Multi-omics analysis reveals the involvement of origin recognition complex subunit 6 in tumor immune regulation and malignant progression
Abstract
Background:
Origin recognition complex 6 (ORC6) is one of the six highly conserved subunit proteins required for DNA replication and is essential for maintaining genome stability during cell division. Recent research shows that ORC6 regulates the advancement of multiple cancers; however, it remains unclear what regulatory impact it has on the tumor immune microenvironment.
Methods:
Unpaired Wilcoxon rank sum and signed rank tests were used to analyze the differences in the expression of ORC6 in normal tissues and corresponding tumor tissues. Multiple online databases have evaluated the genetic alterations, protein expression and localization, and clinical relevance of ORC6. To evaluate the potential prognostic impact and diagnostic significance of ORC6 expression, we carried out log-rank, univariate Cox regression, and receiver operating characteristic curve analysis. The ICGC-LIRI-JP cohort, CGGA-301 cohort, CGGA-325 cohort, CGGA-693 cohort, and GSE13041 cohort were used for external validation of the study findings. The associations between ORC6 expression and immune cell infiltration, immune checkpoint expression, and immunotherapy cohorts was further analyzed. To explore the functional and signaling pathways related to ORC6 expression, gene set enrichment analysis was performed. To clarify the expression and function of ORC6 in hepatocellular carcinoma (LIHC) and glioma, we conducted in vitro experiments.
Results:
Expression of ORC6 is upregulated in the majority of cancer types and is associated with poor patient prognosis, notably in cases of LIHC and gliomas. In addition, ORC6 may be involved in multiple signaling pathways related to cancer progression and immune regulation. High expression of ORC6 correlates with an immunosuppressive state in the tumor microenvironment. The results of further immunotherapy cohort analysis suggested that patients in the ORC6 high-expression group benefited from immunotherapy. Inhibiting ORC6 expression suppressed the proliferative and migratory abilities of LIHC and glioma cells.
Conclusion:
High expression of ORC6 may be used as a biomarker to predict the poor prognosis of most tumor patients. The high expression of ORC6 may be involved in the regulation of the tumor immunosuppressive environment, and it is expected to become a molecular target for inhibiting tumor progression.
Article type: Research Article
Keywords: pan-cancer, prognosis, immunotherapy, tumor microenvironment
Affiliations: Department of General Surgery, The Second Xiangya Hospital, Central South University, Changsha, Hunan, China; Clinical Research Center For Breast Disease In Hunan Province, Changsha, Hunan, China
License: Copyright © 2023 Zhu, Chen, Zeng, Gao, Wu, He, Xu, Pang, Peng, Deng, Han and Yi CC BY 4.0 This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
Article links: DOI: 10.3389/fimmu.2023.1236806 | PubMed: 37901236 | PMC: PMC10602784
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (25.1 MB)
Introduction
Worldwide, cancer presents a life-threatening situation and is one of the most economically burdensome diseases (ref. 1). Currently, no treatment for cancer is absolutely effective. As research advances, scientists are increasingly concentrating on the shared characteristics of different malignant tumors to uncover their underlying causes and create targeted inhibitors for cancer therapy (ref. 2). For instance, PD-L1 levels are often increased in different cancer types, and recent studies indicate that many oncogenic signaling pathways lead to this overexpression. Antagonistic antibodies against the inhibitory immune checkpoint receptor PD-1 or its ligand PD-L1 have shown promise in the treatment of various cancers, leading to significant improvement in patient survival rates (ref. 3). Protein tyrosine kinases from the human epidermal growth factor receptor family, such as EGFR and HER2, are important therapeutic targets for many malignancies, including non-small cell lung cancer, breast cancer, and gastroesophageal cancer, particularly colorectal cancer (ref. 4). Aldehyde dehydrogenase (ALDH) serves as a cancer stem cell biomarker across various cancers. Clinically, ALDHs are also regarded as indicators of poor prognosis in solid cancers. Targeting ALDHs may impede cancer stem cells in solid tumors, thereby achieving therapeutic effects (ref. 5). Therefore, analyzing the differential expression of genes across cancers, screening valuable genes, and exploring their correlation with clinical prognosis and the tumor immune microenvironment will promote the further development of tumor-targeted therapy and immunotherapy.
The origin recognition complex (ORC) is a vital six-subunit protein that is highly conserved across species and plays a crucial role in DNA replication. It is essential for maintaining genome stability during cell division (ref. 6). ORC6, the smallest subunit of human ORC, is primarily involved in chromosome segregation, DNA replication, and cell division. It localizes to replication forks to carry out these functions (ref. 7). ORC6 is a cofactor in the mismatch repair (MMR) complex that promotes efficient mismatch repair (ref. 8). In recent years, research has identified a correlation between elevated ORC6 expression and adverse prognostic outcomes in patients with colorectal cancer (ref. 9), renal clear cell carcinoma (ref. 10), gastric adenocarcinoma (ref. 11), and breast cancer (ref. 12). Current research on ORC6 in tumors is restricted to specific types of human cancers. There has been no systematic multi-omics analysis across different types of cancer, notably in liver hepatocellular carcinoma (LIHC) and glioma (GBMLGG).
The research found that ORC6 was frequently overexpressed in various cancer types and was associated with adverse survival outcomes. Additionally, the biological function of ORC6 may be linked to RNA modifications, DNA methylation, and the tumor immune microenvironment. By examining ORC6 across cancers, we observed that it significantly contributes to the development of LIHC and GBMLGG. Our study reveals that ORC6 acts as an independent risk factor for the overall prognosis of LIHC and GBMLGG. Subsequently, we conducted in vitro experiments to elucidate whether ORC6 promotes the progression of LIHC and GBMLGG. Together, our investigation provides a comprehensive understanding of the tumorigenic role of ORC6 in different cancers and indicates that ORC6 could be a dependable biomarker for predicting the clinical prognoses and immune landscapes in patients with LIHC and GBMLGG.
Materials and methods
Data preprocessing and differential expression analysis
We obtained a unified and standardized pan-cancer dataset (TCGA TARGET GTEx, https://xenabrowser.net/) from the UCSC database. Furthermore, we extracted the ORC6 gene values from each sample and applied log2 (x+1) transformation for each value. The Sangerbox (ref. 13) online tool was used for visualization. In addition, we acquired validation cohorts from external sources, including the International Cancer Genome Consortium (ICGC), Chinese Glioma Genome Atlas Project (CGGA), and Gene Expression Omnibus (GEO) databases. These cohorts included the ICGC-LIRI-JP cohort, CGGA-301 cohort, CGGA-325 cohort, CGGA-693 cohort, and GSE13041 cohort. Cancer-type abbreviations are listed in T1.
Table 1: Tumor types and abbreviations.
| Abbreviation | Full name |
|---|---|
| ACC | Adrenocortical carcinoma |
| ALL | Acute Lymphoblastic Leukemia |
| AML | Acute myeloid leukemia |
| AST | Astrocytoma |
| BLCA | Bladder Urothelial Carcinoma |
| BRCA | Breast invasive carcinoma |
| CESC | Cervical squamous cell carcinoma and endocervical adenocarcinoma |
| CHOL | Cholangiocarcinoma |
| CML | Chronic myelogenous leukemia |
| COAD | Colon adenocarcinoma |
| COADREAD | Colon adenocarcinoma/Rectum adenocarcinoma |
| DLBC | Lymphoid Neoplasm Diffuse Large B-cell Lymphoma |
| ESCA | Esophageal carcinoma |
| GBM | Glioblastoma multiforme |
| GBMLGG | Glioma |
| HGG | High-grade glioma |
| HNSC | Head and Neck squamous cell carcinoma |
| KICH | Kidney Chromophobe |
| KIPAN | Pan-kidney cohort (KICH+KIRC+KIRP) |
| KIRC | Kidney renal clear cell carcinoma |
| KIRP | Kidney renal papillary cell carcinoma |
| LAML | Acute Myeloid Leukemia |
| LGG | Brain Lower Grade Glioma |
| LIHC | Liver hepatocellular carcinoma |
| LSCC | Lung squamous cell cancer |
| LUAD | Lung adenocarcinoma |
| LUSC | Lung squamous cell carcinoma |
| MEL | Melanoma |
| MESO | Mesothelioma |
| NSCLC | Non-small cell lung cancer |
| ODG | Oligodendroglioma |
| OV | Ovarian serous cystadenocarcinoma |
| PAAD | Pancreatic adenocarcinoma |
| PCPG | Pheochromocytoma and Paraganglioma |
| PRAD | Prostate adenocarcinoma |
| RB | Retinoblastoma |
| RCC | Renal cell carcinoma |
| READ | Rectum adenocarcinoma |
| SARC | Sarcoma |
| SKCM | Skin Cutaneous Melanoma |
| STAD | Stomach adenocarcinoma |
| STES | Stomach and Esophageal carcinoma |
| TGCT | Testicular Germ Cell Tumors |
| THCA | Thyroid carcinoma |
| THYM | Thymoma |
| UCEC | Uterine Corpus Endometrial Carcinoma |
| UCS | Uterine Carcinosarcoma |
| UM | Uveal Melanoma |
| UVM | Uveal Melanoma |
| WT | High-Risk Wilms Tumo |
Genetic alterations, localization, and interaction network of ORC6
The gene mutation type and frequency of ORC6 in the TCGA pan-cancer dataset were explored by accessing cBioPortal (https://www.cbioportal.org/). We obtained the copy number variation (CNV) dataset at gene level 4 from all TCGA samples processed by GISTIC software (ref. 14) through GDC (https://portal.gdc.cancer.gov/). We used the unpaired Wilcoxon rank sum test or the Wilcoxon signed rank test to compare the values between the two groups and the Kruskal–Wallis test for differences among multiple groups.
The Human Protein Atlas (HPA, https://www.proteinatlas.org/) was utilized to obtain images of the subcellular localization of ORC6 protein in cancer cells (HEK293 and PC-3) by immunofluorescence staining of cells. Furthermore, the subcellular localization of the ORC6 gene was obtained through the Genecards database.
The comPPI website (http://comppi.linkgroup.hu/) was utilized to analyze the protein-protein interaction network of ORC6. The minimum interaction score was 1, and the edge width was scaled based on the interaction score.
The relationship among ORC6 expression levels, clinical characteristics, and prognosis
The correlation of ORC6 expression with clinical features was assessed by Spearman correlation analysis. We performed univariate Cox regression analysis to investigate the prognostic significance of ORC6 expression in predicting the disease-free interval (DFI), progression-free interval (PFI), overall survival (OS), and disease-specific survival (DSS) in pan-cancer cohorts. We then utilized forest plots for a graphical representation of these results.
TCGA data were curated to extract ORC6 expression levels in transcripts per million (TPM) format, followed by data normalization using log2(TPM+1). Survival data of matched samples were integrated and subsequently subjected to optimal grouping truncation using the ‘surv_cutpoint’ function from the ‘survminer’ package. The aim was to distinguish between the high and low ORC6 expression groups. Prognostic differences between the high- and low-expression groups were evaluated using the log-rank test. RNA-seq data in TPM format from TCGA and GTEx were uniformly processed through the Toil pipeline, as sourced from UCSC XENA (https://xenabrowser.net/datapages/). ORC6 expression levels corresponding to TCGA cancer samples and GTEx normal tissue samples for each cancer type were extracted. The data were normalized using Log2(TPM+1). To assess the diagnostic accuracy for tumor detection, we employed the ‘pROC’ package to calculate sensitivity and specificity. Diagnostic value was quantified by the area under the curve (AUC), with a value of 1.0 indicating perfect diagnostics and 0.5 representing no diagnostic value. An AUC greater than 0.85 was considered to possess a high diagnostic value.
Correlation of ORC6 expression with DNA methylation and RNA modification genes
The correlation of ORC6 expression with DNA promoter methylation levels in cancer was explored by UALCAN(https://ualcan.path.uab.edu/) (ref. 15). The correlation of ORC6 expression with marker gene expression associated with three classes of RNA modifications (N1-methyladenosine (m1A), 5-methylcytosine (m5C) and N6-methyladenosine (m6A)) (ref. 16) across cancers was assessed using Spearman correlation analysis.
Identification of corresponding characteristics of ORC6
To clarify the expression of ORC6 and immune-related characteristics, we employed Spearman correlation analysis to calculate the correlation between ORC6 and 5 types of immune-related genes (chemokines, chemokine receptors, immunosimulators, immunoinhibitors, and MHC). TISIDB(http://cis.hku.hk/TISIDB/) (ref. 17) to assess the immune cell infiltration status of ORC6.
To assess the impact of immunotherapy on ORC6 expression, we analyzed the immunotherapy advanced urothelial carcinoma cohort (IMvigor210 cohort) (ref. 18). The R package ‘limma’ was utilized for differential expression analysis of the target gene in the different groups. Additionally, we accessed the CAMOIP database (https://www.camoip.net/) (ref. 19) to obtain the prognostic information of the Auslander-Melanoma (ref. 20) immunotherapy cohort and assessed the effect of ORC6 expression on the survival time of patients after immunotherapy.
The possibility of ORC6 expression as a predictive marker for immunotherapy response was analyzed using the TISMO (http://tismo.cistrome.org/) (ref. 21) and TIDE (http://tide.dfci.harvard.edu/) (ref. 22) databases. To examine the correlation between ORC6 expression and the half-inhibitory concentration (IC50) of the drug, we employed the R package ‘pRRophetic’ (ref. 23) for the analysis.
Single-cell and bulk transcriptome sequencing analysis
Tumor Immune Single-cell Hub (TISCH, http://tisch.comp-genomics.org/) is a scRNA-seq database that has been specifically developed to investigate the single-cell landscape of the tumor microenvironment (TME) (ref. 24). We screened single-cell datasets, including ALL_GSE132509, BRCA_GSE161529, CESC_GSE168652, CHOL_GSE138709, CRC_GSE166555, ESCA_GSE160269, HNSC_GSE103322, LIHC_GSE166635, LSCC_GSE150321, OV_GSE154600, PAAD_ CRA001160, PRAD_GSE141445, STAD_GSE134520, THCA_GSE148673 and UVM_GSE139829. UMAP plots were used for the visualization of cell types and ORC6 expression levels.
Functional enrichment analysis
We utilized single-cell sequence data obtained from CancerSEA(http://biocc.hrbmu.edu.cn/CancerSEA/) (ref. 25) to examine the relationship between ORC6 and 14 distinct cancer functional states.
To investigate the mechanisms underlying the impact of ORC6 expression on the prognosis of tumor patients, we performed gene set enrichment analysis (GSEA) to explore the ORC6-related signaling pathways, as previously described in the literature (ref. 26, ref. 27). We performed differentially expressed gene (DEG) analysis on the ORC6-low and ORC6-high subgroups of each cancer using the “limma” R package. The threshold was set at 30%, and genes with adjusted P values <0.05 were considered DEGs. We selected the h.all.v7.2.symbols.gmt gene set as our reference and employed it to determine the normalized enrichment score (NES) and false discovery rate (FDR). By examining the correlation between the ORC6 gene expression matrix and the known functional genome, we evaluated the impact of coordinated changes in genes within the genome on phenotypic alterations. The presented findings were visualized as bubble plots with the aid of the R package “ggplot2”. The CAMOIP (ref. 19) network server was employed to perform Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) analyses based on ORC6 expression in TCGA-LIHC transcriptome data using the R package “clusterProfiler”.
Cell lines and ORC6 expression detection
The LIHC cell lines HCCLM3 and MHCC97-H and the hepatic epithelial cell line THLE-2 were acquired from BeNa Culture Collection. The HepG2, U-251 MG, and LN229 cell lines were obtained from Procell and cultured according to the manufacturer’s instructions. Transfection was carried out in 6-well plates (NEST Biotechnology) using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s protocol in HepG2, HCCLM3, U-251 MG, and LN229 cells. The siRNA used in this study was synthesized by GenePharma. Supplementary Table S1 lists the sequences of the siRNAs used in this study. The Western blot experimental steps were described in a previous study (ref. 28). The antibodies used in this study were anti-ORC6 (Proteintech, 17784-1-AP, 1:1000) and anti-alpha tubulin (Proteintech, 11224-1-AP, 1:5000).
Cell viability and proliferation assays
Control and experimental cells were placed in 96-well plates at cell densities of 5,000 (HepG2 and HCCLM3) or 3,000 (U251 MG and LN229) cells per well, respectively. After incubation for 0, 24, 48, and 78 hours, cell viability was assessed by using the CCK-8 assay (GlpBio), and the optical density (OD) was measured at 450 nm with a microplate reader.
Cell proliferation was assessed using EdU (5-ethynyl-2’-deoxyuridine) staining. Briefly, control and experimental cells were seeded in 96-well plates and incubated overnight. After incubation with 10 μM EdU (RiboBio) for 2 hours, cells were fixed with 4% paraformaldehyde for 20 minutes, permeabilized with 0.5% Triton X-100 for 15 minutes, incubated with EdU reaction solution for 30 minutes, and finally incubated with Hoechst 33342 for 10 minutes. Images were taken using an inverted fluorescence microscope (Olympus).
Cell migration assays
Control and experimental cells were seeded onto a 6-well plate and cultured until reaching a confluence of 70% before undergoing transfection and continuing to be cultured normally. Scratch assays were performed when cell confluence reached approximately 90%, with images taken at 0 and 36 hours thereafter. For Transwell migration assays, 50,000 (HepG2 and HCCLM3) or 15,000 (U251 MG and LN229) cells were seeded into the chamber and cultured with serum-free medium within the insert and with 10% complete medium outside of the insert. After 24 hours, cells were fixed and stained, and then the cells within the insert were removed by gently swabbing with a cotton tip before imaging.
Statistical analysis
We used Student’s t-test to determine the statistical significance of differences between the two groups. Paired t-tests were conducted to compare the expression levels of ORC6 in tumor tissues with those in their paired normal tissues. We evaluated the prognostic significance of ORC6 by conducting log-rank and univariate Cox regression analyses. Spearman correlation analysis was employed to assess the correlations between ORC6 and its corresponding features. A p value < 0.05 was considered to indicate statistical significance.
Results
Expression landscape of ORC6
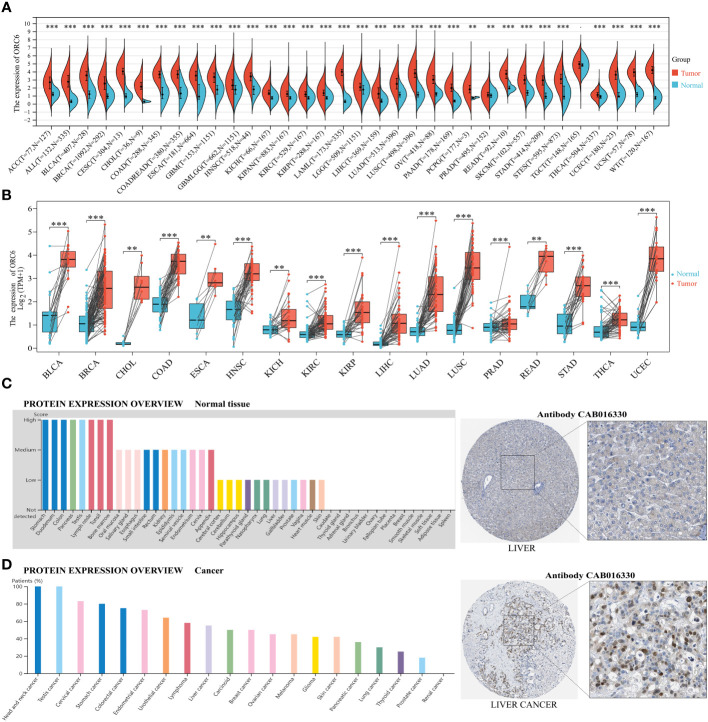
We conducted a comparative analysis of ORC6 expression in tumor vs. normal tissues by merging the TCGA and GTEx databases. ORC6 was found to be significantly upregulated in 33 tumors (all p<0.05), as illustrated in f1. However, no significant changes were observed in TCGT. Then, our analysis of tumor and matched normal tissue samples from the TCGA database revealed that ORC6 expression was significantly elevated in tumor samples from BLCA, BRCA, CHOL, COAD, ESCA, HNSC, KICH, KIRC, KIRP, LIHC, LUSC, LUAD, PRAD, READ, STAD, THCA, and UCEC (f1, all p<0.05). Further protein score hints were provided by the HPA online database. ORC6 showed high protein scores in the stomach, duodenum, colon, pancreas, lymph nodes, testis, and bone marrow but low protein scores in the liver (f1). Regarding its protein expression in tumors, we observed that ORC6 was moderately/highly expressed in 100% of head and neck cancer (3/3) and testicular cancer (11/11) tissues and was moderately/highly expressed in 54.5% (6/11) of liver cancer tissues (f1). To clarify the localization of ORC6 protein expression, we obtained immunofluorescence staining images of ORC6 protein expression in HEK 293 and PC-3 cells through the HPA database (Supplementary Figure S1A). Further access to the Genecards database was performed for validation (Supplementary Figure S1B). We found that ORC6 was mainly concentrated in the nucleus and cytoplasm. Finally, we constructed a PPI network using interaction data sourced from the ComPPI website. The results of our analysis showed that proteins found to be in close interaction with ORC6 were primarily localized within the nucleus, as depicted in Supplementary Figure S1C. In summary, we identified that ORC6 was generally highly expressed in tumors.
ORC6 genetic alterations and epigenetic modifications
The frequency and type of ORC6 gene genetic alterations across cancers were analyzed by the cBioPortal platform. As illustrated in Supplementary Figure S1D, the most frequent type of genetic alteration in ORC6 was “amplification”, followed by “deep deletion”, “mutation” and “structural variation”. SARC exhibited the greatest frequency of ORC6 genetic mutations. These alterations included “deep deletions” in 2.35% of genes and “structural variants” in 0.39% of genes. In PRAD, the gene alteration frequency of ORC6 was 2.63%, of which the frequency of “amplification” reached 2.43%. In both DLBC and UCS, ORC6 genetic alterations were “deeply deleted”. In UCEC and SKCM, the frequency of ORC6 gene “mutation” reached 1.13%. In five different cancer types (ACC, ESCA, LIHC, KIRP, and PAAD), the ORC6 gene only exhibited “amplified” genetic variants. The mutation frequency of the ORC6 gene is generally low, at less than 3%. This could be due to the high conservation of genes within the ORC family (ref. 29). Subsequently, to explore the relationship between ORC6 expression and genomic variations across different cancer types, we employed either the Wilcoxon rank-sum test or the Kruskal–Wallis rank-sum test. We observed differential expression of ORC6 across three distinct variant groups (gain-variant, loss-variant, and no-variant) in 14 different cancers: BLCA, BRCA, CESC, COAD, HNSC, KIPAN, LIHC, LUAD, LUSC, OV, PRAD, STAD, STES, and UCS (Supplementary Figure S1E, all p<0.05). Specifically, ORC6 expression was generally higher in the gain-variant group than in the loss/neutral-variant group.
Numerous reports suggest that abnormal DNA methylation in the promoter region of genes can induce changes in chromatin structure and DNA stability, ultimately leading to the dysregulation of gene expression within the body (ref. 30). Therefore, we analyzed differences in the DNA promoter methylation levels of ORC6 between tumor and normal tissues using UALCAN. As depicted in Supplementary Figure S2A, methylation levels were lower in BLCA, BRCA, HNSC, THYM, UCEC, and PRAD than in normal tissues (all p<0.05). In contrast, methylation levels were higher in PAAD, KIRC, LUSC, and SARC than in normal tissues (Supplementary Figure S2A, all p<0.05). Moreover, RNA modifications are critical in selectively regulating the expression of genes (ref. 31). Our analysis, as illustrated in Supplementary Figure S2B, reveals a strong positive correlation between ORC6 expression and m1A-, m5C-, and m6A-related genes across almost all tumor types. These findings suggest that the ubiquitous overexpression of ORC6 in tumors may be closely associated with its epigenetic modifications and genetic variations. This correlation further supports the potential of ORC6 as a cancer regulatory factor and provides valuable clues for further exploring its role in cancer.
Correlation of ORC6 expression with clinicopathological features
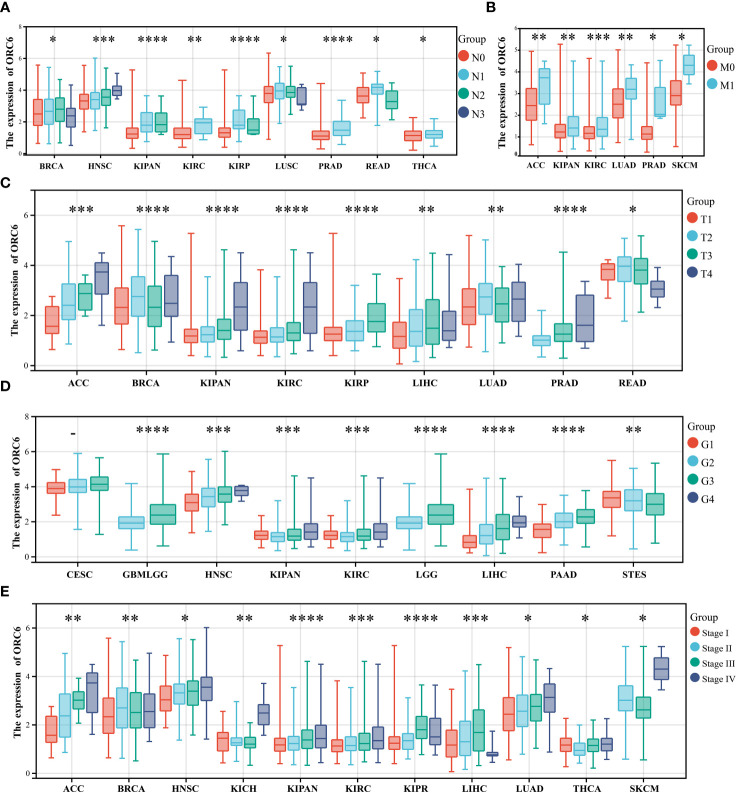
We also investigated the correlations between ORC6 expression and various clinicopathological features. According to the results presented in f2, there was a positive correlation between ORC6 expression and lymph node metastasis in several tumor types, and the correlations in HNSC (p=2.2e-4), KIPAN (p=6.8e-8), KIRC (p=5.6e-3), PRAD (p=4.0e-8), THCA (p=0.02) and other tumor types were the most robust. f2 shows that the increase in ORC6 expression was positively correlated with tumor metastasis in ACC (p=8.2e-3), KIPAN (p=1.9e-3), KIRC (p=4.3e-4), LUAD (p=9.5e-3), PRAD (p=0.01), and SKCM (p=0.03). Furthermore, increased ORC6 expression was positively correlated with the T stage of ACC (p=4.8e-8), KIPAN (p=1.1e-9), KIRC (p=1.4e-5), KIRP (p=2.4e-8) and PRAD (p =7.0e-12) (f2). f2 shows that the increased expression of ORC6 was positively correlated with the histological grade of GBMLGG (p=9.2e-25), HNSC (p=5.6e-9), LGG (p=9.2e-25), LIHC (p=3.0e-15) and PAAD (p=2.9e-9) but negatively correlated with the histological grade of STES (p=6.3e-5). Similarly, increased ORC6 expression was positively correlated with clinical staging (f2), and typical tumor types were ACC (p=6.3e-5), HNSC (p=1.5e-3), KIPAN (p=3.3e-9), KIRC (p=2.4e-4), and LUAD (p=0.01). From the above findings, it can be inferred that ORC6 might play a role in tumor progression and metastasis.
Prognostic and diagnostic value of ORC6
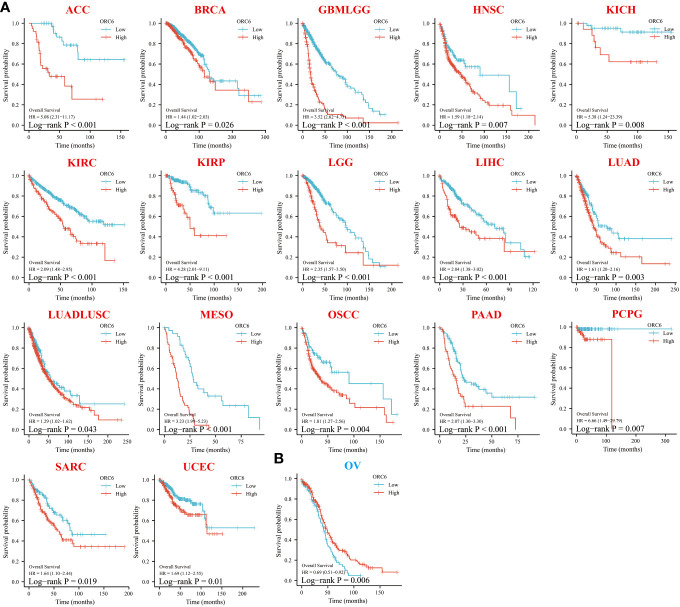
To investigate the effect of ORC6 on tumor prognosis, we plotted survival curves and assessed OS using the Kaplan–Meier method. As shown in f3, in ACC, BRCA, GBMLGG, HNSC, KICH, KIRC, KIRP, LGG, LIHC, LUAD, LUADLUSC, MESO, OSCC, PAAD, PCPG, SARC and UCEC patients, high ORC6 levels were highly correlated with poorer OS (f3; all p<0.05). Moreover, it is worth noting that high ORC6 expression was exclusively correlated with improved OS in OV (f3; p<0.05).
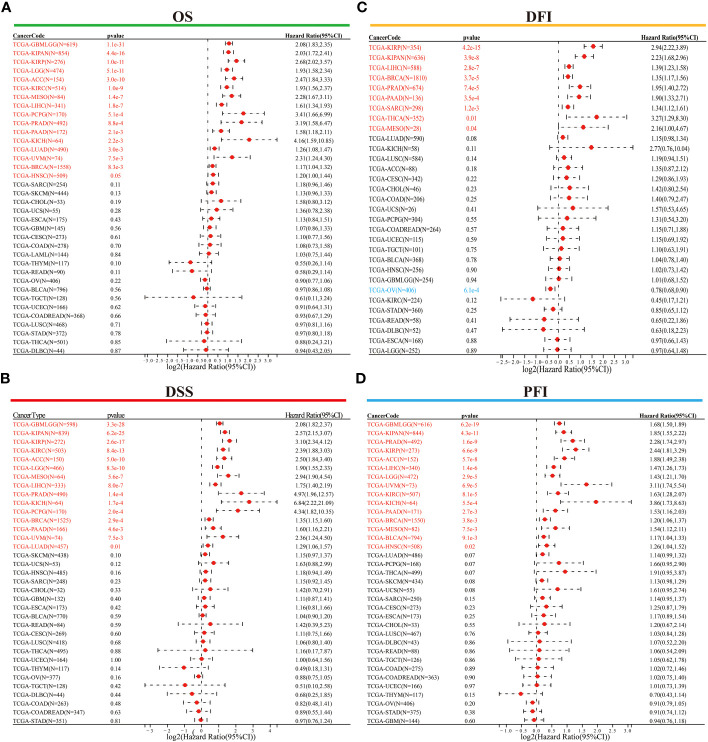
Following this, we conducted Cox regression analysis to assess the correlation between ORC6 expression and several survival outcomes, including OS, DSS, DFI, and PFI, for each tumor type. The results were then presented in the form of a forest plot. As shown in f4, our findings indicate that high ORC6 expression was significantly associated with shorter OS in GBMLGG, KIPAN, KIRP, LGG, ACC, KIRC, MESO, LIHC, PCPG, PRAD, PAAD, KICH, LUAD, UVM, BRCA and HNSC patients (all p<0.05). High ORC6 expression in GBMLGG, KIPAN, KIRP, KIRC, ACC, LGG, MESO, LIHC, PRAD, KICH, PCPG, BRCA, PAAD, UVM, and LUAD patients was associated with poorer DSS (f4, all p<0.05). Regarding DFI, there was a significant association between high ORC6 expression and lower DFI in KIRP, KIPAN, LIHC, BRCA, PRAD, PAAD, SARC, THCA, MESO, and LUAD patients, whereas high ORC6 expression in OV patients was associated with improved prognosis for DFI (f4, all p<0.05). Furthermore, as illustrated in f4, high ORC6 levels were strongly correlated with poorer PFI in GBMLGG, KIPAN, PRAD, KIRP, ACC, LIHC, LGG, UVM, KIRC, KICH, PAAD, BRCA, MESO, BLCA and HNSC (all p<0.05).
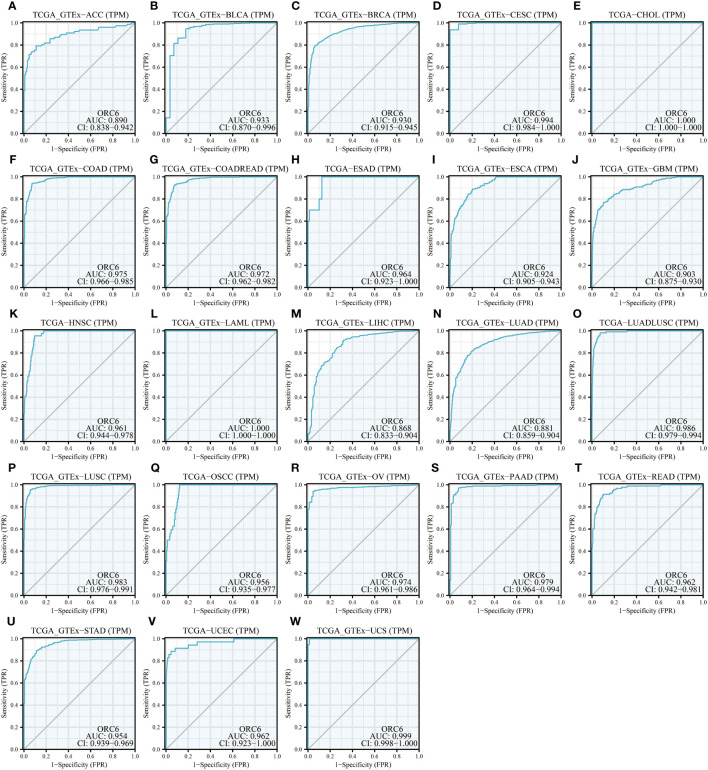
Moreover, we assessed the diagnostic accuracy of ORC6 in different types of cancer using ROC curves. As shown in f5, in ACC, BLCA, BRCA, CESC, CHOL, COAD, COADREAD, ESAD, ESCA, GBM, HNSC, LAML, LIHC, LUAD, LUADLUSC, LUSC, OSCC, OV, PAAD, READ, STAD, UCEC and UCS, ORC6 could be used as a highly accurate diagnostic marker (f5, all AUC>0.85). In DLBC, GBMLGG, KICH, KIRC, KIRP, SKCM, and THYM, ORC6 had moderate diagnostic performance (Supplementary Figures S3A–E, H, K; all AUC=0.7~0.85). In LGG, PRAD, TGCT, and THCA, ORC6 had poor diagnostic values (Supplementary Figures S3F, G, I, J; all AUC=0.5~0.7). In conclusion, our study revealed that high expression of ORC6 is generally linked to unfavorable prognosis in the majority of cancer types, and it has good diagnostic value.
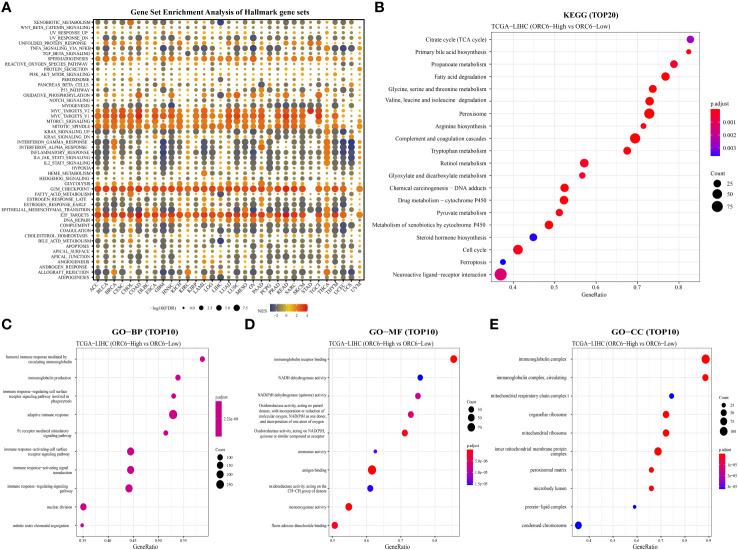
Pathways and functions associated with ORC6 expression
To explore the possible biological pathways influenced by ORC6 that may contribute to tumorigenesis and progression, we conducted GSEA on data obtained from 33 tumors from TCGA. As illustrated in f6, we observed that immune-related pathways, including TNFα signaling via NFκB, IFN-α response, IFN-γ response, inflammatory response, IL-6/JAK/STAT3, IL-2/STAT5, complement and coagulation cascades, and allograft rejection pathways, were significantly enriched across a diverse range of tumors. Moreover, we observed a positive correlation between ORC6 expression and MYC target V2, MYC target V1, MTORC1, mitotic spindle, G2 checkpoint, E2F target, DNA repair, and other pathways across cancers. Furthermore, KEGG analysis revealed that ORC6 was mainly involved in the synthesis and degradation of various substances, drug metabolism, the cell cycle, ferroptosis, and neuroactive ligand-receptor interactions in LIHC (f6). GO analysis, including the BP, CC, and MF categories, indicated that ORC6 was mainly related to immune response regulation and biological enzyme activity in LIHC (f6).
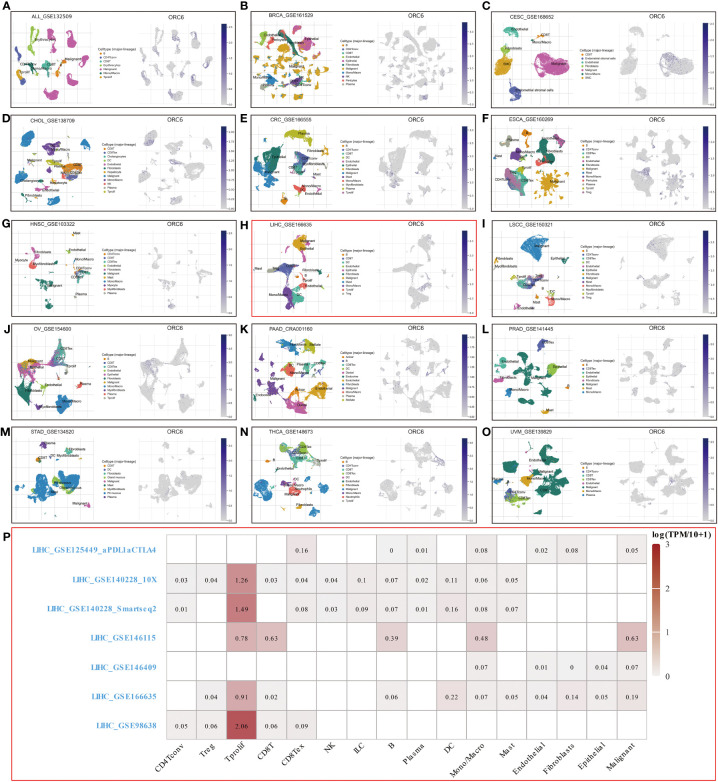
To explore ORC6 expression in diverse TMEs, including ALL, BRCA, CESC, CHOL, CRC, ESCA, HNSC, LIHC, LSCC, OV, PAAD, PRAD, STAD, THCA, and UVM, we investigated their expression distribution (f7). The results were interesting, as they showed that ORC6 was primarily expressed at high levels in the malignant cells of these cancers. In STAD, it was predominantly expressed in pit mucus cells (f7). It is worth noting that in LIHC, ORC6 was also found to be expressed in T-cell proliferation, which demonstrates its potential role in this immune response (f7). To further investigate the relationship between ORC6 and the functional status of different cancers, we analyzed single-cell sequencing data obtained from CancerSEA for 14 types of cancer. In most tumors, ORC6 showed a positive correlation with the cell cycle, proliferation, DNA damage, and DNA repair (Supplementary Figure S4). In contrast, ORC6 was negatively associated with apoptosis, hypoxia, metastasis, and quiescence in most tumors (Supplementary Figure S4). These findings suggest a correlation between abnormal expression of ORC6 and the advancement of cancer as well as the immune response of cancer.

Correlation between ORC6 expression and the tumor immune landscape
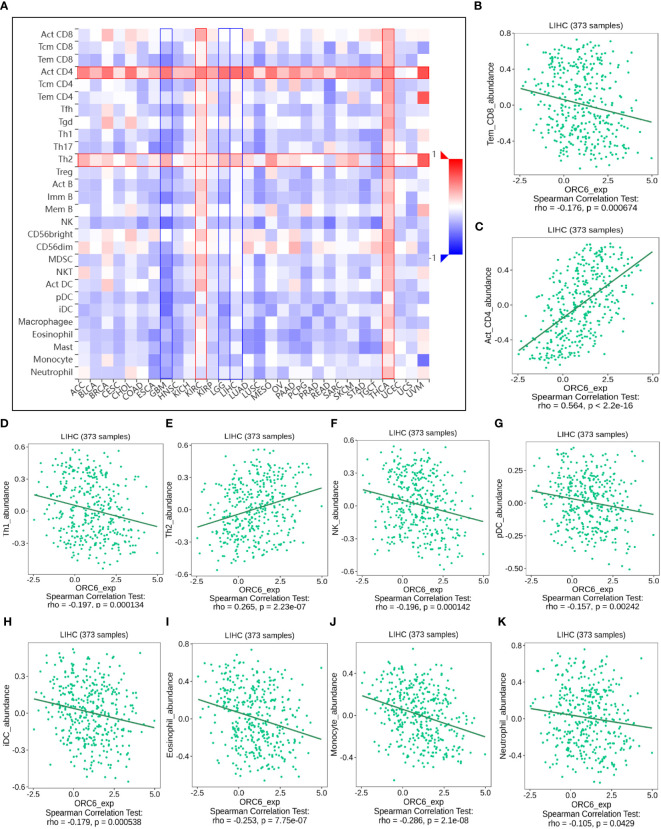
Investigating the possible gene expression within tumors and its connection to immune cells can greatly aid in predicting the clinical outcome for patients with tumors and selecting appropriate diagnostic targets and intervention strategies (ref. 32). To gain further insights into the correlation between ORC6 and immune cells in the TME, we used the TISIDB tool for analysis. The pan-cancer analysis indicated that the expression level of ORC6 displayed an inverse correlation with the infiltration abundance of various immune cells, including Tem CD8 cells, Th1 cells, NK cells, pDCs, iDCs, eosinophils, monocytes, and neutrophils, while showing a positive correlation with the infiltration abundance of Act CD8 cells and Th2 cells (f8). This phenomenon was especially evident in LIHC (f8). Furthermore, in gliomas, which include GBM and LGG, ORC6 expression was negatively correlated with the abundance of infiltrating immune cells, including Tem CD8, Tcm CD4, Tfh, Th1, Th17, Act B, lmm B, NK, MDSC, NKT, Act DC, pDC, iDC, macrophage, eosinophil, mast, monocyte, and neutrophil cells. In contrast, ORC6 expression levels in THCA and KIRC were positively correlated with the majority of immune cell infiltration (f8).
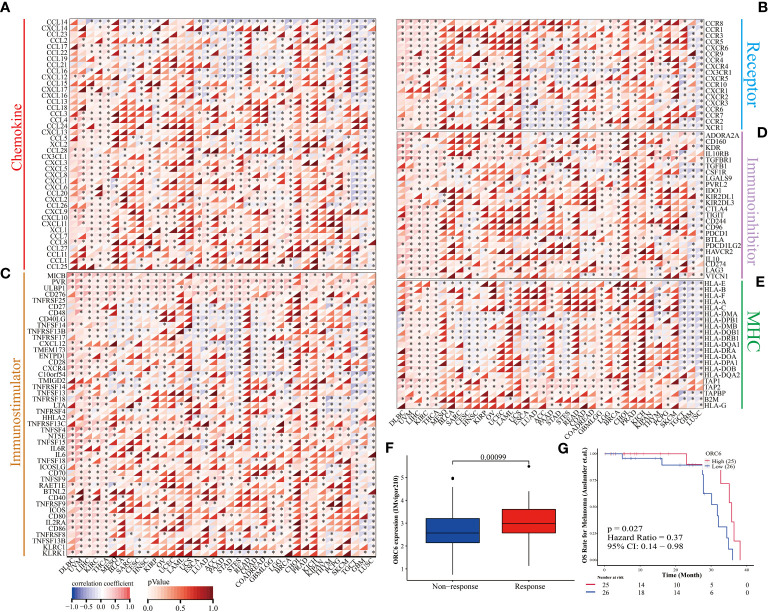
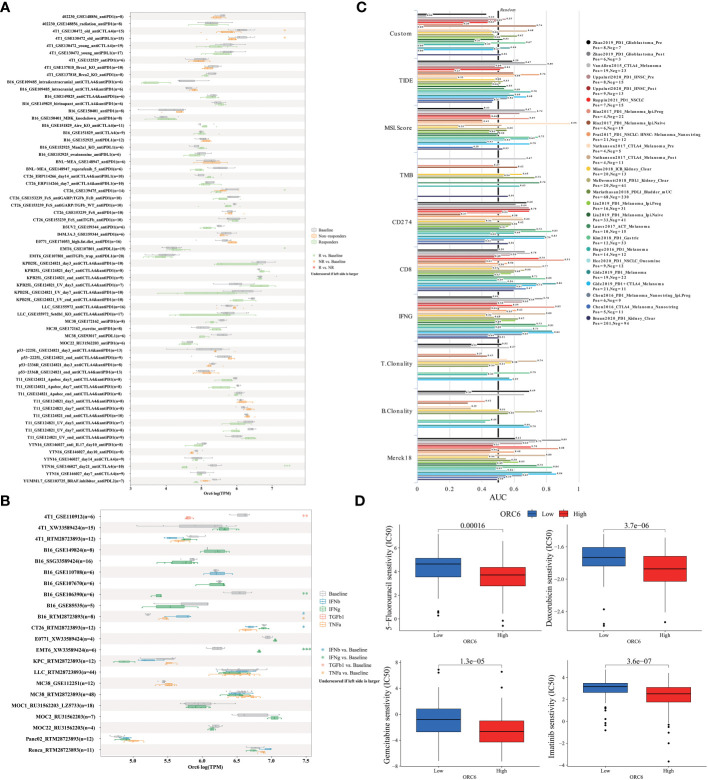
Furthermore, we examined the association between ORC6 expression and the expression of genes related to immune regulation (f9). The heatmap indicated that ORC6 was coexpressed with most immune-related genes across cancers. Especially in DLBC, UVM, LIHC, KIRC, and THCA, ORC6 was roughly positively correlated with 5 immune-related genes. In TGCT, GBM, and LUSC, ORC6 was roughly negatively correlated with immune-related genes. Furthermore, the chemokine CCL14 was negatively correlated with ORC6 expression across cancers (f9). Among the immune activation-related genes, MICB, PVR, ULBP1, CD276, and TNFRSF25 were positively correlated with ORC6 expression in most tumors (f9). There was a positive correlation between the expression of ORC6 and genes related to immunosuppression in several cancer types, such as DLBC, UVM, LIHC, KIRC, THCA, GBMLGG, LGG, and PRAD, as indicated in f9. Notably, in the advanced urothelial carcinoma cohort with immunotherapy, the response group had significantly higher ORC6 expression (f9, p=0.00099). Furthermore, we observed that in the melanoma immunotherapy cohort, the high ORC6 expression group had prolonged survival after immunotherapy (f9, p=0.027). Taken together, these results suggest that ORC6 may be involved in immune cell infiltration and the expression of immunomodulatory genes and that high ORC6 expression may indicate a better response to immunotherapy.
ORC6 predicts immunotherapy response and chemotherapy efficacy
To elucidate the predictive function of ORC6 expression in immunotherapy response, we evaluated it using the TISMO database. As depicted in f10, ORC6 expression was markedly different in 5 subjects before and after ICB treatment and between responder and nonresponder cohorts. Moreover, ORC6 expression was significantly different in the six cell lines before and after cytokine treatment (f10). Furthermore, we performed a biomarker assessment of ORC6 by TIDE. The findings indicated that ORC6 had a better predictive effect in 7 immunotherapy cohorts (f10). In addition, we performed a sensitivity analysis of chemotherapy drugs commonly used to treat LIHC. As shown in f10, the ORC6 high-expression group was closely correlated with the reduction of IC50 of 5-fluorouracil, doxorubicin, gemcitabine, and imatinib (all p<0.001).
External cohort and in vitro experiments clarify the promotional effect of ORC6 on LIHC and GBMLGG
After collating and analyzing the pan-cancer data mentioned above, it was observed that the elevated expression of ORC6 was significantly associated with the unfavorable prognosis and malignancy of LIHC and GBMLGG patients (f1–f4). Consequently, our study will concentrate on LIHC and GBMLGG.
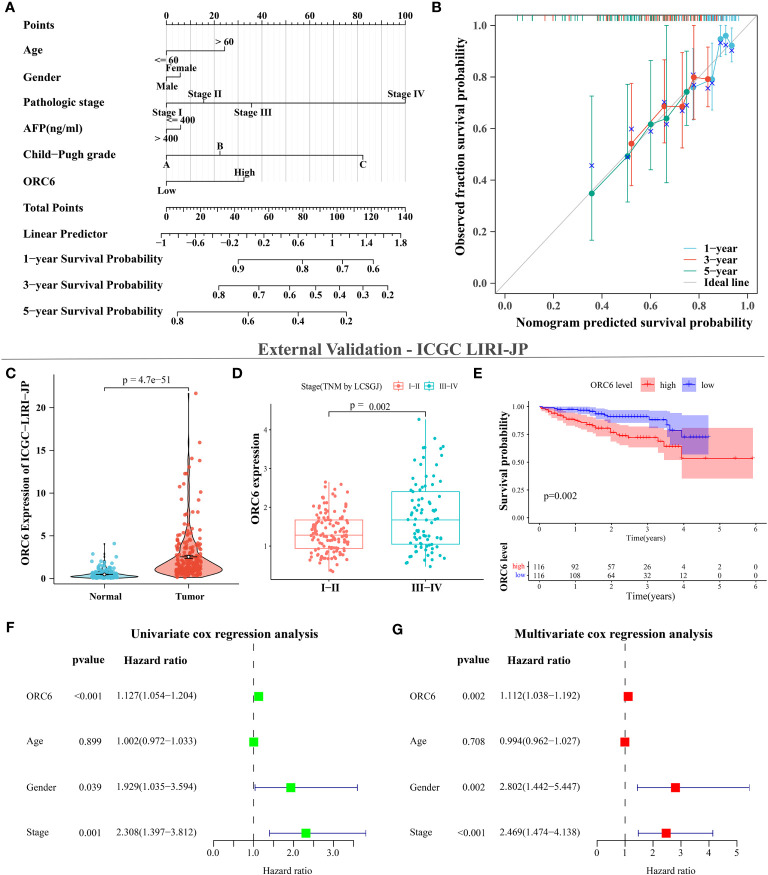
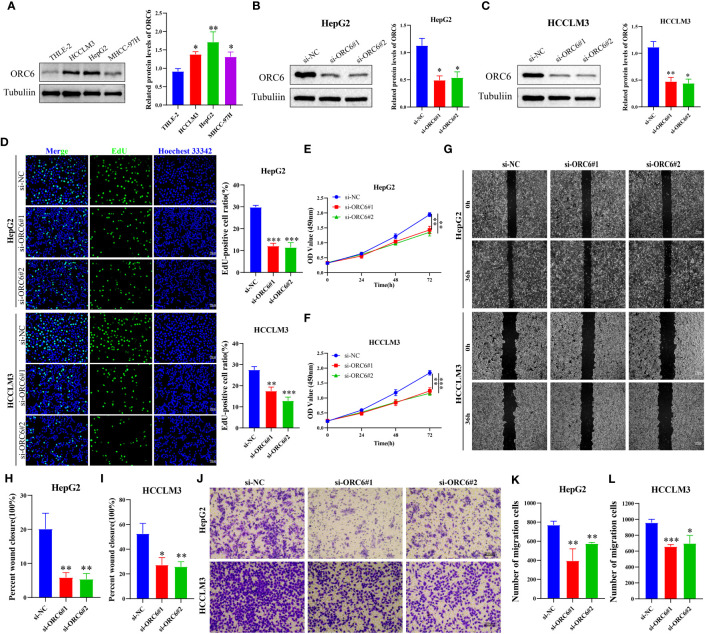
We collected clinical information and ORC6 expression profiles of patients belonging to the TCGA-LIHC cohort. After performing a chi-square test analysis, we discovered a significant correlation between ORC6 expression and tumor histological grade, alpha-fetoprotein (AFP) content, and vascular invasion in LIHC (Supplementary Table S2). Furthermore, via univariate and multivariate Cox regression analysis, we identified ORC6 and pathological stage as independent prognostic risk factors for LIHC (Supplementary Table S3). Subsequently, a nomogram was developed to estimate the survival likelihood of patients at intervals of 1, 3, and 5 years, and its prediction efficiency was confirmed by the calibration curve, as illustrated in f11, f11. This indicates that the model had a high accuracy in its predictive ability. Given that our analysis of ORC6 was solely based on the TCGA database, we conducted external verification by collating clinical information and ORC6 expression profiles of patients belonging to the ICGC-LIRI-JP cohort. The results showed that ORC6 expression in LIHC was significantly higher than that in normal tissues (f11). It was positively correlated with the clinical stage and associated with poor overall survival (f11). Furthermore, univariate and multivariate Cox regression analyses revealed ORC6, sex, and clinical stage as independent risk factors for poor prognosis in LIHC (f11). To investigate the function of ORC6 in LIHC cells, we carried out in vitro experiments. Initially, we assessed the basal expression of ORC6 in LIHC cell lines and normal hepatocytes. Our findings indicate that LIHC cell lines have noticeably increased ORC6 expression compared to normal hepatocytes, as illustrated in f12. To assess the impact of ORC6 downregulation in LIHC cells, we chose two LIHC cell lines with high expression (HepG2 and HCCLM3) and conducted an ORC6 knockout assay (f12). Using the EdU cell proliferation test, a significant decrease in the proliferation of LIHC cells was observed after the knockout of ORC6 expression (f12). In addition, similar results were obtained through CCK-8 analysis (f12). Subsequently, we conducted wound healing and Transwell tests to examine the impact of ORC6 downregulation on the migratory capacity of LIHC cells. The results indicate that compared to the control cells, the downregulation of ORC6 significantly inhibited the migration ability of LIHC cells (f12).
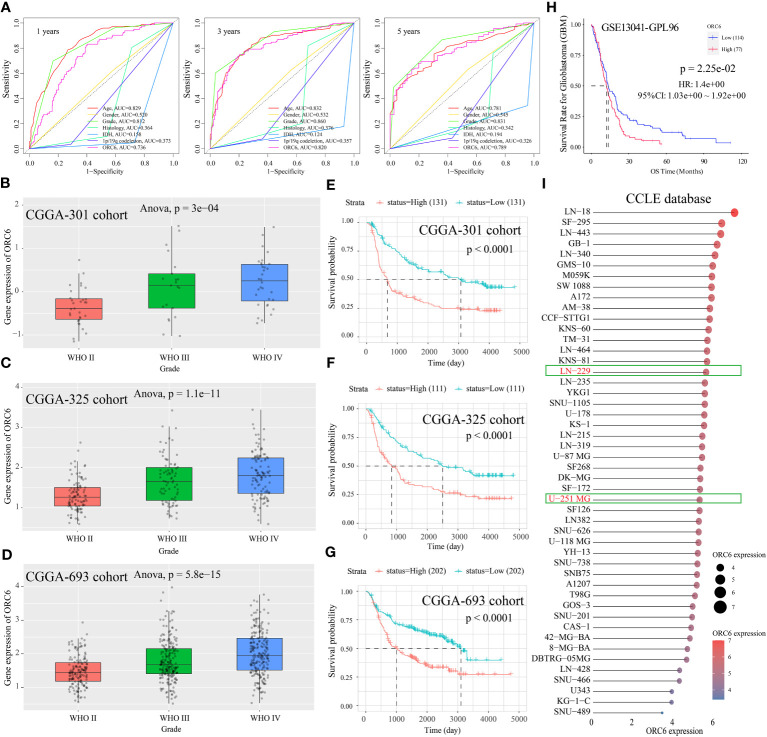
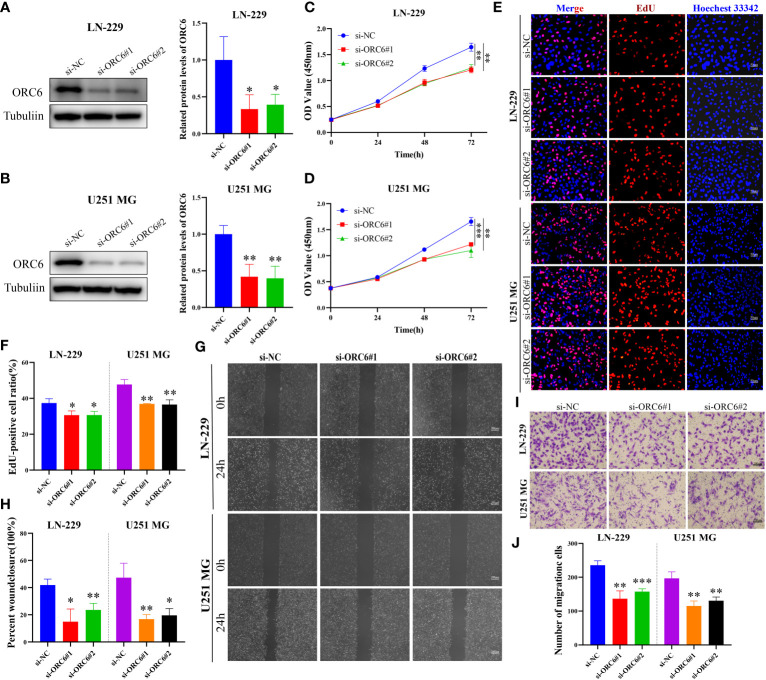
Subsequently, we compiled the ORC6 expression and clinical information of patients in the TCGA-GBMLGG cohort. The chi-square test revealed a significant association between ORC6 expression and age, histological type, WHO grade, IDH status, and 1p/19q codeletion in GBMLGG patients (Supplementary Table S4). Furthermore, our investigation revealed that ORC6, age, WHO grade, and 1p/19q codeletion were independent prognostic risk factors for GBMLGG patients, as confirmed by univariate and multivariate Cox regression analyses (Supplementary Table S5). Time-dependent ROC curve analysis further identified age, WHO grade, and ORC6 as the top three effective predictors for 1-, 3-, and 5-year patient survival (f13, all AUC>0.7). Our results were validated in an external cohort. As demonstrated in f13, ORC6 expression increased concomitantly with WHO grade in the CGGA-301, CGGA-325, and CGGA-693 cohorts. Moreover, high levels of ORC6 expression were significantly associated with an unfavorable prognosis among GBMLGG patients in the CGGA-301, CGGA-325, CGGA-693, and GSE13041 cohorts (f13, all p < 0.05). In addition, the expression profile of ORC6 in the GBMLGG cell line was analyzed using the CCLE database (f13). Further, the expression of ORC6 protein was effectively inhibited in U251 MG and LN229 cells (f14). Consistent with findings in LIHC, the inhibition of ORC6 expression led to decreased proliferation and migration of U251 MG and LN229 cells (f14). In conclusion, our in vitro results strengthen the evidence supporting the carcinogenic effects of ORC6 in LIHC and GBMLGG.
Discussion
Multi-omics data mining analysis is crucial for exploring tumor heterogeneity and complexity and identifying prognostic biomarkers. Prior studies have linked high ORC6 expression to poor tumor prognosis, progression, and drug resistance in some cancers (ref. 33, ref. 34), but its prognostic and biological significance in most cancer types remains unclear. We performed a comprehensive pan-cancer study of ORC6 and revealed its important role in LIHC and GBMLGG. In addition, we verified that ORC6 was highly expressed in LIHC and GBMLGG and could serve as an independent marker of poor prognosis. Further external cohort analysis and in vitro experiments supported our findings.
In this study, we observed that the expression of ORC6 was higher in most tumors than in normal tissues, as well as in paired cancer and paracancerous tissues. Next, we analyzed the correlations between ORC6 expression and clinicopathological features and discovered that ORC6 expression was positively correlated with tumor size, metastasis, histological grade, lymph node metastasis, and clinical analysis, which further implied that ORC6 expression was associated with tumor progression and metastasis. By utilizing log-rank and Cox regression analysis, it was determined that increased expression of ORC6 was significantly linked to unfavorable prognosis in various types of tumors, in concurrence with previous research (ref. 10, ref. 34, ref. 35). Through ROC curves, we also found that ORC6 was a highly accurate diagnostic marker for most tumor types. Single-cell functional analysis also indicated that ORC6 expression was positively correlated with the cell cycle and proliferation of tumor cells. We found that ORC6 and pathologic stage were independent prognostic risk factors for patients with LIHC. ORC6, age, WHO grade, and 1p/19q codeletion are independent risk factors for poor prognosis of GBMLGG, and this result is consistent with previous studies (ref. 36). Furthermore, the time-dependent ROC curves showed that ORC6 was more accurate than sex, histological type, IDH, and 1p/19q colocation in predicting the 1-, 3-, and 5-year survival of GBMLGG patients. After silencing ORC6 expression, we found that the proliferation and migration abilities of LIHC and GBMLGG cells were attenuated. The results of this study suggest that elevated ORC6 levels may serve as a valuable prognostic marker for adverse outcomes in most tumor types. However, the validation of this study was limited to in vitro experiments, and further in vivo studies are needed to fully explore this possibility.
Genetic alterations and altered epigenetic regulation are considered major factors in cancer development and progression (ref. 37, ref. 38). In recent years, there has been growing recognition that RNA not only serves as an intermediary or effector molecule in protein synthesis but also plays a crucial and direct functional role in regulating gene expression. Consequently, the significance of RNA modifications has gained increasing prominence in scientific research and healthcare settings. Extensive evidence has suggested that the perturbation of RNA epigenetic pathways is associated with the development and progression of various human diseases, including cancer (ref. 39). In our study, we found that genetic alterations, DNA promoter methylation, and RNA modifications of ORC6 have important effects on its expression. The main mutation forms of ORC6 in tumors were “amplification” and “deep deletion”, and the amplification was mainly concentrated in PAAD, BRCA, OV, BLCA, ACC, ESCA, LIHC, LUAD, etc. We also noticed that ORC6 expression was markedly linked to CNV, mainly in BRCA, CESC, HNSC, LUAD, LUSC, OV, and STES. In BLCA, BRCA, HNSC, THYM, UCEC, and PRAD, as the levels of ORC6 promoter methylation were reduced compared to those in normal tissues. However, in PAAD, KIRC, LUSC, and SARC, there was a significant increase in ORC6 promoter methylation levels. Furthermore, we identified a positive correlation between ORC6 expression and m1A-, m5C-, and m6A-related genes in almost all of the analyzed tumor types. Our single-cell functional analysis also indicated a close association between ORC6 expression and DNA damage and repair mechanisms, which highlights the underlying mechanisms of aberrant ORC6 expression in cancer from both genetic alteration and epigenetic modification perspectives.
Cancer progression, metastasis, invasion, and resistance to therapy are modulated by bidirectional interactions between cancer cells and the TME (ref. 40). Characteristics of TME include hypoxia, immunosuppression, chronic inflammation, acidosis, high interstitial fluid pressure, increased ECM stiffness, and depletion of essential nutrients. Immunotherapy mainly targets hypoxia and immunosuppression, which are presently active research topics (ref. 41). Precision medicine aims to develop targeted and immunotherapies to enhance the survival rate. Cancer immunotherapy presents an effective and groundbreaking method to fight cancer by manipulating or modulating the immune system to elicit a robust response against tumors (ref. 42). Successful cancer immunotherapy depends on overcoming the immunosuppressive environment in the TME of cancer patients (ref. 43). Increasing evidence suggests that immune dysregulation plays a critical role in allowing tumors to evade the host immune system (ref. 44), involving both innate and adaptive immunity. Research has revealed that tumor-infiltrating lymphocytes tend to exhibit dysfunctional behavior and may remain in a quiescent state near cancerous cells. Despite this, a few patients’ T cells have been found to preserve their ability to proliferate and persist, leading to the complete eradication of sizable tumor deposits (ref. 45). This finding is consistent with our finding in single-cell sequencing that ORC6 is predominantly expressed on Tprolif and malignant cells. Therefore, targeting ORC6 could potentially offer a precise method for identifying Tprolif and malignant cells, leading to novel avenues for tumor immunotherapy (ref. 46). Here, we found an inverse correlation between the expression level of ORC6 and the abundance of immune cells widely believed to contribute to the suppression of tumor infiltration, including Tem CD8 cells, Th1 cells, NK cells, pDCs, iDCs, eosinophils, and monocytes (ref. 47–ref. 52). Interestingly, we noticed that ORC6 expression levels were positively correlated with the abundance of Act CD4 and Th2 cells in the TME. Act CD4 refers to activated CD4 T cells, a key component of the adaptive immune system. Recent studies have found that CD4+ T-cell infiltration defines an immune escape environment and predicts poor patient outcomes (ref. 53). Th2 refers to helper T-cell type 2, which is a specific type of T-cell in the immune system. Th2 cells play a critical role in the adaptive immune response by supporting B-cell function. Interestingly, the accumulation of Th2 cells within tumors, in addition to Hodgkin’s lymphoma, has been associated with a poor prognosis in several types of cancers (ref. 54). At present, researchers are exploring ways to regulate Th2 cells to improve the effect of tumor treatment. Regulatory T cells (Tregs) are a subset of T cells crucial for maintaining immune homeostasis and tolerance. Research suggests that several subtypes of Tregs, including TNFR2+, LAG3+, TIM3+, and CTLA-4+ Tregs, demonstrate potent anticancer capabilities. However, in recent years, researchers have also discovered links between particular highly infiltrated Treg subtypes within tumors and favorable patient outcomes, such as CD30+OX40+ and BLIMP‐1+FOXP3+ Tregs (ref. 55). This might elucidate the favorable correlation between ORC6 and Treg infiltration abundance in BRCA, KIRC, MESO, and THCA, whereas an inverse correlation exists in the majority of other tumor types.
Cancer cells can secrete important cytokines and chemokines for the TME during growth and progression (ref. 56), these cytokines and chemokines can in turn regulate the TME and cell signaling pathways to affect cancer progression (ref. 57, ref. 58). Our study revealed a positive correlation between ORC6 expression and cytokines as well as receptors in different types of tumors. Notably, GSEA revealed a strong association between ORC6 expression and the cytokine-cytokine receptor interaction pathway, and our GSEA further highlights the close association between ORC6 and immune-related pathways in multiple tumor types. Moreover, we discovered a positive association between ORC6 expression and well-known targets for classical immune suppression and activation, such as PVR, MICB, ULBP1, CD276, CTLA4, TNFRSF25, PD-1 (PDCD1), TIGIT, PD-L2 (PDCD1LG2), HAVCR2, PD-L1 (CD274), and LAG3. Strikingly, our findings reveal that ORC6 expression has a certain predictive effect on immunotherapy, cytokine therapy, and chemotherapy response. The current use of multifunctional carriers to deliver therapeutic drugs to lesion sites helps to significantly improve the effect of noninvasive treatment. Multifunctional carriers allow for multiple treatment options, including photodynamic therapy, photothermal therapy, chemotherapy, immunotherapy, or their synergistic treatments (ref. 59). Therefore, molecular probes targeting ORC6 combined with multifunctional carriers are promising cancer treatment strategies (ref. 60). Collectively, these observations provide new insights into the complex regulation of immune cell-mediated tumor suppression and suggest that ORC6 may serve as a promising predictive marker of immunotherapy efficacy in cancer treatment. Nevertheless, the mechanisms by which ORC6 regulates the tumor immune microenvironment and tumor progression need to be further elucidated in the future.
Conclusion
ORC6 emerges as a promising prognostic biomarker across various cancer types, particularly in LIHC and GBMLGG. This study underscores the correlation between high ORC6 expression and the tumor immunosuppressive environment. These findings suggest a potential role for ORC6 in tumor immune regulation, thereby offering further support for advancing the development of cancer immunotherapies.
Data availability statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/Supplementary Material.
Ethics statement
Ethical approval was not required for the study involving humans in accordance with the local legislation and institutional requirements. Written informed consent to participate in this study was not required from the participants or the participants’ legal guardians/next of kin in accordance with the national legislation and the institutional requirements.
Author contributions
JZ conducted the formal analysis and wrote the original draft. WY performed the project administration. QC, LZ, HG, TW, YHe, and YD conducted the experiments. JX, JPang, JPeng, HG, TW, and YHan participated in software analysis. JZ, QC, LZ, and JPeng conducted data curation. JZ and WY contributed to writing, reviewing, and editing the article. WY provided funding acquisition. All authors read and approved the final submitted manuscript.
References
- H Sung, J Ferlay, RL Siegel, M Laversanne, I Soerjomataram, A Jemal. Global cancer statistics 2020: globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries.. CA Cancer J Clin (, 2021. [DOI]
- F Chen, Y Zhang, DL Gibbons, B Deneen, DJ Kwiatkowski, M Ittmann. Pan-Cancer Molecular Classes Transcending Tumor Lineage across 32 Cancer Types, Multiple Data Platforms, and over 10,000 Cases.. Clin Cancer Res (, 2018. [DOI]
- H Yamaguchi, JM Hsu, WH Yang, MC Hung. Mechanisms regulating pd-L1 expression in cancers and associated opportunities for novel small-molecule therapeutics.. Nat Rev Clin Oncol (, 2022. [DOI | PubMed]
- A Friedlaender, V Subbiah, A Russo, GL Banna, U Malapelle, C Rolfo. Egfr and her2 exon 20 insertions in solid tumours: from biology to treatment.. Nat Rev Clin Oncol (, 2022. [DOI | PubMed]
- J Xia, S Li, S Liu, L Zhang. Aldehyde dehydrogenase in solid tumors and other diseases: potential biomarkers and therapeutic targets.. MedComm (, 2023. [DOI | PubMed]
- S Liu, M Balasov, H Wang, L Wu, IN Chesnokov, Y Liu. Structural analysis of human orc6 protein reveals a homology with transcription factor tfiib.. Proc Natl Acad Sci USA (, 2011. [DOI]
- SG Prasanth, KV Prasanth, B Stillman. Orc6 involved in DNA replication, chromosome segregation, and cytokinesis.. Science (, 2002. [DOI]
- YC Lin, D Liu, A Chakraborty, LY Kadyrova, YJ Song, Q Hao. Orc6 is a component of the replication fork and enables efficient mismatch repair.. Proc Natl Acad Sci USA (, 2022. [DOI | PubMed]
- Y Hu, L Wang, Z Li, Z Wan, M Shao, S Wu. Potential prognostic and diagnostic values of cdc6, cdc45, orc6 and snhg7 in colorectal cancer.. Onco Targets Ther (, 2019. [DOI]
- Q Pan, F Li, Y Ding, H Huang, J Guo. Orc6 acts as a biomarker and reflects poor outcome in clear cell renal cell carcinoma.. J Cancer (, 2022. [DOI]
- R Mao, Z Wang, Y Zhang, Y Chen, Q Liu, T Zhang. Development and validation of a novel prognostic signature in gastric adenocarcinoma.. Aging (Albany NY) (, 2020. [DOI]
- S Chen, Z Jin, L Xin, L Lv, X Zhang, Y Gong. Expression and clinical significance of origin recognition complex subunit 6 in breast cancer – a comprehensive bioinformatics analysis.. Int J Gen Med (, 2021. [DOI]
- W Shen, Z Song, X Zhong, M Huang, D Shen, P Gao. Sangerbox: A comprehensive, interaction-friendly clinical bioinformatics analysis platform.. iMeta (, 2022. [DOI]
- CH Mermel, SE Schumacher, B Hill, ML Meyerson, R Beroukhim, G Getz. Gistic2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers.. Genome Biol (, 2011. [DOI | PubMed]
- DS Chandrashekar, SK Karthikeyan, PK Korla, H Patel, AR Shovon, M Athar. Ualcan: an update to the integrated cancer data analysis platform.. Neoplasia (, 2022. [DOI | PubMed]
- S Huang, L Sun, P Hou, K Liu, J Wu. A comprehensively prognostic and immunological analysis of actin-related protein 2/3 complex subunit 5 in pan-cancer and identification in hepatocellular carcinoma.. Front Immunol (, 2022. [DOI | PubMed]
- B Ru, CN Wong, Y Tong, JY Zhong, SSW Zhong, WC Wu. Tisidb: an integrated repository portal for tumor-immune system interactions.. Bioinformatics (, 2019. [DOI]
- AV Balar, MD Galsky, JE Rosenberg, T Powles, DP Petrylak, J Bellmunt. Atezolizumab as first-line treatment in cisplatin-ineligible patients with locally advanced and metastatic urothelial carcinoma: A single-arm, multicentre, phase 2 trial.. Lancet (, 2017. [DOI | PubMed]
- A Lin, C Qi, T Wei, M Li, Q Cheng, Z Liu. Camoip: A web server for comprehensive analysis on multi-omics of immunotherapy in pan-cancer.. Brief Bioinform (, 2022. [DOI | PubMed]
- N Auslander, G Zhang, JS Lee, DT Frederick, B Miao, T Moll. Robust prediction of response to immune checkpoint blockade therapy in metastatic melanoma.. Nat Med (, 2018. [DOI]
- Z Zeng, CJ Wong, L Yang, N Ouardaoui, D Li, W Zhang. Tismo: syngeneic mouse tumor database to model tumor immunity and immunotherapy response.. Nucleic Acids Res (, 2022. [DOI]
- J Fu, K Li, W Zhang, C Wan, J Zhang, P Jiang. Large-scale public data reuse to model immunotherapy response and resistance.. Genome Med (, 2020. [DOI | PubMed]
- P Geeleher, N Cox, RS Huang. Prrophetic: an R package for prediction of clinical chemotherapeutic response from tumor gene expression levels.. PloS One (, 2014. [DOI | PubMed]
- D Sun, J Wang, Y Han, X Dong, J Ge, R Zheng. Tisch: A comprehensive web resource enabling interactive single-cell transcriptome visualization of tumor microenvironment.. Nucleic Acids Res (, 2021. [DOI]
- H Yuan, M Yan, G Zhang, W Liu, C Deng, G Liao. Cancersea: A cancer single-cell state atlas.. Nucleic Acids Res (, 2019. [DOI]
- J Zhu, C Luo, J Zhao, X Zhu, K Lin, F Bu. Expression of lox suggests poor prognosis in gastric cancer.. Front Med (Lausanne) (, 2021. [DOI | PubMed]
- Z Tu, J Peng, X Long, J Li, L Wu, K Huang. Sperm autoantigenic protein 17 predicts the prognosis and the immunotherapy response of cancers: A pan-cancer analysis.. Front Immunol (, 2022. [DOI | PubMed]
- J Zhu, Q Huang, S Liu, X Peng, J Xue, T Feng. Construction of a novel lncrna signature related to genomic instability to predict the prognosis and immune activity of patients with hepatocellular carcinoma.. Front Immunol (, 2022. [DOI | PubMed]
- Z Chen, C Speck, P Wendel, C Tang, B Stillman, H Li. The architecture of the DNA replication origin recognition complex in saccharomyces cerevisiae.. Proc Natl Acad Sci USA (, 2008. [DOI]
- S Saghafinia, M Mina, N Riggi, D Hanahan, G Ciriello. Pan-cancer landscape of aberrant DNA methylation across human tumors.. Cell Rep (, 2018. [DOI | PubMed]
- M Helm, Y Motorin. Detecting rna modifications in the epitranscriptome: predict and validate.. Nat Rev Genet (, 2017. [DOI]
- R Zhang, Y Guo, Z Liu, L Kong, C Li, L He. Identification of irf8 as an immune infiltration-related biomarker in hepatocellular carcinoma by bioinformatics analysis.. MedComm (2020) (, 2022. [DOI | PubMed]
- EJ Gavin, B Song, Y Wang, Y Xi, J Ju. Reduction of orc6 expression sensitizes human colon cancer cells to 5-fluorouracil and cisplatin.. PloS One (, 2008. [DOI | PubMed]
- H Chen, L Bao, J Hu, D Wu, X Tong. Orc6, negatively regulated by mir-1-3p, promotes proliferation, migration, and invasion of hepatocellular carcinoma cells.. Front Cell Dev Biol (, 2021. [DOI | PubMed]
- XK Wang, QQ Wang, JL Huang, LB Zhang, X Zhou, JQ Liu. Novel candidate biomarkers of origin recognition complex 1, 5 and 6 for survival surveillance in patients with hepatocellular carcinoma.. J Cancer (, 2020. [DOI]
- S Qu, O Qiu, Z Hu. The prognostic factors and nomogram for patients with high-grade gliomas.. Fundam Res (, 2021. [DOI]
- M Esteller. Cancer epigenomics: DNA methylomes and histone-modification maps.. Nat Rev Genet (, 2007. [DOI]
- C Lahtz, GP Pfeifer. Epigenetic changes of DNA repair genes in cancer.. J Mol Cell Biol (, 2011. [DOI]
- I Barbieri, T Kouzarides. Role of rna modifications in cancer.. Nat Rev Cancer (, 2020. [DOI]
- D Hanahan, LM Coussens. Accessories to the crime: functions of cells recruited to the tumor microenvironment.. Cancer Cell (, 2012. [DOI]
- S-W Zhang, H Wang, X-H Ding, Y-L Xiao, Z-M Shao, C You. Bidirectional crosstalk between therapeutic cancer vaccines and the tumor microenvironment: beyond tumor antigens.. Fundam Res (, 2022. [DOI]
- Y Gao, M Shen, X Shi. Interaction of dendrimers with the immune system: an insight into cancer nanotheranostics.. View (, 2021. [DOI]
- RA Flavell, S Sanjabi, SH Wrzesinski, P Licona-Limon. The polarization of immune cells in the tumour environment by tgfbeta.. Nat Rev Immunol (, 2010. [DOI]
- S Lario, MJ Ramirez-Lazaro, D Sanjuan-Herraez, A Brunet-Vega, C Pericay, L Gombau. Plasma sample based analysis of gastric cancer progression using targeted metabolomics.. Sci Rep (, 2017. [DOI | PubMed]
- SA Rosenberg, NP Restifo. Adoptive cell transfer as personalized immunotherapy for human cancer.. Science (, 2015. [DOI]
- X Zhang, X Liu, Y Guo, FG Wu. Strategies for visualizing inflammation.. View (, 2021. [DOI]
- B Lv, Y Wang, D Ma, W Cheng, J Liu, T Yong. Immunotherapy: reshape the tumor immune microenvironment.. Front Immunol (, 2022. [DOI | PubMed]
- AM van der Leun, DS Thommen, TN Schumacher. Cd8(+) T cell states in human cancer: insights from single-cell analysis.. Nat Rev Cancer (, 2020. [DOI]
- DS Vinay, EP Ryan, G Pawelec, WH Talib, J Stagg, E Elkord. Immune evasion in cancer: mechanistic basis and therapeutic strategies.. Semin Cancer Biol (, 2015. [DOI]
- J Verneau, C Sautes-Fridman, CM Sun. Dendritic cells in the tumor microenvironment: prognostic and theranostic impact.. Semin Immunol (, 2020. [DOI | PubMed]
- MJ Chusid. Eosinophils: friends or foes?. J Allergy Clin Immunol Pract (, 2018. [DOI]
- S Ugel, S Cane, F De Sanctis, V Bronte. Monocytes in the tumor microenvironment.. Annu Rev Pathol (, 2021. [DOI | PubMed]
- Y Gu, Y Chen, K Jin, Y Cao, X Liu, K Lv. Intratumoral cd103(+)Cd4(+) T cell infiltration defines immunoevasive contexture and poor clinical outcomes in gastric cancer patients.. Oncoimmunology (, 2020. [DOI | PubMed]
- WH Fridman, F Pages, C Sautes-Fridman, J Galon. The immune contexture in human tumours: impact on clinical outcome.. Nat Rev Cancer (, 2012. [DOI | PubMed]
- Z Jiang, H Zhu, P Wang, W Que, L Zhong, XK Li. Different subpopulations of regulatory T cells in human autoimmune disease, transplantation, and tumor immunity.. MedComm (2020) (, 2022. [DOI | PubMed]
- DC Hinshaw, LA Shevde. The tumor microenvironment innately modulates cancer progression.. Cancer Res (, 2019. [DOI]
- A Mantovani, P Allavena, A Sica, F Balkwill. Cancer-related inflammation.. Nature (, 2008. [DOI]
- LA Liotta, EC Kohn. The microenvironment of the tumour-host interface.. Nature (, 2001. [DOI]
- Y Bian, Y Wang, X Chen, Y Zhang, S Xiong, D Su. Image-guided diagnosis and treatment of glioblastoma.. View (, 2023. [DOI]
- H Zhu, C Yang, A Yan, W Qiang, R Ruan, K Ma. Tumor-targeted nano-adjuvants to synergize photomediated immunotherapy enhanced antitumor immunity.. View (, 2023. [DOI]
