Different outer membrane c‐type cytochromes are involved in direct interspecies electron transfer to Geobacter or Methanosarcina species
Abstract
Direct interspecies electron transfer (DIET) may be most important in methanogenic environments, but mechanistic studies of DIET to date have primarily focused on cocultures in which fumarate was the terminal electron acceptor. To better understand DIET with methanogens, the transcriptome of Geobacter metallireducens during DIET‐based growth with G. sulfurreducens reducing fumarate was compared with G. metallireducens grown in coculture with diverse Methanosarcina. The transcriptome of G. metallireducens cocultured with G. sulfurreducens was significantly different from those with Methanosarcina. Furthermore, the transcriptome of G. metallireducens grown with Methanosarcina barkeri, which lacks outer‐surface c‐type cytochromes, differed from those of G. metallireducens cocultured with M. acetivorans or M. subterranea, which have an outer‐surface c‐type cytochrome that serves as an electrical connect for DIET. Differences in G. metallireducens expression patterns for genes involved in extracellular electron transfer were particularly notable. Cocultures with c‐type cytochrome deletion mutant strains, ∆Gmet_0930, ∆Gmet_0557 and ∆Gmet_2896, never became established with G. sulfurreducens but adapted to grow with all three Methanosarcina. Two porin–cytochrome complexes, PccF and PccG, were important for DIET; however, PccG was more important for growth with Methanosarcina. Unlike cocultures with G. sulfurreducens and M. acetivorans, electrically conductive pili were not needed for growth with M. barkeri. Shewanella oneidensis, another electroactive microbe with abundant outer‐surface c‐type cytochromes, did not grow via DIET. The results demonstrate that the presence of outer‐surface c‐type cytochromes does not necessarily confer the capacity for DIET and emphasize the impact of the electron‐accepting partner on the physiology of the electron‐donating DIET partner.
Article type: Research Article
Keywords: direct interspecies electron transfer (DIET), extracellular electron transfer
Affiliations: Department of Microbiology University of Massachusetts‐Amherst Amherst Massachusetts USA; Department of Physical and Biological Science Western New England University Springfield Massachusetts USA; Institute for Advanced Study Shenzhen University Shenzhen China; Department of Biomolecular Sciences Central Connecticut State University New Britain Connecticut USA; College of Environment Zhejiang University of Technology Hangzhou China; College of Environmental Science and Engineering Beijing Forestry University Beijing China
License: © 2022 The Authors. mLife published by John Wiley & Sons Australia, Ltd. on behalf of Institute of Microbiology, Chinese Academy of Sciences. CC BY 4.0 This is an open access article under the terms of the http://creativecommons.org/licenses/by/4.0/ License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Article links: DOI: 10.1002/mlf2.12037 | PubMed: 38818222 | PMC: PMC10989804
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (879 KB)
INTRODUCTION
Direct interspecies electron transfer (DIET) is proposed to play a major role in the global cycling of methane, an important greenhouse gas, and in the generation of methane biofuel from organic wastes. Syntrophic communities producing or consuming methane in anaerobic soils and sediments share electrons via DIETref. mlf212037-bib-0001, ref. mlf212037-bib-0002. Promoting DIET in anaerobic digesters through changes in digester design or with the addition of electrically conductive materials can accelerate anaerobic digestionref. mlf212037-bib-0003, ref. mlf212037-bib-0004. Thus, understanding the mechanisms for electron exchange during DIET could contribute to better modeling of carbon and electron flux in natural anaerobic environments and could suggest better strategies for enhancing DIET during organic waste treatment.
Until now, studies on the routes for interspecies electron transfer during DIET have primarily focused on the mechanisms of electron uptake by the electron‐accepting partnerref. mlf212037-bib-0005, ref. mlf212037-bib-0006, ref. mlf212037-bib-0007, ref. mlf212037-bib-0008, ref. mlf212037-bib-0009. These studies have demonstrated that the electrical connects on the outer cell surface of electron‐accepting partners can be dramatically different. For example, Geobacter sulfurreducens displays a network of electrically conductive pili (e‐pili) and outer‐surface multiheme c‐type cytochromes to facilitate electron uptake during DIETref. mlf212037-bib-0005, ref. mlf212037-bib-0006, ref. mlf212037-bib-0010. In contrast, some methanogens can express electrically conductive archaellaref. mlf212037-bib-0011, but archaella are not known to participate in DIET and do not form a multifiber conductive extracellular mesh, like Geobacter e‐pili. Furthermore, some methanogens, such as Methanothrix harundinacea, Methanobacterium electrotrophus strain YSL and Methanosarcina barkeri, are capable of DIET despite a lack of outer‐surface c‐type cytochromesref. mlf212037-bib-0007, ref. mlf212037-bib-0008, ref. mlf212037-bib-0012, ref. mlf212037-bib-0013, ref. mlf212037-bib-0014, ref. mlf212037-bib-0015, ref. mlf212037-bib-0016. The electrical contacts for DIET in these methanogens are unknown. However, another methanogen, M. acetivorans, expresses an outer‐surface multiheme c‐type cytochrome, MmcA, which is a key electrical contact for DIETref. mlf212037-bib-0009.
This major difference in mechanisms for DIET‐based electron uptake between M. barkeri and M. acetivorans is just one of the physiological dissimilarities between two clades of Methanosarcina species. Type I Methanosarcina species, like M. barkeri, predominate in waste digesters and other high‐energy environments and have different mechanisms for energy conservation than Type II Methanosarcina, such as M. acetivorans, which are better adapted for growth in organic‐poor soils and sedimentsref. mlf212037-bib-0015. It might be expected that differences in the diversity of electrical contacts on electron‐accepting partners for DIET could influence the routes for electron transfer from electron‐donating partners.
Furthermore, the majority of studies on DIET mechanisms have focused on cocultures in which fumarate‐reducing G. sulfurreducens was the electron‐accepting partner. DIET to a fumarate‐reducing partner is unlikely to be important in anaerobic environments because fumarate is not a commonly abundant electron acceptor. In addition, most simple organic substrates can be directly metabolized within individual fumarate‐respiring microbes, which means that syntrophy is not required. In contrast, the inability of most methanogens to use multicarbon electron donors other than acetate necessitates syntrophic metabolism with interspecies electron transfer. The electron carriers that provide electrons to support fumarate reduction during DIET could be much different than those required to promote the reduction of carbon dioxide to methane because the mid‐point potential of the carbon dioxide/methane redox couple is much more negative than the fumarate/succinate couple.
A diversity of electron‐donating partners for DIET have been identified, but in most instances, genetically tractable isolates for mechanistic studies are not available. For example, pure cultures are not available for the archaea proposed to function as electron‐donating partners for DIET in consortia catalyzing anaerobic methane oxidation coupled to sulfate reductionref. mlf212037-bib-0017, ref. mlf212037-bib-0018, ref. mlf212037-bib-0019. Tools for genetic manipulation have not been developed for bacteria, such as Syntrophus aciditrophicus ref. mlf212037-bib-0020, Rhodoferax ferrireducens ref. mlf212037-bib-0008 and Desulfovibrio strain JYref. mlf212037-bib-0021, shown to serve as the electron‐donating partner for DIET in defined cocultures.
Therefore, G. metallireducens, which is genetically tractableref. mlf212037-bib-0022, has served as the primary pure culture model for an electron‐donating partner in DIET. G. metallireducens is an attractive model for DIET studies because of its environmental relevance. Geobacter species are important electron‐donating DIET partners in some anaerobic waste digestersref. mlf212037-bib-0004, ref. mlf212037-bib-0023 and in rice paddy soils, an important source of atmospheric methaneref. mlf212037-bib-0024. G. metallireducens is a good candidate for studying DIET in defined systems because it can oxidize ethanol and other short‐chain alcohols with electron transfer to extracellular electron acceptors without generating H2 or formateref. mlf212037-bib-0005, ref. mlf212037-bib-0006, ref. mlf212037-bib-0010, ref. mlf212037-bib-0025, ref. mlf212037-bib-0026.
Here, we report that DIET transcriptomes suggest that the physiology of the electron‐accepting partner directly impacts the physiology of G. metallireducens serving as the electron‐donating partner. Gene deletion studies suggested different routes for electron transport, depending on which partner G. metallireducens was paired with. We also demonstrate that in many instances, G. metallireducens/Methanosarcina cocultures adapted over time to gene deletions that initially disrupted DIET. Attempts to simplify the study of DIET extracellular electron exchange by initiating DIET cocultures with Shewanella oneidensis, an electroactive microbe with a less complicated array of outer‐surface electron transport options, were unsuccessful.
RESULTS AND DISCUSSION
Gene expression patterns suggest that the electron‐accepting partner influences routes for DIET
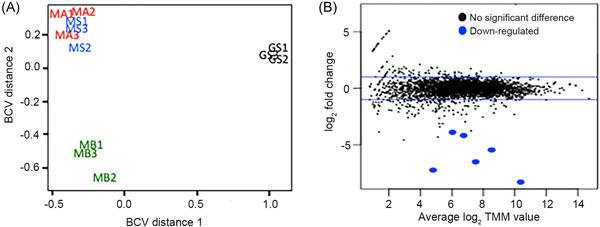
DIET between G. metallireducens and G. sulfurreducens was compared to DIET between G. metallireducens and three Methanosarcina species: M. barkeri (MB), a Type I Methanosarcina that lacks outer‐surface c‐type cytochromesref. mlf212037-bib-0015 and M. acetivorans (MA) and M. subterranea (MS), Type II Methanosarcina with the outer‐surface multiheme c‐type cytochrome MmcA that serves an essential electrical contact for DIETref. mlf212037-bib-0009. Multidimensional scaling analysis generated with the biological coefficient of variation (BCV) method and mean difference plots (MD plots) revealed that the transcriptomes of G. metallireducens grown in DIET‐based coculture with Methanosarcina species were significantly different from those of G. metallireducens grown in coculture with G. sulfurreducens (Figures 1 and S1, Tables S1–, S3). Furthermore, the transcriptomes of G. metallireducens grown with the Type I Methanosarcina, M. barkeri, were significantly different from those of G. metallireducens grown with the Type II Methanosarcina species, M. acetivorans and M. subterranea. Both MDS (Figure 1A) and MD plots (Figure 1B) revealed that the transcriptomes of G. metallireducens grown with either of the Type II Methanosarcina were similar. These results indicate that the properties of the electron‐accepting DIET partner substantially influence the physiological status of G. metallireducens during growth via DIET. Differences in gene expression suggested substantial differences in many aspects of G. metallireducens physiology (Figure S2). Here, we focus on the expression of possible outer‐surface electrical connects for DIET.
Some of the most significant differences in G. metallireducens gene expression patterns during growth with the different types of electron‐accepting partners were for genes coding for multiheme c‐type cytochromes (Figure 2, Tables 1, 2, 3, S1 and S2). The pattern of cytochrome gene expression was most similar in G. metallireducens grown with the two Type II Methanosarcina and the expression patterns between G. metallireducens and the Type I and Type II Methanosarcina were more similar to each other than to G. metallireducens grown with G. sulfurreducens as the electron‐accepting partner. Therefore, similarities and differences in G. metallireducens cytochrome gene expression and the impact of targeted gene deletions on the ability of G. metallireducens to establish DIET with different electron‐accepting partners were further evaluated.
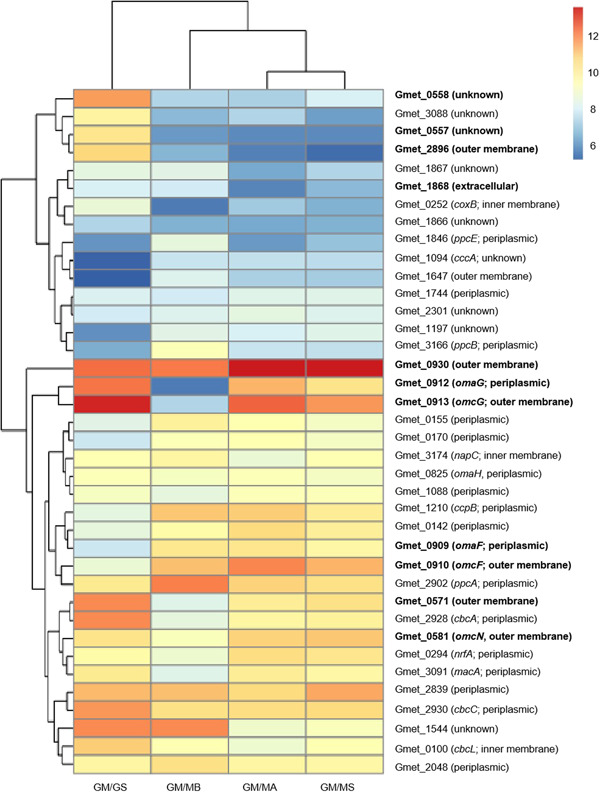
Table 1: Differences in expression of genes coding for electron transfer proteins in Geobacter metallireducens that were important for DIET when G. sulfurreducens was the electron‐accepting partner.
| Locus ID | Annotation | Gene | Location | GS vs. MB | GS vs. MA | GS vs. MS |
|---|---|---|---|---|---|---|
| Gmet_0558 | c‐type cytochrome protein, 27 hemes | Unknown | 18.28 | 19.68 | 13.50 | |
| Gmet_0557 | c‐type cytochrome protein, 4 hemes | Unknown | 18.02 | 22.42 | 24.19 | |
| Gmet_2896 | c‐type cytochrome protein, 4 hemes | Outer membrane/surface | 17.01 | 33.12 | 46.11 | |
| Gmet_0930 | c‐type cytochrome, 8 hemes | Outer membrane/surface | NS | −2.80 | −2.57 | |
| Gmet_1868 | c‐type cytochrome, 4 hemes | Outer membrane/surface | NS | 4.12 | 2.31 | |
| Gmet_0571 | c‐type cytochrome protein, 34 hemes | Outer membrane/surface | 12.63 | 3.29 | 2.72 | |
| Gmet_0581 | c‐type cytochrome protein, 34 hemes | omcN | Outer membrane/surface | 2.31 | NS | NS |
| Gmet_0911 | Porin protein from PccG complex | ombG | Outer membrane/surface | 80.67 | NS | 4.0 |
| Gmet_0912 | 8 heme periplasmic c‐type cytochrome from PccG complex | omaG | Periplasmic | 96.91 | 2.01 | 3.58 |
| Gmet_0913 | 9 heme outer membrane c‐type cytochrome from PccG | omcG | Outer membrane/surface | 70.53 | 2.34 | 3.43 |
| Gmet_0908 | Porin protein from PccF complex | ombF | Outer membrane/surface | −3.57 | −4.76 | −2.41 |
| Gmet_0909 | 9 heme periplasmic c‐type cytochrome from PccF complex | omaF | Periplasmic | −6.50 | −7.08 | −3.81 |
| Gmet_0910 | 10 heme c‐type cytochrome from PccF complex | omcF | Outer membrane/surface | −6.41 | −11.42 | −6.30 |
| Gmet_1399 | Type IV major pilin subunit, PilA | pilA | Outer membrane/surface | −4.31 | 1.64 | 1.76 |
| Gmet_1400 | Short pilin chaperone protein, Spc | spc | Outer membrane/surface | −7.85 | NS | NS |
| Gmet_2163 | Multicopper oxidase protein | ompB | Outer membrane/surface | 3.89 | 7.66 | 10.72 |
| Gmet_2928 | 7 heme c‐type cytochrome protein from CbcABCDE | cbcA | Periplasmic | 11.09 | 4.34 | 4.13 |
| Gmet_2929 | b‐type cytochrome from CbcABCDE complex | cbcB | Inner membrane | 7.85 | 6.08 | 5.22 |
| Gmet_2930 | 11 heme c‐type cytochrome protein, CbcABCD complex | cbcC | Periplasmic | 2.16 | 2.0 | 2.14 |
| Gmet_2931 | Monoheme c‐type cytochrome protein from CbcABCDE complex | cbcD | Periplasmic | 3.18 | 2.21 | 2.50 |
| Gmet_2932 | Membrane protein from CbcABCDE complex | cbcE | Inner membrane | 4.46 | 1.74 | 1.80 |
| Gmet_0252 | Monoheme c‐type cytochrome protein | coxB | Inner membrane | 6.65 | 2.46 | 4.09 |
| Gmet_0325 | 4 heme c‐type cytochrome protein from CbcSTU‐2 complex | cbcS‐2 | Periplasmic | 5.37 | 11.80 | 3.18 |
| Gmet_0327 | b‐type cytochrome from CbcSTU‐1 complex | cbcU‐2 | Inner membrane | 2.25 | 9.48 | 4.64 |
| Gmet_3519 | 4Fe‐4S ferredoxin from CbcSTU‐1 complex | cbcT‐1 | Periplasmic | NS | 3.47 | 3.44 |
| Gmet_3520 | b‐type cytochrome from CbcSTU‐2 complex | cbcU‐1 | Inner membrane | 3.86 | 9.98 | 4.14 |
| Gmet_0100 | Quinone oxidoreductase cytochrome, 9 hemes | cbcL | Inner membrane | 2.96 | 4.51 | 3.37 |
| Gmet_3091 | c‐type cytochrome protein, 2 hemes | macA | Periplasmic | 3.82 | NS | NS |
| Gmet_3165 | c‐type cytochrome protein, 3 hemes | ppcC | Periplasmic | 3.31 | 2.29 | NS |
| Gmet_0580 | c‐type cytochrome protein, 21 hemes | Outer membrane/surface | 3.02 | NS | NS | |
| Gmet_0600 | c‐type cytochrome protein, 19 hemes | Outer membrane/surface | 2.21 | NS | NS | |
| Gmet_0575 | c‐type cytochrome protein, 27 hemes | Outer membrane/surface | 2.01 | 3.03 | 2.86 | |
| Gmet_3088 | c‐type cytochrome protein, 8 hemes | Unknown | 8.84 | 5.59 | 12.98 | |
| Gmet_2898 | c‐type cytochrome protein, 8 hemes | Unknown | 6.81 | NS | NS | |
| Gmet_2470 | c‐type cytochrome protein, 34 hemes | Unknown | 5.67 | 2.05 | 1.70 | |
| Gmet_2899 | c‐type cytochrome protein, 9 hemes | Unknown | 5.43 | NS | NS | |
| Gmet_1087 | c‐type cytochrome protein, 1 heme | Unknown | 4.46 | 8.40 | 6.56 | |
| Gmet_0601 | c‐type cytochrome protein, 8 hemes | Unknown | 2.34 | NS | NS | |
| Gmet_0534 | c‐type cytochrome protein, 5 hemes | cbcR | Periplasmic | NS | 3.36 | 3.10 |
| Gmet_0121 | c‐type cytochrome protein, 2 hemes | Unknown | NS | 2.12 | 2.09 |
Values represent fold difference between cocultures grown with G. sulfurreducens (GS) compared to cocultures grown with the three different Methanosarcina species (M. barkeri [MB]; M. acetivorans [MA]; M. subterranea [MS]). Negative values show that the gene was more highly expressed by G. metallireducens grown in coculture with the methanogen. p values are available in Table S2 but all comparisons shown have p values that are <0.05. NS, no significant difference.
Table 2: Genes coding for electron transport proteins in Geobacter metallireducens that had greater than two‐fold differences in expression in cocultures grown with Methanosarcina barkeri compared to cocultures grown with G. sulfurreducens (GS) and the Type II Methanosarcina (M. acetivorans [MA] and M. subterranea [MS]).
| Locus ID | Annotation | Gene | Location | MB vs. GS | MB vs. MA | MB vs. MS |
|---|---|---|---|---|---|---|
| Gmet_2119 | Cytochrome b from PcmABCDEFG complex | pcmC | Inner membrane | 104.99 | 9.93 | NS |
| Gmet_2117 | Membrane protein from PcmABCDEFG complex | pcmA | Inner membrane | 10.83 | 9.17 | 7.19 |
| Gmet_1019 | c‐type cytochrome, 2 hemes | narC | Inner membrane | 8.44 | NS | NS |
| Gmet_0330 | Monoheme c‐type cytochrome | narH | Inner membrane | 5.37 | NS | NS |
| Gmet_1809 | c‐type cytochrome, 5 hemes | actA | Inner membrane | 5.15 | NS | NS |
| Gmet_1923 | b‐type cytochrome from CbcWXV complex | cbcW | Inner membrane | 3.84 | 1.54 | NS |
| Gmet_1922 | Rieske 2Fe‐2S protein from CbcWXV complex | cbcV | Inner membrane | 1.92 | 1.64 | 2.13 |
| Gmet_2120 | Cytochrome b from PcmABCDEFG complex | pcmD | Inner membrane | NS | 4.66 | NS |
| Gmet_2122 | 4 heme c‐type cytochrome from PcmABCDEFG complex | pcmF | Periplasmic | 123.26 | 6.85 | 83.58 |
| Gmet_2123 | 4 heme c‐type cytochrome from PcmABCDEFG complex | pcmG | Periplasmic | 8.94 | 20.04 | 47.60 |
| Gmet_3166 | c‐type cytochrome, 2 hemes | ppcB | Periplasmic | 7.65 | 3.13 | 3.37 |
| Gmet_1210 | c‐type cytochrome, 2 hemes | ccpB | Periplasmic | 7.59 | NS | 2.11 |
| Gmet_2156 | c‐type cytochrome, 9 hemes | Periplasmic | 7.19 | 2.87 | NS | |
| Gmet_1703 | c‐type cytochrome, 7 hemes | Periplasmic | 6.71 | 7.63 | 3.59 | |
| Gmet_0909 | 9 heme c‐type cytochrome from PccF complex | omaF | Periplasmic | 6.50 | NS | NS |
| Gmet_1846 | c‐type cytochrome, 2 hemes | ppcE | Periplasmic | 6.12 | 5.52 | 3.33 |
| Gmet_0328 | c‐type cytochrome, 2 hemes | narC | Periplasmic | 5.80 | 2.62 | 2.50 |
| Gmet_0335 | c‐type cytochrome, 3 hemes | ppcF | Periplasmic | 5.52 | 2.95 | 2.64 |
| Gmet_2902 | c‐type cytochrome, 2 hemes | ppcA | Periplasmic | 4.31 | 2.42 | 3.16 |
| Gmet_1924 | 5 heme c‐type cytochrome from CbcWXV complex | cbcX | Periplasmic | 4.24 | 2.98 | 2.85 |
| Gmet_0155 | Monoheme c‐type cytochrome | Periplasmic | 3.50 | NS | NS | |
| Gmet_0170 | c‐type cytochrome, 10 hemes | Periplasmic | 3.12 | NS | NS | |
| Gmet_0142 | c‐type cytochrome, 8 hemes | Periplasmic | 2.42 | NS | NS | |
| Gmet_0910 | 10 heme c‐type cytochrome from PccF complex | omcF | Outer membrane/surface | 6.41 | NS | NS |
| Gmet_0908 | Porin protein from PccF complex | ombF | Outer membrane/surface | 3.57 | NS | NS |
| Gmet_1868 | c‐type cytochrome, 4 hemes | Outer membrane/surface | NS | 4.41 | 2.35 | |
| Gmet_2121 | Rieske 2Fe‐2S protein from PcmABCDEFG complex | pcmE | Unknown | 53.94 | 11.03 | NS |
| Gmet_2118 | 4Fe‐4S ferredoxin protein from PcmABCDEFG complex | pcmB | Unknown | 18.16 | 3.76 | NS |
| Gmet_1647 | c‐type cytochrome, 2 hemes | Unknown | 9.35 | NS | NS | |
| Gmet_1094 | c‐type cytochrome, 2 hemes | cccA | Unknown | 6.02 | NS | NS |
| Gmet_1197 | c‐type cytochrome, 5 hemes | Unknown | 5.64 | NS | NS | |
| Gmet_1191 | Monoheme c‐type cytochrome | Unknown | 4.49 | NS | NS | |
| Gmet_1814 | Monoheme c‐type cytochrome | actE | Unknown | 4.16 | NS | NS |
| Gmet_3506 | c‐type cytochrome, 2 hemes | Unknown | 3.49 | NS | NS | |
| Gmet_2432 | Monoheme P460 c‐type cytochrome | Unknown | 2.71 | NS | NS | |
| Gmet_2896 | c‐type cytochrome, 4 hemes | Unknown | NS | 2.01 | 2.63 |
p values are available in Table S2 but all comparisons shown have p values that are <0.05. NS, no significant difference.
Table 3: Genes coding for electron transport proteins in Geobacter metallireducens that had greater than two‐fold differences in expression in cocultures grown with the Type II Methanosarcina (M. acetivorans [MA]) than with G. sulfurreducens (GS), M. barkeri (MB) or M. subterranea (MS).
| Locus ID | Annotation | Gene | Location | MA vs. GS | MA vs. MB | MA vs. MS |
|---|---|---|---|---|---|---|
| Gmet_0533 | Transmembrane protein from CbcMNOPQR complex | cbcQ | Inner membrane | 6.59 | 5.36 | NS |
| Gmet_1019 | c‐type cytochrome, 2 hemes | narC | Inner membrane | NS | 3.71 | NS |
| Gmet_0252 | Monoheme c‐type cytochrome | coxB | Inner membrane | NS | 2.68 | NS |
| Gmet_0539 | b‐type cytochrome from CbcMNOPQR complex | cbcP | Inner membrane | NS | 2.28 | NS |
| Gmet_0330 | Monoheme c‐type cytochrome | narH | Inner membrane | 4.02 | NS | NS |
| Gmet_0538 | Transmembrane protein from CbcMNOPQR complex | cbcO | Inner membrane | NS | NS | NS |
| Gmet_0912 | 8 heme periplasmic c‐type cytochrome from PccG complex | omaG | Periplasmic | NS | 54.54 | NS |
| Gmet_3091 | c‐type cytochrome, 2 hemes | ccpA | Periplasmic | NS | 3.92 | NS |
| Gmet_0294 | c‐type cytochrome, 4 hemes | nrfA | Periplasmic | NS | 3.83 | NS |
| Gmet_0232 | Monoheme c‐type cytochrome | Periplasmic | NS | 2.64 | NS | |
| Gmet_2928 | 7 heme c‐type cytochrome from CbcABCD complex | cbcA | Periplasmic | NS | 2.52 | NS |
| Gmet_0537 | 10 heme c‐type cytochrome from CbcMNOPQR complex | cbcN | Periplasmic | NS | 2.21 | NS |
| Gmet_0142 | c‐type cytochrome, 8 hemes | Periplasmic | 5.15 | 2.09 | NS | |
| Gmet_0155 | Monoheme c‐type cytochrome | Periplasmic | 2.31 | NS | NS | |
| Gmet_2122 | 4 heme c‐type cytochrome from PcmABCDEFG complex | pcmF | Periplasmic | NS | NS | 15.46 |
| Gmet_0909 | 9 heme periplasmic c‐type cytochrome from PccF complex | omaF | Periplasmic | 7.08 | NS | NS |
| Gmet_0534 | 5 heme c‐type cytochrome from CbcMNOPQR complex | cbcR | Periplasmic | NS | NS | NS |
| Gmet_0911 | Porin protein from PccG complex | ombG | Outer membrane/surface | NS | 41.47 | NS |
| Gmet_0913 | 9 heme outer membrane c‐type cytochrome from PccG complex | omcG | Outer membrane/surface | NS | 37.16 | NS |
| Gmet_0580 | c‐type cytochrome, 21 hemes | Outer membrane/surface | NS | 4.92 | NS | |
| Gmet_0908 | Porin protein from PccF complex | ombF | Outer membrane/surface | 4.76 | NS | NS |
| Gmet_0571 | c‐type cytochrome, 34 hemes | Outer membrane/surface | NS | 3.79 | NS | |
| Gmet_0930 | c‐type cytochrome, 8 hemes | omcZ | Outer membrane/surface | 2.80 | 2.75 | NS |
| Gmet_0600 | c‐type cytochrome, 19 hemes | Outer membrane/surface | NS | 2.20 | NS | |
| Gmet_0910 | 10 heme c‐type cytochrome from PccF complex | omcF | Outer membrane/surface | 11.42 | NS | NS |
| Gmet_1647 | c‐type cytochrome, 2 hemes | Outer membrane/surface | 5.03 | NS | NS | |
| Gmet_2626 | Monoheme P460 c‐type cytochrome | Unknown | NS | 5.50 | NS | |
| Gmet_2899 | c‐type cytochrome, 9 hemes | Unknown | NS | 5.15 | NS | |
| Gmet_2898 | c‐type cytochrome, 8 hemes | Unknown | NS | 4.77 | NS | |
| Gmet_0581 | c‐type cytochrome, 34 hemes | Unknown | NS | 3.40 | NS | |
| Gmet_0679 | c‐type cytochrome, 5 hemes | Unknown | 2.60 | 3.20 | NS | |
| Gmet_2470 | c‐type cytochrome, 34 hemes | Unknown | NS | 2.75 | NS | |
| Gmet_0733 | Monoheme c‐type cytochrome | Unknown | NS | 2.41 | NS | |
| Gmet_0601 | c‐type cytochrome, 8 hemes | Unknown | NS | 2.34 | NS | |
| Gmet_1191 | Monoheme c‐type cytochrome | Unknown | 6.24 | NS | NS |
p values are available in Table S2 but all comparisons shown have p values that are <0.05. NS, no significant difference.
Different outer‐surface c‐type cytochromes are required for DIET with G. sulfurreducens than DIET with Methanosarcina
Multiheme, c‐type cytochromes positioned on the outer cell surface can serve as a key electrical contact for interspecies electron transferref. mlf212037-bib-0027. Gene expression patterns and the differential impact of deleting genes for outer‐surface c‐type cytochromes suggested that the relative importance of individual G. metallireducens outer‐surface cytochromes depended upon which microbe served as the electron‐accepting partner for DIET.
For example, a major difference between G. sulfurreducens or Methanosarcina species serving as the electron‐accepting partner was the apparent role of the multiheme c‐type cytochromes Gmet_0558, Gmet_0557 and Gmet_2896. Software (PSORTb version 3.0) designed to predict cellular localizationref. mlf212037-bib-0028 did not specify a location for these cytochromes. However, the presence of a transmembrane helix in Gmet_0557 suggests that it is membrane‐associated and the Gmet_2896 cytochrome was recovered in outer membrane preparations of G. metallireducens ref. mlf212037-bib-0029, ref. mlf212037-bib-0030. The large number of hemesref. mlf212037-bib-0027 predicted for Gmet_0558 and the fact that it has high homology (67% amino acid identity) to OmcO, an outer membrane c‐type cytochrome in G. sulfurreducens ref. mlf212037-bib-0031, also suggests that it may be an outer‐surface electrical contact.
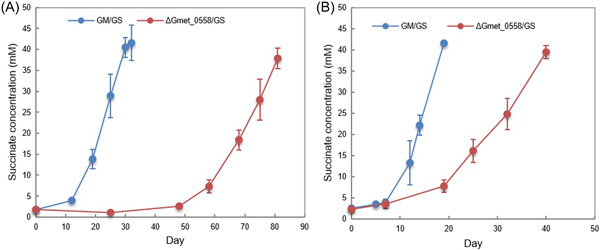
The genes for all three of these cytochromes were more highly expressed when G. sulfurreducens was the electron‐accepting partner than with any of the Methanosarcina species (Table 1). Deleting any of these three genes had no impact on establishing cocultures with the three Methanosarcina species (Figure S3). However, cocultures initiated with G. sulfurreducens and Gmet_0557‐ or Gmet_2896‐deficient strains did not grow and cocultures initiated with a Gmet_0558‐deficient mutant had a more extensive lag period than cocultures initiated with wild‐type G. metallireducens (Figure 3A). Even after further adaption with three more transfers, the cocultures with the Gmet_0558‐deficient strain reduced fumarate at rates that were 2.3 times slower (p = 1.15 × 10−5) than cocultures with wild‐type G. metallireducens (Figure 3B). These results suggest that Gmet_0558, Gmet_0557 and Gmet_2896 are all important for extracellular electron transfer when G. sulfurreducens is the electron‐accepting partner, but they are not essential for DIET with Methanosarcina with or without outer‐surface c‐type cytochrome electrical connects. Previous gene‐deletion studies indicated that Gmet_0558, Gmet_0557 and Gmet_2896 are required for growth with insoluble Fe(III) oxide as the electron acceptorref. mlf212037-bib-0029. Thus, the concept that Geobacter extracellular electron transfer to other microbial species requires the same outer‐surface electron transport components as electron transport to Fe(III) oxide does not appear to hold when Methanosarcina species are the partner for DIET.
Gmet_0930 represents another instance in which an outer‐surface c‐type cytochrome was essential for DIET to G. sulfurreducens and Fe(III) oxide reduction, but not for DIET with Methanosarcina species. Gmet_0930 encodes an eight‐heme outer‐surface c‐type cytochrome that is homologous to OmcZ (GSU2076) in G. sulfurreducens; they have 45% amino acid identity, they both have eight hemes and are predicted to be extracellular and have putative immunoglobulin‐like fold domains. Deletion of the gene for OmcZ had no impact on Fe(III) oxide reduction in G. sulfurreducens ref. mlf212037-bib-0032, but G. metallireducens requires Gmet_0930 to reduce Fe(III) oxideref. mlf212037-bib-0029.
Gmet_0930 was one of the most highly expressed G. metallireducens c‐type cytochrome genes with all electron‐accepting partners (Figure 2 and Table S1). Cocultures could not be established with G. sulfurreducens and a strain of G. metallireducens in which Gmet_0930 was deleted. Initiating cocultures with the Gmet_0930‐deficient strain delayed methane production by all three of the Methanosarcina strains (Figure 4). Although Gmet_0930 was not as essential for DIET with Methanosarcina species as it was for DIET with G. sulfurreducens, it did appear to optimize DIET because even after four transfers, the Methanosarcina cocultures initiated with the Gmet_0930‐deficient strain grew 1.5–2‐fold slower than cocultures initiated with wild‐type G. metallireducens (p value range, 0.001–0.009; Figure S4). The finding that deletion of Gmet_0930 inhibits DIET with G. sulfurreducens, as well as Fe(III) oxide reductionref. mlf212037-bib-0029, further supports the conclusion from the study of Gmet_0558, Gmet_0557 and Gmet_2896 that there are differences between G. metallireducens electron transport to Fe(III) oxide and G. sulfurreducens compared to electron transport to Methanosarcina species.
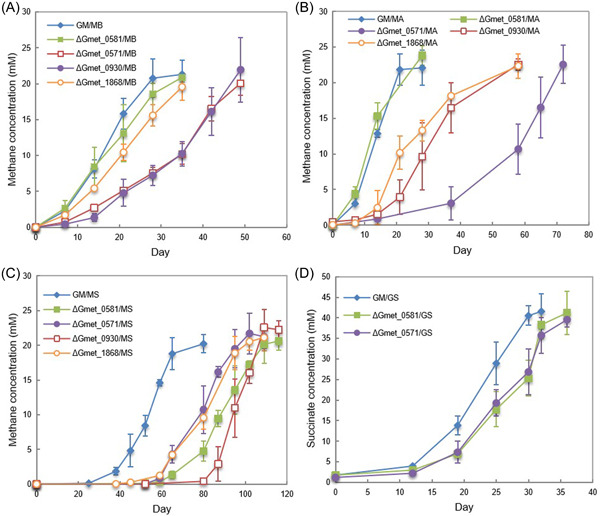
In a similar manner, the four‐heme outer‐surface c‐type cytochrome Gmet_1868, which G. metallireducens requires for Fe(III) oxide reductionref. mlf212037-bib-0029, appeared to be essential for DIET with G. sulfurreducens, but only helpful for DIET with the Methanosarcina species. Transcript abundance for Gmet_1868 was 4.12 (p = 2.39 × 10−5) and 2.31 (p = 0.001) times greater in GM/GS cocultures than it was in coculture with either of the Type II Methanosarcina (GM/MA and GM/MS) (Table S2). Cocultures initiated with a Gmet_1868‐deficient strain and G. sulfurreducens did not grow. The lag phase for the Gmet_1868‐deficient strain and Methanosarcina cocultures was longer than the wild‐type cocultures (Figure 4) and even after four transfers, the cocultures initiated with the Gmet_1868‐deficient strain and the Methanosarcina species grew at rates that were 1.4‐ to 3.3‐fold slower (p value range, 0.001–0.03) than the wild‐type coculture (Figure S4).
The 34‐heme outer‐surface c‐type cytochrome Gmet_0571, which was highly expressed during growth with all electron‐accepting partners (Figure 2) appeared to be more important for DIET with the Methanosarcina species than G. sulfurreducens. Cocultures initiated with G. sulfurreducens and a strain of G. metallireducens in which the Gmet_0571 gene was deleted grew similarly to wild‐type cocultures (Figures 4 and S4). In contrast, cocultures with the Gmet_0571‐deficient strain and Methanosarcina were substantially delayed in methane production compared to cocultures initiated with wild‐type G. metallireducens (Figure 4). With the subsequent transfer, the ∆Gmet_0571/MB, ∆Gmet_0571/MA and ∆Gmet_0571/MS cocultures adapted somewhat to the loss of Gmet_0571, but they continued to grow 1.6–3.0 times slower (p value range, 7.6 × 10−6 − 0.04) than the cocultures established with wild‐type G. metallireducens (Figure S4). These results suggest that Gmet_0571 is required for optimal DIET with Methanosarcina species, but is not necessary for DIET with G. sulfurreducens or for growth with Fe(III) oxide provided as the electron acceptorref. mlf212037-bib-0029.
A different pattern of gene deletion phenotypes was observed with Gmet_0581, another putative 34‐heme c‐type cytochrome likely to be positioned on the outer cell surface. Gmet_0581 shares 63% amino acid identity with the outer membrane c‐type cytochrome OmcN of G. sulfurreducens and has a surface‐associated immunoglobulin‐like domain. It was highly expressed by all electron‐accepting partners. Deletion of Gmet_0581 impacted coculture establishment with Type II M. subterranea but not the other partners. However, after four transfers, cocultures with M. subterranea grew at rates that were comparable to cocultures established with wild‐type G. metallireducens (Figure S4).
Porin–cytochrome complexes (Pcc) are required for DIET
Transcriptomic and gene deletion studies of the Pcc complexes also suggested that there are different requirements for DIET to Methanosarcina species than DIET to G. sulfurreducens. G. metallireducens has genes for three Pcc complexes: PccF (Gmet_0908‐0910), PccG (Gmet_0911‐0913) and PccH (Gmet_0825‐0827). Pcc complexes are thought to be necessary for electron transport across the outer membrane of Geobacter speciesref. mlf212037-bib-0027, ref. mlf212037-bib-0033. The full suite of genes coding for components of the PccH complex was not highly expressed by G. metallireducens in any of the coculture conditions, indicating that PccH was not important for DIET. However, some of the most substantial differences in G. metallireducens gene transcript abundance in cocultures with G. sulfurreducens versus M. barkeri as the electron‐accepting partner were for the genes that code for components of the PccG porin–cytochrome complex (Table 1). The genes from the PccF complex, on the contrary, were more highly expressed by G. metallireducens grown in the three Methanosarcina cocultures (Table 1).
Initiating cocultures with strains of G. metallireducens in which the gene for OmcF or OmcG, the outer‐membrane cytochromes of the PccF and PccG conduits, was deleted delayed the initiation of methane production in cocultures established with each of the Methanosarcina species (Figure 5A–C). There was also a slight lag in cocultures established with G. sulfurreducens and the OmcF‐ and OmcG‐deficient strains of G. metallireducens (Figure 5D). Deletion of the gene coding for the periplasmic‐facing c‐type cytochrome, omaG (Gmet_0912) also delayed methane production with all three Methanosarcina species (Figure 5A–C) but did not have an impact on coculture growth with G. sulfurreducens (Figure 5D).
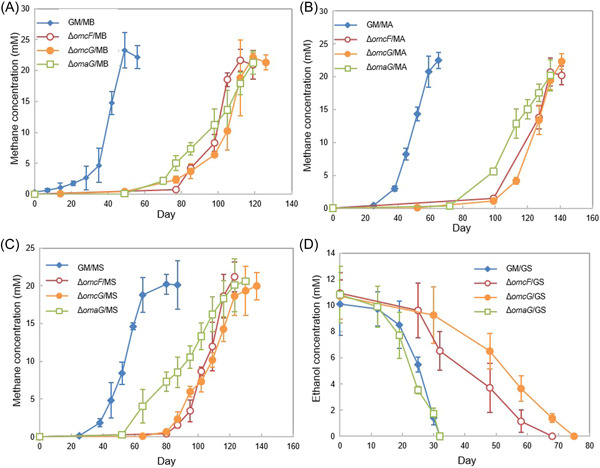
After the cocultures were passed through three additional sequential transfers, the cocultures of G. metallireducens strain ∆omcF and the Methanosarcina species, strain ∆omcF/G. sulfurreducens and strain ∆omcG/G. sulfurreducens, adapted to grow at rates that were similar to the wild‐type (Figure S5). However, even after this long‐term adaption through multiple transfers, the cocultures that had been initiated with G. metallireducens strain ∆omcG or strain ∆omaG and the Methanosarcina species grew ca. 2‐4‐fold slower (p value range, 2.6 × 10−5 – 0.003) than cocultures initiated with wild‐type G. metallireducens (Figure S5). These results suggest that the PccG porin–cytochrome conduit has features that are optimized for DIET with Methanosarcina species, but that either porin–cytochrome conduit is suitable for DIET with G. sulfurreducens. Strains lacking the gene coding for either of the porin proteins, OmbF or OmbG (Gmet_0908 and Gmet_0911) were not tested for growth in coculture because this study focused on redox proteins required for electron transport out of the cell.
e‐Pili are important for DIET but are not always required
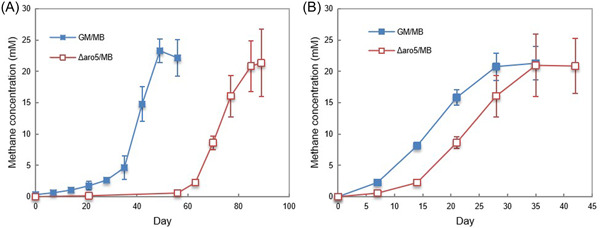
The G. metallireducens gene for PilA, the pilin monomer that is assembled into e‐piliref. mlf212037-bib-0034 and the adjacent gene for Spc, a putative chaperone protein that facilitates pilus assemblyref. mlf212037-bib-0035, as well as many of the pilin accessory proteinsref. mlf212037-bib-0036, were highly expressed in G. metallireducens grown with each of the electron‐accepting partners evaluated (Tables 1 and S4). Constructing Geobacter strains that express poorly conductive pili is a method for evaluating the importance of pili conductivity in extracellular electron exchange without disrupting the proper expression of outer‐surface cytochromes that also play a key role in DIETref. mlf212037-bib-0037, ref. mlf212037-bib-0038, ref. mlf212037-bib-0039, ref. mlf212037-bib-0040, ref. mlf212037-bib-0041. A strain of G. metallireducens that expresses poorly conductive pili (strain Aro5) was previously found to be defective in establishing DIET cocultures with G. sulfurreducens ref. mlf212037-bib-0042 and the Type II Methanosarcina species, M. acetivorans ref. mlf212037-bib-0009. A coculture initiated with G. metallireducens expressing poorly conductive pili and the Type I Methanosarcina species, M. barkeri, was delayed in converting ethanol to methane but adapted to grow in coculture at rates that were similar to the wild‐type cocultures after four transfers (Figure 6). These results suggest that e‐pili are helpful, but not essential for establishing DIET with M. barkeri. Eliminating G. metallireducens e‐pili expression had no impact on DIET between G. metallireducens and Methanobacteium electrotrophus strain YSL, which like M. barkeri lacks outer‐surface c‐type cytochromesref. mlf212037-bib-0012. Thus, it appears that e‐pili are more important for establishing electrical contact in species that rely on outer‐surface c‐type cytochromes (i.e., G. sulfurreducens, M. acetivorans) than species that do not (i.e., M. barkeri, M. electrotrophus strain YSL).
There were also substantial differences in the expression of G. metallireducens genes coding for additional surface‐associated proteins, with different electron‐accepting partners (Table S2). In particular, Gmet_2163, coding for a multicopper protein homologous to OmpB, which G. sulfurreducens requires for Fe(III) oxide reductionref. mlf212037-bib-0043, was more highly expressed in G. metallireducens/G. sulfurreducens cocultures than it was in G. metallireducens cocultures with the Methanosarcina species (Table 1). Further evaluation of the function of the multicopper protein in DIET awaits the construction of the appropriate mutant.
Failure of S. oneidensis to participate in DIET
In addition to illustrating likely differences between routes for DIET from G. metallireducens to G. sulfurreducens versus Methanosarcina species, the results above demonstrate that elucidating the routes for DIET with G. metallireducens is complex because of the high abundance of redox‐active proteins on the outer surface, providing opportunities for adapting alternative routes for extracellular electron transfer. A simpler, yet still genetically tractable system to study DIET is desirableref. mlf212037-bib-0027. Therefore, the possibility for S. oneidensis to serve as the electron‐donating partner in DIET was investigated. Cocultures were initiated with lactate as the electron donor and S. oneidensis as the electron‐donating partner and either M. barkeri, M. acetivorans, M. horonobensis or M. subterranea as the electron‐accepting partner. None of the cocultures grew or produced methane (data not shown). Effective extracellular electron transfer by S. oneidensis is limited to soluble electron acceptors, such as chelated Fe(III), metal ions and electron shuttles, such as flavinsref. mlf212037-bib-0027, ref. mlf212037-bib-0044, ref. mlf212037-bib-0045. S. oneidensis lacks e‐pili and primarily reduces extracellular particulate electron acceptors, such as Fe(III) oxides and electrodes, with flavins as an electron shuttleref. mlf212037-bib-0046, ref. mlf212037-bib-0047, ref. mlf212037-bib-0048. Although soluble electron shuttles have been shown to facilitate interspecies electron transfer between Geobacter species with fumarate serving as the electron acceptorref. mlf212037-bib-0049, the redox potential of reduced flavins may be too positive to support the reduction of carbon dioxide by methanogens. The apparent inability of S. oneidensis to participate in DIET with methanogens is consistent with the finding that Shewanella species are typically not reported to be abundant members of methanogenic communities.
Implications
The results demonstrate that there may be substantial differences in the electron transport routes for DIET between G. metallireducens and G. sulfurreducens versus that between G. metallireducens and methanogens. Possible reasons for this include variations in key outer‐surface electrical contacts and the difference in the redox potential required for the reduction of fumarate versus carbon dioxide. These distinctions may also account for the observation that all the G. metallireducens mutant strains eventually adapted to grow in coculture with Methanosarcina species but not with G. sulfurreducens. Sequencing the genomes of the adapted cocultures might reveal additional mutations associated with these adaptations.
Differences in G. metallireducens transcriptomes and the impact of gene deletions on coculture growth suggested that dissimilarities in electron uptake mechanisms between Type I and Type II Methanosarcina might also influence G. metallireducens physiology during DIET. The results also indicate that the common assumption that extracellular electron transfer to other cells goes through the same pathways as electron transfer to Fe(III) oxide is incorrect, especially for DIET to Methanosarcina species. A number of gene deletions previously shown to inhibit G. metallireducens growth on Fe(III) oxide had no impact on DIET‐based growth with Methanosarcina.
These considerations highlight the need to develop models for DIET with microbes that are relevant to the environment of interest. Although G. metallireducens/G. sulfurreducens cocultures with fumarate as the electron acceptor were convenient to develop the concept of DIETref. mlf212037-bib-0005, ref. mlf212037-bib-0006, ref. mlf212037-bib-0010, ref. mlf212037-bib-0026, ref. mlf212037-bib-0050, ref. mlf212037-bib-0051, DIET is most likely to be environmentally significant under methanogenic conditions. Evaluation of G. metallireducens growing via DIET with other methanogens, such as Methanobacterium ref. mlf212037-bib-0012 and Methanothrix ref. mlf212037-bib-0013 species, which are physiologically and phylogenetically distinct from Methanosarcina species are required to more fully understand potential electron transport pathways for DIET. Further analysis of differences in gene expression patterns in diverse electron‐donating microbes and electron‐accepting methanogens will help identify the important electron transport pathways for DIET in methanogenic environments.
The finding that S. oneidensis was not capable of DIET with Methanosarcina species further demonstrates the often‐ignored substantial differences in the extracellular electron transport capabilities of Shewanella and Geobacter species. Although S. oneidensis is commonly regarded as a model microbe equivalent to Geobacter species for the study of anaerobic biogeochemical processes and bioremediation, it lacks many of the metabolic capabilities of Geobacter speciesref. mlf212037-bib-0027, ref. mlf212037-bib-0052, ref. mlf212037-bib-0053. More broadly, these results have implications for the inference for DIET based on metagenomic analyses and suggest that the mere presence of genes for proteins, such as multiheme c‐type cytochromes, known to be key components for some forms of extracellular electron transfer, is not necessarily sufficient to confer the capacity for DIET.
MATERIALS AND METHODS
Culture media and growth conditions
Methanosarcina acetivorans strain WWM1 (∆hpt)ref. mlf212037-bib-0054 was routinely cultured under strict anaerobic conditions at 37°C in MA medium as previously describedref. mlf212037-bib-0009. M. barkeri MS (DSM 800), M. horonobensis HB‐1 (DSM 21571) and M. subterranea DH‐2ref. mlf212037-bib-0015 were cultivated in the same medium with 1 g/l NaCl as opposed to the 4 g/l NaCl ordinarily found in the MA medium. All Methanosarcina cultures were incubated in an N2–CO2 atmosphere (80:20, vol/vol) with acetate (40 mM) and methanol (20 mM) provided as substrates for growth.
Geobacter metallireducens GS‐15 (ATCC 53774), G. sulfurreducens PCA (ATCC 51573) and S. oneidensis MR‐1 (ATCC 700550) were routinely cultured at 30°C under anaerobic conditions (N2:CO2, 80:20, vol/vol). Eighteen different G. metallireducens c‐type cytochrome deletion mutant strains constructed as previously describedref. mlf212037-bib-0029 were obtained from our laboratory culture collection: ∆Gmet_2896, ∆Gmet_0558, ∆Gmet_0534, ∆Gmet_2928, ∆Gmet_2930, ∆Gmet_0557, ∆Gmet_1868, ∆Gmet_0930, ∆Gmet_0679, ∆Gmet_0825, ∆Gmet_0913, ∆Gmet_0910, ∆Gmet_0912, ∆Gmet_0571, ∆Gmet_0232, ∆fliC, ∆Gmet_2029 and ∆Gmet_0581. All G. metallireducens strains were grown in freshwater mediumref. mlf212037-bib-0055 with ethanol (20 mM) provided as the electron donor and Fe(III) citrate (56 mM) provided as the electron acceptor. For the growth of G. sulfurreducens, acetate (20 mM) was the electron donor and fumarate (40 mM) was the electron acceptor. S. oneidensis was grown with lactate (20 mM) provided as the electron donor and Fe(III) citrate (56 mM) provided as the electron acceptor. Growth of Shewanella required supplementation of 22 mg/l l‐arginine, 22 mg/l l‐glutamine and 44g/l dl‐serine to the medium.
For coculture experiments with Methanosarcina species, G. metallireducens strains and either M. acetivorans, M. barkeri or M. subterranea were grown with 20 mM ethanol provided as the electron donor and carbon dioxide as the electron acceptor at 30°C as previously describedref. mlf212037-bib-0009, ref. mlf212037-bib-0014, ref. mlf212037-bib-0015. Attempted cocultures with S. oneidensis and the various Methanosarcina were grown under the same conditions but lactate (20 mM) was provided as the electron donor and the medium was supplemented with the three amino acids mentioned above. For coculture experiments with Geobacter species, G. metallireducens strains and G. sulfurreducens were grown with ethanol (20 mM) provided as the electron donor and fumarate (45 mM) as the electron acceptor in a freshwater medium as previously describedref. mlf212037-bib-0005. Although G. sulfurreducens can disproportionate fumarateref. mlf212037-bib-0056, previous studies have demonstrated that fumarate is not disproportionated in G. metallireducens/G. sulfurreducens DIET coculturesref. mlf212037-bib-0006. Furthermore, we monitored ethanol oxidation by the coculture to ensure that growth was not due to fumarate disproportionation. All growth rate p values were calculated with an analysis of the variance model using the R Stats package, version 4.0.5ref. mlf212037-bib-0057.
Analytical techniques
A gas chromatograph equipped with a headspace sampler and a flame ionization detector (Clarus 600; PerkinElmer Inc.) was used to monitor ethanol concentrations. Methane in the headspace was measured by gas chromatography with a flame ionization detector GC‐8A (SHIMADZU Scientific Instruments Inc.) as previously describedref. mlf212037-bib-0058 and acetate concentrations were measured with a SHIMADZU high‐performance liquid chromatograph with an Aminex™ HPX‐87H Ion Exclusion column (300mm × 7.8 mm; Bio‐Rad) and an eluent of 8.0 mM sulfuric acid.
Microscopy
Cells were routinely examined by phase‐contrast and fluorescence microscopy (BV‐2A filter set) with a Nikon E600 microscope (Nikon Instruments Inc.).
RNA extraction
Cells were harvested from triplicate 50 ml cultures of G. metallireducens grown via DIET with M. barkeri, M. acetivorans or M. subterranea during the mid‐exponential phase when ~18 mM methane was detected in the headspace.
Cells were split into 50 ml conical tubes (BD Sciences), mixed with RNA Protect (Qiagen Sciences Inc.) in a 1:1 ratio and pelleted by centrifugation at 3000g for 15 min at 4°C. Pellets were then immediately frozen in liquid nitrogen and stored at −80°C. Total RNA was extracted from all samples as previously describedref. mlf212037-bib-0059 and messenger RNA (mRNA) was further enriched from all samples with the MICROBExpress kit (Ambion Inc.).
Illumina sequencing and data analysis
Directional multiplex libraries were constructed with RNA extracted from G. metallireducens/M. barkeri DIET cocultures (GM/MB), G. metallireducens/M. acetivorans DIET cocultures (GM/MA) and G. metallireducens/M. subterranea DIET cocultures (GM/MS). The ScriptSeq™ v2 RNA‐Seq Library Preparation Kit (Epicentre Technologies) was used to make the libraries according to the manufacturer’s instructions and paired‐end sequencing was performed on a Hi‐Seq. 2000 platform at the Deep Sequencing Core Facility at the University of Massachusetts Medical School in Worchester, MA.
Raw data were quality‐checked with FASTQC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/). All read information from libraries constructed for this study (GM/MB, GM/MA, GM/MS, GM/GS) is provided in Table S5 Trimmomaticref. mlf212037-bib-0060 was used to trim and filter all raw paired‐end reads that were then merged with FLASHref. mlf212037-bib-0061. Ribosomal RNA (rRNA) reads were removed from the libraries with SortMeRNAref. mlf212037-bib-0062.
Mapping of mRNA reads
Trimmed and filtered mRNA reads from the triplicate samples for the various coculture conditions were mapped against the G. metallireducens GS‐15 (NC_007517) genome downloaded from IMG/MER (img.jgi.doe.gov) using ArrayStar software (DNAStar). Reads were normalized and processed for differential expression studies using the edgeR package in Bioconductorref. mlf212037-bib-0063. All foldchange and p values were obtained using the glmQLFTest function in edgeR, which is a genewise negative binomial generalized linear model that implements the quasi‐likelihood (QL) methods of Lund et al.ref. mlf212037-bib-0064. All trimmed means of M values were calculated using the calcNormFactors function in edgeR. The plotMDS and plotMD functions in the LIMMA package in RStudioref. mlf212037-bib-0065 were used to generate the MDS and MD plots.
AUTHOR CONTRIBUTIONS
Dawn Holmes and Jinjie Zhou conducted experiments, organized the study and wrote the manuscript. Jessica Smith, Caiqin Wan, and Xinying Liu conducted experiments. Derek Lovley helped to organize the study and to write the manuscript.
ETHICS STATEMENT
No animal or human research was involved in this study.
CONFLICT OF INTERESTS
The authors declare no conflict of interests.
Supplementary Materials
References
- Syntrophy goes electric: direct interspecies electron transfer.. An Rev Microbiol., 2017
- Reverse methanogenesis and respiration in methanotrophic archaea.. Archaea., 2017. [PubMed]
- Direct interspecies electron transfer in anaerobic digestion: a review.. Adv Biochem Eng Biot., 2015
- Sparking anaerobic digestion: promoting direct interspecies electron transfer to enhance methane production.. iScience., 2020. [PubMed]
- Direct exchange of electrons within aggregates of an evolved syntrophic coculture of anaerobic bacteria.. Science., 2010. [PubMed]
- Syntrophic growth with direct interspecies electron transfer as the primary mechanism for energy exchange.. Environ Microbiol Rep., 2013. [PubMed]
- Electron and proton flux for carbon dioxide reduction in Methanosarcina barkeri during direct interspecies electron transfer.. Front Microbiol., 2018. [PubMed]
- Extracellular electron uptake in Methanosarcinales is independent of multiheme c‐type cytochromes.. Sci Rep., 2020. [PubMed]
- Mechanisms for electron uptake by Methanosarcina acetivorans during direct interspecies electron transfer.. mBio., 2021. [PubMed]
- Transcriptomic and genetic analysis of direct interspecies electron transfer.. Appl Environ Microbiol., 2013. [PubMed]
- The Archaellum of Methanospirillum hungatei is electrically conductive.. mBio., 2019. [PubMed]
- Methanobacterium capable of direct interspecies electron transfer.. Environ Sci Technol., 2020. [PubMed]
- A new model for electron flow during anaerobic digestion: direct interspecies electron transfer to Methanosaeta for the reduction of carbon dioxide to methane.. Energy Environ Sci., 2014
- Direct interspecies electron transfer between Geobacter metallireducens and Methanosarcina barkeri .. Appl Environ Microbiol., 2014. [PubMed]
- Correlation of key physiological properties of Methanosarcina isolates with environment of origin.. Appl Environ Microbiol., 2021. [PubMed]
- Extracellular electron uptake by two Methanosarcina species.. Front Energy Res., 2019
- Methane‐fueled syntrophy through extracellular electron transfer: uncovering the genomic traits conserved within diverse bacterial partners of anaerobic methanotrophic archaea.. mBio., 2017. [PubMed]
- Single cell activity reveals direct electron transfer in methanotrophic consortia.. Nature., 2015. [PubMed]
- Intercellular wiring enables electron transfer between methanotrophic archaea and bacteria.. Nature., 2015. [PubMed]
- Syntrophus conductive pili demonstrate that common hydrogen‐donating syntrophs can have a direct electron transfer option.. ISME J., 2020. [PubMed]
- Desulfovibrio feeding Methanobacterium with electrons in conductive methanogenic aggregates from coastal zones.. Water Res., 2021. [PubMed]
- A genetic system for Geobacter metallireducens: role of the flagellin and pilin in the reduction of Fe(III) oxide.. Environ Microbiol Rep., 2012. [PubMed]
- Potential for direct interspecies electron transfer in methanogenic wastewater digester aggregates.. mBio., 2011. [PubMed]
- Metatranscriptomic evidence for direct interspecies electron transfer between Geobacter and Methanothrix species in methanogenic rice paddy soils.. Appl Environ Microb., 2017
- Characterization and modelling of interspecies electron transfer mechanisms and microbial community dynamics of a syntrophic association.. Nat Commun., 2013. [PubMed]
- Expanding the diet for DIET: electron donors supporting direct interspecies electron transfer (DIET) in defined co‐cultures.. Front Microbiol., 2016. [PubMed]
- Electromicrobiology: the ecophysiology of phylogenetically diverse electroactive microorganisms.. Nat Rev Microbiol., 2022. [PubMed]
- PSORTb 3.0: improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes.. Bioinformatics., 2010. [PubMed]
- Outer cell surface components essential for Fe(III) oxide reduction by Geobacter metallireducens .. Appl Environ Microb., 2013
- Syntrophic growth with direct interspecies electron transfer between pili‐free Geobacter species.. ISME J., 2018. [PubMed]
- Proteins involved in electron transfer to Fe(III) and Mn(IV) oxides by Geobacter sulfurreducens and Geobacter uraniireducens .. Microbiology., 2013. [PubMed]
- Anode biofilm transcriptomics reveals outer surface components essential for high density current production in Geobacter sulfurreducens fuel cells.. PLoS One., 2009. [PubMed]
- Extracellular electron transfer mechanisms between microorganisms and minerals.. Nature Rev Microbiol., 2016. [PubMed]
- Protein nanowires: the electrification of the microbial world and maybe our own.. J Bacteriol., 2020
- A pilin chaperone required for the expression of electrically conductive Geobacter sulfurreducens pili.. Environ Microbiol., 2019. [PubMed]
- The electrically conductive pili of Geobacter species are a recently evolved feature for extracellular electron transfer.. Microb Gen., 2016
- Aromatic amino acids required for pili conductivity and long‐range extracellular electron transport in Geobacter sulfurreducens .. mBio., 2013. [PubMed]
- Geobacter sulfurreducens strain expressing Pseudomonas aeruginosa Type IV pili localizes OmcS on pili but is deficient in Fe(III) oxide reduction and current production.. Appl Environ Microbiol., 2014. [PubMed]
- Mechanistic stratification in electroactive biofilms of Geobacter sulfurreducens mediated by pilus nanowires.. Nat Commun., 2016. [PubMed]
- The low conductivity of Geobacter uraniireducens pili suggests a diversity of extracellular electron transfer mechanisms in the genus Geobacter .. Front Microbiol., 2016. [PubMed]
- Direct observation of electrically conductive pili emanating from Geobacter sulfurreducens .. mBio., 2021. [PubMed]
- Geobacter strains expressing poorly conductive pili reveal constraints on direct interspecies electron transfer mechanisms.. mBio., 2018. [PubMed]
- Evidence that OmcB and OmpB of Geobacter sulfurreducens are outer membrane surface proteins.. FEMS Microbiol Lett., 2007. [PubMed]
- Molecular underpinnings of Fe(III) oxide reduction by Shewanella oneidensis MR‐1.. Front Microbiol., 2012. [PubMed]
- Mechanisms for Fe(III) oxide reduction in sedimentary environments.. Geomicrobiol J., 2002
- Shewanella oneidensis MR‐1 nanowires are outer membrane and periplasmic extensions of the extracellular electron transport components.. Proc Natl Acad Sci USA., 2014. [PubMed]
- Secretion of flavins by Shewanella species and their role in extracellular electron transfer.. Appl Environ Microbiol., 2008. [PubMed]
- The colorful world of extracellular electron shuttles.. Annu Rev Microbiol., 2017. [PubMed]
- Syntrophic growth via quinone‐mediated interspecies electron transfer.. Front Microbiol., 2015. [PubMed]
- Magnetite compensates for the lack of a pilin‐associated c‐type cytochrome in extracellular electron exchange.. Environ Microbiol., 2015. [PubMed]
- Promoting direct interspecies electron transfer with activated carbon.. Energy Environ Sci., 2012
- Geobacter: the microbe electric’s physiology, ecology, and practical applications.. Adv Microb Phys., 2011
- Cleaning up with genomics: applying molecular biology to bioremediation.. Nat Rev Microbiol., 2003. [PubMed]
- Development of a markerless genetic exchange method for Methanosarcina acetivorans C2A and its use in construction of new genetic tools for methanogenic archaea.. Appl Environ Microbiol., 2004. [PubMed]
- Geobacter metallireducens gen. nov. sp. nov., a microorganism capable of coupling the complete oxidation of organic compounds to the reduction of iron and other metals.. Arch Microbiol., 1993. [PubMed]
- Fumarate disproportionation by Geobacter sulfurreducens and its involvement in biocorrosion and interspecies electron transfer.. Sci Total Environ., 2022. [PubMed]
- 57 Team R Core . A language and environment for statistical computing. Austria: Vienna; 2012.
- Methane production from protozoan endosymbionts following stimulation of microbial metabolism within subsurface sediments.. Front Microbiol., 2014. [PubMed]
- Genome‐scale analysis of anaerobic benzoate and phenol metabolism in the hyperthermophilic archaeon Ferroglobus placidus .. ISME J., 2012. [PubMed]
- Trimmomatic: a flexible trimmer for Illumina sequence data.. Bioinformatics., 2014. [PubMed]
- FLASH: fast length adjustment of short reads to improve genome assemblies.. Bioinformatics., 2011. [PubMed]
- SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data.. Bioinformatics., 2012. [PubMed]
- edgeR: a bioconductor package for differential expression analysis of digital gene expression data.. Bioinformatics., 2010. [PubMed]
- Detecting differential expression in RNA‐sequence data using quasi‐likelihood with shrunken dispersion estimates.. Stat Appl Genet Mol Biol., 2012
- LIMMA powers differential expression analyses for RNA‐sequencing and microarray studies.. Nucl Acids Res., 2015
