CIPK‐B is essential for salt stress signalling in Marchantia polymorpha
Abstract
Calcium signalling is central to many plant processes, with families of calcium decoder proteins having expanded across the green lineage and redundancy existing between decoders. The liverwort Marchantia polymorpha has fast become a new model plant, but the calcium decoders that exist in this species remain unclear.
We performed phylogenetic analyses to identify the calcineurin B‐like (CBL) and CBL‐interacting protein kinase (CIPK) network of M. polymorpha. We analysed CBL‐CIPK expression during salt stress, and determined protein–protein interactions using yeast two‐hybrid and bimolecular fluorescence complementation. We also created genetic knockouts using CRISPR/Cas9.
We confirm that M. polymorpha has two CIPKs and three CBLs. Both CIPKs and one CBL show pronounced salt‐responsive transcriptional changes. All M. polymorpha CBL‐CIPKs interact with each other in planta. Knocking out CIPK‐B causes increased sensitivity to salt, suggesting that this CIPK is involved in salt signalling.
We have identified CBL‐CIPKs that form part of a salt tolerance pathway in M. polymorpha. Phylogeny and interaction studies imply that these CBL‐CIPKs form an evolutionarily conserved salt overly sensitive pathway. Hence, salt responses may be some of the early functions of CBL‐CIPK networks and increased abiotic stress tolerance required for land plant emergence.
Article type: Research Article
Keywords: calcineurin B‐like protein, calcium signalling, CBL‐interacting protein kinase, evolution, liverwort, salt stress
Affiliations: School of Biological Sciences University of East Anglia Norwich Research Park Norwich NR4 7TJ UK; School of Computing Sciences University of East Anglia Norwich Research Park Norwich NR4 7TJ UK
License: © 2022 The Authors. New Phytologist © 2022 New Phytologist Foundation CC BY 4.0 This is an open access article under the terms of the http://creativecommons.org/licenses/by/4.0/ License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Article links: DOI: 10.1111/nph.18633 | PubMed: 36660914 | PMC: PMC10953335
Relevance: Relevant: mentioned in keywords or abstract
Full text: PDF (4.8 MB)
Introduction
Salinisation of arable land is a leading threat to sustainable global food security and plants demonstrate reduced yield at concentrations as low as 40 mM NaCl (Munns, ref. 2005). It is estimated that 33% of irrigated land globally is afflicted by high salinity, and, with salinised areas increasing by 10% a year, salinisation is predicted to affect > 50% of arable land by 2050 (Jamil et al., ref. 2011). To prepare for increasing agricultural land salinisation, better knowledge of abiotic stress response systems in plants is needed. Release or influx of calcium at different subcellular and tissue localisations acts as an essential secondary messenger in plants, where it is important for cell polarity, directional growth, fertilisation and responses to abiotic and biotic stresses (Knight & Knight, ref. 2001; Li et al., ref. 2006; Boudsocq et al., ref. 2010; Boudsocq & Sheen, ref. 2013; Edel & Kudla, ref. 2015). Hence, understanding calcium signalling responses under salt stimuli is essential for developing new approaches to maintain or increase current global food supply in response to increasing pressures from abiotic stress.
Salt stress initiates calcium signals in plants (Knight et al., ref. 1997; Choi et al., ref. 2014; Huang et al., ref. 2017) that are decoded by various calcium‐binding proteins (Allan et al., ref. 2022). These decoder proteins detect calcium fluctuations by binding calcium through intrinsic helix–loop–helix domains, known as EF‐hands, and fall broadly into sensor‐relay and sensor‐responder types. Sensor‐responders, such as calcium‐dependent protein kinases, detect calcium changes through EF‐hands and then directly phosphorylate downstream proteins via intrinsic kinase domains (Bredow & Monaghan, ref. 2022). Sensor‐relays, such as calcineurin B‐like proteins (CBLs) and calmodulins, have no intrinsic kinase activity but undergo a conformational change upon binding of calcium which allows them to interact with downstream partners to regulate the activity of other signalling proteins (Zhang & Lu, ref. 2003; Kolukisaoglu et al., ref. 2004; Hashimoto et al., ref. 2012). CBL‐interacting protein kinases (CIPKs) are essential to transduce calcium signals in plants through the interactions that they form with CBLs (Albrecht et al., ref. 2001).
Calcineurin B‐like proteins and CIPKs are important for salt stress signalling and were initially characterised in the salt overly sensitive (SOS) pathway of Arabidopsis thaliana. In this pathway, SOS3 (AtCBL4) is activated by a salt‐induced calcium signal in roots. SOS3 then activates SOS2 (AtCIPK24), which transduces the signal by phosphorylating the Na+/H+ antiporter SOS1 (Qiu et al., ref. 2002; Ji et al., ref. 2013) to increase Na+ extrusion, thereby stopping Na+ from entering the transpiration stream (Quan et al., ref. 2007; Quintero et al., ref. 2011). The interaction between SOS3 and SOS2 is dependent on the SOS2 NAF domain and removal of this domain renders SOS2 auto‐active but less capable of activating SOS1 (Albrecht et al., ref. 2001; Quintero et al., ref. 2011). The SOS pathway is in fact more complex as ScaBP8 (AtCBL10) functions in place of SOS3 (AtCBL4) in the shoot (Quan et al., ref. 2007) to interact with SOS2 and activate SOS1 for salt extrusion, and AtCIPK8 can functionally replace SOS2 in the same pathway (Yin et al., ref. 2020). This discovery of many CBL‐CIPKs that can effectively fulfil the same role demonstrates the level of functional redundancy between these proteins and highlights the complexity of calcium signalling during salt stress.
Functional redundancy is common in calcium decoders and can arise from whole genome duplication, but maintenance of selection is required for retaining redundancy. Analysis of the evolution of calcium signalling components has demonstrated that the increase in calcium encoding machineries (such as ion channels) in early land plants was followed by a subsequent increase in decoder components (Edel et al., ref. 2017). To increase the number of calcium decoders, mechanisms to increase gene number would have been required during evolution. Indeed, whole genome duplication and hybridisation are common in later parts of the green lineage (Clark & Donoghue, ref. 2018). Thus far, studies have focussed on these later diverging groups, such as A. thaliana and crops, which have functional redundancy likely due to these whole genome duplication events and maintenance of selection. Studies have analysed the CBL‐CIPK network of the moss Physcomitrium patens (Physcomitrella) (Kleist et al., ref. 2014), but even P. patens has a rather extensive network of six CBLs and eight CIPKs. One of these CBLs (PpCBL6) is not expressed, and an additional CBL (PpCBL5) and CIPK (PpCIPK8) are likely pseudogenes. Together, this implies that there are only four PpCBLs and seven PpCIPKs that are functional and demonstrates the need for maintaining selection to retain redundancy. Of these remaining P. patens CBLs and CIPKs, most form cognate pairs at genomic loci, which suggests that CBLs and CIPKs duplicated in the whole genome duplication events proposed in P. patens (Rensing et al., ref. 2007; Kleist et al., ref. 2014). Most of the PpCIPKs fall into the ‘algal‐type’ clade, which includes AtCIPK24 (SOS2) and AtCIPK8, and hence it could be proposed that these CIPKs function in salt tolerance responses. Indeed, PpCIPK1 has been characterised to function in salt tolerance (Xiao et al., ref. 2021). However, bryophytes diverged from the rest of the green lineage > 400 million years ago and it is therefore essential to characterise multiple bryophytes before salt stress tolerance can be proposed as an ancient function of CBL‐CIPK networks.
Marchantia polymorpha has been recently established as another bryophyte model organism and a range of molecular biology and genetic tools have been developed for this species, including Agrobacterium‐mediated transformation and CRISPR/Cas9 mutagenesis (Bowman et al., ref. 2017; Sauret‐Güeto et al., ref. 2020). Key plant signalling mechanisms are conserved in M. polymorpha, including both auxin and cytokinin signalling, albeit with fewer genes involved (Flores‐Sandoval et al., ref. 2015; Kato et al., ref. 2015; Aki et al., ref. 2019). Similarly, jasmonate signalling components have been discovered in M. polymorpha (Monte et al., ref. 2019). M. polymorpha can also respond to ethylene, but it cannot synthesise ethylene and instead produces the precursor 1‐aminocyclopropane‐1‐carboxylic acid (Li et al., ref. 2020). Although many of the key plant hormone signalling components are present and functional in M. polymorpha, calcium signalling components have not yet been explored. Furthermore, the similarities and differences between signalling in M. polymorpha and other land plants highlight the need to understand signalling, including calcium signalling, in more than one bryophyte. This is particularly necessary to also propose plausible functions for calcium signalling in the last universal common ancestor of land plants. Here, we identify the CBL‐CIPK network of calcium decoders in M. polymorpha and show that specific CBL‐CIPK protein–protein interactions are not found in this bryophyte species. We also demonstrate that M. polymorpha has a CIPK (MpCIPK‐B) with a salt stress signalling role, specifically in response to ionic stress rather than osmotic stress. We propose that M. polymorpha has an evolutionarily conserved SOS pathway and that salt responses may be some of the early functions of CBL‐CIPK signalling networks in land plants.
Materials and Methods
Phylogeny and bioinformatics
Arabidopsis thaliana CBL and CIPK protein sequences were used as a Blast query against predicted protein sequences from Marchantia polymorpha L. ssp. ruderalis (v.5.0, marchantia.info) with an e‐value cut‐off of 1e−50. Sequences were filtered for unique IDs and full length sequences were retrieved. Sequences from P. patens (v.3.0, Phytozome), Klebsormidium nitens (NIES‐2285 v.1.1, plantmorphogenesis.bio.titech.ac.jp), A. thaliana and M. polymorpha (Table S1) were used for multiple alignment with Mafft (v.5.0) followed by trimming with Trimal (v.1.3) and maximum likelihood trees were generated with Iq‐tree (1.6.12, iqtree.cibiv.univie.ac.at) with 1000× bootstrapping (model: JTTDCMut+G4; Kalyaanamoorthy et al., ref. 2017). Trees were visualised with the Interactive Tree of Life (Letunic & Bork, ref. 2021). To identify specific protein domains in the retrieved sequences, scans were performed with the following ProSite IDs: PS50222 (EF‐hand), PS50011 (kinase domain) and PS50816 (NAF domain).
Plant growth and phenotyping
Four accessions of M. polymorpha were used in this study: Takaragaike‐1 (male), Takaragaike‐2 (female), Cambridge‐1 (male) and Cambridge‐2 (female), supplied by the Haseloff laboratory (University of Cambridge). Marchantia polymorpha was maintained on ½MS media (Murashige & Skoog, ref. 1962) with 1% sucrose and 0.8% agar (pH 6.0) and grown under a 16 h photoperiod at 23°C. Phenotyping was carried out with adult plants (>3 wk) by taking c. 5 × 5 mm cuttings and placing them on ½MS plates supplemented with the indicated concentrations of NaCl or sorbitol. Five cuttings were used for each replicate and grown for 7 d. A pooled fresh weight was taken for the five plants and the plant material was then snap frozen in liquid nitrogen for subsequent RT‐qPCR analysis or used fresh to assess chlorophyll content.
Expression analysis by RT‐qPCR
RNA was extracted from plant material using Qiagen RNeasy Plant Minikit following the manufacturer’s instructions. DNA was removed by DNase treatment (Ambion Turbo DNase). RNA quality was confirmed by measuring the absorbance at 260 and 280 nm using a NanoDrop™ 8000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). To test for the absence of contaminating genomic DNA, a PCR was performed using the GoTaq® G2 Green Master mix (Promega) and primers specific for the housekeeping gene Actin1 (ACT1; Table S2). The PCR conditions were as follows: 98°C for 30 s, followed by 30 cycles of 98°C for 10 s, 53°C for 20 s and 72°C for 45 s, followed by 72°C for 5 min. Genomic DNA from wild‐type (WT) plants was used as a positive control. The absence of contaminating genomic DNA was confirmed by the lack of a 154 bp band when the PCR product was analysed by agarose gel electrophoresis. Approximately 1 μg of purified RNA from each sample was also analysed by agarose gel electrophoresis to confirm the presence of the 18S and 28S rRNA double bands. Complementary DNA was synthesised using Superscript™ II (Invitrogen) and oligo‐dT17 with at least 500 ng of RNA as input. RNase activity was inhibited during cDNA synthesis using 1 μl RNAsin. Quantitative RT‐PCR was carried out using SYBR Green (Jumpstart Taq ReadyMix; Sigma‐Aldrich) in 10 μl volumes for three biological and three technical replicates with 1 : 10 diluted cDNA. Measurements were taken on an AriaMX Real‐Time qPCR system (Agilent, Santa Clara, CA, USA). Actin1 and Adenine phosphoribosyl transferase (APT) were used as housekeeping genes. Conditions for RT‐qPCR were as follows: 95°C for 4 min; 40 cycles of 94°C for 30 s, 55°C for 30 s, 72°C for 30 s, followed by cooling and melt curve. Results for each RT‐qPCR were expressed as threshold cycle (C t) values. Sequences of the gene‐specific primers are listed in Table S2. Efficiencies of primer pairs were tested using a dilution series of 10−1–10−5 ng μl−1 of each gene‐specific PCR product and confirmed as 90–110%. It was also confirmed that primers did not amplify control samples without cDNA template. C t values for each gene were averaged across the technical replicates for each cDNA sample. C t values for the two housekeeping genes (ACT1 and APT) were averaged and used as a control. The fold induction (relative expression) for each biological replicate was calculated for treated samples relative to untreated samples using the Pfaffl method (Pfaffl, ref. 2001) and primer efficiencies were calculated for each RT‐qPCR plate (as described earlier). The final fold induction was calculated by averaging the results from all three biological replicates.
Golden Gate cloning
All constructs were assembled using the MoClo Golden Gate cloning system (Weber et al., ref. 2011) according to Feike et al. (ref. 2019). Briefly, Level 0 modules were ordered as synthesised DNA parts (Invitrogen GeneArt) or domesticated from cDNA clones from Tak‐1 to remove internal BsaI and BpiI restriction enzyme sites using the overlap extension PCR or PCR amplification and scarless Golden Gate assembly. Level 1 parts were assembled in one‐step restriction–ligation reactions using BsaI, transformed into chemically competent Escherichia coli DH5α cells and selected with 100 μg ml−1 ampicillin and blue/white screening. Level 2 multigene constructs were made using BpiI in one‐step restriction–ligation reactions, transformed into chemically competent E. coli DH10B cells and selected with 25 μg ml−1 kanamycin and red/white screening. All final Golden Gate constructs (Table S3) were validated by colony PCR using gene‐specific primers, diagnostic restriction enzyme digestions and Sanger sequencing. Constructs for bimolecular fluorescence complementation (BiFC), split luciferase assays and CRISPR/Cas9 mutagenesis were transformed into Agrobacterium tumefaciens strain GV3101, with resulting colonies selected by colony PCR using gene‐specific primers.
Yeast two‐hybrid assays
Saccharomyces cerevisiae was transformed following the lithium acetate transformation procedure (Gietz & Woods, ref. 2002). Yeast strain AH109 was transformed with constructs containing the GAL4‐BD fused to the N‐terminus of each CBL (based on pDEST‐GBKT7), and strain Y187 was transformed with constructs containing GAL4‐AD fused to the N‐terminus of each CIPK (based on pDEST‐GADT7). AH109 and Y187 strains were mated to generate diploid yeast strains containing different pairs of GAL4‐BD‐CBL and GAL4‐AD‐CIPK constructs. Five biological replicates from each mating were taken and yeast colony PCR, with gene‐specific primers, was used to verify the presence of both constructs. To determine protein–protein interactions, yeast growth was assessed on synthetic dropout (SD) media lacking Leu and Trp (SD‐LW; nonselective growth on control plate) and SD media lacking Leu, Trp and His (SD‐LWH; selective growth for interaction test). Diploid yeast cells were grown in YPAD at 30°C overnight, then diluted to an OD600 of 1.5, and a 1 : 10 serial dilution was spotted onto control and interaction test plates. Yeast growth was assessed after 4 d growth at 30°C. Three technical replicates were performed, with all replicates showing similar results.
Western blotting
The five independent biological replicates used for each yeast two‐hybrid interaction test were grown at 30°C in a 50 ml selective culture (SD‐LW with 100 μg ml−1 carbenicillin) until an OD600 of 0.4–0.6 was reached. The cells were pelleted by centrifugation (10 min, 200 g), washed by re‐suspension in 20 ml of water and re‐pelleted again (10 min, 200 g). OD units were calculated by multiplying the final OD600 by the ml of culture. Per OD unit, 10 μl of Laemmli buffer (Laemmli, ref. 1970) was added to the pelleted cells to make a normalised re‐suspension for each interaction. The yeast cells and Laemmli buffer were then boiled at 95°C for 30 min to lyse the cells. SDS‐PAGE was performed on the samples using a 10% resolving gel run at 130 V for 2 h. The proteins were transferred to the PVDF membrane via wet transfer at 100 V for 80 min at 4°C. Membranes were blocked with TBS‐T containing 5% milk for 1 h and then washed with TBS‐T. Membranes were then incubated overnight at 4°C with 10 ml of TBS‐T containing either anti‐c‐myc‐peroxidase antibody (1 : 10 000, A5598; Sigma) or anti‐HA primary antibody (1 : 10 000, H6908; Sigma). Anti‐HA gels were washed three times with 20 ml of TBS‐T and then incubated with peroxidase‐conjugated secondary antibody (1 : 10 000, A0545; Sigma) in TBS‐T for 1 h at room temperature. Peroxidase activity was detected using the ECL Select Western Blotting Detection Reagent (Cytiva, Amersham, UK) following the manufacturer’s instructions.
Protein–protein interaction tests in Nicotiana benthamiana
Nicotiana benthamiana plants were grown at 23°C under long‐day (16 h : 8 h, light : dark) conditions in compost for 4–5 wk before infiltration. Plant transformation was performed by infiltration using Agrobacterium tumefaciens strain GV3101 transformed with constructs of interest via freeze–thaw transformation or electroporation, as described by Feike et al. (ref. 2019). Bimolecular fluorescent complementation constructs contained the C‐terminal half of Venus fused to each CIPK (VenusC‐CIPK), or the N‐terminal half of Venus fused to each CBL (CBL‐VenusN). The Venus halves used were as described by Waadt et al. (ref. 2008), and full length cDNA sequences (from ATG start codon) of CIPKs (with stop codon) and CBLs (no stop codon) were used. Co‐infiltration with two A. tumefaciens strains (GV3101), diluted to OD600 of 0.5, was performed to assess CBL‐CIPK interactions in a pairwise manner. Split luciferase complementation assay constructs contained the SmBiT NanoLuc luciferase (Promega) fused to the CIPK of interest (SmBiT‐CIPK), the LgBiT NanoLuc luciferase (Promega) fused to the CBL of interest (CBL‐LgBiT) and a GUS transformation marker. Protein–protein interactions were assessed 2–3 d after infiltration with A. tumefaciens. Bimolecular fluorescence complementation was assessed by confocal microscopy (LSM 880 with Airyscan; Zeiss, Oberkochen, Germany) with 488 nm excitation, and emission detected from 490 to 543 nm. Images were analysed using ZEN Black 2.1.fluorescence microscopy software. For split luciferase assays, luciferase activity measurements were performed as described by Feike et al. (ref. 2019), although using Nano‐Glo® Live Cell Assay System kit (Promega) containing substrate, dilution buffer and water in a 1 : 50 : 49 ratio and a Hidex Sense microplate reader.
CRISPR/Cas9 mutagenesis
The CRISPR/Cas9 construct contained the Cas9 enzyme (NLS‐pcoCas9), two single guide RNAs designed to target the first exon of CIPK‐B (expressed from the M. polymorpha U6 promoter) and a hygromycin resistance cassette (p35S‐hptI‐tNOS) for selection of positive transformants. Marchantia polymorpha transformation was carried out using A. tumefaciens strain GV3101. Cam‐2 gemmae were co‐cultivated with A. tumefaciens for 3 d in infiltration buffer (10 μM MgCl2, 10 μM MES buffer (pH 5.6), 200 μM acetosyringone) with shaking (200 rpm) in the dark at 23°C. Positive transformants were selected on hygromycin (10 μg ml−1), grown to the T1 generation by propagation of gemmae and confirmed to contain a deletion in CIPK‐B by PCR amplification of exon 1 from extracted genomic DNA. Sanger sequencing was performed on PCR products to determine the precise mutation present in each cipk‐b mutant line.
Chlorophyll content
Chlorophyll content was assessed using an adapted protocol from Caesar et al. (ref. 2018). Briefly, 10 μl of 200 mM Na2CO3 was added to weighed fresh plant tissue, followed by 500 μl dimethyl sulfoxide (DMSO). The plant tissue was then homogenised using a micro‐pestle and incubated at 65°C for 90 min. Another 500 μl DMSO was added and the incubation was repeated. The samples were then centrifuged (2500 g for 10 min) and 200 μl of each sample was added to a clear 96‐well plate in triplicate. The absorbance was then measured at 648, 665 and 700 nm using a Hidex Sense microplate reader. In the case of the 665 nm absorbance reading being > 0.8, the sample was diluted in a 1 : 1 ratio of DMSO to sample. Chlorophyll content was calculated using the following equation:
where DF is the dilution factor (usually 1) and S is the amount of solvent (in this case 0.2 ml), and then normalised per unit fresh weight.
Results
Marchantia polymorpha has a simplified CBL‐CIPK network
To determine the complexity of the CBL‐CIPK network in M. polymorpha, we performed phylogenetic analyses to identify CBL‐CIPK proteins in the v.5.0 release of the M. polymorpha genome. Our analysis identified three CBLs in M. polymorpha with representation in the two main types of CBLs (Fig. 1a). MpCBL‐A was grouped with A. thaliana CBLs known to localise to the tonoplast, but did not have the motif proposed to confer tonoplast localisation (Batistič et al., ref. 2012; Tang et al., ref. 2012). MpCBL‐A also did not contain the MGCxxS/T myristoylation and palmitoylation motif for localisation to the plasma membrane. MpCBL‐B/C both fall into the group of AtCBLs known to localise to the plasma membrane and have the required MGCxxS/T motif for localisation (Fig. S1). None of the three M. polymorpha CBLs group closely with the A. thaliana salt‐responsive CBLs, that is, AtCBL4 (SOS3) or AtCBL10 (ScaBP8), although MpCBL‐B/C are both Type I CBLs like AtCBL4. The three MpCBLs all contain the 14 amino acid‐long first EF‐hand loop, which is a key feature of CBLs, as well as a phosphorylation site (within a conserved FPSF motif) that is known to be regulated by CIPKs (Hashimoto et al., ref. 2012; Fig. S1) and therefore it is likely that this regulation via phosphorylation is present in early‐diverging land plants.
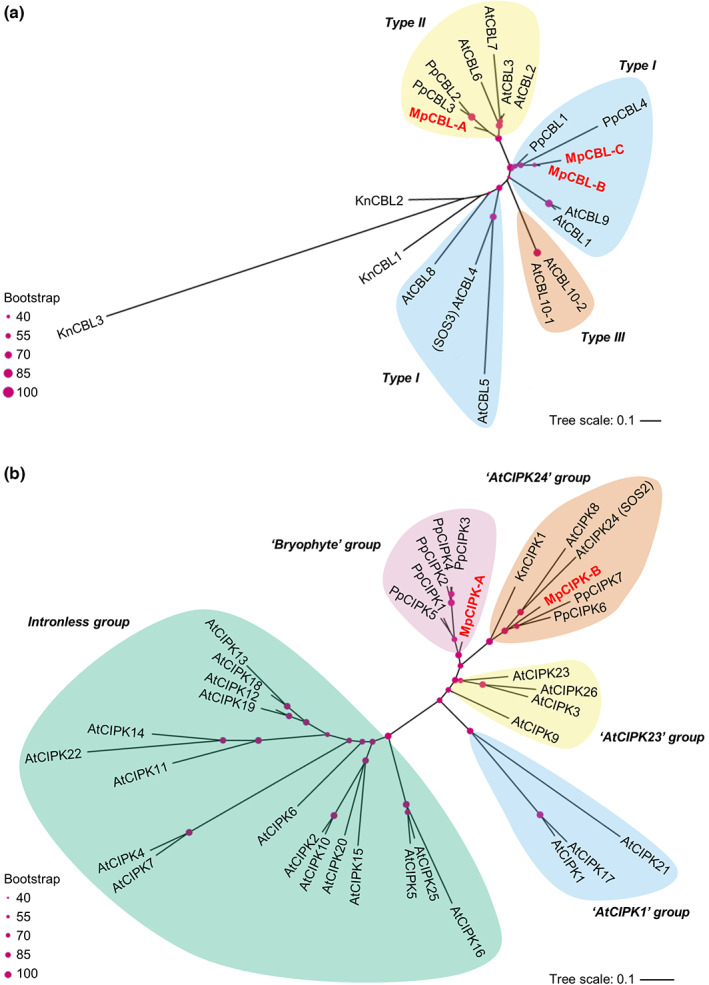
Two MpCIPKs were identified in v.5.0 of the M. polymorpha genome (Fig. 1b). Two clades of CIPKs are known and defined based on intron number, with the intronless clade seemingly arising in angiosperms (Zhu et al., ref. 2016). Both MpCIPKs fall into the intron‐rich clade of CIPKs, as would therefore be expected, and both MpCIPKs contain the NAF domain that is both necessary and sufficient for CBL‐CIPK interactions (Albrecht et al., ref. 2001; Fig. S2). Seven other potential MpCIPKs were identified but excluded based on the lack of a detectable NAF domain. MpCIPK‐A falls into the ‘bryophyte’ group with PpCIPK1/2/3/4/5, which seems to represent a group that diverged in bryophytes and has no representatives from A. thaliana or K. nitens (Fig. 1b). MpCIPK‐B is similar to PpCIPK7 from P. patens and falls into the ‘AtCIPK24’ group, which harbours AtCIPK8 and AtCIPK24 (SOS2). Klebsormidium nitens has a single CIPK that falls into the ‘AtCIPK24’ group which may imply that this is the earliest clade of CIPKs present in land plants (Kleist et al., ref. 2014). Both AtCIPK8/24 and PpCIPK1 operate in salt sensitivity pathways (Quan et al., ref. 2007; Yin et al., ref. 2020; Xiao et al., ref. 2021), implying that salt stress signalling might be the original function of CIPKs in the ‘AtCIPK24’ group and in fact the main function of the CIPKs in M. polymorpha.
Different M. polymorpha accessions show similar salt stress tolerance responses
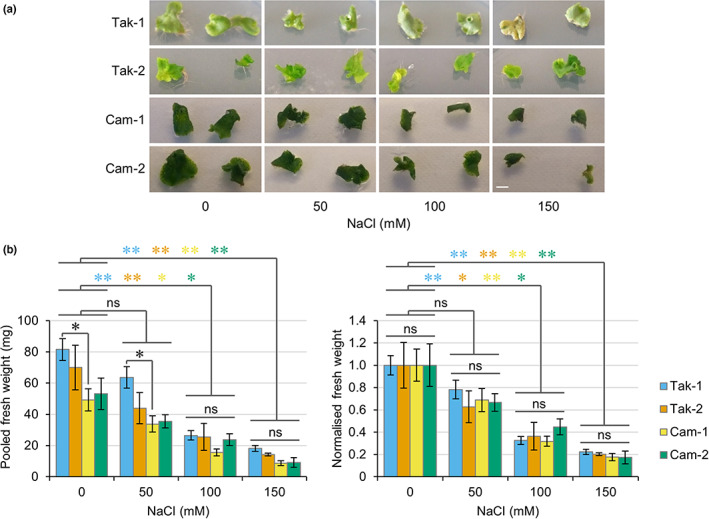
To establish whether common salt tolerance mechanisms exist in M. polymorpha, we compared the salt stress responses of different plant accessions. Two M. polymorpha accessions from different regions have been widely used in research to date: Tak‐1/2 from Takaragaike Park in Kyoto (Japan) and Cam‐1/2 from Cambridge (United Kingdom). Utilising common nomenclature, male plants are labelled 1 (e.g. Tak‐1) and female plants are labelled 2 (e.g. Tak‐2). Male and female plants of both accessions were grown in the presence of different concentrations of salt for 1 wk, after which growth was assessed and total fresh weight was measured. All plants survived the treatment of 150 mM NaCl for 1 wk and decreased growth was observed in response to increasing salt concentration (Fig. 2a,b). Treatments with 100 and 150 mM NaCl caused significant decreases in fresh weight relative to control plants, but treatments with 50 mM NaCl did not cause a significant decrease (Fig. 2b). Since statistical differences in fresh weight were observed between thallus tissue of the same size from Tak‐1 and Cam‐1 plants, normalised fresh weight was used to compare between accessions (Fig. 2b). These comparisons reveal that Tak‐1, Tak‐2, Cam‐1 and Cam‐2 plants all demonstrate similar salt tolerance responses from 0 to 150 mM salt, with no significant differences observed between male and female plants or between the Takaragaike and Cambridge accessions (Fig. 2b).
CIPK‐B expression is regulated by salt stress
To determine whether any of the M. polymorpha CIPKs and CBLs function in a salt sensitivity pathway, expression of the MpCBLs and MpCIPKs was investigated in plants grown for 1 wk in the presence of 0–150 mM NaCl. In A. thaliana, the SOS pathway is upregulated in the first 24 h of salt stress but then downregulated after day 6 (Ji et al., ref. 2013; Rolly et al., ref. 2020). We therefore expected early salt‐responsive CBLs and CIPKs in M. polymorpha to also show downregulated expression after a salt stress treatment of 1 wk. Cam‐2 (Fig. 3) and Tak‐1 (Fig. S3) plants showed strong upregulation of the abiotic stress marker gene LEA‐like4 upon salt treatments and both accessions had salt‐responsive MpCIPKs and MpCBLs. In Cam‐2, both MpCIPK‐A/B were downregulated in response to salt treatments from 100 to 150 mM NaCl (Fig. 3a). A similar downregulation of MpCIPK‐A/B was also observed in Tak‐1 at 50 mM NaCl (Fig. S3). Interestingly, in both accessions, downregulation of CIPK‐B was observed in a greater number of salt stress treatments than CIPK‐A. Therefore, MpCIPK‐A/B likely maintain functions in either salt‐ or drought‐responsive pathways, with CIPK‐B possibly more important in these processes. MpCBL‐C was downregulated in response to all salt stress treatments in Cam‐2 (Fig. 3a), and MpCBL‐C was also downregulated in Tak‐1 under 50 and 100 mM NaCl treatments (Fig. S3). MpCBL‐A/B showed some downregulation in response to salt stress, but this was more modest than MpCBL‐C downregulation and somewhat variable between accessions. The consistent and strong downregulation of MpCBL‐C expression in both accessions implies that MpCBL‐C may function in salt‐ or drought‐responsive tolerance in M. polymorpha more widely. By contrast, changes in MpCBL‐A/B expression were not consistent between the two accessions and therefore MpCBL‐A/B may have subtly different functions or regulation between the accessions.
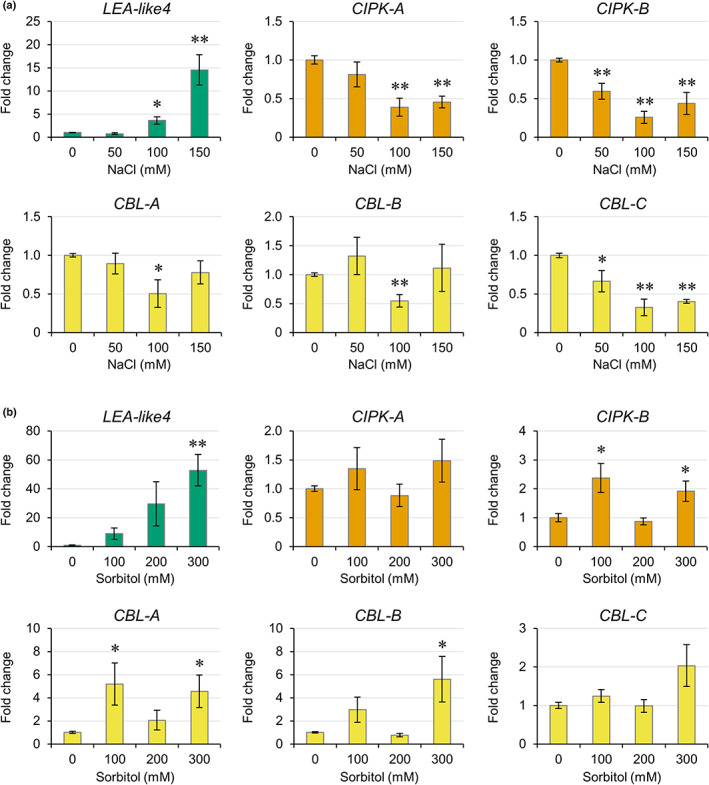
As MpCBL‐C and MpCIPK‐B show more pronounced transcriptional changes in two accessions grown under salt stress, it is most likely that these are the salt‐responsive CBL and CIPK in M. polymorpha. However, these transcriptional responses could also be caused by the drought component of salt stress. To determine whether CBL‐C and CIPK‐B therefore play roles in osmotic stress, we grew Cam‐2 plants for 1 wk in the presence of isoosmolar concentrations of the osmoticum sorbitol and assessed gene expression via RT‐qPCR. The abiotic stress marker gene LEA‐like4 was again strongly induced by growth on treatment plates (Fig. 3b). MpCIPK‐A showed no significant changes in gene expression, but MpCIPK‐B showed significant upregulation with sorbitol (Fig. 3b). Of the CBLs, MpCBL‐A/B both showed significant upregulation, while MpCBL‐C expression was unchanged with sorbitol (Fig. 3b). This suggests MpCBL‐A/B may play roles in osmotic stress, while the likely function of MpCBL‐C in salt stress may be due to ionic rather than osmotic stress. Overall, the data show that the transcriptional responses of CIPK‐B and CBL‐C are different upon salt stress and osmotic stress, and that the downregulation of these genes during salt stress is specific to ionic stress.
CBLs and CIPKs from M. polymorpha do not form specific protein–protein interactions
Protein–protein interactions between different CBLs and CIPKs were assessed, including AtSOS3 (AtCBL4) and AtSOS2 (AtCIPK24), to determine which MpCBLs and MpCIPKs can interact and whether any of the MpCBLs and MpCIPKs may constitute a SOS pathway in M. polymorpha. Yeast two‐hybrid assays revealed that MpCIPK‐A could interact with all MpCBLs and AtCBL4, while MpCIPK‐B interacted with CBL‐A and CBL‐B but not CBL‐C or AtCBL4 (Figs 4a, S4). AtCIPK24 (SOS2) specifically interacted with CBL‐B and CBL‐C. Removal of the NAF domain abolished all CBL‐CIPK interactions (Fig. 4a), confirming that M. polymorpha CBL‐CIPK interactions occur in an equivalent manner to A. thaliana, where the NAF domain is necessary and sufficient for CBL‐CIPK interactions (Albrecht et al., ref. 2001). Based on these protein–protein interactions, we would expect that CBL‐C or CBL‐B could be AtCBL4 homologues. Moreover, MpCIPK‐A could be a potential SOS2 homologue as it can interact with AtCBL4. However, since MpCIPK‐A interacts with all MpCBLs and AtCBL4, it may be that MpCIPK‐A simply forms nonspecific interactions in the yeast system. MpCBL‐B also interacts with both MpCIPKs and AtCIPK24, and may therefore also be nonspecific for the interactions it forms. The similar protein–protein interactions observed for MpCBL‐C and AtCBL4, alongside the MpCBL‐C expression data, support the hypothesis that MpCBL‐C may be a salt‐sensitive AtSOS3 homologue. However, it is notable that CIPK‐B and CBL‐C do not interact in our yeast two‐hybrid assays. This appears to contradict our hypothesis that CIPK‐B and CBL‐C function in salt stress signalling, so we sought to validate all protein–protein interactions assessed by yeast two‐hybrid using additional in planta assays.
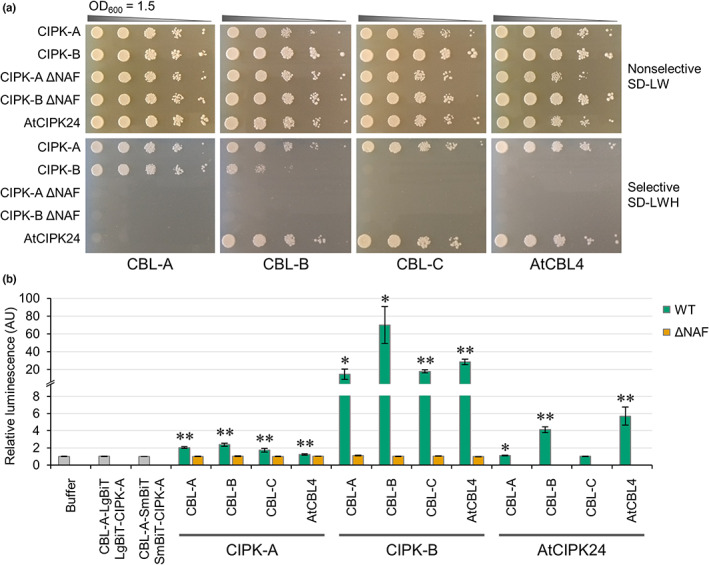
To confirm the results from the yeast two‐hybrid system, a split luciferase assay was performed to determine the interactions formed between MpCBLs and MpCIPKs in N. benthamiana. Importantly, all interactions between MpCBLs and MpCIPKs were detectable in this N. benthamiana system (Fig. 4b), and all interactions were dependent on the presence of the NAF domain. AtCIPK24 (SOS2) formed interactions with MpCBL‐A and MpCBL‐B, but not with MpCBL‐C (Fig. 4b). Bimolecular fluorescence complementation assays were also performed in N. benthamiana leaves, confirming interactions between all MpCBLs and MpCIPKs (Fig. S5). Since the results of the two in planta assays support each other and identified interactions that could not be detected in the yeast two‐hybrid assays, we conclude that MpCBLs do not form specific protein–protein interactions with MpCIPKs. Therefore, all CBL‐CIPK interactions are likely possible in M. polymorpha.
CIPK‐B is involved in tolerance to salt stress but not osmotic stress
Two independent knockout lines of CIPK‐B from M. polymorpha were generated using the CRISPR/Cas9 system. The cipk‐b‐1 mutant contains a 35 bp deletion and the cipk‐b‐2 mutant contains two small deletions and a single nucleotide substitution (Fig. 5), both leading to missense mutations and introduction of a premature stop codon (resulting in predicted proteins containing 29 and 39 amino acids, respectively; Fig. S6). Both cipk‐b mutant lines were smaller than WT (Cam‐2) plants (Fig. 5b), suggesting a possible role for CIPK‐B in plant growth and development. To test whether cipk‐b mutants were impaired in responses to salt stress, equal‐sized pieces of thallus tissue were excised from mature plants, grown in the presence of different concentrations of NaCl for 1 wk, and the fresh weight was measured. To account for any differences in the starting weight between WT and cipk‐b mutant plants, the fresh weight for each line was also normalised to growth on control plates (0 mM NaCl). Increased salt in the growth media resulted in decreased growth of WT plants and both cipk‐b mutants (Fig. 5c,d). However, both cipk‐b mutant lines demonstrated reduced salt tolerance, showing decreased growth and biomass under mild salt stress (50 mM NaCl) compared with WT plants (Fig. 5c,d). Both cipk‐b mutants also showed increased chlorosis and death from as low as 50 mM NaCl compared with WT plants, which showed tolerance up to 150 mM NaCl (Fig. 5c). Chlorophyll content was lower in both cipk‐b mutant lines than in WT plants under all salt stress conditions (Fig. 5e).
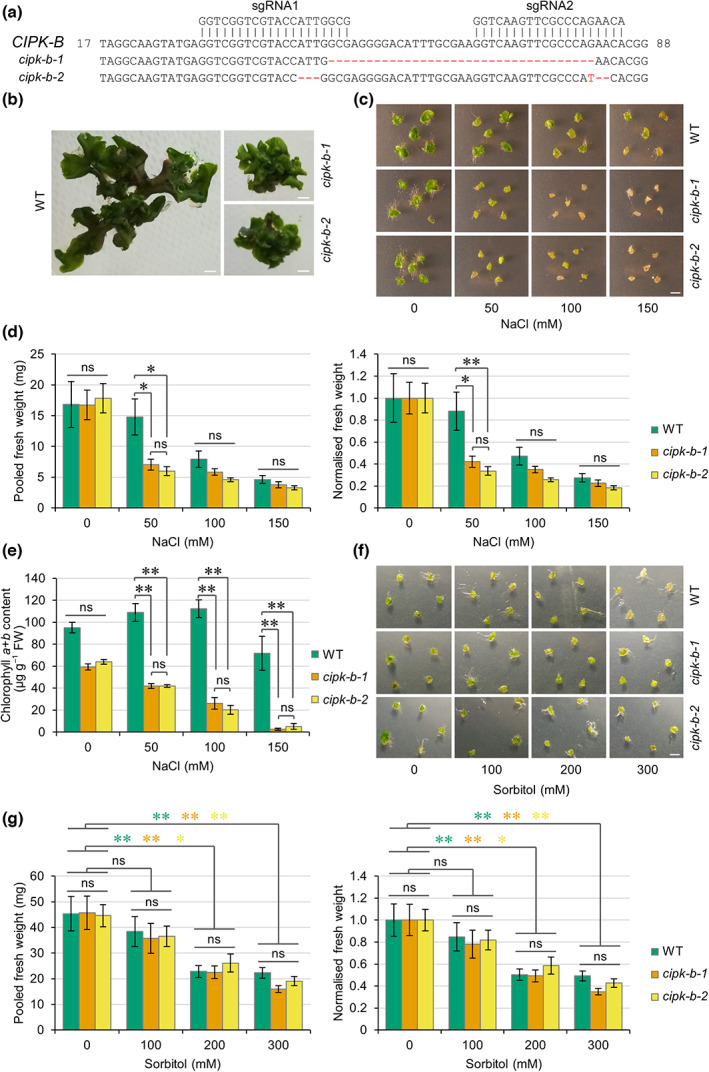
To test whether CIPK‐B is involved in the ionic or osmotic component of salt stress, treatments were also performed using isoosmolar concentrations of sorbitol. Wild‐type and both cipk‐b mutant lines showed decreased growth in response to increasing concentrations of sorbitol (Fig. 5f,g), but no significant difference was observed in the pooled or normalised fresh weight between WT plants and either cipk‐b mutant. We therefore conclude that CIPK‐B does not have a role in osmotic stress and that the salt stress phenotype observed for cipk‐b mutant plants is rather due to ionic responses associated with salt stress.
Discussion
A number of distinct signalling pathways have been characterised in M. polymorpha, including those using auxin, cytokinin, jasmonate and ethylene. Signalling in these pathways is similar to that in angiosperms, but with fewer pathway components or instead utilising precursors of the hormone. Many of these reduced signalling pathways may represent signalling present in the last common ancestor between bryophytes and other land plants. However, there is some evidence of reductive evolution in M. polymorpha, including stomata loss and subsequent development of air pores (Harris et al., ref. 2020). Therefore, to have confidence in inferring the state of the last universal common ancestor of land plants, multiple bryophytes need to be examined. This is also true for understanding calcium signalling in bryophytes. Indeed, calcium signals have been identified in P. patens in response to various stimuli, including salt (Qudeimat et al., ref. 2008; Kleist et al., ref. 2017; Storti et al., ref. 2018). To date, two studies have investigated CBL‐CIPKs in P. patens (Kleist et al., ref. 2014; Xiao et al., ref. 2021), but, to our knowledge, this is the first study of CBL‐CIPK calcium decoders in M. polymorpha. Our work demonstrates a clear role for MpCIPK‐B in salt stress signalling. MpCIPK‐B falls into the same clade as AtCIPK24 (SOS2; Fig. 1b) and cipk‐b knockout mutants showed increased salt sensitivity (Fig. 5). Since cipk‐b mutants did not respond to sorbitol of isoosmolar concentrations, we conclude that MpCIPK‐B is ionic specific and not drought‐responsive more generally. The contrasting expression changes with salt and sorbitol (Fig. 3) suggest that CIPK‐B may even have opposite roles during ionic and osmotic stress.
Previously, the P. patens genome was examined and six PpCBLs and eight PpCIPKs were identified; however, two of these CBLs and one CIPK showed no evidence of being expressed or were pseudogenes (Kleist et al., ref. 2014). Utilising v.5.0 of the M. polymorpha genome, we identified three CBLs and two CIPKs. MpCBL‐B/C were in the Type I CBL group with identifiable MGCxxS/T palmitoylation and myristoylation motifs for plasma membrane targeting, with close grouping to PpCBL4 in a branch containing AtCBL1/9 and PpCBL1. MpCBL‐A fell into the Type II CBL group with no detectable motif for tonoplast targeting, defined as multiple cysteine residues in the first 20 amino acids for S‐acylation (Batistič et al., ref. 2012; Tang et al., ref. 2012). The closest related CBLs to MpCBL‐A include PpCBL2/3 and AtCBL2/3/6/7, which are known to localise to the tonoplast. Whether MpCBL‐A localises to the tonoplast remains to be determined, but this may be interesting to explore further given that MpCBL‐A lacks a canonical tonoplast targeting motif and subcellular targeting motifs in M. polymorpha may be different from angiosperms. No Type III CBLs were identified in this study or previously for PpCBLs or MpCBLs (Kleist et al., ref. 2014; Edel & Kudla, ref. 2015), indicating that this group of CBLs may not be present in bryophytes, or was subsequently lost by both P. patens and M. polymorpha.
CBL‐interacting protein kinases in the last universal common ancestor of land plants were likely to be intron‐rich and subsequent evolution has led to much wider diversification. Two MpCIPKs were identified here and in a previous study (Edel & Kudla, ref. 2015). Of these, MpCIPK‐A falls into a clade of CIPKs alongside PpCIPK1/2/3/4/5 and likely represents a bryophyte group of signalling functions of CIPKs (Kleist et al., ref. 2014). MpCIPK‐B falls into a group including PpCIPK6/7 and AtCIPK8/24, which likely represents an early group that arose with progression onto land. AtCIPK24 (SOS2) is known to confer salt tolerance through phosphorylation‐based activation of SOS1, a Na+/H+ antiporter downstream of calcium decoding, by SOS3 (AtCBL4) or ScaBP8 (AtCBL10). AtCIPK8 has also been characterised in salt tolerance through the same pathway through interaction with ScaBP8 (AtCBL10). In addition to MpCIPK‐B, the only other characterised bryophyte CBL or CIPK involved in salt tolerance of which we are aware is PpCIPK1 (Xiao et al., ref. 2021). Interestingly, PpCIPK1 falls into the bryophyte CIPK clade (with MpCIPK‐A) and not the AtCIPK24 clade (with MpCIPK‐B; Fig. 1b), suggesting that the general function of several of the early land plant CIPKs may also be in response to salt tolerance.
Since CIPK‐B is involved in a SOS pathway in M. polymorpha, we would also expect to observe calcium signals induced upon salt stress in this species and subsequent activation of a SOS1 homologue. These aspects of the M. polymorpha SOS pathway still need to be resolved, but initial blast searches with the A. thaliana SOS1 protein sequence identify two possible SOS1 homologues in M. polymorpha. Whether either of these potential SOS1 homologues are activated by MpCIPK‐B remains to be established. Interestingly, M. polymorpha shows considerable expansion in the number of transporters encoded in its genome (Bowman et al., ref. 2017), so it is also possible that MpCIPK‐B may target other (as yet uncharacterised) transporters to mediate salt stress tolerance. Determining the targets of MpCIPK‐B in future will therefore help elucidate the full SOS pathway in bryophytes and shed light on the evolution of this pathway and stress tolerance mechanism more generally in the green lineage.
Previous studies in A. thaliana have investigated CBL and CIPK expression after 24 h of salt treatment and demonstrated that components of the SOS pathway are upregulated within 24 h but then downregulated at later time points (Ji et al., ref. 2013; Rolly et al., ref. 2020). A number of CBLs and CIPKs are involved in ionic balance in angiosperms (such as AtCBL1/9 and AtCIPK23; Li et al., ref. 2006; Cheong et al., ref. 2007; Ragel et al., ref. 2015) and these too are upregulated within 24 h, despite not being directly involved in Na+ tolerance but rather in Na+/K+ equipoise. We have analysed a previously published RNA sequencing dataset and confirmed that all MpCBLs and MpCIPKs are upregulated within 24 h of salt stress (Wu et al., ref. 2021). As our RT‐qPCR confirms downregulation of CBL‐C, CIPK‐A and CIPK‐B in both the Cam‐2 and Tak‐1 accessions after 1 wk of salt stress (Figs 3, S3), we are confident that either or both MpCIPKs have some involvement in drought and salt tolerance responses alongside MpCBL‐C.
The finding that all MpCBLs and MpCIPKs interact in planta is important. Our M. polymorpha finding is different from the studies on PpCBLs and PpCIPKs, where specific protein–protein interactions have been identified (Xiao et al., ref. 2021), and suggests that either specificity was lost or greater versatility of the CBL‐CIPK network arose following whole genome duplication. Many previous studies have utilised the yeast two‐hybrid method to test CBL‐CIPK interactions and then confirmed positive protein–protein interactions in planta. From our data, it is evident that such an approach may fail to detect important CBL‐CIPK interactions due to the prominence of false‐negative results with the yeast two‐hybrid method. For instance, yeast two‐hybrid assays revealed that MpCIPK‐B could not interact with AtCBL4 and MpCBL‐C (Fig. 4a), which we believe to be the main salt signalling M. polymorpha CBL based on phylogeny (Fig. 1a) and gene expression (Figs 3, S3), but positive interactions were detected between these proteins when tested in planta (Figs 4b, S5). The comparatively small CBL‐CIPK network of M. polymorpha means that we have been able to analyse carefully in planta the protein–protein interactions formed by an entire CBL‐CIPK network. This important analysis reveals a surprising lack of specificity for CBL‐CIPK interactions in M. polymorpha. This lack of specificity may not necessarily mean that all CBL‐CIPK combinations are capable of forming in a single cell at the same time, to transduce different calcium signals into kinase activity and substrate phosphorylation. For example, some CBLs or CIPKs may show different spatial or temporal expression profiles, but more work is needed in future to explore this further. It would also be interesting to investigate P. patens and M. polymorpha CBL‐CIPK interactions further to determine how the specificity of these interactions arose, and whether M. polymorpha CBLs can interact with P. patens CIPKs (or vice versa). Such work may also determine whether evolution of specificity of the protein–protein interaction was primarily driven by the CBL or the CIPK. Previous structural work implies that the specificity of CBL‐CIPK interactions comes from the tertiary structure of the CBL, either allowing or disallowing interactions (Sánchez‐Barrena et al., ref. 2005); however, how phosphorylation causes structural changes that can facilitate interactions, for example as in AtCBL10, is still unknown (Xie et al., ref. 2009).
Together, our work suggests that the general function of several of the early land plant CIPKs is in salt tolerance responses. We suggest that additional functions to regulate other ionic stresses may then have evolved later and possibly as a result of genome duplication that gave rise to subsequent expansion and increased versatility of the CBL‐CIPK network. Additional tests are required to interrogate these early groups of CIPKs and CBLs to understand fully the original functions of calcium signalling and decoder proteins in the evolution of land plants. Further investigation of other calcium decoder families will give an understanding of the evolution of unique calcium decoder function, and how neofunctionalisation occurs in a complex system of encoding, decoding and transducing calcium signals to give rise to tolerance responses. Modulating these signalling processes in plants will be key to future solutions to engineer tolerance responses in crops in the face of global problems of salinisation in agriculture.
Competing interests
None declared.
Author contributions
CT, TW and JBM conceptualised the study. CT and TW carried out the phylogenetic analyses. CT performed the phenotyping of wild‐type plants (Takaragaike and Cambridge accessions) grown under salt stress. CT and AMER carried out gene expression analyses by RT‐qPCR during salt and osmotic stress, respectively. CT performed yeast two‐hybrid analysis, western blotting and BiFC assays. JH carried out split luciferase assays. CT performed CRISPR/Cas9 knockout mutant generation. CT and JH performed the phenotyping of wild‐type and cipk‐b mutant lines grown under salt stress. JH and RDP carried out the chlorophyll measurements. AMER and BW performed the phenotyping of wild‐type and cipk‐b mutant lines grown under osmotic stress. CT and JBM wrote the manuscript, with input and critical feedback from all co‐authors. JH and AMER contributed equally to this work.
References
- Cytokinin signaling is essential for organ formation in Marchantia polymorpha .. Plant and Cell Physiology, 2019. [PubMed]
- The NAF domain defines a novel protein‐protein interaction module conserved in Ca2+‐regulated kinases.. EMBO Journal, 2001. [PubMed]
- Encoding, transmission, decoding, and specificity of calcium signals in plants.. Journal of Experimental Botany, 2022. [PubMed]
- S‐acylation‐dependent association of the calcium sensor CBL2 with the vacuolar membrane is essential for proper abscisic acid responses.. Cell Research, 2012. [PubMed]
- CDPKs in immune and stress signaling.. Trends in Plant Science, 2013. [PubMed]
- Differential innate immune signalling via Ca2+ sensor protein kinases.. Nature, 2010. [PubMed]
- Insights in to land plant evolution garnered from the Marchantia polymorpha genome.. Cell, 2017. [PubMed]
- Cross‐kingdom regulation of calcium‐ and/or calmodulin‐dependent protein kinases by phospho‐switches that relieve autoinhibition.. Current Opinion in Plant Biology, 2022. [PubMed]
- Revisiting chlorophyll extraction methods in biological soil crusts – methodology for determination of chlorophyll a and chlorophyll a + b as compared to previous methods.. Biogeosciences, 2018
- Two calcineurin B‐like calcium sensors, interacting with protein kinase CIPK23, regulate leaf transpiration and root potassium uptake in Arabidopsis.. The Plant Journal, 2007. [PubMed]
- Salt stress‐induced Ca2+ waves are associated with rapid, long‐distance root‐to‐shoot signaling in plants.. Proceedings of the National Academy of Sciences, USA, 2014
- Whole‐genome duplication and plant macroevolution.. Trends in Plant Science, 2018. [PubMed]
- Increasing complexity and versatility: how the calcium signaling toolkit was shaped during plant land colonization.. Cell Calcium, 2015. [PubMed]
- The evolution of calcium‐based signalling in plants.. Current Biology, 2017. [PubMed]
- Characterizing standard genetic parts and establishing common principles for engineering legume and cereal roots.. Plant Biotechnology Journal, 2019. [PubMed]
- A simple auxin transcriptional response system regulates multiple morphogenetic processes in the liverwort Marchantia polymorpha .. PLoS Genetics, 2015. [PubMed]
- Gietz DR , Woods RA . 2002. Transformation of yeast by lithium acetate/single‐stranded carrier DNA/polyethylene glycol method. In: Guthrie C , Fink GR , eds. Methods in enzymology. Cambridge, MA, USA: Academic Press, 87–96.
- Phylogenomic evidence for the monophyly of bryophytes and the reductive evolution of stomata.. Current Biology, 2020. [PubMed]
- Phosphorylation of calcineurin B‐like (CBL) calcium sensor proteins by their CBL‐interacting protein kinases (CIPKs) is required for full activity of CBL‐CIPK complexes toward their target proteins.. Journal of Biological Chemistry, 2012. [PubMed]
- Cytosolic and nucleosolic calcium signaling in response to osmotic and salt stresses are independent of each other in roots of Arabidopsis seedlings.. Frontiers in Plant Science, 2017. [PubMed]
- Gene expression profiling of plants under salt stress.. Critical Reviews in Plant Sciences, 2011
- The salt overly sensitive (SOS) pathway: established and emerging roles.. Molecular Plant, 2013. [PubMed]
- ModelFinder: fast model selection for accurate phylogenetic estimates.. Nature Methods, 2017. [PubMed]
- Auxin‐mediated transcriptional system with a minimal set of components is critical for morphogenesis through the life cycle in Marchantia polymorpha .. PLoS Genetics, 2015. [PubMed]
- Genetically encoded calcium indicators for fluorescence imaging in the moss physcomitrella: GCaMP3 provides a bright new look.. Plant Biotechnology Journal, 2017. [PubMed]
- Comparative phylogenomics of the CBL‐CIPK calcium‐decoding network in the moss Physcomitrella, Arabidopsis, and other green lineages.. Frontiers in Plant Science, 2014. [PubMed]
- Abiotic stress signalling pathways: specificity and cross‐talk.. Trends in Plant Science, 2001. [PubMed]
- Calcium signalling in Arabidopsis thaliana responding to drought and salinity.. The Plant Journal, 1997. [PubMed]
- Calcium sensors and their interacting protein kinases: genomics of the Arabidopsis and rice CBL‐CIPK signaling networks.. Plant Physiology, 2004. [PubMed]
- Cleavage of structural proteins during the assembly of the head of bacteriophage T4.. Nature, 1970. [PubMed]
- Interactive Tree of Life (iTOL) v.5: an online tool for phylogenetic tree display and annotation.. Nucleic Acids Research, 2021. [PubMed]
- Ethylene‐independent functions of the ethylene precursor ACC in Marchantia polymorpha .. Nature Plants, 2020. [PubMed]
- A Ca2+ signaling pathway regulates a K+ channel for low‐K response in Arabidopsis .. Proceedings of the National Academy of Sciences, USA, 2006
- A single JAZ repressor controls the jasmonate pathway in Marchantia polymorpha .. Molecular Plant, 2019. [PubMed]
- Genes and salt tolerance: bringing them together.. New Phytologist, 2005. [PubMed]
- A revised medium for rapid growth and bio assays with tobacco tissue cultures.. Physiologia Plantarum, 1962
- A new mathematical model for relative quantification in real‐time RT‐PCR.. Nucleic Acids Research, 2001. [PubMed]
- Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3.. Proceedings of the National Academy of Sciences, USA, 2002
- SCABP8/CBL10, a putative calcium sensor, interacts with the protein kinase SOS2 to protect Arabidopsis shoots from salt stress.. Plant Cell, 2007. [PubMed]
- A PIIB‐type Ca2+‐ATPase is essential for stress adaptation in Physcomitrella patens .. Proceedings of the National Academy of Sciences, USA, 2008
- Activation of the plasma membrane Na/H antiporter Salt‐Overly‐Sensitive 1 (SOS1) by phosphorylation of an auto‐inhibitory C‐terminal domain.. Proceedings of the National Academy of Sciences, USA, 2011
- The CBL‐interacting protein kinase CIPK23 regulates HAK5‐mediated high‐affinity K+ uptake in Arabidopsis roots.. Plant Physiology, 2015. [PubMed]
- An ancient genome duplication contributed to the abundance of metabolic genes in the moss Physcomitrella patens .. BMC Evolutionary Biology, 2007. [PubMed]
- Salinity stress‐mediated suppression of expression of salt overly sensitive signaling pathway genes suggests negative regulation by AtbZIP62 transcription factor in Arabidopsis thaliana .. International Journal of Molecular Sciences, 2020. [PubMed]
- The structure of the Arabidopsis thaliana SOS3: molecular mechanism of sensing calcium for salt stress response.. Journal of Molecular Biology, 2005. [PubMed]
- Systematic tools for reprogramming plant gene expression in a simple model, Marchantia polymorpha .. ACS Synthetic Biology, 2020. [PubMed]
- Systemic calcium wave propagation in Physcomitrella patens .. Plant and Cell Physiology, 2018. [PubMed]
- Tonoplast calcium sensors CBL2 and CBL3 control plant growth and ion homeostasis through regulating V‐ATPase activity in Arabidopsis .. Cell Research, 2012. [PubMed]
- Multicolor bimolecular fluorescence complementation reveals simultaneous formation of alternative CBL/CIPK complexes in planta .. The Plant Journal, 2008. [PubMed]
- A modular cloning system for standardized assembly of multigene constructs.. PLoS ONE, 2011. [PubMed]
- Evolutionarily conserved hierarchical gene regulatory networks for plant salt stress response.. Nature Plants, 2021. [PubMed]
- Protein kinase PpCIPK1 modulates plant salt tolerance in Physcomitrella patens .. Plant Molecular Biology, 2021. [PubMed]
- Roles of SCaBP8 in salt stress response.. Plant Signaling & Behavior, 2009. [PubMed]
- The protein kinase complex CBL10‐CIPK8‐SOS1 functions in Arabidopsis to regulate salt tolerance.. Journal of Experimental Botany, 2020. [PubMed]
- Calmodulin‐binding protein kinases in plants.. Trends in Plant Science, 2003. [PubMed]
- Evolution of an intron‐poor cluster of the CIPK gene family and expression in response to drought stress in soybean.. Scientific Reports, 2016. [PubMed]
