The New SH3b_T Domain Increases the Structural and Functional Variability Among SH3b-Like CBDs from Staphylococcal Phage Endolysins
Abstract
Supplementary Information:
The online version contains supplementary material available at 10.1007/s12602-024-10309-0.
Article type: Research Article
Keywords: Bacteriophage, Endolysin, Cell wall binding, Binding specificity, SH3b
Affiliations: https://ror.org/00cv9y106grid.5342.00000 0001 2069 7798Department of Biotechnology, Ghent University, Ghent, Belgium; https://ror.org/0119pby33grid.512891.6Centro de Investigación Biomédica en Red de Enfermedades Respiratorias (CIBERES), Madrid, Spain; https://ror.org/00bnagp43grid.419120.f0000 0004 0388 6652Instituto de Productos Lácteos de Asturias (IPLA-CSIC), Villaviciosa, Asturias Spain; https://ror.org/05xzb7x97grid.511562.4DairySafe Group. Instituto de Investigación Sanitaria del Principado de Asturias (ISPA), Oviedo, Spain
License: © The Author(s) 2024 CC BY 4.0 Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
Article links: DOI: 10.1007/s12602-024-10309-0 | PubMed: 39080103 | PMC: PMC12634807
Relevance: Relevant: mentioned in keywords or abstract
Full text: PDF (619 KB)
Introduction
Endolysins are one of the gene products that dsDNA (bacterio)phages use to release the viral progeny from their host bacterial cells. They contain a catalytic, peptidoglycan-degrading activity, and thus when released to the periplasm via different tightly regulated mechanisms, they provoke bacterial cell lysis by osmotic shock due to the disruption of the peptidoglycan [ref. 1]. Besides their natural key role in the phage infection cycle, endolysins have sparked interest since they can be purposed as alternative antimicrobial agents [ref. 2–ref. 4]. Due to the escalating burden of antibiotic resistance among clinically relevant bacteria [ref. 5], the discovery and development of novel antimicrobials is one of the current main scientific priorities set up by many healthcare authorities and international organizations [ref. 6, ref. 7]. Recombinantly produced endolysins have been extensively shown to be effective as exogenous antibacterial agents in vitro and in vivo, and different clinical trials have been conducted [ref. 8–ref. 10]. Besides their lesser probability to cause resistance in bacteria, the most interesting features of lysins both from an applied and a fundamental perspective are probably their great natural variability and modularity [ref. 11, ref. 12]. Both characteristics are intertwined, as phages rapidly and dynamically evolve in a modular manner, exchanging functionally autonomous modules of genetic information between each other and with their bacterial hosts [ref. 13]. At the endolysin level, their modularity means that they may comprise different functional domains: one or more enzymatically active domains (EADs) and a cell wall-binding domain (CBD). Usually, phages that infect Gram-negative hosts bear lysins with a single EAD, whereas those from Gram-positive hosts typically have several domains, at least one of each kind [ref. 11, ref. 14]. The preferential presence of CBDs in endolysins from a Gram-positive background is hypothetically explained either by (i) the need for a tropism of the enzyme towards its insoluble, non-diffusible substrate (as is the case for many enzymes acting on polymeric substrates [ref. 15]); (ii) for the endolysin to remain tightly bound to the cell debris of the lysed host thus preventing the killing of neighboring cells that are potential new hosts for the phage progeny; or by a combination of both reasons [ref. 16]. Importantly, due to their typically high affinity, CBDs are thought to be the main determinant for the observed endolysin specificity, as proven, for example, by domain swapping experiments in endolysins derived from phages infecting Streptococcus or Listeria [ref. 17–ref. 19].
The case of endolysins from staphylococcal phages has been extensively studied due to the prominent role of many staphylococcal species in human or animal microbiota and disease [ref. 20]. For example, Staphylococcus aureus is one of the most burdensome human bacterial pathogens globally [ref. 21], and Staphylococcus epidermidis plus some other so-called coagulase-negative staphylococci are widespread components of the human skin microbiota that are also responsible for nosocomial infections [ref. 22]. Endolysins from staphylococcal phages have a typical bicatalytic structure [ref. 11], with evolutionarily conserved CBDs belonging to the bacterial SH3 (SH3b) superfamily [ref. 23]. SH3b (bacterial Src Homology 3) is a superfamily of widespread ligand-binding domains that appear in many bacterial and phage proteins and are also related to homologous ligand-binding domains in other kingdoms [ref. 24]. The SH3 fold is one of the simplest and oldest ones [ref. 25] and its main structural feature is a β-barrel layout usually devoted to a ligand-binding function. The SH3b domains in particular are mainly known to bind cell wall motifs, thus playing a prominent role in cell wall-remodeling enzymes, autolysins and phage endolysins. In the particular case of staphylococcal endolysins, the studied CBDs from SH3b have been classified into the SH3_5 (PF08460) family, and are assumed to specifically bind the peptide moiety of staphylococcal peptidoglycan, including the peptide stem and the peptide cross-link, as recently shown for the SH3_5 CBD of lysostaphin [ref. 26, ref. 27]. However, SH3b domains comprise representatives that, while sharing the characteristic SH3-like β-barrel topology, have evolved to recognize a variety of cell wall ligands. For example, the SH3_5 from Lactiplantibacillus plantarum major autolysin Acm2 is a broad-range CBD that recognizes many different peptidoglycan chemotypes [ref. 28], the SH3b-like, PSA_CBD (PF18341) domain from Listeria phage endolysins recognizes serovar-specific motifs at the cell wall teichoic acids [ref. 29], and the SH3b CBD from the endolysin of Bacillus phage PBC5 binds to the glycan chain [ref. 30].
In this work, we aimed at providing insights on the specificity range of SH3b-like CBDs from staphylococcal endolysins and how they impact the antibacterial spectrum of the lysins in which they are inserted. To this end, we focused on three Staphylococcus phage endolysins: LysRODI, LysC1C and LysIPLA5 [ref. 31, ref. 32]. The binding profiles of the selected CBDs were experimentally characterized both as a standalone and in connection with their ability to modulate the activity range of EADs derived from LysRODI and LysC1C. In this way, we expect this work contributes to understand how the structural diversity of staphylococcal CBDs connects to their peptidoglycan-binding function, and how this ability cooperates with intrinsic features of EADs to produce the experimentally observed activity spectra in lysins purposed for exogenous lysis.
Materials and Methods
Bacterial Strains and Culture Conditions
The staphylococcal strains used in this work (Table 1) were grown in tryptic soy broth (TSB) at 37 °C with shaking (200 rpm) or on TSB plates containing 2% (w/v) bacteriological agar. Escherichia coli TOP10 was used for cloning and E. coli BL21(DE3) for protein expression. Acinetobacter baumannii RUH 134 [ref. 33] was used as a control strain. All the former Gram-negative bacteria were grown in LB medium at 37 °C with shaking (200 rpm). For the positive selection of pVTEIII or pVTD3 E. coli transformants, 100 μg/ml ampicillin or 50 μg/ml kanamycin were used, respectively, together with 5% (w/v) sucrose to negatively select against plasmids lacking insertion (as explained in [ref. 34]). Then, 100 μg/ml ampicillin was used to select transformants of vectors based on pET21(a). Bacterial stocks were made by adding 20% v/v glycerol to grown bacterial cultures and were kept at – 80 °C.
Table 1: Staphylococcal strains used in this work
| Species | Strain | Source |
|---|---|---|
| S. epidermidis | F12 | [ref. 35] |
| B | ||
| DG2n | ||
| YLIC13 | ||
| LO5RB1 | ||
| DH3LIk | ||
| Z2LDC14 | ||
| S. aureus | Sa9 | [ref. 36] |
| IPLA1 | [ref. 37] | |
| IPLA16 | ||
| 15981 | [ref. 38] | |
| V329 | [ref. 39] | |
| MRSAE10 | Pig skin isolate (unpublished) | |
| S. hominis | ZL31-13 | [ref. 40] |
| S. xylosus | ZL61-2 | |
| S. haemolyticus | ZL89-3 | |
| S. gallinarum | ZL90-5 | |
| S. kloosi | ZL74-2 |
Plasmid Construction and DNA Manipulation
The sequences encoding LysRODI, LysC1C and LysIPLA5 were codon optimized (GenSmart Codon optimization), synthetized and cloned into a pET21(a) vector (between NdeI and XhoI restriction sites) by GenScript (Rijswijk, Netherlands). For all other proteins used in this work, the expression vectors were constructed through the VersaTile workflow as described in [ref. 34]. In brief, each individual domain (EAD, CBD or eGFP) was PCR-amplified from its source plasmid with specific primers including BpiI and BsaI recognition sites at both the 5′ and 3′ end, according to the VersaTile method. A restriction/ligation reaction with BpiI was carried out with these amplicons to insert them into the entry vector pVTEIII (AmpR, SucS). The ligation products were subsequently used for transformation of E. coli TOP10 by electroporation and transformants bearing pVTEIII plasmids with the inserted tile (AmpR, SucR) were selected on LB plates with ampicillin and sucrose. The TOP10 cells were used as a source for tiles, which were all confirmed by Sanger sequencing (LGC Genomics) and stored at the VersaTile repository of Ghent University. Tile ligation into the destination vector pVTD3 (KanR, SucS) was conducted by setting up restriction/ligation reactions with BsaI and the appropriate tiles from the repository (e.g. eGFP plus RODI_CBD and a 6×His tag). All chimeric coding sequences were designed with a C-terminal 6×His tag for purification unless otherwise stated. Final constructs were used for transformation of E. coli BL21(DE3), selecting the transformants with kanamycin and sucrose, and their sequence was verified by Sanger sequencing. A list of the tiles used in this work, including their source NCBI entry and the delineation coordinates, can be found in Table 2
Table 2: Tiles used in this work
| Tile name | Source | Delineation (start:end in full protein sequence) |
|---|---|---|
| IPLA5_CHAP | AFM73732.1 (LysIPLA5) | 232:379 |
| IPLA5_Ami2 | 2:231 | |
| IPLA5_CBD | 380:574 | |
| RODI_CHAP | YP_009195893.1 (LysRODI) | 2:188 |
| RODI_CBD | 406:496 | |
| C1C_CHAP | YP_009214649.1 (LysC1C) | 2:167 |
| C1C_CBD | 381:484 | |
| eGFP | AFA52650.1 | 3:238 |
Protein Expression and Purification
The fusion proteins used throughout this work were expressed in E. coli BL21(DE3) strains bearing the corresponding pVTD3 or pET21(a) vectors prepared as described in the previous section. Protein expression and purification was performed as previously described [ref. 41]. After purification, the buffer was exchanged to 50 mM sodium phosphate buffer pH 7.4 using Zeba™ Spin Desalting Columns, 7K MWCO, 5 ml (Thermo Fisher Scientific) following the supplier’s recommendations. Finally, proteins were sterilized by filtration (0.45 μm PES membrane filters, VWR).
Protein concentration was quantified using the Quick Start Bradford Protein assay (BioRad). Relevant information on the proteins used in this work is in Table 3.
Table 3: Predicted features (molecular weight, isoelectric point, extinction coefficient) of the proteins used in this work using Expasy ProtParam (https://web.expasy.org/protparam/) along with their experimentally verified purification yield
| Protein | MW (kDa) | pI | ε (M−1 cm−1) | Purification yield (mg/L) |
|---|---|---|---|---|
| LysIPLA5 | 66.940 | 9.89 | 156455 | 0.2 |
| IPLA5_Ami2 | 27.078 | 9.54 | 48485 | 0.1 |
| IPLA5_CHAP | 18.175 | 8.92 | 40005 | 0.2 |
| IPLA5_Ami2-CBD | 50.140 | 10.11 | 116450 | 0.1 |
| IPLA5_CHAP-CBD | 41.242 | 10.80 | 107970 | 0.1 |
| RODI_CHAP | 22.047 | 10.09 | 45380 | 1.1 |
| RODI_CHAP-CBD | 31.950 | 9.87 | 69580 | 3.1 |
| RODI_CHAP-IPLA5_CBD | 45.100 | 10.33 | 113345 | 0.4 |
| C1C_CHAP | 20.205 | 9.99 | 43890 | 0.7 |
| C1C_CHAP-C1C_CBD | 32.100 | 9.69 | 88475 | 0.2 |
| C1C_CHAP-IPLA5_CBD | 43.263 | 10.30 | 111855 | 0.2 |
| eGFP-RODI_CBD | 37.890 | 6.84 | 46090 | 1.5 |
| eGFP-C1C_CBD | 39.960 | 6.45 | 39967 | 0.3 |
| eGFP-IPLA5_CBD | 51.052 | 9.70 | 89980 | 2.1 |
Quantification of Bacterial Binding
Binding of CBDs to bacterial substrates was measured by recording the fluorescence of eGFP fusions of the different domains. Such fusions were prepared using VersaTile and comprised eGFP at the N-terminal, the CBD at the central and a 6×His tag at C-terminal position. To perform the binding assay, exponential (OD600 ≈ 0.5–0.6) or stationary phase (OD600 ≈ 1–2) cultures of the strains to be tested were centrifuged (10,000×g, 1 min) and the pellets were washed with PBS (137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4, pH 7.4). The bacterial suspensions were adjusted to OD600 ≈ 1.0 and dispensed on dark flat bottom 96-well plates (180 µl per well). Further, 20 µl of 20 µM solutions of the eGFP-CBD fusion proteins (or just buffer for bacterial autofluorescence controls) were added to each well and then the plates were incubated for 10 min in the dark at room temperature. Then the plates were centrifuged (1000×g, 5 min), the supernatants were removed, and the pellets washed once with PBS and finally suspended in 200 µl of PBS. Then, 200 µl of 10 µM fluorescein were added to the plate as internal control to automatically optimize gain, as well as positive fluorescence controls for each eGFP-CBD fusion protein (180 µl PBS plus 20 µl of the 20 µM protein stock solution). Fluorescence was then measured in a TECAN Infinite 200 PRO plate reader (TECAN, Männedorf, Switzerland) with excitation/emission wavelengths of 485 nm and 530 nm, respectively. Fluorescence measurements were then corrected for comparability between proteins by applying a correction factor Fmax/Fprot in which Fmax is the maximum fluorescence recorded and Fprot is the fluorescence of each eGFP-CBD at 2 µM. Fluorescence measurements were acquired for three biological replicates.
Minimum Inhibitory Concentration
The minimum inhibitory concentrations (MICs) of the antimicrobial proteins in this work were determined by the broth microdilution assay as described before [ref. 41]. The MIC values reported correspond to the mode of three independent biological replicates.
Bioinformatic Analyses
Two complementary approaches were taken to build a protein sequence dataset of SH3b-like domains related to staphylococcal lysins (Supplementary Fig. S1, Online Resource 1). To find representative sequences of the IPLA5_CBD family, termed SH3b_T (PF24246), a phmmer search [ref. 42] was conducted against Reference Proteomes restricted to viral taxa (taxid: 10239) and using the first IPLA5_CBD as query (UniProt I6T7G5, from coordinate 380 to 458). The 38 significant hits (i.e. sequences comprising only the SH3b_T-like domains) were clustered with CD-HIT and an exclusion cutoff of 97% identity was applied to decrease redundancy [ref. 43]. A second phmmer iteration was conducted, now against the full UniProt database, using two queries: the first repeat of the LysIPLA5 CBD and the SH3b_T domain from A0A499SIE6 (positions 169 to 250), displaying the lowest % identity, in amino acid sequence, with the former (39%). This yielded 366 significant hits. A length cutoff was applied to the sequences (source full proteins > 100 amino acids; query coverage > 60 amino acids) and a second CD-HIT redundancy reduction with 97% identity cutoff was applied, reducing the final set to 59 representatives for which additional metadata (domain predictions for the full-length sequences, bacterial hosts) were mined from UniProt or predicted using hmmscan against the Pfam database. Alternatively, representative domain sequences from the SH3_5 family were retrieved from PhaLP database of phage lysins [ref. 12], restricting the search to endolysins from host genus Staphylococcus and domain name SH3_5 (236 entries total). The CBD sequences were extracted from these entries using the delineating coordinates from the Pfam SH3_5 predictions stored in PhaLP, and then the same length and % identity cutoffs were imposed to obtain 53 final representative SH3_5 domain sequences. The same PhaLP-based pipeline was used to retrieve PSA_CBD examples (type: ‘endolysin’, domain name: ‘PSA_CBD’), while the phmmer pipeline was used to retrieve PBC5_CBD examples. The first SH3b-like repeat of LysPBC5 (A0A218KCJ1, residues 230 to 279) was used as query for the phmmer searches. The dataset built this way can be accessed as Online Resource 2.
Multiple sequence alignments (MSAs) and phylogenetic analyses were performed and represented in R. The packages ‘msa’ and ‘ggmsa’ were used respectively to build the MSAs (with algorithm ClustalW) and to represent them [ref. 44, ref. 45]. The phylogenetic trees were built based on the MSAs for which a sequence similarity-based pairwise distance matrix was computed, which was then used as input for building an UPGMA phylogenetic tree and representing it [ref. 46–ref. 48].
Protein three-dimensional structure predictions were performed using AlphaFold2 algorithm as implemented in ColabFold [ref. 49] with default parameters plus Amber relaxation, and models were analyzed and rendered with DeepView [ref. 50], which was also used for structural alignments.
Statistical Analyses and Experimental Data Representation
Representation of experimental data was performed using R package ‘ggplot2’ [ref. 51]. Error bars in bar plots represent the standard deviation of three independent replicates. Additional statistical analyses (Dunn’s test) were performed in R.
Results
LysIPLA5 has an Atypical Bicatalytic Architecture with an SH3b-like C-Terminal End
The endolysins from phages phiIPLA-RODI and phiIPLA-C1C (i.e. LysRODI and LysC1C) are examples of the canonical CHAP (PF05257):Amidase_2 (PF01510):SH3_5 architecture described for a majority of staphylococcal endolysins. The S. epidermidis phage vB_SepiS-phiIPLA5, however, bears an endolysin (LysIPLA5) with two predicted EADs, Amidase_2 and CHAP, in the reverse order. In addition, LysIPLA5 has a C-terminus where usually a CBD would be present, but no CBD could be predicted (Fig. 1a). In a preliminary functional study of LysIPLA5 domains, neither the full protein nor any of its individual domains showed in vitro antimicrobial activity against S. epidermidis F12, probably due to very low expression yields and subsequently low working concentrations, at least for the full protein (Supplementary Fig. S2, Online Resource 1). The C-terminal domain, henceforth referred to as IPLA5_CBD, was nevertheless efficiently expressed and a concentration as high as 34.16 µM was unable to exert any growth inhibition. Nevertheless, whereas ~ 6 µM IPLA5_CHAP was also inactive, it was possible to observe a MIC (0.75 µM) for the fusion protein comprising the IPLA5_CHAP EAD fused to the native C-terminal domain IPLA5_CBD. These results initially supported the possibility of IPLA5_CBD being a CBD.
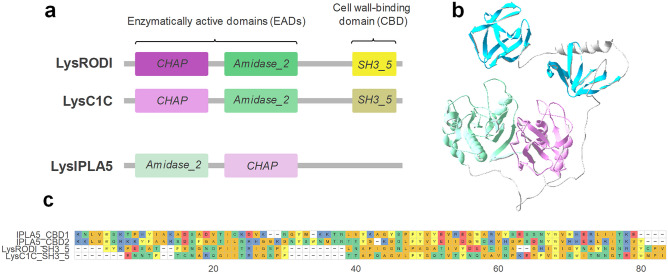
According to a three-dimensional structure prediction of LysIPLA5, this C-terminal stretch contains two repeats with a fold akin to the typical SH3 β-barrels (Fig. 1b). The sequences of these repeats substantially differ from the SH3_5 CBDs of LysRODI and LysC1C (percent identities between 16 and 21%), while the repeats are 44% identical to each other (Fig. 1c). Thus, it was concluded that LysIPLA5 bears a C-terminal CBD that belongs to the SH3b superfamily but that constitutes a different family than the usual SH3_5, which was then termed SH3b_T (present in the Pfam database as PF24246).
CBDs from SH3b_T Family Show Differences in Their Bacterial Distribution with Respect to the SH3_5 ones
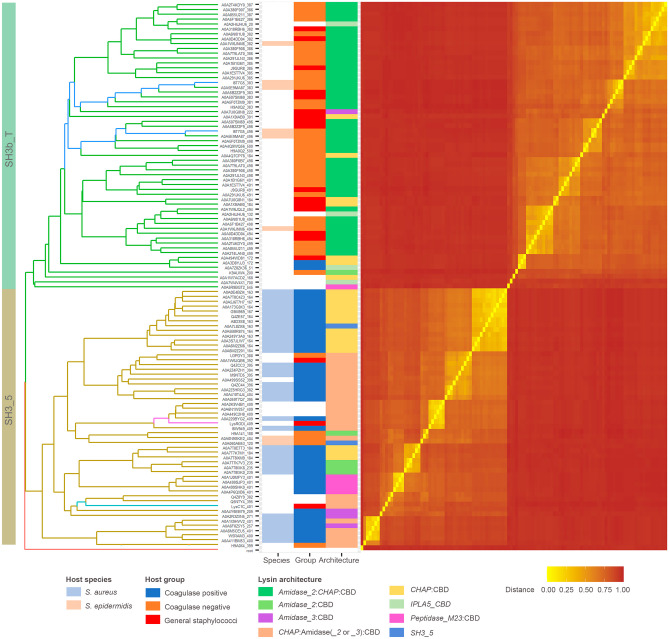
The tree shown in Fig. 2 indicates that the 59 SH3b_T-like sequences retrieved by an iterative phmmer search substantially differ in sequence from the 53 established SH3_5 examples obtained from PhaLP, as they are allocated in different clades and are clearly distinguished in the similarity-based distance matrix heatmap. They also display a marked difference in the bacterial species to which they are associated. In the case of SH3_5, they mostly appear in phages whose host is annotated as S. aureus, whereas the preferred host for IPLA5_CBD is S. epidermidis or other coagulase-negative staphylococci, with only a few exceptions to this rule. In addition, a trend can also be established with respect to the preferred full endolysin architecture: while the preferential architecture for endolysins with an SH3_5 CBD is the canonical CHAP:Amidase_2:CBD, the order of the EADs is mostly reversed for lysins with a IPLA5_CBD, as in LysIPLA5 itself.
RODI_CBD, C1C_CBD and IPLA5_CBD Present Different Binding Profiles
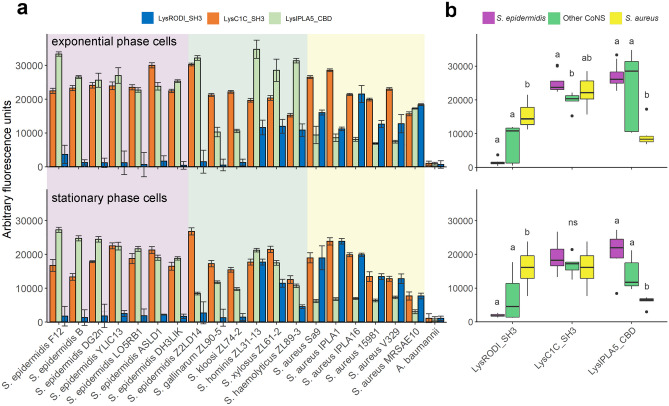
To confirm the nature of IPLA5_CBD as a true CBD and explore the specificity differences to which the differential taxonomical distributions in Fig. 2 point, eGFP fusions of IPLA5_CBD, RODI_CBD and C1C_CBD were obtained. These eGFP-tagged domains were then assayed for binding capacity against a set of staphylococcal strains (Fig. 3a). Well-marked trends could be observed in the binding specificity of the three different domains. RODI_CBD bound almost exclusively to S. aureus and a few other coagulase-negative staphylococci (CoNS), C1C_CBD was able to bind generally to all the tested staphylococci, and IPLA5_CBD bound preferentially to S. epidermidis plus other CoNS, although not in such a specific manner as RODI_CBD. A summary and the statistical significance supporting these trends are available in Fig. 3b. Although generalizations should be made with care, these results provide an experimental explanation to the preferred association of SH3b_T family to S. epidermidis and CoNS, while SH3_5 is more commonly associated to S. aureus and only in some cases to CoNS or staphylococci in general (and LysC1C is an example of the latter). An additional conclusion of the results in Fig. 3 is that binding (i.e. the magnitude of the recorded fluorescence value) seems to be generally lower when cells in stationary phase are used versus using exponential phase ones. An explanation for this may be provided by the nature of the ligand that has been described before for SH3b domains in anti-staphylococcal lysins, which has been usually identified as the peptidoglycan peptide moiety [ref. 26, ref. 27, ref. 52, ref. 53]. Stationary phase S. aureus cells are known to have fewer cross-links in the peptidoglycan [ref. 54], which would then mean a lower number of potential binding ligands for SH3b-like CBDs, thus explaining the results in Fig. 3.
The Specificity Profile of SH3b-Like CBDs Modulates the Antibacterial Spectrum of Accompanying EADs
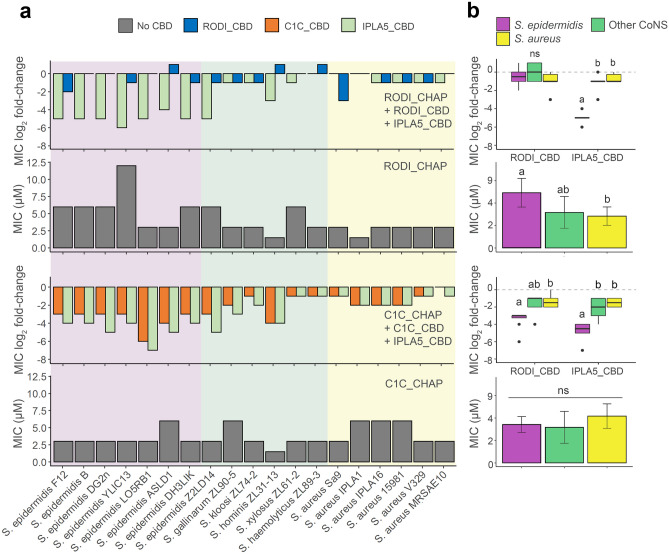
To better understand the contribution of the sole-CBD specificity profile to the activity spectrum of full lysins, fusions of the CHAP EADs of LysRODI and LysC1C with either their wild-type CBD or IPLA5_CBD were obtained. Then, the MIC for each of the constructs, including the EADs only, was calculated against the full set of staphylococcal strains (Fig. 4). A general conclusion from this experiment is that fusing a CBD normally improves the antimicrobial activity since all mean MIC log2 fold-change values in Fig. 4b are negative, reflecting a decrease in the MIC as a result of fusing CBDs to the EADs. This is in accordance with the common notion that CBDs are necessary for the efficient action of endolysins. There are, however, a few cases in which the EAD-CBD fusion underperforms when compared with the sole EAD, namely RODI_CHAP is less active against some CoNS and a S. epidermidis strain when fused to RODI_CBD. In fact, the MIC improvement is not too impressive when fusing RODI_CBD to RODI_CHAP against any strain (with mean log2 fold-change values of about −1, in contrast with the 5 log2 fold decrease achieved by the IPLA5_CBD fusion against S. epidermidis, for example). This may be explained by the fact that RODI_CHAP already seems a highly optimized EAD against S. aureus, with clearly lower MIC values when compared with S. epidermidis or even the other coagulase negative staphylococci. The RODI_CHAP-IPLA5_CBD fusion does exhibit a remarkably lower MIC against S. epidermidis strains, but not against S. aureus, a behavior correlating to the specificity spectrum shown by IPLA5_CBD in Fig. 3. In contrast with the S. aureus-specialized RODI_CHAP, C1C_CHAP is a broad-range EAD, as much as its native CBD is also broad range, although with a slight preference towards S. epidermidis (however non-significant). This intrinsic optimization towards S. epidermidis is more apparent when the effect of fusing C1C_CHAP to the different CBDs is considered: while, as expected, adding IPLA5_CBD improves the performance against S. epidermidis, the fusion with the broad-range C1C_CBD does not decrease the MIC value equally against S. aureus and S. epidermidis; in fact, it shows a similar effect to the IPLA5_CBD fusion (Fig. 4b). This confirms a concomitant specialization of C1C_CHAP towards S. epidermidis peptidoglycan rather than that of S. aureus.
The Difference in Specificity Between RODI_CBD and C1C_CBD Could be Explained by Variability in Key Residues
The structures of the CBDs under investigation in this work were analyzed to find determinants for the perceived functional differences shown in Figs. 3 and 4. To this end, MSAs were obtained using the representative sequence sets from Fig. 2. A direct MSA-based comparison between the SH3_5 and the SH3b_T domains was not possible due to their low reciprocal identity (Fig. 1c); thus, the analysis was split to focus first on SH3_5 domains (Supplementary Fig. S3, Online Resource 1) and, therefore, on the structural differences that explain the different specificities of RODI_CBD and C1C_CBD. A set of key residues for peptidoglycan binding in SH3_5 domains was determined from the thorough, previously published works on the SH3_5 CBD of lysostaphin [ref. 26, ref. 27]. Then, the corresponding residues in RODI_CBD and C1C_CBD (Table 4) were identified assisted by MSA in Supplementary Fig. S3 and the comparison of the predicted 3D models of RODI_CBD and C1C_CBD with the experimentally determined structure of lysostaphin CBD (Fig. 5a).
Table 4: Key residues in SH3_5 domains associated with binding the peptide stem or the cross-bridge according to [ref. 26, ref. 27]. Cells are colored to highlight different types of amino acids (green = polar non-charged, orange = aliphatic, yellow = aromatic, red = negatively charged, blue = positively charged). Residues are numbered according to the full sequence of the source proteins (lysostaphin = UniProt P10547)
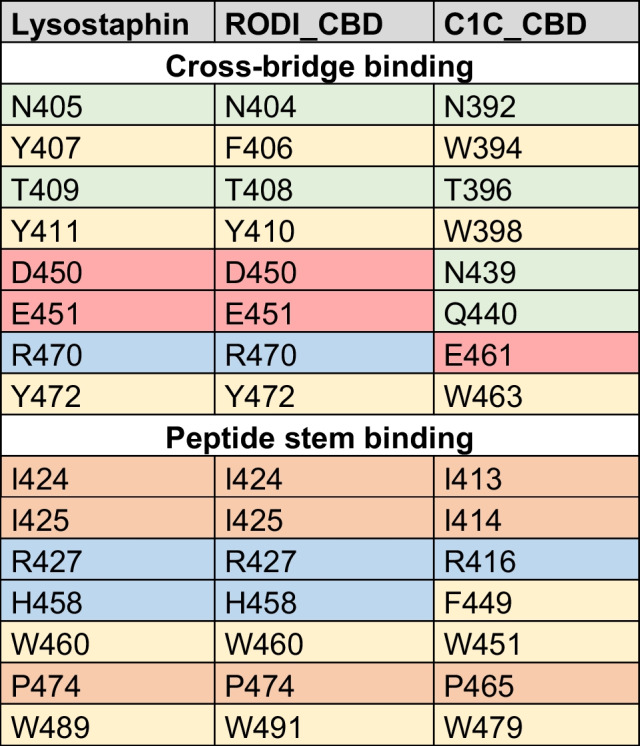
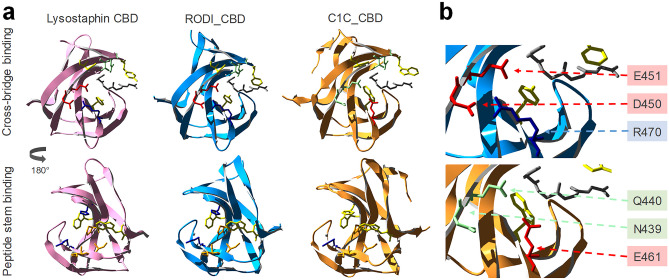
Table 4 shows little or no change in the chemical nature of the residues known to bind the peptide stem of the peptidoglycan across the three CBDs. Conversely, C1C_CBD displays radical shifts in amino acid properties at positions 439, 440 and 461, within the peptidoglycan cross-bridge binding site, with respect to lysostaphin and RODI_CBD. Whereas C1C_CBD contains two polar, non-charged amino acids (N439, Q440) and a negatively charged one (E461), RODI_CBD and lysostaphin bear two negatively charged residues (D450, E451) and a positively charged one (R470) at the corresponding sites. Zooming into the analogous structures of these CBDs, it can be concluded that these changes imply a substantial rearrangement of the chemical environment at the groove that is assumed to bind the peptidoglycan cross-bridge. Particularly, D450 and R470 in RODI_CBD are at a distance between 2.98 and 4.13 Å (depending on the atoms considered), which makes it possible for them to form a salt bridge [ref. 55]. This possibility is disrupted in C1C_CBD by the presence of N439 and E461 (Fig. 5b).
The facts that (i) the key residues in RODI_CBD and lysostaphin CBD are relatively unchanged but (ii) there are obvious differences in C1C_CBD only at the cross-bridge-binding region, are interpretable in the light of the experimental results presented in Figs. 3 and 4. While RODI_CBD binds S. aureus specifically, as lysostaphin CBD does, C1C_CBD seems to have no clear preference between binding S. aureus or S. epidermidis. Assuming that the three SH3_5 CBDs compared bind to the same ligand, the peptide moiety of peptidoglycan, which seems plausible given their sequence similarity (Supplementary Fig. S2, Online Resource 1) and the conservation of the residues putatively devoted to binding the peptide stem (Table 4), then the differences found at the cross-bridge binding pocket of C1C_CBD must explain its promiscuous binding profile. In fact, the major difference between the peptidoglycans of S. aureus and S. epidermidis is the structure of their cross-bridges. While the cross-bridge of S. aureus is the well-known pentaglycine bridge, the cross-bridging peptide in S. epidermidis is either GGSGG or AGGGG [ref. 56]. Since the introduction of a central serine residue in the cross-bridge is a known resistance mechanism to lysostaphin binding [ref. 57], it is reasonable that the RODI_CBD, structurally equivalent to lysostaphin CBD, is unable to bind the serine-containing S. epidermidis peptidoglycan cross-bridge. The specificity mechanism in lysostaphin CBD is thought to be one of steric constraint (the cross-bridge binding pocket can only accommodate a pentaglycine peptide [ref. 26]). Therefore, in C1C_CBD, the variants N394, Q440 and E461 should provide a greater flexibility for ligand placing at the cross-bridge binding site. This increased flexibility may be achieved by the disruption of the D450-R470 salt bridge in C1C_CBD (Fig. 5b), although this remains to be experimentally proven.
The IPLA5_CBD Fold is Closer to PSA_CBD or PCD5_CBD and May Bind a Different Ligand than SH3_5 CBDs
Given the low sequence similarity between the SH3_5 and the SH3b_T sequences, a structure-based comparison was attempted using a few bona fide experimentally determined examples from the wider SH3b CBD superfamily (i.e. lysostaphin CBD, PSA_CBD and PCD5_CBD), which were aligned to the predicted structure of a single IPLA5_CBD repeat (Fig. 6a). This comparison initially suggested that IPLA5_CBD repeats were predicted in an SH3b fold more similar to that of the PSA_CBD family (PF18341) or the still poorly described PCD5_CBD family. When sets of representative sequences from PSA_CBD and PCD5_CBD families were added to a phylogenetic tree together with SH3b_T and SH3_5, this was made apparent by the clustering of the former three apart from the latter (Fig. 6b).
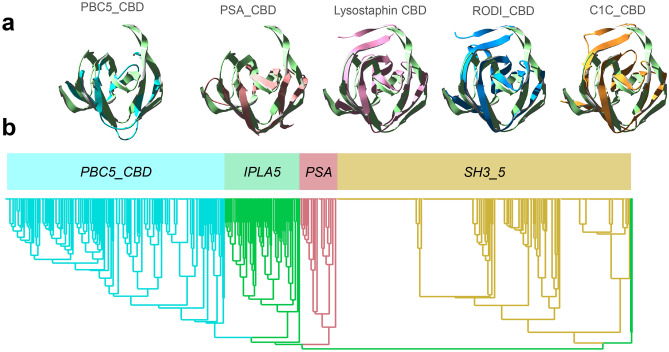
A preliminary conclusion from this observation is that the ligand for SH3b_T family may be different from that of SH3_5 (i.e. not the peptidoglycan cross-bridge or peptide moiety) given that PSA_CBD domains are known to bind sugar moieties in the teichoic acids of Listeria cells [ref. 29], while the ligand for PCD5_CBD has been described as the peptidoglycan glycan strands [ref. 30]. Since SH3b_T family is closer to PSA_CBD and PCD5_CBD than to SH3_5 (Fig. 6b), it may bind a glycan moiety (like PSA_CBD and PCD5_CBD) rather that a peptidic one (like SH3_5). Given that the composition of the cell wall teichoic acids are also a differential feature between S. aureus and S. epidermidis [ref. 58], these might be the ligand of IPLA5_CBD, explaining its preference for binding S. epidermidis in detriment of S. aureus.
Discussion
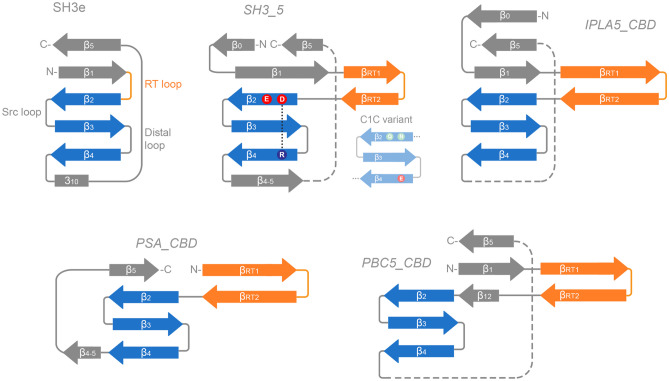
In this work, we have identified a new family of SH3b-like domains that binds elements of the staphylococcal cell wall, with a preference towards S. epidermidis, and we have set a context for part of the underexplored diversity of the versatile SH3b-like folds among endolysins. Here, we have compared four SH3b families present in phage endolysins: the SH3_5 family commonly found in staphylococcal lysins, the newly described SH3b_T present in LysIPLA5, the listerial PSA_CBD and the PBC5_CBD domains found in Bacillus and their phages. All of them share the common β-barrel structure, typical of SH3 folds (Fig. 6a), although with differing topologies (Fig. 7). These differences may correlate with the type of cell wall ligands they recognize, namely (i) the peptidoglycan peptide moiety (or more specifically the cross-bridge) for SH3_5 CBDs or (ii) different glycan moieties for PSA_CBD, PBC5_CBD and, perhaps, SH3b_T, which remains to be validated experimentally. Regarding their topology, all of them conserve the general structure of the SH3 fold, and share the common SH3b trait of an extended RT β-hairpin (equivalent of the RT loop in SH3e, the topology of eukaryotic SH3 domains) plus the conserved central antiparallel β-sheets 2 to 4 (‘β-core’, Fig. 7). Their main topological differences are located (i) at the N- and C-proximal β-sheets, such as the presence/absence of an additional N-terminal β-sheet (β0, present in SH3_5 and SH3b_T) or an additional β-sheet connecting β4 and β5 (β4-5, in SH3_5 and PSA_CBD); and also (ii) at the RT β-hairpin, which is clearly more elongated in PSA_CBD, PBC5_CBD and SH3b_T, with the latter having the longest version. These differences should somehow account for the different ligands of the families, although this work does not provide concrete insights about the possible ligand.
On a different note, the results hereby presented show the versatility of the SH3b domains for evolving slightly different structures that bind different ligands, which is particularly evident in the comparison between RODI_CBD and C1C_CBD. While both belong to the same SH3_5 family, their divergence only in a small set of key residues sets them rather radically apart in their specificity, determined by the nature of the peptidoglycan cross-bridge on the ligand side. This suggests that not only the SH3 fold can evolve towards topologically diverse families, each of them recognizing a different kind of ligand, but also, within each family, SH3b domains can fine-tune the residues at the binding pockets to select the specific ligands they bind. Thus, the wide-spread presence of SH3b-like domains in endolysins from phages that infect very diverse groups of bacteria can be explained on the basis of this versatility of SH3 folds as ‘raw material’ to evolve CBDs targeted at very specific ligands. However, the binding specificity dictated by CBDs, as we have shown, does not fully determine the activity spectrum of the derived lysins in which they are allocated (Fig. 4). For example, the increased activity against S. epidermidis provided by IPLA5_CBD was more prominent when fused to an EAD with poor activity against this bacterium (RODI_CHAP) than when accompanying an EAD already optimized against S. epidermidis (C1C_CHAP). Thus, our set of results imply that, while the final antibacterial outcome correlates in terms of specificity to the observed specificity profile of the CBDs, it is also true that such outcome is influenced by the intrinsic activity range displayed by the EAD. Therefore, while the CBD can be a specificity determinant, it must be considered that it does not fully determine endolysin specificity, at least in the case of anti-staphylococcal lysins, and this may be of greater importance when engineering new tailor-made lysins.
Supplementary Materials
References
- R Young. Phage lysis: three steps, three choices, one outcome. J Microbiol, 2014. [DOI | PubMed]
- R Vázquez, E García, P García. Phage lysins for fighting bacterial respiratory infections: a new generation of antimicrobials. Front Immunol, 2018. [DOI | PubMed]
- D Dams, Y Briers. Enzybiotics: enzyme-based antibacterials as therapeutics. Adv Exp Med Biol, 2019. [DOI | PubMed]
- SB Linden, AB Alreja, DC Nelson. Application of bacteriophage-derived endolysins to combat streptococcal disease: current state and perspectives. Curr Opin Biotechnol, 2021. [DOI | PubMed]
- CJL Murray, KS Ikuta, F Sharara. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. The Lancet, 2022. [DOI]
- 6.O’Neill J (2016) Tackling drug-resistant infections globally: final report and recommendations. The Review on Antimicrobial Resistance, United Kingdom
- E Tacconelli, E Carrara, A Savoldi. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect Dis, 2018. [DOI | PubMed]
- K Abdelkader, H Gerstmans, A Saafan. The preclinical and clinical progress of bacteriophages and their lytic enzymes: the parts are easier than the whole. Viruses, 2019. [DOI | PubMed]
- VG Fowler, AF Das, J Lipka-Diamond. Exebacase for patients with Staphylococcus aureus bloodstream infection and endocarditis. J Clin Invest, 2020. [DOI | PubMed]
- 10.Fowler VG Jr, Das AF, Lipka-Diamond J et al (2024) Exebacase in addition to standard-of-care antibiotics for staphylococcus aureus bloodstream infections and right-sided infective endocarditis: a phase 3, superiority-design, placebo-controlled, randomized clinical trial (DISRUPT). Clin Infect Dis ciae043. 10.1093/cid/ciae043
- R Vázquez, E García, P García. Sequence-function relationships in phage-encoded bacterial cell wall lytic enzymes and their implications for phage-derived product design. J Virol, 2021. [DOI | PubMed]
- B Criel, S Taelman, W Van Criekinge. PhaLP: a database for the study of phage lytic proteins and their evolution. Viruses, 2021. [DOI | PubMed]
- BJ Smug, K Szczepaniak, EPC Rocha. Ongoing shuffling of protein fragments diversifies core viral functions linked to interactions with bacterial hosts. Nat Commun, 2023. [DOI | PubMed]
- H Gerstmans, B Criel, Y Briers. Synthetic biology of modular endolysins. Biotechnol Adv, 2018. [DOI | PubMed]
- D Guillen, S Sanchez, R Rodriguez-Sanoja. Carbohydrate-binding domains: multiplicity of biological roles. Appl Microbiol Biotechnol, 2010. [DOI | PubMed]
- MJ Loessner, K Kramer, F Ebel, S Scherer. C-terminal domains of Listeria monocytogenes bacteriophage murein hydrolases determine specific recognition and high-affinity binding to bacterial cell wall carbohydrates. Mol Microbiol, 2002. [DOI | PubMed]
- R Diez-Martinez, HD De Paz, E Garcia-Fernandez. A novel chimeric phage lysin with high in vitro and in vivo bactericidal activity against Streptococcus pneumoniae. J Antimicrob Chemother, 2015. [DOI | PubMed]
- M Schmelcher, VS Tchang, MJ Loessner. Domain shuffling and module engineering of Listeria phage endolysinsfor enhanced lytic activity and binding affinity. Microb Biotechnol, 2011. [DOI | PubMed]
- R Vázquez, M Domenech, M Iglesias-Bexiga. Csl2, a novel chimeric bacteriophage lysin to fight infectionscaused by Streptococcus suis, an emerging zoonotic pathogen. Sci Rep, 2017. [DOI | PubMed]
- D Gutierrez, L Fernandez, A Rodriguez, P Garcia. Are phage lytic proteins the secret weapon to kill Staphylococcus aureus?. mBio, 2018. [DOI | PubMed]
- KS Ikuta, LR Swetschinski, GR Aguilar. Global mortality associated with 33 bacterial pathogens in 2019: a systematic analysis for the Global Burden of Disease Study 2019. Lancet, 2022. [DOI | PubMed]
- K Becker, C Heilmann, G Peters. Coagulase-negative staphylococci. Clin Microbiol Rev, 2014. [DOI | PubMed]
- H Oliveira, M Sampaio, LDR Melo. Staphylococci phages display vast genomic diversity and evolutionary relationships. BMC Genomics, 2019. [DOI | PubMed]
- S Kamitori, H Yoshida. Structure-function relationship of bacterial SH3 domains. SH Domains: Structure, Mechanisms and Applications, 2015
- C Alvarez-Carreño, P Penev, A Petrov, L Williams. Fold evolution before LUCA: common ancestry of SH3 domains and OB domains. Mol Biol Evol, 2021. [DOI | PubMed]
- P Mitkowski, E Jagielska, E Nowak. Structural bases of peptidoglycan recognition by lysostaphin SH3b domain. Sci Rep, 2019. [DOI | PubMed]
- LS Gonzalez-Delgado, H Walters-Morgan, B Salamaga. Two-site recognition of Staphylococcus aureus peptidoglycan by lysostaphin SH3b. Nat Chem Biol, 2020. [DOI | PubMed]
- A Beaussart, T Rolain, MC Duchene. Binding mechanism of the peptidoglycan hydrolase Acm2: low affinity, broad specificity. Biophys J, 2013. [DOI | PubMed]
- Y Shen, I Kalograiaki, A Prunotto. Structural basis for recognition of bacterial cell wall teichoic acid by pseudo-symmetric SH3b-like repeats of a viral peptidoglycan hydrolase. Chem Sci, 2021. [DOI]
- KO Lee, M Kong, I Kim. (2019) Structural basis for cell-wall recognition by bacteriophage PBC5 endolysin. Struct Lond Engl, 1993. [DOI]
- D Gutiérrez, B Martínez, A Rodríguez, P García. Isolation and Characterization of bacteriophages infecting Staphylococcus epidermidis. Curr Microbiol, 2010. [DOI | PubMed]
- D Gutiérrez, D Vandenheuvel, B Martínez. Two phages, phiIPLA-RODI and phiIPLA-C1C, Lyse Mono- and dual-species Staphylococcal biofilms. Appl Environ Microbiol, 2015. [DOI | PubMed]
- L Dijkshoorn, H Aucken, P Gerner-Smidt. Comparison of outbreak and nonoutbreak Acinetobacter baumannii strains by genotypic and phenotypic methods. J Clin Microbiol, 1996. [DOI | PubMed]
- H Gerstmans, D Grimon, D Gutierrez. A VersaTile-driven platform for rapid hit-to-lead development of engineered lysins. Sci Adv, 2020. [DOI | PubMed]
- S Delgado, R Arroyo, E Jiménez. Staphylococcus epidermidis strains isolated from breast milk of women suffering infectious mastitis: potential virulence traits and resistance to antibiotics. BMC Microbiol, 2009. [DOI | PubMed]
- P García, C Madera, B Martínez. Prevalence of bacteriophages infecting Staphylococcus aureus in dairy samples and their potential as biocontrol agents. J Dairy Sci, 2009. [DOI | PubMed]
- D Gutiérrez, S Delgado, D Vázquez-Sánchez. Incidence of Staphylococcus aureus and analysis of associated bacterial communities on food industry surfaces. Appl Environ Microbiol, 2012. [DOI | PubMed]
- J Valle, A Toledo-Arana, C Berasain. SarA and not σB is essential for biofilm development by Staphylococcus aureus. Mol Microbiol, 2003. [DOI | PubMed]
- C Cucarella, C Solano, J Valle. Bap, a Staphylococcus aureus Surface Protein Involved in Biofilm Formation. J Bacteriol, 2001. [DOI | PubMed]
- V Martín, A Maldonado-Barragán, L Moles. Sharing of bacterial strains between breast milk and infant feces. J Hum Lact Off J Int Lact Consult Assoc, 2012. [DOI]
- D Gutiérrez, V Garrido, L Fernández. Phage lytic protein LysRODI prevents Staphylococcal mastitis in mice. Front Microbiol, 2020. [DOI | PubMed]
- SC Potter, A Luciani, SR Eddy. HMMER web server: 2018 update. Nucleic Acids Res, 2018. [DOI | PubMed]
- Y Huang, B Niu, Y Gao. CD-HIT Suite: a web server for clustering and comparing biological sequences. Bioinformatics, 2010. [DOI | PubMed]
- U Bodenhofer, E Bonatesta, C Horejš-Kainrath, S Hochreiter. msa: an R package for multiple sequence alignment. Bioinformatics, 2015. [DOI | PubMed]
- L Zhou, T Feng, S Xu. ggmsa: a visual exploration tool for multiple sequence alignment and associated data. Brief Bioinform, 2022. [DOI | PubMed]
- D Charif, JR Lobry. SeqinR 1.0-2: a contributed package to the R project for statistical computing devoted to biological sequences retrieval and analysis. Structural Approaches to Sequence Evolution: Molecules, Networks, Populations, 2007
- KP Schliep. phangorn: phylogenetic analysis in R. Bioinformatics, 2011. [DOI | PubMed]
- S Xu, L Li, X Luo. Ggtree: a serialized data object for visualization of a phylogenetic tree and annotation data. iMeta, 2022. [DOI | PubMed]
- M Mirdita, K Schütze, Y Moriwaki. ColabFold: making protein folding accessible to all. Nat Methods, 2022. [DOI | PubMed]
- N Guex, MC Peitsch, T Schwede. Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: a historical perspective. Electrophoresis, 2009. [DOI | PubMed]
- 51.Wickham H (2016) ggplot2 elegant graphics for data analysis, 2nd ed. Springer International Publishing
- A Grundling, O Schneewind. Cross-linked peptidoglycan mediates lysostaphin binding to the cell wall envelope of Staphylococcus aureus. J Bacteriol, 2006. [DOI | PubMed]
- JZ Lu, T Fujiwara, H Komatsuzawa. Cell wall-targeting domain of glycylglycine endopeptidase distinguishes among peptidoglycan cross-bridges. J Biol Chem, 2006. [DOI | PubMed]
- X Zhou, L Cegelski. Nutrient-dependent structural changes in S. aureus peptidoglycan revealed by solid-state NMR spectroscopy. Biochemistry, 2012. [DOI | PubMed]
- S Kumar, R Nussinov. Close-range electrostatic interactions in proteins. ChemBioChem, 2002. [DOI | PubMed]
- KH Schleifer, O Kandler. Peptidoglycan types of bacterial cell walls and their taxonomic implications. Bacteriol Rev, 1972. [DOI | PubMed]
- N Batool, KS Ko, AK Chaurasia, KK Kim. Functional identification of serine hydroxymethyltransferase as a key gene involved in Lysostaphin resistance and virulence potential of Staphylococcus aureus strains. Int J Mol Sci, 2020. [DOI | PubMed]
- X Du, J Larsen, M Li. Staphylococcus epidermidis clones express Staphylococcus aureus-type wall teichoic acid to shift from a commensal to pathogen lifestyle. Nat Microbiol, 2021. [DOI | PubMed]
