SignatureFinder enables sequence mining to identify cobalamin‐dependent photoreceptor proteins
Abstract
Photoreceptors control cellular processes in response to light. Most photoreceptors sense blue or red light, but the recent discovery of the cobalamin‐dependent photoreceptor, CarH, has expanded the wavelength range of photoreception to other regions of the electromagnetic spectrum to include the green light region. Further identification of cobalamin‐dependent green light‐sensitive photoreceptors has been hampered owing to poor annotation of the light responsiveness of cobalamin‐binding domains (CBDs) in public databases. Here we report a computational workflow, SignatureFinder, that uses a combination of sequence and structural analyses to identify new light‐responsive CBD‐containing proteins. The light response of exemplar proteins containing the proposed signature were confirmed experimentally. A structural analysis of these new photoreceptors, including the crystal structure of a new CBD domain, highlights how the signature elements interact with the cobalamin chromophore to sense light. Database mining of 128 000 CBD‐containing sequences using the identified signature revealed more diverse CBD‐containing photoreceptors, thereby expanding the family of green‐light photoreceptors. A SignatureFinder web server is available (https://enzymeevolver.com) for wider applications, including the identification of signature sequences of other biological ligands of interest.
Article type: Research Article
Keywords: bioinformatics, cobalamin, photoreceptors, sequence motif, structure
Affiliations: Department of Chemistry The University of Manchester, Manchester Institute of Biotechnology UK; Present address: Astra Zeneca Cambridge UK; Department of Chemistry The University of Manchester UK; Present address: Department of Biology and Chemistry, College of Sciences National University of Defense Technology Changsha China
License: © 2024 The Author(s). The FEBS Journal published by John Wiley & Sons Ltd on behalf of Federation of European Biochemical Societies. CC BY 4.0 This is an open access article under the terms of the http://creativecommons.org/licenses/by/4.0/ License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Article links: DOI: 10.1111/febs.17377 | PubMed: 39718193 | PMC: PMC11796333
Relevance: Relevant: mentioned in keywords or abstract
Full text: PDF (14.5 MB)
Introduction
In cell biology, optogenetics empowers researchers to use light to perturb and control cellular processes with fine spatial and temporal resolution [ref. 1, ref. 2, ref. 3, ref. 4, ref. 5]. Optogenetics has also provided valuable insights into cell malfunction for the development and improvement of therapeutics [ref. 2, ref. 6, ref. 7, ref. 8]. Photoreceptors are at the core of optogenetics where they play important roles in maintaining cellular activities in response to light [ref. 9, ref. 10, ref. 11, ref. 12, ref. 13]. Photoreceptors typically comprise sensor domains and coupled effector domains [ref. 1, ref. 14, ref. 15]. They sense light using a bound chromophore located in the sensor domain and transmit this signal to the coupled effector domains, which undergo conformational changes to trigger a downstream response [ref. 6, ref. 16, ref. 17, ref. 18, ref. 19, ref. 20, ref. 21]. Photoreceptors are highly diverse in that different sensor domains combine with a range of available effector domains. Additionally, artificial photoreceptor fusion proteins can combine sensors and targeted effector domains to expand this diversity, which forms the basis of a multifunctional optogenetics toolbox [ref. 1].
CarH was the first discovered photoreceptor to use adenosylcobalamin as a light‐sensing chromophore [ref. 20]. This interesting protein regulates the expression of the genes required for the synthesis of carotenoids, which can protect cells from photo‐oxidative stress [ref. 20, ref. 22, ref. 23]. In the dark, CarH is a tetramer that binds to target DNA to inhibit transcription. Upon illumination with green light, adenosylcobalamin photochemistry leads to dissociation of the tetramer, which causes the release of the DNA and initiation of transcription. As a rare example of a green‐light‐induced photoreceptor, CarH has been exploited in light‐dependent cell release/recovery and regulation of cell adhesion [ref. 24, ref. 25, ref. 26, ref. 27]. Future applications of CarH in establishing cell or mini‐organ culturing platforms, and optical therapeutic treatments, are also envisioned [ref. 28]. The use of CarH in medical hydrogels is limited by the permeability of green light through the skin barrier. Therefore, cobalamin‐dependent photoreceptors sensitive to other wavelengths of light may provide for further optogenetic tools, particularly proteins sensitive to red light [ref. 29]. To assist in this goal, reliable identification of new cobalamin‐binding photoreceptors is needed. A major challenge is that most cobalamin‐binding domains (CBDs) are not naturally light‐responsive, making it difficult to identify new photoreceptor proteins. For example, CBD‐containing enzymes, such as methylcobalamin‐dependent methionine synthase, adenosylcobalamin‐dependent enzymes glutamate mutase and methylmalonyl‐CoA mutase, utilise cobalamin as a nonphotoactive cofactor. As such, these thermally activated enzymes do not show the typical CarH‐like cobalamin photochemical response [ref. 30, ref. 31, ref. 32, ref. 33, ref. 34, ref. 35]. There are 128 000 sequences containing CBDs in the protein database (InterPro [ref. 36] classification number: IPR006158), and there are no annotations for the light‐responsive features of these proteins. Methods are now required to identify signature sequences for light‐responsive CBD‐containing proteins to distinguish potential light‐responsive sequences from the thousands of other CBD‐containing proteins in the database to enable identification and experimental characterisation of cobalamin‐dependent photoreceptor proteins. We address this need here.
Results and discussion
Overview of approach
We set out to combine phylogenetic analysis and structural predictions to classify photoreceptors and nonphotoreceptors. Our hypothesis is that proteins similar to CarH at both the evolutionary and structural level have a strong likelihood of being photoreceptors, which was then validated by experimental analysis. First, we identified signature sequences using sequence alignments and performed biochemical analysis of potential photoreceptor‐AdoCbl models, as well as determining the crystal structure and biophysical/chemical analysis of the predicted photoreceptor CtMerR to confirm light‐responsivity. Subsequent database mining based on the identified signature sequence was then used to expand the photoreceptor family to 1500 potential light‐responsive CBD‐containing photoreceptors of wide‐ranging function. A sequence similarity network (SSN) was constructed and published on NDEx [ref. 37, ref. 38, ref. 39] with an interactive interface to reveal the protein clusters, and representatives from each cluster were modelled by AlphaFold for annotation of functional domains. Our computational workflow is available as the web server SignatureFinder alongside our previous tool IREDFisher at https://enzymeevolver.com which will allow community use to explore signature sequences in protein families of interest [ref. 40]. This work also led to the characterisation of proteins that contain two chromophores allowing this photocobilin protein family to sense light across the UV–Vis spectrum [ref. 29].
SignatureFinder web server
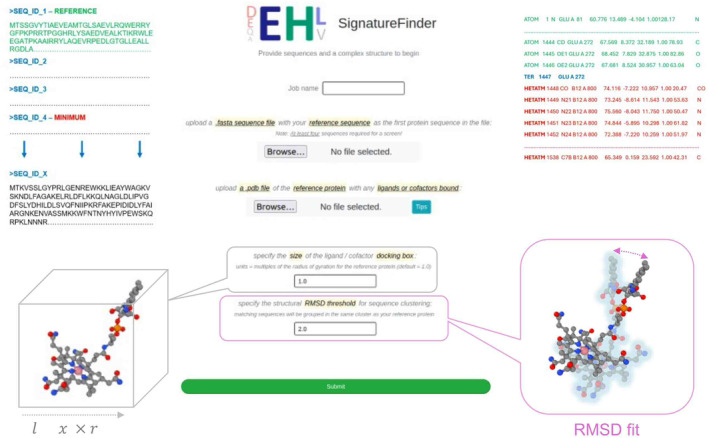
We demonstrate the application of the SignatureFinder web server by using it to identify light‐responsive signatures in CBD‐containing proteins (Fig. 1). A reference protein known to have the function of interest is needed prior to use of the web server. The web server requires the upload of a document containing the reference FASTA sequence and at least three query FASTA sequences. A ligand‐bound structure of a reference protein is also needed for structural comparison with predicted models of the homologues. Choosing an appropriate reference protein sequence and structure are key to the success of this method and web server. In this study, CarH from Thermus thermophilus (TtCarH) was the reference protein and CBD‐containing sequences collected by a BLAST [ref. 41] homology search based on the CBD in TtCarH (TtCBD) were the query sequences. The crystal structure of the CBD with a bound adenosylcobalamin molecule (PDB code: 5C8A) was used as the reference structure. To ensure that the models don’t diverge from the reference proteins, a 2.0 Å cut‐off was applied to the root‐mean‐square deviation (RMSD) between the models and reference. The size of the docking box was calculated based on gyration radius of the ligand (Rg) in the reference structure. It is a cubic box with Rg * Rg * Rg. The user can specify the size by filling out the ratio from the web server. A ratio of 1.0 was used in this case and can be used as a default by users.
The query sequences go through the workflow in a stepwise manner (Fig. 2): in the first step, a phylogenetic tree file is generated to give a general picture of the evolutionary relationship between the query sequences and the reference protein. Sequences in the same branch of the phylogenetic tree are defined as photoreceptors. Next, the 3D structure of each sequence is modelled and compared with the reference structure. Sequences where the RMSD of the model is under the cut‐off values are defined as photoreceptors and vice versa. Then, sequences of predicted photoreceptors are aligned and the light‐sensor adenosylcobalamin (AdoCbl) is docked into the structure of each predicted photoreceptor. The amino acid residues that interact with AdoCbl and are also conserved in the sequence alignment are defined as the signature for light response.
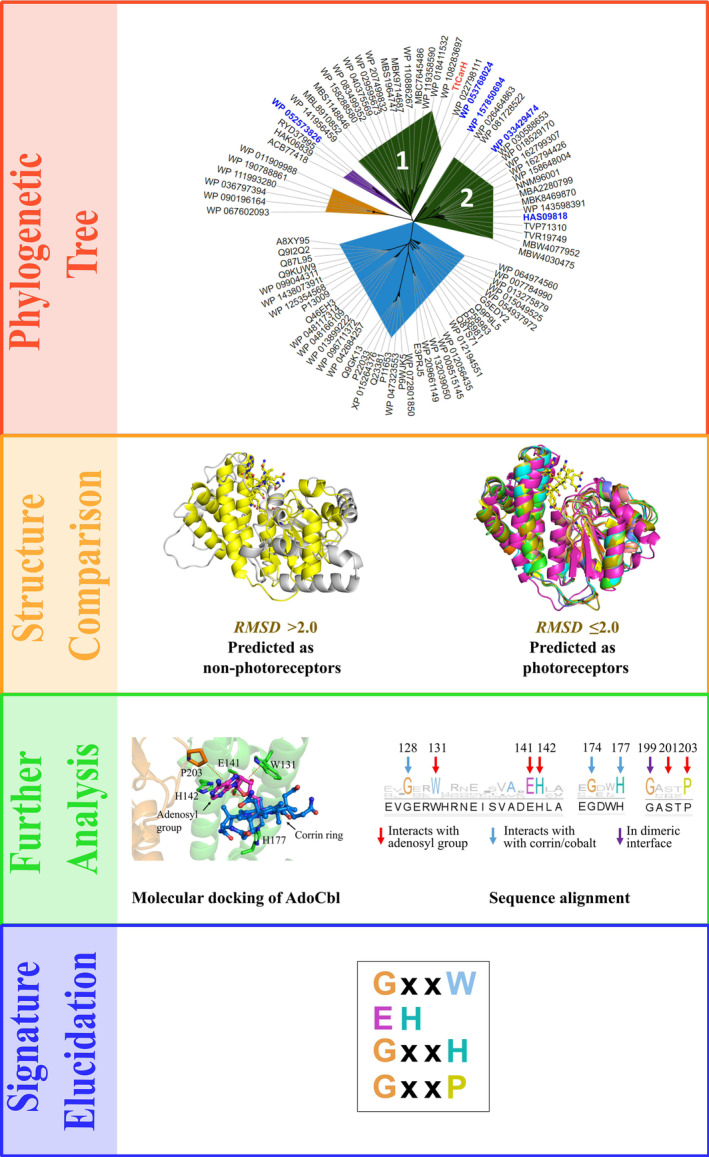
Identification of signatures for light response in cobalamin‐binding proteins by SignatureFinder workflow
The phylogenetic tree file generated by SignatureFinder was visualised by ITOL [ref. 42] (Fig. 2) to show the evolutionary relationship between TtCBD and input sequences. The input sequences show diversity by nesting in different branches of the phylogenetic tree, although they are all homologous CBDs. Sequences in blue, orange and purple areas are distant from TtCarH in both evolution and structure (for structural annotation, see Table S1): generally, structures in blue resemble CBD‐dependent enzymes; structures in orange resemble flavoprotein AppA; sequences in purple were not modelled because sequence identity to the template protein was lower than 20%. AppA is an interesting photoreceptor first discovered in Rhodobacter sphaeroides which contains a BLUF domain that uses FAD to sense blue light [ref. 43]. This protein is therefore a good contrasting protein to use against TtCarH as it uses different cofactors and is structurally very different. There are two groups of sequences resembling CarH in structure (dark green area). Group 1, which contains TtCarH, is presumably from the MerR family, like TtCarH itself. Intriguingly, Group 2 shows a difference in evolution but structural resemblance to TtCarH. The two groups are both defined to be putative light‐responsive CBDs by SignatureFinder. The AdoCbl was docked into the binding site of each putative CBD model. Sequences were then aligned, and conservation of each site was viewed by Jalview [ref. 44] (Fig. 2). By comparison with other decoy CBDs, there are several conserved motifs that have been identified between residues 128 and 177 (GxxW, EH, GxxH, GxxxP), which interact with different regions of the AdoCbl cofactor in light‐responsive CBD‐containing proteins. Among them, W131, E141, H142 and H177 have been proven to be key residues in binding AdoCbl in CarH [ref. 20]. It is noteworthy that not only sequences from the MerR family, but also sequences with ambiguous annotations, such as ‘hypothetical protein’, were predicted to be light‐responsive by SignatureFinder. We reason that these putative unknown sequences could be new photoreceptors.
Structural modelling and experimental validation of new CBD‐containing photoreceptors
Five sequences were selected for experimental validation (marked in blue font in Fig. 2): 1 sequence from group 1 which shares the same branch with TtCarH, 3 new sequences from group 2 which are ambiguously annotated in public databases and 1 sequence from the branch close to group 1 which does not contain the light‐responsive signature as a negative control. All 5 genes were synthesised and expressed in E. coli (Table S1). Full‐length structures of the 5 proteins were modelled by AlphaFold [ref. 45] to view the functional domains along with CBD sensor domains (Fig. 3).
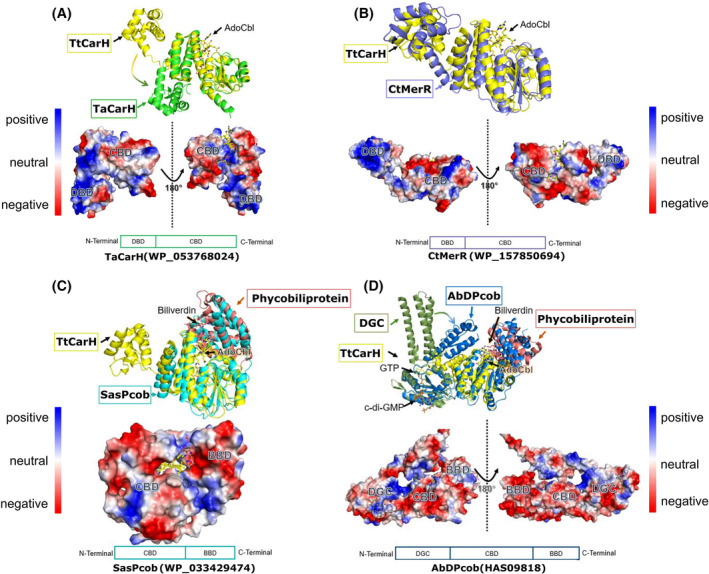
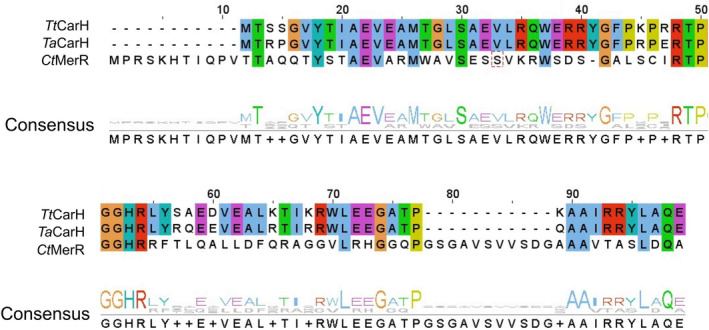
Both TaCarH (WP_053768024) and CtMerR (WP_157850694) show positively charged N‐terminal domains for binding DNA (Fig. 3A,B), indicating they are likely transcription regulators from the MerR family. TaCarH is presumably a CarH protein as it shares high sequence identity with TtCarH (75% for full length and 87% for DNA‐binding domain). However, CtMerR shows obvious variance in sequence particularly in the DNA‐binding domain with 32% sequence identity for full‐length protein and 25% for the DNA‐binding domain (Fig. 4). There are also significant structural differences to CarH (full‐length RMSD = 5.8 Å), suggesting that CtMerR is a new light‐responsive transcriptional regulator with an alternative DNA‐binding site. For SasPcob (WP_033429474), there is a predicted biliverdin‐binding region (BBD) on the C‐terminal side of the CBD (Fig. 3C) and in the case of AbDPcob (HAS09818) a diguanylate cyclase (DGC) domain was predicted (Fig. 3D), followed by a CBD and a BBD. We characterised these two proteins probing their cofactor binding and structural changes in response to light in our recent manuscript [ref. 29]. These two proteins found using SignatureFinder are members of the newly identified Photocobilin family. Using the biliverdin cofactor, these proteins can activate the adenosylcobalamin photochemistry observed in TtCarH even using red light. The identification of these proteins highlights the ability of SignatureFinder to identify light‐sensitive proteins.
The full‐length structures of CtMerR, SasPcob and AbDPcob suggest they are new light‐induced photoreceptors, although they are all ambiguously annotated in the public database (Fig. 3; Table S1). On the other hand, WP_052573826 (HaAI‐2E), which does not contain the light‐responsive signature showed a quorum‐sensing signal autoinducer‐2 exporter (AI‐2E) domain, CBD domain and a DGC domain in the AlphaFold model (Fig. S1). The CBD of HaAI‐2E has limited space for accommodating the AdoCbl in comparison with TtCarH as there is an arginine in the equivalent position of the conserved H177 residue that ligates to the cobalt. The light responsiveness of the CBD is not indicated in the annotation from the public database, and it is not predicted to be a photoreceptor by SignatureFinder.
Biophysical analysis of Signaturefinder‐identified photoreceptor proteins
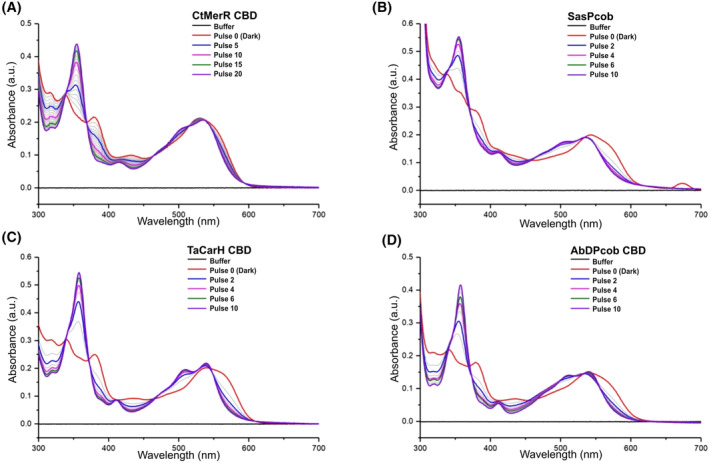
Encouraged by the in silico analysis, the selected proteins were expressed (Fig. S2) and their light response validated by absorbance spectroscopy measurements (Fig. 5; Fig. S3). For improved yield and spectral experiments, all relevant proteins were produced as truncated constructs with the DNA‐binding domain or the DGC domain removed (Fig. S2). TaCarH, CtMerR, SasPcob and AbDPcob, all show a clear shift in the absorbance maximum compared to free AdoCbl indicating that AdoCbl is able to bind to these proteins. This is highlighted by the shift to 540 nm with a shoulder at 560 nm in the dark form (Fig. 5). In each case, upon illumination with green light, there is a decrease in absorbance at 560 nm and the appearance of a new peak at 356 nm. This indicates the formation of hydroxocobalamin (OHCbl) or a water‐ligated cobalamin, which forms after Co‐C bond photolysis and confirms that all these proteins respond to green light. It appears that TaCarH, SasPcob and AbDPcob require lower levels of illumination than CtMerR to reach the final light state, suggesting that they have different quantum efficiencies that are likely to arise from different arrangements of the AdoCbl binding site. For protein HaAI‐2E, which does not have the light response signature, only OHCbl binding was observed and neither adenosylcobalamin nor methylcobalamin (MeCbl) were able to bind (Fig. S3), indicating it is a light‐independent CBD. These spectroscopy measurements are consistent with the predictions from SignatureFinder. HaAI‐2E may not be able to bind AdoCbl or MeCbl due to steric constraints or unfavourable interactions with nearby residues.
To investigate the oligomerisation state of the new photoreceptors in solution, we conducted analytical size exclusion chromatography (Fig. S4), multi‐angle light scattering (SEC‐MALS) (Fig. S5) and native mass spectrometry (Fig. S6) on the CBD of AbDPcob, CtMerR, TaCarH and the full‐length SasPcob proteins. Analytical size exclusion indicated after light exposure the AbDPcob CBD and SasPcob proteins did not change oligomerisation state whereas CtMerR CBD and TaCarH CBD did change oligomerisation state (Fig. S4). SEC‐MALS suggested CtMerR and TaCarH are stable as a tetramer in the dark and disassemble into monomers after light illumination, in a similar manner to TtCarH. Surprisingly, SasPcob remains as a monomer in both states, whereas AbDPcob forms a dimer in both dark and light conditions (Fig. S5). Though powerful analysis tools, SEC‐MALS and analytical size exclusion assume proteins are compact and spherical, and therefore, masses can be inaccurate [ref. 46, ref. 47]. However, these results were consistent with those attained by native mass spectrometry for all proteins except TaCarH. Interestingly, SEC‐MALS and analytical size exclusion calculated the protein mass to be approximately 13 kDa lower than the mass calculated from native mass spectrometry and the predicted mass from the FASTA sequence. This suggests this protein has a more compact conformation than a protein of this size typically has (Fig. S6).
Native mass spectrometry showed TaCarH and CtMerR form tetramers which monomerise in response to light. In addition, we observed the masses of the tetramers corresponded to the dark state with adenosylcobalamin bound and the light state with hydroxocobalamin bound (Fig. S6). This suggested that CtMerR and TaCarH are likely to control gene expression in a similar way to TtCarH in response to light. However, SasPcob and AbDPcob behave differently, and their function and behaviour regarding the light regulation required further investigation as discussed in our additional manuscript [ref. 29]. Interestingly, it was discovered that SasPcob does change oligomerisation state from a dimer to monomer in response to light when biliverdin is bound [ref. 29]. For all experiments described in this paper, biliverdin was not bound to any protein. Without biliverdin bound, we observed SasPcob was unable to dimerise and remained a monomer in the dark and light with slight changes in mass corresponding to adenosylcobalamin and hydroxocobalamin bound in the dark and light states, respectively (Fig. S6). The changes in oligomerisation state and the lack of stability observed by SasPcob and AbDPcob without biliverdin bound suggests this cofactor is needed for proper protein folding, particularly in the dark state. All masses observed are summarised in Table S2.
Structural analysis of conserved interactions responsible for light sensing in CBD
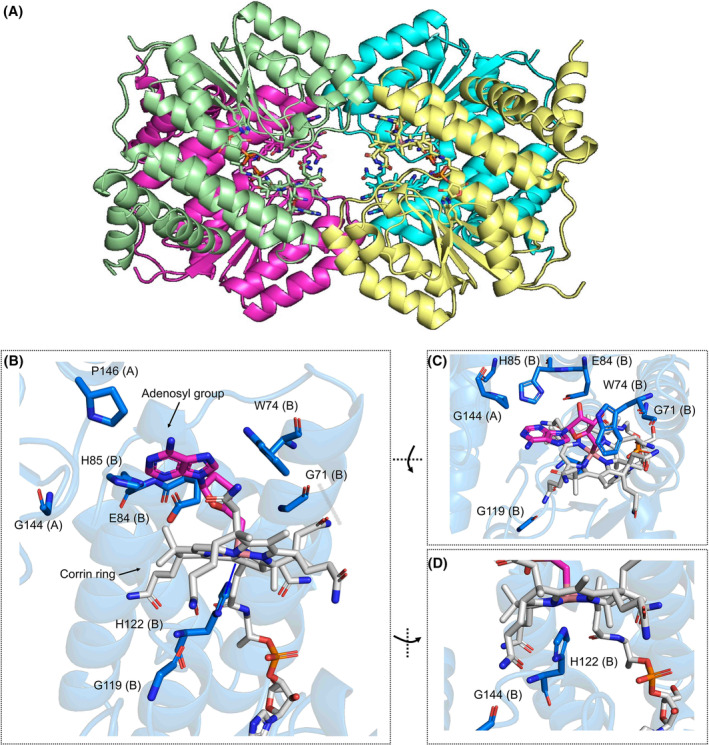
Although it was not possible to obtain crystal structures for all the selected proteins, we were able to crystallise in the dark CBD domain of CtMerR (Figs 6, 7; Table S3). CtMerR crystallised in the P 31 2 1 space group with a dimer in the asymmetric unit. The functional tetramer can be generated with the symmetry elements (Fig. 6A). The structure also highlights how the identified light‐responsive signature elements (GxxW, EH, GxxH, GxxxP) interact with AdoCbl (Figs 6, 8; Fig. S7). His204 forms the lower axial ligand to the cobalt. Trp156 and Pro228 from the neighbouring monomer interact by hydrophobic interactions (Fig. 6A) while Glu84 forms a hydrogen bond (Fig. 6B) with the hydroxyl moiety of the upper adenosyl group. Main chain carbonyl groups Gly153 and Gly201 form hydrogen bonds with the corrin ring amide moieties (Fig. 6B,C). There are no obvious interactions between Gly226 and AdoCbl, although it presumably plays a role in modulation of the flexibility of the loop at the dimer interface. An alignment between the CtMerR and TtCarH crystal structures show the corrin ring and adenosyl group is accommodated by a conserved region formed by a helical bundle in the N‐terminal domain and the nearby Rossmann fold (Fig. S8), despite clear differences in more distant regions of the Rossmann fold structure.
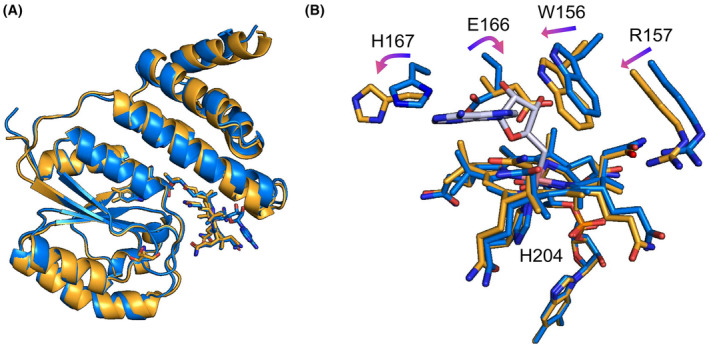
Attempts to expose dark‐state crystals to light led to loss of diffraction, while light‐exposed protein would not crystallise. Light exposure of dark‐state crystals was undertaken in anaerobic conditions, following observations that WT and a variant forms of TtCarH required oxygen for effective monomerisation resulting in large conformation changes that likely affect crystal packing [ref. 20]. For CtMerR, light exposure under anaerobic conditions led to structural changes without monomerisation. The alpha helices within the Rossman fold show some movement when the protein is exposed to light, up to 2.0 Å around Q190 (Fig. 7A). However, the largest changes are present in the environment surrounding the AdoCbl cofactor. The dark and light‐adapted monomers overlaid with a r.m.s.d of 0.525 Å (calculated for all backbone Cɑ atoms). The cobalt‐carbon bond is cleaved, and the adenosyl moiety is absent from the structure leaving aquo/hydroxocobalamin bound (Fig. 7B). This is observed in TtCarH [ref. 20] and SasPcob [ref. 29] upon light exposure suggesting a similar mechanism of photoactivation. Several key residues also rotate in this binding pocket including the signature residues E166 and H167.
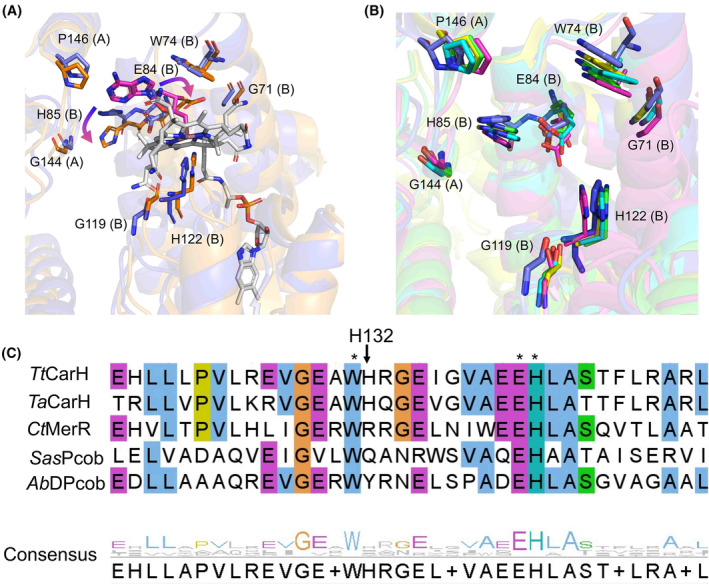
The TtCarH structural studies showed a horizontal shift of the cobalamin to form a bis‐His ligated cobalamin upon illumination [ref. 20]. However, this is not observed in the anaerobic CtMerR light‐exposed structure, which contains an arginine rather than a histidine in the equivalent position (Fig. 8C). A similar observation is made for the TtCarH H132A mutant, which undergoes modest structural rearrangement after light exposure but is unable to monomerise [ref. 48]. Sequence alignments show variation at this position; the equivalent residue for SasPcob is a glutamine and for AbDPcob is a tyrosine (Fig. 8C). Instead of a bis‐His ligation to stabilise the cobalamin in the light state, a reorientation of the Glu166 residue in CtMerR which allows formation of water‐mediated hydrogen bonding interaction with the cobalt (Fig. 7B). This suggests that the bis‐His ligated light state observed in TtCarH is representative of the CBD light‐responsive state and that different photochemical mechanisms may be used in CtMerR, SasPcob and AbDPcob. Furthermore, the alignment of AlphaFold models of SasPcob, AbDPcob and TaCarH to the crystal structures of TtCarH and CtMerR highlighted a similar pattern of interactions with the adenosyl group of AdoCbl and the key signature residues (Fig. S8). The adenosyl group is vital for the nature of these proteins due to the cobalt–carbon bond cleavage induced by light resulting in structural rearrangement [ref. 48]. This provided further confirmation of the proposed signature for light sensing in CBD‐containing proteins.
Database mining for further green‐light‐sensitive proteins
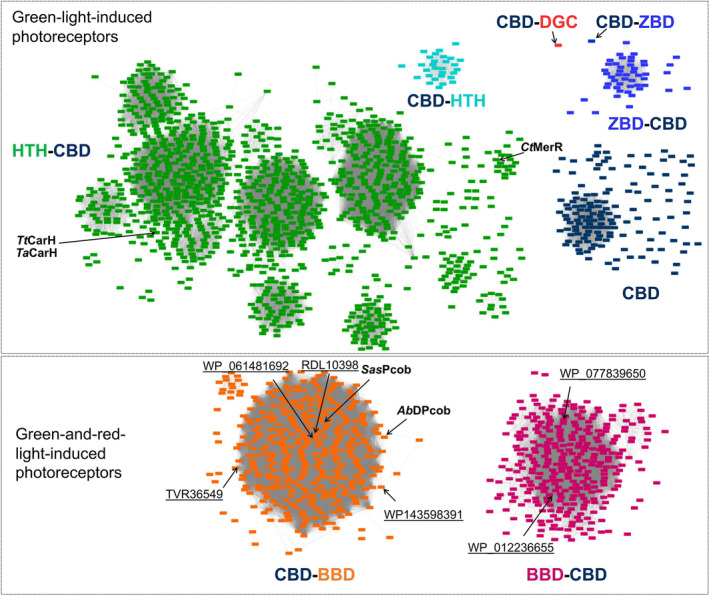
The light‐responsive CBD signature was used to search for further green‐light‐sensitive proteins in the public database. In this analysis, 1561 out of 128 000 CBD‐containing sequences (InterPro database entry: IPR006158) were found to have the proposed signature after clustering using a sequence identity of 80%. The sequence similarity network was constructed to show the diversity of the sequences (Fig. 9). A 3D model from each cluster was predicted by AlphaFold, and the putative functional domains were annotated according to the model (Fig. S9). The new photoreceptors CtMerR, SasPcob and AbDPcob identified in the current work and the previously reported CBD‐containing photoglobins [ref. 49] were also retrieved from the public database by using this approach, further highlighting the effectiveness of the signature as a filter for retrieving light‐responsive CBDs. Light‐responsive CBDs can be found fused to output domains such as helix‐turn‐helix (HTH) domains, diguanylate cyclase (DGC) domain, biliverdin‐binding domain (BBD), GAF domains and MEthanogen/methylotroph DcmR Sensory (MEDS) at both the N‐terminal and C‐terminal region, indicating it is a versatile building block for multi‐domain proteins to regulate manifold activities inside cells in response to light. Interestingly, database mining revealed more BBD‐CBD fusion proteins, the recently discovered photoglobin family, thus expanding the toolbox of dual‐light‐regulated photoreceptors [ref. 49]. The interactive sequence similarity network for identifying novel green‐light‐induced photoreceptors is now accessible to the public.
Conclusions
As the only characterised green‐light‐induced B12‐dependent photoreceptor, CarH has shown exciting potential in light‐dependent biotechnological applications [ref. 24, ref. 25, ref. 26, ref. 27]. The discovery of new green‐light‐sensitive photoreceptors to diversify and expand the optogenetics toolbox is now a major bottleneck in this process. Although (Ado)Cbl is the chromophore used by CarH for sensing green light, not all cobalamin‐binding proteins are photoreceptors and the characteristic features responsible for photoreception are currently unknown. We have now developed a computational workflow, SignatureFinder, to identify the signature for binding a ligand of interest by combining structural and sequence analysis of input sequences, which facilitates further database mining of the resulting signature. The power of this approach has been demonstrated by applying SignatureFinder to identify light‐responsive signatures (GxxW, EH, GxxH, GxxxP) in CBD‐containing proteins, which was then validated by spectroscopic and structural analysis. Additionally, we characterised a new family of proteins, called the photocobilins, containing adenosylcobalamin and biliverdin [ref. 29]. Encouraged by these results, SignatureFinder has now been constructed into a web server freely accessible to the public and is expected to find signature sequences for binding any ligand of interest, such as adenosylcobalamin. Using SignatureFinder, we have identified many more possible green‐light‐sensitive proteins that could be used for future optogenetics applications and have published the sequence similarity network on NEDx, which has a user‐friendly interface that is freely accessible. The tools and information provided in this study will save a huge amount of time and cost in the discovery of new photoreceptors to significantly expand the optogenetics toolbox.
Materials and methods
Signaturefinder workflow
The Signaturefinder workflow is based on the combination of homology modelling, molecular docking, sequence alignment and phylogenetic tree construction. The details for each step are described below:
Homology modelling
A three‐dimensional structure for each sequence was built by using MODELLER [ref. 50]. The structure from the protein data bank with the highest sequence identity was used as a template for constructing the model. Every model was aligned by PyMol to the input reference structure and root‐mean‐square deviation (RMSD) values between each model and the reference structure were computed using the Cα atoms. An RMSD threshold was set for picking sequences that have similar structures to the reference structure. Outliers were removed from further data processing. Outliers were identified as proteins with RMSD values over 2.0 Å between the models and reference.
Molecular docking
The ligand from the input reference structure was extracted. The centre of mass of the ligand was used as the centre of the docking box. The docking box size was calculated by using script eBoxSize.pl [ref. 51] considering the gyration radius of the substrate, which improves the docking accuracy [ref. 51]. In the case of CBDs, a cubic docking box with the putative models were aligned to the reference structure and the ligand was docked to the docking box. Molecular docking was carried out by AutoDock vina [ref. 52] with exhaustiveness value 10.
Sequence alignment
MUSCLE [ref. 53] was used to produce sequence alignments for the putative sequences. The conservation of each site was viewed by Jalview [ref. 44].
Phylogenetic tree
MUSCLE [ref. 53] was used to make sequence alignments for the input sequences. Raxml‐ng [ref. 54] based on maximum likelihood (ML) methods was used to calculate the phylogenetic tree for the input sequence. Popular protein evolutionary models DAYHOFF, DCMUT, LG, JTT, MTREV, WAG, RTREV, CPREV, VT, BLOSUM62 and MTMAM were evaluated and the best model which gives the lowest AIC (Akaike Information Criterion) score were used to generate the final tree for the input sequences [ref. 55]. The best tree out of 100 ML trees in newick tree format were used to view the evolutionary relationship of all sequences by ITOL [ref. 42].
SignatureFinder web server
The web server SignatureFinder is accessible to the public by https://enzymeevolver.com/SignatureFinder. The example input files are attached as supplementary files.
Protein expression and purification
All chemicals were ordered from Sigma‐Aldrich (St. Louis, MI, USA). The selected genes were synthesised by GeneArt (Thermo Fisher, Loughborough, UK) and subcloned into the pET21a vector (MilliporeSigma (Novagen), Malvern, UK). The recombinant plasmids were then transformed into E. coli BL21(DE3) cells for protein expression. Transformed cells were grown in auto induction LB medium (Formedium™, glucose/lactose ratio 1:4) containing 50 μg·mL−1 ampicillin for 24 h at 25 °C. Cells were harvested by centrifugation at 6000 g for 10 min at 4 °C, resuspended in lysis buffer (20 mm HEPES pH 7.0, 500 mm NaCl, 25 mm imidazole) supplemented with protease inhibitor cocktail and lysed by a cell disruptor (Constant Systems). The cell lysate was centrifuged at 51 000 g for 1 h at 4 °C to remove cell debris. The soluble cell lysate was loaded onto a His‐trap column, and bound protein was eluted with elution buffer (20 mm HEPES pH 7.0, 500 mm NaCl, 250 mm imidazole). The peak fractions were collected and incubated with AdoCbl for at least 2 h. The sample was then loaded onto the size exclusion column (HiLoad 16/600 Superdex 200) for further purification and removal of free ligands. The protein fractions with ligand bound were collected for further experiment.
Light titration with 530 nm LED
The absorbance spectra of all the samples were collected using a Cary 50 spectrophotometer (Agilent Technologies, Cheadle, UK). The TDS3032C 300 MHz Digital Phosphor Oscilloscope (Tektronix, Bracknell, UK) and TGP110 10 MHz Pulse Generator with Delay (Thurlby Thandar Instruments, Huntingdon, UK) were used to generate a 120 ms 530 nm LED (Thorlabs Inc. Ely, UK) pulse. After each LED pulse, the spectrum was collected until there were no further significant changes in absorbance. The experiment was carried out under a dim red light to prevent any light illumination of the sample.
Sec‐MALS
Size exclusion chromatography coupled with multi‐angle light scattering was carried out to investigate the oligomeric state of new photoreceptors. For analysis of light‐activated oligomeric changes, protein was analysed in the dark. Light‐exposed samples were exposed for 5 min using a 530 nm LED as described above. MiniDAWN TREOS MALS detector and Optilab rEX refractive index meter (Wyatt, Santa Barbara, CA, USA) were used to collect light scattering signals. An Agilent G7110B HPLC pump, degasser and autoinjector (Agilent, Santa Clara, CA, USA) was used to auto load the samples (50 μL each run, 1 mg·mL−1). For size exclusion separation, a Superdex 200 10/300 GL column using 20 mm HEPES, pH 6.8, 150 mm NaCl buffer at 1 mL·min−1. All results were processed according to referenced protocol [ref. 47]. Raw data were exported and plotted using Origin 9.0 software (OriginLab, Northampton, MA, USA). Further details are available in [ref. 29].
Analytical size exclusion
Protein was purified as detailed previously and kept at −80°C before analysis. Samples were defrosted, diluted to 20 μm and 500 μL loaded onto a Superdex 200 pg 10/300 GL column from Cytiva Life Sciences. Protein was exposed to ambient light for 10 min at room temperature. The column was equilibrated in 20 mm HEPES, 150 mm NaCl pH 8. The mass of the protein was determined from a calibration curve generated using two molecular weight kits. The first was a Gel Filtration Calibration Kit from Cytiva Life Sciences containing Ferritin (440 000 Da), Aldolase (158 000 Da), Conalbumin (75 000 Da), Ovalbumin (44 000 Da), Carbonic Anhydrase (29 000 Da) and Ribonuclease A (13 700 Da). The second was the HPLC Protein Molecular Weight Markers kit from Sigma‐Aldrich containing cytochrome c (12,400 Da), myokinase (32 000 Da), enolase (67 000 Da) lactate dehydrogenase (142 000 Da) and glutamate dehydrogenase (290 000 Da).
Native mass spectrometry
Protein was analysed as previously described [ref. 48]. Briefly, protein was desalted into 200 mm ammonium acetate pH 8.0. NanoESI capillaries were prepared in house from thin‐walled borosilicate capillaries (inner diameter 0.9 mm, outer diameter 1.2 mm) (World Precision Instruments, Hitchin, UK) using a Flaming/Brown P‐97 micropipette puller (Sutter Instrument Company, Hitchin, UK). Native MS data were acquired using the Thermo Scientific Q Exactive Hybrid Quadrupole‐Orbitrap mass spectrometer (Thermo Fisher Scientific, Loughborough, UK). For all spectra generated, the spray current was kept between 0.2 and 0.3 μA with the spray voltage varying between 0.9 and 1.3 kV accordingly. The resolution used was 25 000 for all proteins except AbDPcob which used 12 500 due to high salt and instability of the protein. For MS settings, see previous work [ref. 48]. Proteins were exposed to ambient light for 5 min for light‐exposed data collection. For analysis, 5 min spectra were averaged and processed in Thermo Xcalibur (Thermo Fisher Scientific, Loughborough, UK) before further data analysis with UniDec [ref. 56].
Protein crystallisation, data collection and structure determination
The protein after size exclusion was concentrated to 10 mg·mL−1 in 20 mm HEPES pH 7.5, 150 mm NaCl. Crystallisation was performed using the sitting drop vapour diffusion technique (200 nL crystallisation reagent mixed with 200 nL protein). The dark state of CtMerR CBD crystals was obtained from 0.1 m Amino acids, 0.1 m Bicine and Tris pH 8.5, 50% v/v Glycerol and Poly(ethylene glycol) 4000. To obtain the light state structure of CtMerR CBD, the dark crystals were transferred into glove box and degassed for at least 3 days. Both crystals were cryo protected with the addition of 20% PEG 200 to the reservoir solution prior to flash cooling in liquid nitrogen. Individual datasets were collected from single cryo‐protected crystals at beamlines i03, i04 and i04‐1 (Diamond Light Source). All data were indexed, scaled and subsequently integrated with Xia2 [ref. 57]. Structure determination was initially performed by molecular replacement in Phaser [ref. 58] using a search model generated by AlphaFold2 [ref. 45]. A combination of automated and manual rebuilding and refinement in Refmac [ref. 59] and COOT [ref. 60] were used to produce the refined models. Validation with both Molprobity [ref. 61] and PDB_REDO [ref. 62] was integrated into the iterative rebuild process. Complete data collection and refinement statistics are available in Table S3. The atomic coordinates and experimental data have been deposited in the Protein Data Bank (www.pdb.org). All figures were made using open‐source PyMOL software (Version 3.0, Schrödinger, LLC, New York, USA).
Conflict of interest
The authors declare no conflict of interest.
Author contributions
NSS, DL and DJH initiated project. NSS and SZ coordinated the project. YY, AH and LJ constructed SignatureFinder workflow and the web server. YY, LJ, and FD performed bioinformatics analysis and produced the online SignatureFinder tool. LNJ, SZ, YY and MS performed protein purification. SZ and YY performed spectroscopy experiment, SEC and crystallisation experiment. LNJ performed native spectrometry and analytical gel filtration experiments. SZ and HP performed and analysed the structural elucidation of CtMerR. LNJ and YY wrote the manuscript and generated the figures. All authors discussed the results and commented on the manuscript.
Peer review
The peer review history for this article is available at https://www.webofscience.com/api/gateway/wos/peer‐review/10.1111/febs.17377.
References
- Natural photoreceptors as a source of fluorescent proteins, biosensors, and optogenetic tools.. Annu Rev Biochem, 2015. [PubMed]
- Temporally precise single‐cell‐resolution optogenetics.. Nat Neurosci, 2017. [PubMed]
- Restoration of visual function by transplantation of optogenetically engineered photoreceptors.. Nat Commun, 2019. [PubMed]
- Illuminating cell signalling with optogenetic tools.. Nat Rev Mol Cell Biol, 2014. [PubMed]
- Optogenetic approaches to study the mammalian brain.. Curr Opin Struct Biol, 2019. [PubMed]
- Natural photoreceptors and their application to synthetic biology.. Trends Biotechnol, 2015. [PubMed]
- Millisecond‐timescale, genetically targeted optical control of neural activity.. Nat Neurosci, 2005. [PubMed]
- Current topics of optogenetics for medical applications toward therapy.. Adv Exp Med Biol, 2021. [PubMed]
- Plant photoreceptors: multi‐functional sensory proteins and their signaling networks.. Semin Cell Dev Biol, 2019. [PubMed]
- Lrit1, a retinal transmembrane protein, regulates selective synapse formation in cone photoreceptor cells and visual acuity.. Cell Rep, 2018. [PubMed]
- Phytochromes and other (photo)receptors of information in plants.. Russian J Plant Physiol, 2019
- The role of cGMP‐signalling and calcium‐signalling in photoreceptor cell death: perspectives for therapy development.. Pflugers Arch, 2021. [PubMed]
- Spatiotemporal control of cell signalling using a light‐switchable protein interaction.. Nature, 2009. [PubMed]
- Structure and function of plant photoreceptors.. Annu Rev Plant Biol, 2010. [PubMed]
- Photoreceptor proteins, “star actors of modern times”: a review of the functional dynamics in the structure of representative members of six different photoreceptor families.. Acc Chem Res, 2004. [PubMed]
- Structural basis of a Phototropin light switch.. Science, 2003. [PubMed]
- Signal amplification and transduction in phytochrome photosensors.. Nature, 2014. [PubMed]
- Light‐activated DNA binding in a designed allosteric protein.. Proc Natl Acad Sci USA, 2008
- Two open states with progressive proton selectivities in the branched channelrhodopsin‐2 photocycle.. Biophys J, 2010. [PubMed]
- Structural basis for gene regulation by a B12‐dependent photoreceptor.. Nature, 2015. [PubMed]
- A new facet of vitamin B12: gene regulation by cobalamin‐based photoreceptors.. Annu Rev Biochem, 2017. [PubMed]
- Light‐dependent gene regulation in nonphototrophic bacteria.. Curr Opin Microbiol, 2011. [PubMed]
- Involvement of CarA/LitR and CRP/FNR family transcriptional regulators in light‐induced carotenoid production in Thermus thermophilus .. J Bacteriol, 2011. [PubMed]
- B12‐dependent photoresponsive protein hydrogels for controlled stem cell/protein release.. Proc Natl Acad Sci USA, 2017. [PubMed]
- Injectable, photoresponsive hydrogels for delivering neuroprotective proteins enabled by metal‐directed protein assembly.. Sci Adv, 2020. [PubMed]
- Spatiotemporal control over multicellular migration using Green light reversible cell–cell interactions.. Adv Biol, 2021
- Turning cell adhesions ON or OFF with high spatiotemporal precision using the Green light responsive protein CarH.. Chemistry, 2020. [PubMed]
- B12‐dependent photoreceptor protein as an emerging tool for materials synthetic biology.. Smart Mater Med, 2022
- Photocobilins integrate B(12) and bilin photochemistry for enzyme control.. Nat Commun, 2024. [PubMed]
- Cobalamin‐ and corrinoid‐dependent enzymes.. Met Ions Life Sci, 2009. [PubMed]
- Cobalamins and cobalamin‐dependent enzymes in Candida utilis .. J Bacteriol, 1979. [PubMed]
- Cobalamin‐dependent methyltransferases.. Acc Chem Res, 2001. [PubMed]
- Adenosylcobalamin‐dependent methylmalonyl‐CoA mutase isozymes in the photosynthetic protozoon Euglena gracilis Z.. Microbiology (Reading), 1996. [PubMed]
- The stereochemistry of the formation of the methyl group in the glutamate mutase‐catalysed reaction in clostridium tetanomorphum.. FEBS Lett, 1984. [PubMed]
- Cobalamin‐dependent methionine synthase.. FASEB J, 1990. [PubMed]
- InterPro: the integrative protein signature database.. Nucleic Acids Res, 2009. [PubMed]
- NDEx: a community resource for sharing and publishing of biological networks.. Methods Mol Biol, 2017. [PubMed]
- NDEx 2.0: a clearinghouse for research on cancer pathways.. Cancer Res, 2017. [PubMed]
- NDEx, the network data exchange.. Cell Syst, 2015. [PubMed]
- Structure‐based design of small imine reductase panels for target substrates.. ACS Catal, 2023. [PubMed]
- NCBI BLAST: a better web interface.. Nucleic Acids Res, 2008. [PubMed]
- Interactive tree of life (iTOL) v5: an online tool for phylogenetic tree display and annotation.. Nucleic Acids Res, 2021. [PubMed]
- A single flavoprotein, AppA, integrates both redox and light signals in Rhodobacter sphaeroides .. Mol Microbiol, 2002. [PubMed]
- Jalview version 2—a multiple sequence alignment editor and analysis workbench.. Bioinformatics, 2009. [PubMed]
- Highly accurate protein structure prediction with AlphaFold.. Nature, 2021. [PubMed]
- Considerations for sample preparation using size‐exclusion chromatography for home and synchrotron sources.. Adv Exp Med Biol, 2017. [PubMed]
- Characterization of proteins by size‐exclusion chromatography coupled to multi‐angle light scattering (SEC‐MALS).. J Vis Exp doi:, 2019. [DOI]
- Redox driven B(12)‐ligand switch drives CarH photoresponse.. Nat Commun, 2023. [PubMed]
- Photoglobin, a distinct family of non‐heme binding globins, defines a potential photosensor in prokaryotic signal transduction systems.. Comput Struct Biotechnol J, 2022. [PubMed]
- Comparative protein structure modeling using MODELLER.. Curr Protoc Bioinformatics, 2016
- Calculating an optimal box size for ligand docking and virtual screening against experimental and predicted binding pockets.. J Chem, 2015
- AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading.. J Comput Chem, 2010. [PubMed]
- MUSCLE: a multiple sequence alignment method with reduced time and space complexity.. BMC Bioinformatics, 2004. [PubMed]
- RAxML‐NG: a fast, scalable and user‐friendly tool for maximum likelihood phylogenetic inference.. Bioinformatics, 2019. [PubMed]
- A primer on model selection using the Akaike information criterion.. Infect Dis Model, 2020. [PubMed]
- Bayesian deconvolution of mass and ion mobility spectra: from binary interactions to polydisperse ensembles.. Anal Chem, 2015. [PubMed]
- xia2: an expert system for macromolecular crystallography data reduction.. J Appl Cryst, 2010
- Phaser crystallographic software.. J Appl Cryst, 2007. [PubMed]
- Refinement of macromolecular structures by the maximum‐likelihood method.. Acta Crystallogr D Biol Crystallogr, 1997. [PubMed]
- Features and development of Coot.. Acta Crystallogr D Biol Crystallogr, 2010. [PubMed]
- MolProbity: all‐atom structure validation for macromolecular crystallography.. Acta Crystallogr D Biol Crystallogr, 2010. [PubMed]
- The PDB_REDO server for macromolecular structure model optimization.. IUCrJ, 2014
