Prognostic and clinicopathological significance of TRIM21 in various cancers: A meta and bioinformatic analysis
Abstract
Methods:
We performed a systematic literature retrieval in various electronic databases including PubMed, Embase, Web of Science, Wanfang and China National Knowledge Infrastructure. Besides, the hazard ratio (HR) and the pooled relative risk (RR) were integrated in the assessment of cancer incidence and cancer mortality by Stata SE15.1. Additionally, we used an online database based on The Cancer Genome Atlas (TCGA) to further validate our results.
Results:
A total of 17 studies were included, totaling 7239 participants. High expression of TRIM21 was significantly correlated with better OS (HR = 0.74; 95% CI: 0.57–0.91; P < .001) and progression-free survival (PFS) (HR = 0.66; 95% CI: 0.42–0.91; P < .001). We found that high TRIM21 expression predicted significant impact on clinical characteristics like decreased lymph node metastasis (RR = 1.12; 95% CI: 0.97–1.30; P < .001), tumor stage (RR = 1.06; 95% CI: 0.82–1.37; P < .001) and tumor grade (RR = 1.07; 95% CI: 0.56–2.05; P < .001). However, TRIM21 expression had no significant impact on other clinical characteristics such as age (RR = 1.06; 95% CI: 0.91–1.25; P = .068), sex (RR = 1.04; 95% CI: 0.95–1.12; P = .953), or tumor size (RR = 1.14; 95% CI: 0.97–1.33; P = .05). Based on the Gene Expression Profiling Interactive Analysis (GEPIA) online analysis tool, TRIM21 was significantly downregulated in 5 cancers while significantly upregulated in 2 cancers, and the descending expression of TRIM21 predicted shorter OS in 5 cancers, worse PFS in 2 malignancies, while the elevated expression of TRIM21 predicted shorter OS and worse PFS in 2 carcinomas.
Conclusions:
TRIM21 could serve as a new biomarker for patients with solid malignancies and could be a potential therapeutic target for patients.
Article type: Research Article
Keywords: bioinformatic analysis, meta-analysis, neoplasms, prognosis, TRIM21
Affiliations: a Department of General Surgery, The Second Affiliated Hospital of Nanchang University, Nanchang, Jiangxi, People’s Republic of China; b The First School of Clinical Medicine of Nanchang University, Jiangxi Medical College of Nanchang University, Nanchang, Jiangxi, People’s Republic of China; c The Second School of Clinical Medicine of Nanchang University, Jiangxi Medical College of Nanchang University, Nanchang, Jiangxi, People’s Republic of China.
License: Copyright © 2023 the Author(s). Published by Wolters Kluwer Health, Inc. CC BY 4.0 This is an open access article distributed under the Creative Commons Attribution License 4.0 (CCBY), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Article links: DOI: 10.1097/MD.0000000000034012 | PubMed: 37335642 | PMC: PMC10256428
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (1.5 MB)
1. Introduction
Cancer refers to one of the leading causes of death globally that has conquered researchers’ attention for decades. According to the latest statistics, worldwide, an estimated 19.3 million new cancer cases and almost 10.0 million cancer deaths occurred in 2020.[1] With the advance of research process, many tumor markers have been proven to play an important role in various cancers, but only a few have been used in clinical practice. Therefore, identifying valuable tumor biomarkers is essential for both cancer screening and the prognosis.
Tripartite motif containing protein 21 (TRIM21), also named as Ro52,[2] is a member of the ubiquitin ligase group belonging to RING family which can be featured by the existence of a tripartite motif.[3] Multiple studies have shown that the expression level of TRIM21 is reduced in the poor prognosis and the deterioration of malignancies, such as diffuse large B-cell lymphoma,[2] renal cell cancer,[4] hepatic cell cancer,[5] breast cancer,[6,7] and ovarian cancer.[8] By contrast, in glioma[9] and pancreatic cancer,[10] patients with poor prognosis, the TRIM21 level is elevated, which indicates the additional function of TRIM21 in diverse types of cancers. Though present articles have manifested the relationship between TRIM21 and several malignancies, the significance of TRIM21 in prognosis has not been evaluated in meta-analysis. Therefore, we systematically collected and analyzed data from published literature to reveal the relationship between TRIM21 expression and overall survival (OS), progression-free survival (PFS), and clinicopathological features in cancer patients to assess the prognostic value of the possibly new biomarker TRIM21 in diverse types of cancers.
2. Materials and methods
2.1. Literature retrieval strategy
Web of Science, PubMed, Embase, China National Knowledge Infrastructure, and Wanfang databases are included in the scoepe of systematic computer literature retrieval. The supporting information are shown in the table, using the keywords: (“TRIM21” OR “Tripartite motif containing protein 21” OR “ubiquitin ligase”) AND (“cancer” OR “tumor” OR “neoplasm” OR “carcinoma” OR “malignancy”) AND “prognosis.” Besides, we also performed a manual screen to obtain potentially relevant studies.
2.2. Selection criteria
Eligible materials were selected into this meta-analysis based on the following criteria: the correlation between TRIM21 expression and OS and disease-free survival (DFS) in cancer patients; provided clinicopathological parameters; studies divided patients into 2 or more groups based on TRIM21 expression level; reported hazard ratio (HR) with 95% confidence interval (CI) or given sufficient data to estimate the HR with 95% CI. The exclusion criteria were: studies with insufficient data; letters, reviews, case reports, overlapping data, or expert opinions.
2.3. Date extraction and quality assessment
The extracted data and information were as follows: the first author name, year of publication, country, cancer type, number of patients, gender, method of assessing TRIM21 expression, cutoff value, follow-up times (months), outcome measures, analysis type, clinicopathological characteristics, HRs with 95% CI, prognostic value (OS, PFS), and P values. If the studies only provide the survival curve in figures, the estimated survival data were extracted by Engauge Digitizer.
2.4. Validation of bioinformatics database
Gene Expression Profiling Interactive Analysis (GEPIA), a web-based tool to deliver fast and customizable functionalities based on TCGA and GTEx data, was used. Survival plots of the correlation between TRIM21 expression and OS or DFS were retrieved as Kaplan–Meier curves based on different cancer datasets from the GEPIA online database. Median was set for cutoff value. Differential expression analysis between cancer and normal tissues was conducted based on GEPIA online analysis. All P value < .05 was regarded as statistically significant.
2.5. Statistical methods
All statistical analyses were carried out by Stata SE12.0. HRs with 95% CI for OS and DFS and relative risks (RRs) for clinicopathological parameters are included in the calculation. Heterogeneity was assessed among studies using χ2 tests and I2 statistics. We applied a random effects model to pool studies because of significant heterogeneity, as determined by the inconsistency index (I2 ≥ 50%) and χ2-test (P ≤ .10). The publication bias was evaluated by Begg funnel plot and Egger test, in which case P < .05 was considered significant.
3. Results
3.1. Main information of the enrolled studies
In total, 764 studies were retrieved after searching keywords on PubMed, Web of Science, Embase, China National Knowledge Infrastructure, and Wanfang databases. Among these, 310 studies were removed as duplicates. A further 333 were excluded after screening the title and abstract, being studies not regarding cancers, studies performed on animals, review papers, or studies not evaluating the TRIM21 gene. A total of 84 studies were successively excluded for being unable to view full text and no available data were reported. Finally, 17 studies[2,4–9,11–20] were included with total 7239 patients in this meta-analysis (Fig. 1). The flow chart maps out the number of studies identified, screened, included, and excluded.
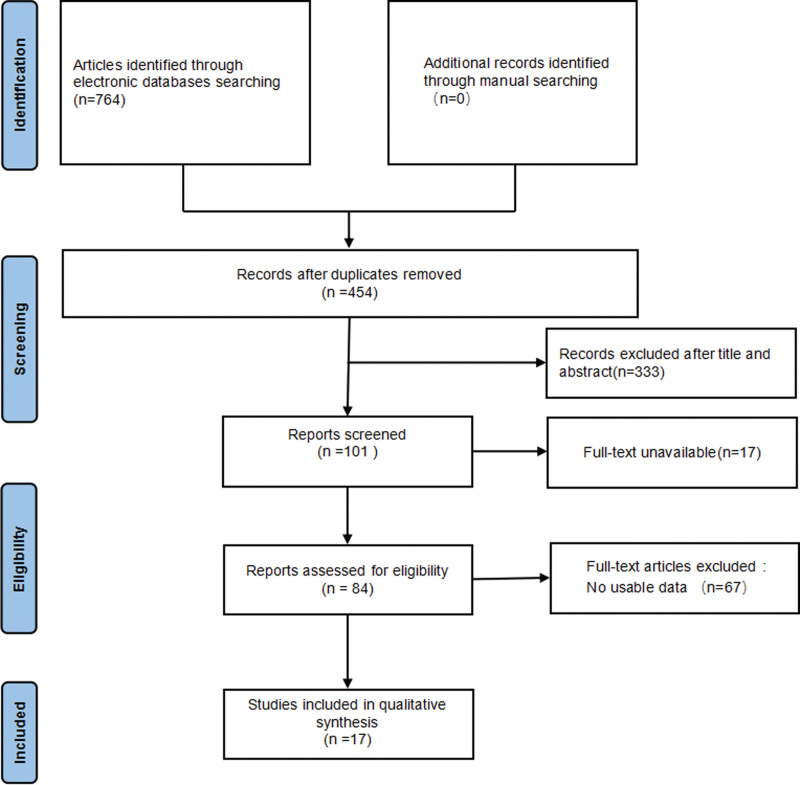
The malignant neoplasms assessed in these studies included breast cancer, ovarian cancer, HCC (hepatocellular carcinoma), gastric cancer, pancreatic cancer, renal cell carcinoma, NSCLS (non-small cell lung cancer), HNSCC (head and neck squamous cell carcinoma), diffuse large B-cell lymphoma, and glioma. Sixteen studies were from China and 1 from Sweden. Ten were used to evaluate the HR of OS for cancer patients and 6 were used to evaluate the HR of both OS and DFS. The expression levels of TRIM21 gene were evaluated by immunohistochemistry in 10 studies, while several other studies adopted western blotting and Quantitative real-time PCR (qRT-PCR). The main information of the enrolled studies can be found in Table 1.
Table 1: Main characteristics of studies included in the meta-analysis.
| Study | Country | Cancer type | Sample size | Follow-up | Tumor stage | Outcome measures | Analysis type | Gender (f/m) | NOS |
|---|---|---|---|---|---|---|---|---|---|
| Zhang ZH (2022) | China | HCC | 198 | 167 | NA | OS | U | NA | 7 |
| Zhang J (2021) | China | NSCLC | 132 | NA | 100/32 (1/2–3) | NA | U | 31/35 | 7 |
| Yan G (2021) | China | RCC | 157 | 150 | NA | OS | U | NA | 6 |
| Xue CF (2017) | China | Ovarain cancer | 1656 | 240 | NA | OS | U | 1656/0 | 6 |
| Wang JL (2021) | China | GC | 80 | 36 | 33/28/19 (2/3/4) | OS | U | 23/57 | 7 |
| Wang JL (2022) | China | Pancreatic cancer | 87 | 36 | 27/26/27/7 | OS | U | 32/55 | 7 |
| He D (2016) | China | Ovarain cancer | 90 | NA | 42/48 | OS | U | 90/0 | 7 |
| Zhou W (2018) | China | Breast cancer | 169 | 74 1/2 | 67/87/15 | OS/DFS | U/M | 169/0 | 8 |
| Zhao Z (2020) | China | Glioma | 246 | NA | NA | OS/DFS | U/M | NA | 8 |
| Sun J (2022) | China | Ovarain cancer | 558 | 200 | 410/84/70/24 | OS/DFS | U | 558/0 | 6 |
| Si W (2020) | China | Brest cancer | NA | 300 | NA | OS | U | NA | 5 |
| Jin Y (2019) | China | Brest cancer | 2136 | NA | 1085/1051 | NA | NA | 2136/0 | 6 |
| Ding Q (2015) | China | HCC | 242 | 80 | 20/152/24/29 | OS/DFS | U | 31/211 | 8 |
| Dai W (2015) | China | HCC | 343 | 133 | NA | OS | U | NA | 6 |
| Chuang C (2021) | China | HNSCC | 518 | 200 | NA | OS | U | NA | 7 |
| Chen X (2021) | China | RCC | 246 | 60 | NA | OS/DFS | U | 44/79 | 6 |
| Brauner S (2015) | Sweden | Lymphoma | 381 | 100 | NA | OS/DFS | U | NA | 7 |
DFS = disease-free survival, GC = gastric cancer, HCC = hepatocellular carcinoma, HNSCC = head and neck aquamous cell carcinoma, NSCLC = non-small cell lung cancer, OS = overall survival, RCC = renal cell carcinoma.
3.2. Correlation of TRIM21 expression level with OS
Data from the 17 studies were used for OS analysis with a random-effects model because of significant heterogeneity (I2 = 93.7%, P < .001). From the result, we could observe that compared with low TRIM21 expression, the high level of TRIM21 may predict better OS, with the pooled HR being 0.74 (95% CI: 0.57–0.91; P < .001; Fig. 2).
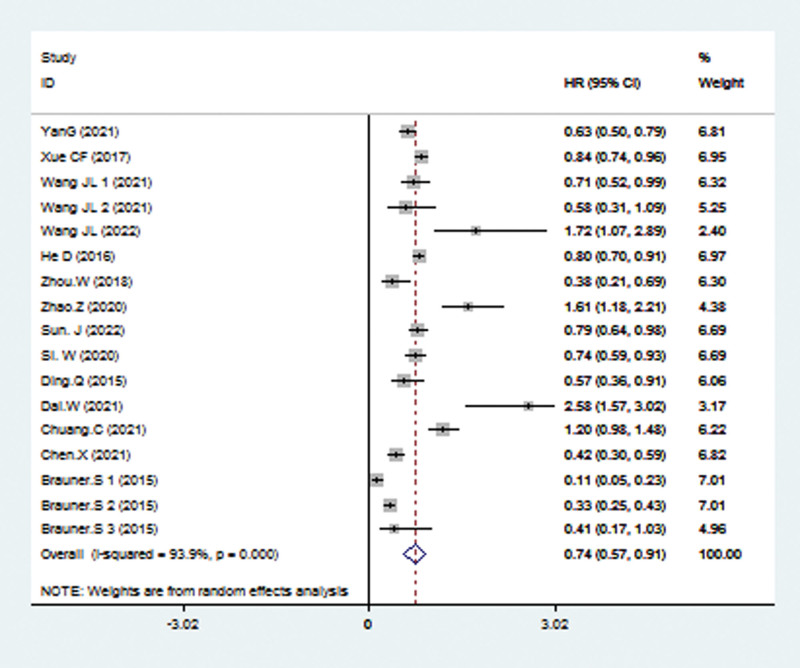
3.3. Subgroup analysis for OS
The included studies were analyzed by stratifying them according to subgroups of data source, pathology type, detection method, and tumor sites. Stratified analysis according to different tumor sites revealed a significant association between the high level of TRIM21 and better OS in reproductive system tumors (random effects model: combined HR = 0.74; 95% CI: 0.62–0.86; P = .015) and other systems (random effects model: combined HR = 0.50; 95% CI: 0.27–0.72; P < .001; Fig. 3). While a significant association between the high level of TRIM21 and worse OS was found in digestive system cancer (random effects model: combined HR = 1.10; 95% CI: 0.69–1.52; P < .001; Fig. 3); subgroup analysis according to pathological type showed a combined HR of 0.74 (95% CI: 0.63–0.85; P = .013) for adenocarcinoma and pooled HR (HR = 0.74; 95% CI: 0.49–1.00; P < .001) for another group (Supplementary Fig. 1, http://links.lww.com/MD/J126). In subgroup analysis data source (Supplementary Fig. 2, http://links.lww.com/MD/J127), a significantly better OS was observed both in reported data group (random-effects model: pooled HR = 0.79; 95% CI: 0.64–0.93; P < .001) and in data extracted from survival curve (random-effects model: pooled HR = 0.66; 95% CI: 0.43–0.89; P < .001). Finally, subgroup analysis according to different assay method showed that a high level of TRIM21 was associated with a significantly better OS in immunohistochemistry methods group (random-effects model: pooled HR = 0.50; 95% CI: 0.33–0.66; P < .001) and other group (random-effects model: pooled HR = 0.90; 95% CI: 0.061–1.20; P < .001) while worse OS in qRT-PCR group (random-effects model: pooled HR = 1.44; 95% CI: 0.90–1.99; P < .001) (Supplementary Fig. 3, http://links.lww.com/MD/J128).
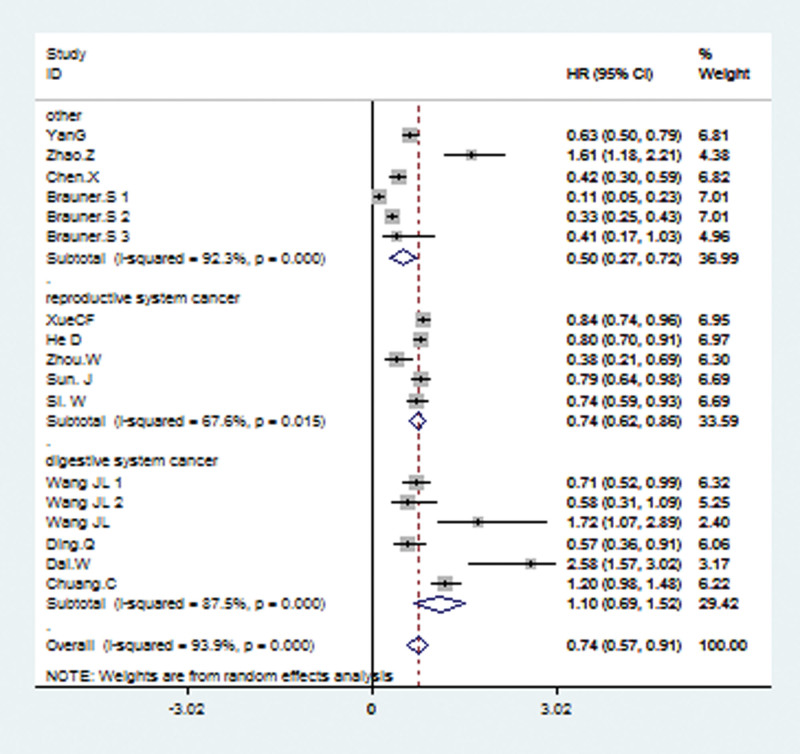
3.4. Correlation of TRIM 21 expression level with PFS
HR for PFS was used in 6 studies, including 1594 patients The pooled HR confirmed a significant correlation between high TRIM21 gene and better PFS (HR = 0.66; 95% CI: 0.42–0.91; P < .001; Fig. 4). However, substantial heterogeneity was observed across studies, so the random-effects model was applied (I2 = 81.9; P < .001).
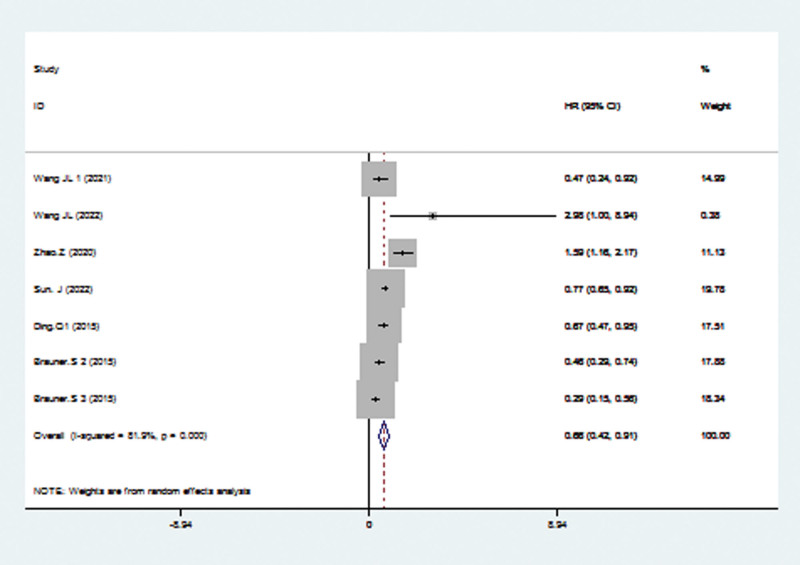
3.5. Association between the expression of the TRIM21 gene and clinical parameters
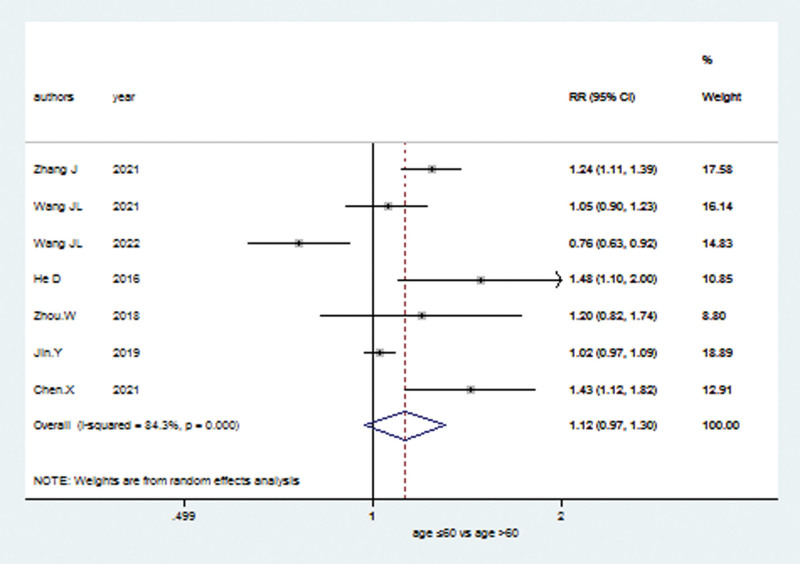
The relationship between the expression of TRIM21 gene and clinical outcomes were delineated in the meta-analysis. The information of Clinical Parameters was shown with pooled RRs and 95% CI (Table 2). We found that high TRIM21 expression predicted significant impact on clinical characteristics like lymph node metastasis (RR = 1.12; 95% CI: 0.97–1.30; P < .001; Fig. 5), tumor stage (RR = 1.06; 95% CI: 0.82–1.37; P < .001, random effects model; Supplementary Fig. 4, http://links.lww.com/MD/J129) and grade (RR = 1.07; 95% CI: 0.56–2.05; P < .001, random effects model; Supplementary Fig. 5, http://links.lww.com/MD/J130). However, no significant correlation was observed between high TRIM21 level and age (RR = 1.06; 95% CI: 0.91–1.25; P = .068, random effects model; Supplementary Fig. 6, http://links.lww.com/MD/J131), sex (RR = 1.04; 95% CI: 0.95–1.12; P = .953; Supplementary Fig. 7, http://links.lww.com/MD/J132), or tumor size (RR = 1.14; 95% CI: 0.97–1.33; P = .05, random effects model; Supplementary Fig. 8, http://links.lww.com/MD/J133).
Table 2: RRs and 95% CIs for patient survival or disease progression in association with TRIM21 expression in enrolled studies.
| Heterogeneity | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Parameters | Studies (n) | Number of patients (n) | RR | LCI | UCI | P | I2 | Ph | Model |
| Age | 6 | 3434 | 1.06 | 0.91 | 1.25 | .068 | 51.40% | 0.45 | Random |
| Sex | 5 | 787 | 1.04 | 0.95 | 1.11 | .95 | 0.00% | 0.41 | Fixed |
| Lymph node metastasis | 7 | 940 | 1.12 | 0.97 | 1.30 | <.001 | 84.30% | 0.13 | Random |
| Tumor size | 5 | 876 | 1.14 | 0.97 | 1.33 | .05 | 57.90% | 0.11 | Random |
| Stage | 7 | 3282 | 1.06 | 0.82 | 1.37 | <.001 | 86.70% | 0.65 | Random |
| Tumor grade | 7 | 3282 | 1.07 | 0.56 | 2.1 | <.001 | 96.10% | 0.83 | Random |
RR = relative risk, TRIM21 = Tripartite motif-containing protein 21.
3.6. Publication bias and sensitivity analysis
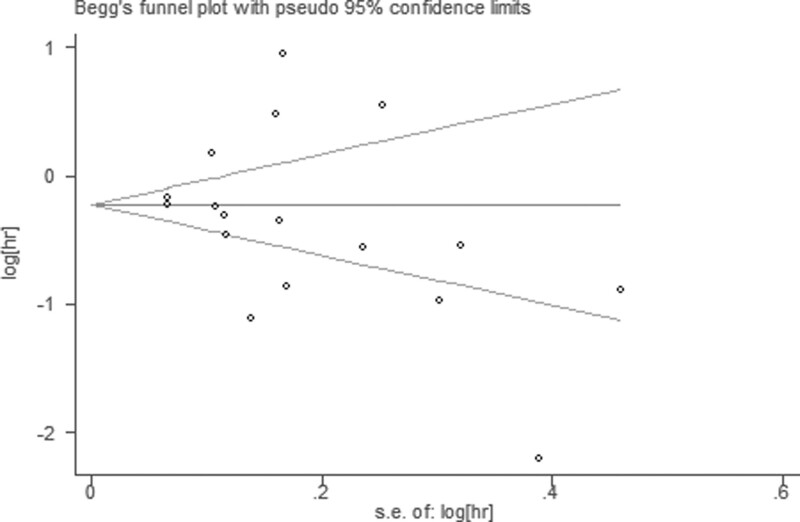
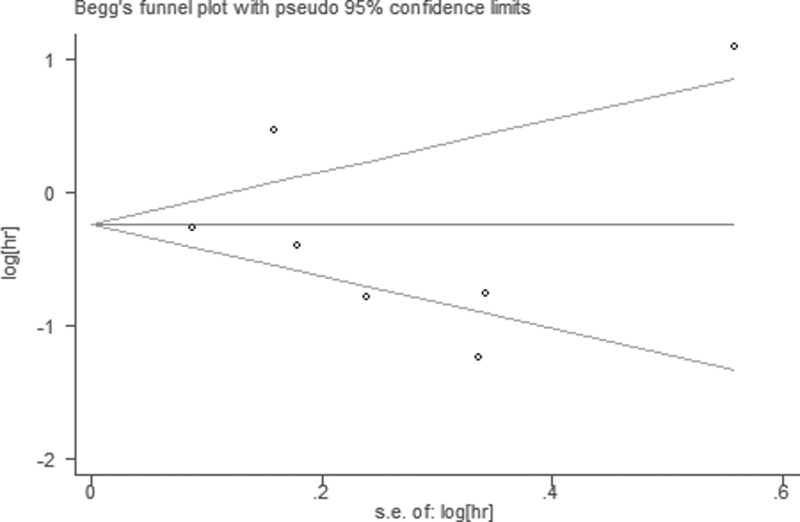
Publication bias was evaluated by Begg funnel plot and Egger test. Among 17 cohorts evaluating OS, no obvious asymmetry was observed in Begg funnel plots (Z = 1.77; Pr >|Z| = 0.077) and the Egger P value (P = .396; Fig. 6), but asymmetry was found among 7 cohorts evaluating PFS (Z = 0.60; Pr > |Z| = 0.548) and the Egger P value (P = .752; Fig. 7).
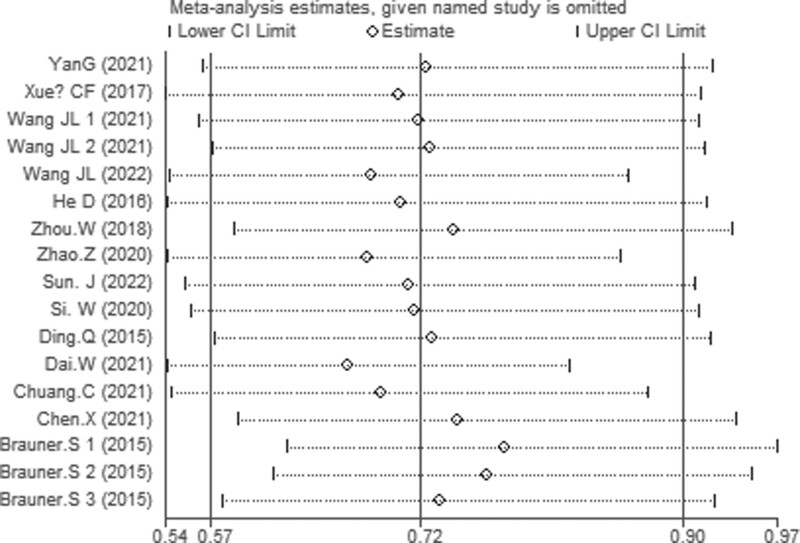
A sensitivity analysis was applied to verify its accuracy of pooled HR of OS. The findings revealed that excluding any single research had no important effect on pooled HR, indicating the result relative (Fig. 8).
3.7. Validation of the results in the GEPIA database
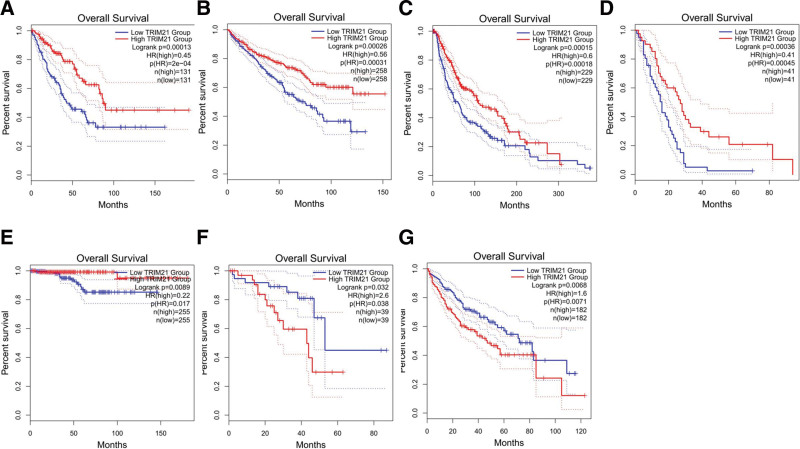
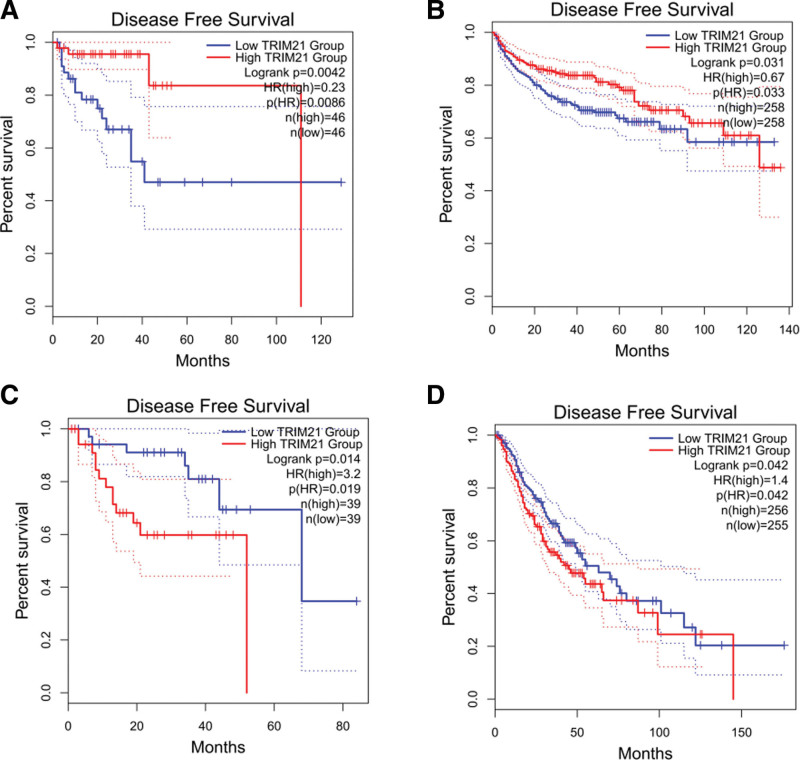
GEPIA online analysis tool was adopted to further strengthen our conclusion (http://gepia2.cancer-pku.cn/). In terms of TRIM21 dysregulation, low TRIM21 expression was identified in KIRC (Kidney renal clear cell carcinoma), MESO (Mesothelioma), SARC (Sarcoma), SKCM (Skin Cutaneous Melanoma), READ (Rectum adenocarcinoma) and THCA (Thyroid carcinoma). Regarding the association between and prognosis, decreased TRIM21 expression was correlated with worse OS in KIRC, MESO, SARC, SKCM, THCA (Fig. 9) and with worse PFS in KIRC and READ (Fig. 10). These results support our results and indicate that TRIM21 could be a novel prognostic biomarker for various cancers.
4. Discussion
TRIM21 gene is located on the short arm of chromosome 11 (11p15.5), the locus associated with malignancies in many organs. Previous studies have indicated that TRIM21 play important roles in inflammation,[21] autoimmunity and cancer. TRIM21 was first discovered as an auto antigen associated with autoimmune diseases, such as SLE (Systemic Lupus Erythematosus).[22] The up-regulation of TRIM21 can reduce cell proliferation and promote cell apoptosis in SLE, which indicated the TRIM21 could regulate the cell proliferation and apoptosis.[23] After that, studies have shown pathogen infection such as viruses and bacteria can upregulate TRIM21 expression and further achieve its effective anti-pathogen protection by activating related transcription factor pathways, such as NF-κB, activator protein 1 (AP-1), interferon regulators IRF3, IRF5, and IRF7.[24]
Besides, the role of TRIM21 in various cancers has been observed in many studies, but the conclusions are inconsistent. These inconsistent outcomes might hint at dissimilar potential mechanisms that affect cancer recurrence. Like other E3 ligases, TRIM21 functions by ubiquitination of target substrates.[25] A high expression of TRIM21 arrests the HCC cells cycle and inhibits cells proliferation by promoting ATG4B ubiquitylation,[26] and suppresses the invasion of HCC cells by promoting β-catenin ubiquitylation and degradation.[27] TRIM21 negatively regulates the NF-κB which is known as lymphoma proliferation and survival factor by the ubiquitylation and translocation of IKKβ to autophagosomes.[28,29] Therefore, it might be the mechanism of TRIM21. High TRIM21 expression is associated with better prognosis in patients with diffuse large B-cell lymphoma (DLBCL).[2] By contrast, high TRIM21 levels were associated with lower OS and poor prognosis in pancreatic cancer patients.[10] The specific physiological role and detailed molecular mechanism by which TRIM21 contributes to tumor cells remain unclear. The exact functions of TRIM21 have not been fully understood and there are a series of mysteries that need to be explored and studied.
The results of the subgroup analysis showed that the relationship between TRIM21 expression levels and prognosis of patients was influenced by a variety of factors, including tumor types, the ethnicity of the study population, the method of detection and the source of the data analysis. Firstly, we performed subgroup analysis based on the different systemic origins of the tumors. The results showed that high levels of TRIM21 expression were significantly associated with better prognosis in patients with reproductive tumors or other cancers, but worse OS with digestive system cancers. In light of this, the prognostic value of TRIM21 may vary in different cancers because of different mechanisms of TRIM21. Secondly, we performed a subgroup analysis based on different pathological types to investigate the impact of TRIM21 expression levels on patient prognosis. The analysis showed that high levels of TRIM21 were significantly associated with better OS in adenocarcinoma, suggesting that detecting TRIM21 expression levels in patients with adenocarcinoma may help to predict their prognosis.
Our meta-analysis comprehensively and systematically reviewed the prognostic value of TRIM21 in various cancers. In this study, OS analysis revealed a pooled HR of 0.74, demonstrating that high TRIM21 expression is related with a better prognosis; however, the heterogeneity is significant (I2 = 93.7%; P < .001). PFS analysis also showed that increased TRIM21 expression is associated with better outcome significantly (HR = 0.76; P < .05). In addition, increased TRIM21 expression is negatively associated with lymph node metastasis with pooled RR (RR = 1.12; 95% CI: 0.97–1.30; P < .001), but no significant correlation was observed between TRIM21 and age, sex, tumor size, tumor grade or stage. GEPIA and TCGA databases were further used to validate our results as broadly as possible. Low TRIM21 expression levels were observed in KIRC, MESO, SARC, SKCM, READ, and THCA. What more, decreased TRIM21 expression was associated with worse OS in KIRC, MESO, SARC, SKCM, THCA, and with worse PFS in KIRC, READ. However, high TRIM21 expression levels were observed in LIHC (Liver hepatocellular carcinoma) and UVM (Uveal Melanoma), and elevated TRIM21 was associated with worse OS and PFS in these 2 cancers. Taken together, the research of TRIM21 may provide new ideas for the diagnosis and treatment of malignant tumors, and the research as a targeted treatment site of malignant tumors has achieved initial success. TRIM21 will bring new prospects for the clinical treatment of tumors.
There is no denying that several limitations existed in this meta-analysis. Firstly, the included tumor types had a narrow distribution range, mainly focusing on few cancer studies, which could not well represent the whole tumors. Secondly, the sample size was evenly distributed, and the difference in sample size would have some impact on HR. Thirdly, the TRIM21 expression level intercept was not uniform, and the difference between the low TRIM21 expression and the high TRIM21 expression would increase the heterogeneity of this study. Fourthly, the HR of some studies was extracted by KM survival curve, and there are errors in the extraction process, which will further affect the results of HR. Fifthly, this meta-analysis includes some retrospective studies. Therefore, these factors should be considered when drawing a conclusion. Finally, this study was nearly constrained to studies published in China, so our results may best illustrate the association between TRIM21 and Asian patients.
In conclusion, TRIM21 may affect the prognosis of cancer in some extent. It might be an effective indicator for predicting prognosis and tumor progression in the future. More clinical studies should be carried out to get a more accurate evaluation of the prognostic role of TRIM21 in patients with cancers.
Author contributions
Conceptualization: Feng Hu, Yan Liu, Feiyang Wang, Xinyi Fu, Zhenhong Zou.
Data curation: Feng Hu, Yan Liu, Feiyang Wang, Xinyi Fu, Xiangjun Liu, Zhenhong Zou, Bin Zhou.
Formal analysis: Feng Hu, Yan Liu, Bin Zhou.
Writing – original draft: Feng Hu, Yan Liu, Feiyang Wang, Xinyi Fu, Xiangjun Liu.
Writing – review & editing: Feng Hu, Yan Liu, Zhenhong Zou, Bin Zhou.
Supplementary Materials
References
- H Sung, J Ferlay, RL Siegel. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries.. CA Cancer J Clin., 2021. [PubMed]
- S Brauner, W Zhou, C Backlin. Reduced expression of TRIM21/Ro52 predicts poor prognosis in diffuse large B-cell lymphoma patients with and without rheumatic disease.. J Intern Med., 2015. [PubMed]
- D Tomar, R Singh. TRIM family proteins: emerging class of RING E3 ligases as regulator of NF-κB pathway.. Biol Cell., 2015. [PubMed]
- X Chen, Z Li, H Yong. Trim21-mediated HIF-1α degradation attenuates aerobic glycolysis to inhibit renal cancer tumorigenesis and metastasis.. Cancer Lett., 2021. [PubMed]
- Q Ding, D He, K He. Downregulation of TRIM21 contributes to hepatocellular carcinoma carcinogenesis and indicates poor prognosis of cancers.. Tumour Biol., 2015. [PubMed]
- Y Jin, Y Zhang, B Li. TRIM21 mediates ubiquitination of Snail and modulates epithelial to mesenchymal transition in breast cancer cells.. Int J Biol Macromol., 2019. [PubMed]
- W Si, J Zhou, Y Zhao. SET7/9 promotes multiple malignant processes in breast cancer development via RUNX2 activation and is negatively regulated by TRIM21.. Cell Death Dis., 2020. [PubMed]
- J Sun, X Chen, X Ji. TRIM21 deficiency promotes cell proliferation and tumorigenesis via regulating p21 expression in ovarian cancer.. Bioengineered., 2022. [PubMed]
- Z Zhao, Y Wang, D Yun. TRIM21 overexpression promotes tumor progression by regulating cell proliferation, cell migration and cell senescence in human glioma.. Am J Cancer Res., 2020. [PubMed]
- JQ Nguyen, R Irby. TRIM21 is a novel regulator of Par-4 in colon and pancreatic cancer cells.. Cancer Biol Ther., 2017. [PubMed]
- CY Chuang, YC Chien, CW Lin. TRIM21 polymorphisms are associated with susceptibility and clinical status of oral squamous cell carcinoma patients.. Int J Med Sci., 2021. [PubMed]
- W Dai, J Wang, Z Wang. Comprehensive analysis of the prognostic values of the TRIM family in hepatocellular carcinoma.. Front Oncol., 2021. [PubMed]
- W Zhou, Y Zhang, C Zhong. Decreased expression of TRIM21 indicates unfavorable outcome and promotes cell growth in breast cancer.. Cancer Manag Res., 2018. [PubMed]
- D He. Expression and function of TRIM21 protein in ovarian cancer [D];, 2016
- JL Wang, ZY Yu, ZT Chen. Correlation of TRIM21 with clinicopathological features and immune infiltration in pancreatic cancer.. J Chongqing Med Univ., 2022
- JL Wang, ZY Yu, S Ade. Value of TRIM21 in the diagnosis and prognosis of gastric cancer.. Chin J Bases Clin Gen Surg., 2021
- CF Xue. The value of TRIM21 and TRIM24 expression in the prognosis of ovarian cancer [J].. World Latest Med Inf Dig., 2017
- G Yan, RY Wei, XT Chen. Effect of Trim21 on the proliferation, migration and invasion ability of kidney cancer cells and its mechanism.. J Xuzhou Med Univ., 2021
- J Zhang. The expression and clinical significance of triple domain protein 21 in non-small cell lung cancer [D]., 2021
- ZH Zhang, ZR Zhu, HL Sheng. TRIM21 inhibits the invasive ability of hepatocellular carcinoma cells: degradation of β-catenin based on the ubiquitination pathway.. J South Med Univ., 2022
- Y Ahn, JH Hwang, Z Zheng. Enhancement of Th1/Th17 inflammation by TRIM21 in Behçet’s disease.. Sci Rep., 2017. [PubMed]
- DA Rhodes, J Trowsdale. TRIM21 is a trimeric protein that binds IgG Fc via the B30.2 domain.. Mol Immunol., 2007. [PubMed]
- Y Kunishita, R Yoshimi, R Kamiyama. TRIM21 dysfunction enhances aberrant B-cell differentiation in autoimmune pathogenesis.. Front Immunol., 2020. [PubMed]
- WA Mcewan, JC Tam, RE Watkinson. Intracellular antibody-bound pathogens stimulate immune signaling via the Fc receptor TRIM21.. Nat Immunol., 2013. [PubMed]
- A Espinosa, W Zhou, M Ek. The Sjogren’s syndrome-associated autoantigen Ro52 is an E3 ligase that regulates proliferation and cell death.. J Immunol., 2006. [PubMed]
- T Li, LX Chen, LB Sun. Experimental study on the inhibition of hepatocellular carcinoma cell proliferation by triple structural domain protein 21 through downregulation of autophagy-related protein 4B.. J Third Mil Med Univ., 2019
- Z Zhang, Z Zhu, H Sheng. [TRIM21 suppresses invasion of hepatocellular carcinoma cells by promoting β-catenin ubiquitylation and degradation].. Nan Fang Yi Ke Da Xue Xue Bao., 2022. [PubMed]
- K Wada, M Niida, M Tanaka. Ro52-mediated monoubiquitination of IKK{beta} down-regulates NF-{kappa}B signalling.. J Biochem., 2009. [PubMed]
- S Balaji, M Ahmed, E Lorence. NF-κB signaling and its relevance to the treatment of mantle cell lymphoma.. J Hematol Oncol., 2018. [PubMed]
