Clinicopathological Significance and Prognostic Values of Long Noncoding RNA BCYRN1 in Cancer Patients: A Meta-Analysis and Bioinformatics Analysis
Abstract
Background:
Although combination therapies have substantially improved the clinical outcomes of cancer patients, the prognosis and early diagnosis remain unsatisfactory. As a result, it is critical to look for novel indicators linked to cancer. Despite a number of recent studies indicating that the lncRNA brain cytoplasmic RNA1(BCYRN1) may be a potential predictive biomarker in cancer patients, BCYRN1‘s prognostic value is still being debated.
Methods:
We utilized PubMed, Embase, Web of Science, and the Cochrane Library to search for studies related to BCYRN1 until October 2021. Valid data were extracted after determining the articles according to the inclusion and exclusion criteria, and forest plots were made using Stata software. We used hazard ratios (HRs) or odds ratios (ORs) with 95% confidence intervals to evaluate the relationship between abnormal BCYRN1 expression and patient prognosis and clinicopathological characteristics.
Results:
Meta-analysis revealed that increased BCYRN1 expression was associated with both overall tumor survival (OS; HR = 1.84, 95% CI 1.51–2.25, p < 0.0001) and disease-free survival (DFS; HR = 1.65, 95% CI 1.20–2.26, p=0.002). Furthermore, a strong association was discovered between increased BCYRN1 expression and tumor invasion depth (OR = 2.11, 95% CI 1.49–2.99, p=0.000), clinical stage (OR = 2.52, 95% CI 1.18–5.37, p=0.017), and distant tumor metastasis (OR = 4.19, 95% CI 1.45–12.05, p=0.008).
Conclusions:
We found that high BCYRN1 expression was associated with poor survival prognosis and aggressive clinicopathological characteristics in various cancers, indicating that it is a potential prognostic indicator as well as a therapeutic target. Further research is needed on pan-cancer cohorts to determine the clinical relevance of BCYRN1 in distinct cancer types.
Affiliations: Graduate School, Ning Xia Medical University, Yinchuan 750004, Ning Xia, China; General Surgery Clinical Medical Center, Gansu Provincial Hospital, Lanzhou 730000, Gansu, China; Key Laboratory of Molecular Diagnostics and Precision Medicine for Surgical Oncology in Gansu Province, Gansu Provincial Hospital, Lanzhou 730000, Gansu, China; Gansu University of Chinese Medicine, First Clinical Medical College, Lanzhou 730000, Gansu, China; The First Clinical Medical College of Lanzhou University, Lanzhou 730000, Gansu, China; Evidence-Based Medicine Center, School of Basic Medical Sciences, Lanzhou University, Lanzhou 730000, China; NHC Key Laboratory of Diagnosis and Therapy of Gastrointestinal Tumor, Gansu Provincial Hospital, Lanzhou 730000, China
License: Copyright © 2022 Xiaoyong Han et al. CC BY 4.0 This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Article links: DOI: 10.1155/2022/8903265 | PubMed: 35874631 | PMC: PMC9303157
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (1.1 MB)
1. Introduction
Since the turn of the new century, cancer incidence and mortality have gradually exceeded that of other chronic diseases [ref. 1]. According to the most recent CA, a cancer journal, estimates, 1.9 million new cases of cancer were diagnosed in the United States in 2021, with an estimated 610,000 deaths [ref. 2]. Although combination therapies, such as surgery, chemotherapy, radiation therapy, targeted therapy, and immunotherapy, have substantially improved the clinical outcomes of cancer patients, the prognosis and early diagnosis remain unsatisfactory [ref. 3]. In recent years, advances in screening techniques, targeted therapies, immunotherapies, bioinformatics, and cancer biology have identified novel biomarkers for early diagnosis and prognosis prediction [ref. 4], and novel tumor marker detection techniques are important [ref. 5–ref. 8].
Long non-coding RNAs (lncRNAs) have gained considerable interest as cancer biomarkers in recent years with the advent of next-generation sequencing technologies [ref. 9]. The lncRNAs have a length of more than 200 nucleotides and lack any protein-coding activity [ref. 10]. They regulate gene expression at the transcriptional level through chromatin remodeling and miRNA sponging, at the post-transcriptional level by affecting RNA splicing and stability and at the translational level by controlling signal transmission [ref. 11, ref. 12]. Studies increasingly show that aberrant lncRNA expression is linked to biological processes such as tumor growth, angiogenesis, metastasis, and invasion, and lncRNAs can be exploited as tumor suppressor genes or oncogenes for cancer therapy and prevention [ref. 12, ref. 13].
The lncRNA brain cytoplasmic 200 (BC200), also known as brain cytoplasmic RNA1 (BCYRN1), is normally expressed in neurons and is implicated in cancer and neurological diseases [ref. 14]. Studies show that BCYRN1 is overexpressed in non-small-cell lung cancer [ref. 15–ref. 17], hepatocellular carcinoma [ref. 18–ref. 20], colorectal cancer [ref. 21–ref. 24], bladder cancer [ref. 25, ref. 26], esophageal squamous cell carcinoma [ref. 27, ref. 28], gastric cancer [ref. 29, ref. 30], cervical cancer [ref. 31], ovarian cancer [ref. 32], and breast cancer [ref. 33] tissues compared to matched normal tissues. However, no systematic review has been conducted so far on the pan-cancer data of BCYRN1. To this end, we performed a meta-analysis of the relevant studies to further evaluate whether BCYRN1 is a reliable prognostic biomarker and therapeutic target for different cancers by evaluating the correlation between BCYRN1 expression levels and cancer-related clinicopathological features and patient prognosis. Finally, the clinicopathological and prognostic value of BCYRN1 in cancer patients was validated by bioinformatics analysis of cancer databases.
2. Materials and Methods
2.1. Search Strategy for Literature
All procedures mentioned below were performed in accordance with PRISMA Checklist protocols [ref. 34]. Prior to October 1, 2021, PubMed, Web of Science, Embase, and the Cochrane Library were used to search for relevant papers studying the association between lncRNA BCYRN1 expression and clinical outcomes in cancer patients. Medical Subject Headings (MESH) keywords and free terms were merged in this search. Our search keywords are as follows: (“LncRNA” OR “Long non-coding RNA”) AND (“BCYRN1” OR “BC200” OR “BC200a” OR “LINC00004” OR “NCRNA00004” OR “Brain cytoplasmic RNA1”) AND (“Neoplasms” OR “Carcinoma” OR “Tumor” OR “Cancer”). To guarantee accuracy and consistency, two writers independently assessed the database search approach and discussed the results.
2.2. Inclusion and Exclusion Criteria
The duplicate articles were first eliminated, and the titles and abstracts of the remaining studies were screened on the basis of the following inclusion criteria: (1) patients with histopathologically proven cancer; (2) analysis of cancer tissues and adjacent normal tissues; (3) detection of BCYRN1 levels by qRT-PCR; (4) the paper included clinical factors such as age, gender, tumor size, TNM stage, clinical stage, lymph node metastasis, or distant metastases, as well as prognostic markers such as overall survival (OS), disease-free survival (DFS), or progression-free survival (PFS); (5) demarcation of patients into BCYRN1 low and BCYRN1 high expression groups based on the cut-off value, with the number of patients in each group explicitly specified; (6) survival hazard ratios (HRs) and 95% confidence interval (CI) by multivariate analysis or Kaplan–Meier (K-M) curves; and (7) published in the English language. Exclusion criteria are as follows: (1) studies describing other lncRNAs or lncRNAs unrelated to cancer; (2) duplicate articles; (3) other types of literature, such as reviews, letters, conference abstracts, meta-analyses, case reports, and so on; (4) articles focusing on biological functions and related mechanisms; and (5) a lack of sufficient HR and 95% CI to extract data.
2.3. Data Extraction and Quality Evaluation
The following information should be extracted from eligible literature: first author, publication year, country, tumor type, sample type, sample size (high/low), cutoff of BCYRN1 expression, analysis method, survival (OS/RFS/PFS), HR availability, HR (95%CI) with p value, month of follow-up, and Newcastle–Ottawa Scale score (NOS). Survival HRs (95% CI) were retrieved indirectly from K-M curves using the Engauge Digitizer tool in case multivariate analysis had not been performed. The NOS scoring criteria (scores from 0 to 9) were used to assess the quality of the included studies, and those with scores >6 were included in the meta-analysis.
2.4. Statistical Analysis
We used log HR and SE to summarize survival outcomes, while OR and corresponding 95% CI were applied to summarize clinicopathological parameters. In addition, between-study heterogeneity was assessed by the x2 test and I2 statistic. Q test (PQ) p value < 0.05 and I2 > 50% indicated that there was statistical heterogeneity among studies, and a random-effects model was used to analyze the results. In other cases, a fixed-effects model was employed. We used forest plots to present the meta-analysis results and used the Begg test to assess any prospective bias in the publications. Sensitivity analyses were performed by sequentially removing individual included studies to test whether the overall pooled estimate was stable. Analyses were performed using Stata 12.0 for Windows (Stata, College Station, TX, USA), and p < 0.05 was considered statistically significant.
2.5. To Identify the Differential Expression of BCYRN1 Gene in Human Cancers
UCSC Xena (https://xena.ucsc.edu/ originated from TCGA database) was used to retrieve RNA sequences, somatic mutations (SNPs and short INDELs), clinicopathologic, and survival data for 33 malignancies. We picked the ONCOMINE database (http://www.oncomine.org/) to acquire a complete knowledge of BCYRN1 expression in pan-cancers utilizing many data sets. As a result, the levels of BCYRN1 in various cancer types were assessed after a particular threshold was set (p value = 0.05, fold change = 1.5). Perl software was used to extract and combine BCYRN1 expression levels for TCGA pan-cancer analysis. The “Wilcoxon test” method was used to investigate the differential expression of BCYRN1 in various cancer types. As a threshold, a false discovery rate (FDR) of 0.05 was chosen. “∗”, “∗∗”, and “∗∗∗” represent FDR <0.05, <0.01, and <0.001, respectively. Box plots were then created with the R-package “ggpubr.” The cBioPortal database (https://www.cbioportal.org/) was used to assess changes in BCYRN1 expression in various cancer types [ref. 35].
2.6. Association between BCYRN1 Expression and TMB or MSI in Pan-Cancer
Tumor mutational burden (TMB) was computed using the Perl script and divided by the entire length of the exons to count the number of mutations in each tumor sample (i.e., 33 tumors using somatic mutation data and corrected to a number of mutated bases per 1 million bases). The microsatellite instability (MSI) score came from the TCGA website. The “cor. test” command was used to do Spearman’s method correlation study between cancer gene expression and TMB or MSI. A radar map was created using the R-package “fmsb” to view both indications.
2.7. Verification of Survival Outcomes in the GEPIA Database
Gene expression profiling interaction analysis (GEPIA) was performed according to the Cancer Genome Atlas (TCGA) data set to further validate the prognostic relevance of BCYRN1 overexpression in tumor tissues. TCGA and GTEx data were matched in various tumors, with a cutoff of p < 0.01. OS and DFS of BCYRN1 in pan-cancer were plotted using the Kaplan–Meier method.
3. Results
3.1. Screening Process for Eligible Literatures
The cancer-related gene BCYRN1 was thoroughly searched in four major English databases: PubMed (n = 53), Web of Science (n = 93), Embase (n = 73), and Cochrane Library (n = 0). After deleting duplicates (n = 106), the remaining papers’ titles and abstracts (n = 113) were examined and appraised. Sixty-three articles were rejected owing to the aims, article type (reviews, case studies, or conference abstracts), or unrelated findings. Forty-three full-text articles were downloaded, of which 31 were rejected following preliminary analysis due to lack of significant data or unsatisfactory quality of the data. Finally, 12 studies with sufficient data on survival and clinical features were included in the meta-analysis. The procedure is outlined in Figure 1.
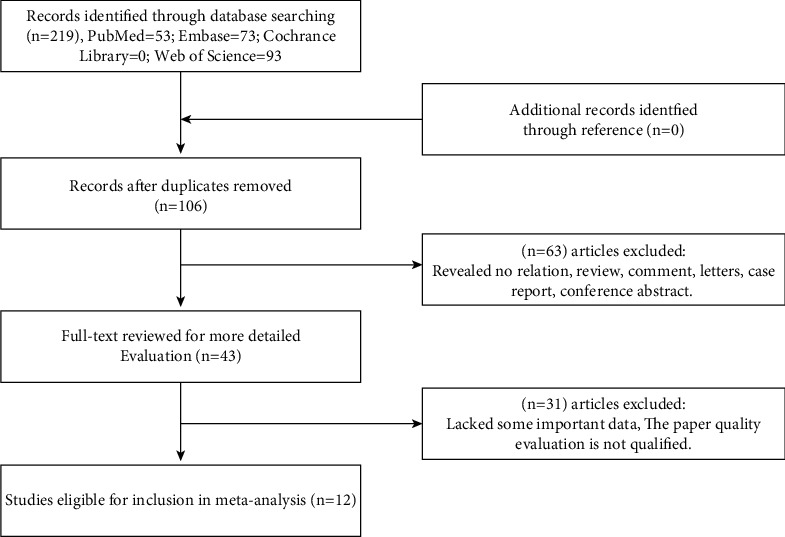
3.2. Characteristics of Included Research Projects
All included studies had been conducted in China and comprised 1,284 patients. The articles were published between 2016 and 2021. Two studies looked into hepatocellular carcinoma; two looked into colorectal cancer; and the remaining studies looked into bladder cancer, extranodal NK/T-cell lymphoma, glioblastoma, gastric cancer, prostate cancer, colon cancer, cancer, and esophageal squamous cell carcinoma. BCYRN1 expression in cancer and para-cancer tissues was detected by qRT-PCR. The patients were demarcated into the BCYRN1 low and BCYRN1 high groups, and the cutoff was the median expression level in five studies and mean expression in three studies. No cutoff value was indicated in the remaining four studies. Only two studies included the DFS and one PFS, whereas 8 studies provided OS. The HR and 95% CI of three studies were obtained directly from the multivariate regression analysis, and that for the remaining six were extracted from the K-M survival curves using Engauge Digitizer software. The duration of follow-up ranged from 40 to 96 months. The NOS scores of the studies were 6 to 8. The data are summarized in Table 1.
Table 1: Characteristics of the included studies.
| Author | Year | Country | Cancer type | Sample size (high/low) | Sample | Survival analysis | Detection method | Cutoff value | Extract method of HR | Follow-up time | NOS score |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Zheng et al. [ref. 25] | 2021 | China | BLC | 210 (105/105) | Tissue | OSDFS | qRT-PCR | Median | Data in paper | 96 months | 8 |
| Wang et al. [ref. 36] | 2021 | China | ENKTCL | 40 (20/20) | Tissue | PFS | qRT-PCR | Median | Survival curves | 40 months | 7 |
| Su et al. [ref. 37] | 2020 | ChinaTaiwan | GB | 48 (25/23) | Tissue | NR | qRT-PCR | NR | NR | NR | 6 |
| Huo et al. [ref. 26] | 2020 | China | PRCA | 72 (36/36) | Tissue | NR | qRT-PCR | NR | NR | NR | 7 |
| Zhai and Li [ref. 29] | 2019 | China | GC | 127 (63/64) | Tissue | OS | qRT-PCR | Median | Data in paper | 90 months | 8 |
| Yu and Chen [ref. 23] | 2019 | China | CRC | 150 (79/71) | Tissue | OS | qRT-PCR | Mean | Survival curves | 60 months | 8 |
| Ming et al. [ref. 20] | 2019 | China | HCC | 73 (37/36) | Tissue | OS | qRT-PCR | Median | Survival curves | 50 months | 7 |
| Gao and Wang [ref. 16] | 2019 | China | NSLC | 76 (32/44) | Tissue | OS | qRT-PCR | Mean | Survival curves | 70 months | 7 |
| Lin [ref. 38] | 2018 | China | HCC | 240 | Tissue | OS | qRT-PCR | Mean | Survival curves | 80 months | 8 |
| Wu et al. [ref. 21] | 2018 | China | CC | 82 | Tissue | OS | qRT-PCR | Mean | Survival curves | 80 months | 7 |
| Gu et al. [ref. 24] | 2018 | China | CRC | 96 (63/33) | Tissue | NR | qRT-PCR | NR | NR | NR | 6 |
| Zhao et al. [ref. 28] | 2016 | China | ESCC | 70 (35/35) | Tissue | OSDFS | qRT-PCR | Median | Data in paper | 50 months | 8 |
Note. BLC: bladder cancer, ENKTCL: extranodal NK/T-cell lymphoma, GB: glioblastoma, PRCA: prostate cancer, GC: gastric cancer, CRC: colorectal cancer, HCC: hepatocellular carcinoma, NSLC: non-small-cell lung cancer, CC: colon cancer, ESCC: esophageal squamous cell carcinoma, qRT-PCR: quantitative real-time polymerase chain reaction, and NR: not reported.
3.3. Association of BCYRN1 Level with Survival Outcome
Eight studies including 1,028 cancer patients investigated the link between BCYRN1 levels and OS. Since no significant heterogeneity was found (I2 = 0.0%, p=0.964), we performed a pooled analysis using a fixed-effect model. Pooled HRs indicated that high BCYRN1 levels were strongly associated with worse OS (HR = 1.84, 95% CI 1.51–2.25, p < 0.001; Figure 2(a)). In addition, only two studies (280 patients) were included to assess the association of BCYRN1 expression with DFS. Consistent with the OS results, increased BCYRN1 expression was found to be associated with unfavorable DFS (HR = 1.65, 95% CI 1.20–2.26, p=0.002; Figure 2(b)). Furthermore, we conducted subgroup analyses to look into the relationship between BCYRN1 expression levels and OS based on the cancer type (digestive or other systems; Figure 2(c)), sample size (≥100 or <100 tissues; Figure 2(d)), follow-up time (≥80 or <80 months; Figure 2(e)), and article quality (NOS score ≥8 or≤7; Figure 2(f)). There was no evidence of considerable heterogeneity within groups, and the findings of the subgroup analysis had no effect on BCYRN1‘s ability to predict OS in these malignancies.
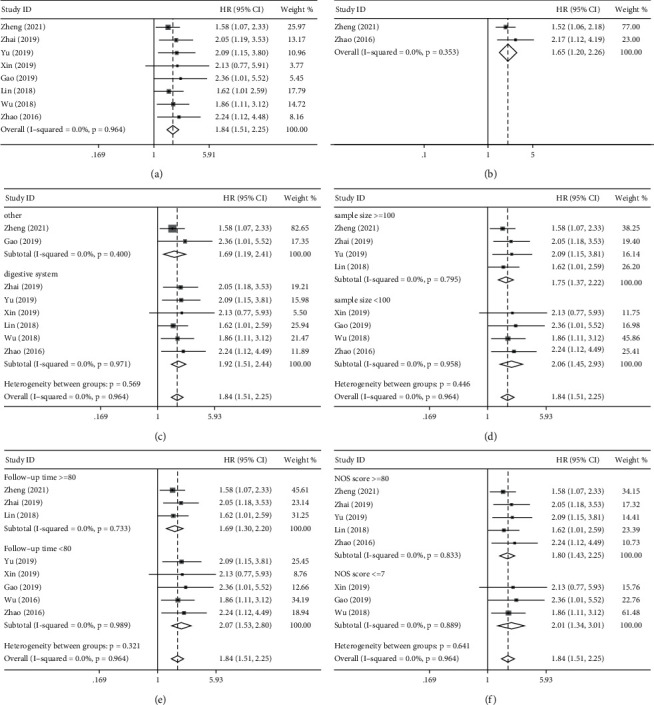
3.4. Association of BCYRN1 Expression with Clinicopathologic Parameters
The results showed that overexpression of BCYRN1 was associated with age (≥60 vs.<60, OR = 1.12, 95% CI 0.82–1.15, p=0.475; Figure 3(a)), gender (male vs. female, OR = 0.89, 95% CI 0.63–1.24, p=0.568; Figure 3(b)), tumor size (large vs. small, OR = 1.61, 95% CI 0.82–3.15, p=0.166; Figure 3(c)), lymph node metastasis (positive vs. negative, OR = 2.09, 95% CI 0.79–5.51, p=0.135; Figure 3(d)), and tumor differentiation (poor vs. good, OR = 1.10, 95% CI 0.59–2.05, p=0.774; Figure 3(e)) that were not significantly associated, and the results were not found to be statistically significant. However, high expression of BCYRN1 was observed to be significantly associated with some advanced clinical features, including TNM stage (III-IV vs. I-II, OR = 2.52, 95% CI 1.18–5.37, p=0.017; Figure 3(f)), T stage of the tumor (III-IV vs. I-II, OR = 2.11, 95% CI 1.49–2.99, p=0.000; Figure 3(g)), and tumor distant metastasis (positive vs. negative, OR = 4.19, 95% CI 1.45–12.05, p=0.008; Figure 3(h)). A fixed-effects model was used for low heterogeneity (0–50%), while a random-effects model was used for large heterogeneity (>50%). Data pertaining to the forest plot of survival prognosis and clinical pathology are recorded in Table 2.
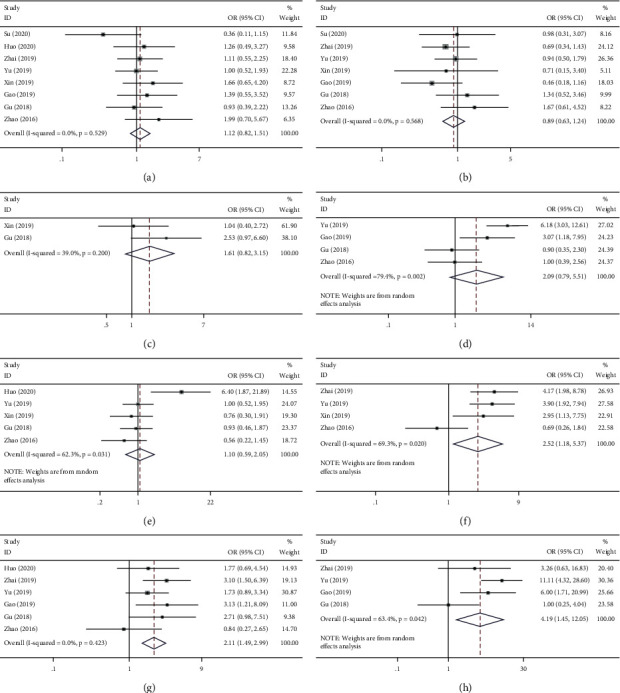
Table 2: Association of BCYRN1 expression with clinicopathological features and survival prognosis.
| Outcome | Studies (n) | OR/HR | 95% CI | p value | Model | Heterogeneity | ||
|---|---|---|---|---|---|---|---|---|
| Chi2 | I2 | p value | ||||||
| Age (≥60 vs.<60) | 8 | 1.12 | 0.82–1.15 | 0.475 | Fixed | 6.09 | 0.0% | 0.529 |
| Gender (male vs. female) | 7 | 0.89 | 0.63–1.24 | 0.568 | Fixed | 4.81 | 0.0% | 0.568 |
| Tumor size (large vs. small) | 2 | 1.61 | 0.82–3.15 | 0.166 | Fixed | 1.64 | 39% | 0.200 |
| Lymph node metastasis (positive vs. negative) | 4 | 2.09 | 0.79–5.51 | 0.135 | Random | 14.55 | 79.4 | 0.002 |
| Tumor differentiation (bad vs. well) | 5 | 1.10 | 0.59–2.05 | 0.774 | Random | 10.61 | 62.3% | 0.031 |
| TNM stage (III-IV vs. I-II) | 4 | 2.52 | 1.18–5.37 | 0.017 | Random | 9.79 | 67.3% | 0.020 |
| Tumor T stage (III-IV vs. I-II) | 6 | 2.11 | 1.49–2.99 | 0.000 | Fixed | 4.94 | 0.0% | 0.423 |
| Distant metastasis (positive vs. negative) | 4 | 4.19 | 1.45–12.05 | 0.008 | Random | 8.19 | 63.4% | 0.042 |
| Overall survival (OS) | 8 | 1.84 | 1.51–2.25 | 0.000 | Fixed | 1.91 | 0.0% | 0.964 |
| Disease-free survival (DFS) | 2 | 1.65 | 1.20–2.26 | 0.002 | Fixed | 0.86 | 0.0% | 0.353 |
3.5. Publication Bias and Sensitivity Analysis
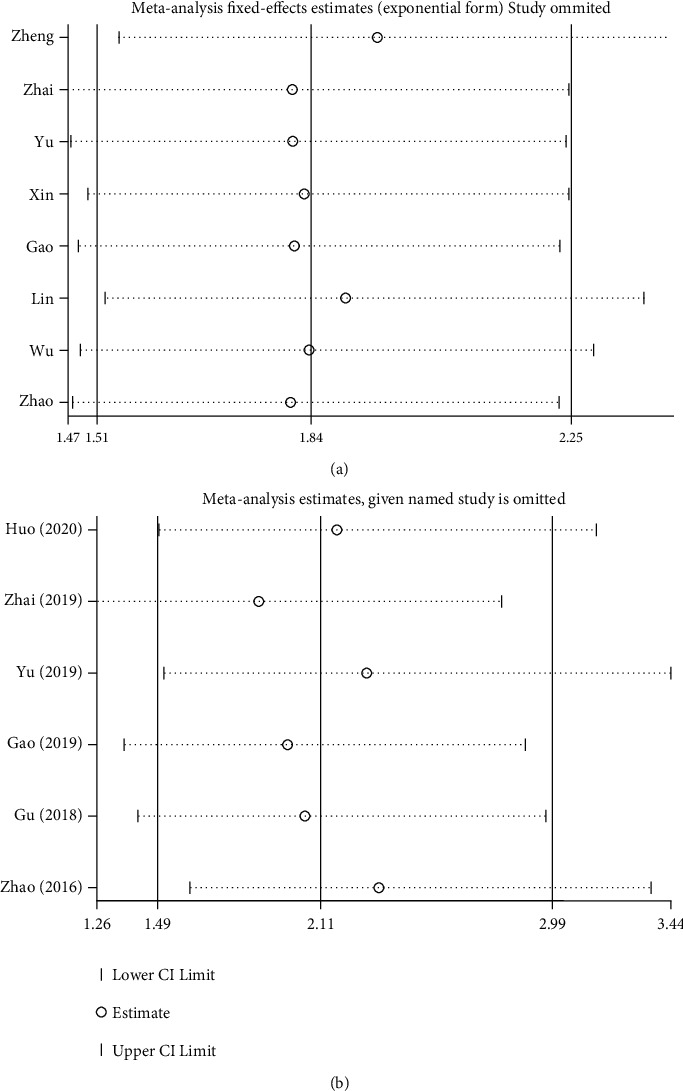
Begg’s test was used to analyze potential publication bias. For OS, the funnel plot appeared asymmetric, and the Begg test (p > |t| = 0.019; Figure 4(a)) indicated some publication bias. Using the scissors method, after filling out three imaginary unpublished papers, the funnel plot became symmetrical, and the pooled HR and 95% CI remained stable (HR = 1.768, 95% CI 1.473–2.123, p < 0.001; Figure 4(b)) [ref. 39]. For pathological parameters with significant differences in pooled ORs, Begg plot data showed TNM stage (p > |t | = 0.231; Figure 4(c)), distant metastasis (p > |t| = 0.237; Figure 4(d)), and tumor T stage (p > |t| = 0.605; Figure 4(e)), indicating no significant publication bias. A sensitivity analysis was performed for OS (Figure 5(a)) and tumor T stage (Figure 5(b)), and the pooled HR and OR changed within a limited range without significant change after deletion of each study, indicating that our results were stable. From this, it can be seen that the relevant conclusions we draw are stable and reliable.
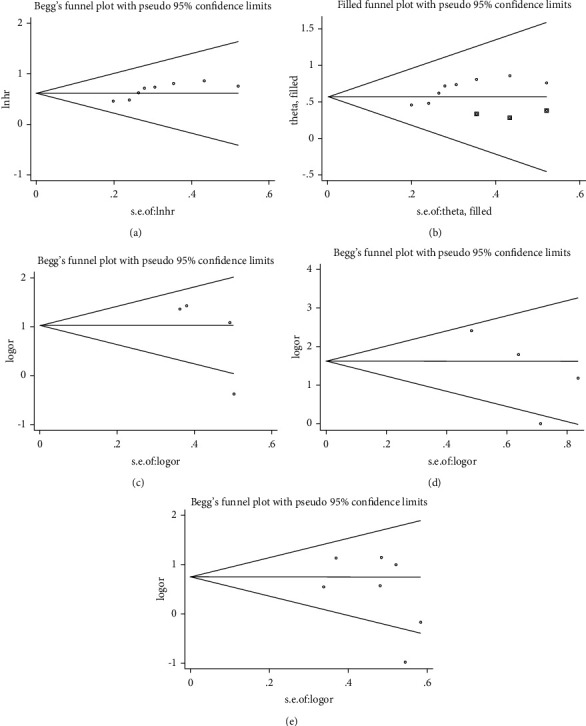
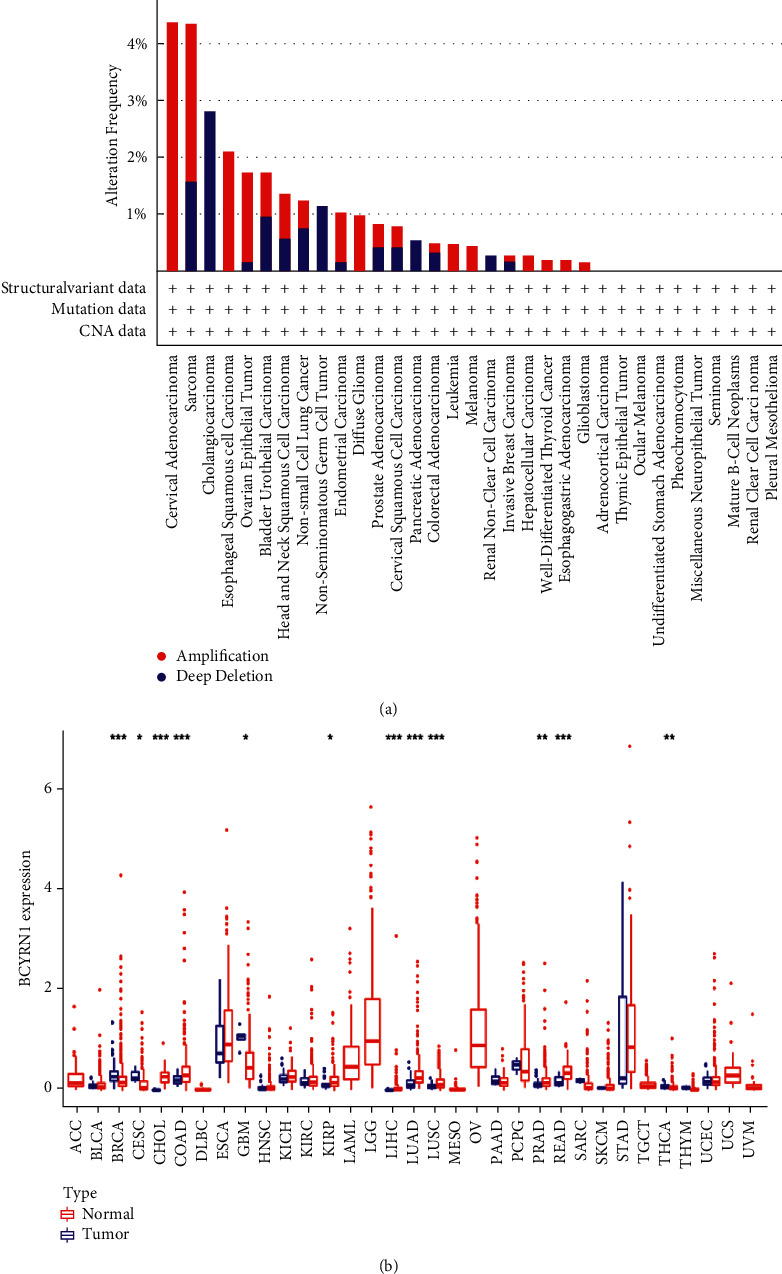
3.6. Expression of BCYRN1 in Pan-Cancers
We used R software to examine RNA sequencing data in the TCGA database to further investigate the differential expression of BCYRN1 in pan-cancers. According to our findings, BCYRN1 is significantly expressed in multiple cancer types, including CHOL, COAD, KIRP, LIHC, LUAD, LUSC, PRAD, and READ. However, low BCYRN1 expression was observed in BRCA, CESC, GBM, and THCA (Figure 6(a)). Using the cBioPortal database, we observed the variation of BCYRN1 in various types of cancer. The correlation results showed that the variation was mainly significant amplification, followed by deep deletion. Among all malignancies, cervical adenocarcinoma had the highest frequency of variants, followed by sarcoma (Figure 6(b)).
3.7. Association of BCYRN1 Expression with TMB and MSI in Pan-Cancer
High TMB is a newly identified class of biomarkers related to sensitivity to immune checkpoint inhibitors, including PD-1/PD-L1 inhibition, which can assess the efficacy of immunotherapy in cancer patients [ref. 40, ref. 41]. Therefore, it is interesting to investigate the relationship between TMB and BCYRN1 expression in different types of cancer. The results indicate that BCYRN1 expression correlates with TMB in a significant number of cancers. BCYRN1 expression was positively correlated with TMB in six cancer types, including BLCA, BRCA, HNSC, LUAD, LUSC, and THYM. In contrast, BCYRN1 expression was inversely correlated with TMB in six other cancer types, which included COAD, GBM, LGG, LIHC, STAD, and UCEC (Figure 7(a)).
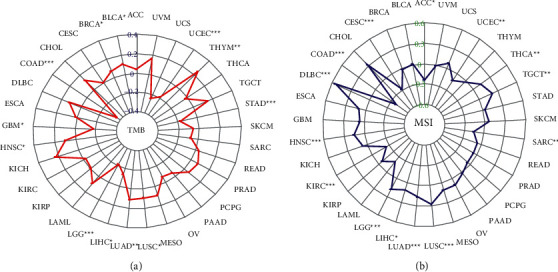
Recently, it has been found that MSI can be detected in numerous tumors (such as colorectal cancer) and has the potential to be a marker of PD-1 blockade [ref. 42, ref. 43]. Therefore, further verification of whether BCYRN1 expression is associated with MSI in different types of cancer is warranted. The results showed that BCYRN1 expression was significantly correlated with MSI in 14 cancer types. BCYRN1 expression was positively correlated with MSI in 8 of the cancer types (DLBC, HNSC, LGG, LIHC, LUAD, LUSC, TGCT, and THCA). In addition, BCYRN1 expression was inversely correlated with MSI in six other cancer types (ACC, CESC, COAD, KIRC, SARC, and UCEC; Figure 7(b)).
3.8. Correlation Analysis between BCYRN1 Expression and TNM Staging of Pan-Cancer
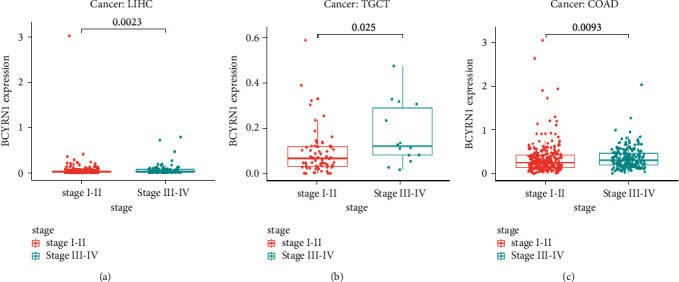
BCYRN1 expression was associated with the clinical stage in several cancers (Figure 8). For LIHC (p=0.0023; Figure 8(a)), TGCT (p=0.025; Figure 8(b)), and COAD (p=0.0093; Figure 8(c)), BCYRN1 was highly expressed in stage III-IV, but lowly expressed in stage I-II. From this, it can be seen that in the above cancers, high expression of BCYRN1 is associated with clinical stage progression of cancer and has the potential to be a predictor of tumor prognosis and progression.
3.9. Verification of Survival Outcomes in the GEPIA Database
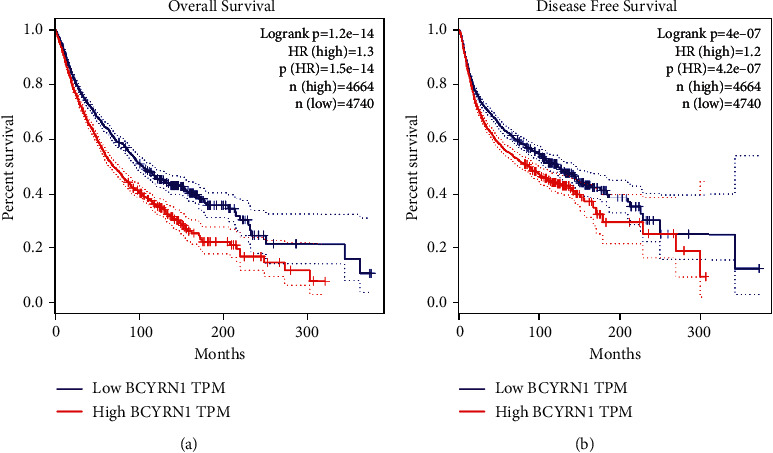
Regarding the relationship between BCYRN1 expression and prognosis, in the GEPIA cohort, 33 malignancies in 4,740 patients were divided into high and low expression groups according to the median value, and the survival curve showed that upregulation of BCYRN1 expression was associated with deterioration of OS ((HR = 1.3, log rank p < 0.05)) and DFS (HR = 1.2, log rank p < 0.05; Figure 9), which confirmed the results of our meta-analysis. These results support our conclusion and suggest that BCYRN1 may become a novel prognostic biomarker in multiple cancers.
4. Discussion
Given the steady increase in the annual rates of cancer incidence and mortality throughout the world, it is estimated that cancer will overtake chronic diseases as the primary cause of death and a major impediment to increasing life expectancy [ref. 44]. Despite the recent advances in cancer therapies, most cancer patients have a poor prognosis. Therefore, early diagnosis and treatment are critical to improving patient prognosis. However, the biomarkers currently used in clinical practice lack sensitivity and specificity, thereby necessitating the identification of novel tumor markers [ref. 45]. LncRNAs are transcribed by RNA polymerase II, and their expression levels vary significantly between tumors and the corresponding normal tissues. Studies show that lncRNAs regulate gene expression through X chromosome silencing, chromatin modification, transcriptional interference, and activation, which in turn regulate various physiological and pathological processes [ref. 46]. The lncRNA BCYRN1 is upregulated in multiple cancers and is therefore a potential diagnostic biomarker and therapeutic target. In addition, aberrant BCYRN1 expression is also related to the neurodegeneration underlying Alzheimer’s disease [ref. 47]. We conducted a meta-analysis of 12 studies including 1,284 cancer patients and 10 distinct cancer types and found that BCYRN1 overexpression in the tumors correlated significantly to poor survival, worse clinical stage, distant tumor metastasis, and advanced tumor T stage with greater invasiveness. Our findings are in line with previous reports indicating the prognostic relevance of BCYRN1 in cancer. Finally, we further evaluated the prognostic and pathological value of BCYRN1 by downloading relevant data using public databases, and the results were consistent with our meta-analysis.
Three studies included in the meta-analysis reported increased expression of BCYRN1 in lung cancers, which correlated with poor outcomes. Wang et al. showed that BCYRN1 promoted the proliferation and metastasis of NSCLC cells by activating the Wnt/β-catenin signaling pathway [ref. 15]. Another study reported an association between BCYRN1 and advanced tumor stage and metastasis in NSCLC patients. BCYRN1 augmented the malignant development by targeting H1299/DDP-induced apoptosis [ref. 16]. Furthermore, Hu and Lu found that c-myc-activated BCYRN1 controlled NSCLC cell metastasis by upregulating MMP9 and MMP13 [ref. 17]. Three studies analyzed the relationship between BCYRN1 and liver cancer and reported upregulation of BCYRN1 in the tumor tissues. Ding et al. identified the BCYRN1/miR-490-3p/POU3F2 ceRNA regulatory network mediating reduced survival and increased tumor cell proliferation and metastasis in HCC patients [ref. 18]. Tan et al. found that BCYRN1 influences hepatoma cell proliferation and migration by modulating the expression of the c-Myc protein [ref. 19], and Ming et al. showed that BCYRN1 regulates tumor-associated pathways and promotes hepatocarcinogenesis via lncRNA-miRNA-mRNA networks [ref. 20]. The upregulation of BCYRN1 in colorectal cancer was reported in four studies. Wu et al. found that knocking down BC200 decreased invasion and epithelial-mesenchymal transition (EMT) in HCT-116 and HT29 cells via the downregulation of MMP-2 and MMP-9 [ref. 21]. Yang et al. showed that BCYRN1 functioned as an oncogene in colorectal cancer via the miR-204-3p/KRAS axis [ref. 22]. In addition, Yu and Chen reported that the aberrantly high expression of BCYRN1 in colorectal cancer tissues increased metastasis and worsened patient prognosis [ref. 23]. Likewise, BCYRN1 overexpression was linked to larger tumors and advanced pathological stages in colorectal cancer patients [ref. 24]. Two studies so far have analyzed the relationship between BCYRN1 expression and prostate cancer. Zheng et al. showed that the high expression of BCYRN1 in prostate cancer tissues induced BCA lymphatic metastasis by activating VEGF-C/VEGFR3 signaling [ref. 25]. Huo et al. found that BCYRN1 enhanced HDAC11 levels and promoted prostate cancer cell proliferation, glucose metabolism, and survival by targeting miR-939-3p [ref. 26]. Two studies showed a link between BCYRN1 expression and gliomas. Mu et al. showed that BCYRN1 is downregulated in gliomas and controls CUEDC2 expression and the PTEN/AKT/p21 pathway to suppress tumor progression by competitively binding to miR-619-5p [ref. 48]. Su et al. on the other hand reported overexpression of BCYRN1 in gliomas and found that it targets the BC200/miR218-5p signaling axis to overcome T mozolamide resistance and inhibit tumor growth [ref. 37]. The link between BCYRN1 expression and ESCC has been reported in two studies. Zhao et al. showed that BC200 enhances esophageal cancer cell metastasis and controls the expression of ATF4 and its downstream genes [ref. 27] and that patients with high BC200 expression exhibited worse disease-free and overall survival [ref. 28]. There are two reports investigating the link between BCYRN1 expression and stomach cancer. Zhai and Li found that BCYRN1 is highly expressed in gastric cancer tissues and controls gastric cancer cell proliferation, cell cycle, migration, and invasion by targeting miR-204-5p [ref. 29]. Ren et al. reported similar findings [ref. 30]. Peng et al. discovered that BCYRN1 was highly expressed in cervical cancer and that miR-138 inhibition increased cervical cancer proliferation and invasion [ref. 31]. In addition, BCYRN1 is upregulated in extranodal lymphomas and may enhance ASP resistance by activating autophagy [ref. 36]. BC200 is expressed at low levels in ovarian cancer and may inhibit tumor cell proliferation [ref. 32]. Singh et al. found that BC200 is upregulated in breast cancer and is a potential target for estrogen-dependent breast cancer [ref. 33]. Except for gliomas and ovarian cancer, BCYRN1 is highly expressed in most malignancies, and the oncogenic mechanisms need future investigation. The mechanism and research progress of BCYRN1 in various types of cancer are shown in Table 3.
Table 3: Summary of lncRNA BCYRN1 functional roles and related genes.
| Cancer types | Expression | Potential targets | Pathways and mechanisms | Related microRNAs | References |
|---|---|---|---|---|---|
| NSCLC | Up | β-Catenin/c-Myc/cyclin D1 | Cell proliferation and migration↑; Wnt/β-catenin signaling | NR | [ref. 15] |
| PI3K/AKT/STAT3 | Cell proliferation, invasion, and migration↑; PI3K/AKT pathway | NR | [ref. 16] | ||
| c-MYC | Cell metastasis↑; promoting the expressions of MMP9 and MMP13 | NR | [ref. 17] | ||
| HCC | Up | POU3F2 | Cells growth, clone formation, and movement abilities↑ | miR-490-3p | [ref. 18] |
| c-MYC/Bcl-xL | Affected the proliferation and migration of HepG2 cells; reduced the expression of Bcl-xL protein | NR | [ref. 19] | ||
| Colon cancer | Up | STAT3/β-catenin | Proliferation↑; apoptosis↓; reduction of the phosphorylation of STAT3 | NR | [ref. 21] |
| CRC. | Up | KRAS | Proliferation, migration, and invasion ↑; apoptosis↓ | miR-204-3p | [ref. 22] |
| CCA T2 | CCA T2; miR-320a axis | miR-320a | [ref. 23] | ||
| NPR3 | proliferation↑; apoptosis↓ | NR | [ref. 24] | ||
| BLC | UP | WNT5A/VEGF-C/VEGFR3 | Activates WNT5A/vegf-c/vegfr3 feedforward loop to drive lymphatic metastasis | NR | [ref. 25] |
| Up | HDAC11 | Sponged miR939-3p to upregulate histone deacetylase 11 (HDAC11) expression | miR-939-3p | [ref. 26] | |
| Glioma | Down | CUEDC2 | Regulate CUEDC2 expression and the PTEN/akt/p21 pathway | miR-619-5p | [ref. 48] |
| ESCC | Up | ATF4 | Cell invasion and migration↑ | NR | [ref. 27] |
| GC | Up | NR | Cell proliferation, cell cycle, migration, and invasion↑ | miR-204-5p | [ref. 29] |
| EpCAM | Cell proliferation and metastasis↑; apoptosis↓ | NR | [ref. 30] | ||
| Cervical cancer | Up | NR | The proliferation and invasion ↑of cervical cancer via targeting miR-138 | miR-138 | [ref. 31] |
| ENKTCL | Up | PI3K/AKT/mTOR/p53 | PI3K/AKT/mTOR/p53 pathways | NR | [ref. 36] |
| Breast cancer | Up | Bcl-xL | Apoptosis↓ | NR | [ref. 33] |
Note. NSCLC: non-small-cell lung cancer, HCC: hepatocellular carcinoma, CRC: colorectal cancer, BLC: bladder cancer, ESCC: esophageal squamous cell carcinoma, ENKTCL: extranodal NK/T-cell lymphoma, NR: not reported, ↑: promote, and ↓: inhibit.
There are several limitations to our study that ought to be considered. Since all studies had been conducted in China, our findings may only apply to Asian patients. Second, only a tiny percentage of cases were included, and several cancer types had limited sample sizes. To overcome these restrictions, we analyzed the gene in an existing public database to validate and increase the reliability of the results. Third, manually deriving HRs for OS and PFS from Kaplan–Meier curves might lead to operational mistakes. Fourth, we did not have a uniform threshold value for high and low BCYRN1 expression, since some studies used median and others used mean. Fifth, this study used meta-analysis and bioinformatics analysis to make a preliminary summary and judgment of the prognosis and expression of BCYRN1 in tumors, providing a theoretical basis for future research in this area, but the lack of specific laboratory validation is a great regret. Sixth, this paper is an analysis of pan-cancer, but there is heterogeneity between each cancer, and BCYRN1 can be specifically analyzed in separate cancer types in the future. Finally, all studies were published in English, which may have led to selection bias.
5. Conclusion
Elevated BCYRN1 expression is correlated to worse prognosis and clinicopathological features (including T stage, clinical stage, and distant tumor metastasis) in cancer patients. There was no significant relationship between high BCYRN1 expression and patient age, gender, tumor differentiation, lymphatic metastasis, or tumor size. Thus, BCYRN1 is a potential diagnostic biomarker and therapeutic target in various cancers, although the underlying mechanisms and clinical significance have to be corroborated further with large-scale, multicenter cohort studies.
References
- F. Bray, M. Laversanne, E. Weiderpass, I. Soerjomataram. The ever‐increasing importance of cancer as a leading cause of premature death worldwide. Cancer, 2021. [DOI | PubMed]
- R. L. Siegel, K. D. Miller, H. E. Fuchs, A Jemal. Cancer statistics. CA: A Cancer Journal for Clinicians, 2021. [DOI | PubMed]
- R. L. Siegel, K. D. Miller, A. Jemal. Cancer statistics. CA: A Cancer Journal for Clinicians, 2017. [DOI | PubMed]
- L. W. Elmore, S. F. Greer, E. C. Daniels. Blueprint for cancer research: critical gaps and opportunities. CA: A Cancer Journal for Clinicians, 2021. [DOI | PubMed]
- R. I. Stefan-van Staden, I. R. Comnea-Stancu, H. Yanık, M. Göksel, A. Alexandru, M. Durmuş. Phthalocyanine-BODIPY dye: synthesis, characterization, and utilization for pattern recognition of CYFRA 21-1 in whole blood samples. Analytical and Bioanalytical Chemistry, 2017. [DOI | PubMed]
- R. I. Stefan-van Staden, R. M. Ilie-Mihai, S. Gurzu. The fast screening method of biological samples for early diagnosis of gastric cancer. Multidisciplinary Cancer Investigation, 2020. [DOI]
- R.-I. Stefan-van Staden, R. M. Ilie-Mihai, F. Pogacean, S. M. Pruneanu. Needle stochastic sensors for on-site fast recognition and quantification of biomarkers for gastric cancer in biological samples. New Journal of Chemistry, 2020. [DOI]
- I. R. Comnea-Stancu, R. I. Stefan-van Staden, J. F. van Staden, C. Stanciu-Gavan. Stochastic sensors based on maltodextrins for screening of whole blood for neuron specific enolase, carcinoembryonic antigen and epidermal growth factor receptor. Microsystem Technologies, 2016. [DOI]
- N. P. T. Huynh, B. A. Anderson, F. Guilak, A. McAlinden. Emerging roles for long noncoding RNAs in skeletal biology and disease. Connective Tissue Research, 2017. [DOI | PubMed]
- M. Gugnoni, A. Ciarrocchi. Long noncoding RNA and epithelial mesenchymal transition in cancer. International Journal of Molecular Sciences, 2019. [DOI]
- E. Anastasiadou, L. S. Jacob, F. J. Slack. Non-coding RNA networks in cancer. Nature Reviews Cancer, 2018. [DOI | PubMed]
- S. Geisler, J. Coller. RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. Nature Reviews Molecular Cell Biology, 2013. [DOI | PubMed]
- X. Chen, C. C. Yan, X. Zhang, Z. H. You. Long non-coding RNAs and complex diseases: from experimental results to computational models. Briefings in Bioinformatics, 2017. [DOI]
- S. Ghafouri-Fard, S. Dashti, B. M. Hussen, M. Farsi, M. Taheri. BCYRN1: an oncogenic lncRNA in diverse cancers. Pathology, Research and Practice, 2021. [DOI]
- Y. Wang, W. Bai, M. Wang, T. Yu, W. Zhang. Long non-coding RNA brain cytoplasmic RNA 1 acts as an oncogene and regulates cell proliferation and metastasis in non-small cell lung cancer. Journal of Nanoscience and Nanotechnology, 2019. [DOI | PubMed]
- B. B. Gao, S. X. Wang. LncRNA BC200 regulates the cell proliferation and cisplatin resistance in non-small cell lung cancer via PI3K/AKT pathway. European Review for Medical and Pharmacological Sciences, 2019. [DOI | PubMed]
- T. Hu, Y. R. Lu. BCYRN1, a c-MYC-activated long non-coding RNA, regulates cell metastasis of non-small-cell lung cancer. Cancer Cell International, 2015. [DOI]
- S. Ding, Y. Jin, Q. Hao, Y. Kang, R. Ma. LncRNA BCYRN1/miR-490-3p/POU3F2, served as a ceRNA network, is connected with worse survival rate of hepatocellular carcinoma patients and promotes tumor cell growth and metastasis. Cancer Cell International, 2020. [DOI]
- N. Tan, B. Zhu, H. Shu. Effect of lncRNA‑BC200 on proliferation and migration of liver cancer cells in vitro and in vivo. Oncology Reports, 2020. [DOI | PubMed]
- X. L. Ming, Y. L. Feng, D. D. He. Role of BCYRN1 in hepatocellular carcinoma pathogenesis by lncRNA-miRNA-mRNA network analysis and its diagnostic and prognostic value. Epigenomics, 2019. [DOI | PubMed]
- K. Wu, K. Xu, K. Liu. Long noncoding RNA BC200 regulates cell growth and invasion in colon cancer. The International Journal of Biochemistry & Cell Biology, 2018. [DOI | PubMed]
- L. Yang, Y. Zhang, J. Bao, J. F. Feng. Long non-coding RNA BCYRN1 exerts an oncogenic role in colorectal cancer by regulating the miR-204-3p/KRAS axis. Cancer Cell International, 2020. [DOI]
- J. H. Yu, Y. Chen. Clinical significance of lncRNA BCYRN1 in colorectal cancer and its role in cell metastasis. European Review for Medical and Pharmacological Sciences, 2019. [DOI | PubMed]
- L. Gu, L. Lu, D. Zhou, Z. Liu. Long noncoding RNA BCYRN1 promotes the proliferation of colorectal cancer cells via up-regulating NPR3 expression. Cellular Physiology and Biochemistry, 2018. [DOI | PubMed]
- H. Zheng, Y. Chen, M. Luo. Tumor‐derived exosomal BCYRN1 activates WNT5A/VEGF‐C/VEGFR3 feedforward loop to drive lymphatic metastasis of bladder cancer. Clinical and Translational Medicine, 2021. [DOI]
- W. Huo, F. Qi, K. Wang. Long non‑coding RNA BCYRN1 promotes prostate cancer progression via elevation of HDAC11. Oncology Reports, 2020. [DOI | PubMed]
- R. Zhao, S. Cao, R. Jin. LncRNA BC200 promotes esophageal squamous cell cancer migration and invasion and can regulate ATF4 expression. Frontiers in Oncology, 2020. [DOI]
- R. H. Zhao, X. K. Li, W. Cao. BC200 LncRNA a potential predictive marker of poor prognosis in esophageal squamous cell carcinoma patients. OncoTargets and Therapy, 2016. [DOI | PubMed]
- H. Zhai, Y. Li. BCYRN1 is correlated with progression and prognosis in gastric cancer. Bioscience Reports, 2019. [DOI]
- H. Ren, X. Yang, Y. Yang. Upregulation of LncRNA BCYRN1 promotes tumor progression and enhances EpCAM expression in gastric carcinoma. Oncotarget, 2018. [DOI | PubMed]
- J. Peng, F. Hou, J. Feng, S. X. Xu, X. Y. Meng. Long non-coding RNA BCYRN1 promotes the proliferation and metastasis of cervical cancer via targeting microRNA-138 in vitro and in vivo. Oncology Letters, 2018. [DOI | PubMed]
- D. I. Wu, T. Wang, C. Ren. Downregulation of BC200 in ovarian cancer contributes to cancer cell proliferation and chemoresistance to carboplatin. Oncology Letters, 2016. [DOI | PubMed]
- R. Singh, S. C. Gupta, W. X. Peng. Regulation of alternative splicing of Bcl-x by BC200 contributes to breast cancer pathogenesis. Cell Death and Disease, 2016. [DOI]
- A. Liberati, D. G. Altman, J. Tetzlaff. The PRISMA statement for reporting systematic reviews and meta-analyses of studies that evaluate healthcare interventions: explanation and elaboration. BMJ, 2009. [DOI]
- Y. Miao, J. Wang, Q. Li. Prognostic value and immunological role of PDCD1 gene in pan-cancer. International Immunopharmacology, 2020. [DOI]
- L. Wang, J. Yang, H. Wang. LncRNA BCYRN1-induced autophagy enhances asparaginase resistance in extranodal NK/T-cell lymphoma. Theranostics, 2021. [DOI | PubMed]
- Y. K. Su, J. W. Lin, J. W. Shih. Targeting BC200/miR218-5p signaling axis for overcoming temozolomide resistance and suppressing glioma stemness. Cells, 2020. [DOI]
- Y. H. Lin, M. H. Wu, Y. H. Huang. Thyroid hormone negatively regulates tumorigenesis through suppression of BC200. Endocrine-Related Cancer, 2018. [DOI | PubMed]
- M. D. Jennions, A. P. Møller. Publication bias in ecology and evolution: an empirical assessment using the “trim and fill” method. Biological Reviews of the Cambridge Philosophical Society, 2002. [DOI | PubMed]
- Z. R. Chalmers, C. F. Connelly, D. Fabrizio. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Medicine, 2017. [DOI]
- M. Yarchoan, A. Hopkins, E. M. Jaffee. Tumor mutational burden and response rate to PD-1 inhibition. New England Journal of Medicine, 2017. [DOI | PubMed]
- J. C. Dudley, M. T. Lin, D. T. Le, J. R. Eshleman. Microsatellite instability as a biomarker for PD-1 blockade. Clinical Cancer Research, 2016. [DOI | PubMed]
- C. R. Boland, A. Goel. Microsatellite instability in colorectal cancer. Gastroenterology, 2010. [DOI | PubMed]
- F. Bray, J. Ferlay, I. Soerjomataram, R. L. Siegel, L. A. Torre, A. Jemal. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians, 2018. [DOI | PubMed]
- C. Deng, X. Hu, K. Wu, J. Tan, C. Yang. Long non‑coding RNA XIST predicting advanced clinical parameters in cancer: a meta‑analysis and case series study in a single institution review. Oncology Letters, 2019. [DOI | PubMed]
- F. R. Kong, Y. H. Lv, H. M. Yao, H. Y. Zhang, Y. Zhou, S. E. Liu. LncRNA PCAT6 promotes occurrence and development of ovarian cancer by inhibiting PTEN. European Review for Medical and Pharmacological Sciences, 2019. [DOI | PubMed]
- E. Mus, P. R. Hof, H. Tiedge. Dendritic BC200 RNA in aging and in Alzheimer’s disease. Proceedings of the National Academy of Sciences, 2007. [DOI]
- M. Mu, W. Niu, X. Zhang, S. Hu, C. Niu. Correction: LncRNA BCYRN1 inhibits glioma tumorigenesis by competitively binding with miR-619-5p to regulate CUEDC2 expression and the PTEN/AKT/p21 pathway. Oncogene, 2021. [DOI | PubMed]
