adenocarcinoma |
| DLBC | Lymphoid Neoplasm Diffuse Large B-cell Lymphoma |
| ESCA | Esophageal carcinoma |
| GBM | Glioblastoma multiforme |
| HNSC | Head and Neck squamous cell carcinoma |
| KICH | Kidney Chromophobe |
| KIRC | Kidney renal clear cell carcinoma |
| KIRP | Kidney renal papillary cell carcinoma |
| LAML | Acute Myeloid Leukemia |
| LGG | Brain Lower Grade Glioma |
| LIHC | Liver hepatocellular carcinoma |
| LUAD | Lung adenocarcinoma |
| LUSC | Lung squamous cell carcinoma |
| MESO | Mesothelioma |
| OV | Ovarian serous cystadenocarcinoma |
| PAAD | Pancreatic adenocarcinoma |
| PCPG | Pheochromocytoma and Paraganglioma |
| PRAD | Prostate adenocarcinoma |
| RCC | Renal cell carcinoma |
| READ | Rectum adenocarcinoma |
| SARC | Sarcoma |
| SKCM | Skin Cutaneous Melanoma |
| STAD | Stomach adenocarcinoma |
| TGCT | Testicular Germ Cell Tumors |
| THCA | Thyroid carcinoma |
| THYM | Thymoma |
| UCEC | Uterine Corpus Endometrial Carcinoma |
| UCS | Uterine Carcinosarcoma |
| UVM | Uveal Melanoma |
Expression analysis
The expression data of the TMTC1 gene (ENSG00000133687) were extracted from downloaded transcriptome data. Following TPM normalization, we conducted a log2(x+1) conversion. In order to enhance reliability and control for possible batch effects, data from the same sample, but acquired from different test batches, were averaged. Furthermore, we filtered the samples from the following sources: Solid Tissue Normal, Primary Solid Tumor, Primary Tumor, Normal Tissue, Primary Blood Derived Cancer – Bone Marrow, and Primary Blood Derived Cancer – Peripheral Blood. We also excluded cancer types with less than 3 samples, resulting in a final dataset of expression data from 28 cancer and their corresponding adjacent normal tissues. We used R software (version 4.0.2) to calculate the expression differences of TMTC1 between tumor and normal samples in each tumor type. Additionally, we assessed the expression differences of TMTC1 in different clinical stage samples using Wilcoxon’s test for significance analysis. The results were visualized using the “ggpubr” R package [ref. 27].
The Human Protein Atlas (HPA) database [ref. 28] (https://www.proteinatlas.org/) is a comprehensive database of human protein information, aiming to provide a detailed description of human gene and protein expression patterns. This database integrates proteomic data from various tissue and cell types, including immunohistochemistry, immunofluorescence, and high-throughput antibody preparation techniques. Utilizing this database, we obtained immunohistochemical data of TMTC1 protein in distinct cancerous and normal tissues, enabling us to examine the differential protein expression profiles at a protein level. Furthermore, to elucidate the cellular localization of the TMTC1 protein, we also obtained immunofluorescence data from U-251-MG cell line treated with TMTC1 antibodies from this database.
Survival analysis
The “survival” R package [ref. 29] was employed to conduct survival analysis on the patient data. To establish an optimal cutoff point for continuous TMTC1 expression, the surv_cutpoint function was utilized. Subsequently, an analysis was performed to correlate the expression levels of TMTC1 with OS. To visualize the results, forest plots and Kaplan-Meier (KM) curves plots were generated using the “forestplot” [ref. 30] and “survminer” [ref. 31] R packages, respectively.
Genetic alteration analysis
In order to explore the specific characteristics of genetic alterations in TMTC1 across various cancers. we conducted a comprehensive study on the incidence and number of mutations, amplifications, and deep deletions of TMTC1 in 32 different types of cancers. Furthermore, the impact of TMTC1 genetic alterations on patient survival was assessed using the "survival" R package. Survival analyses comparing patients with TMTC1 mutations, amplifications, and deep deletions to those without any alterations were conducted, and the results were visualized using Kaplan-Meier (KM) curve plots.
Immune infiltration analysis
To investigate the relationship between TMTC1 expression and immune infiltration, we initially employed the "Immune Infiltration" module of the SangerBox3.0 [ref. 32] platform. This module allowed us to compute the Pearsons’s correlation coefficient between TMTC1 expression and immune infiltration scores, including stromal, immune, and ESTIMATE scores [ref. 33], across various tumor samples, resulting in the identification of significant correlations. The correlation analysis results were then visualized as a heatmap using the "pheatmap" [ref. 34] R package.
To explore the relationship between TMTC1 expression and the abundance of cancer-associated fibroblasts and endothelial cells in the tumor microenvironment, we utilized the "Gene" module in TIMER2.0 (http://timer.cistrome.org/), which provides several immune deconvolution methods for calculating the abundance of cancer-associated fibroblasts and endothelial cells including EPIC [ref. 35], MCP-counter [ref. 36], XCELL [ref. 37], and TIDE. The Spearman’s correlations were calculated and a heatmap with numbers showed the purity-adjusted spearman’s rho across various cancer types. By clicking on specific cells within the heatmap, scatter plots illustrating the relationship between infiltrate estimation values and gene expression in different types of cancer were generated.
Single-cell analysis of TMTC1
To investigate the correlation between TMTC1 and the tumor microenvironment at the single-cell level, we utilized the "Gene Exploration" module in the Tumor Immune Single-cell Hub (TISCH) database (http://tisch.comp-genomics.org/home/) [ref. 38]. This module enabled quantification of TMTC1 expression in immune cells and stromal cells within the tumor microenvironment. The following main parameters were set: gene selected as “TMTC1”, cell-type annotation selected as “Celltype (maior-lineage)”, cancer type selected as “BLCA, BRCA, CHOL, KIRC, LIHC, UVM, UCEC, STAD, SKCM, SARC, PAAD, OV, and HHSC”. To explore the expression of TMTC1 in single-cell data of a specific cancer, the "Dataset" module provided the option to click on a specific dataset name, followed by inputting "TMTC1″ in the "Gene" module, resulting in the generation of heatmaps and violin plots displaying TMTC1 expression.
Immunotherapy outcome analysis
Immunotherapy has emerged as a promising approach for treating various diseases, including cancer [ref. 39]. To explore the potential prognostic value of TMTC1 in the context of immunotherapy, we collected data from six immunotherapy cohorts of four types of cancer, along with their transcriptomic profiles and clinical information (see the data acquisition section above for more details). We then performed survival analysis on patients with different expression levels of TMTC1 (see the survival analysis section for methods).
Drug sensitivity analysis
The GDSC2 (Genomics of Drug Sensitivity in Cancer version 2) database (https://www.cancerrxgene.org) is a comprehensive database aimed at studying the sensitivity of cancer drugs [ref. 40]. This database integrates various data on cancer cell lines, including genomics, transcriptomics, and drug responses. To analyze the correlation between drug sensitivity and TMTC1 expression, IC50 values and transcriptomic data for cell lines treated with 198 drugs were downloaded from the GDSC2 database. The “oncoPredict” [ref. 41] R package was used to calculate the IC50 for each sample from TCGA, using the transcriptomic data from GDSC2 as the training group and transcriptomic data from TCGA as the verification group. Pearson’s correlation coefficient was computed to assess the relationship between drug IC50 values and TMTC1 expression across pan-cancer samples. Heatmaps, scatter plots, and box plots were generated using the "pheatmap" [ref. 34] and "ggpubr" R packages for visualization.
TMTC1-related gene enrichment analysis
The LinkedOmics database (http://www.linkedomics.org/login.php) [ref. 42] was utilized to identify TMTC1-related genes in ovarian serous cystadenocarcinoma. The following main parameters were set: cancer type selected as ovarian serous cystadenocarcinoma, search dataset selected as "RNA-seq data type, HiSeq RNA platform", search dataset attribute narrowed down to TMTC1, target dataset specified as "RNA-seq data type, HiSeq RNA platform", and statistical method chosen as Pearson’s correlation test. The top 100 genes, with 50 positively correlated and 50 negatively correlated with TMTC1 in ovarian cancer, were displayed using a heatmap. The gene set enrichment analysis (GSEA) function module was applied to examine KEGG pathways and GO_BP terms, using a rank criterion of FDR <0.05 and performing 1000 simulations.
Statistical analysis
Statistical analyses were performed using R 3.6.2 (https://www.r-project.org/). Differences between the two groups and among multiple groups were analyzed using the default Wilcoxon’s test and one-way analysis of variance (ANOVA), respectively. The differences in overall survival between groups were determined by Kaplan-Meier analysis and a log-rank test. P value < 0.05 was considered to be statistically significant if not otherwise stated.
Results
Analysis of TMTC1 expression in pan-cancer
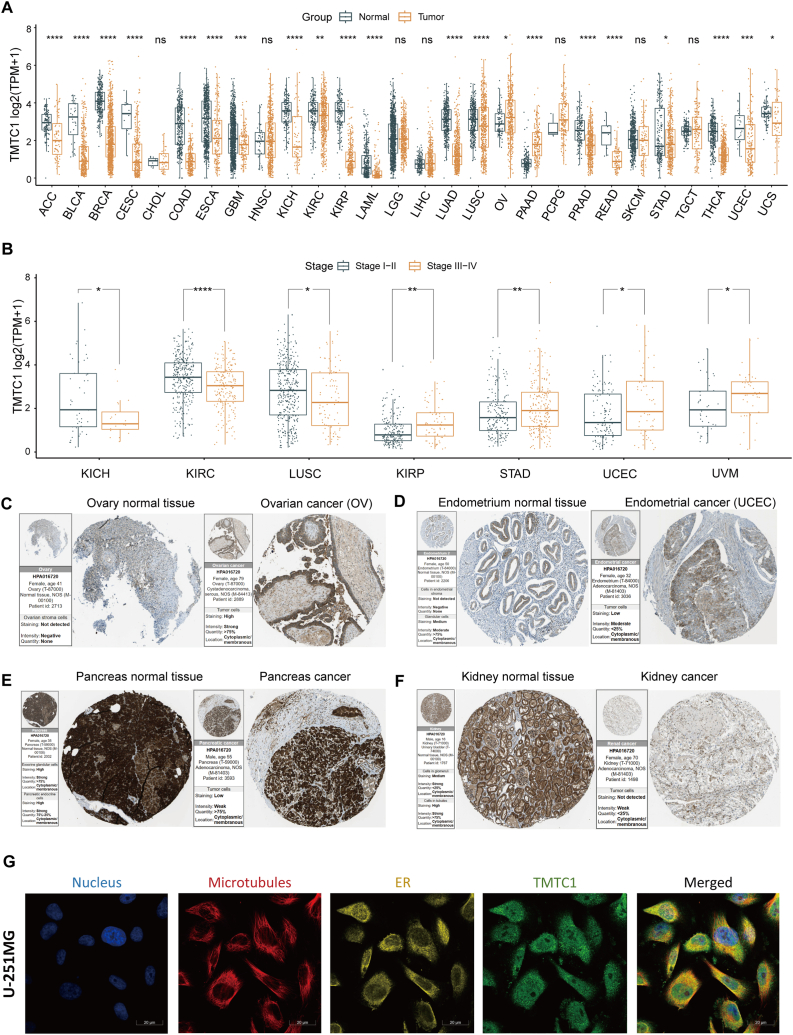
To investigate the expression of TMTC1 in different types of cancer, we firstly conducted a comprehensive study by comparing mRNA expression levels in cancer tissues and normal tissues using data from the TCGA and GTEx databases. As shown in Fig. 1A, noteworthy downregulation of TMTC1 expression was observed in the majority of tumor tissues, including ACC, BLCA, BRCA, CESC, COAD, ESCA, GBM, KICH, KIRC, KIRP, LAML, LUAD, LUSC, PRAD, READ, STAD, THCA, UCEC, and UCS. Conversely, TMTC1 was found to be upregulated in OV and PAAD. Additionally, we investigated whether TMTC1 had an impact on cancer staging. As shown in Fig. 1B, our findings demonstrated significantly higher TMTC1 expression in the early stages (Stage I and Stage II) of LUSC, KICR, and KIRC, while it was significantly higher in the late stages (Stage III and Stage IV) of KIRP, STAD, UCEC, and UVM. Moreover, analysis of TMTC1 protein expression using immunohistochemistry data from the HPA database revealed a marked increase in OV (Fig. 1C) and UCEC (Fig. 1D) when compared to normal tissues. Conversely, TMTC1 expression was significantly lower in pancreas (Fig. 1E) and kidney (Fig. 1F) cancer tissues compared to normal tissues, thus confirming the differential expression observed at the mRNA level. Finally, immunofluorescence data from the HPA revealed that TMTC1 primarily was primarily located in the endoplasmic reticulum and microtubules.
Prognostic significance of TMTC1 expression in pan-cancer
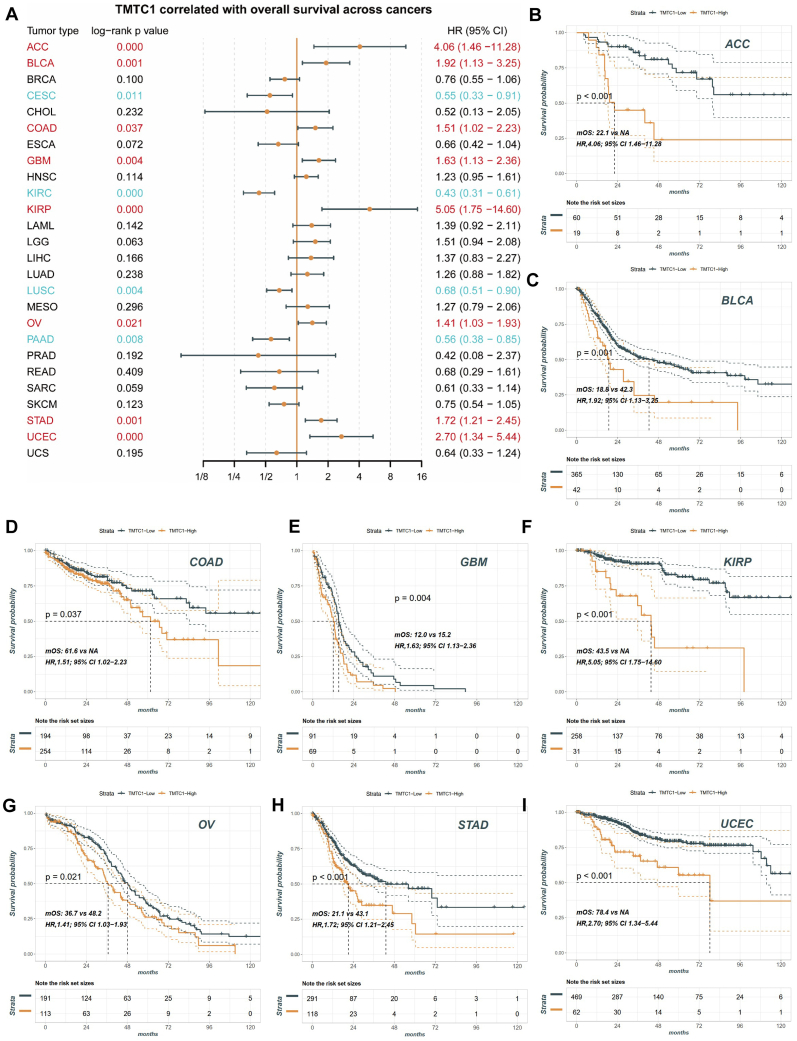
To investigate the potential prognostic value of TMTC1 expression in pan-cancer patients, data from the TCGA database were analyzed. The patients were divided into two groups based on the level of TMTC1 expression. Initially, we performed univariate Cox regression analysis to determine the association between TMTC1 expression and survival. As shown in Fig. 2A, our findings revealed that TMTC1 expression served as an independent risk factor for survival in patients with ACC (HR = 4.06, 95 % CI 1.46–11.28), BLCA (HR = 1.92, 95 % CI 1.13–3.25), COAD (HR = 1.51, 95 % CI 1.02–2.23), GBM (HR = 1.63, 95 % CI 1.13–2.36), KIRP (HR = 5.05, 95 % CI 1.75–14.60), OV (HR = 1.41, 95 % CI 1.03–1.93), STAD (HR = 1.72, 95 % CI 1.21–2.45), and UCEC (HR = 2.70, 95 % CI 1.34–5.44) (all log-rank p value < 0.05). Conversely, TMTC1 expression appeared to have a protective effect on survival in patients with CESC (HR = 0.55, 95 % CI 0.33–0.91), KIRC (HR = 0.43, 95 % CI 0.31–0.61), LUSC (HR = 0.68, 95 % CI 0.51–0.90), and PAAD (HR = 0.56, 95 % CI 0.38–0.85) (all log-rank p value < 0.05). Moreover, Kaplan–Meier survival curves showed that TMTC1 were associated with poorer OS in patients with ACC (Fig. 2B), BLCA (Fig. 2C), COAD (Fig. 2D), GBM (Fig. 2E), KIRP (Fig. 2F), OV (Fig. 2G), STAD (Fig. 2H), and UCEC (Fig. 2I).
Analysis of TMTC1 genetic alterations in pan-cancer
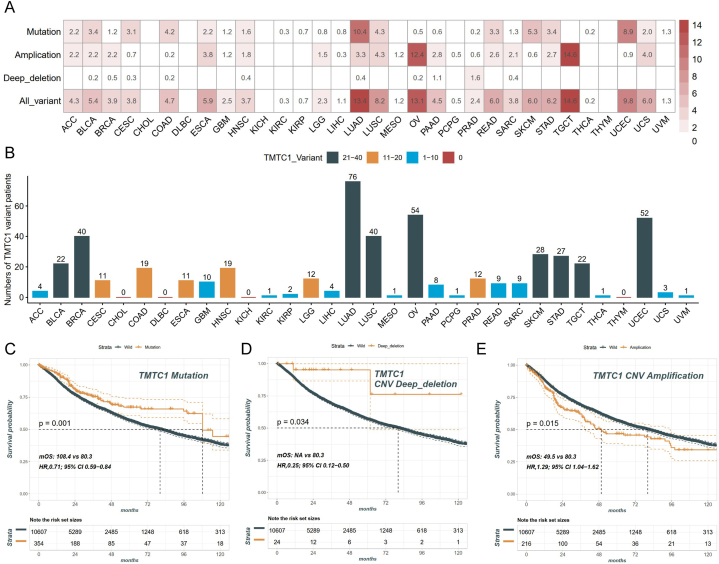
It is widely recognized that gene mutations are closely associated with tumorigenesis. To examine the genetic variations in TMTC1 across different types of cancers, we conducted a comprehensive analysis using genomic data obtained from the TCGA database across various types of cancer, As shown in Fig. 3A and B, among the 32 types of cancer patients, a total of 499 patients presented the TMTC1 variant. Except for CHOL, DLBC, KICH, and THYM, all other 28 types of cancer patients showed the TMTC1 variant. These variants occurred primarily in the form of mutation and amplification. The highest frequency of TMTC1 variant was observed in LUAD, OV, and TGCT. Among these, LUAD had mutations as the primary alteration, while OV and TGCT presented amplification as the main change. Finally, our analysis examined the impact of TMTC1 genetic alterations on the prognosis of cancer patients. The results revealed in Fig. 3A and B demonstrated that patients with TMTC1 mutation and deep deletion had significantly better OS than the wild-type group. Conversely, patients with TMTC1 amplification exhibited significantly worse OS compared to the wild-type group (Fig. 3C).
Immune infiltration analysis of TMTC1
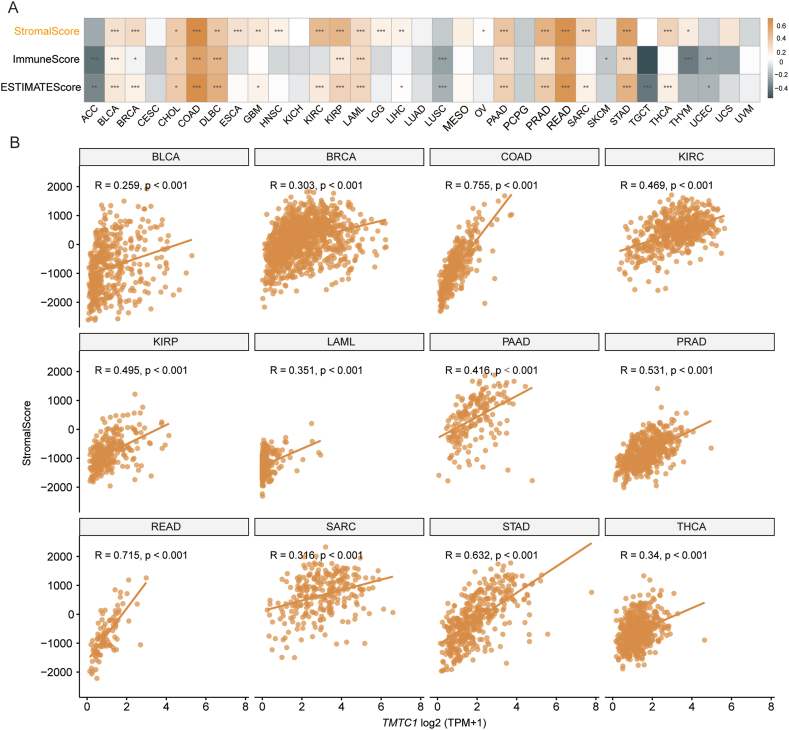
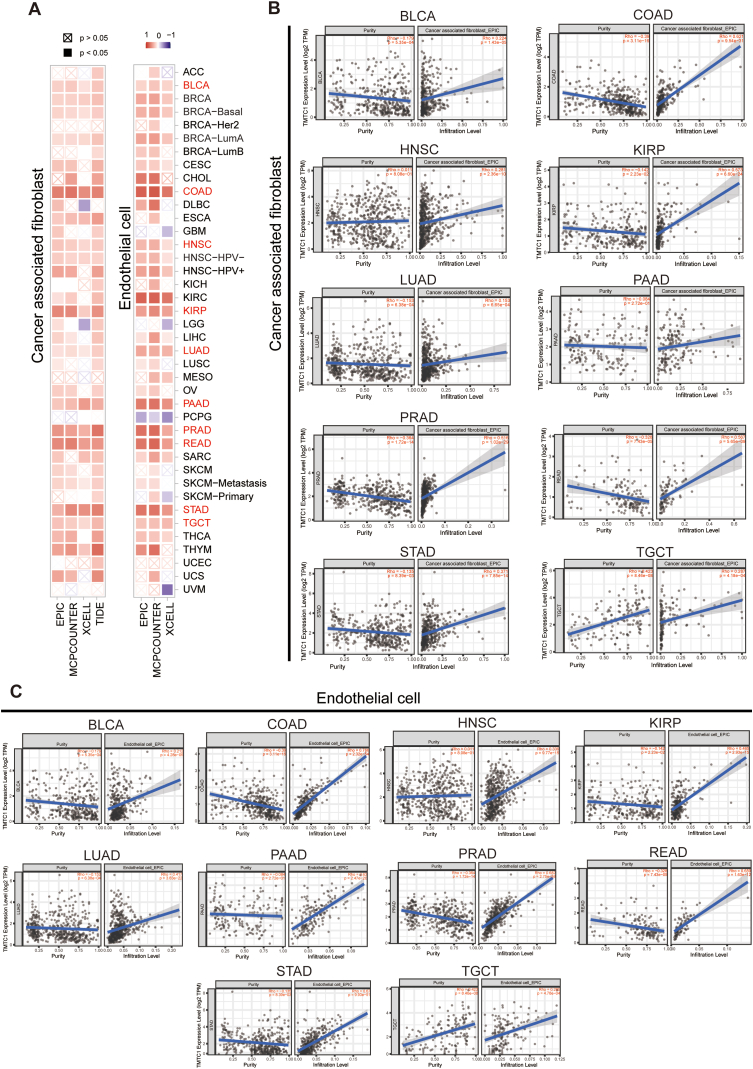
To investigate the relationship between TMTC1 expression and immune infiltration, we utilized SangerBox3.0 to compute immune infiltration scores for 33 types of cancer. Subsequently, we examined the Pearson’s correlation between TMTC1 expression and the stromal score, immune score, and ESTIMATE score for these 33 cancers. As shown in Fig. 4A, a significant positive correlation was observed between TMTC1 expression and stromal score across 21 cancers. Scatter plots in Fig. 4B further illustrated this strong correlation between TMTC1 expression and stromal score in 12 specific cancers (BLCA, BRCA, COAD, KIRC, KIRP, LAML, PAAD, PRAD, READ, SARC, STAD, and THCA). Given the significant correlation between TMTC1 expression and stromal score in most cancers, we further evaluated the relationship between TMTC1 expression and infiltration levels of stromal cells, such as cancer-associated fibroblasts (CAFs) and endothelial cells, which were important components of the tumor microenvironment and contributed to tumor development. As shown in Fig. 5A, a significant positive correlation was observed between TMTC1 expression and the infiltration levels of these two cell types in most cancers through different algorithms (EPIC, MCP-COUNTER, XCELL, and TIDE). Scatter plots in Fig. 5B and C demonstrate significant correlations between TMTC1 expression and infiltration levels of CAFs and endothelial cells, respectively, in 10 specific cancer types (BLCA, COAD, HNSC, KIRP, LUAD, PAAD, PRAD, READ, STAD, and TGCT) through EPIC algorithm.
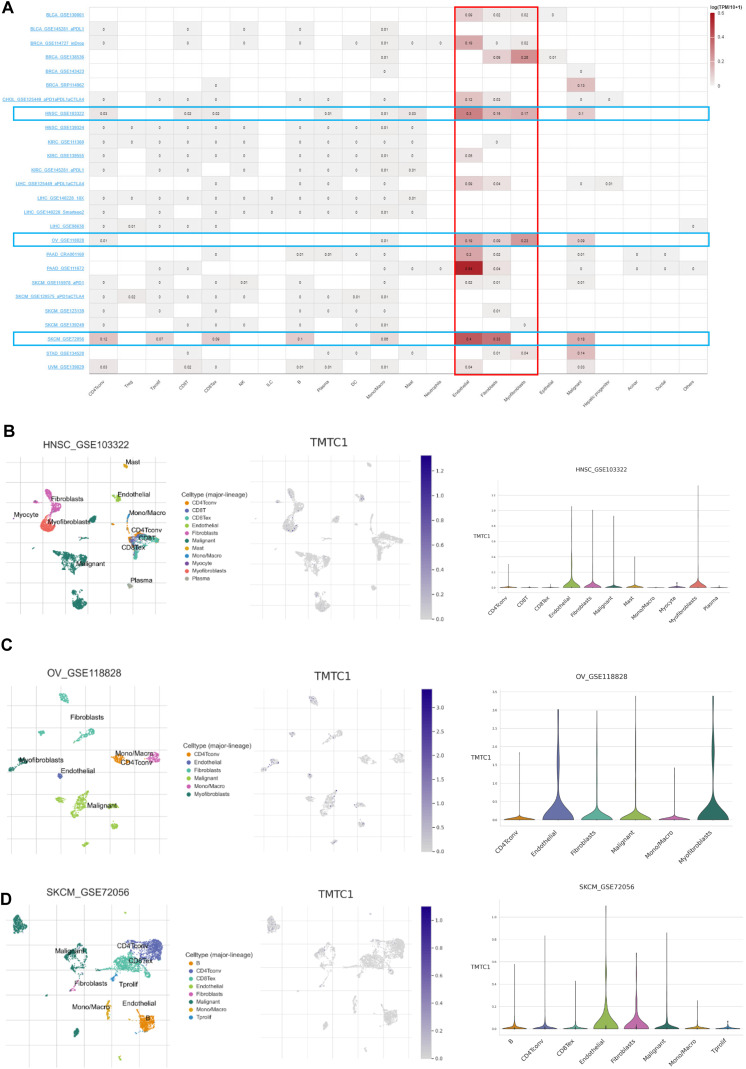
Single-cell analysis of TMTC1
The tumor microenvironment consists of a heterogeneous collection of cell types, including immune cells, endothelial cells, fibroblasts, and cancer cells. To investigate the expression of TMTC1 in different cell types within the tumor microenvironment, we utilized the TISCH database. As shown in Fig. 6A, TMTC1 demonstrates higher expression levels in stromal cells, particularly fibroblasts, endothelial cells, and myofibroblasts, compared to immune cells in most cancers. Furthermore, the scatter plots in Fig. 6B–D also illustrated that TMTC1 is highly expressed in fibroblasts, endothelia cells and myofibroblasts in the tumor microenvironment in HNSC, OV, and SKCM.
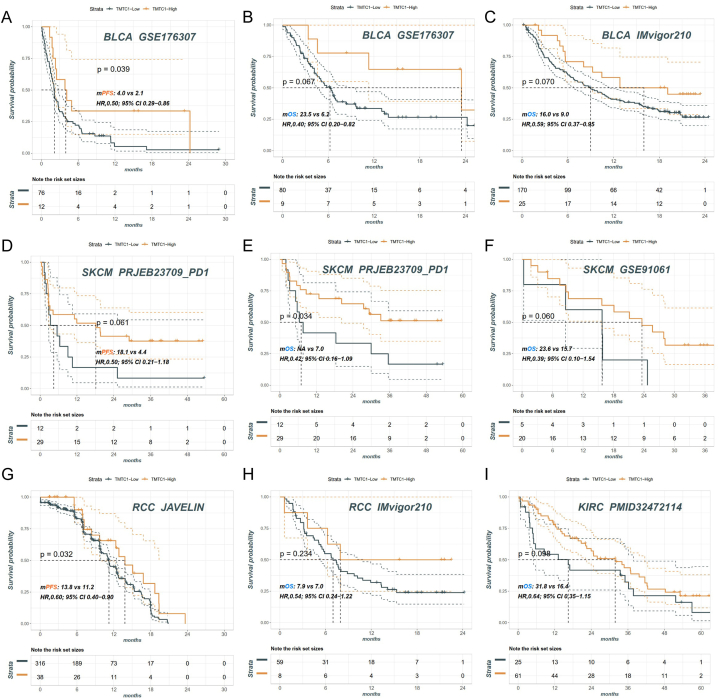
Immunotherapy analysis of TMTC1
To investigate the correlation between the expression level of TMTC1 and the efficacy of immunotherapy in tumors, we conducted a comparative analysis of prognostic outcomes among cancer patients with high and low TMTC1 expression following treatment with immune checkpoint inhibitors. Our analysis included data from various cohorts: the BLCA cohort from GSE176307 (Fig. 7A and B) and IMvigor210 (Fig. 7C), the SKCM cohort from PRJEB23709 (Fig. 7D and E) and GSE91061 (Fig. 7F), the RCC cohort from JAVELIN (Fig. 7G) and IMvigor210 (Fig. 7H), and the KIRC cohort from PMID32472114 (Fig. 7I). The results consistently indicated that patients with high TMTC1 expression had a longer PFS or OS compared to those with low TMTC1 expression.
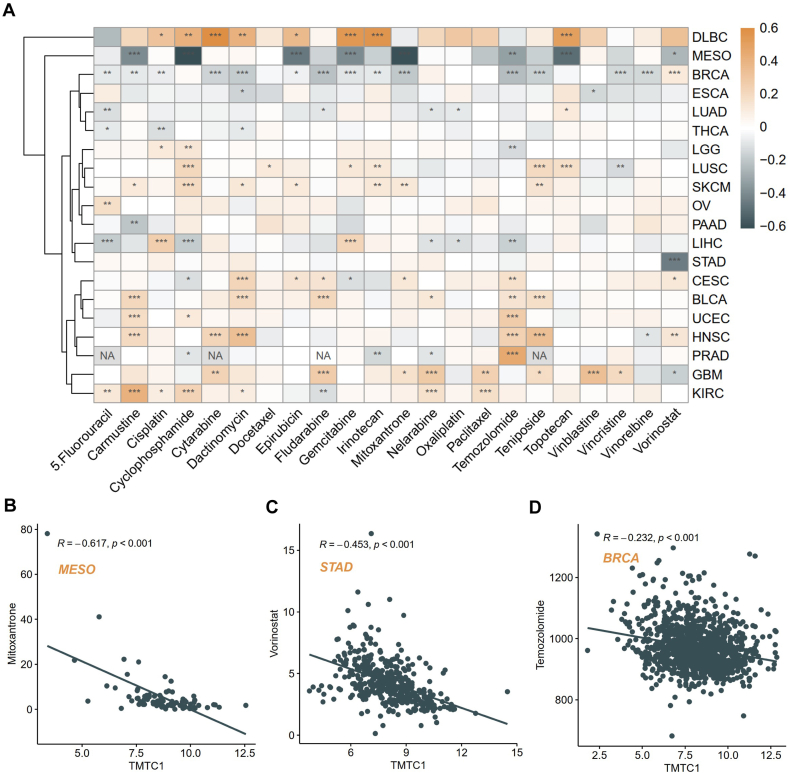
Drug sensitivity analysis of TMTC1
To examine the potential influence of TMTC1 on patients’ response to chemotherapy, we downloaded the IC50 (half-maximal inhibitory concentration) and expression data in cancer cell lines from the Genomics of Drug Sensitivity in Cancer (GDSC) database and then utilized the ‘OncoPredict’ package to predict the drug sensitivity of patients from TCGA across 20 cancer types. Subsequently, we examined the correlation between TMTC1 expression levels and the predicted drug sensitivity. As shown in Fig. 8A, our analysis revealed significant positive associations between TMTC1 expression and the sensitivity of 22 common antitumor drugs in multiple tumor types, including DLBC, SKCM, BLCA, and GBM. Conversely, a marked negative correlation was observed between TMTC1 expression and the sensitivity of these drugs in MESO and RBCA. Specifically, the overexpression of TMTC1 significantly decreased the IC50 values of mitoxantrone in MESO (Fig. 8B), vorinostat in STAD (Fig. 8C), and temozolomide in BRCA (Fig. 8D). This suggests that elevated TMTC1 expression reduces the resistance of tumor cells to these drugs, thereby enhancing their effectiveness in treatment.
TMTC1-related gene enrichment analysis
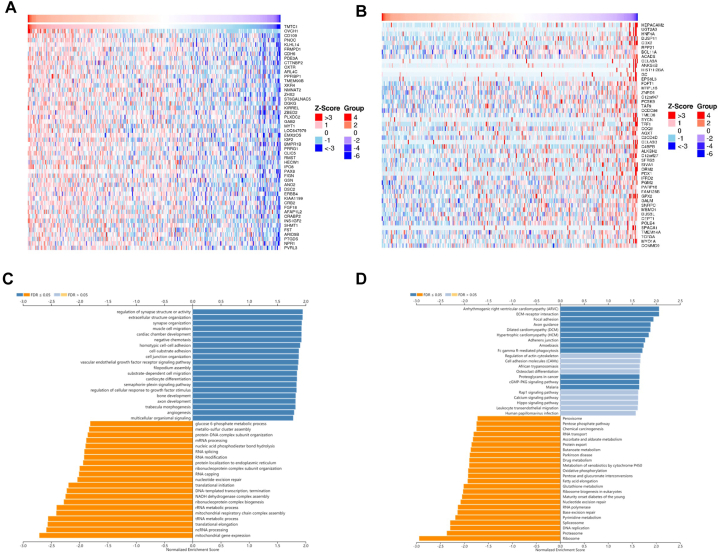
To investigate the functions of TMTC1 in cancer, we utilized the LinkedOmics tool to identify 100 genes that were related with TMTC1 in ovarian serous cystadenocarcinoma. Fig.s 9A and 9B present heat maps of 50 genes that are significantly positively correlated and 50 genes that are significantly negatively correlated with TMTC1 expression, respectively. Through Gene Set Enrichment Analysis, which was conducted using the 100 TMTC1-related genes in ovarian serous cystadenocarcinoma, it was found that GO biological process terms primarily showed enrichment in regulation of synapse structure or activity, extracellular structure organization, synapse organization, muscle cell migration, cardiac chamber development, negative chemotaxis, homotypic cell-cell adhesion, cell-substrate adhesion, cell junction organization, vascular endothelial growth factor receptor signaling pathway, filopodium assembly, substrate-dependent cell migration, cardiocyte differentiation, semaphorin-plexin signaling pathway, regulation of cellular response to growth factor stimulus, bone development, axon development, trabecula morphogenesis, and angiogenesis (Fig. 9C). Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis showed that the major enriched pathways were arrhythmogenic right ventricular cardiomyopathy (ARVC), ECM-receptor interaction, focal adhesion, axon guidance, dilated cardiomyopathy (DCM), hypertrophic cardiomyopathy (HCM), adherens junction, amoebiasis, Fc gamma R-mediated phagocytosis, proteoglycans in cancer, and GMP-PKG signaling pathway (Fig. 9D).
Discussion
TMTC1 is an essential ER protein that helps maintain ER calcium balance [ref. 5] and is involved in protein glycosylation, crucial for protein folding, stability, and function [ref. 7,ref. 9,ref. 43]. Dysregulation of TMTC1 leads to altered intracellular Ca2+ levels and impaired glycosylation, both linked to cancer development and progression [ref. [44], ref. [45], ref. [46], ref. [47]]. Although TMTC1 dysregulation is associated with several cancers. [ref. 10,ref. 12,ref. 16,ref. 17], a comprehensive pan-cancer analysis was lacking. Our study addresses this by examining TMTC1‘s expression, prognosis, genetic alterations, immune infiltration, immunotherapy response, drug sensitivity, and regulatory mechanisms across various cancers, aiming to explore the prognostic significance of TMTC1 and its potential as an immunotherapeutic target.
Firstly, our analysis revealed that TMTC1 exhibits lower expression in 19 cancers (ACC, BLCA, BRCA, CESC, COAD, ESCA, GBM, KICH, KIRC, KIRP, LAML, LUAD, LUSC, PRAD, READ, STAD, THCA, UCEC, and UCS) compared to normal tissues. This finding was consistent with previous research on TMTC1 in GBM [ref. 17], BRCA [ref. 15], and STAD [ref. 14]. The downregulation of TMTC1 in cancer may have significant implications for tumor progression., as reduced TMTC1 expression may lead to a decrease in O-mannosylation of the tumor suppressor protein E-cadherin, which promoted tumor spread [ref. 7,ref. 46]. Additionally, our analysis demonstrated that elevated expression of TMTC1 is associated with poor prognosis in several cancers, including ACC, BLCA, COAD, GBM, KIRP, OV, STAD, and UCEC. These findings were consistent with previous studies conducted by Yeh et al. in ovarian cancer [ref. 8], Xin Chen et al. in gastric cancer [ref. 14], and McInerney et al. in GBM [ref. 17]. Conversely, TMTC1 expression showed a protective effect on survival in patients with CESC, KIRC, LUSC, and PAAD. These findings highlight the heterogeneity of TMTC1‘s prognostic significance across different cancer types and emphasize its potential as a biomarker for predicting patient outcomes.
Mutations or functional loss in TMTC proteins are closely associated with various diseases, such as hearing loss [ref. [48], ref. [49], ref. [50]], neuronal cell migration disorders [ref. 51,ref. 52], brain development abnormalities [ref. 51], and brain malformations [ref. 52]. Our study is the first to analyze the relationship between TMTC1 genetic alterations and cancer. Our findings show that TMTC1 alterations, primarily mutations and amplifications, are most frequent in LUAD, OV, and TGCT. These genetic alterations of TMTC1 might be related to the mechanisms and molecular characteristics underlying different tumor types. Additionally, TMTC1 genetic alterations significantly affect patient survival across various cancers, suggesting TMTC1 mutations could be potential prognostic markers for survival and treatment response.
Our analysis found a significant positive correlation between TMTC1 expression and stromal score across various cancers, suggesting TMTC1‘s role in stromal cell infiltration. TMTC1 expression also correlated positively with cancer-associated fibroblasts (CAFs) and endothelial cell infiltration. CAFs promote tumor cell proliferation, migration, and invasion by secreting growth factors, cytokines and extracellular matrix proteins [ref. 53]. Endothelial cells support tumor angiogenesis by producing factors like VEGF and PDGF. The association between TMTC1 expression and the infiltration of CAFs and endothelial cells emphasizes its potential involvement in cancer metastasis. Single-cell analysis using the TISCH database showed higher TMTC1 expression in stromal cells, particularly fibroblasts, endothelial cells, and myofibroblasts, compared to immune cells. This suggests TMTC1 plays a significant role in the non-immune components of the tumor microenvironment.
Immunotherapy has emerged as a promising treatment approach for various cancers [ref. 54]. One of the key strategies in immunotherapy is the use of immune checkpoint inhibitors, which are drugs that block the inhibitory signals that allow cancer cells to evade the immune system [ref. 55]. Our study reveals that high TMTC1 expression is associated with improved prognosis in cancer patients receiving immune checkpoint inhibitors. Specifically, patients with elevated TMTC1 levels show longer progression-free survival (PFS) and overall survival (OS) compared to those with low TMTC1 expression. These results suggest that TMTC1 could be an important factor in predicting and enhancing the effectiveness of immunotherapy.
Chemotherapy is a common treatment approach for cancer, which utilizes drugs to destroy or inhibit the growth of cancer cells in the body [ref. [56], ref. [57], ref. [58]]. Our analysis explored the potential influence of TMTC1 on patients’ response to chemotherapy. We observed significant associations between TMTC1 expression and the sensitivity of several common antitumor drugs in different tumor types. This analysis highlights the potential role of TMTC1 in mediating drug sensitivity in various cancer types. It suggests that targeting TMTC1 and modulating its expression could be a promising strategy to improve the efficacy of chemotherapy in certain tumor types. Further studies are warranted to elucidate the underlying mechanisms and validate these findings in clinical settings.
Recent research has revealed that TMTC1 promotes the occurrence and progression of ovarian cancer, indicating its potential as a therapeutic target [ref. 8]. To further investigate the molecular mechanisms of TMTC1 in ovarian cancer, we conducted gene enrichment analysis using TMTC1-related genes in ovarian serous cystadenocarcinoma. The results revealed significant enrichment in biological processes and pathways associated with cell migration, adhesion, and angiogenesis. Cell migration played a crucial role in the development and progression of cancer by facilitating the movement of cancer cells from the primary tumor site to distant organs, thereby promoting metastasis [ref. 59]. Cell adhesion refers to the ability of cancer cells to attach to neighboring cells or the surrounding extracellular matrix. Alterations in the adhesion properties of cancer cells endow them with an invasive and migratory phenotype [ref. 60]. Moreover, angiogenesis played a critical role in tumor growth by promoting the formation of new blood vessels to provide nutrients and oxygen to the growing tumor [ref. 61]. The significant correlation between TMTC1 and these processes suggested that TMTC1 may contribute to cancer aggressiveness and metastasis by directly or indirectly influencing cell migration, adhesion, and angiogenesis.
Our study is the reliance on publicly available datasets, which may introduce biases and inconsistencies in data collection and processing. Additionally, the regulatory mechanisms underlying the dysregulation of TMTC1 in cancer remain largely unknown. Therefore, future studies should focus on elucidating these mechanisms and exploring potential therapeutic strategies targeting TMTC1. Furthermore, the functional role of TMTC1 in cancer was primarily explored through bioinformatics analysis, and further experimental validation is necessary to confirm these findings.
Conclusion
In conclusion, this study comprehensively analyzed the expression, prognosis, genetic alterations, immune infiltration, immunotherapy response, drug sensitivity, and regulatory mechanisms of TMTC1 in pan-cancer. The findings highlight the prognostic significance of TMTC1 and its potential as an immunotherapeutic target.
Data availability statement
The RNA sequencing and paired OS outcomes of thirty-three cancer types form TCGA were acquired from the UCSC Xena database (https://xenabrowser.net/datapages/). The PD1/PD-L1 therapy cohorts (IMvigor210 cohort and GSE176307) from the GEO database and studies conducted by Balar [ref. 62]. The IC50 values and transcriptomic data for cell lines treated with 198 drugs were downloaded from the GDSC2 database (https://www.cancerrxgene.org). Readers could download all data included in this study by searching accession numbers mentioned in Data acquisition of Method.
Funding
This work did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Ethics statement
All the data included in the analysis were obtained from public databases without the need of permissions from local ethical committees.
CRediT authorship contribution statement
Ying Zhang: Writing – review & editing, Writing – original draft. Dan Wu: Writing – review & editing, Writing – original draft. Tiantian Yu: Formal analysis, Data curation. Yao Liu: Methodology, Formal analysis, Data curation. Chunbo Zhao: Writing – review & editing, Visualization, Conceptualization. Ruihong Xue: Conceptualization.
Declaration of competing interest
The authors declare the following financial interests/personal relationships which may be considered as potential competing interests:There no any other relationship or activity that may be interpreted as a conflict of interest by the reader. If there are other authors, they declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- F. Bray. The ever‐increasing importance of cancer as a leading cause of premature death worldwide. Cancer, 2021. [PubMed]
- H. Sung. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA A Cancer J. Clin., 2021
- B. Eisenhaber. Conserved sequence motifs in human TMTC1, TMTC2, TMTC3, and TMTC4, new O-mannosyltransferases from the GT-C/PMT clan, are rationalized as ligand binding sites. Biol. Direct, 2021
- J.B. Graham. Endoplasmic reticulum transmembrane protein TMTC3 contributes to O-mannosylation of E-cadherin, cellular adherence, and embryonic gastrulation. Mol. Biol. Cell, 2020. [PubMed]
- J.C. Sunryd. TMTC1 and TMTC2 are novel endoplasmic reticulum tetratricopeptide repeat-containing adapter proteins involved in calcium homeostasis. J. Biol. Chem., 2014. [PubMed]
- J. Li. Deletion of Tmtc4 activates the unfolded protein response and causes postnatal hearing loss. J. Clin. Invest., 2018. [PubMed]
- I.S.B. Larsen. Discovery of an O-mannosylation pathway selectively serving cadherins and protocadherins. Proc. Natl. Acad. Sci. USA, 2017. [PubMed]
- T.-C. Yeh. TMTC1 promotes invasiveness of ovarian cancer cells through integrins β1 and β4. Cancer Gene Ther., 2023. [PubMed]
- I.S.B. Larsen. Multiple distinct O-Mannosylation pathways in eukaryotes. Curr. Opin. Struct. Biol., 2019. [PubMed]
- T.-C. Yeh. TMTC1 promotes invasiveness of ovarian cancer cells through integrins β1 and β4. Cancer Gene Ther., 2023
- X. Shen. Circular RNA profiling identified an abundant circular RNA circTMTC1 that inhibits chicken skeletal muscle satellite cell differentiation by sponging miR-128-3p. Int. J. Biol. Sci., 2019. [PubMed]
- A. Verma. Association of inflammatory bowel disease with arthritis: evidence from in silico gene expression patterns and network topological analysis. Interdiscipl. Sci. Comput. Life Sci., 2017
- R.G. Mealer. Glycobiology and schizophrenia: a biological hypothesis emerging from genomic research. Mol. Psychiatr., 2020
- X. Chen, Q. Zhang, T. Chekouo. Filtering high-dimensional methylation marks with extremely small sample size: an application to gastric cancer data. Front. Genet., 2021
- R. Ali. Identification of new key genes and their association with breast cancer occurrence and poor survival using in silico and in vitro methods. Biomedicines, 2023
- F. Moccia. Breast and renal cancer—Derived endothelial colony forming cells share a common gene signature. Eur. J. Cancer, 2017. [PubMed]
- C.E. McInerney. Using AI-based evolutionary algorithms to elucidate adult brain tumor (glioma) etiology associated with IDH1 for therapeutic target identification. Curr. Issues Mol. Biol., 2022. [PubMed]
- Y. Zhao. CircTMTC1 contributes to nasopharyngeal carcinoma progression through targeting miR-495-MET-eIF4G1 translational regulation axis. Cell Death Dis., 2022
- M.J. Goldman. Visualizing and interpreting cancer genomics data via the Xena platform. Nat. Biotechnol., 2020. [PubMed]
- G. Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science, 2015. [PubMed]
- N. Riaz. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell, 2017. [PubMed]
- T.N. Gide. Distinct immune cell populations define response to anti-PD-1 monotherapy and anti-PD-1/anti-CTLA-4 combined therapy. Cancer Cell, 2019. [PubMed]
- S. Mariathasan. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature, 2018. [PubMed]
- T.L. Rose. Fibroblast growth factor receptor 3 alterations and response to immune checkpoint inhibition in metastatic urothelial cancer: a real world experience. Br. J. Cancer, 2021. [PubMed]
- R.J. Motzer. Avelumab plus axitinib versus sunitinib in advanced renal cell carcinoma: biomarker analysis of the phase 3 JAVELIN Renal 101 trial. Nat. Med., 2020. [PubMed]
- D.A. Braun. Interplay of somatic alterations and immune infiltration modulates response to PD-1 blockade in advanced clear cell renal cell carcinoma. Nat. Med., 2020. [PubMed]
- A. Kassambara, M.A. Kassambara. Package ‘ggpubr’. R package version 0.1, 2020
- F. Pontén, K. Jirström, M. Uhlen. The human protein atlas—a tool for pathology. J. Pathol.: A Journal of the Pathological Society of Great Britain and Ireland, 2008
- T.M. Therneau, T. Lumley. Package ‘survival’. R Top Doc, 2015
- M. Gordon, T. Lumley, M.M. Gordon. 2019
- A. Kassambara. 2017
- W. Shen. Sangerbox: a comprehensive, interaction‐friendly clinical bioinformatics analysis platform. Imeta, 2022. [PubMed]
- K. Yoshihara. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun., 2013. [PubMed]
- R. Kolde, M.R. Kolde. Package ‘pheatmap’. R package, 2015
- J. Racle, D. Gfeller. EPIC: a tool to estimate the proportions of different cell types from bulk gene expression data. Bioinformatics for Cancer Immunotherapy: Methods and Protocols, 2020
- E. Becht. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome biology, 2016. [PubMed]
- D. Aran, Z. Hu, A.J. Butte. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome biology, 2017. [PubMed]
- D. Sun. TISCH: a comprehensive web resource enabling interactive single-cell transcriptome visualization of tumor microenvironment. Nucleic acids research, 2021. [PubMed]
- K. Esfahani. A review of cancer immunotherapy: from the past, to the present, to the future. Curr. Oncol., 2020
- W. Yang. Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic acids research, 2012. [PubMed]
- D. Maeser, R.F. Gruener, R.S. Huang. oncoPredict: an R package for predicting in vivo or cancer patient drug response and biomarkers from cell line screening data. Briefings Bioinf., 2021
- S.V. Vasaikar. LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic acids research, 2018. [PubMed]
- B. Eisenhaber. Conserved sequence motifs in human TMTC1, TMTC2, TMTC3, and TMTC4, new O-mannosyltransferases from the GT-C/PMT clan, are rationalized as ligand binding sites. Biol. Direct, 2021. [PubMed]
- A. Bergner, R. Huber. Regulation of the endoplasmic reticulum Ca2+-store in cancer. Anti Cancer Agents Med. Chem., 2008
- N. Prevarskaya. Remodelling of Ca2+ transport in cancer: how it contributes to cancer hallmarks?. Phil. Trans. Biol. Sci., 2014
- S. Carvalho. O-mannosylation and N-glycosylation: two coordinated mechanisms regulating the tumour suppressor functions of E-cadherin in cancer. Oncotarget, 2016
- B.N. Vajaria, P.S. Patel. Glycosylation: a hallmark of cancer?. Glycoconj. J., 2017. [PubMed]
- C.L. Runge. Association of TMTC2 with human nonsyndromic sensorineural hearing loss. JAMA Otolaryngology–Head & Neck Surgery, 2016. [PubMed]
- H. Guillen‐Ahlers. TMTC2 variant associated with sensorineural hearing loss and auditory neuropathy spectrum disorder in a family dyad. Molecular Genetics & Genomic Medicine, 2018. [PubMed]
- J. Li. Deletion of Tmtc4 activates the unfolded protein response and causes postnatal hearing loss. The Journal of clinical investigation, 2018. [PubMed]
- J. Jerber. Biallelic mutations in TMTC3, encoding a transmembrane and TPR-containing protein, lead to cobblestone lissencephaly. Am. J. Hum. Genet., 2016. [PubMed]
- S.M. Farhan. Identification of a novel synaptic protein, TMTC3, involved in periventricular nodular heterotopia with intellectual disability and epilepsy. Hum. Mol. Genet., 2017. [PubMed]
- B. Erdogan, D.J. Webb. Cancer-associated fibroblasts modulate growth factor signaling and extracellular matrix remodeling to regulate tumor metastasis. Biochem. Soc. Trans., 2017. [PubMed]
- K.Y. Helmy. Cancer immunotherapy: accomplishments to date and future promise. Ther. Deliv., 2013. [PubMed]
- R.W. Jenkins, D.A. Barbie, K.T. Flaherty. Mechanisms of resistance to immune checkpoint inhibitors. Br. J. Cancer, 2018. [PubMed]
- D.L. Gustafson, R.L. Page. Cancer chemotherapy. Withrow and MacEwen’s small animal clinical oncology, 2013
- B.A. Chabner, T.G. Roberts. Chemotherapy and the war on cancer. Nat. Rev. Cancer, 2005. [PubMed]
- Jr V.T. DeVita, E. Chu. A history of cancer chemotherapy. Cancer Res., 2008. [PubMed]
- H. Yamaguchi, J. Wyckoff, J. Condeelis. Cell migration in tumors. Curr. Opin. Cell Biol., 2005. [PubMed]
- N. Makrilia. Cell adhesion molecules: role and clinical significance in cancer. Cancer Invest., 2009. [PubMed]
- N. Nishida. Angiogenesis in cancer. Vasc. Health Risk Manag., 2006. [PubMed]
- A.V. Balar. Atezolizumab as first-line treatment in cisplatin-ineligible patients with locally advanced and metastatic urothelial carcinoma: a single-arm, multicentre, phase 2 trial. Lancet, 2017. [PubMed]
