Imaging plant metabolism in situ
Abstract
Mass spectrometry imaging (MSI) has emerged as an invaluable analytical technique for investigating the spatial distribution of molecules within biological systems. In the realm of plant science, MSI is increasingly employed to explore metabolic processes across a wide array of plant tissues, including those in leaves, fruits, stems, roots, and seeds, spanning various plant systems such as model species, staple and energy crops, and medicinal plants. By generating spatial maps of metabolites, MSI has elucidated the distribution patterns of diverse metabolites and phytochemicals, encompassing lipids, carbohydrates, amino acids, organic acids, phenolics, terpenes, alkaloids, vitamins, pigments, and others, thereby providing insights into their metabolic pathways and functional roles. In this review, we present recent MSI studies that demonstrate the advances made in visualizing the plant spatial metabolome. Moreover, we emphasize the technical progress that enhances the identification and interpretation of spatial metabolite maps. Within a mere decade since the inception of plant MSI studies, this robust technology is poised to continue as a vital tool for tackling complex challenges in plant metabolism.
Article type: Review Article
Keywords: Biochemistry, chemical imaging, desorption ionization mass spectrometry (DESI), mass spectrometry imaging (MSI), matrix-assisted laser desorption/ionization (MALDI), metabolism, metabolome, primary metabolism, spatial maps, specialized metabolism
Affiliations: BioDiscovery Institute and Department of Biological Sciences, University of North Texas, Denton TX 76203, USA
License: © The Author(s) 2023. Published by Oxford University Press on behalf of the Society for Experimental Biology. CC BY 4.0 This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted reuse, distribution, and reproduction in any medium, provided the original work is properly cited.
Article links: DOI: 10.1093/jxb/erad423 | PubMed: 37889862 | PMC: PMC10938046
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (283 KB)
IntroductionMass spectrometry imaging: a brief overview
The evaluation of a plant metabolome usually begins with the extraction of the mixtures of molecules from some tissue(s) of a plant in an appropriate solvent compatible with further separation and/or analysis by mass spectrometry (MS) (ref. Mushtaq ). The extraction process is often optimized for the target class or category of metabolites to be studied, and invariably mixes the metabolites uniformly in a solvent extract as a starting point for metabolomics analysis. Although this approach facilitates the comprehensive qualitative and quantitative analysis of the metabolome (ref. Alseekh ; ref. Aharoni ), it also completely disrupts the spatial location of those metabolites in the original tissue sample. In contrast, mass spectrometry imaging (MSI) provides an approach for the analysis of metabolite compositions in situ, directly in tissue sections or on plant parts (ref. Boughton ; ref. Sturtevant ). This general MSI approach has been adapted for a variety of applications in plant metabolism studies, using various ionization sources coupled to high-resolution mass spectrometers, as detailed in other recent reviews (ref. Hu ; ref. Unsihuay ; ref. Dong and Aharoni, 2022; ref. Yin ; ref. Borisjuk ).
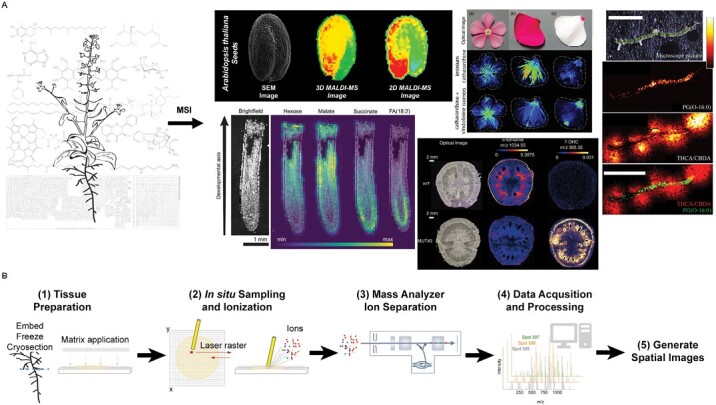
One of the most widely employed MSI techniques involves matrix-assisted laser desorption/ionization (MALDI), and can be accomplished under vacuum or at atmospheric pressure depending upon the specific capabilities of the ionization source (ref. Liu ; ref. Zhu ). Several high-quality reviews have detailed critical specimen preparation and analysis (ref. Dong ; ref. Bokhart ; ref. Buchberger ; ref. Sturtevant ; ref. Caleb Bagley ), with a general workflow for MALDI-MSI shown in Fig. 1. Briefly, MALDI-MSI can be performed on essentially any plant tissue sections. Most often samples are flash-frozen in an embedding medium (e.g. gelatin), sectioned on a cryo-microtome (e.g. 10–50 μm in thickness), and then lyophilized to preserve the subcellular organization of all small molecules. A compatible chemical matrix [e.g. 2,5-dihydroxybenzoic acid (DHB)] is then applied to the surface of the tissue section enabling metabolite ionization, with high-resolution mass spectra collected at each location on a tissue section. The final tissue distributions of all selected metabolites can be reconstructed in two dimensions with commercially available or open-source software. The spatial resolution is dependent upon laser spot and raster step size, with 1–10 μm spatial resolution achievable with lasers on most contemporary commercial instruments (ref. Hansen and Lee, 2018). However, most studies in plants routinely use spatial resolutions of 20–50 μm (i.e. around cellular resolution) that balance sampling time and metabolite signal. Chemical mass-to-charge (m/z) resolution varies among high-resolution mass spectrometers, but resolutions of >1 000 000 for lipid samples are possible with Fourier-transformed ion cyclotron resonance (FTICR) MS (ref. Tiquet ), making it feasible for experimental plans to involve isotope labeling of hundreds of isotopomer species.
While MALDI is the most used ionization method, desorption electrospray ionization (DESI)-MSI has been increasingly used in plant imaging studies. DESI does not use a chemical matrix but instead relies on a charged solvent for ionization at ambient conditions, enabling the analysis of additional plant metabolites (ref. Ferreira ; ref. Xue ; ref. He ). Although DESI can be applied directly to a plant’s surface, thereby minimizing sample pre-treatment, most plant DESI studies still rely on sample embedding, freezing, and cryo-sectioning. Imaging plant metabolites can be challenging due to tissue fragility and high water content, as well as tissues with thick epidermal layers that cannot easily be penetrated by soft ionization techniques. Tissue imprinting offers one approach for addressing these issues while maintaining spatial integrity. While the absence of a chemical matrix reduces errors in metabolite spatial assignment, DESI typically operates at much lower spatial resolutions (50–250 μm) and lower ion sensitivity relative to MALDI. As will be illustrated below, while most recent plant MSI studies have used MALDI instead of DESI, both techniques have contributed to unique insights in plant metabolite imaging.
Expanding the spatial metabolome coverage
In the past decade, an increasing number of metabolites have been spatially mapped in situ (Table 1 with additional details in Supplementary Table S1). Early applications of MSI targeted readily ionizable molecules in primary metabolism, such as glycerolipids, amino acids, and carbohydrates, with fewer reports on specialized metabolism [also known as secondary metabolites or natural products metabolites, as reviewed elsewhere (ref. Lee ; ref. Boughton ; ref. Sturtevant ; ref. Qin )]. However, recent reports have suggested that the technology is increasingly used to study specialized metabolism, thereby drastically expanding the number of metabolites profiled. Although the entire plant kingdom’s metabolome is thought to lie in the range 0.2–1 million molecules, a given plant species may still produce several thousand distinct compounds (ref. Fang ). In this section, we review progress towards chemical imaging of the plant metabolome, highlighting recent studies in the last 5 years as summarized in Table 1. The subsections below are organized by metabolite category where practical. While many of these studies analyzed multiple metabolite classes, not all metabolites mapped will be described. MSI remains limited in the number of simultaneous metabolites analyzed within a single biological sample, requiring additional technical advances (discussed further in the Conclusions). While it is difficult to accurately quantify the number of metabolites covered to date, it is no doubt still much less than conventional metabolomics studies performed on tissue extracts. Overall, however, the chemical diversity of metabolites is highly encouraging in terms of expanding metabolite coverage and providing technical conditions for future studies. Finally, it also appears that MSI instrumentation is increasingly accessible to diverse research groups, supporting its utility as a core technique and providing continual insights into in situ metabolism.
Table 1.: Representative plant metabolites analyzed by mass spectrometry imaging
| Metabolite class | MetabolitesT1Fn1 | Plant tissue | Reference | MethodT1Fn2 |
|---|---|---|---|---|
| Lipids | Triacylglycerols (TAGs), phosphatidylcholines (PCs) | Pennycress seed (Thlaspi arvense L) | ref. Johnston ; ref. Romsdahl | MALDI |
| TAGs, PCs | Rapeseed (Brassica napus) | ref. Lu | MALDI | |
| PC- and TAG-cyclopropane fatty acids (FAs) | Camelina seed (Camelina sativa) | ref. Yu | MALDI | |
| TAGs, wax esters | Jojoba seed (Simmondsia chinensis) | ref. Sturtevant | MALDI | |
| TAGs with hydroxy FAs | Castor seed (Ricinus communis L.) | ref. Sturtevant | MALDI | |
| PCs | Barrel medic nodule (Medicao truncatula) | ref. Dokwal | MALDI | |
| PCs, lysophospholipids, phosphatidylglycerol (PG) | Tomato leaf (Solanum lycopersicum) | ref. Veličković | MALDI | |
| PCs, lysoPLs, phosphatidylethanolamine (PE), phosphatidic acid (PA), sulfoquinovosyl diacylglycerol (SQDG) | Barley root (Hordeum vulgare L.) | ref. Sarabia | MALDI | |
| PA, PE, PG, phosphatidylinositol (PI) | Arabidopsis leaf (Arabidopsis thaliana) | ref. Mugume | MALDI | |
| Oxylipins, FAs, ergosterol, glycerolipids | Wheat grain (Triticum spp.) | ref. Righetti | AP-MALDI | |
| Stigmastanol | Carrot root (Daucus carota) | ref. Xiang | AP-MALDI | |
| Carbohydrates | Sucrose and non-sucrose disaccharides | Onion bulb (Allium cepa) | ref. Zhan | MALDI |
| Hexoses, sorbitol, sucrose | Apple fruit (Malus domestica) | ref. Horikawa | MALDI | |
| Hexoses, sucrose | Strawberry fruit (Fragaria × ananassa) | ref. Wang ;ref. Enomoto, (2021) | MALDIDESI | |
| (Enzymatically degraded) cellulose, hemicellulose | Maize stem (Zea mays) | ref. Arnaud | MALDI | |
| Sucrose | Grapevine leaf (Vitis vinifera) | ref. Maia | MALDI | |
| Amino acids | Cys, Asn, GABA, Gln, Lys | Strawberry fruit (Fragaria × ananassa) | ref. Nizioł | SALDI |
| Ala, Asn, Gly, Gln Leu/Ile, Val | Maize root (Zea mays) | ref. O’Neill and Lee (2020) | MALDI | |
| Leu, Asn, Pro, His, Arg, Trp | Lentil seedling (Lens culinaris L) | ref. Y.-X. Zhang | AP-MALDI | |
| Spermidine, spermine | Soybean seed (Glycine max) | ref. Sagara | AP-MALDI | |
| Spermidine, spermine | Danshen root (Salvia miltiorrhiza) | ref. Sun | MALDI | |
| N1,N10-diferuloylspermidine | Pineapple fruit (Ananas comosus) | ref. Suarez | MALDI | |
| Aconitate, (iso)citrate, succinate, fumarate, malate | Maize root (Zea mays) | ref. T. Zhang | DESI | |
| Malate, citrate | Arabidopsis seedling (Arabidopsis thaliana),liverwort seedling (Marchantia polymorpha) | ref. Gomez‐Zepeda | MALDI | |
| Pyruvate, lactate, 2-ketobutyrate, maleate/fumarate, oxaloacetate, malate | Strawberry fruit (Fragaria × ananassa) | ref. Enomoto (2021) | DESI | |
| Malate, citrate, gluconate | Notoginseng root (Panax notoginseng) | ref. Sun | MALDI | |
| Malate, maleate, citrate | Banlangen root (Isatis tinctoria L.) | ref. Nie | AP-MALDI | |
| Malate, citrate | Mango fruit (Mangifera indica L.) | ref. Zhao | AFAI | |
| Phenolics | Liquiritigenin, apigenin, naringenin, luteolin, dihydrokaempferol, daidzein, quercetin, taxifolin, kaempferol, isorhamnetin, myricetin, catechin, quercetin 3-β-d-glucoside, baicalin, rutin | Litchi seed (Litchi chinensis Sonn.) | ref. Y. Liu | MALDI |
| Quercetin, kaempferol, isorhamnetin | Rapeseed stem (Brassica napus) | ref. Krysa | MALDI | |
| Caffeic acid, rosmarinic acid, other phenolic acids | Danshen stem, root, leaf, flower (Salvia miltiorrhiza) | ref. Tong | DESI | |
| Flavonoid aglycones, biflavonoids, flavonoid glycosides, biginkgosides | Ginkgo leaf (Ginkgo biloba) | ref. Li | MALDI | |
| Pinoresinol, phillygenin, forsythoside A, forsythoside E, rutin, caffeic acid | Weeping Forsythia dried fruit (Forsythia suspensa) | ref. Jing | MALDI | |
| Puerarin and derivatives, daidzin, mirificin, ambocin, hesperidin, ononin | Puerariae sp. dried root (Puerariae lobata and P. thomsonii) | ref. Guo | AFA-DESI | |
| Nobiletin, tangeretin, tetramethoxyflavone, and feruloylputrescine, guaiacol | Citrus leaf (Citrus sinensis-limonia) | ref. de Moraes Pontes | DESI | |
| Apigenin, cannaflavin A, cannaflavin B, cannaflavin C, kaempferol, luteolin, orientin, quercetin, vitexin, isovitexsin | Cannabis leaf (Cannabis sativa) | ref. Lorensen | AP-MALDI | |
| Alkaloids | Strictosidine, reserpine and derivatives, ajmalicine and derivatives, ajmaline and derivatives, serpentine | Devil pepper root, stem, leaf, fruit (Rauvolfia tetraphylla L.) | ref. Mohana Kumara | DESI |
| Reserpine and rescinnamine, and associated biosynthetic intermediates | Devil pepper root, stem, leaf, fruit (Rauvolfia tetraphylla L.) | ref. Lorensen | AP-MALDI | |
| Atharanthine, vindolinine, serpentine, vindoline, anhydrovinblastine | Periwinkle petal (Catharanthus roseus) | ref. Dutkiewicz | SALDI | |
| Gelsemine-, koumine-, gelsedine, humantenine-, yohimbine, sapargine-type alkaloids | Heartbreak grass stem, root, leaf (Gelsemiume legans) | ref. Z.-H. Wu | DESI | |
| Arecoline, arecaidine, caffeine, cotinine, guvacine, guvacoline, hordenine, sophoridine, trigonelline, vicine | Areca fruit (Areca catechu) | ref. J. Wu | MALDI | |
| (Pseudo)ephedrine, methyl(pseudo)ephedrine | Ephedra shoot (Ephedra sinica) | ref. Yun | DART | |
| Atharanthine, vindolinine, serpentine, vindoline, and anhydrovinblastine | Peyote crown and stem (Lophophora williamsii) | ref. Lin | AP-MALDI | |
| Cocaine, truxilline, benzoylecgonine, cinnamoylcocaine | Coca plant (Erythroxylum coca) | ref. dos Santos | MALDI/LDI | |
| (Dehydro)tomatine and dervatives, (dehydro) esculeoside A | Tomato fruit (Solanum lycopersicum) | ref. Dong ; ref. Kazachkova | MALDI | |
| Lupanine, 13-hydroxylupanine, angustifoline | Narrow-leafed lupin seed (Lupinus angustifolius) | ref. Otterbach | MALDI | |
| Tomatine and derivatives, lycoperoside A-C and H | Cherry tomato fruit (Lycopersicon esculentum) | ref. Bednarz | MALDI | |
| Solasodine and derivatives, chaconine | Black nightshade fruit (Solanum nigrum) | ref. Bednarz | MALDI | |
| Tomatidenol, soladulcidine and derivatives | Bittersweet nightshade fruit (Solanum dulcamara) | ref. Bednarz | MALDI | |
| Terpenes | Vitexilactone, vitetrifolin B/E/F, rotundifuran | Chaste tree fruit (Vitex agnus-castus L.) | ref. Heskes | MALDI |
| Carnosol and tanshinone plus pathway intermediates, other diterpenes | S. grandifolia and Danshen root (Salvia grandifolia and S. miltiorrhiza) | ref. Y.-X. Zhang | AP-MALDI | |
| Kaurane diterpenes, xylopic acid | Ethiopian pepper fruit (Xylopia aethiopica) | ref. Kyekyeku | MALDI | |
| Platycodin D, platycodin D3, platycoside E | Balloon flower root (Platycodon grandiflorum) | ref. Tang | MALDI | |
| Carnosol, kahweol, lactaroviolin, squalene, toxoids | Taxus leaf (Taxus mairei) | ref. Zhan | MALDI | |
| Momilactone-A/B, phytocassane-A–E | Thai rice leaf (Oryza sativa) | ref. Komkleow | MALDI | |
| Soyasaponins | Soybean root nodules (Glycine max) | ref. Agtuca | LAESI | |
| Nobilomethylene, dendronobilin F/K, rupestonic acid G, isopetasol, dendroside G, dendroterpene C | Noble dendrobium stem (Dendrobium nobile) | ref. Q. Liu | MALDI | |
| Vitamins and pigments | (Vit E) tocopherols, tocotrienols | Upland cotton seed (Gossypium hirsutum) | ref. Salimath | MALDI |
| Vit A1, Vit B1, Vit B6, Vit C | Raw and dried persimmon fruit (Diospyros kaki) | ref. Shikano | MALDI | |
| 7-Dehydrocholesterol (provitamin D3), cholesterol | Tomato fruit (Solanum lycopersicum) | ref. Li | MALDI | |
| Anthocyanins | Strawberry fruit (Fragaria × ananassa) | ref. Wang | MALDI | |
| Betalains, chlorophyll a | Tobacco leaf (Nicotiana benthamiana) | ref. Dong | MALDI | |
| β-Carotene | Carrot root (Daucus carota) | ref. Xiang | AP-MALDI | |
| Polyacetylenes | Cangzhu root (Atractylodes lancea) | ref. Jiang | DESI/PI | |
| Cannabinoids | Δ9-THCA, CBNA | Cannabis leaf (Cannabis sativa) | ref. dos Santos | MALDI/LDI |
| Δ9-THCA, CBN(A), CBE(A), CBGA | Cannabis leaf (Cannabis sativa) | ref. Lorensen | AP-MALDI | |
| Glucosinolates | Indol-3-ylmethyl glucosinolate (GSL),4-methylthiobutyl GSL, methylthiooctyl GSL, and other GSLs | Arabidopsis leaf (Arabidopsis thaliana) | ref. Morikawa-Ichinose | MALDI |
| Isatindigoside F GSL and other GSLs | Banlangen root (Isatis tinctoria L.) | ref. Nie | DESI | |
| Hormones | Brassinosteroid, salicylic acid, 1-aminocyclopropane-1-carboxylic acid, abscisic acid, cytokinin auxin | Rice root (Oryza sativa) | ref. Shiono and Taira (2020) | Nano-PALDI |
| Nucleotide bases | Guanine, adenine, adenosine | Banlangen root (Isatis tinctoria L.) | ref. Nie , ref. 2022) | DESIAP-MALDI |
a Not all metabolites are listed for each study. Studies are generally categorized by predominant and/or unique metabolites imaged.
b Techniques: matrix-assisted laser desorption/ionization (MALDI), atmospheric pressure-MALDI (AP-MALDI), surface-assisted laser desorption/ionization (SALDI), desorption electrospray ionization (DESI), air flow-assisted ionization (AFAI), direct analysis in real-time (DART), DESI with post-photoionization assembly (DESI/PI).
Lipids
Many early MSI studies were focused on plant lipids because they are easily ionizable molecules via MALDI (e.g. using a DHB matrix) and abundant throughout many plant tissue types (ref. Horn , ref. 2013; ref. Horn and Chapman, 2014). Lipids have numerous biological functions, such as membrane compartmentation, intracellular communication, plant defense, and physical hydrophobic barriers (ref. Li-Beisson ). The heterogenous spatial distributions observed in oilseed tissues were largely unexpected, as recently reviewed (ref. Borisjuk ) and briefly highlighted here. MSI has substantially enhanced our understanding of the spatial precursor–product relationships among membrane and storage lipids [i.e. phosphatidylcholine (PC) and triacylglycerol (TAG), respectively]. For example, MSI of pennycress (an emerging bioenergy cover crop) seed embryos labeled with 13C (see ‘Spatial maps of metabolism using stable isotope labeling’) revealed complex spatial and temporal, tissue-specific labeling patterns of PC and TAG (ref. Romsdahl ). Enhancing oil content and tailoring the fatty acid (FA) composition of oilseeds is a longstanding goal within the lipid research community (ref. Horn and Benning, 2016; ref. Park ). In two independent, comparative imaging studies of high- versus low-oil lines in pennycress (ref. Johnston ) and rapeseed (ref. Lu ), the extensive heterogeneity of major TAG and PC molecular species was clearly evident; however, the spatial profiles of high-oil lines were relatively similar to those of low-oil lines. Additional differences to explain the total oil variations became evident by combining MSI with transcriptomics and conventional metabolomics, which supports the use of integrated approaches.
Some oilseeds accumulate uncommon (or unusual) FAs with important industrial and nutritional properties (ref. Ohlrogge ). For example, MSI of castor bean, which can accumulate up to 90% hydroxy-FAs, showed heterogeneous tissue distribution of mono-, di-, and tri-hydroxy-TAGs in the embryo and endosperm (ref. Sturtevant ). Likewise, in jojoba seeds, wax esters (rarely stored in high amounts in other seeds) accumulated in the cotyledons whereas TAG accumulated in the embryonic axis (ref. Sturtevant ). In both studies, integration of MSI with transcriptomics suggested tissue-specific mechanisms and primary metabolic pathways for storage lipid accumulation of selected classes and species. Imaging uncommon FAs in their host plants can offer valuable insights into metabolic bottlenecks, as it has been challenging to engineer agronomically superior oilseeds to accumulate these FAs. In metabolically engineered oilseeds, novel spatial patterns often emerge that represent metabolic changes in endogenous (and introduced) pathways, such as the production of cyclopropane FAs or very long chain polyunsaturated FAs in camelina seeds (ref. Usher ; ref. Yu ). Collectively, MSI continues to be valuable in understanding the evolution of seed biochemistry and informing metabolic engineering strategies.
MSI of lipids is not restricted to oilseeds and has expanded to most lipid classes. For example, PC molecular species were revealed to have a heterogenous distribution in nodules of the legume Medicago (ref. Dokwal ). The distributions changed in response to phosphate-deficient conditions, revealing a potential role for lipid remodeling in phosphate stress within legume–rhizobia interactions. In tomato leaves subjected to physical wounding, MSI showed variable lipid patterns in response to wounding especially in lysophospholipids (which are not commonly imaged due to potential formation via hydrolysis and technical artifacts). LysoPCs were abundant in the injured zone while some lysophosphatidylglycerol (lysoPG) species accumulated in the apex of the injured zone or were depleted (ref. Veličković ). These results led to a proposed mechanism of differential spatial hydrolysis of PC and PG in response to wounding. MSI analysis of wheat grains subjected to a fungal disease demonstrated a rapid accumulation of several defense metabolites, including oxylipins in the endosperm consistent with the activation of the 13-lipoxygenase pathway after pathogen assault (ref. Righetti ). Several membrane lipid classes profiled in barley roots under salt stress showed distinct profile shifts in molecular species of PC and lysoPC, phosphatidic acid (PA), phosphatidylethanolamine (PE), and sulfoquinovosyldiacylglycerol (SQDG) (ref. Sarabia ). Alternatively, in a study on the role of autophagy in Arabidopsis leaves, most phospholipid molecular species in autophagy mutants and under different nitrogen replete/deplete conditions showed unchanged spatial profiles (ref. Mugume ). This was surprising considering changes observed in individual lipids from total leaf extracts. However, spatial imaging revealed findings otherwise lost in extract-based analysis; that is, the lipid responses in cells (within sections imaged) must be triggered simultaneously in response to autophagic status. Many studies continue to highlight the role of lipid remodeling in abiotic and biotic stress responses (ref. Yu ), where MSI can play an important role in addressing spatial metabolic alterations and mechanisms.
Carbohydrates (sugars)
Carbohydrates are primary photosynthates with a multitude of physiological functions in plants, such as energy and transport, structural building blocks, developmental and stress signaling responses, and flavor indicators (ref. Pigman, 2012). While MSI can detect most monosaccharides and disaccharides using DHB or compatible matrices, other non-invasive chemical imaging methods, such as MRI, are particularly powerful for visualizing the in situ distribution of sugars (ref. Borisjuk ). Sweetness is a major factor in consumer preference for many fruits. MSI of different maturity stages within strawberry sections showed accumulation of hexoses (glucose and fructose) and sucrose during ripening (ref. Wang ). While hexoses were preferentially located in the pith and receptacle structures, sucrose was more uniformly detected throughout the sections and more abundant in the red maturity stage. Similar hexose and sucrose spatial distributions of mature strawberries were observed using DESI-MSI, along with unique profiles of many other metabolite classes (ref. Enomoto, 2021). The spatial distribution of specific sugars is probably linked to the regulation of ripening through abscisic acid (ABA; along with other metabolites such as organic acids and anthocyanins). In contrast, for apple, where sorbitol is the primary carbohydrate transported from leaves to fruits, this sugar alcohol accumulated preferentially at the center of the fruit whereas sucrose was higher in the cortex (ref. Horikawa ). As excessive sorbitol accumulation leads to an abiotic stress disorder called ‘watercore’ (ref. Itai, 2015), additional MSI studies may be beneficial to examining mutants in sorbitol transport.
Resolving isomers in MSI can be difficult, and many times an m/z is associated with sucrose when in fact it may be one of several disaccharide isomers. Tandem MS has been used to resolve sucrose and non-sucrose disaccharide isomers in cross-sections of onion bulb, showing distinct isomer patterns in inner versus outer epidermis, results that may reveal additional functional or metabolic roles of disaccharides (ref. Zhan ). Polysaccharides are predominantly used for storage and assembly of cell walls in essentially all plant tissues, but their size and chemical linkages render polysaccharides difficult for ionization and detection by current MSI methods. Mapping the distribution of enzymatically degraded cell wall polysaccharides offers an alternative approach, as shown in maize stems (ref. Arnaud ). Imaging cellulose, mixed linked β-glycans, and heteroxylans showed distinct patterns within lignified and non-lignified tissues which may help better address the recalcitrance of producing cellulosic biofuels. Recently, grapevine leaf discs infected with the fungal pathogen Plasmopara viticola showed the accumulation of sucrose mainly on the veins of infection sites, contributing to new knowledge on the fungal infection structures (ref. Maia ). In this study, MSI facilitated the observations of spatial changes in the carbohydrate distribution that is associated with pathogen interaction, and also provided evidence for the authors to hypothesize that the pathogen itself may have gained access to the metabolite for completing its life cycle. Future studies that focus on the detection of sugars associated with signaling and developmental cues such as trehalose 6-phosphate could expand the applications of MSI in the field of carbohydrates.
Amino acids
In addition to building blocks for protein synthesis, many amino acids are precursors for specialized metabolites and have active roles in plant development and stress responses (ref. Trovato ). Free amino acids typically have poor ionization efficiency for MALDI-MSI analysis; however, on-tissue derivatization with coniferyl aldehyde improves the ionization efficiency (ref. Manier ) such that all 20 canonical amino acids have been detected and most visualized in situ (ref. O’Neill and Lee, 2020). For example, MSI of young roots within maize B73 and Mo17 inbred lines showed a development-associated shift in amino acid signal from the cortex to the root center, and an eventual decrease in overall tissue signal. These spatial profiles are consistent with the transport of certain amino acids from the root to other parts of the maize plant. Furthermore, spatial differences between the inbred lines may point to differences in contributions of direct metabolic synthesis in roots versus transport. The inbred lines also had differences in abundance and localization of amino acids, with hybrid offspring inheriting spatial profile characteristics from both parents, suggesting that spatial metabolic profiling may include biomarkers for breeding programs.
In germinating seedlings of lentil, spatial differences were observed for multiple amino acids from 12 h to 72 h post-germination, hypothesized to be a reflection of protease-mediated hydrolysis of seed storage proteins for germination and seedling establishment (ref. Y.-Z. Zhang ). Profiling of amino acids has also been enhanced using a matrix of cationic silver nanoparticles (ref. Nizioł ). In strawberry fruits, differences in spatial distributions of amino acids suggests that there is cell-specific metabolic regulation of several amino acid pathways, for example (i) achene-enriched (containing the seed), nitrogen-rich Gln, Lys, and Asn as an energy source for germination; (ii) periphery-enriched, the non-proteinogenic amino acid γ-aminobutyric acid (γ-GABA) important for tissue protection; and (iii) dispersive, Cys the most abundant biothiol in strawberry for oxidative protection. Future studies would benefit from continuing to improve the detection of amino acids as well as establishing precursor–product pathways with spatially distinct metabolites.
Polyamines are aliphatic polycations derived from amino acids (Arg and ornithine) with numerous roles in plant health and stress responses (ref. Handa ). In soybean seed cross-sections, two polyamines (spermidine and spermine) were more abundant in the embryo axis relative to the cotyledons, with the highest concentrations in root and shoot meristem tissues (ref. Sagara ). Spatial profiles of metabolites reflect enzymatic activity and regulation, and, here, the spatial profiling of polyamines was consistent with immunohistochemistry and fluorescent microscopy studies of the respective polyamine-producing aminopropyltransferase enzymes (ref. Belda-Palazón ). In roots of the medicinal plant Salvia miltiorrhiza, spermine and spermidine were enriched in the xylem and outer cortex roots of repeatedly cropped plants relative to other root tissue regions (ref. Sun ). The authors were surprised to find that normal cropped roots had much higher polyamine levels, which may explain some of the altered growth physiology observed in continuously cropped plants. Finally, a polyamine derivative N1,N10-diferuloylspermidine, a potential HMG-CoA reductase (HMGCR) inhibitor with the potential to lower cholesterol biosynthesis, was localized in pineapple tissue near the peel/shell and gradually decreased towards the flesh and ovary (ref. Suarez ). Cells on the periphery of plant tissues are exposed to UV light and probably regulate the production of protective metabolites such as polyamines, which may explain the distribution of polyamine derivatives in pineapple fruits. Additional MSI studies of polyamines are likely to yield new insights into plant growth and stress responses.
Organic acids
Organic acids have important roles in plant physiology as central intermediates in carbon metabolism, pH regulation, and redox regulation (ref. Igamberdiev and Eprintsev, 2016), with many types having been spatially mapped in situ. For example, DESI-MSI of maize roots in longitudinal sections showed that tricarboxylic acid (TCA) cycle intermediates were enriched in developmentally distinct regions, with implications for plant growth and development (ref. T. Zhang ). While succinate was enriched in the meristem, three TCA intermediates, aconitate, fumarate, and malate, were enriched in the differentiation zone. These results were consistent with tissue-specific gene expression profiles [e.g. genes encoding aconitase (producing aconitate), succinate dehydrogenase (producing fumarate), and fumarase (producing malate) were enriched in the differentiation zone]. Disruption of the normal expression patterns in addition to experiments exogenously treating roots with TCA intermediates showed that metabolite localization was highly predictive of molecular functions in stem cell behavior.
Root extrusion is important for plant nutrition, tolerance to cation toxicity, and plant–microbe interactions (ref. Gomez‐Zepeda ). An MSI method developed for visualizing organic acids in roots and root exudates of Arabidopsis seedlings in different environments has enabled new ways of examining how localization of these metabolites is linked to physiological responses. Furthermore, in root cross-sections of the medicinal plants, ginseng and Banlangen, multiple organic acids were enriched in the phloem and medulla rather than the xylem (ref. Nie ; ref. Sun ). The enrichment patterns of these metabolites in ginseng may be reflective of organic acids contributing to the rhizosphere exudate (ref. Goodwin, 2022). In fruits, organic acids also serve as flavoring agents that impact acidity and organoleptic qualities. In mango cross-sections analyzed by air flow-assisted ionization, it was shown that malate was distributed throughout the slice whereas citrate was enriched in the pulp (ref. Zhao ). Given their role in central metabolism, and other pathways, applying isotope labeling MSI methods may yield unique insights into the spatial regulation of these metabolic pathways and storage endpoints.
Phenolics
Phenolics are a large group of structurally diverse, specialized metabolites ranging from simple phenolic compounds to more complex molecules and polymers. These compounds contribute to diverse functions such as plant–herbivore interactions, antioxidants, and the color, taste, and flavor of many herbs, foods, and plant-derived beverages (ref. Rehab and Amira, 2018). Several studies have used MSI to visualize flavonoid-rich plants with promising pharmacological activity. For example, MALDI-MSI of litchi seed tissue sections using the 2-mercaptobenzothiazole (2-MBT) matrix revealed 15 detectable flavonoids grouped into four tissue-distinct spatial patterns: four embryo-enriched flavonoids (e.g. luteolin), three cotyledon-enriched (e.g. rutin), seven distributed throughout both embryo and cotyledon (e.g. apigenin, catechin, quercetin), and one in the testa and cotyledon periphery (e.g. taxifolin) (ref. Y. Liu ). Each of these unique tissue enrichment patterns may be attributed to their proposed specialized functional roles in litchi seed and a result of regulation at pathway branch points of shared phenolic substrates.
Similarly, a DESI-MSI study of diverse tissues including roots, flowers, stems, and leaves from Chinese medicinal herb S. miltiorrhiza showed phenolic acids abundant in the root cortex and vasculature, in contrast to the abietane diterpene, tanshinone, restricted to the periderm (ref. Tong ). Standard metabolomics is an important tool for pathway elucidation. The integration of MSI data can provide spatial context to complex pathways and, in some cases, help address multiple biosynthetic routes. For example, there are multiple possible metabolic pathways for the biosynthesis of rosmarinic acid. MSI showed similar co-localization of two potential substrates—caffeic acid and danshensu—that, when combined with metabolomics-based 13C trace studies, suggested this route as the core biosynthesis pathway to produce rosmarinic acid in Salvia species.
Other recent MSI studies on medicinal plants have spatially mapped diverse phenolics in Cannabis sativa leaves (discussed below with ‘Other metabolites’) (ref. Lorensen ), Ginkgo biloba leaves (ref. Li ), dried fruits of Forsythia suspensa (ref. Jing ), dried roots of two Radix puerariae species (ref. Guo ), and Paeonia sp. roots (ref. Li ). Finally, imaging of phenolics is not restricted to medicinal plants and is likely to be important to understanding abiotic and biotic interactions. For example, MSI of rapeseed treated with Nod factor-based biofertilizers (flavonoid-induced signaling molecules produced by rhizobia in legume root nodule formation) for testing their impact on non-legumes showed an alteration in stem flavonoid patterns, with quercetin and kaempferol derivatives increasing in concentration post-treatment (ref. Krysa ). MSI of metabolites may also be used to screen for pathogenicity, for example specific flavonoids (and other metabolites) accumulated in citrus in response to yellow dragon disease (ref. de Moraes Pontes ). These studies on flavonoid spatial distribution lay the groundwork for spatial pathway elucidation, trait selection in human health applications, and genetic engineering of specific phenolic compounds in alternative crops.
Terpenes
Terpenes are a large and diverse class of isoprene-derived compounds with roles in plant defense, cell development, membrane permeability, light harvesting, and protection, as well as contributing to plant fragrance, taste, and pigments (ref. Rehab and Amira, 2018). Relative to other specialized metabolites, many terpenes without polar groups are not easily ionized. However, there are several recent reports on imaging diterpenoids. For example, DESI-MSI revealed the spatial maps of 33 metabolites involved in diterpenoid tanshinone and carnosol biosynthetic pathways in S. miltiorrhiza and S. grandifolia roots (ref. Xia ). Most of the diterpenoid molecules were visualized in the periderm (with similar results of tanshinone to ref. Tong ), although some compounds showed differential accumulation between the two related species. In roots of the medicinal plant Platycodon grandiflorum, three bioactive oleanane-type triterpenoid saponins were observed enriched in the R2 versus R1 xylem layer of cells, a rather unusual spatial distribution, that led to discovery of a glycosyltransferase involved in the biosynthesis (ref. Tang ). In shoot tissues, some terpenes accumulate in leaves while others accumulate in glandular trichomes (along with other specialized metabolites). For example, in the plant that produces compounds for the anti-cancer drug taxol, Taxus sp., a few bioactive terpenes accumulated in the spongy mesophyll layer (e.g. toxoids, carnosol) while squalene was located in palisade mesophyll cells (ref. Zhan ). MSI of the medicinal plant Vitex agnus-castus L. showed the diterpenoid vitexilactone, linked to lowering prolactin in treating female reproductive conditions, localized on the periphery of fruits presumed to be glandular trichomes (ref. Heskes ). Other recent studies analyzing terpenes include kaurane diterpenes in Ethiopian pepper (ref. Kyekyeku ), triterpenoid soyasaponins in root nodules disrupted in symbiosis (ref. Agtuca ), accumulation of diterpenoid phytoalexins in Thai rice in response to blast disease (ref. Komkleow ), and distribution of sesquiterpenes in mature Dendrobium nobile stem (ref. Q. Liu ). Increasing the amounts of (or engineering new tissues to) produce bioactive compounds requires a better understanding of the mechanisms used to naturally accumulate these compounds. Improvements in terpene ionization and identification should be prioritized to broaden our knowledge of the spatial organization of metabolism for this largest class of specialized metabolites in plants.
Alkaloids
Alkaloids are structurally diverse, specialized metabolites with a heterocyclic ring containing a nitrogen atom. Alkaloids have diverse functions in plants as feeding deterrents, antimicrobial defenses, and germination inhibitors (ref. Jacobowitz and Weng, 2020). Like flavonoids, given their important pharmaceutical, bioactive properties, it is not surprising that many MSI studies have targeted the spatial profiles of alkaloids to characterize these metabolic pathways and their regulation. For example, two recent imaging studies using DESI- and/or MALDI-MSI showed heterogeneous spatial patterns of monoterpenoid indole alkaloids (MIAs) within diverse tissues of Rauvolfia tetraphylla (ref. Mohana Kumara ; ref. Lorensen ). MIAs were enriched in the early stages of mature fruit, with several compounds restricted to the exocarp (e.g. yohimbine), while ajmaline was largely restricted to the mesocarp (ref. Mohana Kumara ). In roots and stems, enrichment was observed for MIAs in distinct tissues: epidermis (e.g. deserpidine), cortex (e.g. ajmalicine), pith only (e.g. reserpine), and epidermis and pith (e.g. serpentine). MIAs were less often detected in leaves. To produce chemically diverse compounds in specific tissues that use similar initial building blocks, such as trypamine (via tryptophan decarboxylation), inherently requires coordinated metabolic regulation and utilization of diverse biochemical reactions (ref. Horn, 2021). Another study focused on using MSI to address the biosynthesis of reserpine, an anti-hypertension and anti-microbial agent (and biochemically related rescinnamine), by analyzing not only the endproduct reserpine (and rescinnamine) but also the theoretical intermediates (ref. Lorensen ). High-resolution MALDI-MSI provided unprecedented detail, detecting most of the proposed intermediates of reserpine in similar cell types among diverse tissues, and this was reinforced using a deuterium-labeled trypamine precursor. Finally, a matrix-free imaging method using tissue imprints on a functionalized TiO2 nanowire solid substrate was highly effective in imaging a range of alkaloid pathways in the fragile petals of periwinkle (ref. Dutkiewicz )
Several alkaloids are also addictive drugs, including nicotine, caffeine, morphine, and the lesser reported arecoline. Arecoline accumulates in the fruit of Areca catechu. MALDI-MSI [using a 3,4-dimethoxycinnamic acid (DMCA) matrix] revealed heterogeneous patterns of arecoline and other bioactive alkaloids (ref. J. Wu ). Understanding how localization impacts bioactivity will be important for therapeutic applications and may inform understanding of tissue-specific functions for these metabolites. Other alkaloid-enriched medicinal plants recently studied by MSI include Gelsemium elegans showing differential accumulation in root and stem cell types and insights into alkaloid diffusion and transfer among tissues (ref. Z.H. Wu ). Also imaged were the hallucinogenic compound mescaline enriched in epidermal cells of peyote crown and stem (ref. Lin ), and cocaine in leaves of coca plant (ref. dos Santos ).
Multiple MSI studies have focused on characterizing the spatial distribution of toxic alkaloids associated with fruit and seed bitterness (an adaptive trait to ward off herbivores prior to the ripening process being finished). For example, as tomato fruits ripen, the bitter compound α-tomatine (an important plant defense compound) is exported from the vacuole to the cytosol and enzymatically converted into non-bitter and non-toxic esculeosides (ref. Kazachkova ). An MSI study using virus-induced gene silencing (VIGS) of the α-tomatine transporter showed a dramatic increase in α-tomatine and a decrease in esculeoside A in virus-infected regions. MSI is powerful in studying parallel pathways. In this manner, using RNAi to silence the conversion of dehydrotomatine to α-tomatine (with dehydrotomatine subsequently converted to dehydroesculeoside during ripening) showed that both metabolites were abundant in fruit epidermis and pericarp, but α-tomatine was higher in the wild type and dehydrotomatine higher in the RNAi lines (ref. Dong ). In a broader study of the Solanaceae family, structurally diverse alkaloids were profiled by MSI (ref. Bednarz ). In Solanum lycopersicum, the major alkaloids showed distinct spatial patterns in different plant tissues (e.g. fruits versus seed versus stalk versus flower), while in S. nigrum and S. dulcamar dramatic shifts were observed in alkaloid distribution during fruit ripening. Advanced tissue-specific developmental profiling, such as single-cell RNA-seq, would probably reveal elaborate gene expression profiles that enable these metabolite distributions. Bitterness is not restricted to the fruit of the Solanaceae family. In the legume Lupinus angustifolius, toxic quinolizidine alkaloids (QAs) have long been known to accumulate in the seed (ref. Otterbach ). MSI was able to visualize these bitter QA compounds in pods (but not seeds) of early developmental stages, whereas upon maturation, the distribution of QAs shifted to seeds (and was absent in pods). These results support a possible mechanism for transport of these bitter QAs, thereby opening up new strategies for engineering new (less bitter) varieties.
Finally, typically MSI studies have focused on increased spatial resolution that can address spatial metabolite patterns at the cellular or even organellar levels (ref. Hansen and Lee, 2018; ref. Dong ). However, high-resolution imaging is time consuming via MALDI-MS and, therefore, alternative methods are needed for imaging large areas, such as whole plants. To address this need, ambient ionization MS was coupled to direct analysis in real-time time-of flight-MS (DART-TOF-MS) on the medicinal plant Ephedra sp. (ref. Yun ). Here, using low resolution imaging, ~1 mm spot size, the alkaloids (pseudo)ephedrine and methyl(pseudo)ephedrine were visualized. In stems, the alkaloids were enriched on the inner section (versus the outer section). The side branches (commonly used for medicine) were enriched in these alkaloids relative to the main stem, with concentration gradients decreasing towards the branch tips. Whole-plant chemical imaging is powerful for studying plant physiology and biochemistry, as revealed using other non-invasive imaging approaches (ref. Borisjuk ). These MSI studies within alkaloids signify a promising path for future alkaloid pathway discovery, localization, and engineering.
Vitamins and pigments
Plant vitamins are essential for metabolism as enzymatic cofactors and their roles in redox chemistry (ref. Asensi-Fabado and Munné-Bosch, 2010). Cottonseed oil is naturally rich in vitamin E components but contains little or no tocotrienols (ref. Salimath ). To improve the oil value, tocotrienols were produced through genetic engineering of upland cotton. MSI showed that engineered tocotrienols accumulated throughout the transgenic embryos while α- and γ-tocopherol were enriched in the cotyledon or embryonic axis, respectively, despite similar upstream substrates. This location contrasted with the polyphenol gossypol located in embryo gland cells. Introducing changes in metabolic pathways via engineering requires careful consideration of endogenous spatial distributions and pathway fluxes, as well as promoters for gene expression of introduced enzymes and other levels of metabolic regulation. Using similar genome editing approaches to address human vitamin D deficiencies and the poor vitamin D content in plants, tomato fruits were engineered to produce 7-dehydrocholesterol (7-DHC or provitamin D3) (ref. Li ). Imaging 7-DHC (and small amounts of downstream cholesterol) showed a relatively equal distribution in the flesh and peel of tomatoes. Surprisingly, α-tomatine levels were also reduced, with shifts in its metabolic distribution providing insights into these integrated pathways. In another study, the provitamin A β-carotene within carrots was widely found in the phloem and cambium regions, while it was less common in the metaxylem and protoxylem regions, pointing to the roles of carotenoid transport and metabolite accumulation (ref. Xiang ). Lastly, many plants are processed prior to human consumption. MSI of raw versus dried persimmon fruit tissues showed a dramatic change in localization and relative amounts of vitamins: lipid-soluble vitamin A1 increased in the pericarp, water-soluble vitamin B1 and B6 increased in the moist mesocarp, and vitamin C underwent photodegradation (ref. Shikano ). Considering the prevalence of nutrient deficiencies and poor dietary habits of many humans, using MSI should assist in efforts for breeding, fortification, and processing of vitamin-rich foods.
Plant pigments are important in light harvesting, photoprotection, and overall growth and development of plants, as well as serving as visible signals to attract insects, birds, and animals for pollination and seed dispersal (ref. Fernández-López ). MSI is frequently used to visualize pigment distribution in plants. For example, the red anthocyanins of strawberries were mainly located in the periphery of fruit at late maturity stages associated with antioxidant roles (ref. Wang ). In the medicinal plant Atractylodes lancea, the color of the rhizoma secretory cavities is often associated with quality, and this color varies widely among accessions (ref. Jiang , Preprint). MSI found that three polyacetylenes were probably contributing to the red color and could serve as a biomarker for trait selection. Although the biosynthesis pathways for these polyacetylenes have not been elucidated (the genome for A. lancea has not been reported), MSI suggests that a transcriptomics analysis of these secretory cavities may reveal gene candidates. Finally, the natural colorants betalains were heterologously produced in tobacco through transient overexpression of key genes, showing accumulation of pigment at the site of infection in contrast to naturally produced chlorophyll which is widely distributed (ref. Dong ). Transient expression studies are powerful for testing and confirming metabolic pathways. The importance of color (and associated functions) necessitates further studies on pigment biosynthesis and localization via MSI.
Other metabolites
Cannabinoids are of broad pharmaceutical interest, with recent demand increases driving trait optimization of the host plant Cannabis sativa and renewed interest in the biosynthesis of these compounds (ref. Lorensen ). An initial MSI study of cannabis leaves determined that 2-mercaptobenzothiazole (MBT) was the optimal matrix for MALDI detection of the acidic cannabinoids Δ9-THCA (Δ-9-tetrahydrocannabinolic acid) and cannabinolic acid (CBNA). Punctate accumulation patterns were observed, albeit at lower resolution. A follow-up study combining the lower resolution DESI-MSI (both tissue imprints and direct analysis) with higher resolution MALDI-MSI [using a 1,5-diaminonaphthalene (DAN) matrix] largely confirmed the accumulation of cannabinoids within glandular trichomes (by comparison with a chloroplast-specific lipid showing the contour of the leaf). This study also expanded the number of cannabinoids of interest detected, such as THCA, CBNA, CBN, CBE(A), and CBGA, to understanding synergistic effects of bioactive compounds. In contrast to the discrete cannabinoid localization, several flavonoids also appeared more dispersed throughout the leaf. It will be interesting to observe how future engineering designs in alternative hosts may address the location of production for cannabinoid biosynthesis.
For most MSI studies, compounds are present and stable in the tissue of interest under multiple sampling conditions. However, for sulfur-containing glucosinolates with roles in plant defense and plant storage, it was observed that these compounds localized to inner regions of leaves and decreased markedly under darkness (ref. Morikawa-Ichinose ). Several glusoinolates were also detected in the dried root of the medicinal plant Isatis tinctoria, for example isatindigoside F which accumulated mostly in phloem (ref. Nie ). Like variations observed in field versus controlled environmental studies, it will be important to prioritize sampling conditions for all MSI studies.
Nucleosides and nucleobases are not commonly imaged due to their lower molecular weight; however, a few studies have captured spatial maps. For example, in dried root of Isatis tinctoria, guanine, adenine, and adenosine were located in the cambium and outer areas of the phloem (ref. Nie ). In a related study on Isatis root, guanine was located almost exclusively in the root xylem (ref. Nie ). In addition to roles in nucleic acid biosynthesis, nucleoside triphosphates (dNTPs) are essential molecules for energy coupling, use building blocks shared with other metabolic pathways, and themselves may be precursors for downstream metabolites. Chemical imaging of these compounds may inform differences in spatial utilization among plant tissues.
Plant hormones are notoriously difficult to image via MALDI-MSI due to matrix effects on low-abundant small molecules (<500 m/z). However, nanoparticle-assisted LDI (nano-PALDI)-MSI has enabled the mapping of multiple hormones in root cross-sections of rice (ref. Shiono and Taira, 2020). In the elongation zone, brassinosteroid, salicylic acid, and the ethylene precursor ACC (1-aminocyclopropane-1-carboxylic acid) were broadly distributed, whereas ABA and cytokinin (CK) were higher in the outer root cells relative to the stele, and auxin (IAA) was detected in the epidermis, cortex, and stele. MSI is particularly well suited to address questions regarding hormone localization given that the metabolic pathway location for synthesis is often distant from the site of action. Imaging the developmental and response patterns for hormones in plant tissues more broadly will probably support major research advances in the future given the wide range of physiological responses regulated by these diverse molecules.
Finally, plants grown in natural environments are exposed to exogenous metabolites, xenobiotics, and contaminants (ref. Singh ). In recent studies, MSI has revealed (i) in maize roots, the fungal mycotoxin (aflatoxin B1) infiltrated the root vascular cells resulting in the disruption of multiple metabolic pathways and their spatial distributions (ref. Righetti ); (ii) in carrot roots, prolonged exposure to a phthalate ester organic pollutant showed its accumulation in vascular cells and associated changes in plant metabolites (ref. Xiang ); (iii) in apples, an applied fungicide imazalil penetrated from the peel to the inner region during storage (ref. Pereira ); (iv) in wheat leaves, penetration and accumulation of the fungicide azoxystrobin in the cells occurred around the vascular bundle; and (v) in cabbage leaves, visualization enabled a proposed translocation mechanism for carrier-mediated modified pesticides (ref. Wu ). MSI enables a fresh technical approach for examining the incorporation and spatial distribution of exogenous compounds and their impact on plant metabolism.
Spatial maps of metabolism using stable isotope labeling
LC/MS-based approaches to analyze stable isotope-labeled tissues has become commonplace to trace metabolites and characterize flux through specific metabolic pathways (ref. Freund and Hegeman, 2017; ref. Allen and Young, 2020). Less commonly employed is the application of MSI to the localization of metabolites following stable isotope labeling. However, recent reports highlight both the promise and challenge of combining MSI with 13C labeling approaches, especially becoming more frequent in biomedical applications (ref. Grey ). The challenges are both analytical and computational, but advances in MS instrumentation and computational workflows have made it possible to map the locations of complex metabolic pathway metabolites, even where there may be many potential isotopomers of each metabolite in a single 13C labeling experiment. For example, in studies aimed at assessing metabolic heterogeneity in glioblastoma tumors, mice fed [13C]glucose were subjected to MSI of brain sections (ref. Schwaiger-Haber ). Here, a combination of DESI- and MALDI-MSI was used to evaluate metabolic pathways induced in tumors. FA synthesis was elevated 8-fold in tumors over nearby healthy cells, and a specific enhancement of elongation of FAs was noted in glioblastoma tumors. In other work, the metabolism of surfactant in lung tissues was examined by MSI, where application of [13C]PC species enabled visualization of lipid remodeling in lung tissues (ref. Ellis ). Other stable isotopes, including 2H and 15N, can be useful in MSI approaches to visualize metabolism. By way of example, MSI (using secondary ion MS, SIMS) demonstrated that the metabolic utilization of [2H]glucose and [15N]glutamine by different tumors in situ was remarkably heterogenous among individual tumors (ref. Zhang ). In addition to monitoring metabolism, MSI by SIMS of [13C]thymidine and [15N]thymidine has been used to image DNA in stem cells (ref. McMahon and Lechene, 2021). Variations of this multi-isotope imaging mass spectrometry (MIMS) using a focused ion beam to generate secondary ions has remarkable lateral resolution of <50 nm and has been used to image processes at subcellular resolution (ref. Gyngard and Steinhauser, 2019), such as protein turnover by lysosomes (ref. Narendra ) and lipid sorting to neuronal lipid droplets (ref. Bailey ). In plants, nanoSIMS has been used in parallel with NMR and MALDI-MSI to examine drought-tolerant mechanisms in plant roots of Piper sp., Hibiscus rosa sinensis, and Clitoria fairchildiana roots (ref. Honeker ). NanoSIMS of [13C]pyruvate-labeled roots showed different 13C distribution patterns, associated with carbohydrate storage, in the presence versus absence of its rhizosphere. The rhizosphere is an important contributor to drought responses in many plants.
Although mostly applied to date to animal tissues, MSI studies of metabolism with stable isotopes are beginning to emerge in plant systems. In one report, siliques of two Brassica species (camelina and pennycress) were fed with [13C]glucose, and the isotopologs of PC species (a key intermediate in TAG accumulation) were imaged in developing embryos by MALDI-MSI at 64 h and 120 h after labeling following transport of labeled precursor into seed tissues. These efforts helped shed light on the metabolic processes for the conversion of carbohydrate into storage lipids in plant embryos, particularly in the later stages of glycerolipid assembly. Here differences in FA desaturation and elongation were noted among different tissues in the embryos (ref. Romsdahl ). In both plant species, embryos incorporated 13C into PC species with different efficiency in cotyledons and embryonic axis tissues of developing seeds (Fig. 2). In addition, in both tissues, 13C isotopes of longer chain, more saturated PC species were enriched over those PC species with polyunsaturated species, suggesting that FA elongation proceeds more rapidly in embryos compared with desaturation activities.
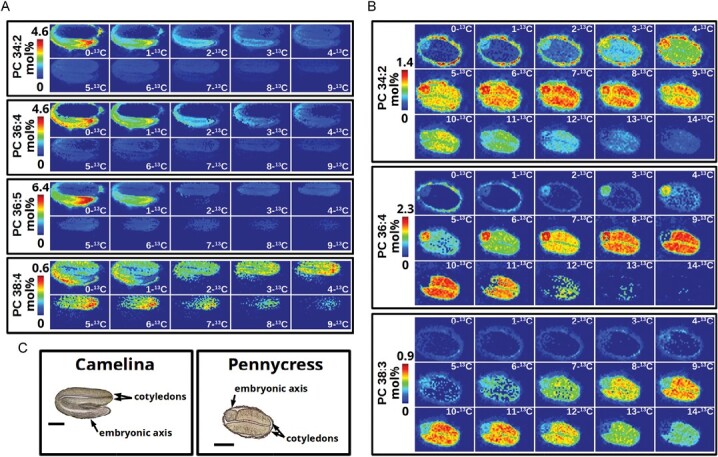
It is likely that future studies with stable isotopes will begin to reveal more nuances about the compartmentalization of plant metabolism. Nonetheless, it remains a significant challenge to combine isotope labeling approaches with MALDI-MSI. Matrix interference can obscure spectra of small metabolites, and larger molecules that are more easily resolved by MALDI-MS have increased complexity with more carbon atoms. For example, a typical PC molecular species with one palmitoyl and one linoleoyl acyl chain has 44 carbons, including the head group, so thus up to 44 potential isotopologs that could be labeled with 13C. Plants typically have ≥20 molecular species of PC. Further, a difference of 0.009 amu separates the m/z of a saturated FA versus an unsaturated FA with two 13C isotopes. Hence, while ref. Romsdahl demonstrated the feasibility of MSI and stable isotope labeling in plant embryos, the experimental data from a 13C labeling experiment can be extremely complex even when just focusing on one class of a particular phospholipid. Most probably advances in computational capabilities and the incorporation of artificial intelligence applications into MS data analysis will make these challenges more tractable, and we will see more examples of isotope labeling studies of spatial metabolism in plant systems in the future.
Conclusions: the future of spatial metabolome imaging
Chemical-based, spatial imaging techniques have enabled researchers to see and study plant metabolism as never before (ref. Borisjuk ). MSI delivers a powerful spatial snapshot revealing metabolite distribution and relative concentration that has resulted from complex mechanisms of gene expression and metabolism. In the past 5 years, there has been a dramatic expansion in the number of metabolite classes, tissue types, and the plant species that have been examined by MSI (Table 1). Through these studies, common themes continue to emerge that reinforce the utility of MSI approaches for plant metabolism research and suggest the following three key questions/areas for future development.
Heterogeneous spatial patterns
Why and how are metabolites visualized by MSI consistently organized in heterogeneous spatial distribution patterns in situ? For some specialized pathways, the answer is obvious in that plants produce and concentrate molecules where they are needed. However, this phenomenon extends to metabolites involved in primary and specialized metabolism, irrespective of their abundance, and across various tissue types and plant species. Furthermore, MSI has now supplied overwhelming evidence that metabolite heterogeneity is a key contributor in engineering bioproducts in plants, and therefore should be considered to achieve targeted outcomes. Consequently, it is imperative to re-evaluate ways to delineate metabolic pathways. Traditionally, metabolic pathways are depicted as discrete biochemical steps that connect substrates, products, enzymatic reactions, and encoded genes, as exemplified by the Plant Metabolic Network pathways (ref. Hawkins ). These representations are invaluable and certainly adequate for many applications. However, they often fail to account for spatial and temporal variations or tissue-specific differences in pathway dynamics. Future investigations, supported by advanced computational techniques and visualization methods, should strive to incorporate these additional dimensions when exploring plant metabolism.
Pathway discovery and elucidation
Can MSI play a broader, systematic role in metabolic pathway elucidation and discovery? Conventional metabolomics, supported by genetic approaches, is critical for characterizing biochemical pathways. Nevertheless, MSI is increasingly used to examine the spatial profiles of metabolic intermediates and endproducts, providing additional insights into metabolic pathways. In some cases, co-localization of metabolites has suggested contributions of a specific alternative pathway that may have been hidden in analysis of extracts by conventional metabolomics. MSI data are highly insightful when integrated with metabolic maps and tissue-specific transcriptomics. In most cases (but not all), the heterogeneous spatial patterns can be attributed to variations in gene expression patterns for individual biochemical steps or pathways. Furthermore, steady-state isotope labeling of plant tissues visualized by MSI, while complex to analyze, could help link active metabolic pathways and metabolite accumulations in situ. Future studies that integrate isotope labeling, single-cell RNA-seq, and single-cell proteomics technologies should enable exciting new findings in the field of plant metabolism.
Overcoming limitations in identification, quantification, and interpretation
What limitations and technical challenges need to be overcome for MSI to be employed as a routine metabolomics tool? MSI still has limitations as a chemical imaging methodology. Other recent reviews have addressed various limitations in more detail (ref. Dong ; ref. Hu ; ref. Unsihuay ; ref. Dong and Aharoni, 2022; ref. Borisjuk ), and a few key limitations are highlighted here. Foremost, like most analytical techniques, sample preparation is key and there is no single methodology that can image all metabolites simultaneously at this time (i.e. the entire plant metabolome). However, an increasing number of proof-of-concept studies for diverse metabolites provide empirical evidence for researchers wishing to incorporate MSI into their analytical toolbox (Table 1). In terms of methods, MALDI-MSI has been far more commonly used than DESI-MSI, contributing to a wider scope of types of metabolites imaged (Supplementary Table S1). Based on these studies, the most common reasons attributed to using MALDI-MSI (versus DESI-MSI) included superior spatial resolution, enhanced metabolite signal, and ionization source availability. DESI-MSI was used more often when imaging fragile tissues and to illustrate technical developments. Across all MALDI studies, >15 types of matrix and nanoparticles were used, with DHB the most common, covering most metabolite classes. Systematic testing and optimization of matrices remains limited in terms of recommendation for use on specific metabolites. Nonetheless, even under ideal sample conditions, MSI is limited in compound identification and verification due to the limited number of ions available for analysis and resolution of isobaric compounds. Advances in MSI sensitivity and resolution [e.g. transmission-mode geometry (t-MALDI-MSI) and laser-induced post-ionization (MALDI-2), (ref. Niehaus )], supported by advanced computational algorithms, should continue to improve the utility of MSI in these areas. While absolute quantification of compounds is difficult, advances have been made incorporating internal standards within chemical matrices (for MALDI) and methods for normalizing differences in ionization efficiencies. In most cases, relative quantification of metabolites remains informative. Finally, MSI raw data and generated images are often complex to interpret and integrate with other multi-omics datasets. Machine learning and artificial intelligence-based algorithms are increasingly used in MSI analysis, and represent a key development that will enhance the processing and interpretation of MSI information (ref. Verbeeck ; ref. Boiko ; ref. Aharoni ; ref. Jetybayeva ).
Supplementary Materials
References
- Metabolomic profiling of wild‐type and mutant soybean root nodules using laser-ablation electrospray ionization mass spectrometry reveals altered metabolism.. The Plant Journal, 2020. [PubMed]
- Plant and microbial sciences as key drivers in the development of metabolomics research.. Proceedings of the National Academy of Sciences, USA, 2023
- Tracing metabolic flux through time and space with isotope labeling experiments.. Current Opinion in Biotechnology, 2020. [PubMed]
- Mass spectrometry-based metabolomics: a guide for annotation, quantification and best reporting practices.. Nature Methods, 2021. [PubMed]
- Imaging study by mass spectrometry of the spatial variation of cellulose and hemicellulose structures in corn stalks.. Journal of Agricultural and Food Chemistry, 2020. [PubMed]
- Vitamins in plants: occurrence, biosynthesis and antioxidant function.. Trends in Plant Science, 2010. [PubMed]
- Antioxidant role for lipid droplets in a stem cell niche of Drosophila.. Cell, 2015. [PubMed]
- Mass spectrometry imaging of the spatial and temporal localization of alkaloids in nightshades.. Journal of Agricultural and Food Chemistry, 2019. [PubMed]
- Aminopropyltransferases involved in polyamine biosynthesis localize preferentially in the nucleus of plant cells.. PLoS One, 2012. [PubMed]
- Fully automated unconstrained analysis of high-resolution mass spectrometry data with machine learning.. Journal of the American Chemical Society, 2022. [PubMed]
- MSiReader v1 0: evolving open-source mass spectrometry imaging software for targeted and untargeted analyses.. Journal of the American Society for Mass Spectrometry, 2017. [PubMed]
- Seeing plants as never before.. New Phytologist, 2023. [PubMed]
- Mass spectrometry imaging for plant biology: a review.. Phytochemistry Reviews, 2016. [PubMed]
- Mass spectrometry imaging: a review of emerging advancements and future insights.. Analytical Chemistry, 2018. [PubMed]
- The development and application of matrix assisted laser desorption electrospray ionization: the teenage years.. Mass Spectrometry Reviews, 2023. [PubMed]
- Mass spectrometry imaging as a potential technique for diagnostic of Huanglongbing disease using fast and simple sample preparation.. Scientific Reports, 2020. [PubMed]
- Phosphorus deprivation affects composition and spatial distribution of membrane lipids in legume nodules.. Plant Physiology, 2021. [PubMed]
- Image to insight: exploring natural products through mass spectrometry imaging.. Natural Product Reports, 2022. [PubMed]
- Sample preparation for mass spectrometry imaging of plant tissues: a review.. Frontiers in Plant Science, 2016. [PubMed]
- High mass resolution, spatial metabolite mapping enhances the current plant gene and pathway discovery toolbox.. New Phytologist, 2020. [PubMed]
- Analysis of Erythroxylum coca leaves by imaging mass spectrometry (MALDI–FT–ICR IMS).. Journal of the American Society for Mass Spectrometry, 2021. [PubMed]
- LDI and MALDI-FT-ICR imaging MS in Cannabis leaves: optimization and study of spatial distribution of cannabinoids.. Analytical Methods, 2019
- Visualizing vinca alkaloids in the petal of Catharanthus roseus using functionalized titanium oxide nanowire substrate for surface-assisted laser desorption/ionization imaging mass spectrometry.. The Plant Journal, 2021. [PubMed]
- Mass spectrometry imaging of phosphatidylcholine metabolism in lungs administered with therapeutic surfactants and isotopic tracers.. Journal of Lipid Research, 2021. [PubMed]
- Adhesive film applications help to prepare strawberry fruit sections for desorption electrospray ionization-mass spectrometry imaging.. Bioscience, Biotechnology, and Biochemistry, 2021. [PubMed]
- Exploring the diversity of plant metabolism.. Trends in Plant Science, 2019. [PubMed]
- New insights into red plant pigments: more than just natural colorants.. RSC Advances, 2020. [PubMed]
- Desorption electrospray ionization mass spectrometry imaging: recent developments and perspectives.. Journal of Biomolecular Techniques, 2019
- Recent advances in stable isotope-enabled mass spectrometry-based plant metabolomics.. Current Opinion in Biotechnology, 2017. [PubMed]
- Mass spectrometry-based quantification and spatial localization of small organic acid exudates in plant roots under phosphorus deficiency and aluminum toxicity.. The Plant Journal, 2021. [PubMed]
- The rhizosphere microbiome of ginseng.. Microorganisms, 2022. [PubMed]
- Applications of stable isotopes in MALDI imaging: current approaches and an eye on the future.. Analytical and Bioanalytical Chemistry, 2021. [PubMed]
- Spatially resolved metabolomics combined with bioactivity analyses to evaluate the pharmacological properties of two Radix puerariae species.. Journal of Ethnopharmacology, 2023. [PubMed]
- Biological explorations with nanoscale secondary ion mass spectrometry.. Journal of Analytical Atomic Spectrometry, 2019. [PubMed]
- Polyamines: bio-molecules with diverse functions in plant and human health and disease.. Frontiers in Chemistry, 2018. [PubMed]
- High-spatial resolution mass spectrometry imaging: toward single cell metabolomics in plant tissues.. Chemical Record, 2018. [PubMed]
- Plant Metabolic Network 15: a resource of genome-wide metabolism databases for 126 plants and algae.. Journal of Integrative Plant Biology, 2021. [PubMed]
- Comparing DESI-MSI and MALDI-MSI mediated spatial metabolomics and their applications in cancer studies.. Frontiers in Oncology, 2022. [PubMed]
- Biosynthesis of bioactive diterpenoids in the medicinal plant Vitex agnus-castus.. The Plant Journal, 2018. [PubMed]
- Elucidating drought-tolerance mechanisms in plant roots through 1H NMR metabolomics in parallel with MALDI-MS, and NanoSIMS imaging techniques.. Environmental Science & Technology, 2022. [PubMed]
- Visualization of soluble carbohydrate distribution in apple fruit flesh utilizing MALDI–TOF MS imaging.. Plant Science, 2019. [PubMed]
- Where do the electrons go? How numerous redox processes drive phytochemical diversity.. Phytochemistry Reviews, 2021
- The plant lipidome in human and environmental health.. Science, 2016. [PubMed]
- Lipidomics in situ: insights into plant lipid metabolism from high resolution spatial maps of metabolites.. Progress in Lipid Research, 2014. [PubMed]
- Spatial mapping of lipids at cellular resolution in embryos of cotton.. The Plant Cell, 2012. [PubMed]
- Imaging heterogeneity of membrane and storage lipids in transgenic Camelina sativa seeds with altered fatty acid profiles.. The Plant Journal, 2013. [PubMed]
- Mass spectrometry imaging for direct visualization of components in plants tissues.. Journal of Separation Science, 2021. [PubMed]
- Organic acids: the pools of fixed carbon involved in redox regulation and energy balance in higher plants.. Frontiers in Plant Science, 2016. [PubMed]
- Watercore in fruits. In:. Abiotic stress biology in horticultural plants., 2015
- Exploring uncharted territories of plant specialized metabolism in the postgenomic era.. Annual Review of Plant Biology, 2020
- A review on recent machine learning applications for imaging mass spectrometry studies.. Journal of Applied Physics, 2023
- Spatial metabolomics reveals potential biomarkers for red secretory cavities in Atractylodes lancea natural accessions.. Authorea doi:, 2022. [DOI]
- Visualizing the spatial distribution of functional metabolites in Forsythia suspensa at different harvest stages by MALDI mass spectrometry imaging.. Fitoterapia, 2022. [PubMed]
- Effective mechanisms for improving seed oil production in pennycress (Thlaspi arvense L) highlighted by integration of comparative metabolomics and transcriptomics.. Frontiers in Plant Science, 2022. [PubMed]
- The GORKY glycoalkaloid transporter is indispensable for preventing tomato bitterness.. Nature Plants, 2021. [PubMed]
- Maldi-mass spectrometry imaging for phytoalexins detection in RD6 Thai rice.. Applied Biochemistry and Microbiology, 2021
- MALDI MSI and Raman spectroscopy application in the analysis of the structural components and flavonoids in Brassica napus stem.. Metabolites, 2023. [PubMed]
- MALDI–HRMS imaging and HPLC–HRESI–MSn characterisation of kaurane diterpenes in the fruits of Xylopia aethiopica (Dunal) A Rich (Annonaceae).. Phytochemical Analysis, 2020. [PubMed]
- Use of mass spectrometry for imaging metabolites in plants.. The Plant Journal, 2012. [PubMed]
- Unveiling spatial metabolome of Paeonia suffruticosa and Paeonia lactiflora roots using MALDI MS imaging.. New Phytologist, 2021. [PubMed]
- Interrogation of spatial metabolome of Ginkgo biloba with high-resolution matrix-assisted laser desorption/ionization and laser desorption/ionization mass spectrometry imaging.. Plant, Cell & Environment, 2018
- Biofortified tomatoes provide a new route to vitamin D sufficiency.. Nature Plants, 2022. [PubMed]
- Acyl-lipid metabolism.. The Arabidopsis Book, 2013. [PubMed]
- How to sample a seizure plant: the role of the visualization spatial distribution analysis of Lophophora williamsii as an example.. Forensic Sciences Research, 2023. [PubMed]
- Matrix-assisted laser desorption/ionization mass spectrometry imaging (MALDI MSI) for in situ analysis of endogenous small molecules in biological samples.. Trends in Analytical Chemistry, 2022
- Metabolic profiling, in-situ spatial distribution, and biosynthetic pathway of functional metabolites in Dendrobium nobile stem revealed by combining UPLC-QTOF-MS with MALDI-TOF-MSI.. Frontiers in Plant Science, 2023. [PubMed]
- Visualizing the distribution of flavonoids in litchi (Litchi chinenis) seeds through matrix-assisted laser desorption/ionization mass spectrometry imaging.. Frontiers in Plant Science, 2023. [PubMed]
- Spatial localization of monoterpenoid indole alkaloids in Rauvolfia tetraphylla by high resolution mass spectrometry imaging.. Phytochemistry, 2023a. [PubMed]
- Leaves of Cannabis sativa and their trichomes studied by DESI and MALDI mass spectrometry imaging for their contents of cannabinoids and flavonoids.. Phytochemical Analysis, 2023b. [PubMed]
- Spatial analysis of lipid metabolites and expressed genes reveals tissue-specific heterogeneity of lipid metabolism in high- and low-oil Brassica napus L seeds.. The Plant Journal, 2018. [PubMed]
- Grapevine leaf MALDI-MS imaging reveals the localisation of a putatively identified sucrose metabolite associated to Plasmopara viticola development.. Frontiers in Plant Science, 2022. [PubMed]
- A derivatization and validation strategy for determining the spatial localization of endogenous amine metabolites in tissues using MALDI imaging mass spectrometry.. Journal of Mass Spectrometry, 2014. [PubMed]
- High-resolution multi-isotope imaging mass spectrometry (MIMS) imaging applications in stem cell biology.. Current Protocols, 2021. [PubMed]
- UPLC and ESI-MS analysis of metabolites of Rauvolfia tetraphylla L and their spatial localization using desorption electrospray ionization (DESI) mass spectrometric imaging.. Phytochemistry, 2019. [PubMed]
- Involvement of BGLU30 in glucosinolate catabolism in the Arabidopsis leaf under dark conditions.. Plant and Cell Physiology, 2020. [PubMed]
- Complex changes in membrane lipids associated with the modification of autophagy in Arabidopsis.. Metabolites, 2022. [PubMed]
- Extraction for metabolomics: access to the metabolome.. Phytochemical Analysis, 2014. [PubMed]
- Coupling APEX labeling to imaging mass spectrometry of single organelles reveals heterogeneity in lysosomal protein turnover.. Journal of Cell Biology, 2019
- Microscopic mass spectrometry imaging reveals the distribution of phytochemicals in the dried root of Isatis tinctoria.. Frontiers in Pharmacology, 2021. [PubMed]
- Desorption electrospray ionization mass spectrometry imaging illustrates the quality characters of Isatidis radix.. Frontiers in Plant Science, 2022. [PubMed]
- Transmission-mode MALDI-2 mass spectrometry imaging of cells and tissues at subcellular resolution.. Nature Methods, 2019. [PubMed]
- Mass spectrometry imaging of low molecular weight metabolites in strawberry fruit (Fragaria × ananassa Duch) cv Primoris with 109Ag nanoparticle enhanced target.. Phytochemistry, 2019. [PubMed]
- Plant FA db: a resource for exploring hundreds of plant fatty acid structures synthesized by thousands of plants and their phylogenetic relationships.. The Plant Journal, 2018. [PubMed]
- Visualizing genotypic and developmental differences of free amino acids in maize roots with mass spectrometry imaging.. Frontiers in Plant Science, 2020. [PubMed]
- Quinolizidine alkaloids are transported to seeds of bitter narrow-leafed lupin.. Journal of Experimental Botany, 2019. [PubMed]
- Toward sustainable production of value‐added bioenergy and industrial oils in oilseed and biomass feedstocks.. GCB Bioenergy, 2021
- Matrix-assisted laser desorption ionization imaging and laser ablation sampling for analysis of fungicide distribution in apples.. Analytical Chemistry, 2019. [PubMed]
- Pigman W. 2012. The carbohydrates: chemistry and biochemistry. Amsterdam: Elsevier.
- Recent advances in matrix-assisted laser desorption/ionisation mass spectrometry imaging (MALDI-MSI) for in situ analysis of endogenous molecules in plants.. Phytochemical Analysis, 2018. [PubMed]
- Plants secondary metabolites: the key drivers of the pharmacological actions of medicinal plants. In:. Herbal medicine., 2018
- Unveiling the spatial distribution of aflatoxin B1 and plant defense metabolites in maize using AP-SMALDI mass spectrometry imaging.. The Plant Journal, 2021. [PubMed]
- Mass spectrometry imaging disclosed spatial distribution of defense-related metabolites in Triticum spp.. Metabolites, 2022. [PubMed]
- Analyzing mass spectrometry imaging data of 13C-labeled phospholipids in Camelina sativa and Thlaspi arvense (Pennycress) embryos.. Metabolites, 2021. [PubMed]
- Spermidine and other functional phytochemicals in soybean seeds: spatial distribution as visualized by mass spectrometry imaging.. Food Science & Nutrition, 2020. [PubMed]
- Production of tocotrienols in seeds of cotton (Gossypium hirsutum L) enhances oxidative stability and offers nutraceutical potential.. Plant Biotechnology Journal, 2021. [PubMed]
- High-mass-resolution MALDI mass spectrometry imaging reveals detailed spatial distribution of metabolites and lipids in roots of barley seedlings in response to salinity stress.. Metabolomics, 2018. [PubMed]
- Using mass spectrometry imaging to map fluxes quantitatively in the tumor ecosystem.. Nature Communications, 2023
- Localization analysis of multiple vitamins in dried persimmon (Diospyros kaki) using matrix-assisted laser desorption/ionization mass spectrometry imaging.. Journal of Oleo Science, 2020. [PubMed]
- Imaging of multiple plant hormones in roots of rice (Oryza sativa) using nanoparticle-assisted laser desorption/ionization mass spectrometry.. Journal of Agricultural and Food Chemistry, 2020. [PubMed]
- Singh A , PrasadSM, SinghRP. 2016. Plant responses to xenobiotics. Singapore: Springer.
- In situ localization of plant lipid metabolites by matrix-assisted laser desorption/ionization mass spectrometry imaging (MALDI-MSI).. Methods in Molecular Biology, 2021. [PubMed]
- Three-dimensional visualization of membrane phospholipid distributions in Arabidopsis thaliana seeds: a spatial perspective of molecular heterogeneity.. Biochimica et Biophysica Acta, 2017. [PubMed]
- Matrix assisted laser desorption/ionization-mass spectrometry imaging (MALDI-MSI) for direct visualization of plant metabolites in situ.. Current Opinion in Biotechnology, 2016. [PubMed]
- The genome of jojoba (Simmondsia chinensis): a taxonomically isolated species that directs wax ester accumulation in its seeds.. Science Advances, 2020. [PubMed]
- Tissue-specific differences in metabolites and transcripts contribute to the heterogeneity of ricinoleic acid accumulation in Ricinus communis L (castor) seeds.. Metabolomics, 2019. [PubMed]
- In situ spatiotemporal mapping of 3-hydroxy-3-methylglutaryl-CoA reductase (HMGCR) inhibitor in pineapple (Ananas comosus) fruit tissue by MALDI mass spectrometry imaging.. Frontiers in Natural Products, 2023. [DOI]
- Visualizing the spatial distribution and alteration of metabolites in continuously cropped Salvia miltiorrhiza Bge using MALDI-MSI.. Journal of Pharmaceutical Analysis, 2022. [PubMed]
- Visualizing the distributions and spatiotemporal changes of metabolites in Panax notoginseng by MALDI mass spectrometry imaging.. Journal of Ginseng Research, 2021. [PubMed]
- MALDI imaging assisted discovery of a di-O-glycosyltransferase from Platycodon grandiflorum root.. Angewandte Chemie, 2023. [PubMed]
- FT-ICR mass spectrometry imaging at extreme mass resolving power using a dynamically harmonized ICR cell with 1ω or 2ω detection.. Analytical Chemistry, 2022. [PubMed]
- Biosynthesis-based spatial metabolome of Salvia miltiorrhiza Bunge by combining metabolomics approaches with mass spectrometry-imaging.. Talanta, 2022. [PubMed]
- Editorial: amino acids in plants: regulation and functions in development and stress defense.. Frontiers in Plant Science, 2021. [PubMed]
- Quantitative mass spectrometry imaging of biological systems.. Annual Review of Physical Chemistry, 2021
- Tailoring seed oil composition in the real world: optimising omega-3 long chain polyunsaturated fatty acid accumulation in transgenic Camelina sativa.. Scientific Reports, 2017. [PubMed]
- Preserved and variable spatial–chemical changes of lipids across tomato leaves in response to central vein wounding reveals potential origin of linolenic acid in signal transduction cascade.. Plant-Environment Interactions, 2021. [PubMed]
- Unsupervised machine learning for exploratory data analysis in imaging mass spectrometry.. Mass Spectrometry Reviews, 2020. [PubMed]
- Visualizing the distribution of strawberry plant metabolites at different maturity stages by MALDI-TOF imaging mass spectrometry.. Food Chemistry, 2021. [PubMed]
- In-situ detection and imaging of Areca catechu fruit alkaloids by MALDI-MSI.. Industrial Crops and Products, 2022
- Nanoparticle-immersed paper imprinting mass spectrometry imaging reveals uptake and translocation mechanism of pesticides in plants.. Nano Research, 2020
- A mass spectrometry imaging approach on spatiotemporal distribution of multiple alkaloids in Gelsemium elegans.. Frontiers in Plant Science, 2022. [PubMed]
- Unveiling the spatial distribution and molecular mechanisms of terpenoids biosynthesis in Salvia miltiorrhiza and S. grandifolia using multi-omics and DESI-MSI.. Horticulture Research, 2023. [PubMed]
- Visualizing the distribution of phthalate esters and plant metabolites in carrot by matrix-assisted laser desorption/ionization imaging mass spectrometry.. Journal of Agricultural and Food Chemistry, 2022. [PubMed]
- Recent advances in ambient mass spectrometry imaging.. Trends in Analytical Chemistry, 2019
- Mass spectrometry imaging techniques: a versatile toolbox for plant metabolomics.. Trends in Plant Science, 2022. [PubMed]
- Mechanisms and functions of membrane lipid remodeling in plants.. The Plant Journal, 2021. [PubMed]
- Identification of bottlenecks in the accumulation of cyclic fatty acids in Camelina seed oil.. Plant Biotechnology Journal, 2018. [PubMed]
- Localization of major Ephedra alkaloids in whole aerial parts of Ephedrae herba using direct analysis in real time-time of flight-mass spectrometry.. Molecules, 2021. [PubMed]
- MALDI-TOF/TOF tandem mass spectrometry imaging reveals non-uniform distribution of disaccharide isomers in plant tissues.. Food Chemistry, 2021. [PubMed]
- Mass spectrometry imaging and single-cell transcriptional profiling reveal the tissue-specific regulation of bioactive ingredient biosynthesis in Taxus leaves.. Plant Communications, 2023. [PubMed]
- Chemical imaging reveals diverse functions of tricarboxylic acid metabolites in root growth and development.. Nature Communications, 2023
- Imaging mass spectrometry reveals tumor metabolic heterogeneity.. Iscience, 2020. [PubMed]
- Tracking spatial distribution alterations of multiple endogenous molecules during lentil germination by MALDI mass spectrometry imaging.. Journal of Agricultural and Food Chemistry, 2023. [PubMed]
- Study on the distribution of low molecular weight metabolites in mango fruit by air flow-assisted ionization mass spectrometry imaging.. Molecules, 2022. [PubMed]
- Advances in MALDI mass spectrometry imaging single cell and tissues.. Frontiers in Chemistry, 2022. [PubMed]
