TRMT13 inhibits the growth of papillary thyroid cancer by targeting ANAPC4
Abstract
The recently discovered gene TRMT13 encodes a type of RNA methylase and is a member of the CCDC family (also called CCDC76). Here, we delineate its role in papillary thyroid cancer (PTC). Bioinformatics analysis shows significant TRMT13 and ANAPC4 downregulation in PTC and reveals that the expression levels of both genes are linearly correlated. Subsequent analyses confirm that both TRMT13 and ANAPC4 expressions are downregulated in PTC tissues and that this change in expression has a significant impact on cancer diagnosis. We conduct assays on PTC cells subjected to TRMT13 and ANAPC4 silencing or overexpression to assess the biological effects of these genes. We also perform rescue experiments to validate the regulatory effects of TRMT13 on ANAPC4. A nude mouse tumor model is used to evaluate the effects of TRMT13 and ANAPC4 on PTC tumorigenesis. TRMT13 expression is decreased in PTC tissues and cell lines and is positively correlated with that of ANAPC4. Cell assays reveal that TRMT13/ANAPC4 attenuates the malignancy of PTC cells by restraining cell proliferation, migration and invasion, while rescue experiments corroborate that ANAPC4 is a downstream target of TRMT13. In the nude mouse xenograft model, both TRMT13 and ANAPC4 inhibit tumor growth, and TRMT13 and ANAPC4 expression levels are significantly associated with survival. Taken together, these findings lead to the conclusion that TRMT13 inhibits PTC growth via ANAPC4, indicating a new role of TRMT13 and providing insights into the tRNA methyltransferase and coiled-coil domain-containing protein families.
Article type: Research Article
Keywords: ANAPC4, CCDC76, CCDC family, methyltransferase, thyroid cancer, TRMT13
Affiliations: Department of Endocrinology and Metabolism Punan Hospital Shanghai 200125 China; PharmaLegacy Laboratories Shanghai 201201 China; Department of Intervention Gongli Hospital Naval Medical University Shanghai 200135 China; Nursing Department Jinshan Branch of Shanghai Sixth People’s Hospital Shanghai 201599 China; Department of Endocrinology and Metabolism Gongli Hospital Naval Medical University Shanghai 200135 China
License: © The Author(s) 2021. CC BY 4.0 0
Article links: DOI: 10.3724/abbs.2024010 | PubMed: 38425244 | PMC: PMC11532200
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (3.3 MB)
Introduction
As of 2020, there were 586,202 new cases of papillary thyroid cancer (PTC) worldwide. PTC accounts for 3.0% of all cancers and is the 11 th most common cancer ref. [1]. In China, the incidence of PTC is approximately 15 cases per 100,000 person-years ref. [2], and the five-year survival rate is approximately 84.3% ref. [3]. Despite the relatively low malignancy of PTC, it is inoperable in approximately 5% of all locally advanced cases, and its rate of local recurrence is high ref. [4]. For these reasons, the mortality rate of this disease is also elevated ref. [5].
TRMT13, also known as coiled-coil domain containing 76 (CCDC76), was recently discovered. Liu et al. ref. [6] reported that cytoplasmic hTrmt13 catalyzes the 2′- O-methylation of tRNAs, whereas nuclear hTrmt13 directly binds to DNA as a transcriptional coactivator. However, these functions are mutually exclusive. ANAPC4 or APC4 is an E3 ubiquitin ligase that regulates mitosis and the G1 phase [ ref. 7‒ ref. 10] . ANAPC4 abnormalities promote proliferation and correlate with tumor diameter in oral squamous cell carcinoma ref. [11]. ANAPC4 inhibition favors docetaxel resistance in breast cancer ref. [12]. ANAPC4 enables esophageal cancer and neural stem cells to proliferate [ ref. 13, ref. 14] . ANAPC4 transcription and expression are regulated by tRNA methylation and affect cell proliferation ref. [15]. To the best of our knowledge, however, the roles of ANAPC4 in PTC and its relationship with TRMT13 have not yet been investigated. Bioinformatics analysis of The Cancer Genome Atlas (TCGA) data revealed that both TRMT13 and ANAPC4 levels are significantly downregulated and that their expression levels are linearly correlated in PTC tissues ref. [16]. Hence, we investigated the functions and interactions of TRMT13 and ANAPC4 in the onset and progression of PTC.
The present study aimed to clarify the biological roles of TRMT13 and ANAPC4 in PTC via cell ( in vitro) and nude mouse ( in vivo) models and to examine the regulatory effects of TRMT13 on ANAPC4. The discoveries made herein could help improve both the diagnosis and treatment of PTC and provide novel insights into the molecular roles of tRNA methyltransferases and coiled-coil domain-containing proteins.
Materials and Methods
Bioinformatics analysis
The Gene Expression Profiling Interactive Analysis (GEPIA) database ( http://gepia.cancer-pku.cn/) was used to examine thyroid cancer (THCA) cases. The GEPIA database is a new interactive web server that uses a standard pipeline to analyze RNA sequencing expression data for 9736 tumors and 8587 normal samples from the TCGA and the GTEx projects. The analysis included 512 tumor and 337 normal tissue samples.
Clinical samples
Samples of papillary thyroid cancer (PTC) and adjacent normal tissues 2 cm from the tumors were collected from 30 patients at Punan Hospital (Shanghai, China). A pathologist microscopically examined the cells for the presence or absence of cancer cells. The patient inclusion criteria included (1) age range, 25–60 years; (2) pathologically confirmed PTC diagnosis; and (3) provided written informed consent. The patient exclusion criteria included (1) prior treatment for PTC; (2) co-infection; or (3) the presence of other malignant tumors. Survival time was calculated from the date of diagnosis to the date of the last visit or death. The median transfer RNA methyltransferase 13 homolog (TRMT13) or anaphase-promoting complex subunit 4 (ANAPC4) expression level was used as the threshold to assign the patients to the “high” and “low” expression groups. Receiver operating characteristic (ROC) was plotted to test the the area under the receiver operating curve (AUC) between PTC tumor and adjacent normal tissues. The research protocol was approved and the trial was supervised by the Ethics Committee of Punan Hospital.
Cell culture
The human normal thyroid cell line HTori-3 and the PTC cell lines TTA1, ACT-1, KTC-1, TPC-1, KMH-2, and CAL-62 (Mlbio, Shanghai, China) were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% (v/v) fetal bovine serum (FBS; Sigma-Aldrich, St Louis, USA). The cells were maintained in a 5% CO 2 incubator at 37°C and 95% relative humidity (RH), authenticated by the short tandem repeat (STR) method, and confirmed to be mycoplasma free.
Cell transfection
TTA1 and CAL-62 cells expressing TRMT13 or ANAPC4 were transfected so that the aforementioned genes would be silenced or overexpressed, respectively. Group 1 included the control, TRMT13 +/+, ANAPC4 +/+, TRMT13 siRNA, ANAPC4 siRNA, plasmid-NC, and siRNA-NC. Group 2 included the control, TRMT13 +/+, TRMT13 +/++ANAPC4 siRNA, TRMT13 siRNA, and TRMT13 siRNA+ANAPC4 +/+. The plasmids were constructed by amplifying the TRMT13/ ANAPC4 sequences via polymerase chain reaction (PCR) and cloning them into pIRES2-EGFP plasmids. Lentiviral transduction was performed to overexpress TRMT13/ ANAPC4. Small interfering RNAs (siRNAs) were designed to target TRMT13/ ANAPC4 sequences and were transfected into the cells with Lipofectamine TM RNAiMAX (Thermo Fisher Scientific, Waltham, USA). The sequences of siRNAs were shown as follows: ANAPC4 sense: GAGGCUCCAGUUUCCUGUAUGCAUU, antisense: AAUGCAUACAGGAAACUGGAGCCUC; and TRMT13 sense: CAGGAUGGUGGUGGCCGCAGGGAAA, antisense: UUUCCCUGCGGCCACCACCAUCCUG. siRNA-NC was purchase from Qiagen (AllStars Negative Control siRNA; Qiagen, Hilden, Germany).
RT-qPCR analysis
Cell and tissue RNAs were extracted using Trizol Reagent (Thermo Fisher Scientific) and reverse-transcribed into cDNA using High Capacity cDNA Reverse Transcription kit (Applied Biosystems, Foster City, USA). The TRMT13 and ANAPC4 mRNA expression levels were measured with SYBR Green-based qPCR ref. [17] and normalized to that of glyceraldehyde 3-phosphate dehydrogenase ( GAPDH) using the 2 ‒ΔΔCt method. The primers used were as follows: TRMT13 forward primer: GCTGCCAGTTTTGAGGAAAG, reverse primer: TGCCCACATAATGTCTCCAA; and ANAPC4 forward primer: CACCCCCTAACACAGAAGGA, reverse primer: CTGGCTTTTGCAAACACTGA.
Western blot analysis
The total proteins were isolated from the cells and tissues with radioimmunoprecipitation assay (RIPA) buffer (Sigma-Aldrich), separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), and transferred to polyvinylidene fluoride (PVDF) membranes. The membranes were blocked with 5% skim milk and then incubated with rabbit primary antibodies against TRMT13 (1/5000; Abcam, Cambridge, USA) or ANAPC4 (2 μg/mL; Abcam). The membranes were then washed with phosphate-buffered saline (PBS; Sigma-Aldrich), followed by incubation with HRP-conjugated secondary antibody (ab205718, 1/5000; Abcam). Finally, the protein bands were visualized with an enhanced chemiluminescence (ECL) kit (Amersham Pharmacia Biotech, Piscataway, USA).
Cell counting kit-8 (CCK-8) assay
One hundred microliters of cell suspension (5×10 4/mL) was added to each well of a 96-well plate (Beyotime, Shanghai, China). Ten microliters of CCK-8 solution (Beyotime) was added to each well after 24, 48, and 72 h. The plates were mixed on an orbital shaker at 37°C for 1 min to ensure uniform dispersion of the cells and reagents. The plate was then incubated for 2 h, and the OD 450 value of each well was subsequently measured with a microplate reader (LEx808, No. 25-315S; Lonza Group, Walkersville, USA).
Wound healing assay
The cells were cultured in 6-well plates (Beyotime) under the aforementioned conditions until they reached 90%–95% confluence. A 200-μL pipette tip (Beyotime) was dragged from the top to the bottom of the well to create vertical scratches. The scratch width was initially measured, the cell debris were removed, and the remaining cells were cultured in serum-free medium (Sigma-Aldrich). Images of the cultures were captured 24 h later with a microscope (Hitachi, Tokyo, Japan) to evaluate wound healing.
Colony formation assay
Two hundred cells were digested and resuspended under the aforementioned conditions for 14 days and then evenly distributed and cultured in medium. The latter was replaced every 2–3 days to ensure optimal cell growth. Macroscopic colony formation was observed, and the supernatant was discarded. The colonies were washed twice with PBS, fixed in 4% (v/v) paraformaldehyde (PFA; Sigma-Aldrich) for 15 min, stained with Giemsa (Sigma-Aldrich) for 20 min, washed again with PBS and air-dried. A gridded transparent film was placed on the plate and inverted to enumerate colonies composed of >20 cells.
Transwell assay
The upper chamber of a 24-well Transwell ® plate (Sigma-Aldrich) was coated with Matrigel ® (1:8; v/v; Corning Life Sciences, Corning, USA). Cells were starved in serum-free medium at 37°C for 24 h, followed by digestion with trypsin. Then, 100 μL of cell suspension (5×10 4 cells /mL) was added to the upper Transwell chamber, and 600 μL complete medium containing 20% (v/v) fetal bovine serum (FBS) was added to the lower chamber. After 24 h of incubation, the non-invading cells were washed away with PBS. The cells that infiltrated to the lower chamber were stained with 0.1% (w/v) crystal violet (Sigma-Aldrich) at 25°C for 20 min and fixed with 95% (v/v) ethanol. The number of cells was counted in five randomly selected fields under 400× magnification under a microscope.
Xenotransplantation assay
Male BALB/C nude mice aged 4 weeks (Charles River Laboratories, Beijing, China) were used to assess tumorigenesis. The animals were housed at 24±1°C and 60%±5% RH. Group 1 ( n=6) included the control, TRMT13 +/+, lentiviral NC, TRMT13 siRNA, and siRNA NC treatments. Group 2 ( n=6) included the control, ANAPC4 +/+, lentiviral NC, ANAPC4 siRNA, and siRNA NC treatments. Group 3 ( n=6) included the control, TRMT13 +/+, TRMT13 +/++ANAPC4 siRNA, TRMT13 siRNA, and TRMT13 siRNA+ANAPC4 +/+ treatments. Five million cells overexpressing TRMT13 or lentiviral NC, TRMT13-silenced, or siRNA NC; overexpressing ANAPC4 or lentiviral NC; or ANAPC4-silenced or siRNA NC were prepared by transfection as above described. The lentivirus infection efficiency was verified by RT-qPCR. The transfected cells were resuspended in 200 μL of PBS and administered as a single injection into the axillary region of each nude mouse. Tumor formation at the injection site was observed on day 7, and body weight and tumor volume were measured every 4 days for the next 20 days. The animals were euthanized via CO 2 inhalation when their tumor diameters were over 2 cm, after which the tumors were excised and weighed. A pathologist examined the tissue sections under a microscope to confirm xenograft thyroid tumor establishment. All animal experiment procedures used was approved by the Ethical Committee of Punan Hospital.
Statistical analysis
All the experiments were conducted independently in triplicate. Data are expressed as the mean±standard deviations (SD) and subjected to analysis of variance (ANOVA) and Tukey’s multiple comparisons tests using GraphPad v. 7.0 (GraphPad Software, La Jolla, USA). Overall survival (OS) curves were plotted by the K-M method, and statistical significance was determined by the log-rank test. P<0.05 was considered statistically significant.
Results
TRMT13 and ANAPC4 expressions are downregulated in PTC tissues and their expressions are correlated with cancer diagnosis
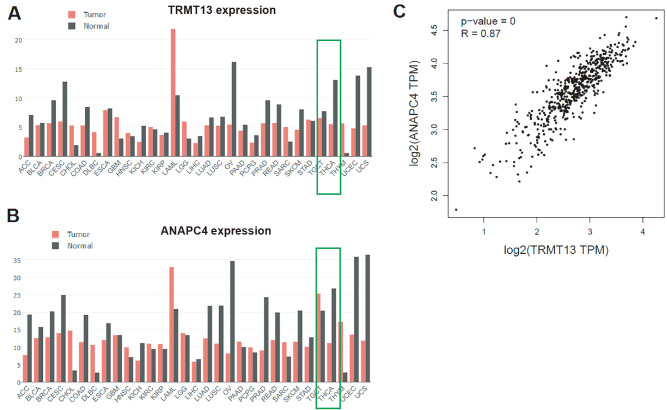
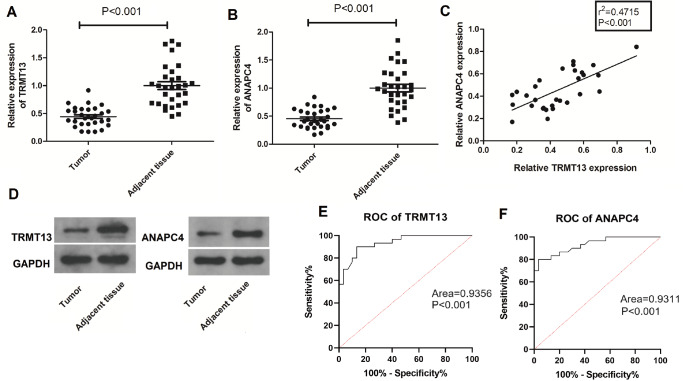
The GEPIA database includes The Cancer Genome Atlas (TCGA; https://www.cancer.gov/ccg/research/genome-sequencing/tcga) data and reveals that both TRMT13 and ANAPC4 expressions are downregulated in PTC relative to normal tissues ( Figure 1A,B). The expression levels of these genes are significantly linearly correlated ( P=0; R=0.87) ( Figure 1C). The TRMT13 and ANAPC4 mRNA levels are significantly lower in the clinical PTC specimens than in the normal tissue specimens ( Figure 2A,B). The TRMT13 and ANAPC4 mRNA levels are positively correlated in the PTC tissue samples (r 2=0.4715; P<0.001) ( Figure 2C). The TRMT13 and ANAPC4 protein levels are lower in the PTC tissues than in the normal tissues ( Figure 2D). The ROC analysis revealed that TRMT13 and ANAPC4 expressions accurately distinguished PTC from normal controls ( Figure 2E,F).
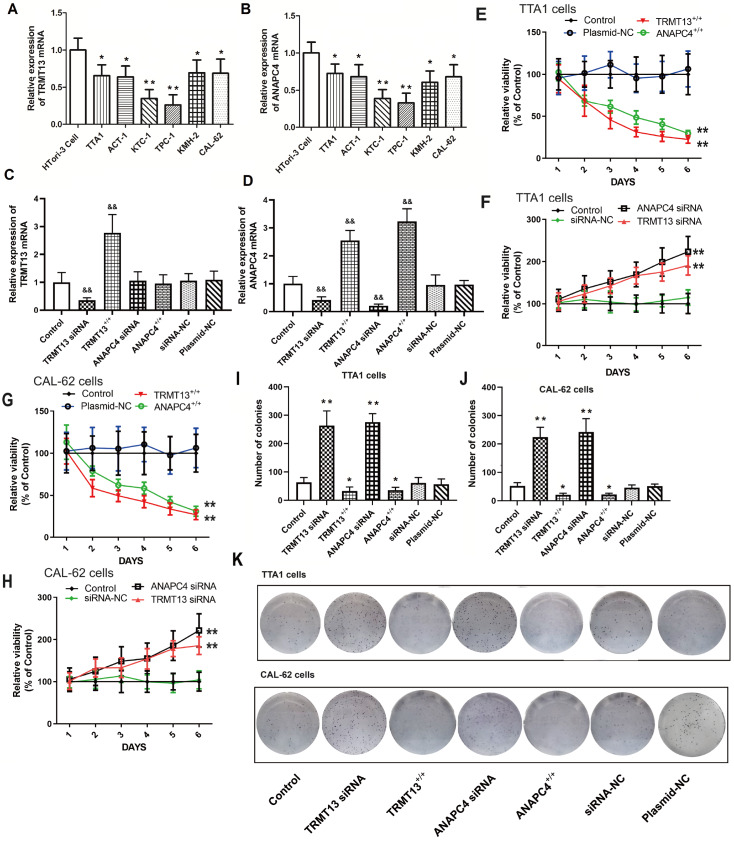
TRMT13 and ANAPC4 expressions are downregulated in PTC cells, which inhibits cell proliferation
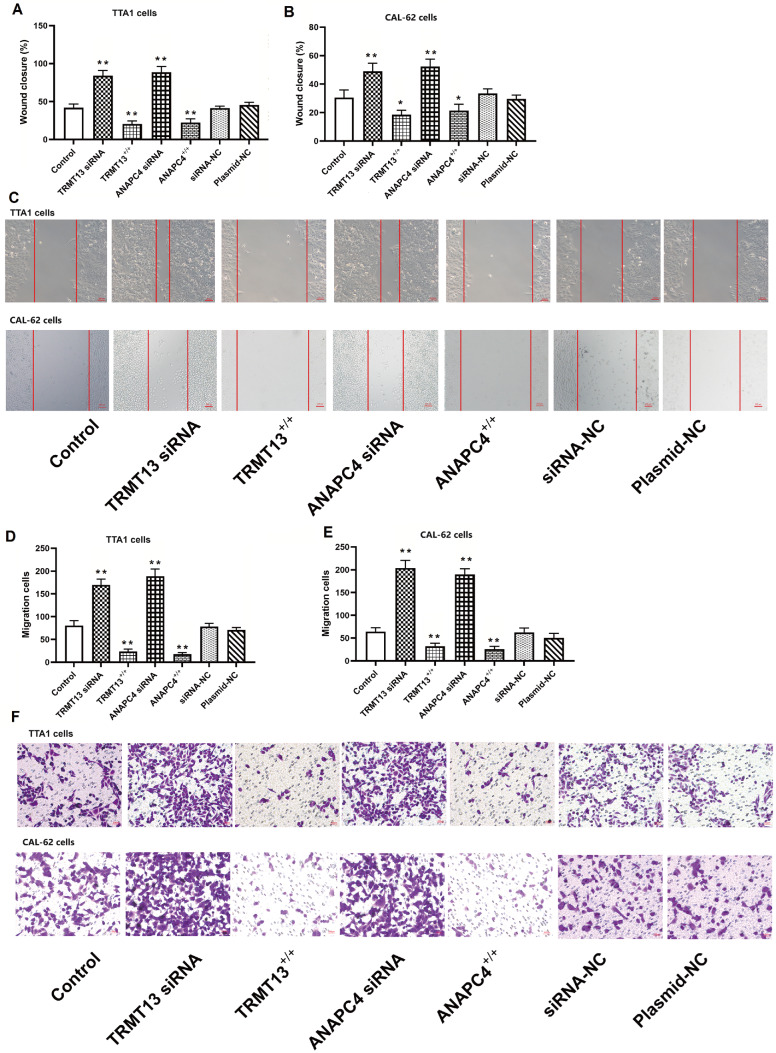
The TRMT13 and ANAPC4 expression levels were significantly lower in all six PTC cell lines than those in the normal HTori-3 cell line ( Figure 3A,B). The TRMT13 and ANAPC4 mRNA levels after plasmid exposure or siRNA treatment are shown in Figure 3C,D. Transfection with the TRMT13 plasmid or siRNA significantly altered the TRMT13 mRNA level relative to that in the untreated cells. However, neither the ANAPC4 plasmid nor the siRNA treatment changed the TRMT13 mRNA level. In contrast, the TRMT13 plasmid and siRNA treatments significantly modified the ANAPC4 mRNA level. We generated cells in which TRMT13 and ANAPC4 were overexpressed or silenced to investigate the effects of these genes and their products. The results showed that TRMT13 or ANAPC4 upregulation inhibited the proliferation of TTA1 and CAL-62 cells, whereas TRMT13 or ANAPC4 downregulation promoted the proliferation of TTA1 and CAL-62 cells ( Figure 3E–H). We then assessed the impact of TRMT13 and ANAPC4 on PTC malignancy and found that upregulation of either or both of these genes inhibited colony formation of TTA1 and CAL-62 cells ( Figure 3I–K). Compared with those in the control group, cancer cell migration and invasion were lower in the TRMT13 +/+ and ANAPC4 +/+ groups but greater in the TRMT13 siRNA and ANAPC4 siRNA groups ( Figure 4).
TRMT13 and ANAPC4 inhibit PTC progression in vivo
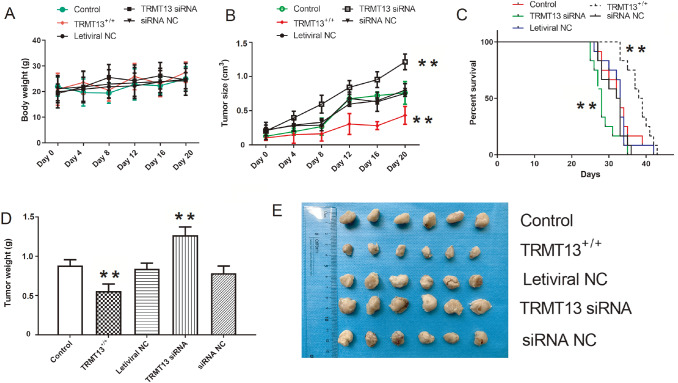
We created tumor-bearing nude mouse models from the control, TRMT13 +/+, lentiviral NC, TRMT13 siRNA, and siRNA NC groups to investigate the impact of TRMT13 on PTC progression in vivo. Body weight did not significantly change within groups during tumor formation or growth ( P>0.05; Figure 5A). However, TRMT13 overexpression significantly reduced tumor volume, whereas TRMT13 silencing significantly promoted tumor growth and reduced survival ( Figure 5B,C). The TRMT13 siRNA and TRMT13 +/+ groups presented larger and smaller mean tumor masses, respectively, than the control group ( Figure 5D). Figure 5E showed the inhibitory effects of TRMT13 on PTC progression in vivo.
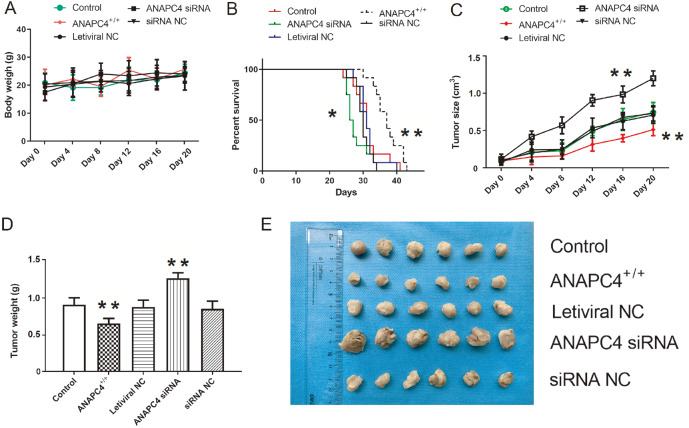
We constructed xenotransplantation models using cells from the control, ANAPC4 +/+, lentiviral NC, ANAPC4 siRNA, and siRNA NC groups to investigate the effects of ANAPC4 on PTC progression in vivo. Body weight did not significantly change within groups during tumor formation or growth ( P>0.05; Figure 6A). However, ANAPC4 overexpression significantly reduced tumor volume, whereas ANAPC4 silencing significantly promoted tumor growth and reduced survival ( Figure 6B,C). The ANAPC4 siRNA and ANAPC4 +/+ groups presented larger and smaller mean tumor masses, respectively, than the control group ( Figure 6D). Figure 6E showed the inhibitory effects of ANAPC4 on PTC progression in vivo.
ANAPC4 reverses the effects of TRMT13 silencing on PTC cells
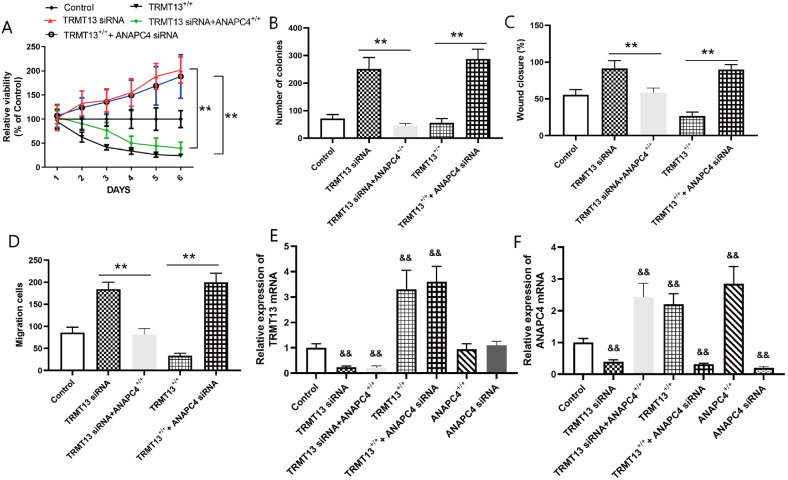
We silenced ANAPC4 combined with overexpressing TRMT13 or silenced TRMT13 combined with overexpressing ANAPC4 to investigate the involvement of ANAPC4 in the mechanism by which TRMT13 regulates PTC progression. We assigned the PTC cells to the control, TRMT13 +/+, TRMT13 +/++ANAPC4 siRNA, TRMT13 siRNA, and TRMT13 siRNA+ANAPC4 +/+ groups. The results demonstrated that silencing of ANAPC4 reversed the inhibition of PTC cell proliferation and colony formation caused by TRMT13 overexpression, while ANAPC4 overexpression impeded PTC progression by silencing TRMT13 ( Figure 7A,B). Moreover, cell migration and invasion were significantly greater in the TRMT13 +/++ANAPC4 siRNA group than in the TRMT13 +/+ group. In contrast, cell migration and invasion were significantly lower in the TRMT13 siRNA+ANAPC4 +/+ group than in the TRMT13 siRNA group ( Figure 7C,D). These findings suggest that ANAPC4 contributes to the inhibitory effect of TRMT13 on PTC. We then assigned the PTC cells to the control, TRMT13 +/+, TRMT13 +/++ANAPC4 siRNA, TRMT13 siRNA, TRMT13 siRNA+ANAPC4 +/+, ANAPC4 +/+, and ANAPC4 siRNA groups and measured their TRMT13 and ANAPC4 mRNA levels. Figure 7E,F showed that the TRMT13 plasmid and siRNA treatments significantly altered the TRMT13 mRNA level, while the ANAPC4 plasmid and siRNA treatments did not. In contrast, TRMT13 plasmid and siRNA treatment significantly altered ANAPC4 mRNA level.
ANAPC4 reverses the effects of TRMT13 silencing on PTC progression in vivo
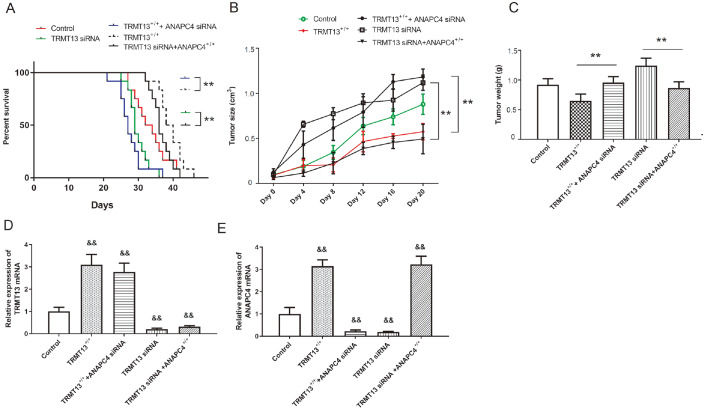
We then constructed xenograft models by injecting nude mice with cells from the control, TRMT13 +/+, TRMT13 +/++ANAPC4 siRNA, TRMT13 siRNA, or TRMT13 siRNA+ANAPC4 +/+ groups. TRMT13 overexpression and silencing improved and worsened survival, respectively, whereas ANAPC4 silencing counteracted the improvement in survival afforded by TRMT13 upregulation. While TRMT13 downregulation worsened mouse survival, ANAPC4 overexpression improved survival ( Figure 8A). Tumor volume and mass were greater in the TRMT13 +/++ANAPC4 siRNA group than in the TRMT13 +/+ group and lower in the TRMT13 siRNA+ANAPC4 +/+ group than in the TRMT13 siRNA group ( Figure 8B,C). The relative TRMT13 and ANAPC4 expression levels in the tumor tissue are shown in Figure 8D,E. While the TRMT13 plasmid and siRNA treatments significantly altered TRMT13 mRNA level, the ANAPC4 plasmid and siRNA treatments had no significant effect on TRMT13 mRNA level. In contrast, the TRMT13 plasmid and siRNA treatment significantly altered ANAPC4 mRNA level.
Discussion
Dysregulated gene expression is characteristic of all malignancies and neoplasms, including PTC. According to GeneCards ( https://www.genecards.org/), TRMT13 is implicated in tRNA methylation. The tRNA methyltransferase family has been extensively studied. METTL1-mediated m7G tRNA modification enhances lenvatinib resistance in hepatocellular carcinoma (HCC) ref. [18]. The N7-methylguanosine (m7G) tRNA modification may modulate autophagy in certain cancers ref. [19]. Methyltransferase-like protein 8-dependent mitochondrial tRNA m3C modification mediates the epitranscriptomic regulation of cortical neurogenesis ref. [20].
TRMT13 ( CCDC76) belongs to a large, extensively studied gene family. CCDC50-mediated autophagy regulates NLRP3 inflammasome activation ref. [21]. HPV-CCDC106 integration promotes cervical cancer progression ref. [22]. To the best of our knowledge, however, the biological function of TRMT13 in PTC has not been reported, and there is limited information on its relationship with cancer onset and progression. The TRMT13 gene ( CCDC76) was first described in a study by Weinmann et al. ref. [23]. The authors performed a wide-ranging chromatin immunoprecipitation (ChIP) assay targeting the αE2F transcription factor (TF) and selected 68 targets, including CCDC76 ref. [23]. In terms of size and domain architecture, hCCDC76 resembles scTrm13 except that the latter has two estimated zinc finger domains, whereas the former has only one ref. [24]. Liu et al. ref. [6] reported that hTrmt13 catalyzes 2′- O-methylation in tRNAs and, in contrast, coactivates transcription ref. [6]. These findings suggest that RNA-modifying enzymes may also have noncatalytic functions.
Here, bioinformatics analyses revealed that TRMT13 is downregulated in PTC and that its expression is inversely correlated with cancer cell growth/tumor progression ( Figure 1). qPCR and western blot analysis demonstrated that TRMT13 was downregulated in clinical cancer tissue specimens. ROC analyses indicated that TRMT13 expression significantly influenced patient cancer diagnosis. Hence, TRMT13 might suppress PTC tumors. We also employed TRMT13-silenced and TRMT13-overexpressing PTC cells to confirm the effects of these treatments on PTC cell malignancy ( Figure 3). Treatment with the TRMT13 plasmid or siRNA significantly altered TRMT13 mRNA level, whereas treatment with the ANAPC4 plasmid or siRNA did not. However, treatment with the TRMT13 plasmid or siRNA significantly modified ANAPC4 mRNA levels. Thus, TRMT13 is upstream of ANAPC4 in the signaling pathway.
Experiments in nude mice demonstrated that TRMT13 significantly suppressed PTC tumors in vivo ( Figure 5). Thus, TRMT13 may participate in PTC onset and progression. However, these findings contradict those reported by Liu et al. ref. [6], who identified hTrmt13 as a breast and cervical cancer oncogene. This apparent discrepancy may be explained by the functional differences in genes among various types of cancer. ANAPC4 is a component of the anaphase-promoting complex/cyclosome (APC/C), which regulates the cell cycle by targeting specific proteins for degradation. APC/C dysregulation has been implicated in different malignancies and neoplasms. Bradfield et al. ref. [25] reported that ANAPC4 expression was significantly associated with favorable prognosis in patients with endometrioid cancer. siRNA screening revealed that ANAPC4 is involved in chromosomal instability tolerance in diploid cells ref. [26]. ANAPC4 inactivation confers resistance to multiple TTK protein kinase inhibitors in triple-negative breast cancer ref. [27]. Here, bioinformatics analysis revealed that ANAPC4 is significantly downregulated in PTC cells ( Figure 1) and tissues. ROC analysis demonstrated that ANAPC4 significantly influences cancer diagnosis. Therefore, ANAPC4 may suppress PTC tumor onset and progression.
To the best of our knowledge, however, there is no known direct relationship between TRMT13 and ANAPC4 expression. qPCR analysis indicated that ANAPC4 expression was significantly lower in PTC tissue than in normal tissue and was positively correlated with TRMT13 expression. Here, ANAPC4 overexpression inhibited PTC cell growth, while ANAPC4 silencing promoted PTC cell growth and tumor growth ( Figures 3 and 6). These findings suggest that ANAPC4 is involved in the molecular mechanism by which TRMT13 inhibits PTC onset and progression. Rescue experiments confirmed that TRMT13 modulates ANAPC4 expression ( Figures 7 and 8). Hence, ANAPC4 partially mediates the inhibitory effect of TRMT13 on PTC. The present work provides preliminary evidence that TRMT13/ ANAPC4 suppresses PTC onset and progression. However, further research is needed to determine the clinical significance of these genes in this type of cancer by investigating the associations between gene alterations and patient outcomes or treatment responses and between gene expression profiles and patient outcomes. Moreover, future research will focus on the regulatory mechanism between TRMT13 and ANAPC4.
In summary, the present work established that TRMT13 and ANAPC4 suppress PTC cell proliferation, migration, and invasion in vitro and reduce PTC tumors in vivo. They also impeded tumor growth in a nude mouse PTC xenograft model. The findings presented herein lay the foundation for future research on the clinical application of TRMT13 and ANAPC4 in the diagnosis and treatment of PTC. This study also facilitates the clarification of the roles of tRNA methyltransferases and coiled-coil domain-containing proteins.
References
- H Sung, J Ferlay, RL Siegel, M Laversanne, I Soerjomataram, A Jemal, F Bray. Global cancer statistics 2020: globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin, 2021. [DOI | PubMed]
- M Li, R Zheng, L Dal Maso, S Zhang, W Wei, S Vaccarella. Mapping overdiagnosis of thyroid cancer in China. Lancet Diabetes Endocrinol, 2021. [DOI | PubMed]
- J Wang, F Yu, Y Shang, Z Ping, L Liu. Thyroid cancer: incidence and mortality trends in China, 2005–2015. Endocrine, 2020. [DOI | PubMed]
- M Schlumberger, S Leboulleux. Current practice in patients with differentiated thyroid cancer. Nat Rev Endocrinol, 2021. [DOI | PubMed]
- V Subbiah, MI Hu, LJ Wirth, M Schuler, AS Mansfield, G Curigliano, MS Brose. Pralsetinib for patients with advanced or metastatic RET-altered thyroid cancer (ARROW): a multi-cohort, open-label, registrational, phase 1/2 study. Lancet Diabetes Endocrinol, 2021. [DOI | PubMed]
- H Li, H Dong, B Xu, QB Xiong, CT Li, WQ Yang, J Li. A dual role of human tRNA methyltransferase hTrmt13 in regulating translation and transcription. EMBO J, 2022. [DOI | PubMed]
- NB Cronin, J Yang, Z Zhang, K Kulkarni, L Chang, H Yamano, D Barford. Atomic-resolution structures of the APC/C subunits Apc4 and the Apc5 N-terminal domain. J Mol Biol, 2015. [DOI | PubMed]
- RC Martinez-Chacin, T Bodrug, DL Bolhuis, KM Kedziora, T Bonacci, A Ordureau, ME Gibbs. Ubiquitin chain-elongating enzyme UBE2S activates the RING E3 ligase APC/C for substrate priming. Nat Struct Mol Biol, 2020. [DOI | PubMed]
- H Yu, JM Peters, RW King, AM Page, P Hieter, MW Kirschner. Identification of a cullin homology region in a subunit of the anaphase-promoting complex. Science, 1998. [DOI | PubMed]
- Furuta T, Tuck S, Kirchner J, Koch B, Auty R, Kitagawa R, et al. EMB-30: an APC4 homologue required for metaphase-to-anaphase transitions during meiosis and mitosis in Caenorhabditis elegans. Mol Bio Cell 2000, 11: 1401–1419
- MG Diniz, J de Fatima Correia Silva, FTA de Souza, NB Pereira, CC Gomes, RS Gomez. Association between cell cycle gene transcription and tumor size in oral squamous cell carcinoma. Tumor Biol, 2015. [DOI]
- Q Hu, W Chen, S Zhong, J Zhang, T Ma, H Ji, M Lv. MicroRNA-452 contributes to the docetaxel resistance of breast cancer cells. Tumor Biol, 2014. [DOI]
- H Ito, H Shiwaku, C Yoshida, H Homma, H Luo, X Chen, K Fujita. In utero gene therapy rescues microcephaly caused by Pqbp1-hypofunction in neural stem progenitor cells. Mol Psychiatry, 2015. [DOI | PubMed]
- E Ali, MJ Kalita, S Kalita, J Talukdar, AJ Deka, J Sultana, BN Choudhury. Upregulation of anaphase promoting complex (APC) 7 as a prognostic marker for esophageal squamous cell carcinoma: a hospital based study. Heliyon, 2022. [DOI | PubMed]
- DS Makeeva, AS Lando, A Anisimova, AA Egorov, MD Logacheva, AA Penin, DE Andreev. Translatome and transcriptome analysis of TMA20 (MCT-1) and TMA64 (eIF2D) knockout yeast strains. Data Brief, 2019. [DOI | PubMed]
- Z Tang, C Li, B Kang, G Gao, C Li, Z Zhang. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res, 2017. [DOI | PubMed]
- R Upadhyay, WZ Ying, Z Nasrin, H Safah, EA Jaimes, W Feng, PW Sanders. Free light chains injure proximal tubule cells through the STAT1/HMGB1/TLR axis. JCI Insight, 2020. [DOI | PubMed]
- M Huang, J Long, Z Yao, Y Zhao, Y Zhao, J Liao, K Lei. METTL1-mediated m7G tRNA modification promotes lenvatinib resistance in hepatocellular carcinoma. Cancer Res, 2023. [DOI | PubMed]
- H Han, S Zheng, S Lin. N7-methylguanosine (m7G) tRNA modification: a novel autophagy modulator in cancer. Autophagy, 2023. [DOI | PubMed]
- F Zhang, K Yoon, DY Zhang, NS Kim, G Ming, H Song. Epitranscriptomic regulation of cortical neurogenesis via Mettl8-dependent mitochondrial tRNA m3C modification. Cell Stem Cell, 2023. [DOI | PubMed]
- P Hou, T Tian, P Jia, Y Lin, Z Li, Y Wang, Y Ye. The regulation of NLRP3 inflammasome activation by CCDC50-mediated autophagy. Autophagy, 2023. [DOI | PubMed]
- W Zhi, Y Wei, C Lazare, Y Meng, P Wu, P Gao, S Lin. HPV- CCDC106 integration promotes cervical cancer progression by facilitating the high expression of CCDC106 after HPV E6 splicing. J Med Virol, 2023. [DOI | PubMed]
- AS Weinmann, PS Yan, MJ Oberley, THM Huang, PJ Farnham. Isolating human transcription factor targets by coupling chromatin immunoprecipitation and CpG island microarray analysis. Genes Dev, 2002. [DOI | PubMed]
- WL Towns, TJ Begley. Transfer RNA methytransferases and their corresponding modifications in budding yeast and humans: activities, predications, and potential roles in human health. DNA Cell Biol, 2012. [DOI | PubMed]
- A Bradfield, L Button, J Drury, DC Green, CJ Hill, DK Hapangama. Investigating the role of telomere and telomerase associated genes and proteins in endometrial cancer. Methods Protoc, 2020. [DOI | PubMed]
- L Sansregret, JO Patterson, S Dewhurst, C López-García, A Koch, N McGranahan, WCH Chao. APC/C dysfunction limits excessive cancer chromosomal instability. Cancer Discov, 2017. [DOI | PubMed]
- KL Thu, J Silvester, MJ Elliott, W Ba-alawi, MH Duncan, AC Elia, AS Mer. Disruption of the anaphase-promoting complex confers resistance to TTK inhibitors in triple-negative breast cancer. Proc Natl Acad Sci USA, 2018. [DOI | PubMed]
