Characterization of Clostridium perfringens Phage Endolysin PlyDolk21
Abstract
Background: Clostridium perfringens is a significant cause of food poisoning. Broad-spectrum antibiotics, commonly used to control C. perfringens, are becoming less effective due to the rise of antibiotic-resistant strains, necessitating alternative control strategies. Methods: A C. perfringens-infecting bacteriophage, Dolk21, and its endolysin, PlyDolk21, were isolated and characterized. The lytic activity of PlyDolk21 was assessed in comparison to its catalytic domain alone. Both PlyDolk21 and its cell wall binding domain (CBD) were evaluated in beef and milk for their antimicrobial activity and cell wall binding activity, respectively. Results: While phage Dolk21 was specific to certain C. perfringens strains, PlyDolk21 exhibited lytic activity against all C. perfringens strains tested. The full-length PlyDolk21 showed stronger lytic activity compared to its catalytic domain alone. PlyDolk21_CBD successfully bound to C. perfringens in vitro and in foods. Additionally, PlyDolk21 effectively reduced the viable cell counts of C. perfringens by 3-log in beef soup and milk samples. Conclusions: This study demonstrates that PlyDolk21 and its CBD hold potential as a biocontrol and detection agent targeting C. perfringens in various food matrices.
Article type: Research Article
Keywords: bacteriophage, endolysin, cell wall-binding domain, biocontrol
Affiliations: Department of Food Science and Biotechnology, Research Institute of Food and Biotechnology, Seoul National University of Science and Technology, Seoul 01811, Republic of Korea; suyoung4330@snu.ac.kr; Department of Food Biotechnology, Dong-A University, Busan 49315, Republic of Korea; bkson@dau.ac.kr
License: © 2025 by the authors. CC BY 4.0 Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Article links: DOI: 10.3390/antibiotics14010081 | PubMed: 39858367 | PMC: PMC11762992
Relevance: Relevant: mentioned in keywords or abstract
Full text: PDF (2.5 MB)
1. Introduction
Clostridium perfringens is a Gram-positive, rod-shaped, spore-forming anaerobe that causes a wide range of human and veterinary diseases such as gas gangrene and non-foodborne gastrointestinal infections [ref. 1,ref. 2]. In addition, C. perfringens is responsible for two distinct foodborne diseases in humans [ref. 3,ref. 4]. One is a common form of foodborne illness, typically leading to mild symptoms, which is classic A diarrhea, while the other is Type C necrotic enteritis, a more severe but rare condition [ref. 3]. In the U.S., C. perfringens is the second most important foodborne pathogen [ref. 5,ref. 6], and is also responsible for over 4 million infections worldwide annually [ref. 7]. Broad-spectrum antibiotics are frequently used to manage C. perfringens, but the emergence of antibiotic-resistant strains has reduced the effectiveness of these traditional treatments [ref. 8]. Therefore, alternative strategies for controlling this bacterium are crucial.
Bacteriophages and their lytic enzyme, endolysins, have garnered significant attention as a potential solution for combating antibiotic-resistant bacteria [ref. 9,ref. 10]. During phage infection, endolysin is synthesized in the late stage of phage multiplication, followed by degrading the peptidoglycan layer of host bacteria [ref. 11,ref. 12]. The use of phages has many advantages including host specificity, self-amplification, and safety [ref. 13,ref. 14]. However, lysogenic phages could carry the risk of transferring virulent or antibiotic-resistant genes [ref. 15]. In contrast, endolysins offer additional benefits over the use of phages, such as broad-spectrum activity, no transduction issues, and the absence of reported bacteria, making them promising candidates for next-generation antimicrobial agents [ref. 10]. The structures of endolysins are different between Gram-positive and Gram-negative phages [ref. 16]. Endolysins from Gram-negative phages are typically single-domain proteins, lacking a cell wall-binding domain (CBD), because the outer membrane limits access to the peptidoglycan, reducing the need for a cell-binding domain. In contrast, most endolysins from Gram-positive phages consist of more than two domains, mainly an N-terminal enzymatic active domain (EAD) and a C-terminal CBD, which are connected by a flexible linker [ref. 17]. Endolysins from C. perfringens phages generally exhibit C. perfringens-specific lytic activity, with the CBD likely contributing to the species specificity of these endolysins [ref. 18]. However, recent X-ray crystal structures and modeling studies of C. perfringens endolysins have suggested that the EAD itself may also have species-specific enzymatic activity because the endolysin cleavage site is located near the unique peptide bridges in the C. perfringens peptidoglycan [ref. 19]. Thus, analysis of each domain is as important as studying full-length endolysin since this approach allows for a better understanding of the specific contributions of both the EAD and CBD to species specificity and lytic activity. Such insights can inform the development of more targeted and effective antimicrobial agents.
There has been a growing number of studies of C. perfringens-targeting endolysins, including those with Generally Recognized as Safe (GRAS, GRN000802) [ref. 20]-approved status [ref. 19,ref. 21]. However, while endolysins have been extensively studied in clinical and veterinary settings, research into their application in food systems remains relatively limited [ref. 22]. This gap highlights the need for further exploration of endolysins as potential antimicrobial agents in food safety and preservation.
In this study, a novel virulent C. perfringens phage Dolk21 was isolated and its biological and genomic characteristics were investigated. An endolysin gene, plyDolk21, was identified in the genome of the phage Dolk21. The antimicrobial spectrum and lytic activity of the full-length PlyDolk21 were compared to those of its enzymatically active domain (PlyDolk21_EAD). Predicted CBD (PlyDolk21_CBD) was purified with EGFP and tested for its cell wall-binding ability. In foods, the full-length endolysin was evaluated for its antimicrobial ability against C. perfringens and the CBD protein for its binding ability to C. perfringens. The results could accelerate developing biocontrol and detection technologies for the effective control of C. perfringens foodborne illnesses.
2. Results and Discussion
2.1. Analysis and Characterization of C. perfringens Phage Dolk21
2.1.1. Lytic Activity and Host Range of Phage Dolk21
A novel bacteriophage, Dolk21, was isolated from a soil sample in Bucheon, Republic of Korea using C. perfringens ATCC 13124 as the host strain. To determine the host range of the phage Dolk21, a spotting assay was conducted, with bacterial strains shown in Table 1. The Dolk21 phage was capable of lysing four out of the ten C. perfringens strains tested, forming clear plaques. All other Gram-positive and Gram-negative bacterial species exhibited resistance to Dolk21. These findings indicate that the phage Dolk21 has highly restricted host specificity, prompting us to explore alternative approaches, such as utilizing its endolysin, to develop an efficient biocontrol agent capable of targeting multiple strains of C. perfringens.
Table 1: Host range of phage Dolk21, lytic activity of PlyDolk21 and its enzymatic active domain PlyDolk21_EAD, and cell wall-binding activity of PlyDolk21_CBD.
| Species | Strain No. a | Dolk21 b | PlyDolk21 c | PlyDolk21_EAD c | PlyDolk21_CBD d |
|---|---|---|---|---|---|
| C. perfringens | 2 | + | +++ | ++ | + |
| C. perfringens | 24 | + | ++ | + | + |
| C. perfringens | 2585 | − | +++ | + | + |
| C. perfringens | 2589 | + | +++ | + | + |
| C. perfringens | ATCC 3624 | − | +++ | + | + |
| C. perfringens | ATCC 13124 | + | +++ | ++ | + |
| C. perfringens | NCCP 15911 | − | ++ | + | + |
| C. perfringens | H3 | − | ++ | + | + |
| C. perfringens | H9 | − | ++ | + | + |
| C. perfringens | FD1 | − | +++ | ++ | + |
| Other Gram-positive | |||||
| Bacillus cereus | ATCC 10987 | − | − | − | − |
| Bacillus subtilis | ATCC 23857 | − | + | ++ | − |
| Staphylococcusaureus | Newman | − | − | − | − |
| Listeriamonocytogenes | ATCC 15313 | − | − | + | − |
| Geobacillusstearothermophilus | ATCC 10149 | − | − | ++ | − |
| Levilactobacillus brevis | ATCC 11433 | − | − | − | − |
| Gram-negative | − | ||||
| E. coli O157:H7 | ATCC 35150 | − | − | − | − |
| Salmonella Typhimurium | LT2 | − | − | − | − |
| Pseudomonasaeruginosa | PAO1 | − | + | + | − |
| Cronobactersakazakii | ATCC 29544 | − | + | + | − |
a ATCC, American Type Culture Collection; NCCP, National Culture Collection for Pathogens. b +, activity of dotting assay; −, no activity. c −, 0–10%; +, 11–40%; ++, 41–70%; +++, 71–100%. d +, activity of cell wall-binding assay; −, no activity.
2.1.2. Morphology and Genome Characterization
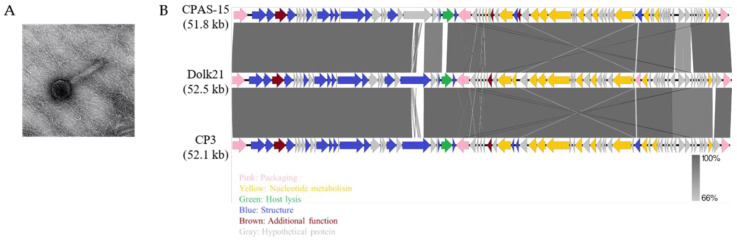
The TEM analysis revealed that the Dolk21 phage belongs to the class Caudoviricetes. Dolk21 has a thick and contractile tail, indicative of a myovirus-like morphology (Figure 1A). Its icosahedral head is 74.49 ± 3.40 nm (n = 13), and the length of its tail is 155.74 ± 8.24 nm (n = 12). The genome sequence analysis of C. perfringens phage Dolk21 indicated that its genome consists of 52,463 base pairs of double-stranded DNA, containing 34% of G+C content The phage’s genome encodes a total of 75 ORFs, of which the functional roles of 31 ORFs were predicted. The functional ORFs were classified into five functional groups of packaging, structure, host lysis, nucleotide metabolism, and additional function (Figure 1B). As there were no lysogeny-related genes found, such as an integrase, a recombinase, or a repressor, this suggested that Dolk21 is likely a virulent phage. BLASTN analysis revealed that the Dolk21 genome shares 97.8% and 98.0% of its DNA sequence identity with Clostridium phages CP3 (GenBank Accession No. ASZ76631.1) and CPAS-15 (GenBank Accession No. QGF20128.1), respectively.
2.2. Production and Characterization of PlyDolk21
2.2.1. Identification and Expression of the Endolysin PlyDolk21
The putative endolysin gene was identified from the phage Dolk21 genome and designated as plyDolk21. Amino acid sequence analysis using BLASTP (https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Proteins, accessed on 7 January 2025) and InterProScan (version 5.72-103.0) revealed that PlyDolk21 consists of two functional domains, an N-terminal amidase_2 domain as the EAD and a C-terminal SH3_3 domain as the CBD (Figure 2A). The segment between these two domains did not match any known functional domain in the current databases. Further experimental or computational studies would be necessary to assign a potential function to this region. BLASTP analysis showed several proteins homologous to PlyDolk21, but most of them have not yet been studied. The amino acid sequence of the PlyDolk21 was aligned with those of endolysins from C. perfringens phages CP3, Clo-PEP-1 (GenBank Accession No. APQ41998.1), CPAS-15, and CPD4 (GenBank Accession No. MK017819.1), which exhibit high similarity to PlyDolk21 (Figure 2B). Notably, the sequence of the putative CBD of PlyDolk21 is highly homologous to that of the phage CPAS-15 endolysin, which was experimentally confirmed to have cell wall-binding activity, while the EAD differs significantly. Based on this information, the PlyDolk21 and each domain were cloned and expressed in E. coli with an N-terminal His-tag. A single band of each purified protein was identified on sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE), and each band corresponded to the expected molecular mass of the respective protein (Figure 2C).
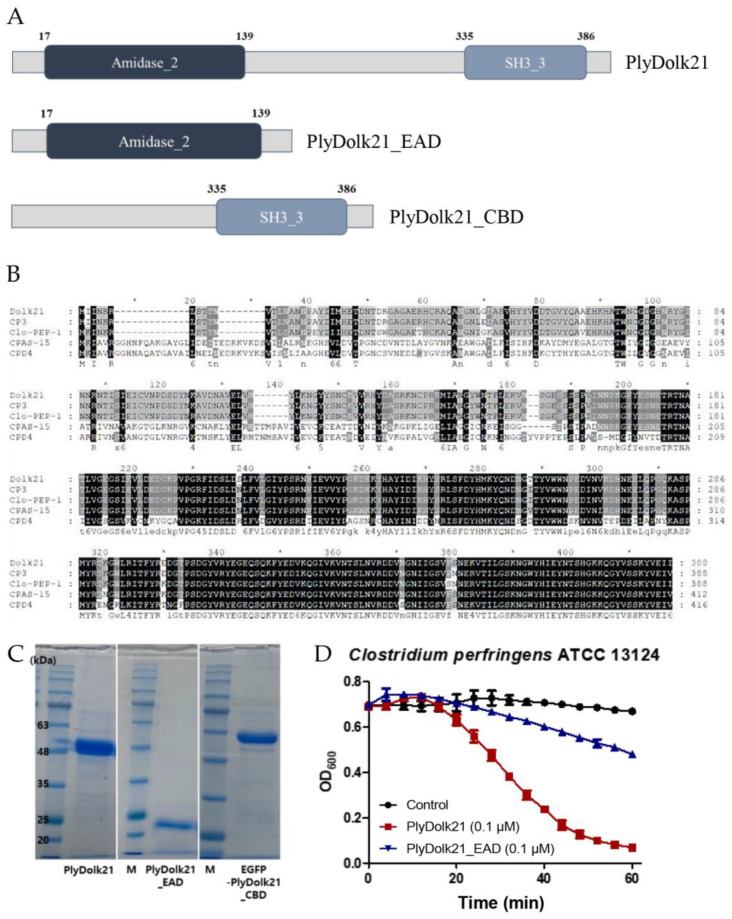
2.2.2. Antimicrobial Activity of PlyDolk21 and PlyDolk21_EAD
The antimicrobial activity of PlyDolk21 and PlyDolk21_EAD was evaluated by a turbidity reduction assay. The treatment of 0.1 μM of PlyDolk21 and PlyDolk21_EAD dropped the turbidity of the C. perfringens cell suspension by 85% and 30%, respectively, indicating that CBD was necessary for PlyDolk21 to have its maximal lytic activity (Figure 2D). This phenomenon was particularly observed when endolysins targeted their natural host bacteria. In a previous study, the lytic activity of LysPBC1 and its enzymatically active domain (LysPBC1_EAD) was compared against both target and non-target bacteria. The findings showed that the C-terminal domain of LysPBC1 was required for full activity against its natural target, but not for non-target bacteria [ref. 23].
The antimicrobial spectral analysis showed that PlyDolk21 and PlyDolk21_EAD inhibited the growth of all tested strains of C. perfringens, B. subtilis, P. aeruginosa, and C. sakazakii, demonstrating a much broader lytic spectrum compared to phage Dolk21 (Table 1). It should be noted that PlyDolk21 and PlyDolk21_EAD showed antimicrobial activity against P. aeruginosa and C. sakazakii without the addition of outer membrane permeabilizers. This lytic activity may be attributed to the attached His-tag, which likely enhanced interactions with the highly negatively charged outer membrane of P. aeruginosa and C. sakazakii [ref. 24]. In particular, the membrane lipid A of P. aeruginosa was modified with 4-amino-4-deoxy-L-arabinose (L-Ara4N) or phosphoethanolamine, further increasing its negative charge and potentially facilitating stronger electrostatic interactions with the His-tag of the proteins tested [ref. 25]. Similarly, C. sakazakii possesses a negatively charged outer membrane with lipid A modifications like L-Ara4N or phosphoethanolamine [ref. 26]. The turbidity reduction assay using PlyDolk21_EAD without the His-tag exhibited abolished antimicrobial activity (Figure S1), further emphasizing the critical role of the His-tag in facilitating interactions with the negatively charged outer membrane of P. aeruginosa and C. sakazakii. However, the exact mechanisms by which these endolysins permeate the outer membrane remain unclear. Notably, PlyDolk21_EAD also exhibited lytic activity against L. monocytogenes and Geobacillus stearothermophilus. It is known that the cell wall-binding domain of endolysins can inhibit lytic activity when not bound to its target [ref. 27], which may explain why the antimicrobial spectrum of the PlyDolk21_EAD appeared broader than that of the full-length endolysin, PlyDolk21. In this case, the presence or absence of the His-tag could not have significantly affected the activity of PlyDolk21_EAD. Indeed, we observed that the lytic rate of PlyDolk21_EAD against G. stearothermophilus was even faster in the absence of the His-tag (Figure S1), suggesting that its activity against L. monocytogenes and Geobacillus may not have strongly relied on the presence of the His-tag.
2.2.3. Stability of PlyDolk21 Under Various Stress Conditions
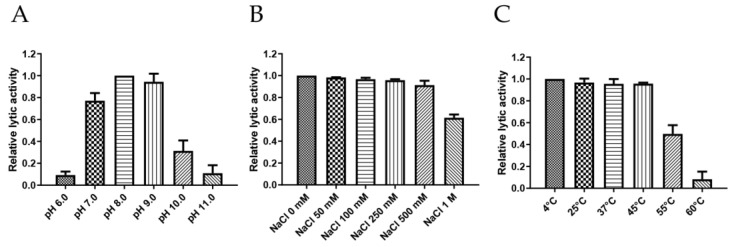
Endolysin should maintain stable activity under various stress conditions, including pH, temperature, and NaCl concentrations, to ensure its broad applicability [ref. 28]. To this end, the stability of PlyDolk21 was tested under a broad range of pH (6.0 to 11.0), temperature (4 °C to 60 °C), and NaCl concentrations (0 mM to 1000 mM) (Figure 3). PlyDolk21 showed maximal lytic activity between pH 7.0 and 9.0, but its activity diminished under acidic conditions (pH < 6.0) (Figure 3A). PlyDolk21 was relatively stable across all tested NaCl concentrations, maintaining high levels of lytic activity (Figure 3B). When tested under varying temperatures, its lytic activity remained stable between 4 °C and 45 °C, but gradually declined at 55 °C (Figure 3C).
2.3. Isolation and Binding Activity Analysis of PlyDolk21_CBD
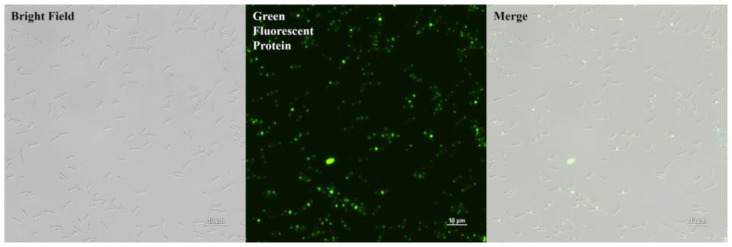
PlyDolk21_CBD was purified as shown in Figure 2C. The specific bacterial binding activity of the putative PlyDolk21_CBD was identified with an enhanced green fluorescent protein (EGFP), demonstrating that it primarily binds to the septal region and pole of the cells (Figure 4). The septal region undergoes active cell wall remodeling during cell division. Since amidases are involved in peptidoglycan hydrolysis in the septal region, its CBD could have high affinity to this region [ref. 29,ref. 30]. All tested C. perfringens strains were detected by EGFP_PlyDolk21_CBD, but not other Clostridium species or other Gram-positive and Gram-negative bacteria (Table 1 and Figure S2). These findings suggest that PlyDolk21_CBD could serve as a valuable material for the development of C. perfringens detection tools.
2.4. Food Applications
2.4.1. Bacterial Control of PlyDolk21 in Food Samples
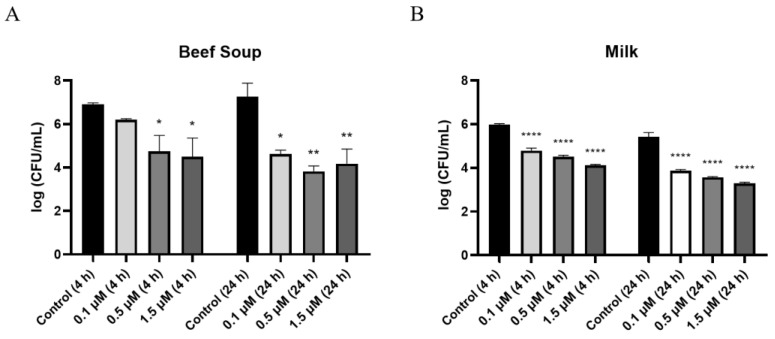
As commercial milk and beef are often reported to be contaminated with C. perfringens [ref. 31], sterilized milk and beef soup were chosen to evaluate the inhibitory activity of PlyDolk21 against C. perfringens in a food environment. In beef soup, treatment with 0.1 μM PlyDolk21 resulted in a 1-log reduction in CFUs after 4 h of incubation and a 3-log reduction after 24 h. (Figure 5A). When the concentration increased to 0.5 μM, a 4-log reduction in CFUs was observed after 24 h of incubation (Figure 5A). In sterilized milk, treatment with 1.5 μM PlyDolk21 resulted in a 3-log reduction in C. perfringens counts after 24 h of incubation (Figure 5B). These findings suggest that PlyDolk21 could be utilized as a natural food preservative to control C. perfringens in milk and beef soup. There are a few studies that explore the potential of using endolysins in foods to inhibit C. perfringens. For instance, LysCP28 from C. perfringens phage vB_CpeS_BG3P was evaluated for its efficacy in reducing C. perfringens in duck meat [ref. 32]. Additionally, the phage endolysin cpp-lys was applied to lettuce, highlighting the promising role of endolysins as biocontrol agents in various food systems [ref. 33].
2.4.2. Cell Wall-Binding Ability of PlyDolk21_CBD in Foods
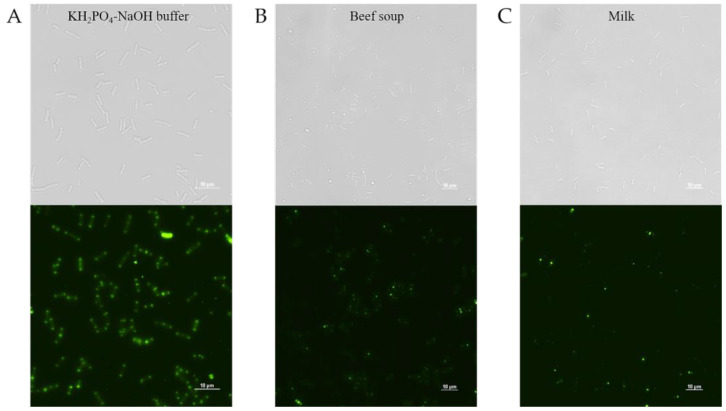
To evaluate the detection ability of PlyDolk21_CBD in foods, PlyDolk21_CBD fused with EGFP was added to a sterilized milk and beef soup artificially contaminated with C. perfringens. Given the presence of various components in food, CBD-bound cells were monitored after thorough washing. Although its binding activity was reduced by approximately 43.2% in milk and 38% in beef soup compared to that in the buffer, the EGFP-fused PlyDolk21_CBD successfully bound to C. perfringens within 5 min (Figure 6 and Figure S3). These findings suggest that the CBD protein may be useful for the rapid detection of specific pathogens in food [ref. 34,ref. 35].
3. Materials and Methods
3.1. Bacterial Strains and Growth Conditions
C. perfringens ATCC 13124 was used as a host strain for the isolation and propagation of the phage Dolk21. All tested strains are listed in Table 1. Clostridium strains and L. monocytogenes ATCC 15313 were grown in Brain Heart Infusion (BHI) broth at 37 °C under anaerobic conditions. B. subtilis ATCC 23857, S. aureus Newman, and C. sakazakii ATCC 29544 were grown in Tryptic Soy (TS) broth at 37 °C under shaking and Geobacillus stearothermophilus ATCC 10149 was grown in the same medium at 50 °C under shaking. B. cereus ATCC 10987 and the rest of the Gram-negative bacteria were grown in Luria–Bertani (LB) broth at 37 °C. E. coli BL21 was grown in LB broth at 37 °C and used for expression of the recombinant PlyDolk21 and its domains.
3.2. Isolation and Propagation of Bacteriophage Dolk21
A soil sample was collected from Bucheon, Republic of Korea. Phage isolation was performed as previously described with some modifications [ref. 23]. A 10 g sample was homogenized with 10 mL 2× BHI broth and 10 mL mixture was incubated with 400 μL cultured bacteria, 5 mM MgCl2, and 5 mM CaCl2. After an overnight anaerobic incubation at 37 °C, the culture was centrifugated (10,000× g at 4 °C for 10 min) and the supernatant was filtered using a 0.45 μm pore size filter (Sartorius AG, Göttingen, Germany). The presence of phages was confirmed using a plaque-forming assay with molten 0.7% DW soft agar inoculated with C. perfringens ATCC 13124 overnight cultures. After incubating at 37 °C overnight, a single plaque was picked with a sterile pipette tip and eluted in 1 mL of SM buffer (50 mM Tris-HCl [pH 7.5], 100 mM NaCl, and 10 mM MgCl2). These plaque isolation and elution steps were repeated five times to purify the single phage. Isolated phages were amplified through serial propagation and concentrated via polyethylene glycol (PEG) precipitation, followed by CsCl density gradient ultracentrifugation (78,500× g at 4 °C for 2 h) [ref. 36]. The concentrated phages were dialyzed with SM buffer for 2 h, with the buffer being changed two times during the dialysis.
3.3. Morphological Analysis by TEM
Purified Dolk21 (1.2 × 1010 PFU/mL) was placed on carbon-coated copper grids and negatively stained with 2% aqueous uranyl acetate (pH 4.0) by pipetting 10 times [ref. 23]. Dolk21 was visualized by energy-filtering transmission electron microscopy (EF-TEM, Thermo Fisher Scientific, Waltham, MA, USA) at 120 kV. Images were scanned at NICEM at Seoul National University (Seoul, Republic of Korea).
3.4. DNA Purification and Whole Genome Sequencing of Bacteriophage Dolk21
The Dolk21 genomic DNA extraction was carried out as previously described [ref. 36]. The purified genomic DNA of Dolk21 was sequenced using the Illumina Miseq platform (San Diego, CA, USA) and assembled with SPAdes v.3.15.2 (Sanigen, Anyang, Republic of Korea).
3.5. Bioinformatic Analysis
Functions of the predicted ORFs were confirmed and annotated using BLASTP and InterProScan [ref. 37] programs. The genome annotation identified an ORF encoding the endolysin PlyDolk21, along with its EAD and CBD. The primary protein structure of PlyDolk21 was compared with the endolysins of Clostridium phages (endolysin of phage CP3, NCBI Accession ID ASZ76631.1; endolysin of phage Clo-PEP-1, NCBI Accession ID APQ41998.1; endolysin of phage CPAS-15, NCBI Accession ID QGF20128.1; endolysin of phage CPD4, NCBI Accession ID KQC91385.1) using ClustalX2.1 and Genedoc v.2.6. To locate the EAD and CBD domains, secondary protein structures of PlyDolk21 were predicted using the NPS@ server (https://npsa.lyon.inserm.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_sopma.html, accessed on 7 January 2025) and PSIPRED v.4.02 tool, and tertiary protein structures of PlyDolk21 were predicted using the Phyre v.2.0 server and PyMOL v.3.0 program.
3.6. Cloning, Expression, and Purification
The endolysin gene (plyDolk21) was amplified from the genomic DNA of the bacteriophage Dolk21 by PCR using primers fPlyDolk21_BamHI (5′-GCG GGATCC ATGATAATCAATAAAAGATTAAGTACTACTAATGTTACCTTAAAC-3′) and rPlyDolk21_HindIII (5′-GCG AAGCTT CTAAATTATCTCTACATATTTTGAACTTACATAACCTTGC-3′). Additionally, the reverse primer rPlyDolk21_EAD_HindIII (5′-GCG AAGCTT CTATGAATTTACTCTTTCTAGGAATGTATTCCAGTAACC-3′) was designed for its EAD domain, and the forward primer fPlyDolk21_CBD_ BamHI (5′-GCG GGATCC GGAGAGAGTTCAAGCTCACCAGTAATTAATAATC-3′) was also designed for its CBD domain. The PCR products of plyDolk21 and plyDolk21_EAD were cloned into pET28a, which has an N-terminal hexahistidine (His6)-tag sequence. To check the binding to the cell wall with fluorescence, the enhanced green fluorescent protein (EGFP) gene was inserted in front of PlyDolk21_CBD using the NheI and BamHI restriction sites. Plasmids with each insert were transformed into a competent E. coli BL21 (DE3) cell. Each protein was overexpressed and purified following the methods described in a previous study [ref. 23]. A Thrombin CleanCleave Kit (Sigma-Aldrich, St. Louis, MO, USA) was used for N-terminus His6-tag cleavage. A total of 100 μL of washed thrombin–agarose resin was resuspended in 100 μL of 10X cleavage buffer (500 mM Tris-HCl, pH 8.0, 100 mM CaCl2), and 800 μg of purified PlyDolk21_EAD was added. The mixture was subsequently allowed to react at room temperature for 3 h, during which it was stirred gently. The cleavage of the His6-tag was confirmed using SDS-PAGE analysis (Figure S1).
3.7. Antimicrobial Spectrum of the Phages Dolk21, PlyDolk21, and PlyDolk21_EAD
The host range of the phage Dolk21 was confirmed, with the strains listed in Table 1. After incubation overnight, 100 μL of each strain was added to 5 mL of 0.7% molten soft agar and overlaid on a suitable agar plate. Then, 10 μL of each serially diluted phage suspension was spotted on the plate and incubated at 37 °C overnight. The antimicrobial spectra of PlyDolk21 and PlyDolk21_EAD were evaluated by turbidity reduction assay [ref. 23]. Bacterial cells were early exponentially grown and resuspended with reaction buffer (20 mM Tris-Cl, pH 8.0) to adjust the OD600 to approximately 0.6. The purified protein was added to a final concentration of 0.4 μM, and the inoculated cell resuspension was cultured at 25 °C. The OD600 values were monitored for 30 min with 1 min intervals at room temperature. The relative lytic activity (%) of PlyDolk21 and PlyDolk21_EAD was calculated using the Equation (1) presented below.
3.8. Stability Under Various Conditions
To evaluate the effect of pH on PlyDolk21, endolysin was added to C. perfringens ATCC 13124 cells suspended with a universal pH buffer. The universal buffer (0.04 M H3PO4, 0.04 M CH3COOH, 0.04 M H3BO3) was adjusted to different pH values, between 6.0 to 11.0, using 5 M NaOH. The effect of temperature on PlyDolk21 was tested at different temperatures (4–60 °C). The endolysin was incubated at each temperature for 20 min and added to cells suspended with reaction buffer. The influence of NaCl on PlyDolk21 was tested with reaction buffer containing 0 to 1000 mM NaCl. In each case, PlyDolk21 was treated to a final concentration of 0.4 μM, and the inoculated cell resuspension (106 CFU/mL) was incubated at 25 °C. The OD600 values were monitored for 30 min at 1 min intervals. The lytic activity under each stress condition was relatively measured based on Equation (2).
3.9. Food Applications of PlyDolk21
Milk and beef soup, which are frequently found with C. perfringens, were selected as model food samples. The antimicrobial activity in each food sample was tested as previously described [ref. 38]. Each sample artificially contaminated with C. perfringens ATCC 13124 (106 CFU/mL) was preincubated at 25 °C for 30 min. Subsequently, purified PlyDolk21 at concentrations of 0.1 μM, 0.5 μM, and 1.5 μM was added to the beef soup and milk. The mixture was incubated at 25 °C for 4 and 24 h. The antimicrobial activity of endolysin was determined by calculating the viable cell count, comparing between the control and test groups. All experiments were repeated three times. One-way analysis of variance (ANOVA) and the Tukey’s multiple comparison test were used for statistical analysis.
3.10. Cell Wall-Binding Assay with Fluorescence Microscopy of PlyDolk21_CBD
The results of the amino acid sequence alignment showed that the CBD region of PlyDolk21 has high similarity to that of C. perfringens phage endolysin LysCPAS15 [ref. 34]. Based on this finding, the CBD part of the gene plyDolk21 was cloned in fusion with a gene encoding EGFP and expressed in E. coli. To confirm the binding activity of PlyDolk21_CBD, 100 μL of purified protein at a final concentration of 1 μM was added to 100 μL of C. perfringens cells (106 CFU/mL) resuspended in KH2PO4-NaOH buffer (pH 7.0), and the mixture was incubated at RT for 5 min. Subsequently, cells were collected by centrifugation (16,000 × g for 1 min) and washed twice with KH2PO4-NaOH buffer (pH 7.0). Resuspended pellet with 10 μL KH2PO4-NaOH buffer (pH 7.0) was dropped on a slide glass and covered with an agarose pad (0.1 g of agarose powder (TransGen) + 13 mL of distilled water). An inverted microscope (ECLIPSE Ti2-E, Nikon, Tokyo, Japan) was used in this assay (DIA: 17.1, Intensilight ND 2, 1 s Auto exposure, 1.0× Analog Gain). For quantification, washed cells were resuspended in 200 μL of PBS buffer and their fluorescence was measured using a SpectraMax i3x plate reader (Molecular Devices, San Jose, CA, USA) with excitation at 485 nm and emission at 520 nm.
3.11. Food Applications of EGFP-Fused PlyDolk21_CBD
The ability of the endolysin PlyDolk21 to control C. perfringens ATCC 13124 was demonstrated in the milk and beef soup. To evaluate the detection ability of PlyDolk21_CBD in each food, a cell wall-binding assay was conducted. Using the same procedure as for the endolysin assay, prepared C. perfringens ATCC 13124 cells were added to sterilized milk and beef soup. To remove the unbound CBD protein and other components in food samples such as fat, the mixtures were washed twice in the beef soup sample and four times in the milk sample using sterilized PBS buffer. The binding ability was confirmed as described above.
3.12. Nucleotide Sequence Accession Number
The complete genome sequence of C. perfringens phage Dolk21 is available in the GenBank database under Accession No. OP730325.
4. Conclusions
In this study, the novel bacteriophage Dolk21 was isolated from a soil sample using C. perfringens ATCC 13124 as the host strain. The phage demonstrated highly restricted host specificity, which prompted further investigation into using its endolysin as an alternative. PlyDolk21, the endolysin encoded by the phage Dolk21, exhibited strong lytic activity against a broader range of C. perfringens strains compared to the phage itself, showing potential as an efficient biocontrol agent in food safety. Its stability under various environmental conditions such as pH, temperature, and NaCl concentration supports its use in foods. Furthermore, the rapid detection ability of its CBD provides an additional advantage for C. perfringens identification. PlyDolk21 shows significant promise as a natural and safe alternative for controlling C. perfringens contamination in food products. A future optimization study of PlyDlk21 could enhance its efficacy and expand its applications in the food industry.
References
- B. Canard, B. Saint-Joanis, S.T. Cole. Genomic diversity and organization of virulence genes in the pathogenic anaerobe Clostridium perfringens. Mol. Microbiol., 1992. [DOI | PubMed]
- J.I. Rood, S.T. Cole. Molecular genetics and pathogenesis of Clostridium perfringens. Microbiol. Rev., 1991. [DOI | PubMed]
- S. Brynestad, P.E. Granum. Clostridium perfringens and foodborne infections. Int. J. Food Microbiol., 2002. [DOI | PubMed]
- S. García, N. Heredia. Clostridium perfringens: A dynamic foodborne pathogen. Food Bioprocess Technol., 2011. [DOI]
- J.C. Freedman, A. Shrestha, B.A. McClane. Clostridium perfringens enterotoxin: Action, genetics, and translational applications. Toxins, 2016. [DOI | PubMed]
- R. Kiu, L.J. Hall. An update on the human and animal enteric pathogen Clostridium perfringens. Emerg. Microbes Infect., 2018. [DOI | PubMed]
- H. Suzuki, K. Hosomi, A. Nasu, M. Kondoh, J. Kunisawa. Development of adjuvant-free bivalent food poisoning vaccine by augmenting the antigenicity of Clostridium perfringens enterotoxin. Front. Immunol., 2018. [DOI | PubMed]
- X. Wang, C. Ning, C. Ji, Y. Li, J. Li, Q. Meng, J. Qiao, L. Wang, K. Cai, J. Zhang. Antimicrobial resistance profiling and molecular typing of ruminant-borne isolates of Clostridium perfringens from Xinjiang, China. J. Glob. Antimicrob. Resist., 2021. [PubMed]
- K. Ho. Bacteriophage therapy for bacterial infections: Rekindling a memory from the pre-antibiotics era. Perspect. Biol. Med., 2001. [DOI | PubMed]
- C. Lee, H. Kim, S. Ryu. Bacteriophage and endolysin engineering for biocontrol of food pathogens/pathogens in the food: Recent advances and future trends. CritiCal Rev. Food Sci. Nutr., 2023. [DOI | PubMed]
- H.-W. Hong, Y.D. Kim, J. Jang, M.S. Kim, M. Song, H. Myung. Combination effect of engineered endolysin EC340 with antibiotics. Front. Microbiol., 2022. [DOI | PubMed]
- Z. Drulis-Kawa, G. Majkowska-Skrobek, B. Maciejewska. Bacteriophages and phage-derived proteins–application approaches. Curr. Med. Chem., 2015. [DOI | PubMed]
- R. Nachimuthu, M.M. Royam, P. Manohar, S. Leptihn. Application of bacteriophages and endolysins in aquaculture as a biocontrol measure. Biol. Control, 2021. [DOI]
- N. Principi, E. Silvestri, S. Esposito. Advantages and limitations of bacteriophages for the treatment of bacterial infections. Front. Pharmacol., 2019. [DOI | PubMed]
- A. Colavecchio, B. Cadieux, A. Lo, L.D. Goodridge. Bacteriophages contribute to the spread of antibiotic resistance genes among foodborne pathogens of the Enterobacteriaceae family—A review. Front. Microbiol., 2017. [DOI | PubMed]
- M. Rahman, W. Wang, Q. Sun, J.A. Shah, C. Li, Y. Sun, Y. Li, B. Zhang, W. Chen, S. Wang. Endolysin, a promising solution against antimicrobial resistance. Antibiotics, 2021. [DOI | PubMed]
- F. Abdelrahman, M. Easwaran, O.I. Daramola, S. Ragab, S. Lynch, T.J. Oduselu, F.M. Khan, A. Ayobami, F. Adnan, E. Torrents. Phage-encoded endolysins. Antibiotics, 2021. [DOI | PubMed]
- M. Zimmer, N. Vukov, S. Scherer, M.J. Loessner. The murein hydrolase of the bacteriophage φ3626 dual lysis system is active against all tested Clostridium perfringens strains. Appl. Environ. Microbiol., 2002. [DOI | PubMed]
- J. Venhorst, J.M. van der Vossen, V. Agamennone. Battling enteropathogenic clostridia: Phage therapy for Clostridioides difficile and Clostridium perfringens. Front. Microbiol., 2022. [DOI | PubMed]
- 20.GRN000802EndolysinU.S. Food and Drug AdministrationSilver Spring, MD, USA2019
- V. Kazanavičiūtė, A. Misiūnas, Y. Gleba, A. Giritch, A. Ražanskienė. Plant-expressed bacteriophage lysins control pathogenic strains of Clostridium perfringens. Sci. Rep., 2018. [DOI | PubMed]
- F.M. Khan, J.-H. Chen, R. Zhang, B. Liu. A comprehensive review of the applications of bacteriophage-derived endolysins for foodborne bacterial pathogens and food safety: Recent advances, challenges, and future perspective. Front. Microbiol., 2023. [DOI | PubMed]
- M. Kong, S. Ryu. Bacteriophage PBC1 and its endolysin as an antimicrobial agent against Bacillus cereus. Appl. Environ. Microbiol., 2015. [DOI | PubMed]
- N.P. Antonova, D.V. Vasina, E.O. Rubalsky, M.V. Fursov, A.S. Savinova, I.V. Grigoriev, E.V. Usachev, N.V. Shevlyagina, V.G. Zhukhovitsky, V.U. Balabanyan. Modulation of endolysin LysECD7 bactericidal activity by different peptide tag fusion. Biomolecules, 2020. [DOI | PubMed]
- J. Shephard, A. McQuillan, P. Bremer. Mechanisms of cation exchange by Pseudomonas aeruginosa PAO1 and PAO1 wbpL, a strain with a truncated lipopolysaccharide. Appl. Environ. Microbiol., 2008. [DOI | PubMed]
- L. Liu, Y. Li, X. Wang, W. Guo. A phosphoethanolamine transferase specific for the 4′-phosphate residue of Cronobacter sakazakii lipid A. J. Appl. Microbiol., 2016. [DOI | PubMed]
- L.Y. Low, C. Yang, M. Perego, A. Osterman, R.C. Liddington. Structure and lytic activity of a Bacillus anthracis prophage endolysin. J. Biol. Chem., 2005. [DOI | PubMed]
- S.A. Ranveer, V. Dasriya, M.F. Ahmad, H.S. Dhillon, M. Samtiya, E. Shama, T. Anand, T. Dhewa, V. Chaudhary, P. Chaudhary. Positive and negative aspects of bacteriophages and their immense role in the food chain. npj Sci. Food, 2024. [DOI | PubMed]
- M. Modi, M. Thambiraja, A. Cherukat, R.M. Yennamalli, R. Priyadarshini. Structure predictions and functional insights into Amidase_3 domain containing N-acetylmuramyl-L-alanine amidases from Deinococcus indicus DR1. BMC Microbiol., 2024. [DOI | PubMed]
- R. Priyadarshini, M.A. de Pedro, K.D. Young. Role of peptidoglycan amidases in the development and morphology of the division septum in Escherichia coli. J. Bacteriol., 2007. [DOI | PubMed]
- M.M. Bendary, M.I. Abd El-Hamid, R.M. El-Tarabili, A.A. Hefny, R.M. Algendy, N.A. Elzohairy, M.M. Ghoneim, M.M. Al-Sanea, M.H. Nahari, W.H. Moustafa. Clostridium perfringens associated with foodborne infections of animal origins: Insights into prevalence, antimicrobial resistance, toxin genes profiles, and toxinotypes. Biology, 2022. [DOI | PubMed]
- R. Lu, B. Liu, L. Wu, H. Bao, P. García, Y. Wang, Y. Zhou, H. Zhang. A broad-spectrum phage endolysin (LysCP28) able to remove biofilms and inactivate Clostridium perfringens strains. Foods, 2023. [DOI | PubMed]
- X. Zhao, L. Li, Q. Zhang, M. Li, M. Hu, Y. Luo, X. Xu, Y. Chen, Y. Liu. Characterization of the Clostridium perfringens phage endolysin cpp-lys and its application on lettuce. Int. J. Food Microbiol., 2023. [DOI | PubMed]
- J.-H. Cho, J.-G. Kwon, D.J. O’Sullivan, S. Ryu, J.-H. Lee. Development of an endolysin enzyme and its cell wall–binding domain protein and their applications for biocontrol and rapid detection of Clostridium perfringens in food. Food Chem., 2021. [DOI | PubMed]
- J. Bai, Y.-T. Kim, S. Ryu, J.-H. Lee. Biocontrol and rapid detection of food-borne pathogens using bacteriophages and endolysins. Front. Microbiol., 2016. [DOI | PubMed]
- E. Ha, B. Son, S. Ryu. Clostridium perfringens virulent bacteriophage CPS2 and its thermostable endolysin LysCPS2. Viruses, 2018. [DOI | PubMed]
- E. Quevillon, V. Silventoinen, S. Pillai, N. Harte, N. Mulder, R. Apweiler, R. Lopez. InterProScan: Protein domains identifier. Nucleic Acids Res., 2005. [DOI | PubMed]
- B. Son, M. Kong, Y. Lee, S. Ryu. Development of a novel chimeric endolysin, Lys109 with enhanced lytic activity against Staphylococcus aureus. Front. Microbiol., 2021. [DOI | PubMed]
