Tumor-immune hybrid cells evade the immune response and potentiate colorectal cancer metastasis through CTLA4
Abstract
Supplementary Information:
The online version contains supplementary material available at 10.1007/s10238-024-01515-9.
Article type: Research Article
Keywords: Circulating hybrid cells, Tumor-immune hybrid cells, CTLA4, Consensus molecular subtypes, Metastasis, Spatial transcriptomic analysis, Spatial proteomic
Affiliations: grid.10223.320000 0004 1937 0490Faculty of Medicine Siriraj Hospital, Siriraj Cancer Center, Mahidol University, Bangkok, 10700 Thailand; grid.10223.320000 0004 1937 0490Department of Surgery, Faculty of Medicine Siriraj Hospital, Mahidol University, Bangkok, 10700 Thailand; grid.10223.320000 0004 1937 0490Siriraj Center of Research Excellent for Systems Pharmacology, Faculty of Medicine Siriraj Hospital, Mahidol University, Bangkok, 10700 Thailand; grid.5288.70000 0000 9758 5690Knight Cancer Institute, Oregon Health & Science University, Portland, OR 97201 USA; https://ror.org/009avj582grid.5288.70000 0000 9758 5690Cancer Early Detection Advanced Research Center, Oregon Health & Science University, Portland , OR 97201 USA; https://ror.org/009avj582grid.5288.70000 0000 9758 5690Department of Molecular and Medical Genetics, Oregon Health & Science University, Portland, OR 97239 USA; https://ror.org/009avj582grid.5288.70000 0000 9758 5690Department of Cell, Developmental and Cancer Biology, Oregon Health & Science University, Portland, OR 97201 USA
License: © The Author(s) 2024 CC BY 4.0 Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
Article links: DOI: 10.1007/s10238-024-01515-9 | PubMed: 39499374 | PMC: PMC11538261
Relevance: Relevant: mentioned in keywords or abstract
Full text: PDF (3.5 MB)
Background
Distant metastases are one of the most common causes of colorectal cancer (CRC)-related mortality [ref. 1]. The mechanism of metastatic cascade, tumor cells must escape the immune system, extravasate, seed distant sites, and establish a permissive microenvironment for colonization and growth [ref. 2]. As many solid cancers exhibit leukocytic traits, it is postulated that tumor cells fuse with immune cells generating tumor-immune hybrid cells (THCs) that can evade the immune system [ref. 3, ref. 4]. THCs are presumably the precursor to circulating hybrid cells (CHCs), which may be important for cancer plasticity and metastasis. To support this notion, THCs have been demonstrated in animal studies and confirmed in patient specimens, which enhance tumor heterogeneity and impact patient prognosis [ref. 4–ref. 8]. However, many questions also remain to be answered, such as the molecular landscape of cells after fusion, the specific markers of THCs, and their correlation with clinicopathological parameters.
THCs can be easily recognized in cell culture and animal tumor model experiments using common epithelial and leukocyte markers [ref. 3, ref. 4]. However, THCs are much more difficult to detect in human cancers. Detection of CHCs in peripheral blood may provide a potential resource to detect THCs. CHCs are defined as cells that express both carcinoma and leukocyte proteins [ref. 4–ref. 8]. Circulating tumor cells (CTCs), which are shed from the primary or metastatic tumor, are defined by their expression of a carcinoma marker but not leukocyte markers [ref. 9, ref. 10]. Furthermore, clusters of CTCs in cancer patients can be found as tumor-associated circulating endothelial cells, which are termed rare cells because they correlate with the features of the underlying tumor vasculature [ref. 11–ref. 13]. Several studies reported that the number of CHC is greater than CTCs in head and neck, ovarian and gastrointestinal cancers [ref. 4–ref. 7, ref. 14]; therefore, the detection of CHC may serve as a better biomarker to inform tumor biology and metastatic cascade.
CRC is a heterogeneous disease with multiple cross-talk signaling pathways between cancer and the tumor microenvironment to drive cancer progression and invasion [ref. 15, ref. 16]. Recently, CRC has been categorized into the four different subtypes, called consensus molecular subtype (CMS; CMS1-CMS4), and provide the best description of disease heterogeneity at the gene expression level [ref. 17]. CMS4, called mesenchymal, is enriched for microsatellite stable (MSS) tumors and is characterized by appreciable immune infiltration, intermediate between CMS1, called MSI-like, and CMS2, called canonical, and CMS3, metabolic subtypes [ref. 17, ref. 18]. CMS4 with tumor growth factor β (TGFβ)-activated stroma is a feature of poor prognosis in this subtype [ref. 17]. Up regulation of TGFβ signaling is associated with activation of the epithelial to mesenchymal transition (EMT) pathway, and is strongly associated with immune escape in the immune tumor microenvironment [ref. 19], however, the immune system in each CMS of CRC is not obvious.
Many studies have investigated the genomic and immunologic features of immunotherapy response in MSS CRC [ref. 20, ref. 21]. Immunotherapy has shown limited efficacy in MSS CRC; however, some CRC patients with MSS were exceptional response to neoadjuvant botensilimab (BOT), an Fc-enhanced next-generation anti–CTLA4 antibody, alongside balstilimab (BAL; an anti-PD-1 antibody) [ref. 20]. The activity of BOT/BAL regimen in patients with MSS CRC showed not only a significant increase but also a diverse array of immune cells [ref. 20]. A clinical trial of first-line durvalumab and tremelimumab with chemotherapy in RAS-mutated metastatic CRC underlined that CTLA4 expression at the tumor site was associated with better response [ref. 21]. Thus, focusing on the identification of the effective and personalized biomarker such as CHCs THCs and CTLA4 may not only increase the anticancer activity but also reduce the adverse events [ref. 22, ref. 23]. Recognition of THCs may represent a window of opportunity to efficiently discover new drugs for the future immunotherapy. Furthermore, CMS subtype can be different in primary tumor versus the matched liver metastases [ref. 17]; therefore, we hypothesized that immune cells and THCs can be different for tumor cell immune escape in each CMS subtype.
Recently, single-cell RNA sequencing (scRNA-seq) was used to decipher cell heterogeneity, identify CTCs and CHCs, and analyze their phenotypes [ref. 24, ref. 25]. CHCs exhibit gene expression patterns distinct from parental cells but retain the expression of critical genes of each parental cell [ref. 25]. However, scRNA-seq alone provided limited information on the functional consequences upstream or downstream of THC formation in tumor tissues [ref. 26, ref. 27]. Currently, there are commercially available platforms for conducting spatial transcriptomics and proteomics, including the Visium platform from 10 × Genomics and the GeoMx® from NanoString Technologies. GeoMx® digital spatial profiling (DSP) is a commercialized approach that enables high-level multiplex spatial profiling of proteins and RNA in tissue samples within a defined region of interest (ROI) [ref. 28]. The commercial Visium platform can profile mRNA levels in tissues, enabling its widespread application, including spatially resolved single-cell data [ref. 29, ref. 30]. Taken together, multiplex spatial bioimaging will be powerful in deciphering THCs in metastatic physical progression for designing efficient therapies.
In this study, we examined the diversity of CHCs, CTCs, and tumor-associated circulating endothelial cells in patients with stage IV CRC. We identified putative biological functions of THCs in primary CRC and metastatic liver tissues. Finally, we identified putative signaling pathways that are upregulated/promoted or suppressed using spatial proteomic and transcriptomic analysis.
Materials and methods
Patients
All patients and healthy controls were enrolled in the protocol approved by Siriraj Institutional Review Board (SIRB), a certificate of approval number Si348/2019 and Si105/2021. Informed consent was obtained from all subjects. Both groups of subjects were diagnosed by surgeons, colonoscope, and tissue biopsy pathological report. Patients with CRC stage IV who treated consecutively at the Faculty of Medicine Siriraj Hospital were enrolled, whether the patient had other diseases that can cause leukocytosis. All patients were collected peripheral blood before receiving any treatments. After surgical resection, patients were diagnosed TNM stage by using pathological (American Joint Committee on Cancer, AJCC, 7th edition) or clinical classification. Healthy control subjects who were completely done colonoscopy or diagnosed by surgeons at the Faculty of Medicine Siriraj Hospital were enrolled.
Patient samples
All human samples, fixed formalin embedded paraffin (FFPE) tissues, and peripheral blood samples were collected and analyzed with approved protocols in accordance with the ethical requirements and regulations of the Siriraj Institutional Review Board (SIRB), a certificate of approval number Si348/2019 and Si105/2021. Peripheral blood was obtained from stage IV CRC patients (N = 10) before surgical resection and healthy controls (N = 10). For each subject, 20 ml of blood sample in a 10 ml Vacuette® K2EDTA tube, EDTA (Greiner Bio-One, Frickenhausen, Germany) and a 10 ml BD Vacutainer® sodium heparin N tube, heparin (BD, NJ, USA) was collected. A total of six primary FFPE colorectal adenocarcinoma tissues and two matched liver metastases of FFPE were collected from three CRC patients analyzed with their peripheral blood in the Siriraj Cancer Center laboratory, Faculty of Medicine Siriraj Hospital.
Detection and quantification of CTCs and CHCs from human peripheral blood using the IsoFlux liquid biopsy system
Initially, 10 ml of peripheral whole blood samples in EDTA tube from stage IV CRC patients (N = 10) and healthy controls (N = 10) were processed using the CTCs enrichment kit from the IsoFlux liquid biopsy system (Fluxion Biosciences, CA, USA), which is based on EpCAM expression [ref. 31]. Briefly, Leucosep® tubes (Greiner, Kremsmünster, Austria) and ficoll-paque (GE Healthcare, IL, USA) were used, according to the manufacturer’s instructions, to obtain the peripheral blood mononuclear cell (PBMC) fraction. The EpCAM coated magnetic bead and the IsoFlux system were used for EpCAM-positive cell selection [ref. 32]. Cells were stained with the circulating tumor cell enumeration kit (Fluxion Bio-Sciences, CA, USA), according to the manufacturer’s instructions. Briefly, immunofluorescence staining was performed using anti-cytokeratin (CK), anti-CD45 (CD45), and Hoechst 33,342 (nucleus) (IsoFlux CTC Enumeration Kit; Fluxion Bio-Sciences, CA, USA). The stained cells were mounted in multiwell plates with sensoplate glass bottom (Greiner Bio-One, Frickenhausen, Germany) for imaging. Imaging was performed using an inverted epifluorescence motorized microscope (Cytation1 imaging reader; Biotex, VT, USA). Automate quantification was performed for whole sample regions by imaging software: Gen5 3.08 (Biotex, VT, USA) with individuals blinded to the clinical status of the patients or healthy controls. CTCs were identified as those with an intact nucleated cell showing CK+/CD45− [ref. 32]. CHCs were identified as those with an intact nucleated cell showing CK+/CD45+.
Tumor-associated circulating endothelial cells (rare cells) isolation and analysis
For rare cell detection, 10 ml of blood samples in a heparin tube from patients with stage 4 CRC (N = 10) and healthy controls (N = 10) were sent to X-Zell (X-Zell, Bangkok, Thailand) within 12 h. The samples were processed according to the X-Zell procedures [ref. 33]. Briefly, each sample was subjected to red blood cell lysis and CD45-based high flow magnetic white blood cell depletion. The remaining cells were subjected to multiplexed cryo-immunostaining with antibodies directed against CD31 (mouse IgG1 with Alexa Fluor594, WM59; Biolegend, CA, USA), CD34 (mouse IgG1 with Brilliant Violet421, 581; BD Bioscience, CA, USA), CD45 (mouse IgG1 with Pacific Orange, HI30; EXBIO, Vestec, Czech Republic), Vimentin (rabbit IgG1 with Alexa Fluor488, EPR3776; Abcam, Cambridge, UK), pan-Cytokeratin (mouse IgG1 with Pacific Blue, C-11; Abcam, Cambridge, UK), and EpCAM (mouse IgG1 with Pacific Blue, VU-1D9 (CD326); EXBIO, Vestec, Czech Republic). The company analyzed and classified atypical cells.
The nCounter® analysis and molecular classification
RNA extraction and the nCounter® analysis
Total RNA was extracted from two sectioned FFPE tissues (5 μm thickness) using a high purity FFPE RNA isolation kit (Roche Diagnostics, IN, USA), strictly according to the manufacturer’s instructions. The nCounter® analysis system was used to perform the assay (Nanostring Technologies, WA, USA). A pan-cancer progression panel kit was used to measure the expression of 770 genes. The raw counts of each target gene were normalized by the geometric mean counts of 11 housekeeping genes (HRNP1, RPL27, RPL9, RPL6, RPL30, OAZ1, PTMA, RPS29, UBC, RPS12 and RPS16) and spiked controls. A threshold count value equal to 20 was used for background thresholding and normalizing the samples for differences in hybridization.
Classification of subtypes of CRC based on deep learning
A gene expression data set from Siriraj Hospital’s CRC cohort was logarithm transformed and converted from genetic information to functional spectra associated with biological pathway activities. Subsequently, a DeepCC model (DeepCC R package version 0.1.1), containing a trained artificial neural network, was performed to extract advantageous features and classify the Siriraj hospital gene expression data into four CMS classes, CMS1, CMS2, CMS3, and CMS4 [ref. 34]
In situ detection of CTCs and CHC from human peripheral blood using spatial proteomic analysis
PBMC from two stage IV CRC patients and two healthy controls were isolated from whole blood samples in a heparin tube using ficoll-paque (GE Healthcare, IL, USA) following the manufacturer’s instructions. Briefly, whole blood was first diluted with Dulbecco’s phosphate-buffered saline (DPBS; Thermo Fisher Scientific, MA, USA), overlaid on ficoll-paque, and then centrifuged at 800 × g for 20 min without brake. After centrifugation, the mononuclear cells at the interface were transferred to a new tube, diluted with DPBS, and pelleted at 800 × g for 10 min. The cells were resuspended with FACS buffer (Thermo Fisher Scientific, MA, USA). Cells then adhered to poly-d-lysine-coated slides (Thermo Fisher Scientific, MA, USA) through incubation at 37 °C for 15 min, permeabilized with Triton-X (Sigma-Aldrich, MO, USA), and fixed with 4% paraformaldehyde (PFA; Thermo Fisher Scientific, MA, USA) [ref. 4]. Before staining, slides were made with 1X citrate buffer pH 6.0 (Sigma-Aldrich, MO, USA) at high pressure and temperature for 15 min. Morphological markers included Syto13 for nuclei, CD45 for immune cells, Pan-CK (CK) for adenocarcinoma cells (GeoMx® morphology markers; NanoString Technologies, WA, USA) and EpCAM (mouse IgG2 with Alexa Fluor647, 9C4; Biolegend, CA, USA). for the epithelial cell adhesion molecule. The slides were scanned and imaged using a GeoMx® instrument (NanoString Technologies, WA, USA). CHCs were identified as those with an intact nucleated cell showing CK+/CD45+.
Exploring THC in patient FFPE tissues using spatial proteomic analysis
5-μm thick FFPE sections from three CRC patients (six FFPE primary colorectal adenocarcinoma tissues and two matched FFPE liver metastases) were strictly prepared for DSP using manual instruction from the GeoMx instrument and the GeoMx immune cell profiling panel kit with 24 proteins (NanoString Technologies, WA, USA). Briefly, FFPEs were deparaffinized by incubating slides in (R)-(+)-Limonene (Sigma-Aldrich, MO, USA), and rehydrated with various concentrations of ethanol. Antigen recovery was performed with 1X citrate buffer pH 6.0 (Sigma-Aldrich, MO, USA) at high pressure and temperature for 15 min. Morphological markers included Syto13, CD45, CK (GeoMx® morphology markers; NanoString Technologies, WA, USA) and EpCAM(mouse IgG2 with Alexa Fluor647, 9C4; Biolegend, CA, USA).
The stained slides were loaded onto a GeoMx® instrument (NanoString Technologies, WA, USA) and scanned before the regions of interest (ROI) (approximately 20 nuclei/ ROI) were selected. The four colors of the morphological marker demarcate the regions of THC, tumor-immune cells, epithelial cells, and stromal cells. UV illumination was performed, and oligonucleotides were released. The photocleaved oligonucleotides released were collected using a microcapillary tube inspiration robotic system and transferred to a 96-microwell plate. The barcodes were counted in the nCounter® analysis system using standard procedures (NanoString Technologies, WA, USA). Normalized counts were calculated using three housekeeping proteins: GAPDH, HISTONE H3 and S6.
Spatial transcriptomic analysis of THC in FFPE tissues at the gene expression level (Visium CytAssist)
For spatial transcriptomic construction and sequencing, FFPE sections were obtained from two CRC patients (two FFPE primary colorectal adenocarcinoma tissues and two matched FFPE liver metastases). FFPE samples that passed the RNA quality control (DV200 > 50%) were strictly prepared according to the Visium CytAssist spatial gene expression for the FFPE tissue preparation guide (CG000518, 10X Genomics, CA, USA). The library preparation was strictly performed according to the Demonstrated Protocol (CG000520, 10X Genomics, CA, USA) and proceeded with the Visium CytAssist Spatial Gene Expression for FFPE-Tissue Preparation Guide (CG000495, 10X Genomics, CA, USA) by our laboratory at Siriraj Cancer Center, Siriraj Hospital Faculty of Medicine, Mahidol University. Libraries were sequenced by Novogene Co., Ltd. (Singapore) using a NovaSeq 6000 platform (Illumina, CA, USA). For data analysis, Space Ranger 2.1.0 May 2023 (10X Genomics, CA, USA) and GRCh38-2020-A reference were used to process FASTQ files. Downstream analyzes were performed using Loupe Browser 7.0 (10X genomics, CA, USA). The p value reported here has been given by the Wilcoxon’s test and adjusted for multiple tests via the Benjamini–Hochberg procedure.
Pathway analysis
Ingenuity Pathway Analysis Software (IPA 41280214 and 41,280,202, Ingenuity® Systems, https://digitalinsights.qiagen.com/, accessed on February 19, 2024) was used to examine the biological pathways. The IPA software (IPA 412480214 and 41,280,202) uses a manually curated database that contains information from several reputable sources, including published journal articles and gene annotation databases. Fisher’s exact test was used to calculate the probabilities between the input gene set and the pathway.
Statistical analysis
The analyzes were performed using Prism 9 software (GraphPad Software, Inc., CA, USA). The Mann–Whitney U test or the T test was used to compare whether there was a difference in the dependent variable between the two independent groups. The T test or Chi-squared was performed to test the statistical significance of clinical characteristics. Statistical significance was established at p value < 0.05.
Results
Detection of CTCs, CHCs, and rare cells in peripheral blood
Before detection of THC in tissues, we examined the number of CTCs, CHCs, and rare cells in peripheral blood of ten enrolled patients with stage IV CRC (patient group) collected before resection. Furthermore, the number of cell counts was compared with those of ten healthy volunteers (healthy control). Table 1 reports the clinical characteristics of each group in this study. The results showed that total white blood cell counts were significantly higher in the patient group compared to healthy control patients (p value = 0.006).
Table 1: Clinical characteristics and demographics
| Patient group Number, N = 10 N (%) | Healthy control group Number, N = 10 N (%) | P value | |
|---|---|---|---|
| Age, years; median [min–max] | 63.5 [54–71] | 44.5 [26–69] | 0.11 |
| White blood cells, cells/ul; median [min–max] | 8155 [6410–10260] | 6340 [5220–8930] | 0.006* |
| CEA, ng/ml; median [min–max] | 252.8 [7.66–18784] | ||
| Sex | 0.136 | ||
| Male | 7 (70) | 4 (40) | |
| Female | 3 (30) | 6 (60) | |
| Moderately differentiated adenocarcinoma | 10 (100) | ||
| Stage IV (TNM stages) colorectal cancer | 10 (100) | ||
| Clinical T classification | |||
| T4b | 1 (10) | ||
| Pathological T classification | |||
| T3 | 5 (50) | ||
| T4 | 1 (10) | ||
| T4a | 2 (20) | ||
| T4b | 1 (10) | ||
| Clinical N Classification | |||
| N1 | 1 (10) | ||
| Pathological N classification | |||
| N0 | 1 (10) | ||
| N1b | 3 (30) | ||
| N2a | 1 (10) | ||
| N2b | 4 (40) | ||
| Liver metastasis | 9 (90) | ||
| Lung metastasis | 1 (10) | ||
| KRAS mutation status | |||
| Wild type | 5 (50) | ||
| Mutation | 2 (20) | ||
| ND | 3 (30) | ||
| NRAS mutation status | |||
| Wild type | 5 (50) | ||
| Mutation | – | ||
| ND | 5 (50) | ||
| BRAF mutation status | |||
| Wild type | 5 (50) | ||
| Mutation | – | ||
| ND | 5 (50) | ||
| PIK3CA mutation status | |||
| Wild type | 3 (30) | ||
| Mutation | – | ||
| ND | 7 (70) | ||
| HER2 mutation status | |||
| Wild type | 3 (30) | ||
| Mutation | – | ||
| ND | 7 (70) |
ND not detected, CEA carcinoembryonic antigen; The T test or Chi-squared was performed to test the statistical significance of clinical characteristics; *, p value < 0.05
After immunofluorescence staining, Fig. 1A showed that the CK and CD45 marker can identify CTCs (CK+/CD45-) and CHCs (CK+/CD45+. We found that the mean number of CTCs and CHCs showed statistical significance (p value = 0.045 and 0.029, respectively) in peripheral blood from patients with stage IV CRC compared to healthy volunteers. The number of CHCs (range 137–1522) was higher than the number of CTCs (range 0–136) and rare cells (range 0–25).
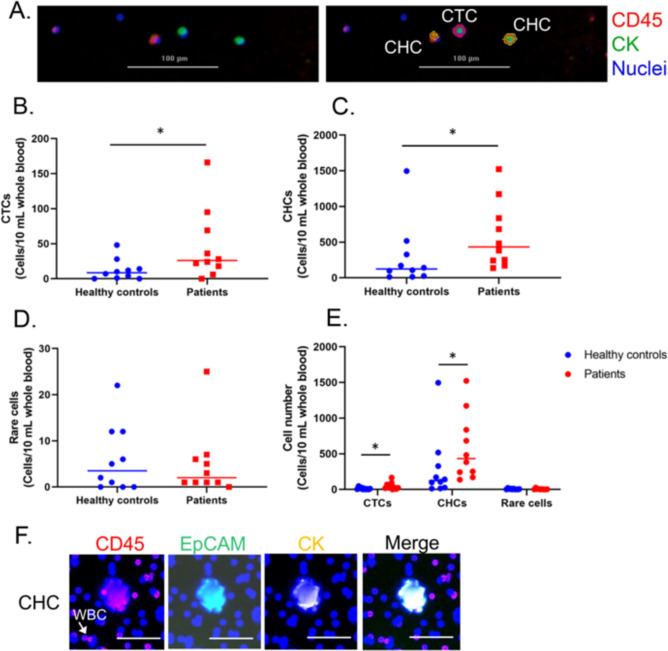
To investigate THC in primary CRC and metastatic liver tissues by spatial proteomic analysis, we added a more specific marker, EpCAM, to identify CHCs. The results showed that the CK, CD45, and EpCAM markers were able to identify CHC using DSP; therefore, these markers were next used to evaluate THC (CK+/EpCAM+/CD45+ in tissues.
Spatial proteomic profiles of THCs in primary CRC and liver metastasis
To examine the biological function of THCs, we compared the THC region with the regions of tumor immune cells, epithelial cells and stromal cells, in primary CRC (Fig. 2A). THC region was defined as the area contained THC and other cells which was approximately 20 nuclei and 2000 surface area. We found that the expression profiles of 21 THC proteins in primary CRC showed region-specific expression patterns highly expressed in stromal cell regions (Fig. 2B). We found that CD3 was significantly down-regulated in THC regions compared to tumor-immune cells (p value = 0.049). To identify the molecular structure of THCs, we determined the varied upregulated proteins in THCs compared to epithelial and stromal cells. We found that CTLA4 was significantly upregulated in THC regions (Fig. 2C); p value = 7.830 × 10–8 and 0.002 for comparing to epithelial and stromal cells, respectively. The THCs showed that KI67 was the same as in the regions of epithelial cells and stromal cells without significant differences (Fig. 2D); p value = 0.804 and 0.981 for comparing to epithelial and stromal cells, respectively. Together, CTLA4 may be a biological characteristic of THCs; while, CD3 proteins may represent immune cells (non-hybrid cells) in tumor tissues.
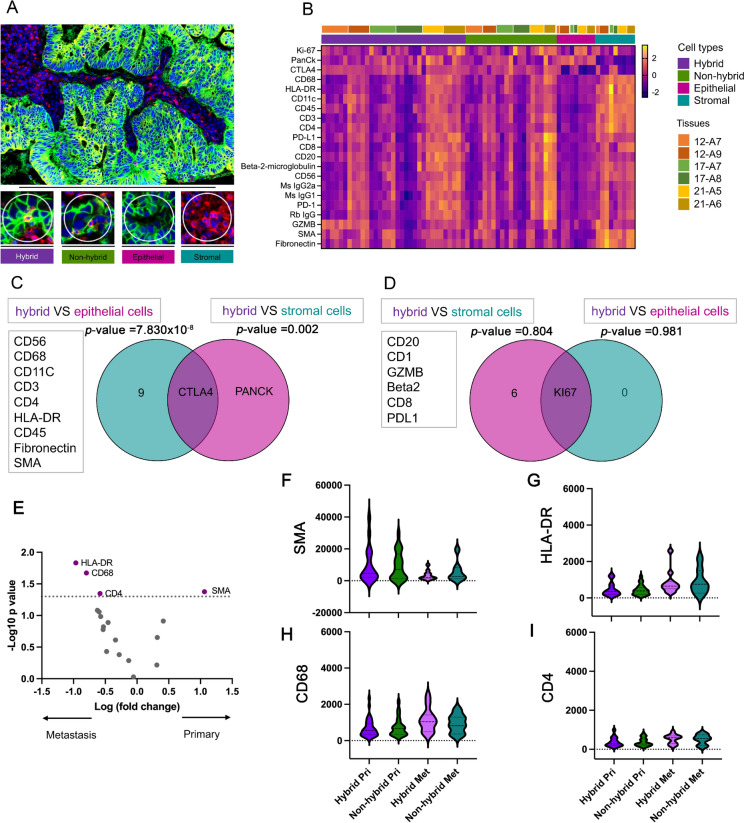
After investigating the biological characteristic of THC in tissues of primary CRC, we determined the characteristics of THCs between primary CRC and liver metastasis. We found that the HLA-DR, CD68, and CD4 proteins were upregulated in the THC region of liver metastasis (Fig. 2E). The SMA protein was significantly overexpressed in the THC region of primary CRC. To characterize the biological characteristics of THC between primary CRC and liver metastasis, we also compared the SMA, HLA-DR, CD68 and CD4 proteins of THC and tumor-immune cell regions between primary CRC and liver metastasis, as shown in Fig. 2F–I. The results showed that tumor-immune cell regions trended toward expressed SMA, HLA-DR, CD68 and CD4 proteins similarly to the THC regions.
Spatial transcriptomic profiles of THCs and tumor-immune cells between primary CRC and matched liver metastases
To comprehensively analyze the prognostic effect of THCs between primary CRC tissues and liver metastatic tissues derived from CRC, we analyzed four tissue samples from two patients (21 and 17) including two primary CRC tissues (C21 and C17) and their matched liver metastatic tissues (L21 and L17). We investigated spatial transcriptomic data, including some spatial proteomic ROI (Fig. 3A). All samples were identified by CMS using their gene expression profiles. The results showed that C21, C17, L21, and L17 were classified as CMS1, CMS2, CMS4 and CMS3, respectively. The cluster of tumor and stromal areas were identified by hematoxylin and eosin (H&E) staining and gene expression profiles of each sample by Visium CytAssist (Fig. 3B). The uniform model approximation and projection (UMAP) analysis indicated the tumor and stromal areas of each sample (Fig. 3C).
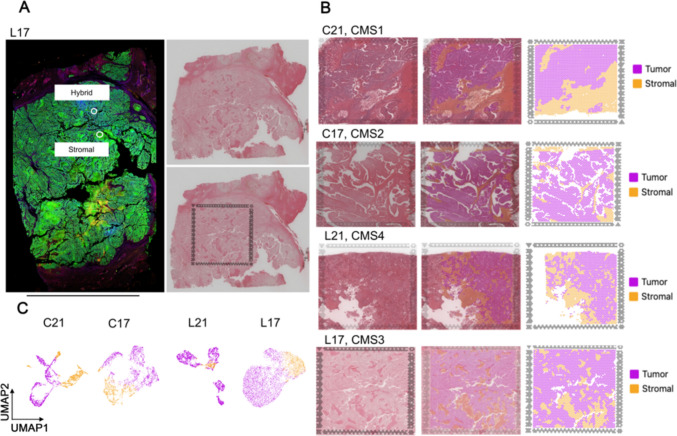
THCs and tumor immune cells between primary CRC and matched liver metastases
In the aforementioned results, we found that CTLA4 protein is a marker of THC regions in primary CRC that may associate with CD68 and CD4 proteins in liver metastatic of CRC. On the contrary, CD3 proteins may be a marker of tumor immune cells in primary CRC. To further explore the characteristic and prognostic value of THCs in primary CRC and in liver metastases, we investigated the distribution of the CD68, CD4 and CTLA4 genes and the distribution of the CD3D, CD3G, and CD3E genes in each tissue section. Spatial transcriptomic analysis showed that the infiltration of CD68, CD4, and CTLA4 genes was higher in patient 21 (Fig. 4A) in which liver metastasis (CMS4) and primary CRC sites (CMS1) was observed compared to the infiltration of these genes in patient 17 (Fig. 4B) in liver metastasis (CMS3) of CRC (CMS2).
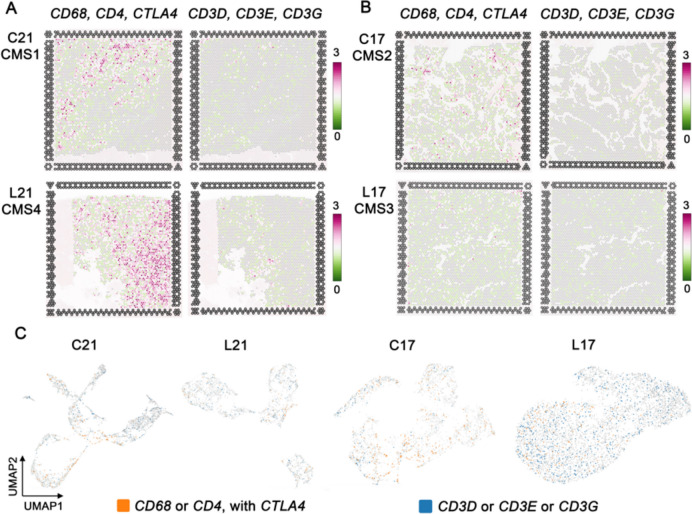
Then we used Loupe Browser 7.0 to group spots with gene characteristics (log2 count > 0) of CD68 or CD4 with CTLA4 as THC, and spots with gene characteristics of CD3D or CD3E or CD3G as tumor immune cells. The feature spot showed highly arranged THCs around the tumor area (Fig. 4C).
Pathway of THCs in the primary and liver metastatic tumors of CRC
To identify significant canonical pathways (p value < 0.05 or -log(p value) > 1.3) between THCs and tumor-immune cells, in the primary and liver metastatic tumors of CRC, we analyzed differentially expressed gene (DEG) between spots with a gene feature of THC and spots with a gene feature of tumor-immune cells using Loupe Browser 7.0 and IPA. We found that the CD68, CD4, and CTLA4 genes are significantly upregulated in spots defined as THC for all tissues in the section. (in section C21, L21, C17 and L17, for CD68 p value = 1.65 × 10–41, 8.70 × 10–45, 7.96 × 10–22, 2.10 × 10–34, for CD4 p-value = 6.44 × 10–28, 5.52 × 10–25, 1.26 × 10–8, 7.65 × 10–35, and for CTLA4 p-value = 2.82 × 10–19, 1.58 × 10–11, 6.65 × 10–16, 6.71 × 10–24, respectively) (Supplementary File), therefore, the clustered spots with those genes could be represented as THC in CRC and metastasis of the matched liver. For patient 21, we found 167 and 60 significant DEG for THC spots in primary CRC and liver metastasis tissues, respectively, however, the CD68, CD4 and CTLA4 genes were only found as significant DEG for THC spots in primary CRC and liver metastasis of patient 17.
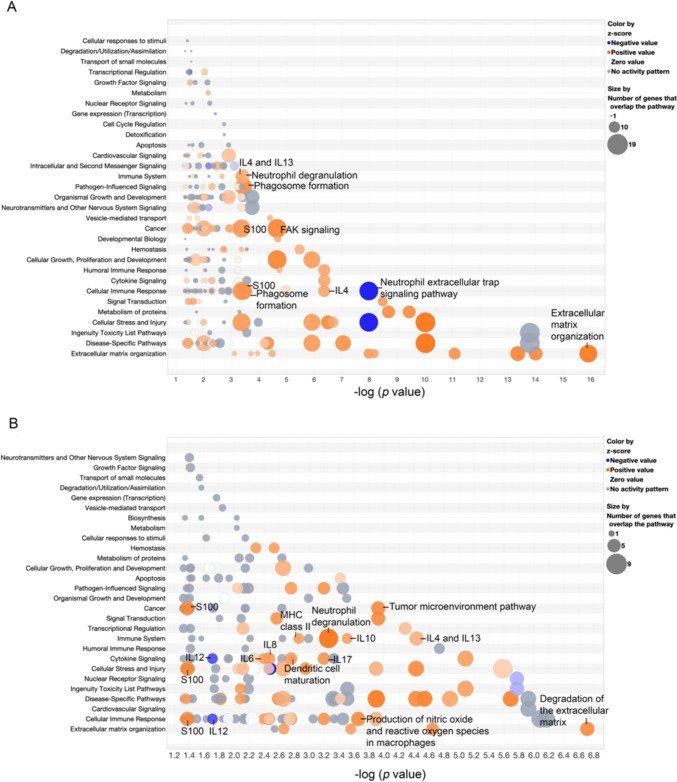
After significant DEGs were imported to the IPA, we analyzed the IPA pathway analysis of THC spots for primary CRC (Fig. 5A, Supplementary File) and for liver metastasis (Fig. 5B, Supplementary File) of patient 21. The results showed that DEGs of THC spots in primary CRC and liver metastases were enriched in activating many critical pathways associated with tumor progression, such as immune system, cancer, cellular immune response, cytokine signaling, and extracellular matrix organization signaling. Interestingly, most of the THC spots’ pathways were involved with myeloid cells, which are especially neutrophils.
Discussion
In this study, we have shown that primary CRC might exploit myeloid and lymphocytes with CTLA4 to facilitate CRC to liver metastases. Currently, there is a wealth of accumulated data to support the hypothesis that THCs are capable of evading the host immune system and progressing to invasive carcinoma [ref. 29]. However, the upstream or downstream functional consequences of THCs in tumor tissues are still missing. Our study tried to close the gaps by elucidating and explaining the molecular pathways and the cell–cell complex between THC and cancer cells.
Our finding indicated that CHCs were found significantly in patients with CRC with liver metastasis outcome, more than in healthy donors. Additionally, lymphocyte cells in patients with CRC have been observed to be higher than those in healthy controls. Chronic inflammation is a common and important contributor to the malignant transformation of many types of solid cancer types, including cholangiocarcinoma, colorectal cancer, lung cancer, and liver cancer [ref. 35–ref. 38]. On the other hand, inflammation has recently reemerged as a paramount target in cancer therapy. Therefore, a significant understanding of immune cells and THCs may benefit for successful immunotherapy in advance cancers.
Here, we reported that CTLA4 exhibits distinct characteristics in THC regions in primary CRC. This result suggests that THC may express CTLA4 or directly increase tumor cells with CTLA4. CTLA4 plays a role in inhibitory receptors (checkpoints) that limit autoreactivity and T cell overactivation [ref. 39], also known as noninflamed tumor or cold tumor [ref. 34]. Previously, similar results were identified in the peripheral blood of patients with advanced stage CRC [ref. 40]. We suggest that CTLA4 may play an important role in the mechanism of immune evasion and metastasis of CHC and THC. Moreover, regions of non-hybrid cells show significant upregulation of CD3. This result shows a differentiation of the cell–cell complex between THC within tumors and between immune cells within tumors. CD3 has been reported to help facilitate T cell receptor signaling activities and the presence of CD3+/CD8+ associated with a good prognosis in colorectal cancer [ref. 41]. Therefore, tumor-immune cells with CD3 can increase a cytotoxic immune response in contrast to the presence of THCs with CTLA4.
Furthermore, HLA-DR, CD68, and CD4 proteins were detected for THC regions in liver metastasis. This result suggested that THC can directly express those proteins or recruit other cells that express those cell surface proteins, such as macrophages (CD68), neutrophils (CD68), and helper T cells (CD4), to facilitate and potentiate liver metastasis. Another interesting finding of our study was the pathway enrichment analysis between THCs and tumor-immune cells. Our results showed that many THC pathways are associated with myeloid cells especially neutrophils. This finding suggested that neutrophils can facilitate the transport of colorectal cancer cells to different tissues [ref. 42].
In addition, we were able to predict the role of the CTLA4, CD68, CD4 genes in the THC spots for each tissue showing different types of CMS. The identification of CMS helps us to understand and explain the molecular characteristic and function of THCs. Previously, critical information identified that immunosuppressive tumor-associated macrophages were enriched in CMS1 and CMS4 [ref. 43]. CD68 expression was positively correlated with phagocytosis and immune cell infiltration, including dendritic cells, monocytes, macrophages, and neutrophils, but not associated with myeloid-derived suppressor cell infiltration [ref. 44]. Anti-CTLA4 immunotherapy can significantly deplete CD68 macrophage in patients with advanced melanoma compared to the untreated sample group [ref. 45]. Immunotherapies targeting CTLA4 have been studied in clinical research against advanced CRC [ref. 20, ref. 21, ref. 46]. In this study, we found that the CTLA4, CD68, and CD4 genes were closely related to THC and CMS1 CRC; therefore, this suggested that CHCs and THCs may act as a new marker of immunotherapies in tumor treatments in the future.
Taken together this finding supported that immune cells and THCs can be different for tumor cell immune escape in each CMS subtype. Our results also supported that bulk transcriptional CMS identification could be a starting point to deepen our knowledge about CRC biology and that its specific immune cell population could support drug discovery and rational combination therapies [ref. 47]. In future studies, it will be important to explore the molecular characteristics of THCs along with CMS classification.
There are some limitations to this study. Firstly, the GeoMx® spatial proteomic and the Visium CytAssist spatial transcriptomic analysis do not provide single-cell colocalization of THCs and immune cells. This study, the analysis area of the spatial proteomic and the spatial transcriptomic analysis include 20 cells and in the range of 1–10 cells, respectively. The biomarkers which were reported here might resulted from the communication among 1–20 cells. However, many of the immune cell types that were evaluated had shown a cell–cell complex in the tumor. Secondly, the probe set of the Visium CytAssist spatial transcriptome lacked the HLA gene family; therefore, HLA was not included in spatial transcriptome analysis. We found the upregulation of HLA-DR in the THC region of liver metastasis, but we could not find HLA gene in our spatial transcriptome analysis. Finally, a small sample size was one of our limitations. In Table 1, the statistically significant "difference" or "not difference" could be due only by chance. Although the small patient cohort limits our study, we envision that CHCs, THCs and CMSs may potentially inform patient-specific treatment strategies. As CHCs and THCs may act as a new marker of immunotherapies in tumor treatments in the future, we anticipate their application to larger CRC cohorts, paving the way toward personalized oncology.
Conclusions
This study explored and addressed a metastatic mechanism of THCs and CHCs. Our data may provide insight into possible improvements in clinical practice to combat metastasis in CRC. We suggest that CHCs and THCs with CTLA4 may act as a new marker of immunotherapies in advance CRC treatments in the future [ref. 40, ref. 41, ref. 45].
Supplementary Materials
References
- RL Siegel, KD Miller, A Goding Sauer, SA Fedewa, LF Butterly, JC Anderson, A Cercek, RA Smith, A Jemal. Colorectal cancer statistics, 2020. CA Cancer J Clin, 2020. [DOI | PubMed]
- CL Chaffer, RA Weinberg. A perspective on cancer cell metastasis. Science, 2011. [DOI | PubMed]
- 3.Henn TE, Anderson AN, Hollett YR, Sutton TL, Walker BS, Swain JR, Sauer DA, Clayburgh DR, Wong MH. Circulating hybrid cells predict presence of occult nodal metastases in oral cavity carcinoma. Head Neck 2021.
- CE Gast, AD Silk, L Zarour, L Riegler, JG Burkhart, KT Gustafson, MS Parappilly, M Roh-Johnson, JR Goodman, B Olson. Cell fusion potentiates tumor heterogeneity and reveals circulating hybrid cells that correlate with stage and survival. Sci Adv, 2018. [DOI | PubMed]
- MS Dietz, TL Sutton, BS Walker, CE Gast, L Zarour, SK Sengupta, JR Swain, J Eng, M Parappilly, K Limbach. Relevance of circulating hybrid cells as a non-invasive biomarker for myriad solid tumors. Sci Rep, 2021. [DOI | PubMed]
- RM Whalen, AN Anderson, JA Jones, Z Sims, YH Chang, MA Nederlof, MH Wong, SL Gibbs. Ultra high content analyses of circulating and tumor associated hybrid cells reveal phenotypic heterogeneity. Sci Rep, 2024. [DOI | PubMed]
- BS Walker, TL Sutton, L Zarour, JG Hunter, SG Wood, VL Tsikitis, DO Herzig, CD Lopez, EY Chen, SC Mayo. Circulating hybrid cells: a novel liquid biomarker of treatment response in gastrointestinal cancers. Ann Surg Oncol, 2021. [DOI | PubMed]
- 8.Parappilly MS, Chin Y, Whalen RM, Anderson AN, Robinson TS, Strgar L, Sutton TL, Conley P, Klocke C, Gibbs SL et al. Circulating neoplastic-immune hybrid cells predict metastatic progression in uveal melanoma. Cancers (Basel) 2022;14(19).
- TA Yap, D Lorente, A Omlin, D Olmos, JS de Bono. Circulating tumor cells: a multifunctional biomarker. Clin Cancer Res, 2014. [DOI | PubMed]
- SH Au, BD Storey, JC Moore, Q Tang, YL Chen, S Javaid, AF Sarioglu, R Sullivan, MW Madden, R O’Keefe. Clusters of circulating tumor cells traverse capillary-sized vessels. Proc Natl Acad Sci U S A, 2016. [DOI | PubMed]
- I Cima, SL Kong, D Sengupta, IB Tan, WM Phyo, D Lee, M Hu, C Iliescu, I Alexander, WL Goh. Tumor-derived circulating endothelial cell clusters in colorectal cancer. Sci Transl Med, 2016. [DOI]
- 12.Bhakdi SC, Suriyaphol P, Thaicharoen P, Grote STK, Komoltri C, Chaiyaprasithi B, Charnkaew K. Accuracy of tumour-associated circulating endothelial cells as a screening biomarker for clinically significant prostate cancer. Cancers 2019;11(8).
- 13.Dudley AC. Tumor endothelial cells. Csh Perspect Med 2012;2(3).
- M Ramakrishnan, SR Mathur, A Mukhopadhyay. Fusion-derived epithelial cancer cells express hematopoietic markers and contribute to stem cell and migratory phenotype in ovarian carcinoma. Cancer Res, 2013. [DOI | PubMed]
- M Schmitt, FR Greten. The inflammatory pathogenesis of colorectal cancer. Nat Rev Immunol, 2021. [DOI | PubMed]
- DC Hinshaw, LA Shevde. The tumor microenvironment innately modulates cancer progression. Cancer Res, 2019. [DOI | PubMed]
- J Guinney, R Dienstmann, X Wang, A de Reynies, A Schlicker, C Soneson, L Marisa, P Roepman, G Nyamundanda, P Angelino. The consensus molecular subtypes of colorectal cancer. Nat Med, 2015. [DOI | PubMed]
- B Soldevilla, C Carretero-Puche, G Gomez-Lopez, F Al-Shahrour, MC Riesco, B Gil-Calderon, L Alvarez-Vallina, P Espinosa-Olarte, G Gomez-Esteves, B Rubio-Cuesta. The correlation between immune subtypes and consensus molecular subtypes in colorectal cancer identifies novel tumour microenvironment profiles, with prognostic and therapeutic implications. Eur J Cancer, 2019. [DOI | PubMed]
- DM Gonzalez, D Medici. Signaling mechanisms of the epithelial-mesenchymal transition. Sci Signal, 2014. [DOI | PubMed]
- PM Kasi, M Hidalgo, MD Jafari, H Yeo, L Lowenfeld, U Khan, ATH Nguyen, D Siolas, B Swed, J Hyun. Neoadjuvant botensilimab plus balstilimab response pattern in locally advanced mismatch repair proficient colorectal cancer. Oncogene, 2023. [DOI | PubMed]
- M Thibaudin, JD Fumet, B Chibaudel, J Bennouna, C Borg, J Martin-Babau, R Cohen, M Fonck, J Taieb, E Limagne. First-line durvalumab and tremelimumab with chemotherapy in RAS-mutated metastatic colorectal cancer: a phase 1b/2 trial. Nat Med, 2023. [DOI | PubMed]
- A Rizzo, M Nannini, M Novelli, A Dalia Ricci, VD Scioscio, MA Pantaleo. Dose reduction and discontinuation of standard-dose regorafenib associated with adverse drug events in cancer patients: a systematic review and meta-analysis. Ther Adv Med Oncol, 2020. [DOI | PubMed]
- A Rizzo, V Mollica, V Tateo, E Tassinari, A Marchetti, M Rosellini, R De Luca, M Santoni, F Massari. Hypertransaminasemia in cancer patients receiving immunotherapy and immune-based combinations: the MOUSEION-05 study. Cancer Immunol Immunother, 2023. [DOI | PubMed]
- 24.Kozuka M, Battaglin F, Jayachandran P, Wang J, Arai H, Soni S, Zhang W, Hirai M, Matsusaka S, Lenz HJ. Clinical significance of circulating tumor cell induced epithelial-mesenchymal transition in patients with metastatic colorectal cancer by single-cell RNA-Sequencing. Cancers (Basel). 2021;13(19).
- 25.Menyailo ME, Zainullina VR, Khozyainova AA, Tashireva LA, Zolotareva SY, Gerashchenko TS, Alifanov VV, Savelieva OE, Grigoryeva ES, Tarabanovskaya NA et al. Heterogeneity of circulating epithelial cells in breast cancer at single-cell resolution: identifying tumor and hybrid cells. Adv Biol (Weinh). 2022:e2200206.
- X Li, CY Wang. From bulk, single-cell to spatial RNA sequencing. Int J Oral Sci, 2021. [DOI | PubMed]
- M Allam, S Cai, AF Coskun. Multiplex bioimaging of single-cell spatial profiles for precision cancer diagnostics and therapeutics. NPJ Precis Oncol, 2020. [DOI | PubMed]
- CR Merritt, GT Ong, SE Church, K Barker, P Danaher, G Geiss, M Hoang, J Jung, Y Liang, J McKay-Fleisch. Multiplex digital spatial profiling of proteins and RNA in fixed tissue. Nat Biotechnol, 2020. [DOI | PubMed]
- E Gracia Villacampa, L Larsson, R Mirzazadeh, L Kvastad, A Andersson, A Mollbrink, G Kokaraki, V Monteil, N Schultz, KS Appelberg. Genome-wide spatial expression profiling in formalin-fixed tissues. Cell Genom, 2021. [DOI | PubMed]
- A Lyubetskaya, B Rabe, A Fisher, A Lewin, I Neuhaus, C Brett, T Brett, E Pereira, R Golhar, S Kebede. Assessment of spatial transcriptomics for oncology discovery. Cell Rep Methods, 2022. [DOI | PubMed]
- W Harb, A Fan, T Tran, DC Danila, D Keys, M Schwartz, C Ionescu-Zanetti. Mutational analysis of circulating tumor cells using a novel microfluidic collection device and qPCR assay. Transl Oncol, 2013. [DOI | PubMed]
- 32.Amado V, Gonzalez-Rubio S, Zamora J, Alejandre R, Espejo-Cruz ML, Linares C, Sanchez-Frias M, Garcia-Jurado G, Montero JL, Ciria R et al. Clearance of circulating tumor cells in patients with hepatocellular carcinoma undergoing surgical resection or liver transplantation. Cancers (Basel) 2021;13(10).
- 33.Bhakdi SC, Suriyaphol P, Thaicharoen P, Grote STK, Komoltri C, Chaiyaprasithi B, Charnkaew K. Accuracy of tumour-associated circulating endothelial cells as a screening biomarker for clinically significant prostate cancer. Cancers (Basel). 2019;11(8).
- 34.Tanjak P, Chaiboonchoe A, Suwatthanarak T, Acharayothin O, Thanormjit K, Chanthercrob J, Suwatthanarak T, Wannasuphaphol B, Chumchuen K, Suktitipat B et al.The KRAS-mutant consensus molecular subtype 3 reveals an immunosuppressive tumor microenvironment in colorectal cancer. Cancers (Basel). 2023;15(4).
- P Tanjak, A Thiantanawat, P Watcharasit, J Satayavivad. Genistein reduces the activation of AKT and EGFR, and the production of IL6 in cholangiocarcinoma cells involving estrogen and estrogen receptors. Int J Oncol, 2018. [PubMed]
- J Terzic, S Grivennikov, E Karin, M Karin. Inflammation and colon cancer. Gastroenterology, 2010. [DOI | PubMed]
- M Levrero, J Zucman-Rossi. Mechanisms of HBV-induced hepatocellular carcinoma. J Hepatol, 2016. [DOI | PubMed]
- DS O’Callaghan, D O’Donnell, F O’Connell, KJ O’Byrne. The role of inflammation in the pathogenesis of non-small cell lung cancer. J Thorac Oncol, 2010. [DOI | PubMed]
- M Shibutani, K Maeda, H Nagahara, T Fukuoka, S Matsutani, S Kashiwagi, H Tanaka, K Hirakawa, M Ohira. A comparison of the local immune status between the primary and metastatic tumor in colorectal cancer: a retrospective study. BMC Cancer, 2018. [DOI | PubMed]
- SM Toor, K Murshed, M Al-Dhaheri, M Khawar, M Abu Nada, E Elkord. Immune checkpoints in circulating and tumor-infiltrating CD4(+) T cell subsets in colorectal cancer patients. Front Immunol, 2019. [DOI | PubMed]
- 41.Alsalman A, Al-Mterin MA, Murshed K, Alloush F, Al-Shouli ST, Toor SM, Elkord E. Circulating and Tumor-Infiltrating Immune Checkpoint-Expressing CD8+ Treg/T Cell Subsets and Their Associations with Disease-Free Survival in Colorectal Cancer Patients. Cancers. 2022; 14(13):3194. https://doi.org/10.3390/cancers14133194
- F Mollinedo. Neutrophil degranulation, plasticity, and cancer metastasis. Trends Immunol, 2019. [DOI | PubMed]
- AM Khaliq, C Erdogan, Z Kurt, SS Turgut, MW Grunvald, T Rand, S Khare, JA Borgia, DM Hayden, SG Pappas. Refining colorectal cancer classification and clinical stratification through a single-cell atlas. Genome Biol, 2022. [DOI | PubMed]
- J Zhang, S Li, F Liu, K Yang. Role of CD68 in tumor immunity and prognosis prediction in pan-cancer. Sci Rep, 2022. [DOI | PubMed]
- A Sharma, SK Subudhi, J Blando, L Vence, J Wargo, JP Allison, A Ribas, P Sharma. Anti-CTLA-4 immunotherapy does not deplete FOXP3(+) regulatory T Cells (Tregs) in human cancers-response. Clin Cancer Res, 2019. [DOI | PubMed]
- AM Monjazeb, A Giobbie-Hurder, A Lako, EM Thrash, RC Brennick, KZ Kao, C Manuszak, RD Gentzler, A Tesfaye, SK Jabbour. A randomized trial of combined PD-L1 and CTLA-4 inhibition with targeted low-dose or hypofractionated radiation for patients with metastatic colorectal cancer. Clin Cancer Res, 2021. [DOI | PubMed]
- HO Lee, Y Hong, HE Etlioglu, YB Cho, V Pomella, B Van den Bosch, J Vanhecke, S Verbandt, H Hong, JW Min. Lineage-dependent gene expression programs influence the immune landscape of colorectal cancer. Nat Genet, 2020. [DOI | PubMed]
