Exploring CISD1 as a multifaceted biomarker in cancer: Implications for diagnosis, prognosis, and immunotherapeutic response
Abstract
CISD1, an outer mitochondrial membrane iron-sulfur cluster protein, regulates intracellular iron levels, oxidative stress, and mitochondrial dynamics, playing critical roles in cellular bioenergetics and redox homeostasis. Although CISD1 has been identified as a prognostic biomarker in specific cancers, its broader implications in tumorigenesis, cancer progression, and immunotherapy remain unclear. Given the heterogeneity of cancer and the need for robust biomarkers across cancers, this study conducts the first comprehensive pan-cancer analysis of CISD1 by evaluating its roles in cancer and treatment. We obtained and analyzed data from databases including TCGA, GTEx, THPA, GEPIA2.0, SangerBox, cBioPortal, TIMER2.0, CAMOIP, DAVID, SRPLOT, and TISIDB. Our findings reveal significant alterations in CISD1 expression at both transcriptional and translational levels, as well as gene mutations across multiple cancers, indicating its potential as a diagnostic biomarker and its involvement in cancer development and progression. CISD1 dysregulation is linked to poor clinical outcomes, as shown through its impact on patient prognosis. GO and KEGG analyses show that CISD1 plays critical roles in cellular bioenergetics. Notably, CISD1 expression is significantly correlated with tumor stemness indices, tumor mutation burden, microsatellite instability, and immune checkpoint proteins in multiple cancers, and altered CISD1 levels are also observed in patients responding to immunotherapy, further supporting its role not only in prognosis but also as a key predictor in immunotherapy responses and outcomes. Our findings demonstrate CISD1 as a reliable and promising diagnostic, prognostic, and immunotherapeutic biomarker for multiple cancers, emphasizing its crucial role in cancer biology and potential to guide personalized cancer therapies.
Article type: Research Article
Keywords: CISD1, Diagnostic biomarker, Immunotherapeutic biomarker, MitoNEET, Pan-cancer analysis, Prognostic biomarker
Affiliations: Department of Interdisciplinary Oncology, School of Medicine, LSU LCMC Cancer Center, Louisiana State University Health Sciences Center, New Orleans, LA 70112, USA; Genetics Graduate Program, School of Medicine, LSU LCMC Cancer Center, Louisiana State University Health Sciences Center, New Orleans, LA 70112, USA; Department of Pathology Laboratory Medicine, School of Medicine, Tulane University, New Orleans, LA 70112, USA; IDP Graduate Program, School of Medicine, LSU LCMC Cancer Center, Louisiana State University Health Sciences Center, New Orleans, LA 70112, USA; Department of Surgery, Colleges of Medicine and Public Health and Health Professions, University of Florida, Gainesville, FL 32611, USA; Department of Biostatistics, Colleges of Medicine and Public Health and Health Professions, University of Florida, Gainesville, FL 32611, USA; Department of Pharmacology and Toxicology, University of Texas Medical Branch, Galveston, TX 77555, USA; Department of Biological Sciences, BMB Division, Louisiana State University, Baton Rouge, LA 70803, USA
License: © 2025 The Authors. Publishing services by Elsevier B.V. on behalf of KeAi Communications Co., Ltdé. CC BY 4.0 This is an open access article under the CC BY license (http://creativecommons.org/licenses/by/4.0/).
Article links: DOI: 10.1016/j.gendis.2025.101677 | PubMed: 40831536 | PMC: PMC12359157
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (5.0 MB)
Introduction
Despite significant advances made in diagnosis and treatment in the past decades, cancer remains a major health challenge and the leading cause of mortality worldwide.ref. bib1 The heterogeneous nature of cancer necessitates the identification of robust biomarkers for early detection, prognosis, and treatment response across multiple cancer types.ref. bib2 Such biomarkers can be classified into diagnostic biomarkers, prognostic biomarkers, and/or predictive biomarkers.ref. bib2 Diagnostic biomarkers help in the early detection of cancer, whereas prognostic biomarkers predict the likely course of diseases, and predictive biomarkers forecast the response to specific therapies.ref. bib2 Current cancer biomarkers typically include genetic mutations such as breast cancer susceptibility gene 1/2 (BRCA1/2) in breast cancer,ref. bib3 protein expression levels such as caveolin-1 in gastrointestinal cancers,ref. bib4 and circulating tumor cells in hepatocellular carcinomaref. bib5 or DNA (ctDNA) in the bloodstream.ref. bib6 However, there is still an urgent unmet need for novel and more effective biomarkers to enhance the accuracy and efficacy of cancer diagnosis and treatment.
The CDGSH iron sulfur domain (CISD) gene family encodes a group of NEET proteins (mitoNEET, NAF-1, and MiNT)ref. bib7 that are a novel class of iron-sulfur ([2Fe–2S]) proteins characterized by a unique CDGSH domain, which binds to their respective [2Fe–2S] clusters.ref. bib8 This domain gives NEET proteins distinctive properties, such as redox activity and lability of the [2Fe–2S] clusters, due to their unique coordination structure (3Cys:1His).ref. bib8,ref. bib9 This family includes three members: CISD1 (mitoNEET), CISD2 (NAF-1), and CISD3 (MiNT).ref. bib7 MitoNEET (CISD1) is located on the outer mitochondrial membrane, and its unique [2Fe–2S] clusters endow mitoNEET with critical redox and pH-sensing properties, enabling it to maintain iron and redox homeostasis in the mitochondria.ref. bib8,ref. bib10 Nutrient-deprivation autophagy factor-1 (NAF-1/CISD2) is found in the outer mitochondrial membrane, endoplasmic reticulum, and mitochondria-associated membranes, and it links mitochondrial and endoplasmic reticulum functions, contributing to calcium signaling, autophagy, and apoptosis regulation; dysregulation of CISD2 is implicated in diseases like Wolfram syndrome and several cancers.ref. bib8 Miner2 (CISD3) is localized in the mitochondrial matrix and primarily regulates mitochondrial iron-sulfur cluster biogenesis and respiratory chain function. Its role in cancer and other pathologies remains less defined compared with CISD1 and CISD2.ref. bib8 Among the CISD family, CISD1 holds a distinct position due to its involvement in regulating mitochondrial dynamics, bioenergetics, ferroptosis, and apoptosis, playing a key role in cellular metabolism and oxidative stress response.ref. bib8,ref. bib10 The ability of the CISD1 protein to modulate mitochondrial iron metabolism and reactive oxygen species production suggests its significant influence on cellular function and survival. Intriguingly, our prior and current studies show that CISD1 functions as a novel redox enzyme to enhance oxidation of NADH and enlarge the NAD+ pool in the cytoplasm,ref. bib11 possibly leading to aberrantly increased glycolysis and ATP/energy production. The role of CISD1 is even more important in cancers, as cancer cells often exhibit alterations in mitochondrial function to support rapid proliferation and resistance to apoptosis. Actually, studies have shown that CISD1 promotes breast cancer survival and proliferation by protecting the mitochondria of cancer cells from iron overaccumulation, enhancing the tolerance of cancer cells to oxidative stress, and suppressing autophagy and apoptosis.ref. bib8 We speculated that by enhancing energy production and glycolysis in somatic cells by enhancing oxidation of NADH and the FMNH2/NADH energy production pathway/axis, CISD1 may act like an oncogene to promote malignant transformation and carcinogenesis.ref. bib12 The role of CISD1 in cancer was first shown in breast cancer, where CISD1 is overexpressed and suppression of CISD1 significantly reduces cell proliferation and tumor growth,ref. bib13,ref. bib14 and it has been further identified as a novel target for breast cancer chemotherapy.ref. bib15 In addition to breast cancer, CISD1 has been demonstrated as a potential novel antileukemic drug target for refractory or relapsed B-cell acute lymphocytic leukemia.ref. bib16 These reports indicate CISD1’s promising role in cancer treatment.
Although CISD1 has been identified as a prognostic biomarker in several cancers, such as breast cancer,ref. bib17,ref. bib18 hepatocellular carcinoma,ref. bib19 gastric cancer,ref. bib20 and bladder cancer,ref. bib21 there is still a lack of systematic pan-cancer analysis of CISD1, which is vital to the understanding of its broad implications in cancer biology and clinical oncology. This manuscript aims to elucidate the expression patterns, genomic alterations, stemness profiles, and clinical significance of CISD1 across multiple cancer types. Using large-scale genomic datasets and advanced bioinformatics tools, we provide insights into the potential of CISD1 as a diagnostic, prognostic, and immunotherapeutic biomarker. Our results indicate that CISD1 expression correlates with patient prognosis and immune cell infiltration in various cancers. This pan-cancer approach highlights the universal and specific roles of CISD1, supporting its candidacy as a robust biomarker for predicting cancer outcomes and tailoring immunotherapy strategies. The comprehensive analyses presented in this study establish the significance of CISD1 in cancer and its potential value as a multi-biomarker for improving cancer prognosis and treatment.
Materials and methods
CISD1 gene and protein expression analysis
The mRNA expression data of CISD1 in 36 types of human normal tissues from the Genotype Tissue Expression (GTEx) cohortref. bib22 and 17 types of tumor tissues from The Cancer Genome Atlas (TCGA) cohort were downloaded from The Human Protein Atlas (THPA) online website.ref. bib23 Specifically, the data was accessed by navigating to the “DATA” section of the website, where the relevant RNA expression dataset was identified and subsequently downloaded from the “DOWNLOADABLE DATA” section, and the data distribution was visualized using GraphPad Prism (version 10). The Gene Expression Profiling Interactive Analysis 2.0 (GEPIA2) is a web server for gene expression analysis based on tumor and normal samples from the TCGA and the GTEx databases; it accesses and processes data using R and Perl scripts,ref. bib24 so we used GEPIA2 to visualize the different mRNA expression level of CISD1 across 33 TCGA tumor tissues versus their normal tissues (TCGA normal tissues combined with GTExref. bib22 normal tissues). Briefly, by navigating to the “Expression Analysis” section and selecting “Expression Profile”, the gene CISD1 was searched in the “Gene” bar. Subsequently, data for all 33 cancer types were selected to obtain the results. The SangerBox portal (version 3.0) is a web-based tool platform that integrates GEO, TCGA, ICGC, and other databases and processes data in batches.ref. bib25 We used this web tool to obtain and visualize the mRNA expression levels of CISD1 in different cancer stages, and in male/female patients of various cancers. The image staining results of protein expression of CISD1 in normal/cancer tissues were obtained from THPA in the “TISSUE"/"CANCER” section.
Genetic alteration analysis
Characteristics of CISD1 genetic alterations were explored using the online cBio Cancer Genomics Portal (cBioPortal) database.ref. bib26 Briefly, by selecting the “Quick Search” section, mutation type, alteration frequency, and copy number alteration data of CISD1 were obtained across the TCGA tumor datasets in the “Cancer Types Summary” module, and mutation types, sites, and numbers were obtained in the “Mutations” module. Genetic mutation ratios and mutation types of CISD1 in different tumors were obtained via the “Gene_Mutation” module on Tumor Immune Estimation Resource 2.0 (TIMER2)ref. bib27 and via SangerBox, respectively. The mRNA expression values of CISD1 and its corresponding copy-number alteration values were analyzed using the SangerBox portal. Correlations of CISD1 expression and somatic mutations were obtained from the Comprehensive Analysis on Multi-Omics of Immunotherapy in Pan-cancer (CAMOIP)ref. bib28 by selecting the “Mutational Landscape” section.
Survival prognosis analysis
Overall survival, disease-specific survival, disease-free survival, and progression-free survival of CISD1 in various tumor types in the TCGA database were evaluated using forest plots by SangerBox online tools according to the website instructions. Overall survival map and the Kaplan–Meier survival plots were generated using GEPIA2 by selecting the “Survival Analysis” and “Survival Map” sections. High-expression and low-expression cohorts of CISD1 were obtained through the expression threshold of the cutoff-high (25%) and cutoff-low (75%) values.
Stemness indices correlation analysis
Values of the correlation coefficient (Pearson’s R) between CISD1 and stemness indices, including DNAss, EREG-METHss, DMPss, ENHss, RNAss, or EREG.EXPss were calculated and downloaded from the SangerBox portal, and the radar charts were generated using Microsoft Excel (Office 365). Correlations between CISD1 and RNA modifications were evaluated, and the chart was generated using Pearson’s R by the SangerBox portal.
CISD1 coexpression gene enrichment analysis
CISD1 coexpression genes and their corresponding Pearson coefficient were downloaded from cBioPortal by selecting the “Co-expression” section. CISD1 positively correlated genes with Pearson’s r > 0.3 were selected to perform Gene Ontology (GO) term enrichment and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis using DAVID.ref. bib29 Briefly, by navigating to the “Start Analysis” section, uploading and submitting the positive gene list, then selecting “Functional Annotation Tool”, the GO and KEGG gene cluster results were downloaded from the website. Venn diagram, GO Chord, and GO/KEGG pathway were visualized using Science and Research Online Plot (SRPLOT),ref. bib30 which is an easy-to-use web server integrating more than a hundred commonly used data visualization and graphing functions together. GEPIA2 was used to visualize the heatmap of CISD1-positive coexpression genes by selecting the “Multiple Genes Comparison” section. Correlations between CISD1 and age were evaluated in the SangerBox portal according to the website instructions.
Immune infiltration and immune-related genes analysis
Using Person’s R correlations of CISD1 expression and StromalScore, ImmuneScore, ESTIMATEscore, or the proportion of tumor infiltration immune cells, including CD4+ T cells, CD8+ T cells, B cells, neutrophils, macrophages, and dendritic cells in tumors across cancer types, were assessed via the SangerBox portal. Correlation coefficient (Pearson’s R) between CISD1 expression and tumor mutation burden (TMB), microsatellite instability (MSI), or neoantigens was downloaded from the SangerBox portal. Radar charts were generated using Microsoft Excel. Also, correlations between CISD1 expression and immune regulation genes, including immunoinhibitor and immunostimulator in different tumors from TCGA cohorts, were assessed via the SangerBox portal. In addition, correlations between CISD1 and chemokine or major histocompatibility complex (MHC) were assessed in TISIDB,ref. bib31 which is a user-friendly web portal integrating multiple types of data resources in oncoimmunology by selecting the “Chemokine” and “Immunomodulator” sections. Expression levels of CISD1 in immunotherapy responders and non-responders were also obtained from TISIDB by selecting the “Immunotherapy” section.
Statistical analysis
Hazard ratio (HR) and p-value were used to evaluate the significance of differences in survival analyses. Pearson’s correlation coefficient and statistical significance were used to assess associations of gene expression, with the absolute value used to determine the strength of correlation. These results were considered statistically significant at ∗p < 0.05, ∗∗p < 0.01, ∗∗∗p < 0.001, and ∗∗∗∗p < 0.0001.
Results
Expression of CISD1 is up-regulated in multiple cancer types
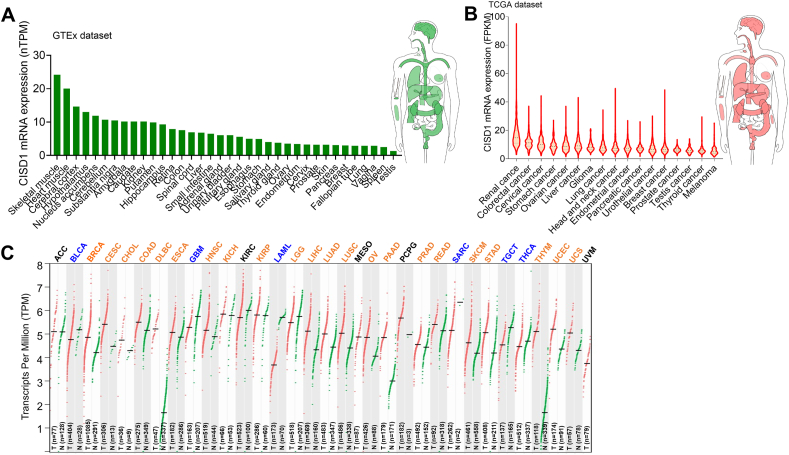
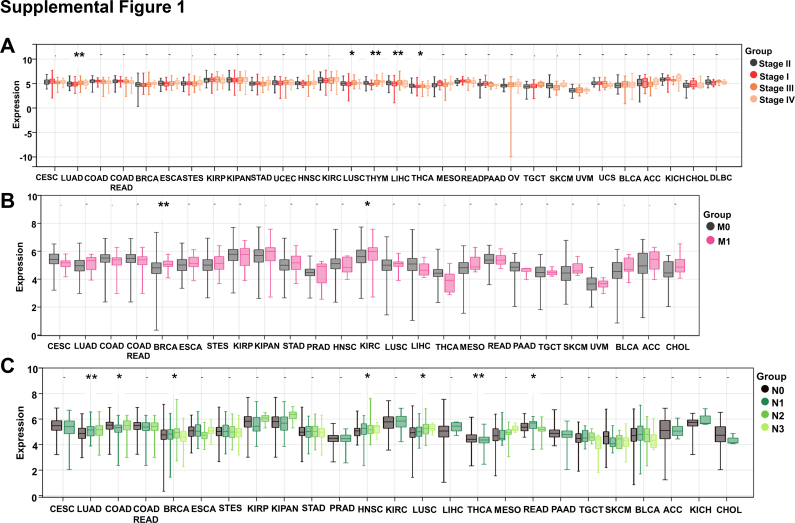
Gene expression alterations often occur in the development and progression of different cancers. CISD1 was reported to be up-regulated in breast cancer and acute lymphoblastic leukemia.ref. bib13,ref. bib15,ref. bib16 We therefore wondered if its expression was also altered across cancer types. We downloaded CISD1 mRNA expression data in different human normal tissues from GTEx datasets and in different tumor tissues from TCGA datasets via the THPA online website and analyzed the CISD1 mRNA expression level. In normal tissues, CISD1 had the highest mRNA expression level in skeletal muscle, followed by heart muscle, cerebral cortex, hypothalamus, etc. (Fig. 1A), suggesting that CISD1 may have a fundamental role in normal physiology in these tissues and organs. In tumor samples, it had the highest mRNA expression level in renal cancer, followed by colorectal cancer, cervical cancer, stomach cancer, etc. (Fig. 1B). Next, we wondered how many tumor tissues showed higher mRNA expression levels of CISD1 compared with corresponding normal tissues. Using GEPIA 2.0 online tools, we analyzed the mRNA expression levels of CISD1 in 33 tumor tissues compared with their corresponding normal tissues (TCGA normal tissue + GTEx normal tissues). We found that the mRNA expression level of CISD1 was prevalently up-regulated in 22 tumor tissues including breast invasive carcinoma (BRCA), cervical squamous cell carcinoma and endocervical adenocarcinoma (CESC), cholangiocarcinoma (CHOL), colon adenocarcinoma (COAD), lymphoid neoplasm diffuse large B-cell lymphoma (DLBC), esophageal carcinoma (ESCA), head and neck squamous cell carcinoma (HNSC), kidney chromophobe (KICH), kidney renal papillary cell carcinoma (KIRP), brain lower grade glioma (LGG), liver hepatocellular carcinoma (LIHC), lung adenocarcinoma (LUAD), lung squamous cell carcinoma (LUSC), ovarian serous cystadenocarcinoma (OV), pancreatic adenocarcinoma (PAAD), prostate adenocarcinoma (PRAD), rectum adenocarcinoma (READ), skin cutaneous melanoma (SKCM), stomach adenocarcinoma (STAD), thymoma (THYM), uterine corpus endometrial carcinoma (UCEC), and uterine carcinosarcoma (UCS) (Fig. 1C), suggesting an oncogenic role for CISD1 in these cancers. Conversely, CISD1 expression was down-regulated in 6 tumor tissues: bladder urothelial carcinoma (BLCA), glioblastoma multiforme (GBM), acute myeloid leukemia (LAML), sarcoma (SARC), testicular germ cell tumors (TGCT), thyroid carcinoma (THCA) (Fig. 1C), suggesting that CISD1 may play tumor suppressive functions in these cancers. CISD1 showed no alteration of mRNA expression level in 5 tumor tissues: adrenocortical carcinoma (ACC), kidney renal clear cell carcinoma (KIRC), mesothelioma (MESO), pheochromocytoma and paraganglioma (PCPG), uveal melanoma (UVM) (all abbreviations are showed in Table S1) (Fig. 1C). Furthermore, from SangerBox online tools analysis, we found a gradual increase in CISD1 mRNA levels across different cancer stages, from stage I to stage IV, in LUAD, LUSC, THYM, and LIHC (Fig. S1A), further confirming the possibility that CISD1 plays an oncogenic role and is associated with cancer progression in these cancers. In addition, CISD1 may be important for metastasis in certain cancer types, as we found that CISD1 mRNA levels were increased in the M1 stage when compared with the M0 stage in BRCA and KIRC (Fig. S1B), and increased across different lymph node stages, from stage N0 to stage N3, in LUAD, BRCA, HNSC, LUSC, and READ (Fig. S1C); while decreased in COAD and THCA (Fig. S1C).
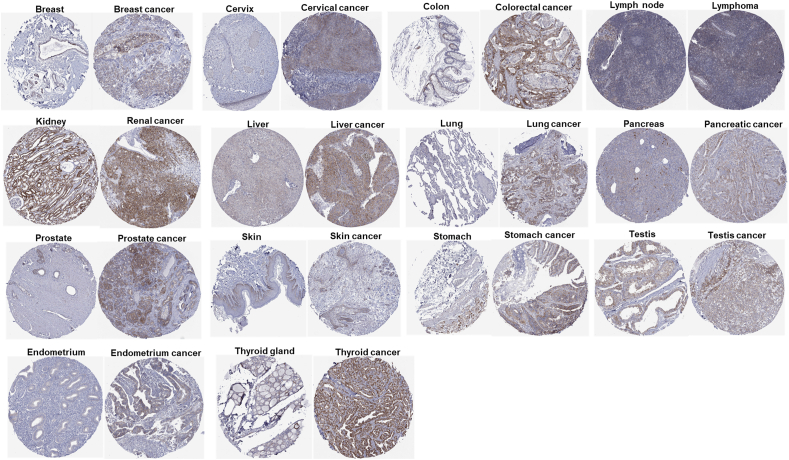
Next, we wondered if the protein expression level of CISD1 was also increased in various cancers. We downloaded the protein staining images from THPA, and found that the protein level of CISD1 was significantly higher in 14 cancers compared with their corresponding normal tissues including breast cancer, cervix cancer, colorectal cancer, lymphoma, renal cancer, liver cancer, lung cancer, pancreatic cancer, prostate cancer, skin cancer, stomach cancer, testis cancer, endometrium cancer, and thyroid cancer (Fig. 2), which is consistent with the mRNA analysis in Figure 1. Taken together, these results reveal significant variations in CISD1 expression, with most cancers showing elevated levels. This suggests CISD1 may be involved in tumorigenesis and is a potential diagnostic biomarker for these cancer types. These findings further highlight the importance of conducting comprehensive pan-cancer analyses to uncover the broader implications of CISD1 in oncology.
Alterations in the CISD1 gene are associated with the development and progression of multiple cancer types
In addition to gene expression, cancer patient outcomes such as survival and disease recurrence are affected by gene alterations and mutations. Activation of oncogenes or inactivation of tumor suppressor genes drives the initiation and progression of cancer.ref. bib32,ref. bib33 Understanding such alterations helps in elucidating the mechanisms of cancer development and disease progression. We next analyzed the genetic profile of CISD1 in cancer tissues. Using cBioPortal, CISD1 showed mutation frequency in several cancers, in which UCEC (>1%) had the highest rate of CISD1 gene mutations frequency, followed by STAD, SARC, LIHC, SKCM, OV, and LUAD (Fig. 3A). Consistent with these results, we found that UCEC showed the highest percentage of patient samples with CISD1 mutations (0.0094%), followed by STAD (0.0046%), SARC (0.0042%), LIHC (0.0027%), OV (0.0024%), SKCM (0.0021%), and LUAD (0.0019%) from exploring CISD1 gene mutation rates on TIMER 2.0 website (Fig. 3B). Using SangerBox online analysis, we further explored the mutation types and mutation sites of CISD1 and found that missense mutations were the main type of CISD1 mutation in cancer (Fig. 3C). According to cBioPortal online analysis (Fig. 3D), the mutation sites frequently occur within the zf-CDGSH domain, which binds to iron to form a redox-active pH-labile 2Fe–2S cluster, and A69 S/V is a hot spot mutation site, suggesting that the mutations of CISD1, especially the hot spot A69 S/V, could be a diagnostic biomarker.
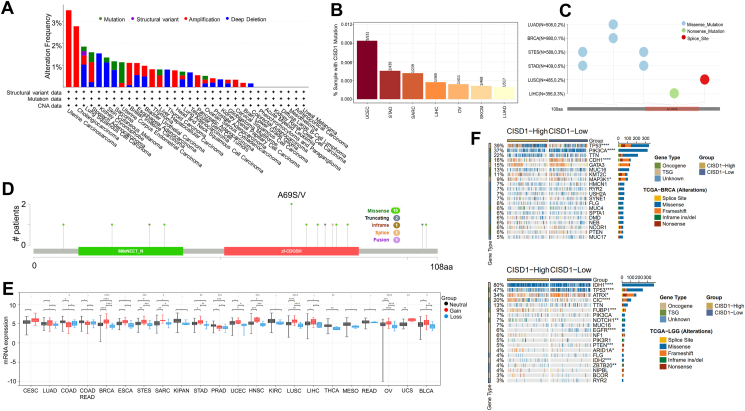
In addition, cBioPortal analysis showed that UCS (>3%), CHOL (>2%), STAD (>1%), and BRCA (>1%) had a higher rate of CISD1 gene amplification frequency (Fig. 3A); while kidney chromophobe (>1%), SKCM (>1%), and lung squamous (>1%) had a higher rate of CISD1 gene deep deletion frequency (Fig. 3A). Further analysis utilizing SangerBox showed that gain copy number of CISD1 is associated with higher mRNA expression levels of CISD1 in 19 cancer types including CESC, LUAD, BRCA, ESCA, STES (Stomach and Esophageal carcinoma), SARC, KIPAN (Pan-kidney cohort (KICH + KIRC + KIRP), STAD, UCEC, HNSC, KIRC, LUSC, LIHC, THCA, MESO, READ, OV, UCS, BLCA (Fig. 3E), suggesting that CISD1 mRNA expression level is mainly due to the copy-number alterations.
Next, the relationship between CISD1 expression and specific genomic characteristics, such as somatic mutations, was analyzed using CAMOIP online analysis. We found that CISD1 was strongly associated with somatic mutations in multiple cancers. The CISD1-high group showed high frequency of somatic mutations in TP53 (39%), PIK3CA (37%), CDH1 (16%), MAP3K1 (9%) in BRCA (Fig. 3F, top), and IDH1 (80%), TP53 (47%), ATRX (34%), CIC (20%), FUBP1 (9%), NOTCH1 (7%), EGFR (6%), PTEN (5%) in LGG (Fig. 3F, bottom). Overall, these results show CISD1 gene mutations, amplifications, and deletions are present in multiple cancers, with missense mutations as the most frequent type of CISD1 gene mutations in various cancers, and copy number alterations as the most frequent genomic change in most of the cancers. Moreover, expression of CISD1 was strongly associated with somatic mutations. Taken together, these results suggest that CISD1 gene alterations may regulate the initiation, growth, and progression of various cancers and can serve as a potential diagnostic biomarker.
CISD1 expression is associated with the prognosis of various tumors
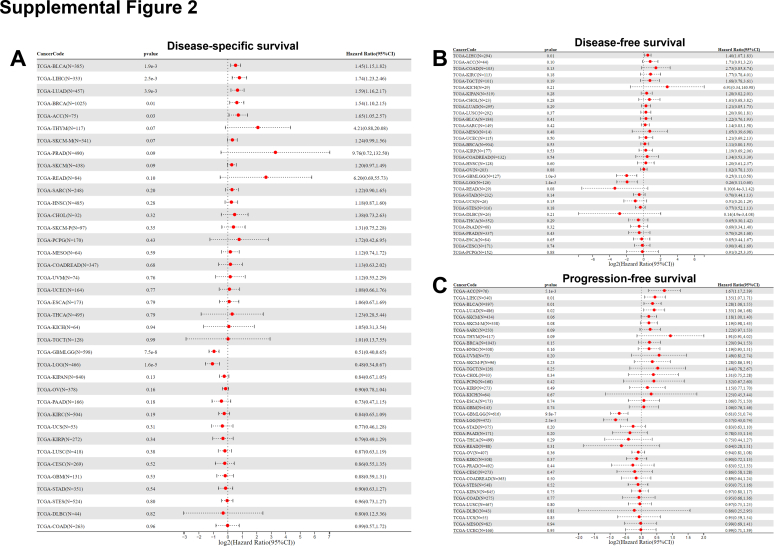
Gene expression profiles can help identify biomarkers that predict patient outcomes, such as overall survival, disease-specific survival, disease-free survival, and progression-free survival.ref. bib34 Our findings above render CISD1 a candidate for further investigation as a prognostic biomarker. Thus, we compared survival in cancer patients with high or low CISD1 expression levels. Using SangerBox online tools, we generated survival forest plots and found that higher CISD1 expression was significantly associated with worse patient overall survival in LIHC (HR = 1.83, p = 3.0e-5), LAML (HR = 1.40, p = 4.6e-5), LUAD (HR = 1.58, p = 4.2e-4), BRCA (HR = 1.53, p = 6.1e-4), THYM (HR = 4.92, p = 3.7e-3), BLCA (HR = 1.32, p = 4.0e-3), SCKM (HR = 1.27, p = 0.03), and ACC (HR = 1.64, p = 0.03) (Fig. 4A). In contrast, higher CISD1 expression was significantly associated with lower risk of death in GBMLGG (Glioma) (HR = 0.54, p = 5.8e-8), LGG (HR = 0.52, p = 8.3e-5), and NB (Neuroblastoma) (HR = 0.58, p = 0.01) (Fig. 4A). We also used GEPIA 2.0 online tools to generate survival heatmaps and Kaplan–Meier survival curves to reveal the relationship between CISD1 expression and overall survival across various cancer types. From the survival heatmaps and the corresponding Kaplan–Meier survival curves, high CISD1 expression is associated with worse patient survival (Fig. 4B) for ACC, HR = 2.8, p = 0.0061; BLCA, HR = 1.5, p = 0.0082; BRCA, HR = 1.6, p = 0.009; KICH, HR = 4, p = 0.02; LAML, HR = 1.9, p = 0.025; LIHC, HR = 2.1, p = 5.7e-05; LUAD, HR = 1.6, p = 0.003); MESO, HR = 2.2, p = 0.004; THCA, HR = 3.5, p = 0.0081; and THYM, HR = 5.7, p = 0.0075. Furthermore, disease-specific survival analysis showed that BLCA (HR = 1.45, p = 1.9e-3), LIHC (HR = 1.74, p = 2.5e-3), LUAD (HR = 1.59, p = 3.9e-3), BRCA (HR = 1.54, p = 0.01), ACC (HR = 1.65, p = 0.03) had significantly stronger association between CISD1 and patient survival (Fig. S2A) where patients with higher expression levels of CISD1 had a higher risk of death. Disease-free survival and progression-free survival analyses are essential tools for assessing treatment efficacy.ref. bib35 Disease-free survival analysis revealed that LIHC patients had a significantly higher risk of death (HR = 1.40, p = 0.01) (Fig. S2B). Similarly, progression-free survival analysis indicated that patients with elevated CISD1 expression faced a heightened risk of death across multiple cancers: ACC (HR = 1.67, p = 5.1e-3), LIHC (HR = 1.35, p = 0.01), BLCA (HR = 1.28, p = 0.01), LUAD (HR = 1.33, p = 0.02), and SKCM (HR = 1.18, p = 0.05) (Fig. S2C), suggesting that the progression of the disease cannot be effectively controlled by the treatment of these cancers. In contrast, patients with higher CISD1 expression showed significantly lower risk of death in GBMLGG (Fig. S2A–C) and LGG (Fig. S2A–C) from disease-specific survival analysis, disease-free survival analysis, and progression-free survival analysis.
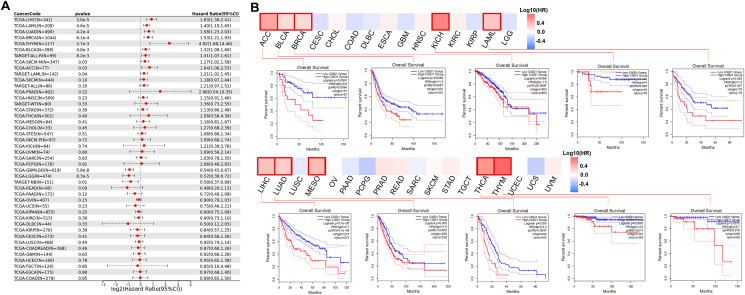
Although the results highlighted that CISD1 expression levels have varying impacts on overall survival across different cancer types, high CISD1 expression is generally associated with worse survival. Taken together, the significant associations between CISD1 expression and patient outcomes strongly suggest that CISD1 can be a potential prognostic biomarker for multiple cancers.
CISD1 is positively correlated with cancer stemness and RNA modifications in multiple cancers
Cancer stem cells have the unique ability to self-renew and differentiate into various cell types that make up the tumor; these cells are thought to be responsible for tumor initiation, progression, therapeutic resistance, and relapse after treatment.ref. bib36 Stemness indices, such as DNA methylation-based stemness score (DNAss), Differentially Methylated Position Stemness Score (DMPss), Enhancer Stemness Score (ENHss), RNA Stemness Score (RNAss), Epigenetically Regulated Gene Methylation Stemness Score (EREG-METHss), and Epigenetically Regulated Gene Expression Stemness Score (EREG.EXPss), quantify the stem-like characteristics of cancer cells, making them valuable tools for studying tumor biology.ref. bib37,ref. bib38 We assessed whether CISD1 is associated with these stemness indices. Using SangerBox online analysis tools, surprisingly, we found that CISD1 was significantly and positively correlated with cancer stemness indices: DNAss in OV, DLBC, THYM, STAD, and STES (Fig. 5A); EREG-METHss in OV, DLBC, THYM, STAD, SKCM, MESO, PAAD, STES, PRAD, and BRCA (Fig. 5B); DMPss in OV, DLBC, THYM, STAD, LAML, LIHC, STES, and LUAD (Fig. 5C); ENHss in OV, DLBC, LAML, LIHC, STAD, and ESCA (Fig. 5D). Intriguingly, CISD1 was significantly and positively correlated with RNAss in multiple cancer types including DLBC, UCEC, LGG, LUAD, KIRC, GBMLGG, THCA, READ, SARC, BRCA, SKCM, PCPG, LAML, LIHC, KIPAN, COADREAD (colon adenocarcinoma/rectum adenocarcinoma), TGCT, ESCA, HNSC, COAD, LUSC, CESC, KIRP, and OV (Fig. 5E); and EREG.EXPss in READ, DLBC, THYM, ACC, THCA, LAML, COADREAD, PRAD, COAD, TGCT, OV, PAAD, BRCA, LUAD, LIHC, GBM, UCEC, ESCA, STEX, STAD, BLCA, and KIPAN (Fig. 5F). These results suggest that higher CISD1 expression is associated with enhanced expression of stemness-related genes, likely contributing to maintaining or enhancing these stem-like properties. This association could explain its potential role in promoting tumor aggressiveness and therapeutic resistance in these cancers. Whereas, CISD1 showed a significant negative correlation with cancer stemness indices: DNAss in KIRP, LGG, KIRC, BLCA, THCA, GBMLGG, KIPAN, and CESC (Fig. 5A); EREG-METHss in KIRP, BLCA, THCA, KIRC, LGG, KIPAN, GBMLGG, and CESC (Fig. 5B); DMPss in KIRP, LGG, BLCA, KIRC, KIPAN, CESE, GBMLGG, and THCA (Fig. 5C); ENHss in LGG, GBMLGG, KIRC, KIRP, BLCA, and KIPAN (Fig. 5D); RNAss in BLCA (Fig. 5E). Overall, the association between CISD1 and cancer cell stemness varies by cancer type.
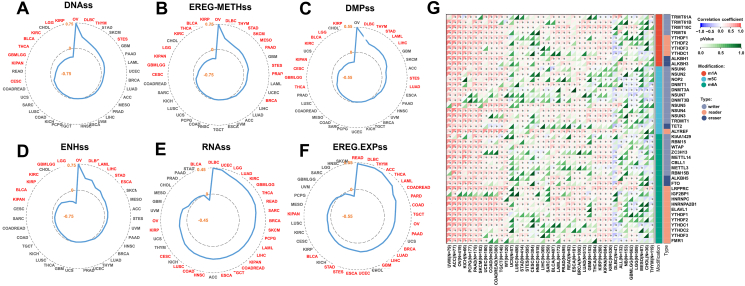
The results above show that CISD1 expression is linked to the methylation status of genes regulated by epigenetic mechanisms that contribute to cancer stemness, especially, higher CISD1 expression is associated with an RNA expression profile that resembles that of cancer stem cells, implying that CISD1 is associated with epigenetic modifications. Epigenetic modifications, such as RNA modifications, the new frontier of this arena, have significant implications for cancer progression and treatment.ref. bib39 We next used SangerBox online tools to analyze the correlation between CISD1 mRNA expression and RNA modification enzymes that catalyze m1A, m5C, and m6A, as these RNA modifications play critical roles in cancer by regulating gene expression, mRNA stability, translation, and tumor progression.ref. bib39 As shown in Figure 5G, CISD1 mRNA expression was significantly and positively correlated with m1A, m5C, and m6A related RNA modification enzymes in UVM, ACC, OV, KICH, PCPG, PAAD, SKCM, UCEC, COAD, COADREAD, TGCT, WT, STAD, STES, HNSC, LIHC, BLCA, LAML, PRAD, READ, ESCA, BRCA, LUAD, THCA, KIRP, KIRC, KIPAN, and KIRC, suggesting that m1A, m5C, and m6A RNA modifications may contribute to gene expression related to tumor progression in these cancers in which when CISD1 expression increases. In contrast, CISD1 mRNA expression was significantly and negatively correlated with m1A, m5C, and m6A related RNA modification enzymes in DLBC (Fig. 5G). Taken together, these results suggest that CISD1 could be a potential biomarker for identifying tumors with high stemness and may serve as a therapeutic target to inhibit these aggressive tumor characteristics.
CISD1 coexpression gene enrichment analysis in various tumors
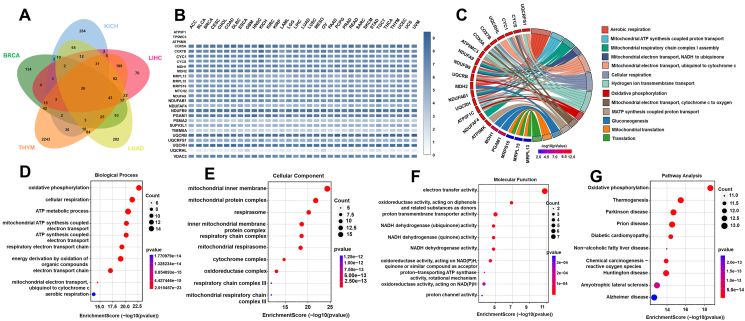
To understand the biological significance of CISD1 and its coexpression genes and their roles within cellular processes, and to elucidate how CISD1 influences cancer progression and patient outcomes, we performed GO and KEGG enrichment analyses for CISD1 and its coexpression genes, as GO and KEGG analyses are crucial in understanding the biological significance of genes and their roles within cellular processes.ref. bib40,ref. bib41 Since BRCA, KICH, LIHC, LUAD, and THYM all showed higher mRNA expression levels of CISD1, high stemness signatures, and high levels of RNA modifications with worse patient survival from the results above, we wondered if there are overlapping CISD1 coexpression genes in these cancers, and if these genes are potentially involved in fundamental processes related to CISD1 in cancers. We first downloaded the CISD1-positive coexpression genes in these five cancers from cBioPortal, and performed Venn analysis using SangerBox online tools, and found that there were 26 overlapped genes in the five cancers analyzed (Fig. 6A). Next, we analyzed the mRNA expression levels of these 26 overlapped genes in different cancers, and the heatmap showed that 20 of these overlapped genes were highly expressed in different cancers (Fig. 6B), suggesting that these genes may play crucial roles in ontogenesis and cancer progression. From GO enrichment analysis, we found that these genes are involved in oxidative phosphorylation, cellular respiration, ATP metabolic process, mitochondrial ATP synthesis coupled electron transport, ATP synthesis coupled electron transport, respiratory electron transport chain, energy derivation by oxidation of organic compounds, electron transport chain, mitochondrial electron transport ubiquinol to cytochrome c, and aerobic respiration (Fig. 6C, D). They were mainly localized to mitochondria, respirasome, respiratory chain complex, cytochrome, and oxidoreductase complex (Fig. 6E), with molecular functions of electron transfer activity, oxidoreductase activity, proton transmembrane transporter activity, NADH dehydrogenase activity, proton-transporting ATP synthase activity, and proton channel activity (Fig. 6F). To further understand how these genes interact with networks and pathways, we also performed KEGG enrichment analysis. These CISD1 coexpressed genes were involved in thermogenesis, Parkinson’s disease, prion disease, diabetic cardiomyopathy, non-alcoholic fatty liver disease, chemical carcinogenesis, reactive oxygen species, Huntington disease, amyotrophic lateral sclerosis, and Alzheimer’s disease (Fig. 6G). Taken together, these results suggest that CISD1 coexpression genes are involved in vital biological processes, particularly related to cellular energy production and metabolic processes, and likely play significant roles in cancer cell survival and proliferation.
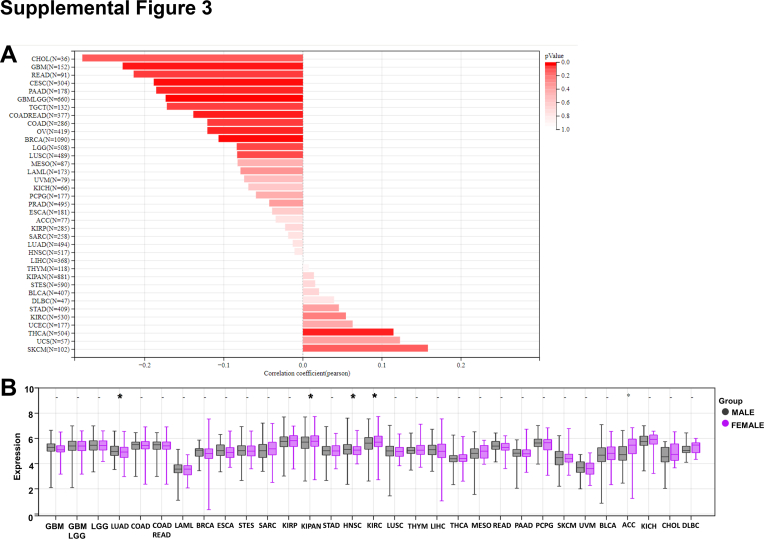
Interestingly, CISD1 mRNA expression levels were negatively correlated with age in CHOL, GBM, READ, CESC, PAAD, GBMLGG, TGCT, COADREAD, COAD, OV, BRCA, LGG, and LUSC (Fig. S3A), suggesting that younger patients have higher mRNA levels of CISD1 in these cancers, further supporting the above results that CISD1 is associated with bioenergetics. Male patients showed higher mRNA expression levels of CISD1 in LUAD, HNSC (Fig. S3B), while female patients showed higher mRNA expression levels of CISD1 in KIPAN and KIRC (Fig. S3B), suggesting that its expression pattern in cancer has limited gender preference.
CISD1 regulates tumor infiltration of immune cells in multiple cancers
The tumor microenvironment (TME) is composed of various immune cells, stromal cells, and extracellular components.ref. bib42 Immune infiltration analysis helps in understanding the composition and functional state of these immune cells within the TME, and whether the TME of a tumor can promote or inhibit tumor growth and metastasis.ref. bib43,ref. bib44 StromalScore quantifies the presence of stromal cells, including fibroblasts, mesenchymal cells, and other connective tissue cells that support the structure of tissues within the tumor microenvironment. A high stromal content is often associated with more aggressive tumor behavior, resistance to therapy, and poorer prognosisref. bib45; ImmuneScore assesses the infiltration of immune cells, particularly lymphocytes like T cells, B cells, and natural killer (NK) cells et al in the tumor microenvironment, and a higher ImmuneScore is often associated with better responses to immunotherapy and better patient outcomes.ref. bib45 An ESTIMATEscore is a composite score derived from the StromalScore and ImmuneScore, where a higher ESTIMATEscore indicates an immunotherapy could be less effective.ref. bib45 We wondered if CISD1 is associated with immune infiltration and immunotherapy outcomes. Raw data were downloaded from SangerBox, and immune scores were analyzed. The results showed that CISD1 was significantly and positively correlated with StromalScore in LAML and BLCA, suggesting that patients with higher expression of CISD1 may be more resistant to therapies in these cancers; while CISD1 showed significantly negative correlation with StromalScore in UCEC, acute lymphocytic leukemia, LGG, GBMLGG, COAD, THCA, SKCM, KIRC, and KIPAN (Fig. 7A), suggesting that patients with higher CISD1 expression might respond better to certain therapies and have better prognoses in these cancers. CISD1 was also significantly and positively correlated with ImmuneScore in BLCA (Fig. 7B), suggesting that higher CISD1 expression is associated with greater immune cell infiltration in the TME; while CISD1 was significantly negatively correlated with ImmuneScore in DLBC, TGCT, acute lymphocytic leukemia, LGG, GBMLGG, STAD, STES, ESCA, LUAD, THCA, and SKCM (Fig. 7B), suggesting that higher CISD1 expression is associated with reduced immune cell infiltration in the TME. CISD1 showed a significant positive correlation with ESTIMATEscore in BLCA, LAML, and PRAD (Fig. 7C), suggesting that CISD1 may contribute to a dense and complex TME. In contrast, CISD1 showed significant negative correlation with ESTIMATEscore in DLBC, TGCT, acute lymphocytic leukemia, LGG, GBMLGG, UCES, THCA, STES, COAD, LUAD, STAD, COADREAD, and SKCM (Fig. 7C), suggesting that CISD1 may be involved in creating a less dense TME. Taken together, these results suggest that CISD1 may serve as a predictive factor in determining how a tumor responds to various treatments.
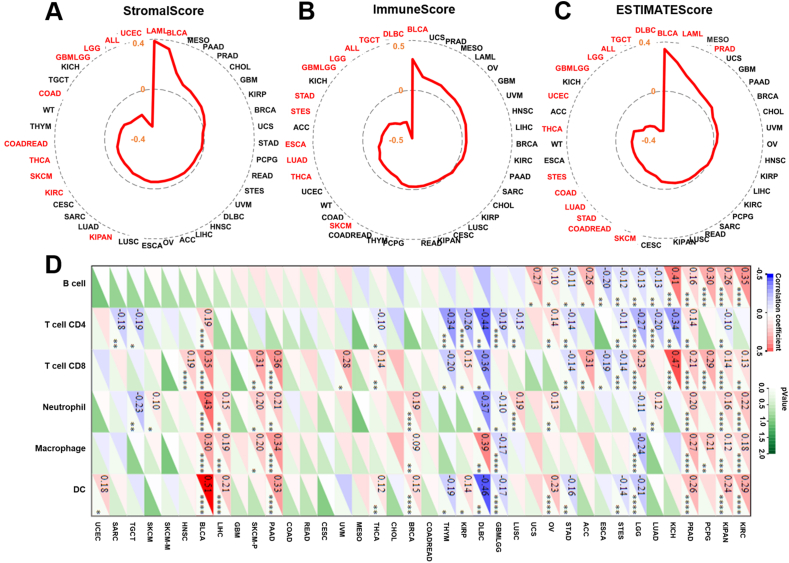
Next, we analyzed the correlation between CISD1 and immune cell infiltration in the TME, including B cells, CD4+ T cells, CD8+ T cells, neutrophils, macrophages, and dendritic cells. As is shown in Figure 7D, CISD1 was significantly and positively correlated with immune cell infiltration in KIRC, KIPAN, PCPG, PRAD, KICH, OV, BRCA, PAAD, SKCM-P (primary SKCM), LIHC, BLCA, and HNSC, suggesting that CISD1 may play a role in facilitating an effective immune response against the tumor, and tumors with higher CISD1 expression might respond better to immunotherapies in these cancers. In contrast, CISD1 was significantly and negatively correlated with immune cells infiltration in STES, ESCA, STAD, GBMLGG, DLBC, THYM, and TGCT, suggesting that CISD1 might contribute to immune suppression and tumor evasion, and tumors with high CISD1 expression may be more resistant to immunotherapies in these cancers.
Taken together, these results suggest that CISD1 may be associated with a more active immune response and potentially better outcomes in those positive correlation cancers, which makes CISD1 a potential marker of immunogenic tumors that may respond well to immunotherapy. In contrast, CISD1 might be involved in immune evasion and tumor progression in those negative correlation cancers, which makes CISD1 a potential target for enhancing immune infiltration and improving the effectiveness of cancer treatments.
CISD1 correlates with immune checkpoint blockade proteins and is a potential biomarker for tumor immunotherapy
The above results suggest that CISD1 is associated with immunotherapies. Thus, we analyzed the correlation between CISD1 and TMB, MSI, or neoantigen burden (NEO), to help understand if CISD1 is associated with generating neoantigens. Specifically, high TMB and NEO load in MSI tumors favor strong antitumor immune responses.ref. bib46,ref. bib47 Data was downloaded from SangerBox and analyzed, and Figure 8A shows that CISD1 was significantly and positively correlated with TMB in GBM, suggesting that higher CISD1 expression is associated with a higher number of mutations. This suggests that CISD1 might be a potential biomarker for predicting the efficacy of immunotherapy in GBM. CISD1 also showed significant positive correlation with MSI in DLBC, HNSC, STAD, LIHC, STES, and GBMLGG (Fig. 8B), suggesting that patients with high CISD1 expression are more likely to benefit from immunotherapies in these cancers. In contrast, CISD1 showed a significant negative correlation with MSI in LUAD, CESC, and PRAD (Fig. 8B), suggesting that patients with high CISD1 expression are less likely to respond to immunotherapies in these cancers. However, CISD1 showed no significant correlation with NEO in all tested cancers (Fig. 8C).
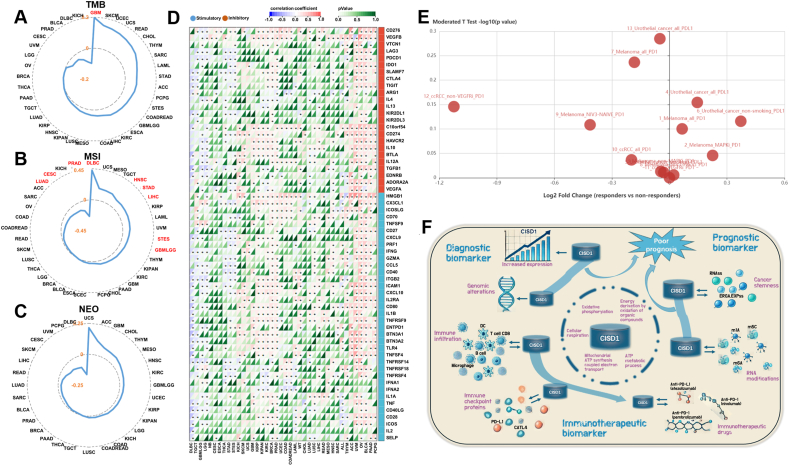
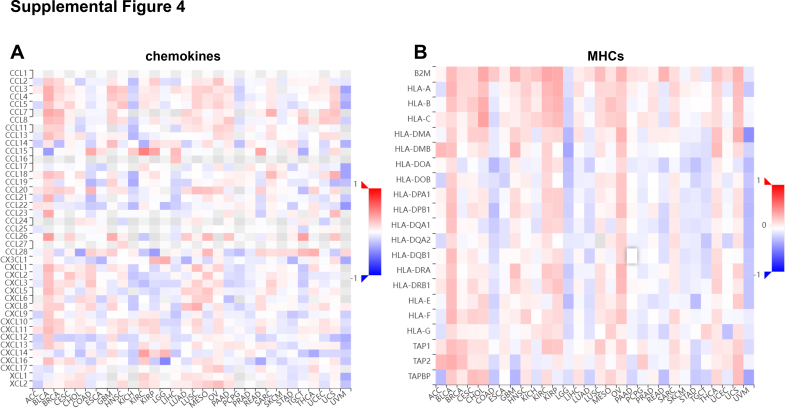
Next, we analyzed the correlation between CISD1 and immune checkpoint blockade proteins, as they are targets for immune checkpoint blockades. CISD1 was significantly and positively correlated with most of the immune checkpoint proteins in PCPG, PAAD, BLCA, OV, UVM, ACC, THYM, HNSC, LUSC, LUAD, PRAD, BRCA, KIRC, KIPAN, KIRP, GBM, and SKCM (Fig. 8D), suggesting that CISD1 might play a role in the immune evasion of tumors, making CISD1 an important biomarker for both prognosis and immunotherapies. Thus, patients with high CISD1 expression may benefit from checkpoint inhibitor therapies in these cancers. Conversely, CISD1 was significantly and negatively correlated with most of the immune checkpoint proteins in STES, STAD, THCA, LGG, GBMLGG, TGCT, and DLBC (Fig. 8D), suggesting that CISD1 might be linked to a more active immune environment within the TME of these tumors. Understanding correlations between CISD1 and immune-related genes can also provide insight into how tumors with high CISD1 expression might respond to immunotherapy. Chemokines help recruit immune cells,ref. bib48 and MHC genes play crucial roles in antigen presentation to immune cells, which is essential for initiating an immune response against tumor cells.ref. bib49 Figure S4A showed that CISD1 was positively correlated with immune chemokines in BLCA, BRCA, CESC, GBM, HNSC, LUSC, MESO, OV, SARC, THCA, and UCS, suggesting that higher CISD1 expression might promote chemokine expression, potentially influencing immune cell recruitment. Figure S4B showed that CISD1 was positively correlated with MHC genes in BLCA, BRCA, CESC, CHOL, GBM, HNSC, KIRC, KIRP, LIHC, LUSC, MESO, OV, SARC, THCA, and UCS, suggesting that high CISD1 expression is associated with up-regulated MHC gene expression, potentially enhancing antigen presentation and immune recognition.
Furthermore, we wondered if CISD1 expression differed between checkpoint blockade (e.g., anti-PDL1 and anti-PD1) responders and non-responders. To investigate this, we used TISIDB, an integrated repository portal for tumor-immune system interactions, which collects data sets of transcriptomic and genomic profiling of pre-treated tumor biopsies from immunotherapeutic responders and non-responders.ref. bib31 We used the TISIDB portal to analyze the relationship between CISD1 expression and immunotherapy response, and found that immunotherapeutic responders had higher CISD1 expression than non-responders in Melanoma_all_PD1 (pembrolizumab and nivolumab), Melanoma_MAPKi_PD1 (pembrolizumab and nivolumab), Urothelial_cancer_all_PDL1 (atezolizumab), and Urothelial_cancer_non-smoking_PDL1 (atezolizumab) (Fig. 8E and Table 1), suggesting that CISD1 is up-regulated in these responders and might serve as a prognostic marker in these cancers; while immunotherapeutic responders had lower CISD1 expression than non-responders in Melanoma_all_PD1 (nivolumab), Melanoma_NIV3-NAÏVE_PD1 (nivolumab), ccRCC_all_PD1 (nivolumab), ccRCC_non-VEGFRi_PD1 (nivolumab), and Urothelial_cancer_all_PDL1 (atezolizumab) (Fig. 8E and Table 1), suggesting that CISD1 is down-regulated in these responders and may be linked with resistance to immunotherapy.
Table 1: The difference in CISD1 expression between immunotherapeutic responders and non-responders.
| CISD1 | No. | PMID | Cancer type | Group | Drug | #Responders | #Non-responders | Log2(fold change) | p value |
|---|---|---|---|---|---|---|---|---|---|
| Up-regulated | 1 | 26997480 | Melanoma | All | Anti-PD-1 (pembrolizumaband nivolumab) | 14 | 12 | 0.066 | 0.794 |
| 2 | 26997480 | Melanoma | MAPKi | Anti-PD-1 (pembrolizumaband nivolumab) | 6 | 5 | 0.225 | 0.9 | |
| 4 | 28552987 | Urothelial cancer | All | Anti-PD-L1 (atezolizumab) | 9 | 16 | 0.145 | 0.701 | |
| 6 | 28552987 | Urothelial cancer | Non-smoking | Anti-PD-L1 (atezolizumab) | 4 | 7 | 0.372 | 0.766 | |
| Down-regulated | 7 | 29033130 | Melanoma | All | Anti-PD-1 (nivolumab) | 26 | 23 | −0.183 | 0.58 |
| 9 | 29033130 | Melanoma | NIV3-NAIVE | Anti-PD-1 (nivolumab) | 11 | 12 | −0.414 | 0.779 | |
| 10 | 29301960 | Clear cell renalcell carcinoma | All | Anti-PD-1 (nivolumab) | 4 | 8 | −0.198 | 0.919 | |
| 12 | 29301960 | Clear cell renalcell carcinoma | Non-VEGFRi | Anti-PD-1 (nivolumab) | 2 | 8 | −1.124 | 0.715 | |
| 13 | 29443960 | Urothelial cancer | All | Anti-PD-L1 (atezolizumab) | 68 | 230 | −0.05 | 0.519 |
Taken together, these results highlight the potential importance of CISD1 in cancer progression and treatment, particularly in the context of the immune response and tumor microenvironment. Given its associations with immune cell infiltration and immunotherapy response, CISD1 can be explored as a potential therapeutic target and/or a biomarker for predicting treatment outcomes in cancer patients.
Discussion
CISD1 is a multifunctional protein localized to the outer membrane of mitochondria.ref. bib9 Up-regulation of CISD1 was reported in various cancer types,ref. bib19,ref. bib20,ref. bib50 and CISD1 was shown to play crucial roles in tumor progression, resistance to apoptosis, and poor prognosis,ref. bib9,ref. bib14 making it an interesting topic of research in understanding carcinogenesis and developing therapeutic strategies. This study is the first comprehensive pan-cancer bioinformatics analysis to determine the functions of CISD1 in multiple cancers using public patient databases, and demonstrate that CISD1 is a potential diagnostic, prognostic and immunotherapeutic biomarker for multiple cancers.
The claim above is supported by the following lines of evidence. First, CISD1 can be a diagnostic biomarker to aid early detection of various cancers. Gene expression level alterations often serve as important diagnostic biomarkers for the early detection of cancer. For example, increased prostate-specific antigen (PSA) protein serves as a biomarker for early detection of prostate cancer.ref. bib51 Elevated alpha-fetoprotein (AFP) levels are used as a diagnostic marker for liver cancer and to monitor disease progression.ref. bib52 The immune checkpoint receptor programmed cell death ligand 1 (PD-L1) is overexpressed in several cancers and can be used to predict response to immune checkpoint inhibitors.ref. bib53 In this study, we found that expression levels of CISD1 were dramatically dysregulated in various cancers. GEPIA2.0 online analysis of TCGA datasets showed that mRNA levels of CISD1 were significantly up-regulated in 22 of 33 total cancer types analyzed (Fig. 1C); and the following THPA analysis showed that protein expression levels of CISD1 were also significantly increased in multiple cancers (Fig. 2). These results demonstrate that CISD1 expression is significantly associated with various cancer types. Elevated levels of CISD1 in specific cancers indicate its possible oncogenic role in tumor growth, and oncogenes play multiple, interconnected roles in cancer progression by driving cell proliferation and promoting metastasis.ref. bib54 Indeed, CISD1 is also associated with cancer stages development, as its mRNA expression levels are increased with the progress of the cancer stages and metastasis in various cancers (Fig. S1). Our analysis is consistent with previous studies that have showed CISD1 is up-regulated in several cancers.ref. bib13,ref. bib15,ref. bib16 Gene alterations can also serve as biomarkers to aid diagnosis of cancers. For instance, mutations in the BRCA1 and BRCA2 genes are strongly associated with an increased risk of breast and ovarian cancers, and genetic testing for BRCA1 and BRCA2 mutations help identify individuals at high risk.ref. bib55 Mutations in TP53 can lead to a loss of its tumor suppressive functions, contributing to cancer development, and early detection of these mutations can lead to early intervention for multiple cancer types.ref. bib56 In our study, CISD1 showed frequent gene mutations, amplifications, and deletions in multiple cancers (Fig. 3A–D). Missense mutations were the most frequent type of CISD1 gene mutations in various cancers (Fig. 3D), highlighting that the mutations of CISD1 could be targets for diagnostic assays. Copy number alterations were the most common genomic change in most of the cancers (Fig. 3A), and cancers with higher rates of CISD1 amplifications had a higher expression level of CISD1, which further confirms that CISD1 is an oncogene in multiple cancers (Fig. 3E). Moreover, expression of CISD1 was strongly associated with somatic mutations, such as TP53 mutations (Fig. 3F). All these genomic alteration profiles indicate that CISD1 is a promising diagnostic biomarker.
Second, CISD1 can be a prognostic biomarker for cancer patients. The expression level of certain oncogenes is often correlated with overall survival rates in cancer patients, and tumors with high oncogene activity often exhibit shorter overall survival and disease-free survival.ref. bib57,ref. bib58 Some mutations can also be directly associated with patient survival. For instance, mutations in tumor suppressor genes like TP53 are often associated with a worse prognosis, as they may lead to more aggressive tumor behavior and poorer overall outcomes.ref. bib56 Our study validated that CISD1 could be a reliable biomarker for cancer prognosis, as patients with higher expression levels of CISD1 had a lower survival probability from the overall survival analysis in multiple cancers (Fig. 4). Moreover, patients with higher expression levels of CISD1 had a higher risk of death from analyses of disease-specific survival, disease-free survival, and progression-free survival (Fig. S2). Our results are also consistent with previous reports that CISD1 is a prognostic biomarker in several cancers.ref. bib18,ref. bib19 This reproducible and significant correlation between CISD1 expression and patient survival across various cancers strengthens its use as a prognostic biomarker in clinical settings. Stemness-related signatures can also be used to predict patient outcomes. High levels of stemness in tumors are often associated with a poor prognosis, as these tumors may be more aggressive and likely to metastasize.ref. bib59 Our study revealed that CISD1 was significantly and positively correlated with stemness indices in multiple cancers (Fig. 5A–F), indicating that CISD1 is involved in promoting or maintaining stem cell-like properties in cancer cells. Stemness is a key feature of cancer aggressiveness, treatment resistance, and metastatic potential. The strong correlation between CISD1 and stemness indices underscores its potential role in driving tumor aggressiveness and therapeutic resistance, and high CISD1 expression may serve as an indicator of tumors with enriched cancer stem cell populations, predicting more aggressive disease and reduced therapeutic efficacy. By modulating cancer stem cell characteristics, CISD1 not only contributes to tumor aggressiveness but also presents as a promising target for innovative therapies aimed at eradicating cancer stem cell-driven disease. Therefore, the correlation could have profound implications for prognosis.ref. bib60 Indeed, patients with high expression of CISD1 (Fig. 1) and high levels of stemness (Fig. 5) have worse prognosis (Fig. 4), such as BRCA, LIHC, LUAD, SKCM, and THYM in our study. Given the association between cancer stem cell-related stemness indices and poor patient outcomes, the correlation with CISD1 highlights its potential as a prognostic biomarker. Targeting CISD1 in cancers with high stemness signatures may offer a promising therapeutic avenue. In addition to stemness, CISD1 is also associated with epigenetic regulation in multiple cancers in our study. Figure 5G showed strong positive correlations between CISD1 and RNA modifications like m1A, m5C, and m6A, indicating that CISD1 could serve as a biomarker for cancer prognosis, as high CISD1 expression, along with altered RNA modifications, might be associated with more aggressive tumor behavior and could predict poor outcomes in certain cancers including KICH, BRCA, LIHC, LUAD, and SKCM.
Understanding the biological functions and pathways associated with CISD1 can potentiate its prognostic value. Our study showed that 26 genes are coexpressed with CISD1 across five types of cancer that have high expression levels of CISD1, high stemness signatures, high levels of RNA modifications, and worse patient survival (Fig. 6A), and 20 of them are highly expressed across a wide range of cancers (Fig. 6B). GO and KEGG enrichment analysis showed that these CISD1 coexpression genes were involved in critical pathways in energy metabolism including oxidative phosphorylation, mitochondrial electron transport, and cellular respiration (Fig. 6C–G). These pathways are often altered in cancer cells to support rapid growth and survival. Our findings are consistent with the previous report that CISD1 plays critical roles in bioenergetics.ref. bib10 Indeed, CISD1 showed a negative correlation with age in multiple cancers (Fig. S3A), indicating that CISD1 expression is higher in younger individuals and tends to decrease in older individuals. The decline in CISD1 expression with age may reflect its role in maintaining cellular bioenergetics.
Third, CISD1 can be an immunotherapeutic biomarker indicating the likelihood of a patient responding to immunotherapy. Immunotherapy has revolutionized cancer treatment by harnessing the immune system to target and destroy cancer cells. Unlike traditional therapies, immunotherapy provides a more personalized approach, with varying outcomes based on patient-specific factors. Certain factors have been identified as immunotherapeutic biomarkers to predict the outcomes of immunotherapy in cancer treatment. For example, high expression of PD-L1 in tumors often correlates with better responses to programmed death-1 (PD-1)/PD-L1 inhibitors such as nivolumab or pembrolizumab.ref. bib61 High TMB or MSI has been reported to link to better response to immune checkpoint inhibitors.ref. bib62,ref. bib63 Further analysis showed that CISD1 expression was positively correlated with immune cells in multiple cancers (Fig. 7D), indicating that tumors with high expression of CISD1 have high immune infiltration. The degree of immune infiltration in a tumor is a critical factor in determining the success of immunotherapy treatments. Indeed, CISD1 is significantly positively correlated with TMB and MSI in several cancers (Fig. 8A, B), indicating that patients with high expression of CISD1, together with high TMB and MSI, can benefit from immunotherapy, as a high TMB or MSI has been linked to better response to immune checkpoint inhibitors.ref. bib62,ref. bib63 Importantly, immune checkpoint analysis showed that CISD1 was significantly and positively correlated with immune checkpoint genes such as programmed cell death 1 (PDCD-1; PD-1 coding gene), CD274 (PD-L1 coding gene), or cytotoxic T-lymphocyte associated protein 4 (CTLA4) in multiple cancers (Fig. 8D), indicating that tumors with high expression of CISD1 have a high expression of immune checkpoint proteins, which serve as targets for immune checkpoint blockades. This finding aligns with a recent study identifying CISD1 as a ferroptosis-related gene in breast cancer, which similarly showed significant correlations between CISD1 and immune checkpoints, including PDCD-1, CTLA4, and lymphocyte activating 3 (LAG3).ref. bib64 Although no direct experimental evidence has yet established a regulatory relationship between CISD1 and these immune checkpoints, CISD1’s association with mitophagy and hypoxia, alongside evidence linking immune checkpoints to mitophagy and tumor hypoxia, suggests an important connection among the three. For example, while inhibition of CISD1 induces mitophagy mediated by Parkin,ref. bib65,ref. bib66 Parkin deficiency (a key mitophagy mediator) reduces mitophagy in the liver, increases PD-1 and CTLA4 expression, creates an immunosuppressive microenvironment, and promotes hepatocarcinogenesis.ref. bib67 Moreover, PD-L1 expression increases under tumor hypoxia, contributing to immune evasion and tumor progression.ref. bib68 Notably, tumor samples from breast cancer patients who responded to combined immunotherapy with immune checkpoint inhibitors and paclitaxel displayed significant PD-L1 distribution in mitochondria.ref. bib69 Taken together, these findings support our speculation that high CISD1 expression may suppress mitophagy, help counteract hypoxia, and maintain a growth advantage for tumor cells, while simultaneously up-regulating immune checkpoints to promote immune evasion. As these elevated immune checkpoints serve as effective immunotherapeutic targets, CISD1 overexpression could be a reliable biomarker for immunotherapy. Furthermore, using published clinical data, we found that CISD1 expression was higher in immunotherapy responders versus non-responders (Fig. 8E). Thus, increased expression of CISD1 can be used to predict response to immune checkpoint inhibitors in cancers. These results strongly support that CISD1 can be a great immunotherapeutic biomarker.
Studies have shown that CISD1 regulates the levels of iron and reactive oxygen species in cancer cells.ref. bib8 It also inhibits mitophagy, apoptosis, and ferroptosis in cancer cells, highlighting its central role in promoting cancer cell proliferation, supporting tumor growth, and facilitating metastasis.ref. bib8 Our pan-cancer analysis further confirms that CISD1 is significantly overexpressed in various cancer types, conferring a survival advantage to cancer cells. These findings underscore the potential of CISD1 as a highly effective target in cancer therapy. Since the function of mitoNEET (CISD1) depends on the relative instability of its [2Fe–2S] cluster and cluster transfer reactions,ref. bib8 drugs that stabilize its iron-sulfur cluster could serve as promising anticancer agents, with pioglitazone being one such example. MitoNEET (CISD1) was first identified as a binding target for pioglitazone,ref. bib70 and pioglitazone binding stabilizes the iron-sulfur cluster of mitoNEET and prevents its transfer.ref. bib71,ref. bib72 Treatment of breast cancer cells with pioglitazone leads to increased mitochondrial iron and reactive oxygen species levels, reducing the cells’ tolerance to oxidative stress and subsequently inducing cancer cell death.ref. bib13,ref. bib73 In human hepatocellular carcinoma cells, pioglitazone inhibits mitochondrial iron uptake and lipid peroxidation, further preventing ferroptosis by stabilizing the CISD1 iron-sulfur cluster.ref. bib74 Building on the success of pioglitazone, novel compounds have been developed to target CISD1 in cancer. For instance, mitoNEET ligand-1 (NL-1), a derivative of pioglitazone designed to target drug-resistant leukemia cells, demonstrates potent anti-cancer activity by stabilizing mitoNEET’s iron-sulfur cluster.ref. bib16 Another promising compound, NTS-01, specifically binds to human mitoNEET protein, stabilizes its [2Fe–2S] cluster under oxidative conditions in vitro, and induces mitochondrial fission, significantly reducing ovarian cancer cell proliferation.ref. bib75 Our pan-cancer analysis highlights CISD1 as not only a reliable biomarker but also a highly promising anti-cancer target due to its overexpression in multiple cancer types. The development of novel therapeutics targeting CISD1’s iron-sulfur cluster or modulating its protein expression holds great potential for improving cancer outcomes.
In addition to CISD1 up-regulation in the vast majority of cancers analyzed, we observed that CISD1 expression was down-regulated in six types of cancer (Fig. 1C), suggesting that it may play a different role in these cancers, potentially acting as a tumor suppressor. Cancer is highly heterogeneous, meaning that the genetic mutations, signaling pathways, and microenvironments vary significantly between cancers, and this heterogeneity can influence how a gene functions.ref. bib76 Genes can play opposing roles in different cancers due to the unique molecular and cellular contexts of each cancer type. For example, the well-known tumor suppressor gene TP53 can lose its tumor-suppressive function and even gain oncogenic functions in certain cancer types.ref. bib56 Depending on the cancer type, NOTCH can act as an oncogene (e.g., in T-cell acute lymphoblastic leukemia) or as a tumor suppressor (e.g., in skin cancer).ref. bib77 Furthermore, CISD1’s expression in some cancers was negatively correlated with tumor indices (Fig. 5A–E), RNA modification enzymes (Fig. 5G), immune cell infiltration (Fig. 7D), or immune checkpoint proteins (Fig. 8D). The relationship between CISD1 and these cancer-related factors can differ across cancer types due to variations in tumor genetic and epigenetic context and tumor microenvironment, as each cancer type has a unique set of genetic alterations and epigenetic modifications that can modulate how a gene interacts with other molecules and signaling pathways,ref. bib78 and tumor microenvironment can alter gene function and expression, leading to different effects of a gene.ref. bib79 A dual role of CISD1 in cancer is not unusual, as many genes exhibit context-dependent functionality.ref. bib56,ref. bib77 This flexibility is shaped by factors such as the specific cancer type, tumor microenvironment, and genetic background. Even with opposing expression levels in different cancers, CISD1 can still provide meaningful diagnostic, prognostic, or predictive information from our findings. CISD1 serving as a biomarker provides valuable information relevant to specific clinical outcomes in our study.
We acknowledge that this study relies on computational analyses based on publicly available datasets, which, while robust and widely used, have inherent limitations. The lack of experimental validation of our findings is a significant limitation, as computational predictions require laboratory-based confirmation to establish causal relationships and functional mechanisms. Furthermore, the dataset heterogeneity, including variations in data processing and normalization across platforms, may introduce biases that could influence the results. Despite these limitations, our integrative analysis provides valuable insights into the potential roles of CISD1 in cancer biology and identifies it as a promising biomarker and therapeutic target. This systematic pan-cancer analysis lays a strong foundation for further exploring the biological functions of CISD1 in cancers. Future studies will focus on experimental validation of CISD1’s role in tumor aggressiveness, stemness, and immunotherapy, and exploration of its molecular mechanisms through in vitro and in vivo experiments.
In summary, our systematic pan-cancer analysis of CISD1, for the first time, shows that its expression is significantly altered in the vast majority of cancers analyzed. Additionally, it undergoes mutations in multiple cancers, suggesting its potential as a diagnostic biomarker for cancers. Moreover, cancer patients with significantly altered expression of CISD1 have lower survival rates and worse prognoses, and its expression is significantly correlated with tumor stemness indices in multiple cancers, highlighting its importance prognostic prediction as a crucial prognostic biomarker. More importantly, it can serve as a biomarker for predicting cancer patients’ response to immunotherapy, as its expression is not only significantly correlated with TMB and MSI but also with immune checkpoints, and its expression is altered in tumors of patients who respond to immunotherapy. In conclusion, all these findings strongly demonstrate that CISD1 is a reliable and promising diagnostic, prognostic, and immunotherapeutic biomarker in multiple cancers (Fig. 8F).
CRediT authorship contribution statement
Caiyue Li: Writing – review & editing, Writing – original draft, Visualization, Validation, Software, Resources, Methodology, Formal analysis, Data curation, Conceptualization. Zhipin Liang: Writing – review & editing. Gabrielle Vontz: Writing – review & editing. Connor Kent: Writing – review & editing. Wenbo Ma: Writing – review & editing, Methodology. Lei Liu: Writing – review & editing. Riya Dahal: Writing – review & editing. Jovanny Zabaleta: Writing – review & editing. Guoshuai Cai: Writing – review & editing, Methodology. Jia Zhou: Writing – review & editing, Resources, Project administration, Funding acquisition. Huangen Ding: Writing – review & editing, Resources, Project administration, Funding acquisition. Qiang Shen: Writing – review & editing, Supervision, Resources, Project administration, Funding acquisition, Conceptualization.
Funding
This project was supported by the Louisiana State University Health Sciences Center (LSUHSC) Startup funds (New Orleans, Louisiana, USA) (to Q.S.), Louisiana State University (LSU) Interinstitutional Cancer Research Funding Initiative (Louisiana, USA) (CCRI; to H.D. and Q.S.), Louisiana Cancer Research Center (LCRC) Strategic Investment in Translational Research Awards (Louisiana, USA) (to Q.S. and H.D.), the John D. Stobo, M.D. Distinguished Chair Endowment (USA) (to J.Z.), and the Edith & Robert Zinn Chair Endowment in Drug Discovery (USA) (to J.Z.).
Conflict of interests
The authors declared no conflict of interests.
References
- Q. Li, C. Xia, H. Li. Disparities in 36 cancers across 185 countries: secondary analysis of global cancer statistics. Front Med, 2024. [PubMed]
- S. Das, M.K. Dey, R. Devireddy, M.R. Gartia. Biomarkers in cancer detection, diagnosis, and prognosis. Sensors (Basel), 2023. [PubMed]
- T.Y. Jin, K.S. Park, S.E. Nam, Y.B. Yoo, W.S. Park, I.J. Yun. BRCA1/2 serves as a biomarker for poor prognosis in breast carcinoma. Int J Mol Sci, 2022. [PubMed]
- K. Kamposioras, P.C. Dinas, J. Barriuoso, V. Trachana, K. Dimas. Caveolin-1 protein expression as a prognostic biomarker of gastrointestinal tumours: a systematic review and meta-analysis. Eur J Clin Invest, 2023
- J.C. Ahn, P.C. Teng, P.J. Chen. Detection of circulating tumor cells and their implications as a biomarker for diagnosis, prognostication, and therapeutic monitoring in hepatocellular carcinoma. Hepatology, 2021. [PubMed]
- S. Sumbal, A. Javed, B. Afroze. Circulating tumor DNA in blood: future genomic biomarkers for cancer detection. Exp Hematol, 2018. [PubMed]
- W.J. Geldenhuys, R. Skolik, M.E. Konkle, M.A. Menze, T.E. Long, A.R. Robart. Binding of thiazolidinediones to the endoplasmic reticulum protein nutrient-deprivation autophagy factor-1. Bioorg Med Chem Lett, 2019. [PubMed]
- R. Mittler, M. Darash-Yahana, Y.S. Sohn. NEET proteins: a new link between iron metabolism, reactive oxygen species, and cancer. Antioxidants Redox Signal, 2019
- S. Tamir, M.L. Paddock, M. Darash-Yahana-Baram. Structure-function analysis of NEET proteins uncovers their role as key regulators of iron and ROS homeostasis in health and disease. Biochim Biophys Acta, 2015. [PubMed]
- W.J. Geldenhuys, T.C. Leeper, R.T. Carroll. mitoNEET as a novel drug target for mitochondrial dysfunction. Drug Discov Today, 2014. [PubMed]
- H. Tasnim, A.P. Landry, C.R. Fontenot, H. Ding. Exploring the FMN binding site in the mitochondrial outer membrane protein mitoNEET. Free Radic Biol Med, 2020. [PubMed]
- Y. Wang, A.P. Landry, H. Ding. The mitochondrial outer membrane protein mitoNEET is a redox enzyme catalyzing electron transfer from FMNH2 to oxygen or ubiquinone. J Biol Chem, 2017. [PubMed]
- Y.S. Sohn, S. Tamir, L. Song. NAF-1 and mitoNEET are central to human breast cancer proliferation by maintaining mitochondrial homeostasis and promoting tumor growth. Proc Natl Acad Sci USA, 2013. [PubMed]
- A.F. Salem, D. Whitaker-Menezes, A. Howell, F. Sotgia, M.P. Lisanti. Mitochondrial biogenesis in epithelial cancer cells promotes breast cancer tumor growth and confers autophagy resistance. Cell Cycle, 2012. [PubMed]
- F. Bai, F. Morcos, Y.S. Sohn. The Fe-S cluster-containing NEET proteins mitoNEET and NAF-1 as chemotherapeutic targets in breast cancer. Proc Natl Acad Sci U S A, 2015. [PubMed]
- W.J. Geldenhuys, R.R. Nair, D. Piktel, K.H. Martin, L.F. Gibson. The MitoNEET ligand NL-1 mediates antileukemic activity in drug-resistant B-cell acute lymphoblastic leukemia. J Pharmacol Exp Therapeut, 2019
- X. Liu, Q. Cui. Identification of CISD1 as a prognostic biomarker for breast cancer. Int J Gen Med, 2022. [PubMed]
- F. Liu, Y. Dong, F. Zhong, H. Guo, P. Dong. CISD1 is a breast cancer prognostic biomarker associated with diabetes mellitus. Biomolecules, 2022. [PubMed]
- T. Lu, C. Li, C. Xiang, Y. Gong, W. Peng, C. Chen. Overexpression of CISD1 predicts worse survival in hepatocarcinoma patients. BioMed Res Int, 2022
- J. Zang, M. Cui, L. Xiao, J. Zhang, R. Jing. Overexpression of ferroptosis-related genes FSP1 and CISD1 is related to prognosis and tumor immune infiltration in gastric cancer. Clin Transl Oncol, 2023. [PubMed]
- Y. Yang, H. Hu, L. Chen, H. Zhang, J. Yang. A new survival model based on ferroptosis-related genes (FRGS) for prognostic prediction in bladder cancer. Actas Urol Esp, 2022. [PubMed]
- G. Consortium. The genotype-tissue expression (GTEx) project. Nat Genet, 2013. [PubMed]
- P.J. Thul, C. Lindskog. The human protein atlas: a spatial map of the human proteome. Protein Sci, 2018. [PubMed]
- Z. Tang, B. Kang, C. Li, T. Chen, Z. Zhang. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res, 2019. [PubMed]
- W. Shen, Z. Song, X. Zhong. Sangerbox: a comprehensive, interaction-friendly clinical bioinformatics analysis platform. Imeta, 2022. [PubMed]
- E. Cerami, J. Gao, U. Dogrusoz. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov, 2012. [PubMed]
- T. Li, J. Fu, Z. Zeng. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res, 2020. [PubMed]
- A. Lin, C. Qi, T. Wei. CAMOIP: a web server for comprehensive analysis on multi-omics of immunotherapy in pan-cancer. Briefings Bioinf, 2022
- B.T. Sherman, M. Hao, J. Qiu. DAVID a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res, 2022. [PubMed]
- D. Tang, M. Chen, X. Huang. SRplot: a free online platform for data visualization and graphing. PLoS One, 2023
- B. Ru, C.N. Wong, Y. Tong. TISIDB: an integrated repository portal for tumor-immune system interactions. Bioinformatics, 2019. [PubMed]
- S. Chakraborty, M. Balan, A. Sabarwal, T.K. Choueiri, S. Pal. Metabolic reprogramming in renal cancer: events of a metabolic disease. Biochim Biophys Acta Rev Cancer, 2021
- H. Wang, M. Guo, H. Wei, Y. Chen. Targeting p53 pathways: mechanisms, structures, and advances in therapy. Signal Transduct Targeted Ther, 2023
- L.J. van ‘t Veer, H. Dai, M.J. van de Vijver. Gene expression profiling predicts clinical outcome of breast cancer. Nature, 2002. [PubMed]
- N. Fatima, A. Zaman, S.M. Azam, W. Shahid, M.U. Zaman. Predictors for disease-free and progression-free survivals in metabolic responders and non-responder on follow-up 18FDG PET/CT after chemoradiation in patients with nasopharyngeal cancer. Asian Pac J Cancer Prev APJCP, 2024. [PubMed]
- X. Chu, W. Tian, J. Ning. Cancer stem cells: advances in knowledge and implications for cancer therapy. Signal Transduct Targeted Ther, 2024
- K. Li, Q. Wang, H. Bian. Comprehensive analysis reveals USP45 as a novel putative oncogene in pan-cancer. Front Mol Biosci, 2022
- B. Zhang, Y. Hao, H. Liu. Interplay of RNA m6A modification-related geneset in pan-cancer. Biomedicines, 2024. [PubMed]
- I. Barbieri, T. Kouzarides. Role of RNA modifications in cancer. Nat Rev Cancer, 2020. [PubMed]
- M. Ashburner, C.A. Ball, J.A. Blake. Gene ontology: tool for the unification of biology. The gene ontology consortium. Nat Genet, 2000. [PubMed]
- M. Kanehisa, S. Goto. KEGG Kyoto encyclopedia of genes and genomes. Nucleic Acids Res, 2000. [PubMed]
- Y. Ruan, H. Ogana, E. Gang, H.N. Kim, Y.M. Kim. Wnt signaling in the tumor microenvironment. Adv Exp Med Biol, 2021. [PubMed]
- H. Sadeghi Rad, J. Monkman, M.E. Warkiani. Understanding the tumor microenvironment for effective immunotherapy. Med Res Rev, 2021. [PubMed]
- Y. Zhang, Z. Zhang. The history and advances in cancer immunotherapy: understanding the characteristics of tumor-infiltrating immune cells and their therapeutic implications. Cell Mol Immunol, 2020. [PubMed]
- K. Yoshihara, M. Shahmoradgoli, E. Martínez. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun, 2013. [PubMed]
- E. Picard, C.P. Verschoor, G.W. Ma, G. Pawelec. Relationships between immune landscapes, genetic subtypes and responses to immunotherapy in colorectal cancer. Front Immunol, 2020. [PubMed]
- A.M. Goodman, E.S. Sokol, G.M. Frampton, S.M. Lippman, R. Kurzrock. Microsatellite-stable tumors with high mutational burden benefit from immunotherapy. Cancer Immunol Res, 2019. [PubMed]
- A. Zlotnik, O. Yoshie. The chemokine superfamily revisited. Immunity, 2012. [PubMed]
- J. Neefjes, M.L.M. Jongsma, P. Paul, O. Bakke. Towards a systems understanding of MHC class I and MHC class II antigen presentation. Nat Rev Immunol, 2011. [PubMed]
- L. Yuan, J. Liu, L. Bao, H. Qu, J. Xiang, P. Sun. Upregulation of the ferroptosis-related STEAP3 gene is a specific predictor of poor triple-negative breast cancer patient outcomes. Front Oncol, 2023
- W.J. Catalona, D.S. Smith, T.L. Ratliff. Measurement of prostate-specific antigen in serum as a screening test for prostate cancer. N Engl J Med, 1991. [PubMed]
- F. Trevisani, P.E. D’Intino, A.M. Morselli-Labate. Serum alpha-fetoprotein for diagnosis of hepatocellular carcinoma in patients with chronic liver disease: Influence of HBsAg and anti-HCV status. J Hepatol, 2001. [PubMed]
- R.S. Herbst, J.C. Soria, M. Kowanetz. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature, 2014. [PubMed]
- S.Y. Lee, M.K. Ju, H.M. Jeon. Oncogenic metabolism acts as a prerequisite step for induction of cancer metastasis and cancer stem cell phenotype. Oxid Med Cell Longev, 2018
- P. Pourmasoumi, A. Moradi, M. Bayat. BRCA1/2 mutations and breast/ovarian cancer risk: a new insights review. Reprod Sci, 2024. [PubMed]
- R. Brosh, V. Rotter. When mutants gain new powers: news from the mutant p53 field. Nat Rev Cancer, 2009. [PubMed]
- M.J. Duffy. Cellular oncogenes and suppressor genes as prognostic markers in cancer. Clin Biochem, 1993. [PubMed]
- K.I. Bland, M.M. Konstadoulakis, M.P. Vezeridis, H.J. Wanebo. Oncogene protein co-expression. Value of Ha-ras, c-myc, c-fos, and p53 as prognostic discriminants for breast carcinoma. Ann Surg, 1995. [PubMed]
- E. Batlle, H. Clevers. Cancer stem cells revisited. Nat Med, 2017. [PubMed]
- L.T.H. Phi, I.N. Sari, Y.G. Yang. Cancer stem cells (CSCs) in drug resistance and their therapeutic implications in cancer treatment. Stem Cell Int, 2018
- S.L. Topalian, F.S. Hodi, J.R. Brahmer. Safety, activity, and immune correlates of anti-PD-1 antibody in cancer. N Engl J Med, 2012. [PubMed]
- M. Palmeri, J. Mehnert, A.W. Silk. Real-world application of tumor mutational burden-high (TMB-high) and microsatellite instability (MSI) confirms their utility as immunotherapy biomarkers. ESMO Open, 2022
- T. Sakakida, T. Ishikawa, T. Doi. Genomic profile and clinical features of MSI-H and TMB-high pancreatic cancers: real-world data from C-CAT database. J Gastroenterol, 2024. [PubMed]
- Z.H. Wu, Y. Tang, H. Yu, H.D. Li. The role of ferroptosis in breast cancer patients: a comprehensive analysis. Cell Death Discov, 2021. [PubMed]
- S. Lee, S. Lee, S.J. Lee, S.W. Chung. Inhibition of mitoNEET induces Pink1-parkin-mediated mitophagy. BMB Rep, 2022. [PubMed]
- T. Zhang, M. Zhang. NL-1 promotes PINK1-parkin-mediated mitophagy through MitoNEET inhibition in subarachnoid hemorrhage. Neurochem Res, 2024. [PubMed]
- G. Ye, X. Sun, J. Li, M. Pu, J. Zhang. Parkin modulates the hepatocellular carcinoma microenvironment by regulating PD-1/PD-L1 signalling. J Adv Res, 2025
- L. Svajda, M. Cserepes, B. Hegyi, T. Niczky, J. Tóvári. Immunomodulation in the tumor microenvironment: therapeutic potential of combined inhibition of tumor hypoxia and PD-1/PD-L1. Magy Onkol, 2024. [PubMed]
- X.Q. Xie, Y. Yang, Q. Wang. Targeting ATAD3A-PINK1-mitophagy axis overcomes chemoimmunotherapy resistance by redirecting PD-L1 to mitochondria. Cell Res, 2023. [PubMed]
- J.R. Colca, W.G. McDonald, D.J. Waldon. Identification of a novel mitochondrial protein ("mitoNEET") cross-linked specifically by a thiazolidinedione photoprobe. Am J Physiol Endocrinol Metab, 2004. [PubMed]
- M.L. Paddock, S.E. Wiley, H.L. Axelrod. MitoNEET is a uniquely folded 2Fe–2S outer mitochondrial membrane protein stabilized by pioglitazone. Proc Natl Acad Sci USA, 2007. [PubMed]
- J.A. Zuris, Y. Harir, A.R. Conlan. Facile transfer of [2Fe-2S] clusters from the diabetes drug target mitoNEET to an apo-acceptor protein. Proc Natl Acad Sci U S A, 2011. [PubMed]
- M. Darash-Yahana, Y. Pozniak, M. Lu. Breast cancer tumorigenicity is dependent on high expression levels of NAF-1 and the lability of its Fe-S clusters. Proc Natl Acad Sci USA, 2016. [PubMed]
- H. Yuan, X. Li, X. Zhang, R. Kang, D. Tang. CISD1 inhibits ferroptosis by protection against mitochondrial lipid peroxidation. Biochem Biophys Res Commun, 2016. [PubMed]
- H.B. Marjault, S. Yang-Sung, K. Zuo, P. Carloni, R. Mittler, R. Nechushtai. Structure-based screening reveals a ligand that stabilizes the [2Fe-2S] clusters of human mitoNEET and reduces ovarian cancer cell proliferation. J Phys Chem B, 2022. [PubMed]
- A. Marusyk, K. Polyak. Tumor heterogeneity: causes and consequences. Biochim Biophys Acta, 2010. [PubMed]
- C. Lobry, P. Oh, I. Aifantis. Oncogenic and tumor suppressor functions of notch in cancer: it’s NOTCH what you think. J Exp Med, 2011. [PubMed]
- H. Shen, P.W. Laird. Interplay between the cancer genome and epigenome. Cell, 2013. [PubMed]
- M.R. Junttila, F.J. de Sauvage. Influence of tumour micro-environment heterogeneity on therapeutic response. Nature, 2013. [PubMed]
