Structural and functional analysis reveals the catalytic mechanism and substrate binding mode of the broad-spectrum endolysin Ply2741
Abstract
The emergence of antibiotic-resistant bacteria has attracted interest in the field of endolysins. Here, we analyzed the diversity of Streptococcus endolysins and identified a new endolysin, Ply2741, that exhibited broad-spectrum bactericidal activity. Our results demonstrated that Ply2741 could effectively eradicate multidrug-resistant gram-positive pathogens in vitro and in vivo. Structural analysis revealed that the bactericidal activity of Ply2741 depends on the classic “Cys-His-Asn” catalytic triad. Site-directed mutagenesis results further identified that the conserved residue Gln29, located near the catalytic triad, also contributes to the lytic activity of Ply2741. Furthermore, the key residues (R189 and W250) in the Ply2741 cell wall binding domain (CBD) responsible for binding to peptidoglycan were revealed by molecular docking and fluorescence-activated cell sorting (FACS) analysis. Ply2741 demonstrates a broad lytic spectrum, with significant bactericidal activity against Enterococcus, Staphylococcus, and Streptococcus and species. To the best of our knowledge, we found that residue Gln29 participated in the lytic activity of endolysin for the first time. Additionally, we systematically elucidate the binding mode and key residues of the Ply2741CBD. This study proposes Ply2741 as a potential antibiotic substitute and provides a structural basis for the modification and design of endolysins.
Article type: Research Article
Keywords: Antibiotic resistance, phage endolysins, antimicrobial agents, lytic activity, structure
Affiliations: National Key Laboratory of Agricultural Microbiology, Hubei Hongshan Laboratory, Huazhong Agricultural University, Wuhan, China; The Cooperative Innovation Centre for Sustainable Pig Production, Huazhong Agricultural University, Wuhan, China; College of Veterinary Medicine, Huazhong Agricultural University, Wuhan, China
License: © 2025 The Author(s). Published by Informa UK Limited, trading as Taylor & Francis Group. CC BY 4.0 This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. The terms on which this article has been published allow the posting of the Accepted Manuscript in a repository by the author(s) or with their consent.
Article links: DOI: 10.1080/21505594.2024.2449025 | PubMed: 39810299 | PMC: PMC11740692
Relevance: Relevant: mentioned in keywords or abstract
Full text: PDF (145 KB)
Introduction
Several species of Streptococcus, Staphylococcus, and Enterococcus are commensal microbiota that are widely present in humans [ref. 1]. However, certain species within these genera can cause serious infections, thereby posing a significant risk to public health. Group B Streptococcus (GBS) is an opportunistic pathogen that can cause neonatal pneumonia, septicemia, and meningitis [ref. 2]. Streptococcus suis, initially recognized as the predominant pathogen in swine, has gradually emerged as an important concern for global public health safety [ref. 3]. Palmieri et al. reported that S. suis has become an antibiotic-resistant reservoir [ref. 4]. Staphylococcus aureus and Enterococcus are important pathogens in community-acquired and hospital infection [ref. 5,ref. 6]. S. aureus can cause blood and respiratory infections and even septicemia [ref. 7]. The prevalence of Methicillin-resistant Staphylococcus aureus (MRSA) has increased significantly in communities and hospitals in various countries [ref. 8–10]. Enterococcus is a common cause of endocarditis and contributes to a series of infections in community environments [ref. 11]. The World Health Organization (WHO) has placed vancomycin-resistant Enterococcus (VRE) at the top of the global priority list of drug-resistant bacteria [ref. 12]. The continuing emergence of antimicrobial resistance (AMR) has been identified as a major threat to humanity by WHO [ref. 13]. With the continued emergence of multidrug-resistant bacteria and the slow progress in the development of new antibiotics, innovative therapeutics are urgently needed to combat bacterial infections.
Bacteriophages and derived enzymes are potential therapeutic agents against antibiotic-resistant bacteria [ref. 14]. However, phage therapeutics present significant challenges such as the development of bacterial resistance, immunogenicity, and horizontal gene transfer [ref. 15–17]. In contrast, endolysins, which are hydrolases produced by bacteriophages, exhibit independent antimicrobial activity, a wider lytic spectrum, and a lower likelihood of bacterial resistance development [ref. 18]. Several studies have highlighted the clinical potential of endolysins. Cpl-1 significantly eliminated Streptococcus pneumoniae from multiple serotypes, inhibited bacterial colonization in the nasopharynx, and effectively prevented acute otitis media [ref. 19,ref. 20]. Moreover, Cpl-1 could effectively rescue mice from infections caused by S. pneumoniae through systemic administration [ref. 21]. In comparison, endolysin SP-CHAP demonstrated more potent bactericidal activity against S. pneumoniae in a mouse nasopharyngeal model, demonstrating superior efficacy to Cpl-1 [ref. 22]. Several endolysins targeting multidrug-resistant Enterococcus have shown excellent therapeutic efficacy in mouse models. Cheng et al. found that LysEF-P10 not only increased the survival rate in mice infected with Enterococcus faecalis, but also restored the balance of the gut microbiota [ref. 23]. Additionally, Ply113 could significantly decrease the bacterial load in a sepsis model of mice co-infected with E. faecium and S. aureus [ref. 24]. These studies highlight the remarkable potential of endolysins in combating bacterial infections.
Endolysins hydrolyze specific sites of bacterial peptidoglycans and are generally classified into three main categories based on different catalytic mechanisms: amidases, peptidases, and glycosidases [ref. 25]. PlyC, an endolysin derived from a streptococcal bacteriophage, was composed of two separate genes, PlyCA and PlyCB. PlyCB can self-assemble into an octameric structure that is responsible for specific binding to the streptococcal cell wall. This unique structure may confer a higher lytic activity [ref. 26]. Gu et al. identified an EF-Hand-Like calcium-binding site near the groove of the active site in the catalytic domain by analyzing the crystal structure of LysGH15 [ref. 27]. The binding domain of endolysins plays a key role in bacterial cell wall attachment [ref. 28]. Zhou et al. analyzed the structure of LysIME-EF1 and revealed a unique tetrameric structure in the binding domain responsible for the translation of the truncated CBD protein via an internal ribosomal-binding site [ref. 29]. In addition, the structure of Lysostaphin SH3b demonstrated that the pentaglycine cross-bridge and peptide stem of Staphylococcus were recognized by independent sites of the SH3b domain [ref. 30]. These studies have elucidated the mechanism of endolysins from the perspective of structural biology. These unique endolysin structures confer distinct biological functions; thus, it is of significant importance to elucidate the structure and mechanism of action of endolysins.
In this study, we identified an endolysin, Ply2741, from an S. suis prophage module and evaluated its lytic activity and spectrum. The results showed that Ply2741 exhibited remarkable bactericidal activity against multidrug-resistant gram-positive bacteria. Notably, the Cys-His-Asn catalytic triad was identified as the key active region of the catalytic domain of Ply2741. Furthermore, to the best of our knowledge, the conserved residue Gln29, located near the catalytic triad, was revealed for the first time to influence the lytic activity of the endolysin. Furthermore, we elucidated the structure of the protein–substrate complex and the binding mode of Ply2741 by AlphaFold v2.0 [ref. 31], autodocking, and molecular dynamics (MD) analysis. These results demonstrate the key residues and mechanisms of the catalytic and binding domains, providing a theoretical basis for further application and modification of endolysins.
Materials and methods
Ethics statement
The animal experiments adhered to the ARRIVE guidelines. The protocol was approved by the Research Ethics Committee of Huazhong Agricultural University (HZAUMO-2023–0336 and HZAUMO-2024–0226). Five-week-old specific-pathogen-free (SPF) BALB/c mice were purchased from the Laboratory Animal Center of Huazhong Agricultural University (HZAU), and the experiments were conducted in the ABSL-2 laboratory at HZAU. During the experiments, mice were maintained under controlled conditions of constant temperature and humidity, with a 12-h light/dark cycle. At the end of the experiments or when mice showed signs of severe distress, euthanasia was performed via CO2 inhalation.
Bacteria and cell culture conditions
All the strains used in this study are listed in Tables S1 and Table S2. Streptococcus, Enterococcus, and Erysipelothrix rhusiopathiae were cultured in TSB and TSA media (BD, USA) containing 10% (v/v) fetal bovine serum (Sigma-Aldrich, USA). Salmonella, Escherichia coli, and Staphylococcus were cultured in Luria-Bertani (LB) broth and LB-agar plates (Solarbio, China). A549 (ATCC: CCL-185), Huh7, and 293T (ATCC: CRL-3216) cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) (Gibco, USA) supplemented with 10% (v/v) fetal bovine serum (Sigma-Aldrich, USA) at 37°C in a 5% CO2 atmosphere.
Phylogenetic analysis of endolysins
Phylogenetic analysis was performed on the amino acid sequences of 270 strains of the Streptococcus endolysins. The amino acid sequences of 247 endolysins were obtained from the prophage modules of Streptococcus species available in the NCBI database, while the remaining 23 prophage endolysin sequences are listed in Table S3. The classification criteria refer to those previously described by Oechslin et al. [ref. 32]. Briefly, open reading frames (ORFs) in Streptococcus prophage genomes annotated as N-acetylmuramoyl-L-alanine amidase, glycoside hydrolase, glucosaminidase, endolysin (lysin), CHAP, or lysozyme were initially identified as potential endolysins. Subsequently, the conserved domains of these potential endolysins were analyzed using SMART (http://smart.embl-heidelberg.de/). Gene fragments containing GH25, amidase, CHAP, or lysozyme in the catalytic domains were explicitly classified as endolysins [ref. 33]. A phylogenetic tree was constructed via Mega 7 using the neighbor-joining method with 1000 bootstrap replications [ref. 34]. The phylogenetic tree was visualized and modified using the Interactive Tree Of Life (https://itol.embl.de/) [ref. 35]. Additionally, the physiochemical characteristics of the endolysin were analyzed using Expasy (https://web.expasy.org/protparam/).
Plasmid construction, protein expression, and purification
The gene encoding the endolysin of Ply2741 (GenBank accession number: PQ213358) was amplified by PCR using S. suis SS2741 as the template with primers specified in Table S4. Subsequently, the fragment was cloned into the pColdTM II plasmid (Takara, Japan), and the recombinant plasmid pCold-Ply2741 was transformed into E. coli BL21 (DE3) competent cells. The recombinant strain was cultured in LB medium supplemented with 100 μg/ml ampicillin at 37°C until OD600 nm = 0.6, and induced for 18 h under conditions by adding 0.6 mm Isopropyl β-D-thiogalactoside (IPTG) at 16°C. After centrifugation at 6000 ×g for 30 min at 4°C, the induced cells were resuspended in binding buffer (20 mm Tris, 150 mm NaCl, pH 7.5) and lysed using a cell pressure crusher (ATS ENGINEERING INC. Canada). The lysed sample was centrifuged at 10,000 ×g for 30 min at 4°C to remove precipitates. The supernatant was then filtered through a 0.22 μm filter, yielding an unpurified protein. For the purification step, all variant proteins were purified using affinity chromatography and size exclusion chromatography (SEC). Briefly, the recombinant protein with 6×His-tag at the N-terminal was initially purified using His-Trap FF column (Cytiva, USA) under elution buffer conditions (20 mm Tris, 150 mm NaCl, 300 mm imidazole, pH = 7.5) and further purified using a HiLoad Superdex 200 pg column (Cytiva, USA) with SEC buffer (20 mm Tris, 150 mm NaCl, pH = 7.5) following the protocols. The eluted proteins were identified using SDS-PAGE.
Lytic spectrum and physicochemical characterization of Ply2741
The lytic spectrum of Ply2741 was determined by spot-test and turbidity reduction assay as described previously [ref. 36]. Briefly, for the spot-test assay, log-phase bacteria were mixed with TSA and TSB media, and the mixture was poured onto TSA or LB-agar plates. Subsequently, 10 μl of endolysin was spotted onto the plate, followed by overnight incubation at 37°C. For the turbidity reduction assay, different bacterial species in the log phase were resuspended in SEC buffer until the OD600 nm reached 0.8–1.2, and then added to a final concentration of 25 μg/ml and 50 μg/ml of Ply2741. After incubating at 37°C for 30 min, the absorbance at OD600 nm was measured to gauge the lytic activity by the reduction in OD600 nm.
The biological characteristics of Ply2741 were determined by a turbidity reduction assay as described previously [ref. 37]. For the optimal temperature assay, log-phase S. suis SS2741 cells were adjusted to OD600 nm of 0.8–1.2. The Ply2741 (25 μg/ml) was added and the mixture was incubated at various temperatures for 1 h (4°C, 20°C, 37°C, 42°C, 50°C, 55°C, 60°C, 70°C, and 80°C). For the optimal pH assay, the lytic activity was tested in different buffers [20 mm sodium acetate buffer (pH 3.0 to 6.0), 20 mm sodium phosphate buffer (pH 7 to 8), and 20 mm Tris-HCl buffer (pH 9.0 to 10.0)]. Finally, the absorbance at OD600 nm was measured and calculated the reduction in OD600 nm. All experiments were performed in triplicate. To explore the effect of EDTA on the lytic activity, Ply2741 (50 μg/ml) was pretreated with various concentration EDTA for 1 h. Then, Ply2741 was mixed with S. suis N15 and determined the OD600 nm for 30 min at 10 min intervals. Furthermore, the effect of Ca2+ was also determined as describe previously [ref. 27]. Briefly, after incubation with 1 mm EDTA, the Ply2741 was dialyzed to remove excess EDTA. Subsequently, CaCl2 was then added to the mixture at concentrations of 0.1 mm and 1 mm. The mixture was incubated at 37°C for 30 min before measuring the reduction in OD600 nm. All experiments were conducted in triplicate. The calculation method for the reduction in OD600 nm is as follows.ReductioninOD600nm=OD600nm Control−OD600nmPly2741
Antimicrobial susceptibility testing and kinetic time kill assay
To explore the lytic activity of Ply2741 against multi-drug resistant bacteria, the MICs of Streptococcus (S. suis N15 and Streptococcus agalactiae ATCC13813), Staphylococcus (S. aureus S8 and Staphylococcus epidermidis Z17), and Enterococcus (E. faecalis 004 and 009) were tested by broth microdilution method according to the guidelines recommended by the Clinical and Laboratory Standards Institute [ref. 38]. Ampicillin, chloramphenicol, erythromycin, tetracycline, oxacillin, and vancomycin were used to test in this assay.
To evaluate the bactericidal activity of Ply2741 against different species of bacteria in vitro, we determined the bactericidal kinetic curve and the number of bacteria after treatment with endolysin, as described previously [ref. 39]. Briefly, the log-phase bacteria were washed three times, and Ply2741 was added (50 μg/ml). After incubation at 37°C for 30 min, samples were followed by ten-fold dilution and cultured on TSA plates for colony counting. Additionally, the bactericidal kinetic curve of Ply2741 was determined. The log-phase bacteria were resuspended in SEC buffer until the OD600 nm reached 0.8 to 1.2, and then different final concentrations of Ply2741 were added. The OD600 nm was determined for 1 h at 5 min intervals. All experiments were repeated three times.
Biofilm eradication assay
The antibiofilm ability of Ply2741 against gram-positive bacteria was evaluated as described previously [ref. 40]. Briefly, log-phase bacteria were inoculated in 96-well plates and incubated at 30°C for 24 h and 48 h. The experimental groups were treated with Ply2741 at final concentrations of 50 μg/ml, followed by incubation at 37°C for 1 h and subsequent washing three times with SEC buffer. Subsequently, in some wells, biofilms were resuspended and counted on plates. For the other wells, crystal violet staining was used to evaluate the ability of Ply2741 to remove bacterial biofilms, as described previously [ref. 41]. In short, after drying at room temperature, the wells were stained by adding 200 μl crystal violet for 30 min. Finally, 33% acetic acid was added for dissolution and the absorbance at OD590 nm was measured. All experiments were performed in triplicates.
Toxicity to mammal cells and mice
To evaluate the toxicity of Ply2741, we performed experiments on cells and mice as described previously [ref. 42]. Briefly, A549, 293T, and Huh7 cells were seeded overnight at a density of 5 × 103cells/well in a 96 well plate. Subsequently, Ply2741 was added to the cells at a final concentration of 200 μg/ml, and SEC buffer served as a negative control. After incubation at 37°C for 24 h, the CellTiter 96® AQueous One Solution Cell Proliferation Assay (Promega, USA) was used to detect the absorbance of OD490 nm according to the protocol, and all experiments were repeated three times.
Mice were used to evaluate the safety of Ply2741 in vivo. Five-week-old specific-pathogen-free (SPF) BALB/c mice were randomly divided into three groups (n = 3). Subsequently, the mice were injected with 500 μg of Ply2741, with a phosphate buffered saline (PBS) buffer serving as control. The mice in the mock group did not receive any treatment. The physiological status and survival rate of mice were monitored daily. After 7 d, the tissues were collected for histopathological analysis.
Transmission electron microscopy (TEM)
To investigate the bacterial morphology after treatment with Ply2741, we observed it by TEM. Briefly, log-phase S. suis SC19 cells were washed and treated with Ply2741. Subsequently, cells were cultured at 37°C for 5 min, 15 min, and 30 min. Untreated bacteria served as controls. Subsequently, the samples were fixed with 2.5% glutaraldehyde, dehydrated in ethanol, and embedded in SPI-Pon 812 resin for cutting using UC6 ultra-microtome (Leica Microsystems, Austria). Observations were conducted using an HT7800 transmission electron microscope (Hitachi, Tokyo, Japan).
Scanning electron microscopy (SEM)
To explore the morphology of bacteria biofilms treated with Ply2741, we performed SEM experiments. Briefly, bacteria were inoculated into a 24-well plate with cell coverslips at the bottom and cultured at 30°C for 48 h. After washing with SEC buffer, Ply2741 at final concentration of 25 μg/ml was added to the experimental groups, and SEC buffer served as a control. After washing with SEC buffer to remove planktonic bacteria, 2.5% glutaraldehyde was added for 30 min. Finally, the coverslips were sprayed with gold and observed by SEM (NTC JSM-6390LV, Japan).
Mouse Streptococcus suis infection model
In a survival experiment, 5-week-old female SPF BALB/c mice were randomly divided into five groups (n = 9 per group) and each group was intraperitoneally challenged with 1 × 109 CFU of S. suis SC19. After 1-h post-infection, mice were treated via intraperitoneal injection with various dose of Ply2741 (100 μg, 200 μg, and 400 μg), ampicillin (5 mg/kg), or PBS, respectively. The survival rate of the mice was monitored for 7 d.
Additionally, the therapeutic efficacy of Ply2741 administered via various injection routes after 3 h challenge was further evaluated. Mice were randomly divided into four groups: control group (n = 9), SC19 group (n = 9), intraperitoneal injection group (i.p.) (n = 9), and intramuscular injection group (i.m.) (n = 9). Each group was inoculated with 5 × 108 CFU of SC19, while the control group received PBS buffer. At 3 h post-infection, mice were treated with 200 μg of Ply2741 via intraperitoneal or intramuscular injection. Mouse organs were collected at 6 h and 5 d post-treatment and homogenized in a PBS buffer to determine the bacterial load by ten-fold serial dilution on TSA plates. Inflammatory mediator levels (TNF-α, IL-6, IL-1β, and IFN-γ) were measured in mouse serum at 6 h post-treatment using an ELISA kit (Wuhan ColorfulGene Biological Technology Co., Ltd., China).
Structure prediction and analysis of Ply2741 domains
To explore the mechanism of action of Ply2741, the three-dimensional structures of CD and CBD were analyzed. Swiss-model was used to identify similar endolysin structures (https://swissmodel.expasy.org/) [ref. 43]. The amino acid sequences exhibiting high similarity to CD and CBD were subjected to multi-sequence alignment using ClustalW v2.1 (https://www.ebi.ac.uk/Tools/msa/clustalo/). The alignment results were modified using ESPript v3.0 (https://espript.ibcp.fr/ESPript/cgi-bin/ESPript.cgi) [ref. 44]. The three-dimensional structures of the wild-type and mutant protein were predicted by artificial intelligence using the Colab server (https://colab.research.google.com/github/sokrypton/ColabFold/blob/main/AlphaFold2.ipynb#scrollTo=11l8k–10q0C) [ref. 45], a simplified version of AlphaFold v2.0. The reliability of all prediction results was evaluated using internally predicted Local Distance Difference Test score (pLDDT). ConSurf (https://consurf.tau.ac.il/consurf_index.php) was used to analyze the conserved surface regions. The PDBsum Generate was used to generate the topology and Ramachandran plot (PDBsum Generate (ebi.ac.uk)). All protein structures were analyzed using PyMol (PyMOL Molecular Graphics System, Schrodinger LLC).
Site-directed mutagenesis and bactericidal activity of mutant endolysin
To further explore the role of the key residues, site-directed mutagenesis was conducted, and the bacterial activity of the mutant protein was determined. Briefly, the residues were modified to alanine using the Mut Express II Fast Mutagenesis Kit (Vazyme, China) according to the manufacturer’s protocol. The primers used are listed in Table S4. The expression and purification protocols for the protein mutants were identical to those of the wild-type.
Circular dichroism analysis
To analyze the secondary structure after key residue mutations in the catalytic domain, circular dichroism analysis was carried out using a JASCO J-1500 spectropolarimeter (JASCO, Japan). In brief, the wild-type and mutant proteins were adjusted to 0.5 mg/ml with SEC buffer, and 0.1 cm quartz cup was used to determine at 200–260 nm absorbance.
Nano differential scanning fluorimetry (nanoDSF)
To explore the difference in thermal stability after the mutation of key residues in the catalytic domain, nanoDSF experiments were carried out using Prometheus NT.48 (Nano Temper, Germany). Briefly, after adjusting the wild type and mutant proteins to 0.5 mg/ml using SEC buffer, Standard Capillary Chips were used to load samples (Nano Temper, Germany). The temperature was increased from 20°C to 95°C at a rate of 1°C/min.
Molecular docking and molecular dynamics analysis
Based on the molecular structure of peptidoglycan, the candidate substrate molecules, L-alanyl-D-isoglutamine (PubChem CID: 7019962), P4 (L-Ala-D-iGln-L-Lys-D-Ala), P4-2A (L-Ala-D-iGln-L-Lys-(L-Ala-L-Ala)-D-Ala), and P4-G5 (L-Ala-D-iGln-L-Lys-(Gly-Gly-Gly-Gly-Gly)-D-Ala) were identified [ref. 46]. To analyze the binding affinities and modes, we conducted molecular docking as described previously [ref. 47]. Briefly, molecular structures were obtained from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/) or drawn using ChemDraw (Cambridge Soft, USA). Subsequently, the structures were converted into PDBQT format. Hydrogen atoms were added after removing all the water molecules. The grid box covers each protein domain, with the docking pocket set as a square pocket of 30 Å × 30 Å × 30 Å and a grid spacing of 0.05 nm. Molecular docking was performed using Autodock Vina 1.2.2 (https://autodock.scripps.edu/). Discovery Studio (Dassault Group, France) was used to analyze the complex structure. Additionally, MD analysis was performed to determine the binding reliability of the substrate molecule with the highest docking score using the GROMACS 5.1.4 program package [ref. 48]. The topological structure of Ply2741CBD as generated using the Amber 99Sb-ILDN force field and TIP3P water model [ref. 49], and the ligand topological structures were prepared using the ACPYPE server [ref. 50]. For each MD analysis system, a simulation was run for 100 ns at 310 K and 1 bar atmospheric pressure. Finally, the root-mean-square fluctuation (RMSF) and root-mean-square deviation (RMSD) were analyzed using GROMACS tools.
GFP-tagged Ply2741CBD wild type and mutant plasmid construction, expression, and purification
Briefly, the Ply2741CBD fragment was amplified by PCR using S. suis SS2741 as the template and the eGFP fragment was cloned by PCR using the pCAG-GFP plasmid (Addgene: #11150) as the template. The Ply2741CBD-eGFP fragment was further amplified by overlap PCR using Ply2741CBD and eGFP fragments as templates, and then inserted into the pColdTM II expression plasmid. Site-directed mutagenesis recombinant plasmids were constructed according to the protocols described above.
Fluorescence-activated cell sorting (FACS) analysis
To explore the binding ability of Ply2741CBD after key residue mutations in the CBD, FACS experiments were performed [ref. 29]. Logarithmic-phase S. aureus S8 was adjusted to 107CFU/ml and washed thrice with SEC buffer. The bacteria were resuspended in eGFP-tagged Ply2741CBD wild-type and mutant proteins, and the SEC buffer served as a control. After the incubation at 37°C for 15 min, the samples were centrifugated at 5000 × g for 2 min and washed six times before measurement. The samples were analyzed using Cytoflex-LX according to the manufacturer’s protocol (Beckman Coulter, USA).
Statistical analysis
Data are expressed as mean ± standard deviation (SD). Data were tested for normality using the Anderson–Darling test (n > 6) and Quantile–Quantile plots (n < 6). Statistical analysis and comparison were performed using the unpaired two-tailed Student’s t-test, one- and two-way analysis of variance (ANOVA) via GraphPad Prism 8 (GraphPad Software, USA). Significant differences in survival experiment were analyzed by log-rank (Mantel-Cox) test. The number of experimental replicates is presented in the figure legends. p values < 0.05 (*) were considered statistically significant.
Results
The diversity of endolysins in Streptococcus prophages genome
To analyze the diversity of phage endolysins within the Streptococcus genus, we conducted a phylogenetic analysis of their amino acid sequences. These endolysins exhibited a broad geographical distribution (Figure 1a). Phylogenetic tree and conserved domain analysis results revealed that the catalytic domains (CDs) of endolysins could be classified into nine distinct types (Figure 1b). These CDs are composed of GH25, CHAP, amidase, and lysozyme-like muramidase in a random manner. Furthermore, the phylogenetic analysis results suggested that there was no consistent pattern in the domain composition of endolysins among different strains of Streptococcus, indicating no correlation between bacterial species and the structural composition of the evaluated endolysins. These findings highlight the extensive diversity of prophage endolysins in the Streptococcus spp. Additionally, in terms of evolutionary relationships, Ply2741 was located on a distinct branch, indicating that it is a new endolysin (Figure 1b). These findings prompted us to conduct an in-depth study of Ply2741.
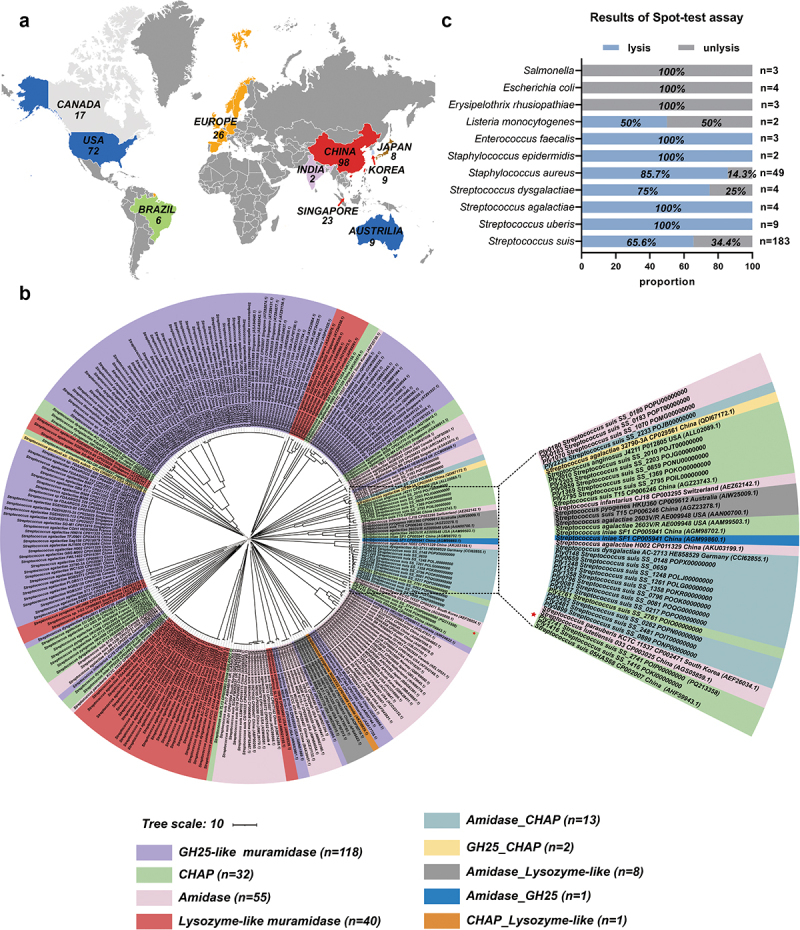
Biological characterization and lytic spectrum of Ply2741
A prophage endolysin, Ply2741, has been identified in S. suis SS2741. The results showed that the theoretical PI, instability index, and GRAVY of Ply2741 were 9.25, 26.96, and −0.344, respectively, implying that Ply2741 was soluble and stable (Table S5). Conserved domain analysis revealed that Ply2741 is composed of cysteine- and histidine-dependent amidohydrolase/peptidase (CHAP) CD and SH3b CBD (Figure S1a). After size exclusion chromatography purification (Figure S1b), the biological properties of Ply2741 were analyzed, and the activity was maintained at temperatures of 4–50°C and pH levels of 5–8 (Figure S1c and S1d). Additionally, as shown in Figure S1e, EDTA (>0.01 mm) could significantly reduce the lytic activity of Ply2741. Surprisingly, the activity of Ply2741 was fully restored after the re-addition of 0.1 mm CaCl2, indicating that the lytic activity of Ply2741 is dependent on the presence of calcium ions (Figure S1f).
The lytic spectrum of Ply2741 was assessed qualitatively using a spot-test and quantitatively using turbidity reduction assays. The spot-test results demonstrated that Ply2741 exhibited clear inhibitory halos against gram-positive bacteria, including S. suis (65.6%, n = 183), S. aureus (85.7%, n = 49), and E. faecalis (100%, n = 3) (Figure 1c and Table S1). Additionally, as depicted in Figure S1G, for Streptococcus, different serotypes of S. suis were sensitive to Ply2741, exhibiting a reduction in OD600 nm ranging from approximately 0.4 to 0.9 after exposure to Ply2741 at a concentration of 50 µg/mL for 30 min. Other streptococci, including S. agalactiae, S. pneumoniae, and Streptococcus uberis, also showed a reduction in OD600 nm to values greater than 0.7 under similar conditions. However, the activity of Ply2741 against Streptococcus dysgalactiae was comparatively lower, with a reduction in OD600 nm to less than 0.6. With Staphylococcus, Ply2741 could lysed all tested strains, resulting in a reduction in OD600 nm to 0.7. Ply2741 also exhibited lytic against E. faecalis, the reduction in OD600 nm was reached to 0.8. However, no significant effect was observed when Ply2741 was applied to gram-negative bacteria. Collectively, these findings highlight the broad lytic spectrum of Ply2741.
Ply2741 exhibited lytic activity against multidrug-resistant gram-positive bacteria in vitro
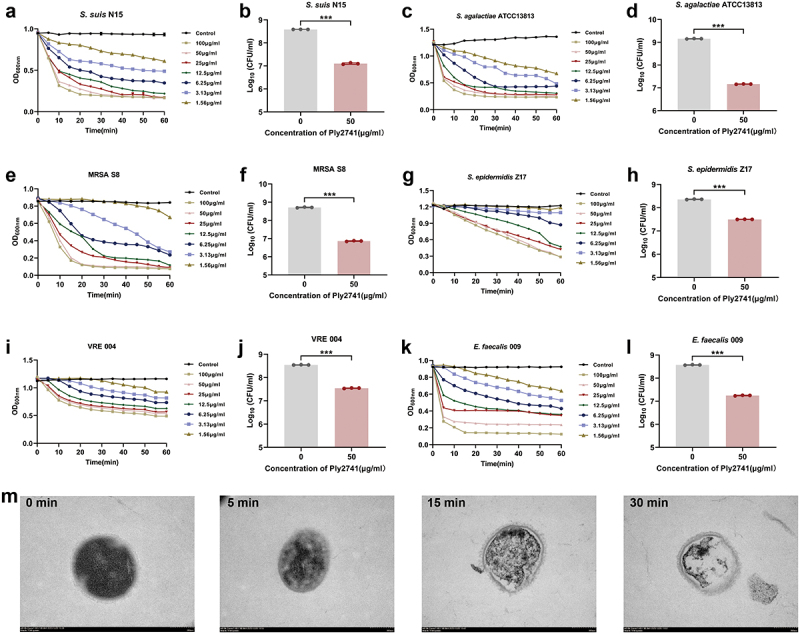
As shown in Table S6, S. suis N15, S. aureus S8 (MRSA), S. epidermidis Z17, E. faecalis 004 (VRE), and E. faecalis 009 were multidrug-resistant bacteria. To evaluate the lytic efficiency of Ply2741 against multidrug-resistant bacteria, we determined its bactericidal activity against different genera of bacteria at various concentrations and incubation times. Ply2741 rapidly reduced the OD600 nm within 15 min against Streptococcus (including multidrug-resistant S. suis N15 and S. agalactiae ATCC13813) (Figure 2a,c). Specifically, incubation with Ply2741 at a concentration of 50 μg/ml for 30 min significantly reduced the bacterial count, with S. suis N15 decreased from 8.59 log10CFU/ml to 7.10 log10CFU/ml (p < 0.001), and S. agalactiae ATCC13813 decreased from 9.15 log10CFU/ml to 7.17 log10CFU/ml (p < 0.001) (Figure 2b,d). For Staphylococcus, Ply2741 effectively eliminated MRSA S8 within 15 min, and S. epidermidis Z17 within 60 min (Figure 2e,f). Additionally, 50 μg/ml of Ply2741 caused a significant reduction in count of MRSA S8 (p < 0.001) and S. epidermidis Z17 (p < 0.001) (Figure 2f,h). Notably, Ply2741 significantly eradicated VRE 004 and E. faecalis 009 within 20 min (Figure 2i,k). Furthermore, the incubation with Ply2741 led to a significant reduction in bacterial counts: for VRE 004, the count decreased from 8.54 log10CFU/ml to 7.54 log10CFU/ml (p < 0.001), and for E. faecalis 009, the count decreased from 8.57 log10CFU/ml to 7.25 log10CFU/ml (p < 0.001) (Figure 2j,l). The morphological changes in S. suis exposed to Ply2741 were analyzed by TEM, revealing the complete release of internal contents and formation of a classic “ghost” after 30 min of incubation (Figure 2m). In conclusion, these results indicate that Ply2741 exhibited significant dose and time-dependent bactericidal activity against multidrug-resistant gram-positive bacteria.
Eradication effects of Ply2741 against biofilms
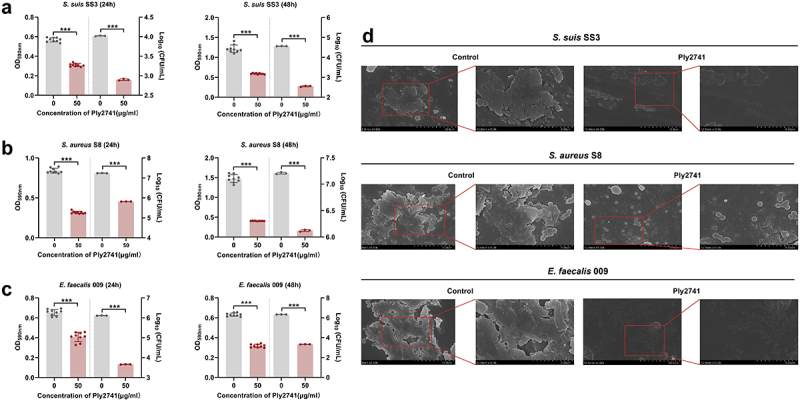
To explore the antibiofilm ability of Ply2741, bacterial counting, crystal violet staining method, and SEM experiments were carried out. Crystal violet experiments at 24 h and 48 h biofilms showed a significant reduction in absorbance at OD590 nm after Ply2741 treatment (p < 0.001) (Figure 3a–c). The bacterial count results revealed a notable decrease in viable bacteria of biofilms after Ply2741 treatment compared to the control. Specifically, for S. suis SS3, the counts decreased from 4.02 log10CFU/ml to 2.90 log10CFU/ml at 24 h and from 4.56 log10CFU/ml to 2.55 log10CFU/ml at 48 h (Figure 3a). Similarly, for S. aureus S8, counts were reduced from 7.24 log10CFU/ml to 5.82 log10CFU/ml at 24 h and from 7.21 log10CFU/ml to 6.12 log10CFU/ml at 48 h (Figure 3b). For E. faecalis 009, reductions were from 6.11 log10CFU/ml to 3.66 log10CFU/ml at 24 h and from 6.33 log10CFU/ml to 3.33 log10CFU/ml at 48 h (Figure 3c). In addition, SEM images further demonstrated that Ply2741 significantly eradicated established biofilms (Figure 3d). Collectively, these results indicate that Ply2741 exhibits a significant ability to eradicate biofilms in vitro.
Therapeutic effect of Ply2741 in a mouse model infected with S. suis
We further explored the therapeutic effects of Ply2741 against S. suis SC19 infection in vivo (Figure 4a). Prior to the experiment, the safety of Ply2741 was evaluated in both mammalian cells and mice. No significant damage was observed in either the cells (Figure S2a) or mouse tissues (Figure S2b), confirming the safety of Ply2741.
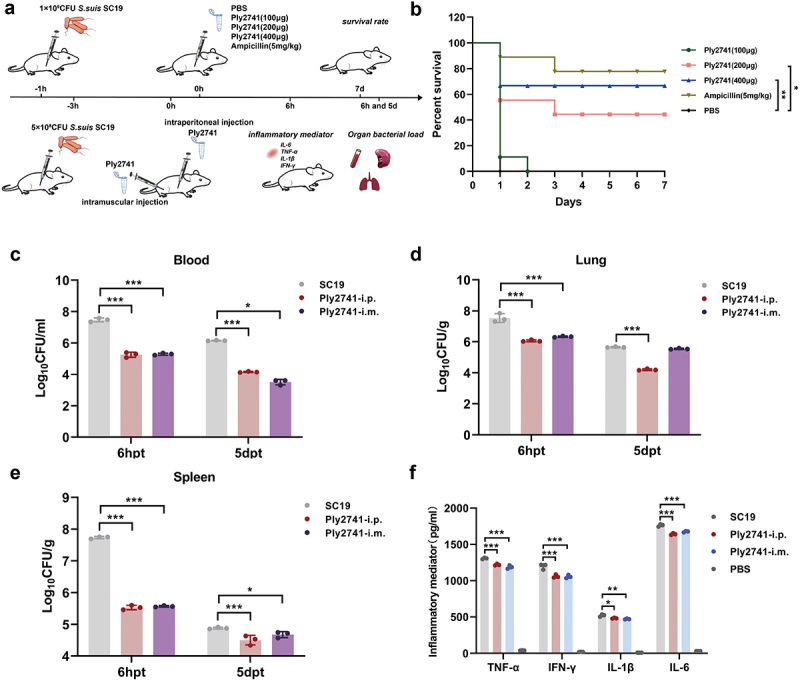
In the survival experiment, all mice challenged with 1 × 109CFU of S. suis SC19 succumbed within 12 h. However, treatment with Ply2741 significantly improved survival rates in a dose-dependent manner (Figure 4b). Ply2741 at doses of 200 μg and 400 μg provided survival rates of 44% (p < 0.05) and 67% (p < 0.01), respectively, while ampicillin exhibited a survival rate of 78% (Ply2741 (400 μg) vs. ampicillin, p > 0.05). Similarly, experiments with different treatment routes at 3 h post-challenge (5 × 108CFU of SC19) showed significant reductions in bacterial load. As shown in the Figure 4c, after 6 h and 5 d of treatment, the bacterial load in the blood of mice treated via intraperitoneal injection decreased by 2.21 log10CFU/ml and 2.17 log10CFU/ml, respectively, while intramuscular injection resulted in decreases of 2.00 log10CFU/ml and 2.66 log10CFU/ml. Comparable therapeutic effects were observed in the lungs and spleen (Figure 4d,e). Furthermore, the inflammatory mediator levels of TNF-α (i.p.: p < 0.001; i.m.: p < 0.001), IL-6 (i.p.: p < 0.001; i.m.: p < 0.001), IL-1β (i.p.: p < 0.05; i.m.: p < 0.01), and IFN-γ (i.p.: p < 0.001; i.m.: p < 0.001) were also decreased significantly (Figure 4f). In conclusion, these results demonstrate that Ply2741 exhibits a significant therapeutic effect against S. suis infection in vivo.
The catalytic activity of Ply2741 depends on the classical “Cys-His-Asn” catalytic triad
To further explore the lytic mechanism of Ply2741, we analyzed the structure of Ply2741CD using ColabFold. The optimal CD structure is shown in Figure 5a, with a pLDDT index of 91.9 (Figure S3a). The Ramachandran plot results showed that 90.2% of residues were located in the most favored regions, with 9.8% in the allowed regions (Figure S3b), indicating accurate modeling of Ply2741CD. In addition, the secondary structure and topological analysis revealed four α-helices and six β-strands in Ply2741CD, with two loop structures between α1 and α2, and between β4 and β5 (Figure 5b). These loops were positioned externally to the core region and oriented on the same side of the overall structure. Structural comparisons with PlyPy (PDB ID: 5udm) revealed similarities (RMSD = 1.534Å), with differences primarily in the flexible loop regions (Figure 5c).
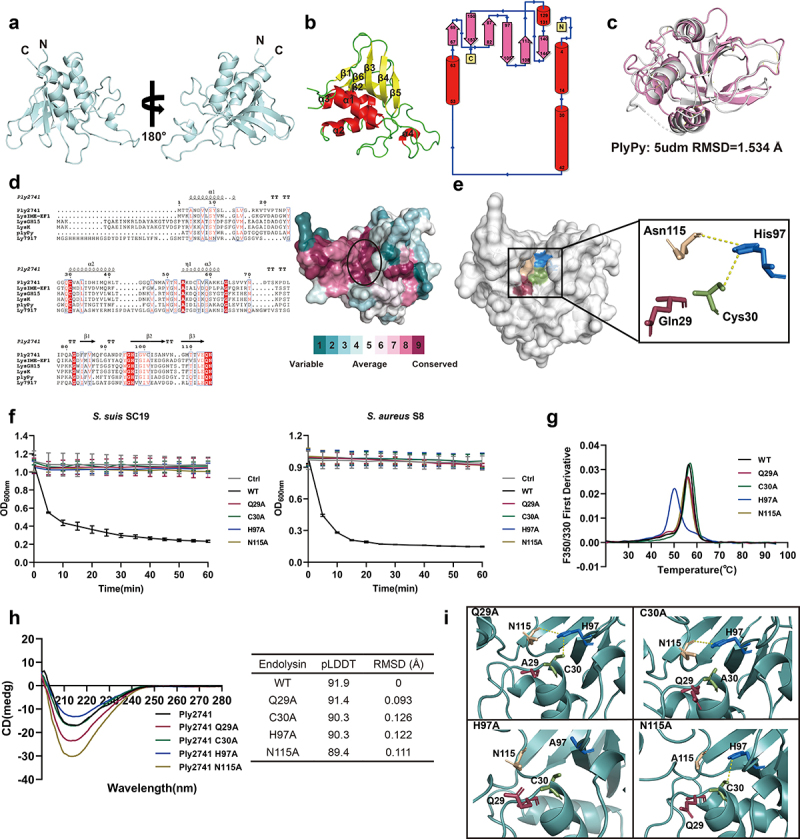
Multiple sequence alignment and structural analysis of conserved surfaces revealed a narrow and deep groove formed by conserved residues within Ply2741CD, suggesting that this region may be associated with catalytic activity (Figure 5d). Further investigation identified a classical catalytic triad consisting of Cys30, His97, and Asn115 within this conserved groove, which are likely catalytic active sites of Ply2741. Additionally, the nearby residue Gln29 is also highly conserved and contributes to the formation of the conserved groove along with the catalytic triad (Figure 5e). To validate the function of these residues, site-directed mutagenesis was performed by mutating Gln29, Cys30, His97, and Asn115 to Alanine, which was identified by SDS-PAGE (Figure S4a and S4b). The lytic activity results indicated that Ply2741 Q29A, C30A, H97A, and N115A were almost completely inactive compared to Ply2741 wild-type (Ply2741WT) (Figure 5f). These results indicate that, in addition to the catalytic triad residues, the conserved residue Gln29 near the catalytic active site also contributes to the bactericidal activity of Ply2741.
Furthermore, nanoDSF results showed that the mutation of His97 within the catalytic triad significantly reduced the thermal stability of Ply2741, decreasing the melting temperature by 6.4°C (Figure 5g). To further investigate the mechanism of active site mutations on protein stability, structural analysis of wild-type and mutant Ply2741CD proteins was performed. Circular dichroism results revealed that there were no significant differences between the wild-type and mutant proteins, with nearly identical overall curves (Figure 5h). The structures of the mutant proteins, as predicted using AlphaFold, were compared to the wild-type and showed no significant differences in overall structure, with a maximum RMSD of 0.126Å, indicating that mutations in the catalytic active sites do not affect the overall structure (Figure 5h). However, structural comparisons of the active center region revealed that the mutation of His97 disrupted hydrogen bonding with Cys30 and Asn115 (Figure 5i), suggesting that the loss of interactions among the catalytic triad residues significantly decreases the overall stability and lytic activity of the endolysin.
Binding mechanism of Ply2741CBD to substrate molecules
The highest-ranked CBD structure is presented in Figure 6a, with a pLDDT score of 97.9 (Figure S3c). The Ramachandran plot results showed that 92.9% of the amino acids were located in the most favored regions (Figure S3d), indicating exceptional accuracy in modeling the Ply2741CBD structure. Additionally, secondary structure and topological analysis showed that the CBD consisted of eight β-strands arranged in an antiparallel fashion (Figure 6b). Interestingly, Ply2741CBD showed significant similarity with Ly7917 SH3b (161-245aa) (PDB ID: 5d74) and Lysostaphin SH3b (402-493aa) (PDB ID: 6rk4) in both sequence (73.49% and 39.51% similarity) and structure (RMSD = 0.533Å and 0.607Å, respectively) (Figure 6c,d), suggesting that Ply2741CBD belongs to the classical SH3b domain and may share a similar binding mechanism.
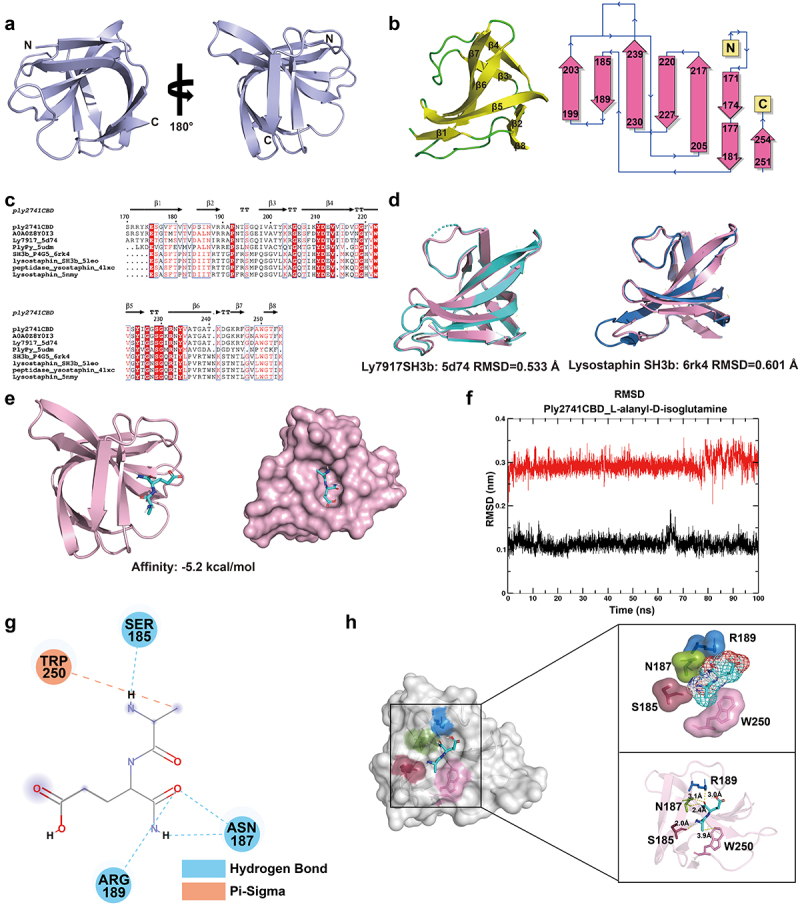
Consequently, we sought to explore the mechanism underlying CBD-peptidoglycan (PG) interactions. The candidate substrate molecules (L-alanyl-D-isoglutamine, P4, P4-2A, and P4-G5) were identified based on the structural composition of peptidoglycan (Figure S5a). Subsequently, molecular docking was performed between Ply2741CBD and the candidate substrates using AutoDock Vina. The binding affinities of Ply2741CBD docked with each substrate are shown in Table S7, and the models with the highest binding affinities are further analyzed. As shown in Figure 6e and S5b, all substrate molecules were predominantly positioned in the pocket of the CBD, suggesting that this pocket may be the main region for CBD to bind with peptidoglycan.
Given the dynamic characteristics of protein-ligand binding, we further analyzed the complexes using molecular dynamics analysis. The RMSD and RMSF of Ply2741CBD and ligands in the complexes were determined by 100 ns MD analysis via GROMACS. The RMSD trajectory serves as a valuable metric for evaluating the dynamics and stability of both the proteins and ligands in the complex. The results illustrated that in the Ply2741CBD_L-alanyl-D-isoglutamine complex structure, both the protein and ligand maintained a highly stable state. Specifically, the L-alanyl-D-isoglutamine ligands exhibited minimal fluctuations, approximately 0.05 nm, from the 75 ns to 100 ns (Figure 6f). Conversely, the stability of ligands in the Ply2741CBD_P4, Ply2741CBD_P4-2A, and Ply2741_P4-G5 structure showed a significant decrease compared to Ply2741CBD_L-alanyl-D-isoglutamine, with fluctuations observed within the range of 0.1 nm to 0.3 nm over 100 ns (Figure S6a–S6c). Additionally, the RMSF results revealed similar RMSF fluctuations in all complexes, yet each residue in the Ply2741CBD_L-alanyl-D-isoglutamine complex exhibited reduced fluctuations (Figure S6d). These findings indicate that the Ply2741CBD_L-alanyl-D-isoglutamine complex was notably more stable and deemed a more suitable substrate-binding molecule for Ply2741CBD.
The key residues in Ply2741CBD that interact with L-alanyl-D-isoglutamine
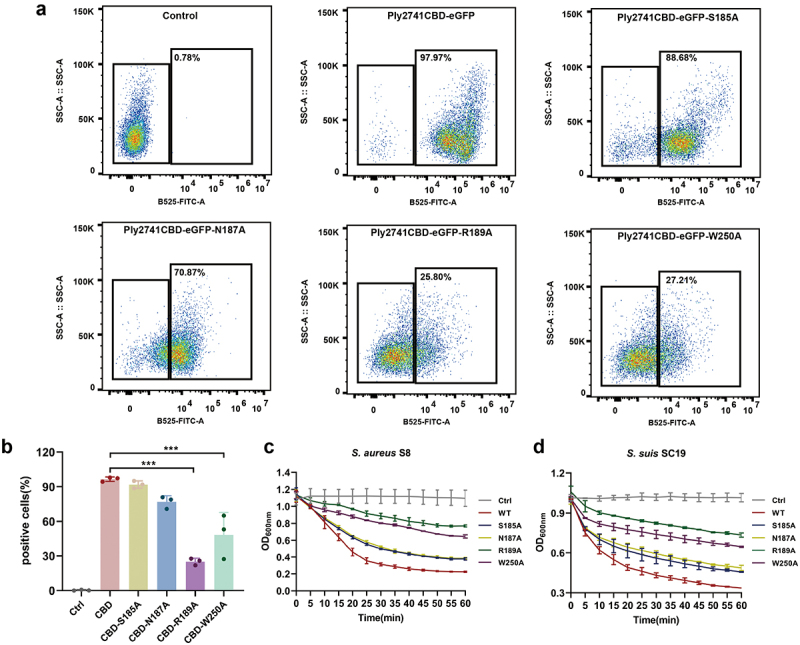
To identify the key residues involved in the interaction between Ply2741CBD and L-alanyl-D-isoglutamine, the structure of Ply2741CBD_L-alanyl-D-isoglutamine complex was analyzed using Discovery Studio. The results revealed that L-alanyl-D-isoglutamine formed hydrogen bond interactions and π–sigma interactions with key residues (Figure 6g). Specifically, the residues Ser185, Asn187, and Arg189 exhibited hydrogen-bonding interactions, whereas Trp250 displayed π–sigma interactions with the substrate, all within a distance of less than 4 Å (Figure 6h). These interactions are crucial for maintaining complex stability and function. To determine the function of the key residues, the eGFP fragment was fused to Ply2741CBD, and site-directed mutagenesis was performed (Figure S4c-S4e). After purifying wild-type Ply2741CBD-eGFP and mutant proteins, the binding activity of CBD to S. aureus S8 was evaluated by fluorescence-activated cell sorting (FACS). Mutations S185A and N187A exhibited a slight impact on the binding ability, whereas R189A and W250A significantly diminished the binding ability of CBD (Figure 7a,b). In addition, the CBD mutation was transferred to a wild-type Ply2741, and the activity of mature protein after binding residue mutation was assessed. The results demonstrated a significant decrease in lytic activity against S. suis SC19 and S. aureus S8 for mutants R189A and W250A compared to the wild-type (Figure 7c,d), which may be attributed to the loss of binding ability. However, the activity of the variant protein was not completely lost, even though R189A and W250A led to the most important efforts. These results were consistent with those of the FACS experiment, indicating that Arg189 and Trp250 play a key role in Ply2741 binding to the bacterial cell wall, but no single mutation completely abolished lytic activity.
Discussion
Phage endolysins, which are emerging as novel antimicrobial agents, have attracted considerable attention in the scientific community [ref. 51]. The modular domain composition of the endolysins can vary [ref. 52]. In this study, phylogenetic analysis was conducted on Streptococcus prophage-derived endolysins, revealing a correlation between evolutionary relationships and structural domain composition. The CDs were recombined and fused to produce nine types of endolysins, indicating a wide range of compositional diversity. Furthermore, our findings suggest that the structural domains of endolysins are not strongly associated with specific bacterial genera. The diversity of endolysins may arise from gene exchanges during bacterial-phage interactions, resulting in distinct endolysin domains in the bacterial genome. Similar results were reported in a study by Oechslin et al. on endolysins derived from Lactobacillus [ref. 32].
Phylogenetic tree analysis showed that the endolysin Ply2741 was located on a separate branch and exhibited inhibition of halos against Streptococcus, Staphylococcus, and E. faecalis. Notably, bactericidal kinetic curves revealed a rapid and dose-dependent reduction in bacterial turbidity, underscoring its broad-spectrum and efficient lytic activity. Endolysins Ly7917 [ref. 53], ply1228 [ref. 54], and ply5218 [ref. 55], derived from S. suis prophage, exhibited lytic activity against Streptococcus but showed no lytic effects on Staphylococcus and Enterococcus. PlySs2 exhibits an exceptionally broad spectrum of bactericidal activity, making it the most versatile lysin reported to date [ref. 56]. In contrast, Ply2741 shows no significant bactericidal activity against Listeria. Furthermore, Ply2741 displays better bactericidal activity against E. faecalis and S. pneumoniae, while PlySs2 demonstrates less efficacy against E. faecalis compared to S. aureus and Streptococcus [ref. 57,ref. 58]. These results suggest potential therapeutic benefits in treating infections caused by VRE. EDTA usually plays the role of chelating metal ions and is able to affect the lytic activity of endolysins with CHAP domain [ref. 27]. PlySs2 showed a significant decrease in lytic activity after the addition of 4 μM EDTA [ref. 56], while Ply2741 was able to tolerate 10 μM EDTA. This result indicates that differences in CHAP domains result in varying metal ion-binding capacities between Ply2741 and PlySs2, which in turn affecting the dependence of endolysins on metal ions. Although the lytic spectrum of Ply2741 is not as extensive as PlySs2, it possesses a rare lytic profile among the other lysins. Consequently, the characterization of Ply2741 enriches the endolysin database and provides additional options comparable to PlySs2 for combating multidrug-resistant infections.
Treatment with Ply2741 significantly improved mouse survival rates in a dose-dependent manner, with the 400 μg dose showing no significant difference in protective efficacy compared to ampicillin. However, although the in vivo therapeutic effect of Ply2741 was not as strong as that of AVPL and Ply1228 [ref. 54,ref. 59], its broad-spectrum bactericidal activity against various bacterial species offers greater potential for further modification. The chimeric lysin ClyQ, compared to the parental lysin LysGH15, not only exhibited significantly enhanced bactericidal activity but also expanded its lytic spectrum [ref. 39]. Additionally, the combination of daptomycin and lysin Cpl-1 demonstrated higher protective efficacy against S. pneumoniae infection compared to each agent used individually [ref. 60]. Therefore, the development of chimeric lysins and the synergistic application of Ply2741 with antibiotics are worth investigation in future studies. Furthermore, treatment with Ply2741 via various routes of administration can significantly reduce bacterial loads and inflammatory cytokines in organs and provide substantial protection. This therapeutic route provides additional options for the clinical application of endolysins.
Considering the lytic activity of Ply2741, investigation of its catalytic mechanism is necessary. Sequence and structural analyses reveal a classic “Cys-His-Asn” catalytic triad within the CHAP domain of Ply2741. This catalytic triad structure is a common active site in various metal-independent enzymes, including cysteine proteinases papain [ref. 61]. Cysteine proteinases typically have a conserved structural core consisting of an α-helix followed by antiparallel β-strands, with the active site located between the helix [ref. 62]. In this structure, the classic catalytic center comprises cysteine and histidine. The thiol group of cysteine is positioned at the N-terminus of the core α-helix and acts as a nucleophile during the catalytic process, while histidine functions as a base to form a reactive thiolate-imidazolium ion pair [ref. 63]. The interaction between the thiolate anion of cysteine and the imidazolium ion is crucial for the catalytic function [ref. 64]. Additionally, the amide oxygen of asparagine forms a hydrogen bond with histidine, allowing the imidazole ring to rotate without disrupting the hydrogen bond, thereby stabilizing the entire catalytic system [ref. 65]. In light of this, we propose that the catalytic triad in Ply2741 has a similar mechanism. Cys30 serves as a nucleophile attacking the carbonyl carbon of the bacterial peptidoglycan amide bond, forming a covalent acyl-enzyme intermediate. His97 can form an imidazolium ion pair to enhance the cysteine nucleophilic attacking ability. Asn115 stabilizes the system by forming hydrogen bonds with His97. The mutation of His97 results in the loss of hydrogen bonding interactions with cysteine and asparagine, which explains the higher reduction in thermal stability observed in the H97A mutant [ref. 66].
Interestingly, a conserved amino acid residue, Gln29, located near the catalytic triad, also affects the bactericidal activity of Ply2741. To our knowledge, this is the first report of a conserved residue near the catalytic center being related to lytic activity of endolysin. Although Gu et al. identified a conserved E134 residue near the catalytic center in LysGH15 and defined it along with “Cys-His-Glu-Asn” as a catalytic quartet, the lytic activity did not completely disappear after E134 mutation [ref. 27]. Multiple sequence alignments reveal that a conserved Gln residue is present in nearly all CHAP endolysins, but its function has not been reported previously. Glutamine in PlyPy and PlySs2, which are highly similar in structure and sequence to Ply2741CHAP, may have similar functions which provide a theoretical basis for further engineering of endolysins. However, the existing evidence does not elucidate the specific role of Gln29 in catalytic activity of Ply2741. We speculate that Gln29 may play a role in stabilizing the catalytic triad. Furthermore, although some studies suggest that Gln residues in certain cysteine proteases might stabilize tetrahedral by forming the oxyanion hole during catalysis [ref. 67], the position of Gln29 in Ply2741 is not consistent with cysteine proteases, indicating the need for further research to verify its potential role in the catalytic mechanism.
The Ply2741CHAP shares a sequence similarity with PlyPy, which demonstrates activity only against Streptococcus [ref. 68]. Therefore, the extensive-binding capabilities of Ply2741CBD are of interest. Previous studies have confirmed that endolysins exert bactericidal effects by destroying peptidoglycans [ref. 68,ref. 69]. Additionally, Eugster et al. found that the CBDs of Ply118, Ply511, and PlyP40 specifically interact with the peptidoglycan backbone of Listeria monocytogenes [ref. 70]. Gonzalez-Delgado et al. further elucidated the structural basis of the binding of Lysostaphin SH3b to Staphylococcus peptidoglycan using NMR [ref. 30]. However, it is noteworthy that other studies suggest that SH3b domains may interact with the bacterial wall teichoic acid (WTA) rather than peptidoglycan. Shen et al. discovered that the Listeria endolysin CBD500 specifically recognizes WTA on the bacterial surface. Structural analysis of CBD500 revealed that the binding cavity formed by β-barrel and pseudo-symmetric SH3b-like repeats serves as the structural basis for WTA recognition, a feature that is absent in Ply2741SH3b [ref. 71]. Conversely, Ply2741SH3b shares a high structural similarity with Lysostaphin SH3b, which binds to the peptidoglycan. Therefore, we explored the mechanism of the Ply2741CBD-PG interaction. However, the potential interaction between Ply2741SH3b and WTA, and its underlying mechanism, remain worthy of further investigation.
Ply2741 exhibits broad-spectrum bactericidal activity and shares sequence and structural similarity with the Lysostaphin SH3b. Although the streptococcal endolysin PlySs2, which also has broad-spectrum activity, does not bind to Staphylococcus, this could be due to limitations in detection sensitivity and resolution [ref. 58]. FACS experiments have demonstrated that Ply2741CBD can significantly bind to S. aureus. Given the composition of Streptococcus and Staphylococcus peptidoglycan, L-alanyl-D-isoglutamine, P4, P4-2A, and P4-G5 of peptidoglycan were selected as candidate substrate molecules. Molecular docking results revealed that all candidate molecules were located in the same pocket region, implying that this region may act as the key binding region of Ply2741. MD analysis revealed an extreme instability in the complex of CBD and substrate P4-2A and P4-G5. Notably, the binding pocket of CBD predominantly interacted with the P4 segment of the P4-2A and P4-G5 molecule. In addition, the substrate molecule P4 contains L-alanyl-D-isoglutamine and compared with L-alanyl-D-isoglutamine, substrate P4 also exhibited instability during the process. Ji et al. determined the crystal structure of the Ly7917 SH3b-substrate complex using X-ray diffraction (PDB id: 5d76). Their findings revealed that Ly7917 SH3b specifically binds to L-alanyl-D-isoglutamine through the residues A176 and R180, which constitute key substrate-binding sites [ref. 30,ref. 72]. Considering their structural similarity, L-alanyl-D-isoglutamine was identified as the substrate of Ply2741CBD. However, it is challenging to determine the binding affinity of CBD with L-alanyl-D-isoglutamine through experiments, likely because of the very weak binding affinity of CBD to the substrate, as found in previous studies [ref. 27,ref. 73,ref. 74]. Analysis of the interaction between L-alanyl-D-isoglutamine and Ply2741CBD revealed the intermolecular interactions between L-alanyl-D-isoglutamine and the residues Ser185, Asn187, Arg189, and Trp250 around the binding pocket. Except for the π–sigma interaction with W250, CBD interacted with the other three amino acids via hydrogen bond interactions. FACS and turbidity reduction assay results showed that the mutation of R189 and W250 significantly affected protein-binding activity. Interestingly, the binding ability of the S185 and N187 mutations and their impact on mature protein activity were not significant. Although mutations at R189 and W250 significantly reduced protein activity, they were not completely abolished. This phenomenon is attributed to the fact that the mutation of a single binding site does not completely lose the binding ability, and the additional-binding site may be compensated, aligning with the findings of Gonzalez-Delgado et al. in the study of lysostaphin SH3b [ref. 30].
In summary, we identified an endolysin, Ply2741, which demonstrated significant broad-spectrum bactericidal efficacy against multidrug-resistant gram-positive bacteria in vitro and in vivo. Through structural analysis of Ply2741 catalytic and cell wall binding domains, we elucidated the mechanism of the “Cys-His-Asn” catalytic triad in the catalytic domain of Ply2741 and identified that the conserved residue Gln29 also affects the lytic activity. Additionally, we systematically characterized the key residues and interaction modes of the Ply2741 CBD involved in binding to bacterial peptidoglycan.
Supplementary Materials
References
- The pathogenesis of streptococcal infections: from tooth decay to meningitis.. Nat Rev Microbiol., 2003. [DOI | PubMed]
- Group B: global incidence and vaccine development.. Nat Rev Microbiol., 2006. [DOI | PubMed]
- Understanding the virulence of Streptococcus suis: a veterinary, medical, and economic challenge.. Med Mal Infect., 2018. [DOI | PubMed]
- Streptococcus suis, an emerging drug-resistant animal and human pathogen.. Front Microbiol., 2011. [DOI]
- Waves of resistance: in the antibiotic era.. Nat Rev Microbiol., 2009. [DOI | PubMed]
- Identification and molecular modification of Staphylococcus aureus bacteriophage lysin LysDZ25.. ACS Infect Dis., 2023. [DOI | PubMed]
- The many faces of Staphylococcus aureus infection.. Postgrad Med., 2001. [DOI]
- Staphylococcus aureus bloodstream infections: risk factors, outcomes, and the influence of methicillin resistance in Calgary, Canada, 2000–2006.. J Infect Dis., 2008. [DOI | PubMed]
- Invasive methicillin-resistant Staphylococcus aureus infections in the United States.. JAMA-J Am Med Assoc., 2007. [DOI]
- Three-year surveillance of community-acquired infections in children.. Clin Infect Dis., 2005. [DOI | PubMed]
- The rise of the enterococcus: beyond vancomycin resistance.. Nat Rev Microbiol., 2012. [DOI | PubMed]
- Ring-fused 2-pyridones effective against multidrug-resistant Gram-positive pathogens and synergistic with standard-of-care antibiotics.. Proc Natl Acad Sci USA., 2022. [DOI | PubMed]
- Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis.. Lancet., 2022. [DOI | PubMed]
- Bacteriophage endolysins: a novel anti-infective to control gram-positive pathogens.. Int J Med Microbiol., 2010. [DOI | PubMed]
- Fitness trade-offs in phage cocktail-resistant Salmonella enterica serovar enteritidis results in increased antibiotic susceptibility and reduced virulence.. Microbiol Spectr., 2022. [DOI | PubMed]
- Alternative therapeutic strategies to treat antibiotic-resistant pathogens.. Nat Rev Microbiol., 2023. [DOI | PubMed]
- Endolysins as antimicrobials.. Adv Virus Res, Vol 83: Bacteriophages, Pt B., 2012
- Phage endolysin LysP108 showed promising antibacterial potential against methicillin-resistant staphylococcus aureus.. Front Cell Infect Microbiol., 2021. [DOI]
- Delivery of the endolysin Cpl-1 by inhalation rescues mice with fatal pneumococcal pneumonia.. J Antimicrob Chemoth., 2013. [DOI]
- Novel strategy to prevent otitis media caused by colonizing streptococcus pneumoniae.. PLOS Pathog., 2007. [DOI | PubMed]
- Systemic use of the endolysin Cpl-1 rescues mice with fatal pneumococcal pneumonia.. Crit Care Med., 2009. [DOI | PubMed]
- SP-CHAP, an endolysin with enhanced activity against biofilm pneumococci and nasopharyngeal colonization.. MBio., 2024. [DOI]
- Endolysin LysEF-P10 shows potential as an alternative treatment strategy for multidrug-resistant Enterococcus faecalis infections.. Sci Rep., 2017. [DOI]
- Bacteriophage endolysin Ply113 as a potent antibacterial agent against polymicrobial biofilms formed by enterococci and staphylococcus aureus.. Front Microbiol., 2023. [DOI]
- Cell wall hydrolases in bacteria: insight on the diversity of cell wall amidases, glycosidases and peptidases toward peptidoglycan.. Front Microbiol., 2019. [DOI]
- PlyC: a multimeric bacteriophage lysin.. Proc Natl Acad Sci USA., 2006. [DOI | PubMed]
- Structural and biochemical characterization reveals LysGH15 as an unprecedented “ef-hand-like” calcium-binding phage lysin.. PLOS Pathog., 2014. [DOI | PubMed]
- Optimization and validation of a simple method using P22: bacteriophage for rapid detection of serotypes A, B, and D in poultry samples.. J Food Protect., 2008. [DOI]
- Structural and functional insights into a novel two-component endolysin encoded by a single gene in Enterococcus faecalis phage.. PLOS Pathog., 2020. [DOI | PubMed]
- Two-site recognition of Staphylococcus aureus peptidoglycan by lysostaphin SH3b.. Nat Chem Biol., 2020. [DOI | PubMed]
- Highly accurate protein structure prediction with AlphaFold.. Nature., 2021. [DOI | PubMed]
- Phage endolysins are adapted to specific hosts and are evolutionarily dynamic.. PLOS Biol., 2022. [DOI | PubMed]
- SMART: recent updates, new developments and status in 2020.. Nucleic Acids Res., 2021. [DOI | PubMed]
- MEGA X: molecular evolutionary genetics analysis across computing platforms.. Mol Biol Evol., 2018. [DOI | PubMed]
- Interactive tree of life (iTOL) v5: an online tool for phylogenetic tree display and annotation.. Nucleic Acids Res., 2021. [DOI | PubMed]
- Novel chimeric lysin with high-level antimicrobial activity against methicillin-resistant Staphylococcus aureus in vitro and in vivo.. Antimicrob Agents Chemother., 2014. [DOI | PubMed]
- Streptococcus suis prophage lysin as a new strategy for combating streptococci-induced mastitis and Streptococcus suis infection.. J Antimicrob Chemother., 2023. [DOI | PubMed]
- Performance standards for antimicrobial susceptibility testing.. CLSI supplement M100., 2021
- Exploiting broad-spectrum chimeric lysin to cooperate with Mupirocin against staphylococcus aureus-induced skin infections and delay the development of Mupirocin resistance.. Microbiol Spectr., 2023. [DOI]
- Antibiofilm potential of purified environmental bacteriophage preparations against early stage Pseudomonas aeruginosa biofilms.. J Appl Microbiol., 2019. [DOI | PubMed]
- Microtiter dish biofilm formation assay.. J Vis Exp., 2011. [DOI]
- A novel tail-associated O91-specific polysaccharide depolymerase from a podophage reveals lytic efficacy of Shiga toxin-producing Escherichia coli.. Appl Environ Microbiol., 2020. [DOI]
- SWISS-MODEL: homology modelling of protein structures and complexes.. Nucleic Acids Res., 2018. [DOI | PubMed]
- Deciphering key features in protein structures with the new ENDscript server.. Nucleic Acids Res., 2014. [DOI | PubMed]
- ColabFold: making protein folding accessible to all.. Nat Methods., 2022. [DOI | PubMed]
- Peptidoglycan structure and architecture.. FEMS Microbiol Rev., 2008. [DOI | PubMed]
- ReverseDock: a web server for blind docking of a single ligand to multiple protein targets using AutoDock Vina.. Front Mol Biosci., 2023. [DOI]
- GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation.. J Chem Theory Comput., 2008. [DOI | PubMed]
- Improved side-chain torsion potentials for the Amber ff99SB protein force field.. Proteins., 2010. [DOI | PubMed]
- The ACPYPE web server for small-molecule MD topology generation.. Bioinformatics., 2023. [DOI]
- Therapeutic potential of bacteriophage endolysins for infections caused by Gram-positive bacteria.. J Biomed Sci., 2023. [DOI]
- Synthetic biology of modular endolysins.. Biotechnol Adv., 2018. [DOI | PubMed]
- A novel endolysin disrupts Streptococcus suis with high efficiency.. FEMS Microbiol Lett., 2015. [DOI | PubMed]
- A novel lysin Ply1228 provides efficient protection against Streptococcus suis type 2 infection in a murine bacteremia model.. Vet Microbiol., 2022. [DOI | PubMed]
- A novel prophage lysin Ply5218 with extended lytic activity and stability against Streptococcus suis infection.. FEMS Microbiol Lett., 2016. [DOI | PubMed]
- Novel bacteriophage lysin with broad lytic activity protects against mixed infection by Streptococcus pyogenes and methicillin-resistant Staphylococcus aureus.. Antimicrob Agents Chemother., 2013. [DOI | PubMed]
- Characterization of the bacteriophage-derived endolysins PlySs2 and PlySs9 with in vitro lytic activity against bovine mastitis streptococcus uberis.. Antibiotics., 2020. [DOI | PubMed]
- Molecular dissection of phage lysin PlySs2: integrity of the catalytic and cell wall binding domains is essential for its broad lytic activity.. Virol Sin., 2015. [DOI | PubMed]
- Aerococcus viridans phage lysin AVPL had lytic activity against streptococcus suis in a mouse bacteremia model.. Int J Mol Sci., 2023. [DOI | PubMed]
- Bactericidal synergism between daptomycin and the phage lysin Cpl-1 in a mouse model of pneumococcal bacteraemia.. Int J Antimicrob Agents., 2013. [DOI | PubMed]
- Understanding nature’s catalytic toolkit.. Trends Biochem Sci., 2005. [DOI | PubMed]
- Identification and classification of papain-like cysteine proteinases.. J Biol Chem., 2023. [DOI | PubMed]
- Modes of inhibition of cysteine proteases.. Acta Biochim Pol., 2004. [DOI | PubMed]
- Catalytic mechanism in papain family of cysteine peptidases.. Method Enzymol., 1994
- Structural and functional roles of Asparagine-175 in the cysteine protease papain.. J Biol Chem., 1995. [DOI | PubMed]
- Structure-based design approach to rational site-directed mutagenesis of β-lactoglobulin.. J Struct Biol., 2020. [DOI | PubMed]
- Contribution of the glutamine-19 side-chain to transition-state stabilization in the oxyanion hole of papain.. Biochemistry., 1991. [DOI | PubMed]
- A highly active and negatively charged Streptococcus pyogenes lysin with a rare D-Alanyl-L-Alanine endopeptidase activity protects mice against streptococcal bacteremia.. Antimicrob Agents Chemother., 2014. [DOI | PubMed]
- Use of 4-sulfophenyl isothiocyanate labeling and mass spectrometry to determine the site of action of the streptococcolytic peptidoglycan hydrolase Zoocin A.. Appl Environ Microbiol., 2009. [DOI | PubMed]
- Wall teichoic acids restrict access of bacteriophage endolysin Ply118, Ply511, and PlyP40 cell wall binding domains to the listeria monocytogenes Peptidoglycan.. J Bacteriol., 2012. [DOI | PubMed]
- Structural basis for recognition of bacterial cell wall teichoic acid by pseudo-symmetric SH3b-like repeats of a viral peptidoglycan hydrolase.. Chem Sci., 2021. [DOI]
- [72]Wenhui J. Structural and functional analysis of endolysin Ly7917 from bacteriophage of Streptococcus suis [PhD dissertation]. Shanghai (China): Shanghai Jiao Tong University; 2015. Chinese.
- Structural and functional insights into lysostaphin–substrate interaction.. Front Mol Biosci., 2018. [DOI]
- Role of SH3b binding domain in a natural deletion mutant of endolysin LysF1 with a broad range of lytic activity.. Virus Genes., 2018. [DOI | PubMed]
