Molecular Characteristics, Oncogenic Roles, and Relevant Immune and Pharmacogenomic Features of EVA1B in Colorectal Cancer
Abstract
Objective:
EVA1B, a protein coding gene, is a critical paralog of EVA1A gene. Herein, our study was conducted to investigate the role of EVA1B in colorectal cancer (CRC) progression and prognosis.
Methods:
Pan-cancer analysis was conducted to analyze expression, genetic and epigenetic alterations, and immunological characteristics of EVA1B. Especially, immunological characteristics and mutational landscape were compared between high and low EVA1B expression groups in the combined TCGA-COAD and TCGA-READ datasets. Through random survival forest analysis, an EVA1B-derived genomic model was developed, and its prognostic value was verified in the external datasets (GSE14333, GSE39582, and GSE87211). Drug sensitivity was compared between high- and low-risk subpopulations. A nomogram was conducted through integrating independent factors.
Results:
EVA1B expression presented a remarkable upregulation in most cancer types, especially CRC. EVA1B expression was significantly correlated to DNA methyltransferases, DNA mismatch repair genes, m6A regulators, TMB, and MSI across pan-cancer. High EVA1B expression indicated an undesirable CRC patients’ prognosis. Additionally, its upregulation was correlated to enhanced immune cell infiltration, increased stromal and immune activation, and elevated activities of cancer immunity cycle. Higher frequencies of amplification and deletion were investigated in high EVA1B expression subpopulation. Following verification, the EVA1B-derived genomic model reliably predicted patients’ prognosis and drug responses. The nomogram (age, stage, EVA1B-derived risk score) was conducted to quantify an individual’s survival probability. Furthermore, our experimental validation based on immunohistochemistry indicated that EVA1B overexpression is correlated with CRC tumorigenesis and poor outcomes in our CRC patients’ cohort.
Conclusion:
Collectively, our findings provided valuable resource for guiding the mechanisms and therapeutic analysis of EVA1B in CRC.
Article type: Research Article
Keywords: colorectal cancer, EVA1B, prognosis, tumor immunity, pharmacogenomic features
Affiliations: Department of Colorectal Surgery, Cancer Hospital of China Medical University, Liaoning Cancer Hospital and Institute, Shenyang, China; Department of Organ Transplantation and Hepatobiliary, the First Affiliated Hospital of China Medical University, Shenyang, China
License: Copyright © 2022 Ma, Wang, Liang, Meng and Li CC BY 4.0 This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
Article links: DOI: 10.3389/fimmu.2022.809837 | PubMed: 35250982 | PMC: PMC8888821
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (29.8 MB)
Introduction
Colorectal cancer (CRC) involving the colon and rectum ranks the third leading cause of cancer mortality globally, with over 1.85 million cases and 850,000 deaths each year (ref. 1). Recently, CRC presents increasing incidence among younger cases (ref. 2). This disease is a heterogeneous disease with distinct pathogenesis mechanisms, involving somatic mutation, genetic fusion, genetic instability, as well as epigenetic alteration (ref. 3). Among newly diagnosed CRC, 20% of cases have occurring metastasis at diagnosis as well as 25% will metastasize following localized disease (ref. 1). Surgical resection is the major therapeutic regimen of CRC. Nevertheless, there are very few treatment regimens for metastatic patients. Although chemotherapy is usually recommended, merely few targeted therapies appropriate for cases who have specific mutation profiling are available, such as EGFR inhibitor and VEGF inhibitor (ref. 3). Hence, it is urgently required to develop novel molecule targets against CRC.
Immunotherapy is a novel alternative against CRC treatment, which utilizes cases’ own immune system to combat tumor cells (ref. 4, ref. 5). A minority of CRC cases present microsatellite instability (MSI), a molecule predictor of defective DNA mismatch repair, and a predictive biomarker of immunotherapeutic response, but most of them are microsatellite-stable (MSS) (ref. 6). Enhanced tumor mutational burden (TMB) and neoantigen load in MSI cancers sustain the infiltrations of immune effector subpopulations, and anti-cancer immune response is strongly in relation to MSS counterparts (ref. 6). Nevertheless, a few MSS cancers present increased TMB as well as infiltrating immune cells to respond to immunotherapy. Hence, novel predictors of immunotherapeutic responses will be required. EVA1B is a protein coding gene, which is a critical paralog of EVA1A gene. It is an endoplasmic reticulum and lysosome-relevant protein involving autophagy and apoptosis (ref. 7). Previously, it triggers papillary thyroid carcinogenesis and epithelial-mesenchymal transition (EMT) through Hippo signaling (ref. 8). Flubendazole exerts an antitumor role through modulating EVA1A-mediated autophagy and apoptosis in breast carcinoma (ref. 9). MiR-125b relieves oxaliplatin resistance in liver carcinoma through negatively modulating EVA1A-mediated autophagy (ref. 10). EVA1B possesses high sequence similar to EVA1A gene, and EVA1B protein presents the similar domain to EVA1A protein (ref. 11). Nevertheless, the role of EVA1B in CRC remains indistinct. This study was conducted to investigate the correlation of EVA1B expression with prognosis, tumor immune and pharmacogenomic features in CRC.
Materials and Methods
Pan-Cancer Analyses
RNA sequencing data (FPKM), somatic mutational data [Mutation Annotation Format (MAF)], copy number variation (CNV) data, and clinical information of 33 cancer types [adrenocortical carcinoma (ACC); bladder urothelial carcinoma (BLCA); breast invasive carcinoma (BRCA); cervical squamous cell carcinoma and endocervical adenocarcinoma (CESC); cholangiocarcinoma (CHOL); colon adenocarcinoma (COAD); lymphoid neoplasm diffuse large B-cell lymphoma (DLBC); esophageal carcinoma (ESCA); glioblastoma multiforme (GBM); head and neck squamous cell carcinoma (HNSC); kidney chromophobe (KICH); kidney renal clear cell carcinoma (KIRC); kidney renal papillary cell carcinoma (KIRP); acute myeloid leukemia (LAML); lower-grade glioma (LGG); liver hepatocellular carcinoma (LIHC); lung adenocarcinoma (LUAD); lung squamous cell carcinoma (LUSC); mesothelioma (MESO); ovarian serous cystadenocarcinoma (OV); pancreatic adenocarcinoma (PAAD); pheochromocytoma and paraganglioma (PCPG); prostate adenocarcinoma (PRAD); rectum adenocarcinoma (READ); sarcoma (SARC); skin cutaneous melanoma (SKCM); stomach adenocarcinoma (STAD); testicular germ cell tumors (TGCT); thyroid carcinoma (THCA); thymoma (THYM); uterine corpus endometrial carcinoma (UCEC); uterine carcinosarcoma (UCS); uveal melanoma (UVM)] were curated from The Cancer Genome Atlas (TCGA) through the GDC data portal (https://portal.gdc.cancer.gov/). Tumor Immune Estimation Resource (TIMER; version 2.0) web tool (http://timer.cistrome.org/) was adopted to analyze the expression of EVA1B across 33 cancer types (ref. 12). For TCGA dataset, FPKM value was transformed to TPM value. The expression profiling of four major DNA methyltransferases (DNMT1, DNMT2, DNMT3A, and DNMT3B), DNA mismatch repair genes (MLH1, MSH2, MSH6, PMS2, and EPCAM), and m6A regulators (CBLL1, VIRMA, METTL3/14, RBM15/15B, WTAP, ZC3H13, ALKBH5, FTO, ELAVL1, FMR1, HNRNPA2B1, HNRNPC, IGF2BP1/2/3, LRPPRC, YTHDC1/2, and YTHDF1/2/3) was extracted from pan-cancer specimens. TMB and MSI of pan-cancer were also harvested from TCGA project. Spearman correlation analysis was conducted to investigate the correlation of EVA1B with above factors across pan-cancer.
Estimation of Immunological Features
The abundance of tumor-infiltrating immune cells was inferred with single-sample gene set enrichment analysis (ssGSEA) derived from gene set variation analysis (GSVA) package (ref. 13). In total, 122 immunomodulatory factors containing MHCs, receptors, chemokines, and immune stimulators were collected from Charoentong et al. (ref. 14). Additionally, immune checkpoint molecules were retrieved from the study of Auslander et al. (ref. 15). Overall infiltration of stromal and immune cells was estimated in CRC tissues with estimation of stromal and immune cells in malignant tumors using expression data (ESTIMATE) algorithm in accordance with mRNA expression data (ref. 16). Cancer immunity cycle was curated from previous research (ref. 17), and the activities of all steps were estimated with ssGSEA (ref. 18).
Collection of CRC Datasets
The GSE14333 (ref. 19), GSE39582 (ref. 20), and GSE87211 (ref. 21) datasets were collected from the Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) project. Batch effects were adjusted with ComBat algorithm derived from sva package (ref. 22). Somatic mutational data were visualized with maftools package (ref. 23). GISTIC2.0 was adopted to analyze amplification and deletion using CNV data (ref. 24). Raw “CEL” files of microarray profiling from Affymetrix platform were downloaded, followed by background correction and quantile normalization via robust multiarray averaging algorithm utilizing affy and simpleaffy packages (ref. 25, ref. 26). Meanwhile, normalized matrix files of microarray profiling from other platforms were directly curated.
Curation of Gene Sets of Known Biological Processes
Gene sets of known biological processes containing CD8+ T effector, DNA damage repair, pan-fibroblast TGF-β response signature (pan-F-TBRS), antigen processing machinery, immune checkpoints, EMT1-3, FGFR3-relevant gene signatures, angiogenesis, KEGG discovered histones, Fanconi anemia, cell cycle, DNA replication, nucleotide excision repair, homologous recombination, mismatch repair, WNT target, as well as cell cycle regulators were curated from previous research (ref. 27–ref. 29). The activities of biological processes were quantified with ssGSEA algorithm.
Functional Enrichment Analysis
GSEA was conducted to investigate the differences in signaling pathways activated in two subpopulations, with the gene set “c2.cp.kegg.v6.2.symbols.gmt” as the reference. Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of EVA1B-relevant genes was conducted with clusterProfiler package (ref. 30). GO categories contained biological process (BP), cellular component (CC), and molecular function (MF). Hallmark gene sets were harvested from the Molecular Signatures Database (ref. 31), and their activities were quantified with ssGSEA algorithm.
Screening EVA1B-Relevant Genes
To identify EVA1B-relevant genes, CRC patients were classified into high and low EVA1B expression subpopulations with the mean value of EVA1B expression. Through empirical Bayesian method from limma package (ref. 32), differentially expressed genes between subpopulations were determined. The significance criteria of EVA1B-relevant genes were set as |fold-change| >1.5 and adjusted p-value <0.01.
Exploitation of an EVA1B-Derived Genomic Model
Through univariate Cox regression models, interaction of EVA1B-relevant genes with CRC prognosis was estimated in TCGA cohort. EVA1B-relevant genes with p-value <0.05 were determined as prognostic factors, which were input into random survival forest analysis (ref. 33). The number of Monte Carlo iterations was set as 100, and the number of steps forward was 5. Thereafter, their relative importance was ranked. In accordance with relative importance >0.28, characteristic EVA1B-relevant genes were determined, which were input into a multivariate Cox regression model. The formula of EVA1B-relevant genomic model was conducted as follows: risk score = , in which n indicated the number of characteristic EVA1B-relevant genes, Expi indicated the expression of characteristic genes, and eHRi indicated the regression coefficients of genes derived from the multivariate Cox regression analysis. In accordance with this formula, risk score of each CRC patient was calculated. Thereafter, CRC patients were clarified into high- and low-risk subpopulations following the mean value of risk score. The risk score distribution, survival state, and heatmap of the expression of characteristic EVA1B-relevant genes were drawn following patients’ risk score. Kaplan–Meier curves of overall survival (OS), disease-specific survival (DSS), and progression-free survival (PFS) were depicted between high- and low-risk subpopulations using survival and survminer packages, followed by log-rank test. Thereafter, receiver operating characteristic (ROC) curves of 1-, 3-, and 5-year survival were established utilizing survivalROC package, and area under the curve (AUC) was calculated to assess the discrimination (ref. 34). Additionally, prognostic value of the EVA1B-derived genomic model was externally verified in the GSE14333, GSE39582, and GSE87211 cohorts.
Assessment of Drug Sensitivity
The Genomics of Drug Sensitivity in Cancer (GDSC) project (https://www.cancerrxgene.org/) offers drug sensitivity information of 138 anticancer agents across approximately 75,000 experiments in 700 cancer cell lines (ref. 35). The half-maximal inhibitory concentration (IC50) that represented drug response was estimated with pRRophetic package (ref. 36).
Nomogram Establishment
Uni- and multivariate Cox regression models were established to evaluate the associations of clinical indicators (age, gender, stage, T, N, and M) and EVA1B-relevant risk score with CRC patients’ OS. ROC curves were conducted to evaluate the prognostic value of this nomogram. Additionally, calibration curves were utilized to assess the consistency between the actual and nomogram-predicted survival probabilities.
Clinical Specimens and Data Source
Formalin-fixed, paraffin-embedded specimens, including primary carcinoma specimens (n = 131), corresponding noncancerous normal tissues (n = 19) and liver metastasis specimens (n = 19) for immunohistochemistry (IHC) were obtained from 131 CRC patients who underwent surgery between 2014 and 2015. CRC primary carcinomas were assessed according to the 8th edition of the American Joint Committee on Cancer (AJCC) staging system. CRC patient data and tissue samples were obtained from the Liaoning Cancer Hospital. Patients included in the current study did not receive preoperative radiotherapy or chemotherapy prior to the study. The current study (20210804GP) was approved by the Ethics Committee of the Liaoning Cancer Hospital, Shenyang, China.
Immunohistochemistry
EVA1B expression levels in samples were determined by IHC. Tissue sections were incubated with EVA1B antibody (Manufacturer: Atlas Antibodies; Catalog number: HPA043537). The level of EVA1B expression was determined by counting the percentages of positively stained immunoreactive cells and evaluating cell staining intensity. EVA1B IHC staining was scored as “−” ~ “+” and “++” ~ “+++,” which represented negative and positive, respectively. All samples were reviewed by two independent, experienced pathologists in who were blinded to the identity of the samples.
Statistical Analysis
Statistical analysis was implemented with R software (v3.4.1; https://www.r-project.org/) and its appropriate packages. Comparison between groups was conducted utilizing Student’s t– or Wilcoxon rank-sum test. Spearman correlation test was adopted to determine the interactions between variables. Chi-square test was performed to analyze the correlation between EVA1B and clinicopathologic characteristics. Kaplan–Meier curves and the log-rank test were used to compare the OS between EVA1B expression level. The prognostic ability of the predictors for OS was evaluated by ROC curves and the AUC values. Univariate and multivariate Cox regression analyses were utilized to evaluate the independent prognostic value of EVA1B regarding OS. p-value <0.05 was set as the threshold.
Results
Analysis of Expression, Genetic and Epigenetic Alterations, and Immunological Characteristics of EVA1B Across Pan-Cancer
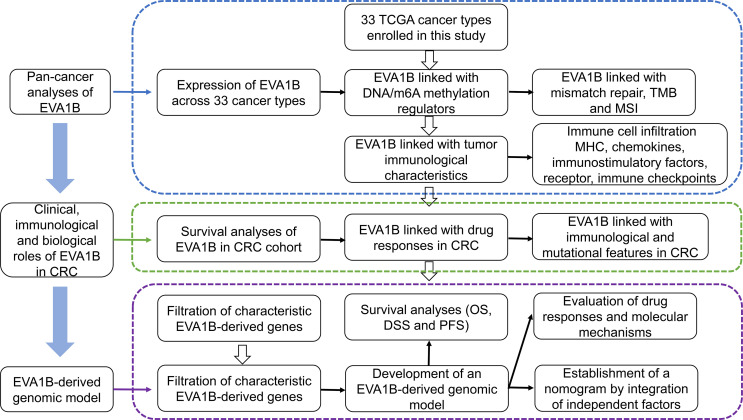
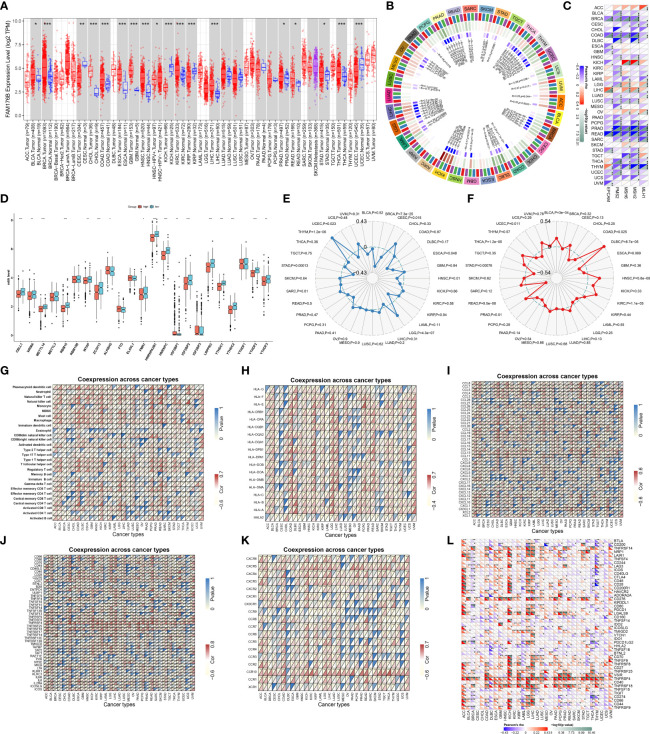
f1 depicts the workflow of this study. EVA1B, as known as FAM176B, displayed remarkably increased expression in most cancer types in comparison with corresponding normal tissues, containing BLCA, BRCA, CHOL, COAD, ESCA, GBM, HNSC, KIRC, LIHC, PRAD, READ, STAD, and THCA (f2). Differently, downregulated EVA1B was investigated in CESC, KICH, KIRP, and UCEC. DNA methyltransferases exert critical roles in changing chromatin structure and gene expression. We noted that EVA1B presented negative interactions with four major DNA methyltransferases containing DNMT1, DNMT2, DNMT3A, and DNMT3B in most cancer types (f2). Additionally, EVA1B was negatively correlated to DNA mismatch repair genes (MLH1, MSH2, MSH6, PMS2, and EPCAM) across cancer types (f2). M6A methylation represents the most common RNA modification, which affects the complexity of cancer progression (ref. 37). Especially, we focused on the interactions of EVA1B with m6A regulators in CRC. With the mean value of EVA1B expression, CRC patients were clustered into high and low EVA1B expression subpopulations. As depicted in f2, downregulated EVA1B was correlated to most m6A regulators like CBLL1, VIRMA, METTL14, and RBM15. TMB becomes a reliable biomarker of sensitivity to immune checkpoint blockage (ref. 38). In f2, EVA1B presented remarkably positive interaction with TBM across THYM, and LGG. Oppositely, it was negatively correlated with TMB across BRCA, CESC, ESCA, HNSC, STAD, and UCEC. MSI is a hypermutation phenotype caused by frequent polymorphisms in short repetitive DNA sequence as well as single nucleotide substitution due to DNA MMR defects (ref. 39). We noted the positive interaction of EVA1B with MSI in THCA, BLCA, DLBC, HNSC, KIRC, and PRAD but negative interaction with MSI in COAD, READ, STAD, and UCEC (f2). Thereafter, we analyzed the immunological characteristics of EVA1B across pan-cancer. Our data suggested that EVA1B was remarkably correlated to immune cell infiltration (f2), MCH molecules (f2), chemokines (f2), immunostimulatory factors (f2), receptors (f2), and immune checkpoint molecules (f2) across pan-cancer.
Prognostic Significance of EVA1B and Its Association With Drug Sensitivity in CRC
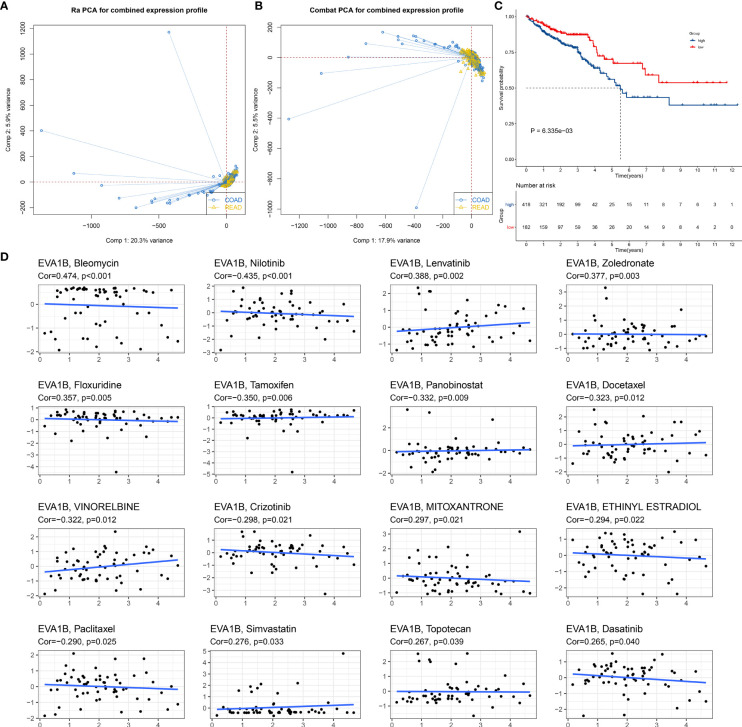
We also focused on the biological role and significance of EVA1B in CRC. We combined the COAD and READ datasets from TCGA project and removed batch effects (f3). Survival analysis was indicative that patients with high EVA1B expression possessed more undesirable OS outcomes than those with low EVA1B expression (f3). Targeted therapies play a key role in CRC management, and genetic alterations that attribute to CRC heterogeneity is correlated to the responses to targeted therapies (ref. 40). Reliable predictive biomarkers for targeted therapies remain greatly lacking. Thus, we evaluated whether EVA1B expression was correlated to drug sensitivity. As a result, EVA1B presented positive association with sensitivity to bleomycin, lenvatinib, zoledronate, floxuridine, mitoxantrone, simvastatin, topotecan, and dasatinib (f3). Differently, EVA1B expression was negatively correlated to sensitivity to nilotinib, tamoxifen, panobinostat, docetaxel, vinorelbine, crizotinib, ethinyl estradiol, and paclitaxel in CRC. Therefore, EVA1B might be a predictive marker of above agents.
Immunological and Biological Significance of EVA1B in CRC
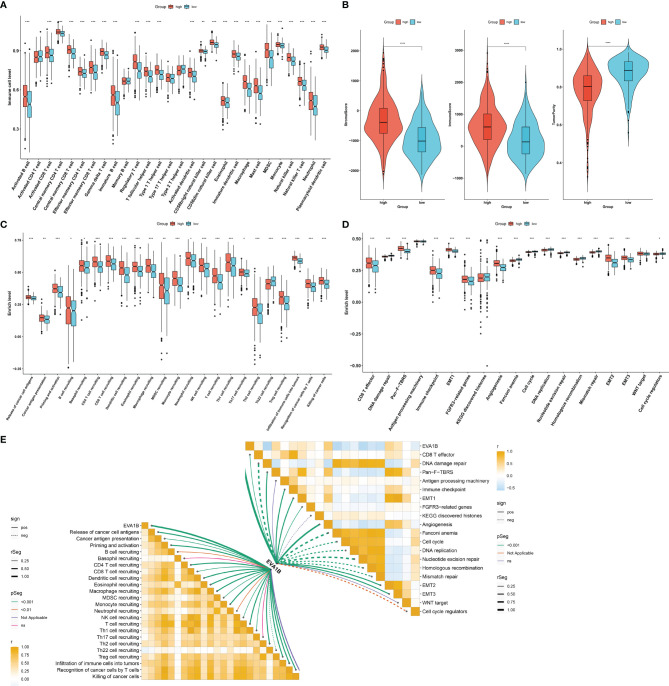
Our ssGSEA results demonstrated the remarkably increased infiltration levels of most immune cells in high EVA1B expression subpopulation (f4). Additionally, high EVA1B expression was correlated to increased stromal and immune score as well as reduced tumor purity in CRC (f4). There were prominently enhanced activities of most steps within cancer immunity cycle in high EVA1B expression subpopulation (f4). We also investigated the increased activities of immune activation processes (like CD8+ T effector, and immune checkpoint) and stromal activation processes (like EMT, pan-F-TBRS, FGFR3-related genes, and angiogenesis) in high EVA1B expression subpopulation (f4). Additionally, the interactions of EVA1B expression with cancer immunity cycle and known biological processes were evaluated across CRC specimens. In f4, EVA1B expression presented positive correlations to most steps within cancer immunity cycle as well as immune and stromal activation processes.
Association of EVA1B With Mutational Landscape in CRC
In f5, we evaluated the prevalence of somatic mutation in high and low EVA1B expression subpopulations. APC, TP53, and TTN ranked the first three mutational genes. Nevertheless, no prominent difference in somatic mutation was investigated in high and low EVA1B expression subpopulations. The GISTIC2.0 results demonstrated that amplification and deletion displayed higher frequencies in high EVA1B expression subpopulation (f5) compared with low EVA1B expression subpopulation (f5). Additionally, we calculated the G-score in accordance with the amplitude of the aberrations and the frequency of their incidence in CRC specimens. Following comparison, mutations occurred in more regions in high EVA1B expression subpopulation (f5).
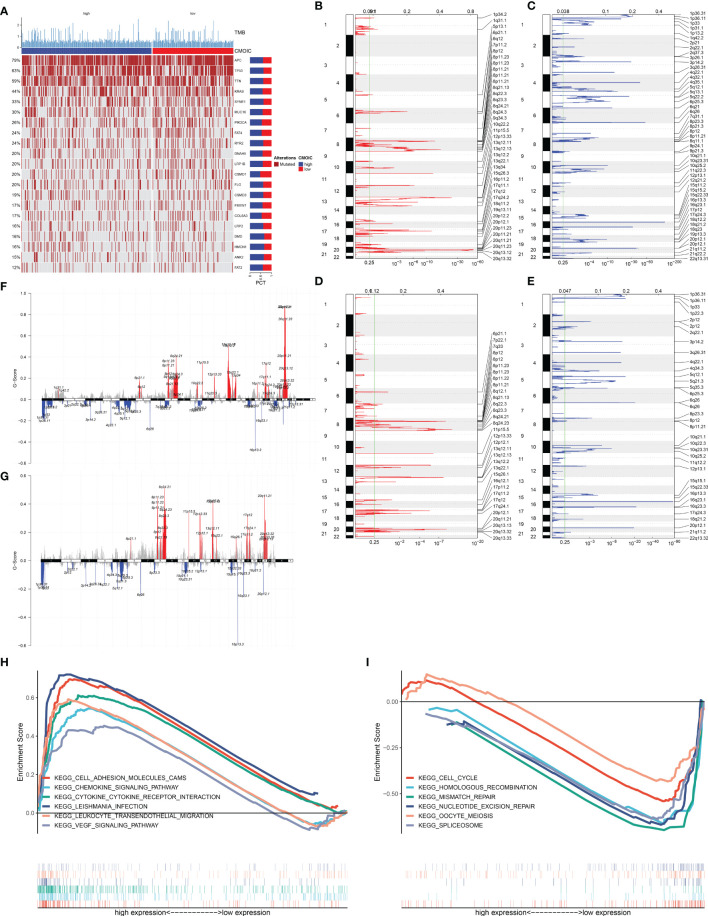
Signaling Pathways Associated With EVA1B
We further analyzed the signaling pathways involving EVA1B via GSEA. In f5, high EVA1B expression was positively correlated to cell adhesion molecules (CAMs), chemokine signaling pathways, cytokine–cytokine receptor interaction, Leishmania interaction, leukocyte trans-endothelial migration, and VEGF signaling pathways. Meanwhile, low EVA1B expression was negatively correlated to cell cycle, homologous recombination, mismatch repair, nucleotide excision repair, oocyte meiosis, and spliceosome (f5). The above data showed that EVA1B was involved in several oncogenic pathways, indicating the oncogenic role of EVA1B.
Identification of EVA1B-Derived Genes and Their Biological Significance
Through the cutoffs of |fold-change| >1.5 and adjusted p-value <0.01, we determined 602 EVA1B-derived genes in CRC individuals (Supplementary Table S1). Their biological significance was further investigated. In f6, EVA1B-derived genes were mainly correlated to immune response in accordance with GO annotation results. Additionally, there were remarkable interactions of EVA1B-derived genes with tumorigenic pathways (such as PI3K-Akt signaling pathway, proteoglycans in cancer, and ECM-receptor interaction) and immune activation pathways (such as IL-17 signaling pathway, Th17-cell differentiation, cytokine–cytokine receptor interaction, Th1- and Th2-cell differentiation, antigen processing and presentation, intestinal immune network for IgA production, complement and coagulation cascades; f6). Overall, EVA1B-derived genes might exert remarkable roles in CRC progression.
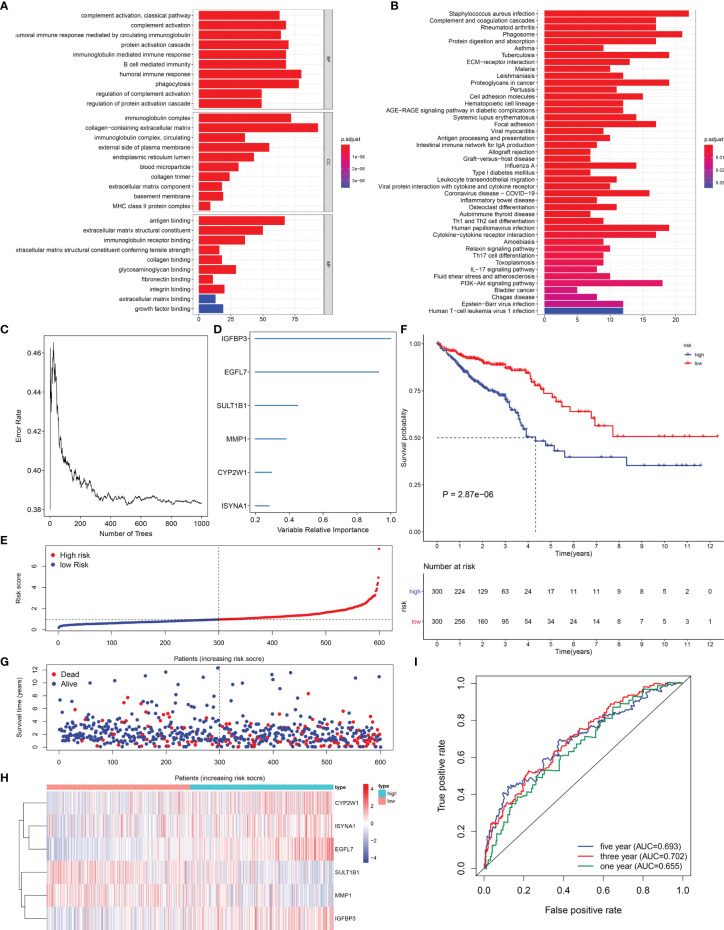
The EVA1B-Derived Genomic Model for CRC Prognosis
In total, 66 EVA1B-derived genes were prominently correlated to CRC patients’ prognosis utilizing univariate-Cox regression models (T1). With random survival forest analysis, we ranked the relative importance of above prognostic EVA1B-derived genes (f6). In accordance with relative importance >0.28, six characteristic EVA1B-derived genes were determined, containing IGFBP3, EGFL7, SULT1B1, MMP1, CYP2W1, as well as ISYNA1 (f6). On the basis of them, a multivariate-Cox regression model was conducted and risk scoring system was developed following the formula: risk score = (−0.117800092) * ISYNA1 expression + 0.067856101 * CYP2W1 expression + (−0.145078003) * MMP1 expression + (−0.107319254) * SULT1B1 expression + 0.427836868 * EGFL7 expression + 0.188926758 * IGFBP3 expression. With the mean value, we classified CRC patients into high- and low-risk subpopulations (f6). Survival analysis uncovered that low-risk CRC individuals displayed a remarkable survival advantage (f6). Nevertheless, no distinct difference in survival status was noted between high- and low-risk subpopulations (f6). Additionally, the heterogeneity in expression of IGFBP3, EGFL7, SULT1B1, MMP1, CYP2W1, as well as ISYNA1 was displayed in two subpopulations (f6). ROC curves confirmed that the EVA1B-derived genomic model possessed the well potency in estimating 1-, 3-, and 5-year OS probabilities (f6).
Table 1: Univariate-Cox regression models determine prognostic EVA1B-derived genes in CRC.
| Gene | HR | Lower 95% CI | Upper 95% CI | p-value | Gene | HR | Lower 95% CI | Upper 95% CI | p-value |
|---|---|---|---|---|---|---|---|---|---|
| EVA1B | 1.2498 | 1.0165 | 1.5367 | 0.0344 | ADAM8 | 1.3206 | 1.0966 | 1.5904 | 0.0034 |
| RCN3 | 1.2516 | 1.0360 | 1.5122 | 0.0200 | SFRP2 | 1.1005 | 1.0178 | 1.1900 | 0.0163 |
| UBTD1 | 1.4292 | 1.1012 | 1.8549 | 0.0073 | GPX3 | 1.2600 | 1.0864 | 1.4613 | 0.0022 |
| C11orf96 | 1.2038 | 1.0175 | 1.4243 | 0.0306 | SCX | 1.2693 | 1.0257 | 1.5707 | 0.0283 |
| SIPA1 | 1.5314 | 1.1049 | 2.1226 | 0.0105 | CAVIN1 | 1.2125 | 1.0151 | 1.4484 | 0.0336 |
| CRIP2 | 1.2970 | 1.0562 | 1.5927 | 0.0131 | RGCC | 1.2466 | 1.0301 | 1.5085 | 0.0235 |
| LTBP3 | 1.3106 | 1.0673 | 1.6093 | 0.0098 | ISYNA1 | 1.2025 | 1.0062 | 1.4372 | 0.0426 |
| CERCAM | 1.1983 | 1.0009 | 1.4346 | 0.0489 | ZNF385A | 1.3667 | 1.1269 | 1.6574 | 0.0015 |
| APOE | 1.1119 | 1.0025 | 1.2333 | 0.0448 | TPM2 | 1.2088 | 1.0411 | 1.4034 | 0.0128 |
| RAB3IL1 | 1.3854 | 1.0918 | 1.7580 | 0.0073 | SPOCK1 | 1.1938 | 1.0173 | 1.4010 | 0.0300 |
| FSTL3 | 1.3310 | 1.1200 | 1.5818 | 0.0012 | G0S2 | 1.1966 | 1.045 | 1.3700 | 0.0094 |
| RGS19 | 1.3237 | 1.0551 | 1.6609 | 0.0154 | GAS1 | 1.1596 | 1.0018 | 1.3423 | 0.0472 |
| IER5L | 1.3062 | 1.0439 | 1.6344 | 0.0195 | MGP | 1.1437 | 1.0137 | 1.2902 | 0.0291 |
| HSD17B14 | 1.3673 | 1.0590 | 1.7655 | 0.0164 | CRYAB | 1.2665 | 1.0729 | 1.4951 | 0.0053 |
| BGN | 1.1667 | 1.0226 | 1.3312 | 0.0219 | CNN1 | 1.1161 | 1.0099 | 1.2333 | 0.0312 |
| GJA4 | 1.2785 | 1.0061 | 1.6246 | 0.0445 | IGFBP3 | 1.2996 | 1.082 | 1.5611 | 0.0051 |
| EGFL7 | 1.4972 | 1.2150 | 1.8449 | 0.0002 | SCT | 1.1864 | 1.0233 | 1.3754 | 0.0235 |
| RTL8B | 1.3178 | 1.0243 | 1.6954 | 0.0318 | DEPP1 | 1.1990 | 1.0193 | 1.4103 | 0.0284 |
| NOTCH3 | 1.2565 | 1.0292 | 1.5341 | 0.0249 | CCL11 | 0.8402 | 0.7174 | 0.9841 | 0.0309 |
| MYL9 | 1.1557 | 1.0126 | 1.3190 | 0.0319 | ARL4C | 1.2237 | 1.0249 | 1.461 | 0.0256 |
| TIMP1 | 1.5239 | 1.2290 | 1.8895 | 0.0001 | LY6E | 1.1578 | 1.0031 | 1.3363 | 0.0452 |
| PRRX2 | 1.2312 | 1.0121 | 1.4977 | 0.0375 | AOC3 | 1.2133 | 1.0483 | 1.4043 | 0.0095 |
| GPC1 | 1.3725 | 1.1201 | 1.6817 | 0.0023 | RAMP1 | 1.1366 | 1.0276 | 1.2571 | 0.0128 |
| PPP1R14A | 1.2016 | 1.0010 | 1.4424 | 0.0487 | TNS1 | 1.1874 | 1.0242 | 1.3766 | 0.0228 |
| HOMER3 | 1.5016 | 1.1820 | 1.9078 | 0.0009 | SPP1 | 1.0872 | 1.0002 | 1.1817 | 0.0495 |
| YPEL3 | 1.3306 | 1.0722 | 1.6512 | 0.0095 | SULT1B1 | 0.8293 | 0.7173 | 0.9588 | 0.0115 |
| SERPING1 | 1.1556 | 1.0055 | 1.3281 | 0.0417 | PRELP | 1.2236 | 1.0686 | 1.401 | 0.0035 |
| CHPF | 1.3780 | 1.0874 | 1.7461 | 0.0080 | RTL8A | 1.1600 | 1.0017 | 1.3435 | 0.0475 |
| FHL3 | 1.3222 | 1.0481 | 1.6679 | 0.0184 | IGLV7-43 | 0.8716 | 0.7825 | 0.9708 | 0.0124 |
| VSIG4 | 1.1643 | 1.0062 | 1.3472 | 0.0410 | MMP1 | 0.8915 | 0.8164 | 0.9736 | 0.0106 |
| COMP | 1.1434 | 1.0333 | 1.2651 | 0.0095 | RNU4-2 | 0.8745 | 0.7652 | 0.9995 | 0.0492 |
| RTL8C | 1.3160 | 1.0979 | 1.5774 | 0.0030 | CYP2W1 | 1.0957 | 1.0049 | 1.1947 | 0.0384 |
| IRF7 | 1.3400 | 1.0801 | 1.6623 | 0.0078 | RN7SL2 | 0.9108 | 0.8295 | 1.0000 | 0.0500 |
Verification of Prognostic Significance of EVA1B-Derived Genomic Model in CRC
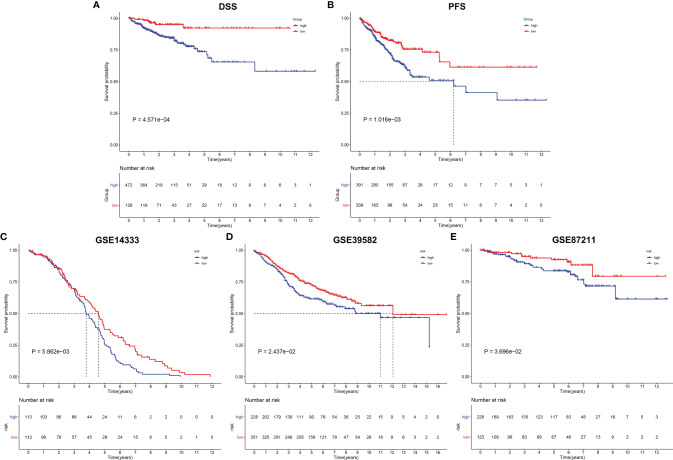
Except for OS, our analysis demonstrated that high-risk subpopulation was indicative of more undesirable DSS and PFS outcomes in comparison with low-risk subpopulation in TCGA cohort (f7). The prognostic significance of the EVA1B-derived genomic model was verified in external cohorts. Our data confirmed that high-risk score was predictive of unfavorable OS outcome in the GSE14333, GSE39582, and GSE87211 datasets (f7).
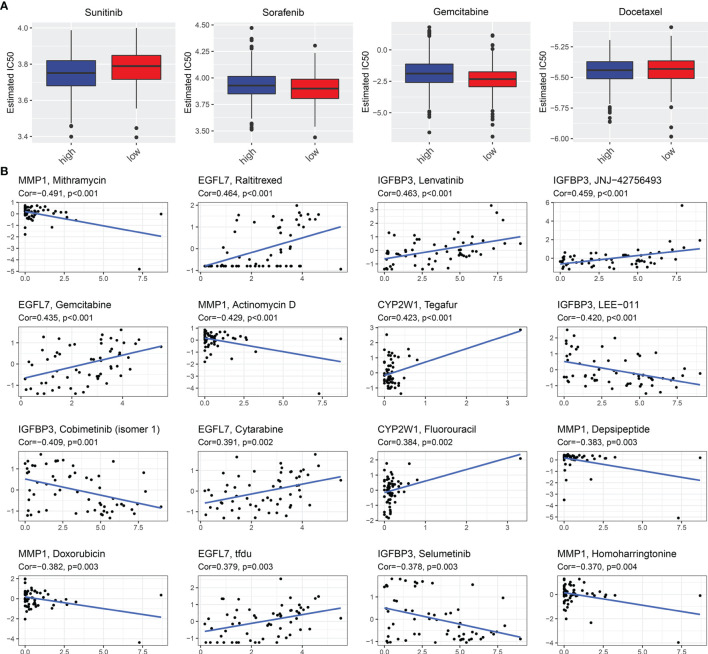
The EVA1B-Derived Genomic Model Predicts Drug Responses of CRC Patients
Further analysis was conducted to uncover whether this EVA1B-derived genomic model was predictive of drug responses of CRC individuals. In f8, we noted that high-risk group had significantly lower estimated IC50 of sunitinib and docetaxel relative to low-risk group, indicating that high-risk subpopulations presented higher sensitivity to sunitinib and docetaxel. Meanwhile, the low-risk group showed significantly lower estimated IC50 of sorafenib and gemcitabine, indicating that low-risk subpopulations were more likely to respond to sorafenib and gemcitabine. We also evaluated the interactions of EVA1B-derived genes with drug responses. As a result, MMP1 was negatively correlated to IC50 of mithramycin, actinomycin D, depsipeptide, and homoharringtonine; EGFL7 presented positive correlation to IC50 of raltitrexed, gemcitabine, cytarabine, and TFdU; IGFBP3 was positively correlated to IC50 of lenvatinib and JNJ-42756493 but negatively correlated to IC50 of LEE-011, cobimetinib (isomer 1), and selumetinib; CYP2W1 displayed positive relationship to IC50 of tegafur and fluorouracil (f8). These data indicated that EVA1B-derived genes might be correlated to sensitivity to above agents.
Establishment of a Reliable Nomogram for Prediction of CRC Prognosis
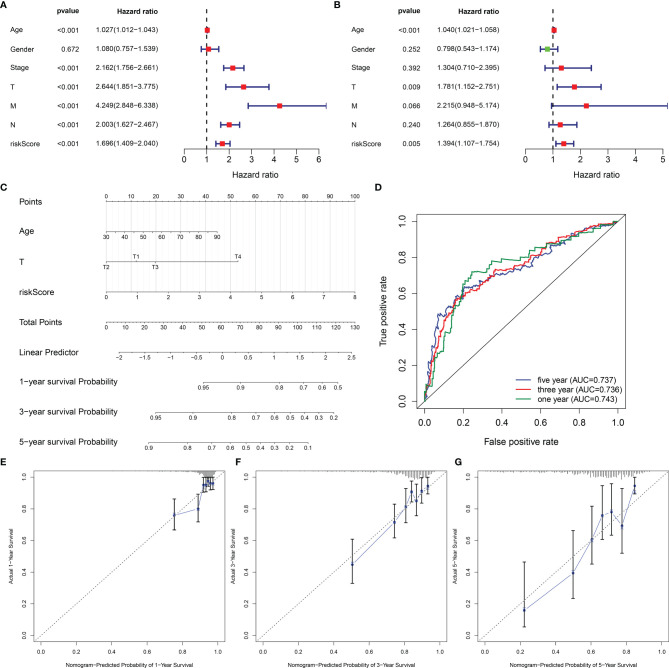
Univariate-Cox regression analysis uncovered the remarkable association of age, stage, T, N, M, and EVA1B-derived risk score with CRC prognosis (f9). Among them, age, T, and EVA1B-derived risk score served as independent prognostic indicators of CRC (f9). By integrating above three independent prognostic indicators, this study exploited a nomogram to estimate CRC patients’ survival outcomes (f9). This EVA1B-derived risk score occupied the most contribution to the prediction of 1-, 3-, and 5-year OS duration. ROC curves confirmed the desirable efficacy in predicting patients’ survival outcomes (f9). Additionally, we evaluated the prediction performance of the nomogram with calibration curves. Our data demonstrated that 1-, 3-, and 5-year predicted by this nomogram was close to the actual survival duration (f9). Above data indicated the superior predictive capacity of this nomogram.
Molecular Mechanisms Involving the EVA1B-Derived Genomic Signature
We further investigated the molecular mechanisms involving the EVA1B-derived genomic signature. As depicted in f10, unfolded protein response, E2F targets, G2M checkpoint, mTORC1 signaling, MYC targets, DNA repair, and oxidative phosphorylation presented higher activities in low-risk subpopulation. Meanwhile, tumorigenic pathways were remarkably activated in high-risk subpopulation, such as P53 pathway, TGF-beta signaling, apoptosis, Hedgehog signaling, EMT, Notch signaling, and angiogenesis, indicative of high-risk subpopulation’s unfavorable survival outcomes. Moreover, immune activation pathways such as allograft rejection, inflammatory response, and complement were significantly activated in high-risk subpopulation.
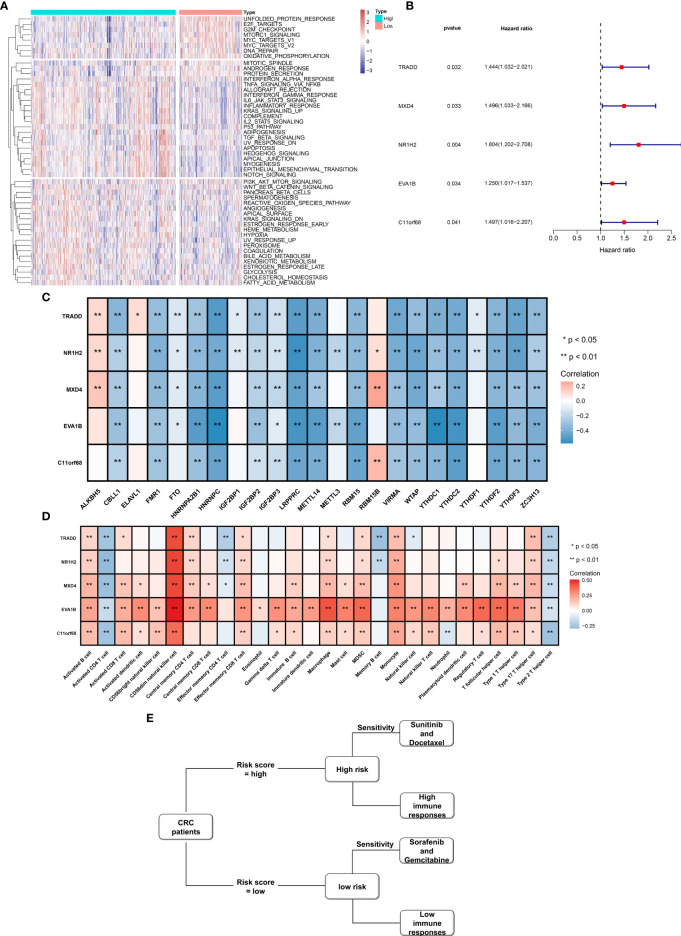
Prognostic Significance of EVA1B-Derived Genes and Their Interaction With m6A Methylation and Immune Cell Infiltration
f10 demonstrates that TRADD, MXD4, NR1H2, EVA1B, and C11orf68 acted as risk factors of CRC outcomes. Additionally, we investigated that above EVA1B-derived genes presented positive correlations to m6A regulator ALKBH5 and RBM15B but negative correlations to other m6A regulators in CRC (f10). In f10, these EVA1B-derived genes were negatively correlated to activated CD4+ T cell, effector memory CD4+ T cell, memory B cell, and type 2 T helper cell but positively correlated to other tumor-infiltrating immune cells in CRC. f10 shows the association among drug sensitivity, predictive immune responses, and risk status of CRC individuals.
EVA1B Overexpression Is Correlated With CRC Tumorigenesis and Poor Outcomes in CRC Patients
EVA1B IHC was performed on 19 pairs of primary CRC, corresponding noncancerous and matching liver metastasis specimens from 19 liver metastasis CRC patients. The findings showed that EVA1B was overexpressed in primary CRC tissues compared with the corresponding noncancerous normal tissues (p < 0.01). In addition, expression level of EVA1B was similar in the liver metastases samples to the expression level in matching primary CRC tissues without statistical significance (f11).
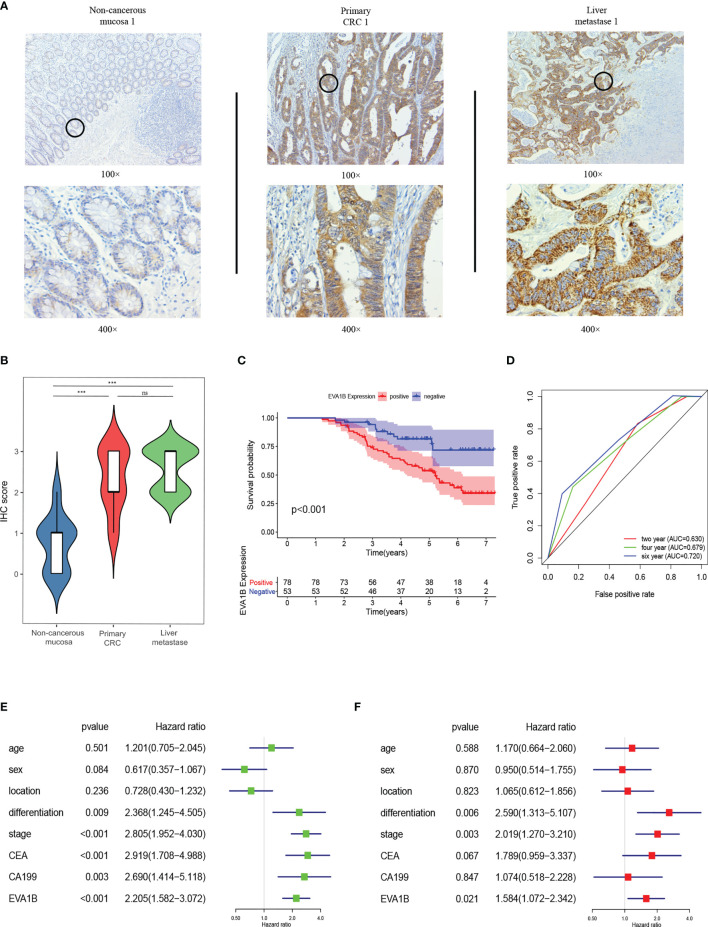
In addition, the relationship between EVA1B expression and clinicopathologic characteristics of 131 patients with CRC was explored (T2). Expression of EVA1B was highly correlated with pN stage (p < 0.001), pM stage (p = 0.005), AJCC stage (p = 0.001), CEA (p = 0.003), and CA199 (p = 0.002). However, the finding showed no significant correlation between EVA1B expression level and age, gender, tumor location, differentiation, and pT stage of CRC patients (T2).
Table 2: EVA1B staining expression associations with clinicopathologic and survival characteristics in CRC patients.
| Variable | No. | Negative | Positive | p-value |
|---|---|---|---|---|
| Clinicopathologic characteristics | ||||
| Age (years) | 0.922 | |||
| <60 | 60 | 24 | 36 | |
| ≥60 | 71 | 29 | 42 | |
| Gender | 0.091 | |||
| Male | 71 | 24 | 47 | |
| Female | 60 | 29 | 31 | |
| Location | 0.596 | |||
| Colon | 63 | 24 | 39 | |
| Rectum | 68 | 29 | 39 | |
| Differentiation | 0.884 | |||
| Poor | 18 | 7 | 11 | |
| Well | 113 | 46 | 67 | |
| pT stage | 0.141 | |||
| T1–T3 | 92 | 41 | 51 | |
| T4 | 39 | 12 | 27 | |
| pN stage | <0.001 | |||
| N0 | 60 | 38 | 22 | |
| N1–N2 | 71 | 15 | 56 | |
| pM stage | 0.005 | |||
| M0 | 109 | 50 | 59 | |
| M1 | 22 | 3 | 19 | |
| AJCC stage | 0.001 | |||
| I | 17 | 12 | 5 | |
| II | 36 | 25 | 11 | |
| III | 56 | 13 | 43 | |
| IV | 22 | 3 | 19 | |
| CEA | 0.003 | |||
| ≤5.0 (negative) | 79 | 40 | 39 | |
| >5.0 (positive) | 52 | 13 | 39 | |
| CA199 | 0.002 | |||
| ≤37.0 (negative) | 114 | 52 | 62 | |
| >37.0 (positive) | 17 | 1 | 16 | |
| Survival characteristics | ||||
| 1-year survival | 131 | 53 (100%) | 78 (100%) | NS |
| 2-year survival | 125 | 52 (98.1%) | 73 (93.6%) | NS |
| 3-year survival | 108 | 50 (94.3%) | 58 (74.4%) | 0.048 |
| 4-year survival | 94 | 44 (83.0%) | 50 (64.1%) | 0.018 |
| 5-year survival | 87 | 44 (83.0%) | 43 (55.1%) | <0.001 |
| 6-year survival | 77 | 42 (79.3%) | 35 (44.9%) | <0.001 |
| 7-year survival | 75 | 42 (79.3%) | 33 (42.3%) | <0.001 |
NS, no significance.
Kaplan–Meier curve was generated to explore the prognosis value of EVA1B in these 131 CRC patients. The findings showed that CRC patients with high expression levels of EVA1B presented with poor OS compared with those with low expression levels of EVA1B (p < 0.001, f11). AUC values for EVA1B expression (0.630, 0.679, and 0.720 for the 2-, 4-, and 6-year OS, respectively) showed that EVA1B accurately discriminated between the high- and low-expression groups (f11). Univariate and multivariate Cox analyses were performed, indicating that EVA1B was an independent as a prognostic factor by adjusting for stage, gender, location, differentiation, stage, CEA, and CA199 (f11).
Discussion
This study presented an integrative analysis of molecular characteristics, oncogenic roles, and relevant immune and pharmacogenomic features of EVA1B across pan-cancer, especially CRC. Our data demonstrated the remarkable upregulation of EVA1B expression in most cancer types. Additionally, its upregulation was significantly correlated to DNA methyltransferases, DNA mismatch repair genes, m6A regulators, TMB, and MSI across pan-cancer. Tumorigenesis is a multistep process where normal cells acquire genetic and epigenetic alterations contributing to tumor initiation and progression (ref. 41, ref. 42). The pathological mechanisms of CRC initiation and progression include chromosomal instability, high MSI and methylation, contributing to oncogene, tumor suppressor gene, and mismatch repair-relevant gene mutations (ref. 43). Our data were indicative of the critical role of EVA1B in rectal and colon carcinogenesis.
High EVA1B expression was indicative of undesirable CRC patients’ clinical outcomes. Nevertheless, we did not note the distinct difference in survival status between high- and low-risk subpopulations due to this retrospective cohort. The role of EVA1B in survival status of CRC should be investigated in a prospective cohort in our future studies. Previous research proposed the upregulation of EVA1B in glioma (ref. 11). At present, the therapeutic regimen is surgical resection plus chemotherapy in more advanced or inoperable patients (ref. 44). Nevertheless, immunotherapy to supplement curative and palliative therapy has been undergoing clinical trials (ref. 45). Our further analysis demonstrated that EVA1B upregulation was correlated to enhanced immune cell infiltration, increased stromal and immune activation, and elevated activities of cancer immunity cycle. Additionally, higher frequencies of amplification and deletion were noted in high EVA1B expression subpopulation. Our data were indicative that EVA1B might be utilized as a promising immunotherapeutic predictor.
We further determined 602 EVA1B-derived genes in CRC that were mainly correlated to immune response, tumorigenic pathways (such as PI3K-Akt signaling pathway, proteoglycans in cancer, and ECM-receptor interaction), and immune activation pathways (such as IL-17 signaling pathway, Th17 cell differentiation, cytokine-cytokine receptor interaction, Th1 and Th2 cell differentiation, antigen processing and presentation, intestinal immune network for IgA production, complement and coagulation cascades), indicative of their remarkable roles in CRC progression. Utilizing random survival forest analysis, we conducted an EVA1B-derived genomic model, composed of IGFBP3, EGFL7, SULT1B1, MMP1, CYP2W1, and ISYNA1. Following verification in diverse external datasets, the genomic model reliably and independently predicted patients’ prognosis and relapse. Additionally, it possessed the potential in estimating drug responses, like sunitinib, sorafenib, gemcitabine, and docetaxel. There was remarkable correlation of characteristic EVA1B-derived genes with responses to small molecular compounds. TRADD, MXD4, NR1H2, EVA1B, and C11orf68 served as remarkable risk factors of CRC outcomes. Previously, modulation of TRADD restores cellular homeostasis as well as alleviates apoptosis (ref. 46). It triggers tumor inhibition through regulation of ULF-dependent p19Arf ubiquitylation (ref. 47). CDK4/6 inhibitor upregulates MXD4 expression that negatively modulates MYC in CD8+ T cells (ref. 48). NR1H2 modulates cholesterol homeostasis within human cells, controlling fitness and function of activated T cells (ref. 49). Nevertheless, more experimental evidence should be conducted to uncover the functions of above EVA1B-relevant genes in CRC.
A nomogram acts as a powerful tool to quantify an individual’s risk in a clinical setting through integrating diverse risk factors (ref. 50). Herein, this study utilized the nomogram in prediction of 1-, 3-, and 5-year OS probabilities via incorporating age, T stage, as well as EVA1B-derived genomic model for CRC individuals. Each indicator was assigned a score on the basis of its contribution to survival risk. Thereafter, ROC curves demonstrated that the nomogram presented the favorable efficiency in predicting an individual patient’ OS outcome. Additionally, calibrated curves confirmed the actual survival duration agreed with the nomogram-estimated survival duration. A few limitations of our research will be pointed out. The functional role of EVA1B in tumor immunity requires in-depth experimental verification. Moreover, prognostic value of EVA1B needs to be verified in larger CRC cohorts.
Conclusion
Overall, our study conducted an integrative analysis to uncover molecular characteristics, oncogenic roles, and relevant immune and pharmacogenomic features of EVA1B in CRC. Our findings indicated that EVA1B acted as a convincing prognostic marker as well as a predictor of therapeutic responses.
Data Availability Statement
Publicly available datasets were analyzed in this study. These data can be found here: https://portal.gdc.cancer.gov/ and GEO, https://www.ncbi.nlm.nih.gov/geo/, under accession numbers GSE14333, GSE39582, and GSE87211.
Ethics Statement
The studies involving human participants were reviewed and approved by the human ethics committees of the Liaoning Cancer Hospital. The patients/participants provided their written informed consent to participate in this study.
Author Contributions
BM and YML conceived and designed this study. YML and KW performed the bioinformatics analyses and visualization. YL collected data and performed the statistical analysis. BM, YML and QM wrote the original draft. BM and YML revised the manuscripts. All authors revised and approved the final manuscript.
Funding
This study was supported by the National Natural Science Foundation of China (81902383); the Revitalizing Liaoning Talents Program (XLYC1907004); the Young and Middle-aged Scientific and Technological Innovation Talent Support Plan of Shenyang City (RC200223); Cultivation Program of National Science Foundation of Liaoning Cancer Hospital (2021-ZLLH-03); Cultivation Program of National Science Foundation of Liaoning Cancer Hospital (2020-ZLLH-44); Natural Guiding Plan Foundation of Liaoning Province (2019-ZD-0584); and Beijing Xisike Clinical Oncology Research Foundation (Y-QL202101-0039).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s Note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
- LH Biller, D Schrag. Diagnosis and Treatment of Metastatic Colorectal Cancer: A Review.. Jama (, 2021. [DOI]
- M Nguyen, S Tipping Smith, M Lam, E Liow, A Davies, H Prenen. An Update on the Use of Immunotherapy in Patients With Colorectal Cancer.. Expert Rev Gastroenterol Hepatol (, 2021. [DOI | PubMed]
- H Xu, L Liu, W Li, D Zou, J Yu, L Wang. Transcription Factors in Colorectal Cancer: Molecular Mechanism and Therapeutic Implications.. Oncogene (, 2021. [DOI]
- J Zhao, S Zhong, X Niu, J Jiang, R Zhang, Q Li. The MHC Class I-LILRB1 Signalling Axis as a Promising Target in Cancer Therapy.. Scand J Immunol (, 2019. [DOI | PubMed]
- X Kong, M Fu, X Niu, H Jiang. Comprehensive Analysis of the Expression, Relationship to Immune Infiltration and Prognosis of TIM-1 in Cancer.. Front Oncol (, 2020. [DOI | PubMed]
- E Picard, CP Verschoor, GW Ma, G Pawelec. Relationships Between Immune Landscapes, Genetic Subtypes and Responses to Immunotherapy in Colorectal Cancer.. Front Immunol (, 2020. [DOI | PubMed]
- S Zhao, H Wang. EVA1A Plays an Important Role by Regulating Autophagy in Physiological and Pathological Processes.. Int J Mol Sci (, 2021. [DOI | PubMed]
- BY Lin, JL Wen, C Zheng, LZ Lin, CZ Chen, JM Qu. Eva-1 Homolog A Promotes Papillary Thyroid Cancer Progression and Epithelial-Mesenchymal Transition via the Hippo Signalling Pathway.. J Cell Mol Med (, 2020. [DOI]
- Y Zhen, R Zhao, M Wang, X Jiang, F Gao, L Fu. Flubendazole Elicits Anti-Cancer Effects via Targeting EVA1A-Modulated Autophagy and Apoptosis in Triple-Negative Breast Cancer.. Theranostics (, 2020. [DOI]
- WW Ren, DD Li, X Chen, XL Li, YP He, LH Guo. MicroRNA-125b Reverses Oxaliplatin Resistance in Hepatocellular Carcinoma by Negatively Regulating EVA1A Mediated Autophagy.. Cell Death Dis (, 2018. [DOI | PubMed]
- S Qu, J Liu, H Wang. EVA1B to Evaluate the Tumor Immune Microenvironment and Clinical Prognosis in Glioma.. Front Immunol (, 2021. [DOI | PubMed]
- T Li, J Fu, Z Zeng, D Cohen, J Li, Q Chen. TIMER2.0 for Analysis of Tumor-Infiltrating Immune Cells.. Nucleic Acids Res (, 2020. [DOI | PubMed]
- S Hänzelmann, R Castelo, J Guinney. GSVA: Gene Set Variation Analysis for Microarray and RNA-Seq Data.. BMC Bioinf (, 2013. [DOI]
- A Sokolov, DE Carlin, EO Paull, R Baertsch, JM Stuart. Pathway-Based Genomics Prediction Using Generalized Elastic Net.. PloS Comput Biol (, 2016. [DOI | PubMed]
- N Auslander, G Zhang, JS Lee, DT Frederick, B Miao, T Moll. Robust Prediction of Response to Immune Checkpoint Blockade Therapy in Metastatic Melanoma.. Nat Med (, 2018. [DOI]
- K Yoshihara, M Shahmoradgoli, E Martínez, R Vegesna, H Kim, W Torres-Garcia. Inferring Tumour Purity and Stromal and Immune Cell Admixture From Expression Data.. Nat Commun (, 2013. [DOI | PubMed]
- DS Chen, I Mellman. Oncology Meets Immunology: The Cancer-Immunity Cycle.. Immunity (, 2013. [DOI | PubMed]
- L Xu, C Deng, B Pang, X Zhang, W Liu, G Liao. TIP: A Web Server for Resolving Tumor Immunophenotype Profiling.. Cancer Res (, 2018. [DOI]
- RN Jorissen, P Gibbs, M Christie, S Prakash, L Lipton, J Desai. Metastasis-Associated Gene Expression Changes Predict Poor Outcomes in Patients With Dukes Stage B and C Colorectal Cancer.. Clin Cancer Res (, 2009. [DOI]
- L Marisa, A de Reyniès, A Duval, J Selves, MP Gaub, L Vescovo. Gene Expression Classification of Colon Cancer Into Molecular Subtypes: Characterization, Validation, and Prognostic Value.. PloS Med (, 2013. [DOI | PubMed]
- Y Hu, J Gaedcke, G Emons, T Beissbarth, M Grade, P Jo. Colorectal Cancer Susceptibility Loci as Predictive Markers of Rectal Cancer Prognosis After Surgery.. Genes Chromosomes Cancer (, 2018. [DOI]
- JT Leek, WE Johnson, HS Parker, AE Jaffe, JD Storey. The Sva Package for Removing Batch Effects and Other Unwanted Variation in High-Throughput Experiments.. Bioinformatics (, 2012. [DOI]
- A Mayakonda, DC Lin, Y Assenov, C Plass, HP Koeffler. Maftools: Efficient and Comprehensive Analysis of Somatic Variants in Cancer.. Genome Res (, 2018. [DOI]
- CH Mermel, SE Schumacher, B Hill, ML Meyerson, R Beroukhim, G Getz. GISTIC2.0 Facilitates Sensitive and Confident Localization of the Targets of Focal Somatic Copy-Number Alteration in Human Cancers.. Genome Biol (, 2011. [DOI | PubMed]
- L Gautier, L Cope, BM Bolstad, RA Irizarry. Affy–Analysis of Affymetrix GeneChip Data at the Probe Level.. Bioinformatics (, 2004. [DOI]
- CL Wilson, CJ Miller. Simpleaffy: A BioConductor Package for Affymetrix Quality Control and Data Analysis.. Bioinformatics (, 2005. [DOI]
- JE Rosenberg, J Hoffman-Censits, T Powles, MS van der Heijden, AV Balar, A Necchi. Atezolizumab in Patients With Locally Advanced and Metastatic Urothelial Carcinoma Who Have Progressed Following Treatment With Platinum-Based Chemotherapy: A Single-Arm, Multicentre, Phase 2 Trial.. Lancet (, 2016. [DOI]
- Y Şenbabaoğlu, RS Gejman, AG Winer, M Liu, EM Van Allen, G de Velasco. Tumor Immune Microenvironment Characterization in Clear Cell Renal Cell Carcinoma Identifies Prognostic and Immunotherapeutically Relevant Messenger RNA Signatures.. Genome Biol (, 2016. [DOI | PubMed]
- S Mariathasan, SJ Turley, D Nickles, A Castiglioni, K Yuen, Y Wang. Tgfβ Attenuates Tumour Response to PD-L1 Blockade by Contributing to Exclusion of T Cells.. Nature (, 2018. [DOI]
- G Yu, LG Wang, Y Han, QY He. Clusterprofiler: An R Package for Comparing Biological Themes Among Gene Clusters.. Omics (, 2012. [DOI]
- A Liberzon, C Birger, H Thorvaldsdóttir, M Ghandi, JP Mesirov, P Tamayo. The Molecular Signatures Database (MSigDB) Hallmark Gene Set Collection.. Cell Syst (, 2015. [DOI]
- ME Ritchie, B Phipson, D Wu, Y Hu, CW Law, W Shi. Limma Powers Differential Expression Analyses for RNA-Sequencing and Microarray Studies.. Nucleic Acids Res (, 2015. [DOI | PubMed]
- H Wang, L Zhou. Random Survival Forest With Space Extensions for Censored Data.. Artif Intell Med (, 2017. [DOI | PubMed]
- PJ Heagerty, T Lumley, MS Pepe. Time-Dependent ROC Curves for Censored Survival Data and a Diagnostic Marker.. Biometrics (, 2000. [DOI]
- W Yang, J Soares, P Greninger, EJ Edelman, H Lightfoot, S Forbes. Genomics of Drug Sensitivity in Cancer (GDSC): A Resource for Therapeutic Biomarker Discovery in Cancer Cells.. Nucleic Acids Res (, 2013. [DOI]
- P Geeleher, N Cox, RS Huang. Prrophetic: An R Package for Prediction of Clinical Chemotherapeutic Response From Tumor Gene Expression Levels.. PloS One (, 2014. [DOI | PubMed]
- S Ma, C Chen, X Ji, J Liu, Q Zhou, G Wang. The Interplay Between M6a RNA Methylation and Noncoding RNA in Cancer.. J Hematol Oncol (, 2019. [DOI | PubMed]
- ZR Chalmers, CF Connelly, D Fabrizio, L Gay, SM Ali, R Ennis. Analysis of 100,000 Human Cancer Genomes Reveals the Landscape of Tumor Mutational Burden.. Genome Med (, 2017. [DOI | PubMed]
- M Baretti, DT Le. DNA Mismatch Repair in Cancer.. Pharmacol Ther (, 2018. [DOI | PubMed]
- WA Messersmith. NCCN Guidelines Updates: Management of Metastatic Colorectal Cancer.. J Natl Compr Canc Netw (, 2019. [DOI | PubMed]
- L Chen, G Wang, X Qiao, X Wang, J Liu, X Niu. Downregulated miR-524-5p Participates in the Tumor Microenvironment of Ameloblastoma by Targeting the Interleukin-33 (IL-33)/Suppression of Tumorigenicity 2 (ST2) Axis.. Med Sci Monit (, 2020. [DOI | PubMed]
- X Niu, J Xu, J Liu, L Chen, X Qiao, M Zhong. Landscape of N(6)-Methyladenosine Modification Patterns in Human Ameloblastoma.. Front Oncol (, 2020. [DOI | PubMed]
- J Li, L Liang, Y Yang, X Li, Y Ma. N(6)-Methyladenosine as a Biological and Clinical Determinant in Colorectal Cancer: Progression and Future Direction.. Theranostics (, 2021. [DOI]
- PG Alexander, DC McMillan, JH Park. A Meta-Analysis of CD274 (PD-L1) Assessment and Prognosis in Colorectal Cancer and its Role in Predicting Response to Anti-PD-1 Therapy.. Crit Rev Oncol Hematol (, 2021. [DOI | PubMed]
- XT Qiu, YC Song, J Liu, ZM Wang, X Niu, J He. Identification of an Immune-Related Gene-Based Signature to Predict Prognosis of Patients With Gastric Cancer.. World J Gastrointest Oncol (, 2020. [DOI]
- D Xu, H Zhao, M Jin, H Zhu, B Shan, J Geng. Modulating TRADD to Restore Cellular Homeostasis and Inhibit Apoptosis.. Nature (, 2020. [DOI]
- II Chio, M Sasaki, D Ghazarian, J Moreno, S Done, T Ueda. TRADD Contributes to Tumour Suppression by Regulating ULF-Dependent p19Arf Ubiquitylation.. Nat Cell Biol (, 2012. [DOI]
- M Heckler, LR Ali, E Clancy-Thompson, L Qiang, KS Ventre, P Lenehan. Inhibition of CDK4/6 Promotes CD8 T-Cell Memory Formation.. Cancer Discovery (, 2021. [DOI]
- AJ Michaels, C Campbell, R Bou-Puerto, AY Rudensky. Nuclear Receptor Lxrβ Controls Fitness and Functionality of Activated T Cells.. J Exp Med (, 2021. [DOI | PubMed]
- Y Miao, Q Li, J Wang, W Quan, C Li, Y Yang. Prognostic Implications of Metabolism-Associated Gene Signatures in Colorectal Cancer.. PeerJ (, 2020. [DOI | PubMed]
