Pan‐cancer analysis and oncogenic implications of MGAM and MGAM2: Toward precision oncology and drug repurposing in colorectal cancer
Abstract
Cancer remains a major global health challenge, with high prevalence and mortality rates emphasizing the urgent need for innovative treatment strategies. Although precision oncology offers tailored therapies based on genetic profiles, the clinical translation of genomic insights has been slow. Drug repurposing, using existing FDA‐approved drugs for new indications, presents a cost‐effective and time‐efficient alternative. This study investigates MGAM as a potential direct target of alpha‐glucosidase inhibitors in colorectal cancer (CRC), explores its biomarker potential, and evaluates gene expression patterns across diverse cancers. Using RNA‐Seq data from Recount3, Firebrowse, and gene set co‐expression analysis databases, we analyzed the differential expression of MGAM and its paralog MGAM2 across 33 cancer types. We examined mutation profiles, methylation status, survival impact, immune cell infiltration, and drug‐mRNA interactions. Validation was performed via real‐time PCR and whole‐exome sequencing (WES) in CRC patients. MGAM and MGAM2 showed differential expression across multiple cancers, with MGAM2 upregulated and MGAM downregulated in gastrointestinal tumors. Both genes were linked to key cancer‐related pathways, including metabolism, apoptosis, cell cycle regulation, and epithelial‐mesenchymal transition. MGAM exhibited frequent mutations and aberrant methylation in several cancers. Their expression correlated with immune cell infiltration and drug sensitivity, highlighting potential for therapy planning. Diagnostic modeling showed over 80% accuracy. In CRC patients, MGAM downregulation was confirmed in 64 samples, and WES revealed a novel MGAM mutation (rs2960746). These findings underscore MGAM and MGAM2 as promising biomarkers and therapeutic targets, supporting their relevance in advancing personalized oncology.
Article type: Research Article
Keywords: acarbose; diagnostic, colorectal cancer, drug repurposing, pan‐cancer, prognostic, RNA‐seq, voglibose
Affiliations: Metabolic Syndrome Research Center Mashhad University of Medical Sciences Mashhad Iran; Department of Medical Biochemistry Faculty of Medicine Mashhad University of Medical Sciences Mashhad Iran; Medical Genetics Research Center Mashhad University of Medical Sciences Mashhad Iran; Department of Medical Oncology Cancer Center Amsterdam Amsterdam U.M.C. VU University Medical Center (VUMC) Amsterdam the Netherlands; Department of Gastroenterology and Hepatology Faculty of Medicine Mashhad University of Medical Sciences Mashhad Iran; Surgical Oncology Research Center Mashhad University of Medical Sciences Mashhad Iran; Department of Biology Payame Noor University Tehran Iran; College of Medicine University of Warith Al‐Anbiyaa Karbala Iraq
License: © 2025 The Author(s). Journal of Cell Communication and Signaling published by John Wiley & Sons Ltd. CC BY 4.0 This is an open access article under the terms of the http://creativecommons.org/licenses/by/4.0/ License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Article links: DOI: 10.1002/ccs3.70042 | PubMed: 40881483 | PMC: PMC12381534
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (4.2 MB)
INTRODUCTION
Cancer is one of the leading causes of death worldwide and poses a significant challenge to global health, carrying a substantial social and economic burden.ref. ccs370042-bib-0001 According to estimates from the International Agency for Research on Cancer (IARC), nearly 20 million new cases of cancer were diagnosed in 2022, with the number of deaths attributed to cancer being estimated to be 9.7 million.ref. ccs370042-bib-0002 Despite significant advancements in surgical, radiation, and immunological‐based treatments that have enhanced the effectiveness of cancer therapies, the intricate and varied nature of the disease continues to pose significant challenges. Hence, there is an urgent need to address the complexities inherent in treating different types of cancer.ref. ccs370042-bib-0003 Precision oncology utilizes specific characteristics of cancer patients, such as their genetic profiles, to recommend new personalized treatment regimens. However, despite extensive pharmaceutical research efforts to develop new drugs targeting these genes, clinical implementation of these discoveries has lagged behind expectations.ref. ccs370042-bib-0004
Drug repurposing, also referred to as drug repositioning, presents an attractive and promising approach to treatment. Its advantages include cost‐effectiveness, expedited development timelines, and enhanced safety profiles, which stem from the comprehensive understanding of the drug’s pharmacokinetics and pharmacodynamics. This method entails discovering novel therapeutic applications for medications that have already received regulatory approval.ref. ccs370042-bib-0003, ref. ccs370042-bib-0005, ref. ccs370042-bib-0006 Drug repurposing employs several methodologies, including disease‐centric, target‐centric, and drug‐centric approaches, each with its unique focus and benefits.ref. ccs370042-bib-0007 The disease‐centric approach initiates with the selection of a specific disease. Researchers then sift through approved drugs used for other ailments to determine if any could offer beneficial effects against the chosen disease.ref. ccs370042-bib-0008 This strategy capitalizes on the extensive knowledge base surrounding the disease, seeking novel therapeutic applications for drugs already recognized for their efficacy in other contexts. The target‐centric approach, conversely, starts with a particular biological target, such as a protein or gene. Researchers pinpoint drugs that engage with this target and assess their capacity to address various diseases. This methodology is deeply grounded in the molecular mechanisms underlying disease pathology, aiming to uncover potential therapeutic agents based on a clear understanding of disease biology.ref. ccs370042-bib-0009 Lastly, the drug‐centric approach selects an individual drug and explores its potential utility across a range of diseases. The goal here is to unlock the full therapeutic potential of a single drug by identifying additional conditions it may effectively treat, thereby maximizing its clinical impact.ref. ccs370042-bib-0003, ref. ccs370042-bib-0010 Each of these methodologies comes with its own set of advantages and challenges. The synergy between them, however, can lead to the development of more effective and innovative treatments for diseases. By harnessing the strengths of each approach, researchers can enhance drug repurposing efforts, ultimately leading to improved patient outcomes.ref. ccs370042-bib-0007, ref. ccs370042-bib-0010
The development of high‐throughput omics analysis techniques and the utilization of tools for big data analysis, facilitated by the Cancer Genome Atlas (TCGA), have progressively outlined the landscape of changes in signaling pathways.ref. ccs370042-bib-0011 With the increasing application of multi‐omics approaches, the cancer research community can explore new opportunities for drug repurposing to target oncogenic stimuli in cancer patients.ref. ccs370042-bib-0012 The likelihood of a drug being repurposed for cancer treatment increases significantly when its initially approved targets align with cancer‐related targets. This may arise when medications or therapies were initially designed and received approval for indications other than cancer, yet they interact with biological pathways or molecules that play a role in cancer development or progression. The rationale behind this potential effectiveness lies in the observation that numerous diseases, including cancer, often share common underlying mechanisms or molecular targets.ref. ccs370042-bib-0013
Recent research has revealed that α‐glucosidase inhibitors such as acarbose, voglibose, and miglitol, commonly prescribed for diabetes management, exhibit antineoplastic properties.ref. ccs370042-bib-0014, ref. ccs370042-bib-0015, ref. ccs370042-bib-0016 It was discovered that these inhibitors exert a beneficial effect on the risk of colorectal cancer (CRC) by influencing the pattern of bile acids in the stool and decreasing the levels of neutral sterols.ref. ccs370042-bib-0017 Moreover, carbohydrate malabsorption has been suggested to potentially offer protection against colon cancer. A specific example of this protective effect comes from research involving the drug acarbose. The study demonstrated that acarbose could significantly lower the risk of developing CRC among diabetic patients. Importantly, the reduction in cancer risk appeared to be directly related to the dosage of acarbose administered, indicating a dose‐dependent relationship between the drug and its protective effects against CRC.ref. ccs370042-bib-0017 On the other hand, cancer cells are recognized to have a higher demand for glucose, necessitating increased energy compared to normal cells.ref. ccs370042-bib-0018 Alpha‐glucosidase inhibitors can effectively curtail the caloric intake of tumor cells by lowering blood glucose levels, ultimately leading to the destruction of tumor cells.ref. ccs370042-bib-0019
The MGAM (maltase‐glucoamylase) gene, which encodes the enzyme maltase‐glucoamylase, is one of the alpha‐glycosidases found in the intestinal brush border membrane. This enzyme plays a crucial role in the final stage of carbohydrate digestion. The MGAM gene is located on chromosome 7q34. Because of its importance in carbohydrate metabolism, MGAM has been a major therapeutic target for treating type 2 diabetes and insulin resistance.ref. ccs370042-bib-0020, ref. ccs370042-bib-0021, ref. ccs370042-bib-0022 MGAM binds to the epithelial cells of the brush border in the small intestine.ref. ccs370042-bib-0023 On the other hand, genetic aberrations in chromosome 7 are commonly observed in a wide range of human diseases, including cancer.ref. ccs370042-bib-0024
This research endeavors to explore the potential of MGAM as a therapeutic agent in cancer treatment through a multifaceted approach. Building upon the established effectiveness of alpha‐glucosidase inhibitors in CRC, as previously outlined in a preliminary literature review, we will first examine the expression of MGAM as a direct target of these drugs in CRC. Subsequently, we will expand our analysis to a broad spectrum of human cancers to identify patterns of MGAM expression, mutations, copy number variations (CNVs), methylation status, immune cell infiltration, and associated drugs across various malignancies. Our goal is to determine whether MGAM could serve as a pan‐cancer therapeutic target based on observed expression/mutation patterns across different cancer types. In addition to our current research, we seek to elucidate the molecular mechanisms that underlie MGAM‘s function and its protein–protein interactions. We will also assess its potential as both a prognostic and diagnostic marker in cancers. This comprehensive approach will provide valuable insights into the therapeutic potential of alpha‐glucosidase inhibitors for cancer treatment.
The findings of this research will contribute significantly to our understanding of how genetic alterations affect drug efficacy across various cancer types. These insights will inform future clinical trials and guide the development of personalized treatment strategies for cancer patients.
MATERIALS AND METHODS
Data collection and identification of differentially expressed genes
To investigate MGAM expression in CRC, we utilized the TCGA‐COAD dataset retrieved from The Cancer Genome Atlas (TCGA) (http://cancergenome.nih.gov/). We focused on the white population to minimize heterogeneity and gain deeper insights by examining gene expression across different stages. After excluding cases with missing clinical data, we conducted gene expression analysis on a total of 214 patient samples and 17 healthy controls from the white population, as detailed in our previous work.ref. ccs370042-bib-0025 We divided patients into three distinct subgroups based on tumor stage: one group combining stages I and II, another with stage III, and the last one with stage IV patients. Subsequently, we utilized the DESeq2 package to perform differential expression analysis within each subgroup separately. The criteria for significant differential expression were set at an adjusted p‐value <0.05 and a log fold change (|logFC|) ≥ 1.5.
In the next step to investigate the expression of MGAM in a pan‐cancer context, we used the recount3 project (RNA.recount.bio), which provided batch‐corrected data comprising 20,753 samples from TCGA and the Genotype‐Tissue Expression (GTEx) portal. This dataset included 11,094 normal and 9569 tumor samples. Using the edgeR software, we identified differentially expressed genes (DEGs) between the normal and tumor tissue samples. The significance threshold was set at |log2 (fold change)| ≥ 1 and adjusted p‐value ≤ 0.05.
The differential expression of candidate genes was further evaluated and confirmed across inter and intra‐cancer types using the Firebrowseref. ccs370042-bib-0026 and gene set cancer analysis (GSCA)ref. ccs370042-bib-0027 online databases. These platforms employ the RSEM method, a software package designed to estimate gene and isoform expression levels from RNA‐Seq data.
Clinical samples
To maintain consistency and avoid redundancy, detailed demographic and clinical characteristics of the patient cohort have been previously published.ref. ccs370042-bib-0025 Briefly, 64 patients diagnosed with non‐hereditary CRC were recruited, with RNA samples collected for qRT‐PCR validation of MGAM expression. Patient consent and ethical approval were obtained as described in the prior publication.ref. ccs370042-bib-0025
Pathway enrichment analysis and PPI
The STRING database (https://string‐db.org)ref. ccs370042-bib-0028 was utilized to identify all known and predicted physical interactions as well as protein–protein functional relationships of MGAM, with a minimum interaction score of 0.7 considered. In the following step, we utilized the GSCA database to examine the relationship between the expression levels of this particular gene and various regulatory elements within signaling pathways across all 33 cancer types.
Mutation profile assessment
The GSCA platform is a web‐based tool that integrates multi‐omics data from the TCGA database. Using this platform, we analyzed the profile of MGAM mutations, as well as the relationship between MGAM mRNA expression, CNVs, and single‐nucleotide variants (SNVs) across different tumor types. Additionally, we investigated the association between MGAM alterations, gene expression, and patient survival in all tumor samples.
Immune cell infiltration analysis
We then conducted an analysis using the GSCA database to examine the correlation between MGAM expression and the infiltration of 24 distinct types of immune cells. The immune cell abundance identity tool within this database provides estimates for the abundance of these immune cells, encompassing 18 subtypes of T cells and six other immune cell types (B cells, NK cells, monocytes, macrophages, neutrophils, and dendritic cells) across various cancer types.
Methylation analysis
The GSCA’s module on differential methylation offers insights into the methylation status in cancer patients versus normal samples. The GSCA database analyzed the correlation between mRNA expression levels, MGAM methylation levels, and their impact on survival across various types of tumors.
Evaluating the prognostic significance of MGAM across different cancer types
Survival analysis was conducted on TCGA datasets utilizing the survival package in R. The study evaluated overall survival (OS) in various cancer types.
ROC curve analysis
The diagnostic capabilities of MGAM, both individually and in conjunction with MGAM2, the paralog of MGAM,ref. ccs370042-bib-0021 were assessed using receiver operating characteristic (ROC) curve analysis. This evaluation emphasized critical performance indicators like sensitivity, specificity, and the area under the ROC curve. The analysis was performed utilizing the CombiROC package within the R programming environment.
Drug‐mRNA interaction
We employed the GSCA online database to explore drug‐mRNA interactions. This tool amalgamates half‐maximal inhibitory concentration (IC50) data from 250 small‐molecule drugs across 860 cell lines, alongside gene expression profiles sourced from the Genomics of Drug Sensitivity in Cancer (GDSC) initiative. It further computes the relationship between mRNA gene expression and drug IC50 values.
Additionally, we examined drug‐mRNA interactions through the drug–gene interaction database (DGIdb). DGIdb is engineered to streamline the identification of potential therapeutic targets within disease‐associated genes, particularly those linked to cancer. It alleviates the challenge of manually sorting through vast amounts of literature, clinical trial data, and databases to discover drug‐gene interactions. By leveraging expert curation and text mining techniques, DGIdb gathers information from reputable sources such as DrugBank, PharmGKB, and ChEMBL. This enables the classification of genes as potential targets based on their involvement in various pathways, molecular functions (MF), and gene families.ref. ccs370042-bib-0029
Invitro validation studies
Real‐time PCR
The expression of MGAM was validated through quantitative PCR (qRT‐PCR) following the isolation of RNA from 64 CRC samples and their corresponding controls using the ParsTous total RNA extraction kit from Tehran, Iran as described previously.ref. ccs370042-bib-0025 Subsequently, the purity and concentration of the total RNA were assessed using a NanoDrop 2000 spectrometer. The extracted RNA was then reverse‐transcribed into cDNA according to the manufacturer’s instructions using the ParsTous cDNA synthesis kit. The expression level of the MGAM gene was quantified using real‐time polymerase chain reaction (RT‐qPCR) with SYBR green master mix from ParsTous, Iran, and the ABI‐PRISM StepOne device. Graphical analysis and visualization were conducted using GraphPad Prism 10.0 software. GAPDH was utilized as an internal reference for RT‐qPCR data analysis, and the 2−ΔΔCt method was employed to standardize the expression levels of the target genes.
The primer pairs utilized in this study were as follows:
MGAM‐Forward: GGCTGCAAGAGGTAATGAGAGAT, MGAM‐Reverse: CTGGGGCTGTTGATTTCAGTG, GAPDH‐Forward: ATCAGCAATGCCTCCTGCAC, and GAPDH‐Reverse: TGGTCATGAGTCCTTCCACG.
Whole exome sequencing (WES) analysis
Somatic mutations in the MGAM gene were investigated through an analysis of the TCGA database using the mutation annotation format (MAF). In the subsequent phase of the study, we also aimed to investigate and confirm MGAM‐related mutations in CRC patients. DNA extraction was carried out utilizing the ParsTous isolation kit from Tehran, Iran. This extraction was performed on blood samples obtained from 15 CRC patients following the manufacturer’s protocol as described completely in our previous work.ref. ccs370042-bib-0025 All identified variants underwent rigorous internal quality control and quality matrix assessment. Variants with a minor allelic frequency (MAF) equal to or exceeding 0.1% for heterozygous variants or 1% for homozygous variants were excluded based on data from the iranome and genome aggregation database (gnomAD). Finally, a suite of prediction tools, including sorting intolerant from tolerant (SIFT),ref. ccs370042-bib-0030 polymorphism phenotyping version 2 (PolyPhen2),ref. ccs370042-bib-0031 Varsome,ref. ccs370042-bib-0032 and COSMIC,ref. ccs370042-bib-0033 were employed to forecast the potential impact of non‐synonymous and synonymous (amino acid) alterations.
Statistical analysis
To assess the efficacy of the treatment, a range of statistical methods were utilized, including ROC curve analysis, Kaplan–Meier survival analysis, independent sample t‐tests, and χ2 tests. These analyses were conducted using R version 4.3.1, with statistical significance determined by a p‐value less than 0.05.
RESULTS
MGAM expression exhibits a stage‐specific marked decrease in CRC
Analysis of the MGAM gene expression across different stages of CRC revealed a notable decline in expression levels. Specifically, MGAM expression decreased by 2.06‐fold in stages I and II, 1.9‐fold in stage III, and approximately 1.3‐fold in stage IV compared to control samples. Figure 1A shows the trend plot of MGAM expression across different stages.
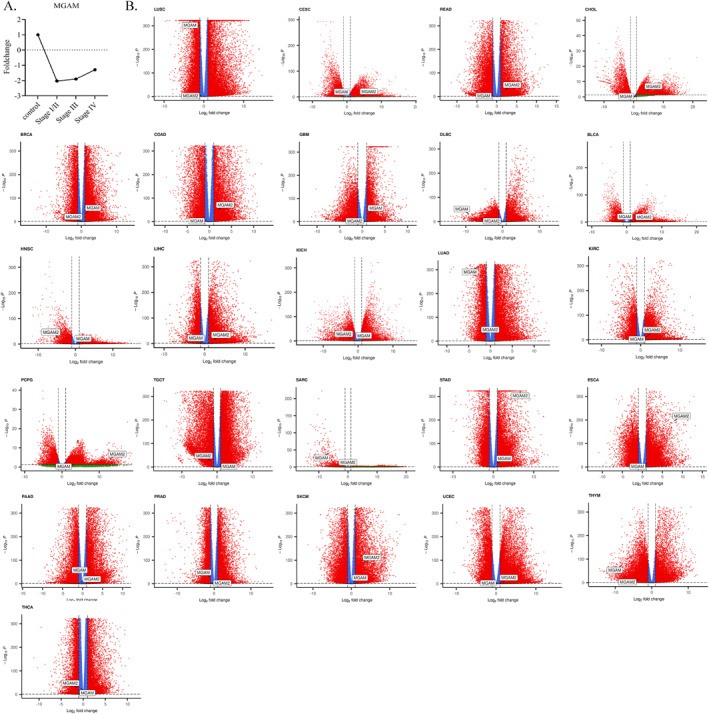
Pan‐cancer analysis revealed differential expression of MGAM and MGAM2 across different GI cancers
We explored the differential expression of MGAM across various cancers, employing R software and online databases. Pan‐cancer analysis of RNA‐seq data highlighted significant differences in MGAM expression in urothelial bladder carcinoma (BLCA), lymphoid neoplasm diffuse large B‐cell lymphoma (DLBC), kidney renal clear cell carcinoma (KIRC), lung adenocarcinoma (LUAD), lung squamous cell carcinoma (LUSC), pancreatic adenocarcinoma (PAAD), sarcoma (SARC), and thymoma (THYM).
Additionally, the investigation extended to MGAM‘s paralog, MGAM2, revealing its elevated expression in conditions such as carcinoma and endocervical adenocarcinoma (CESC), cholangiocarcinoma (CHOL), colon adenocarcinoma (COAD), esophageal squamous cell carcinoma (ESCA), KIRC, liver hepatocellular carcinoma (LIHC), pheochromocytoma and paraganglioma (PCPG), rectum adenocarcinoma (READ), skin cutaneous melanoma (SKCM), stomach adenocarcinoma (STAD), and downregulated expression in DLBC, head and neck squamous cell carcinoma (HNSC), kidney chromophobe (KICH), LUAD, testicular germ cell tumors (TGCT), and THYM (Figure 1B).
As previously mentioned, our RNA‐seq analysis is based on the data from TCGA tumors versus GTEX normal tissues. In this study, we also investigated the expression of these genes through the FireBrowse online database, which used TCGA tumors and matched normal tissue. Based on the results from these databases, a decrease in MGAM expression was also identified, notably in gastrointestinal cancers (GI) such as COAD, READ, CHOL, ESCA, LIHC, and STAD (Supporting Information S1: File 1). Notably, a strong relationship was observed between MGAM expression and subtypes of BRCA, KIRC, STAD, and HNSC, and MGAM2 expression and subtypes of KIRC, BRCA, and BLCA (Supporting Information S1: File 2).
MGAM contributes to metabolic and cancer‐related pathways
The STRING database has elucidated the protein–protein interaction (PPI) network of the MGAM gene, as shown in Figure 2A. Through gene ontology analysis, MGAM is identified as participating in numerous biological processes within metabolic pathways. It plays a significant role in the carbohydrate catabolic and metabolic process, the glucan catabolic process, and the cellular polysaccharide catabolic process, specifically in terms of biological function. Beyond these specific roles, MGAM demonstrated a range of MF in cooperation with MGAM2. These include glucan 1,4‐alpha‐glucosidase activity, alpha‐glucosidase activity, and alpha‐1,4‐glucosidase activity. Furthermore, when considering cellular components (CC), MGAM is linked to various structures such as ficolin‐1‐rich granules, secretory granule lumens, secretory granules, and extracellular exosomes (Figure 2B).
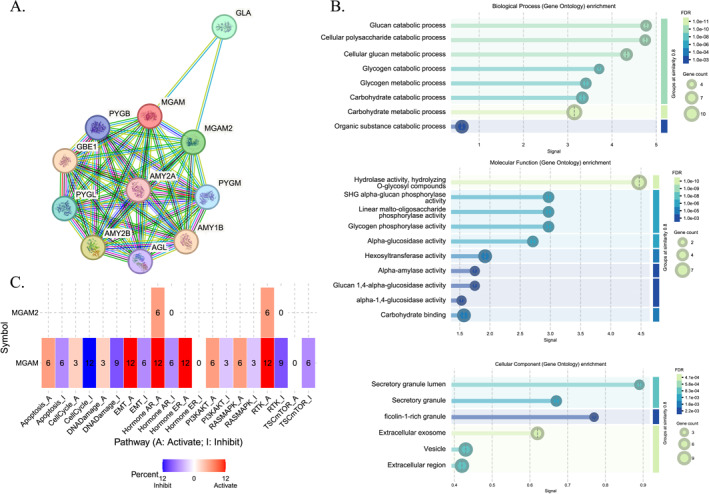
Our comprehensive analysis using the GCSA database revealed intricate relationships between MGAM and its paralog with cancer‐related pathways across various tumor types. Notably, MGAM exhibited divergent impacts on cellular processes in different cancer contexts (Figure 2C). In BRCA cancer, it was linked to the inhibition of key pathways, including apoptosis, cell cycle, DNA damage, RTK, PI3K/AKT, and TSC/mTOR while concurrently stimulating epithelial–mesenchymal transition (EMT), hormone AR, hormone endoplasmic reticulum (ER), and RAS/MAPK pathways. Interestingly, MGAM displayed contrasting effects on EMT in STAD and ESCA cancers compared to LIHC, LUSC, and HNSC. Furthermore, our data indicated that MGAM was associated with increased DNA damage in TGCT cancer and its inhibition in BRCA, KIRC, and LUSC cancers. The gene also demonstrated pathway‐specific activations and inhibitions across different cancer types, including the activation of apoptosis in LGG and LIHC, its inhibition in KIRC, and varied impacts on the cell cycle in ESCA, KIRC, and KIRP.
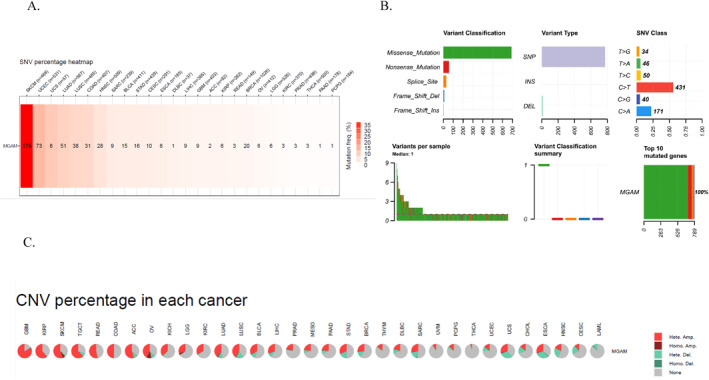
The mutation profile assessment proposed MGAM as a highly mutated gene in cancer
Further analysis of MGAM genetic changes revealed a high mutation rate in SKCM, UCEC, LUSC, LUAD, HNSC, COAD, BRCA, and STAD cancers (Figure 3A). Figure 3B illustrates the SNV classification in MGAM. The data revealed that missense mutations represent the most prevalent classification within MGAM. Notably, C > T transitions comprise the largest category of mutations observed. A significant correlation was observed between MGAM SNVs and OS and progression‐free survival (PFS) in PRAD cancer (Supporting Information S1: File 3). Figure 3C illustrates the frequency of CNVs across various cancers. Notably, a significant correlation was found between survival and CNV of the MGAM gene in UCEC cancer, considering OS, PFS, and disease‐specific survival (DSS) parameters (Supporting Information S1: File 4).
Immune cell infiltration analysis revealed the possible association of MGAM with immunity
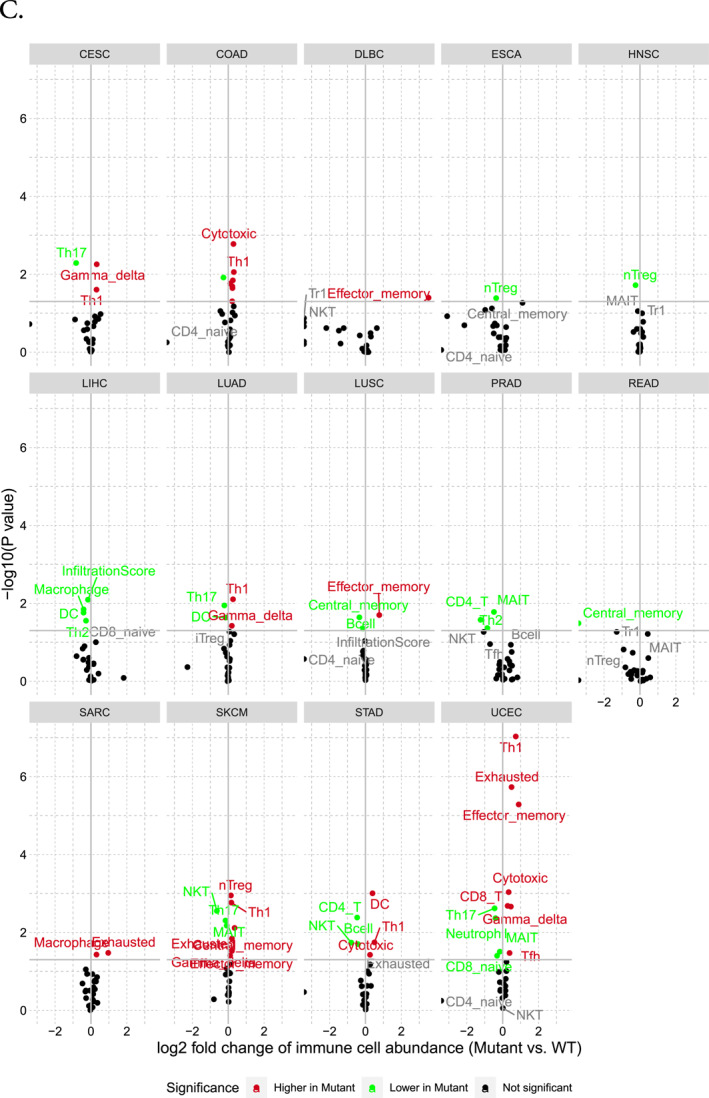
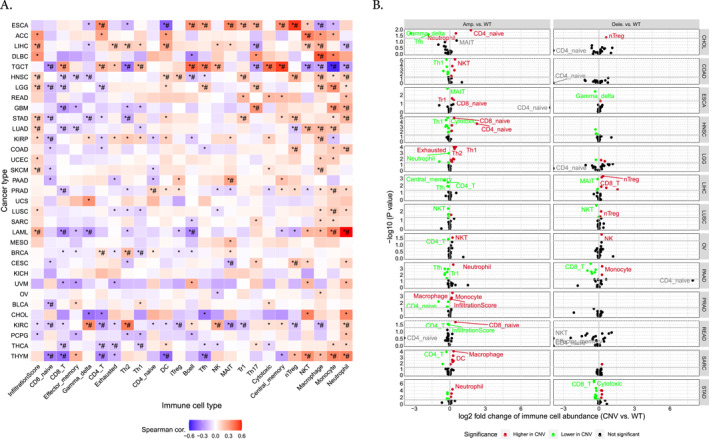
The analysis of immune infiltration concerning the MGAM and MGAM2 genes revealed an overall positive correlation with neutrophil and monocyte infiltration while exhibiting a negative correlation with CD4+, CD8+ T cells, and DC cells across various cancers such as KIRC, THCA, THYM, BRCA, LAML, PRAD, COAD, HNSC, LIHC, LUAD, STAD, and LGG. Particularly noteworthy is the strong correlation observed in LIHC, LGG, STAD, LUAD, SKCM, LAML, and HNSC between cell infiltration frequency and the expression of these genes (Figure 4A). The association of CNVs of MGAM with immune cells in different cancer types is illustrated in Figure 4B. As illustrated, amplification in MGAM seems to have more effect on immune cell abundance than deletion. Furthermore, SNVs of MGAM were identified to be correlated with higher Gamma delta, Th1, effector memory, nTreg, DC, and CD8‐T cells in cancers such as CESC, COAD, DLBC, LUAD, LUSC, SARC, SKCM, STAD, and UCEC, whereas they correlated with lower Th17, nTreg, macrophage, CD1, Th2, NKT, CD4‐Tcell, and Neutrophil in CESC, ESCA, HNSC, LIHC, LUAD, PRAD, SKCM, STAD, and UCEC (Figure 4C).
FIGURE 4(A) The correlation between MGAM expression and tumor infiltration. (B) Overall association of copy number variations in the MGAM across different cancers. (C) Overall association of SNVs in the MGAM across different cancers. All images were generated using the gene set cancer analysis database.
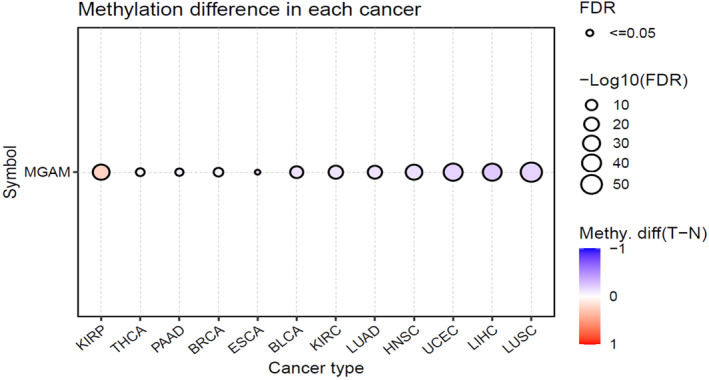
Abnormal methylation pattern in MGAM is associated with several cancers
Abnormal MGAM methylation patterns were observed in several types of cancers, including KIRP, THCA, PAAD, BRCA, ESCA, BLCA, KIRC, LUAD, HNSC, UCEC, LIHC, and LUSC (Figure 5). Furthermore, the relationship between MGAM expression and methylation patterns is evident in various cancers such as KIRC, TGCT, KIRP, CHOL, LGG, THYM, STAD, and PAAD, among others (Supporting Information S1: File 5).
MGAM is a potential prognostic biomarker, especially in GI cancers
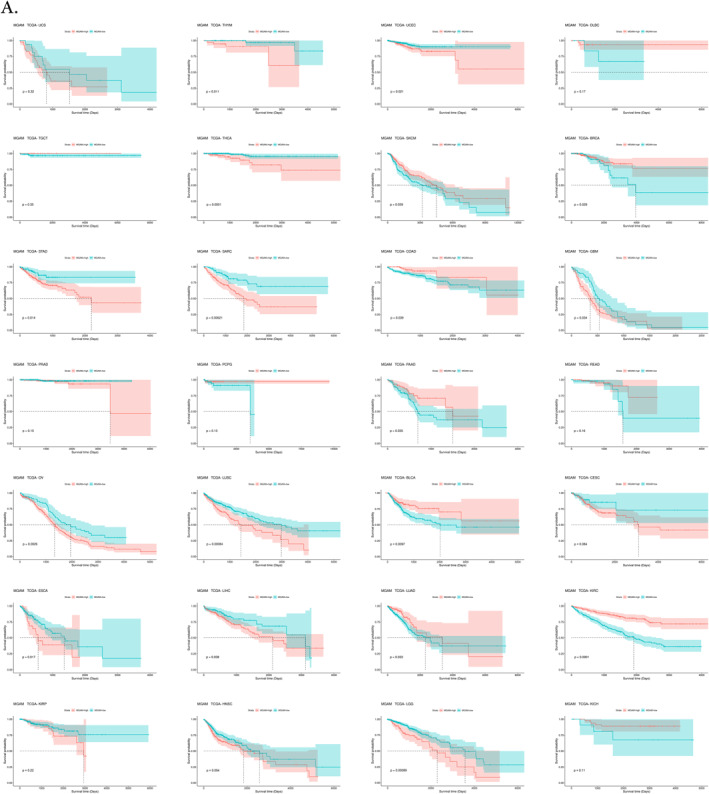
The findings from multiple analyses in this study highlight the potential prognostic significance of MGAM in various cancers, such as BLCA, BRCA, GBM, KIRC, LGG, LUAD, LUSC, OV, SARC, SKCM, THCA, THYM, UCEC, and, interestingly, in most of the GI cancers such as STAD, COAD, ESCA, LIHC, and PAAD (Figure 6). while MGAM2 was identified as a potential prognostic biomarker for cancers, including ACC, BLCA, BRCA, DLBC, ESCA, KIRC, KIRP, LGG, OV, SKCM, THCA, and UCEC (Supporting Information S1: File 6).
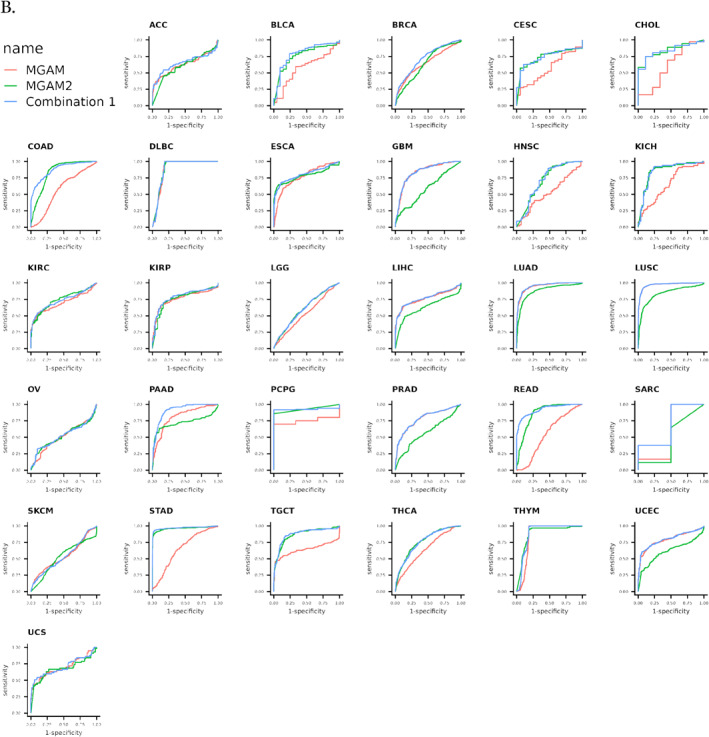
FIGURE 6(A) Pan‐cancer survival analysis of MGAM utilizing the cancer genome atlas datasets and Survival package in R. (B) Receiver operating characteristic curve analysis of MGAM across different cancers using CombiROC in R.
ROC curve analysis showed the diagnostic value of MGAM and MGAM2 in various cancers
ROC curve analysis was conducted to assess the diagnostic efficacy of MGAM and MGAM2 in various cancers. Whereas MGAM alone demonstrated diagnostic power exceeding 70% in DLBC, ESCA, GBM, LIHC, KIRP, LUAD, LUSC, PAAD, PCPG, PRAD, THYM, and UCEC cancers, combining MGAM with MGAM2 resulted in increased diagnostic effectiveness. This combination proved particularly potent, offering diagnostic power of over 80% in a broader range of cancers, including CHOL, DLBC, ESCA, GBM, COAD, LUAD, LUSC, PAAD, PCPG, READ, THYM, and STAD (Figure 6B).
Drug‐mRNA interaction analysis suggested MGAM as a potential target for predicting drug sensitivity and resistance
The expression of MGAM and its correlation with various drugs were investigated using data from the Cancer Therapeutics Response Portal (CTRP) and GDSC databases. The results revealed a negative correlation between MGAM expression and drugs such as PD318088, selumetinib, AICAR, trametinib, tigecycline, (5Z)−7−Oxozeaenol, MLN2480, Dabrafenib, and PLX4720.
Conversely, a positive association was observed with drugs such as avicin D, nintedanib, and BMS‐195614, indicating that increased MGAM expression is linked to increased resistance to these drugs (Figure 7A). Moreover, results from the DGIdb database showed that MGAM and MGAM2 were targets of approved antidiabetic drugs such as VOGLIBOSE, ACARBOSE, MIGLITOL, etc., as demonstrated in Table 1.
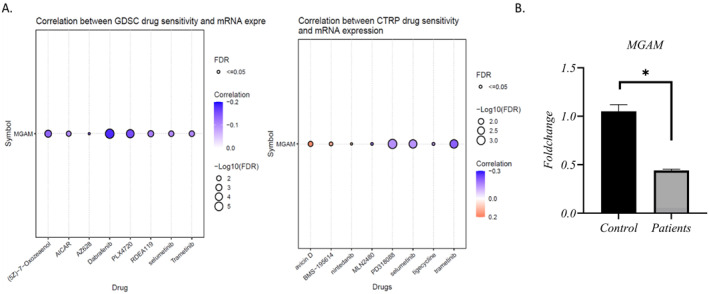
TABLE 1: Drug–target interaction based on DGIdb module (GSCA database).
| Gene | Drug | Regulatory approval | Indication | Interaction score |
|---|---|---|---|---|
| MGAM2 | VOGLIBOSE | Approved | Antidiabetic | 6.563458904 |
| MGAM2 | CELGOSIVIR | Not approved | 6.563458904 | |
| MGAM | MALTOSE | Approved | 2.386712329 | |
| MGAM | SALACINOL | Not approved | 2.386712329 | |
| MGAM | MIGLITOL | Approved | Hypoglycemic agents | 4.773424658 |
| MGAM | VOGLIBOSE | Approved | Antidiabetic | 4.773424658 |
| MGAM | CELGOSIVIR | Not approved | 2.386712329 | |
| MGAM | RESVERATROL 4′‐METHYL ETHER | Not approved | 9.546849316 | |
| MGAM | GLYCOVIR | Not approved | 4.773424658 | |
| MGAM | LYSOSOMAL ALPHA‐GLUCOSIDASE | Approved | 1.591141553 | |
| MGAM | BAICALEIN | Not approved | 0.367186512 | |
| MGAM | DUVOGLUSTAT | Not approved | For the treatment of pompe disease | 1.193356164 |
| MGAM | ACARBOSE | Approved | Antidiabetic | 3.182283105 |
Real‐time PCR validated the low expression of MGAM in CRC patients
Our primary objective was to investigate MGAM as a direct target of alpha‐glucosidase inhibitor drugs in CRC. Furthermore, factors such as the prognostic value of this gene and its diagnostic potential in combination with MGAM2 in CRC, along with the association of gene mutations with CRC, suggested that this gene could serve as a potential biomarker for this cancer. Consequently, at this stage, we validated the gene expression in CRC patients using qRT‐PCR. The results showed downregulation of MGAM in CRC patients versus controls (Figure 7B).
WES analysis revealed a novel mutation of MGAM
First, we conducted a comprehensive analysis of the SNV file from the COAD‐TCGA dataset to identify pathogenic variants. The results revealed several missense mutations with predicted deleterious effects, such as rs782114142, rs1267900787, and rs1554457449, among others, as presented in Table 2.
TABLE 2: Pathogenic variants of MGAM across the COAD‐TCGA dataset.
| Hugo_Symbol | Variant_Classification | dbSNP_RS | HGVSc | HGVSp | HGVSp_Short | Existing_variation | SIFT | PolyPhen | Impact | COSMIC |
|---|---|---|---|---|---|---|---|---|---|---|
| MGAM | Missense_Mutation | rs782114142 | c.1265A > G | p.Asp422Gly | p.D422G | rs782114142; COSV72075101 | deleterious (0) | probably_damaging (0.998) | MODERATE | COSM269579; COSM269580 |
| MGAM | Missense_Mutation | rs1267900787 | c.2351C > T | p.Ala784Val | p.A784V | rs1267900787; COSV101527578 | deleterious (0) | probably_damaging (0.952) | MODERATE | NA |
| MGAM | Missense_Mutation | NA | c.1751A > T | p.Asn584Ile | p.N584I | COSV72074775 | deleterious (0) | probably_damaging (0.965) | MODERATE | COSM1448697; COSM1448698; COSM1448699 |
| MGAM | Missense_Mutation | NA | c.4513G > T | p.Gly1505Trp | p.G1505W | COSV72074484 | deleterious (0) | probably_damaging (1) | MODERATE | COSM1448740; COSM1448741; COSM1448742 |
| MGAM | Missense_Mutation | rs372476876 | c.1847C > T | p.Ala616Val | p.A616V | rs372476876; COSV72073881 | tolerated (0.1) | benign (0.294) | MODERATE | COSM1448700; COSM304505; COSM304506 |
| MGAM | Missense_Mutation | rs1554457449 | c.344G > A | p.Arg115His | p.R115H | rs1554457449; COSV71885052 | deleterious (0.02) | probably_damaging (0.965) | MODERATE | COSM294562; COSM294563; COSM294564 |
| MGAM | Missense_Mutation | rs377213841 | c.3748C > T | p.Arg1250Cys | p.R1250C | rs377213841 | deleterious (0.01) | probably_damaging (1) | MODERATE | NA |
| MGAM | Missense_Mutation | rs377213841 | c.3748C > T | p.Arg1250Cys | p.R1250C | rs377213841 | deleterious (0.01) | probably_damaging (1) | MODERATE | NA |
| MGAM | Missense_Mutation | NA | c.3725G > T | p.Trp1242Leu | p.W1242L | COSV72074456 | deleterious (0.01) | probably_damaging (1) | MODERATE | COSM1448728; COSM1448729; COSM1448730 |
| MGAM | Missense_Mutation | rs778528944 | c.2501G > A | p.Arg834Gln | p.R834Q | rs778528944; COSV72073567 | deleterious (0) | probably_damaging (1) | MODERATE | COSM599595; COSM599596; COSM599597 |
| MGAM | Missense_Mutation | rs1431849178 | c.3287T > G | p.Ile1096Ser | p.I1096S | rs1431849178; COSV101527722 | deleterious (0) | probably_damaging (0.968) | MODERATE | NA |
| MGAM | Missense_Mutation | NA | c.2161C > T | p.Pro721Ser | p.P721S | COSV72074615 | deleterious (0) | possibly_damaging (0.903) | MODERATE | COSM1448701; COSM1448702; COSM1448703 |
| MGAM | Missense_Mutation | rs267601326 | c.4708C > T | p.Arg1570Cys | p.R1570C | rs267601326; COSV72074844 | deleterious (0) | probably_damaging (1) | MODERATE | COSM3634777; COSM3634778; COSM3634779 |
| MGAM | Missense_Mutation | NA | c.3726G > T | p.Trp1242Cys | p.W1242C | COSV72075636 | deleterious (0) | probably_damaging (1) | MODERATE | NA |
| MGAM | Missense_Mutation | NA | c.3368C > A | p.Ser1123Tyr | p.S1123Y | COSV72073742; COSV72074764 | deleterious (0) | probably_damaging (0.999) | MODERATE | COSM1448719; COSM1448720; COSM1448721 |
Next, we performed WES to determine whether any of the identified missense mutations were present in our patient cohort. Interestingly, none of those mutations were detected; however, all 15 individuals consistently harbored a specific variant in the MGAM gene, namely rs2960746. Although this variant is currently classified as a variant of uncertain significance (VUS) in the Varsome database, its potential pathogenicity is supported by deleterious predictions from both the SIFT and SIFT4G algorithms. Ongoing functional studies and routine follow‐up investigations are warranted to better clarify the clinical relevance of this variant.
DISCUSSION
In clinical oncology, where the search for effective treatments against the complex and varied nature of cancer continues, drug repurposing has emerged as a pivotal strategy. This approach leverages existing medications approved for other indications, offering a faster, less costly, and potentially safer alternative to traditional drug development processes. The core objective of drug repurposing is to identify novel therapeutic applications for known compounds, thereby accelerating the delivery of personalized treatments tailored to individual patient needs.ref. ccs370042-bib-0034
The integration of advanced technologies, such as machine learning and omics, plays a crucial role in this endeavor.ref. ccs370042-bib-0035, ref. ccs370042-bib-0036 These tools enable the prediction of disease–drug pairs through a combination of phenotypic studies, mechanistic investigations, chemical genetics, and omics assays. This multidisciplinary approach is particularly critical for addressing the challenges posed by cancer heterogeneity, recurrence, and metastasis, ensuring rapid and personalized interventions for patients. Omics technologies, encompassing genomics, proteomics, metabolomics, and others, offer unparalleled insights into the molecular mechanisms underlying diseases such as cancer.ref. ccs370042-bib-0037 By analyzing these “omic” layers, researchers can uncover potential targets for drug repurposing, identifying pathways and molecules that, when modulated, could lead to therapeutic benefits. This omics‐driven approach facilitates the identification of off‐target effects and polypharmacology, where a single compound interacts with multiple targets simultaneously. Such findings are instrumental in refining phenotyping strategies and tailoring treatments to the unique characteristics of each patient’s tumor.ref. ccs370042-bib-0038, ref. ccs370042-bib-0039, ref. ccs370042-bib-0040
Cancer is increasingly recognized as a metabolic disease, highlighting the importance of metabolic pathways in its development and progression.ref. ccs370042-bib-0041 Drugs originally developed for metabolic disorders, such as type 2 diabetes mellitus (T2DM), are being repurposed for cancer therapeutics because of shared risk factors and pathophysiological mechanisms. For instance, antidiabetic drugs such as metformin have demonstrated a reduced risk of overall cancer incidence and mortality across various types of cancer.ref. ccs370042-bib-0042, ref. ccs370042-bib-0043, ref. ccs370042-bib-0044 Similarly, other antidiabetic agents, such as glyburide and acarbose, showed promise in preclinical models by modulating inflammatory responses and enhancing immune surveillance against cancer.ref. ccs370042-bib-0042 Studies have shown that acarbose can inhibit tumor growth and enhance immune responses in colon cancer, suggesting a synergistic effect when combined with immunotherapeutic approaches.ref. ccs370042-bib-0045
The metabolic landscape of the tumor microenvironment often leads to abnormal glycolysis and the subsequent accumulation of lactic acid within tumor cells. This phenomenon can suppress the response of CD8+ T cells, thereby posing a significant challenge in cancer treatment. This underscores the importance of exploring metabolic pathways as targets for repurposing therapy in colorectal cancer, aiming to develop more effective and personalized treatment strategies.ref. ccs370042-bib-0046, ref. ccs370042-bib-0047, ref. ccs370042-bib-0048 α‐Glucosidase inhibitors, such as acarbose, voglibose, and miglitol, are utilized for their efficacy in managing type 2 diabetes. These drugs act by targeting the maltase–glucoamylase enzyme (MGAM) and play a crucial role in the final stages of starch digestion.ref. ccs370042-bib-0016 Therefore, to gain a clearer understanding of the role and potential significance of MGAM in the molecular mechanisms underlying cancer, with the ultimate goal of identifying a target for drug repurposing strategies in cancer treatment, we conducted a comprehensive pan‐cancer analysis of MGAM. This pioneering analysis encompassed an examination of gene expression levels, mutations, methylation patterns, immune cell infiltration, and the association of drugs with this gene. Following this, we delved into the implicated gene pathways and protein–protein interactions, assessing MGAM‘s and its paralog MGAM2’s potential as a prognostic and diagnostic marker in cancer.
In the present study, the comprehensive analysis of pooled data from multiple resources, including TCGA, GTEX datasets, Firebrowse, and GSCA databases, has unveiled substantial variations in the expression levels of the MGAM gene across a wide spectrum of cancer types. These types encompassed PAAD, ESCA, KIRC, GBM, BLCA, DLBC, LUAD, LUSC, PRAD, THYM, BRCA, SARC, and STAD. It is particularly noteworthy that, based on the results from online databases, MGAM demonstrated a diminished expression trend in gastrointestinal (GI) cancers, which include COAD, READ, CHOL, ESCA, LIHC, and STAD.
To enrich our comprehension of MGAM’s function, we broadened our research to encompass its lesser‐known variant, MGAM2, scrutinizing its differential expression across various forms of cancer. Our results suggested an elevation in MGAM2 expression in conditions such as CESC, CHOL, COAD, ESCA, KIRC, LIHC, PCPG, READ, SKCM, STAD, and downregulation in DLBC, HNSC, KICH, LUAD, TGCT, and THYM. Interestingly, the expression pattern of MGAM and MGAM2 seems to be similar in cancers such as DLBC, KIRC, LUAD, and THYM, whereas it seems to have a converse effect in GI cancers such as COAD, READ, CHOL, ESCA, LIHC, and STAD.
The STRING database mapped the PPI network of the MGAM gene, revealing its involvement in various biological processes and MF. Gene ontology analysis revealed that MGAM is primarily associated with carbohydrate catabolism and cellular polysaccharide breakdown within metabolic pathways. MGAM exhibited specific enzymatic activities, including glucan 1,4‐alpha‐glucosidase, alpha‐glucosidase, and alpha‐1,4‐glucosidase, in collaboration with MGAM2. Furthermore, MGAM interacted with several CC, such as ficolin‐1‐rich granules, secretory granule lumens, and extracellular exosomes, highlighting its multifaceted role within cells. The GSCA database also predicted a significant association between MGAM, MGAM2, and various cancer‐related pathways. In several cancers, including BRCA, STAD, ESCA, LIHC, LUSC, HNSC, TGCT, KIRC, and KIRP, MGAM influenced pathways such as apoptosis, cell cycle, DNA damage, RTK, PI3KAKT, TSCmTOR, EMT, hormone AR, hormone ER, and RASMAPK. Interestingly, MGAM’s role varied across these cancers, exhibiting both stimulatory and inhibitory effects on these pathways. This suggests that MGAM may promote cancer progression through diverse mechanisms, depending on the specific cancer type and its genetic and molecular context which needs further clarification.
Analysis of MGAM mutations showed a high mutation rate across various cancers, including SKCM, UCEC, LUSC, LUAD, HNSC, COAD, BRCA, and STAD. Significant correlations were identified between MGAM‘s SNVs, OS, and PFS in PRAD, and between MGAM‘s CNVs and survival metrics in UCEC. Additionally, MGAM expression correlated strongly with CNVs in OV, HNSC, THCA, SKCM, LUAD, BRCA, and LGG cancers.
Tumor development and occurrence are intricately linked to the infiltration of immune cells. Tumor immune cell infiltration denotes the migration of immune cells from the bloodstream into the tumor tissue to exert their effects. This phenomenon is closely associated with clinical outcomes and has the potential to serve as a target for drug intervention to enhance patient survival.ref. ccs370042-bib-0049
Our investigation into the role of MGAM and MGAM2 genes in various cancers has unveiled intriguing insights into their correlation with immune cell infiltration and their potential as diagnostic and prognostic markers. Specifically, we found a positive correlation with neutrophil and monocyte infiltration, yet a negative correlation with CD4+, CD8+ T cells, and DCs across a broad spectrum of cancers, including KIRC, THCA, THYM, BRCA, LAML, PRAD, COAD, HNSC, LIHC, LUAD, STAD, and LGG. Notably, the strongest correlations were observed in LIHC, LGG, STAD, LUAD, SKCM, LAML, and HNSC, suggesting a significant impact of these genes on immune cell dynamics within tumor microenvironments.
Moreover, abnormal methylation patterns of MGAM were detected in numerous cancer types, underscoring the gene’s involvement in cancer biology beyond its transcriptional regulation. The relationship between MGAM expression and methylation further complicates the gene’s role in cancer progression, highlighting the need for comprehensive epigenetic analyses.
The prognostic implications of MGAM are underscored by our findings, particularly in GI cancers. MGAM showed prognostic capability in BLCA, BRCA, GBM, KIRC, LGG, LUAD, LUSC, OV, SARC, SKCM, THCA, THYM, UCEC, and, interestingly, in most of the GI cancers such as STAD, COAD, ESCA, LIHC, and PAAD, while MGAM2 was identified as a potential prognostic biomarker for cancers, including ACC, BLCA, BRCA, DLBC, ESCA, KIRC, KIRP, LGG, OV, SKCM, THCA, and UCEC. It seemed that MGAM emerged as a promising prognostic biomarker specifically for GI cancers. This finding underscores the critical role of MGAM in predicting outcomes and guiding treatment decisions in GI malignancies.
The diagnostic accuracy of combining MGAM and MGAM2 gene analysis exceeds 80% across multiple cancer types, including DLBC, ESCA, GBM, COAD, LUAD, LUSC, PAAD, PCPG, READ, THYM, and STAD. This high accuracy underscores the potential of these genes as biomarkers for cancer diagnosis.
Investigations into MGAM‘s correlation with drug responses revealed complex associations, with increased MGAM expression correlating negatively with certain drugs’ efficacy and positively with others. These findings suggest a nuanced interplay between MGAM expression and drug sensitivity/resistance profiles, which could inform personalized treatment strategies. Furthermore, MGAM and MGAM2 were identified as targets of approved antidiabetic drugs, adding another layer to their multifaceted roles in cancer biology and therapeutics. This discovery opens avenues for exploring the therapeutic potential of targeting MGAM in conjunction with existing treatments.
Our findings suggested that MGAM could serve as a promising biomarker for CRC. To validate these, we conducted gene expression analysis in CRC patients using qRT‐PCR, which confirmed low expression in CRC patients compared to controls.
The clinical significance of NGS‐based methods is steadily growing. WES stands out as a burgeoning and dependable technology for pinpointing mutational patterns in cancer.ref. ccs370042-bib-0050 In this research, WES analysis revealed a distinct variant of the MGAM gene, specifically rs2960746, consistently present in all 15 CRC patients. This variant was deemed pathogenic through predictions from the SIFT and SIFT4G databases. Conducting functional investigations in this area and regularly updating predictive databases can enhance our understanding of the mutation’s phenotypic impact in the future. Furthermore, we assessed deleterious mutations in this gene based on SNV data from COAD‐TCGA.
Several studies were conducted on the importance of MGAM in cancer: The significance of the MGAM gene in various types of cancer has been extensively explored through network pharmacology and molecular docking technologies. Studies have highlighted MGAM as a critical target for traditional Chinese herbal formulas such as Mahuang Fuzi Xixin Decoction (MFXD) in treating lung cancer, particularly in LUAD, where it is a key mutated generef. ccs370042-bib-0051, ref. ccs370042-bib-0052 In non‐small cell lung cancer (NSCLC), nearly 18% of patients with specific EGFR mutations also presented MGAM–BRAF fusions, underscoring the gene’s importance in this context.ref. ccs370042-bib-0053 Recent advancements in examining driver mutations in cancer patients have provided valuable insights for personalized targeted immunotherapy. Mutations in MGAM have been linked to enhanced response rates, increased PD‐L1 expression, and higher TMB levels in NSCLC patients, suggesting a potential role in modulating tumor‐infiltrating immune cells.ref. ccs370042-bib-0054 The distinct N‐glycosylation pattern of MGAM serves as a biomarker for bladder cancer progression.ref. ccs370042-bib-0055 However, its reduced expression in early intestinal cancer suggests its potential as a serum biomarker for early detection.ref. ccs370042-bib-0056 In a study focusing on three urogenital cancers and benign prostatic hyperplasia (BPH), researchers discovered a strong association between MGAM glycoproteins and aggressive prostate cancer. These glycoproteins were found to be uniquely expressed in urine samples from individuals with prostate cancer.ref. ccs370042-bib-0057 In another study, He et al. proposed a prognostic model comprising seven coding genes, among which MGAM was strongly linked to biochemical recurrence after radical prostatectomy in prostate cancer.ref. ccs370042-bib-0058 MGAM exhibited higher expression levels in castration‐resistant prostate cancer metastatic tumors when compared to primary tumors and normal tissue,ref. ccs370042-bib-0059 which is consistence with our study.
SNV mutations in the MGAM gene are more prominently expressed in the glycolytic subtype of cutaneous melanoma (SKCM), which is associated with the poorest prognosis compared to other subtypes.ref. ccs370042-bib-0060 In gastric cancers, MGAM is among the most aberrantly expressed mRNAs, with decreased expression observed in intestinal‐type gastric cancer.ref. ccs370042-bib-0061 The genome‐wide analysis conducted using complementary DNA microarray has revealed that MGAM is among the genes exhibiting decreased expression in intestinal‐type gastric cancer, suggesting its involvement in carcinogenesis.ref. ccs370042-bib-0062 Genomic comparative hybridization analysis using cDNA microarray was conducted on 30 patients with gastric cancer. The analysis revealed an increased proliferation of regions on chromosome 7, including the MGAM gene, which is a target for drug development.ref. ccs370042-bib-0024 Vincent‐Chong et al.’s study identified copy number alterations (CNA) potentially linked to cancer in oral squamous cell carcinoma (OSCC) through high‐resolution array comparative genomic hybridization (aCGH). Genomic analyses have identified MGAM as a target for drug development because of its amplified expression in OSCC. Furthermore, the study demonstrated a 6.6‐fold increase in MGAM expression in OSCC.ref. ccs370042-bib-0063 Shi et al. have reported that MGAM exhibits a distinct expression pattern in breast cancer among Caucasian and Asian Americans.ref. ccs370042-bib-0064 In research conducted by Shutan Xu and colleagues, it was revealed that the expression of MGAM2 is predominantly found in BLBCs. Interestingly, this expression correlates with improved survival rates among breast cancer patients. In contrast, MGAM is more commonly associated with luminal A breast cancers. The identification of MGAM2 and other ER resident proteins, such as ERV1 homologs (EHVs), has implications for cancer immunotherapy and the treatment of BLBCs. The study further expanded on the distribution of MGAM2, demonstrating its presence across various tissues beyond just the breast. These tissues include the blood and the GI tract, indicating a broader role for MGAM2 in the body. Notably, there was a significant correlation between MGAM2 expression and gene signatures indicative of immune cell activity, especially neutrophils within the bloodstream. Moreover, the findings underscored the relevance of MGAM2 expression in GI cancers, suggesting a potential link between MGAM2 and these types of malignancies.ref. ccs370042-bib-0020
WES analysis conducted on patients with anal canal squamous cell carcinoma (ACSCC) revealed that MGAM2 ranks among the top three most commonly mutated genes in this uncommon form of cancer.ref. ccs370042-bib-0065 MGAM2 has been linked to cancers associated with the carbohydrate metabolism pathway. Additionally, this gene was connected to patients with hereditary CRC and simultaneous GC.ref. ccs370042-bib-0066 Recently, a study has identified deleterious variants in the MGAM gene that may be linked to monogenic or oligogenic inheritance in cases of IBS. This discovery has positioned MGAM as a potential target for personalized treatment in individuals with this syndrome.ref. ccs370042-bib-0067
Whereas our study focuses on CRC, the widespread involvement of MGAM in various cancer types suggests its crucial role in cancer biology. The association of MGAM with cancers linked to carbohydrate metabolism pathways underscores the importance of metabolic reprogramming in cancer. This connection highlights the potential benefits of targeting metabolic pathways in cancer therapy. However, the complexity of cancer biology means that no single gene can fully predict treatment outcomes. Future research with functional studies should aim to integrate MGAM expression data with other molecular markers to develop more comprehensive predictive models for cancer therapy.
In light of the comprehensive analysis of MGAM and MGAM2 gene expressions within various cancer contexts, it is clear that these genes represented pivotal players in the complex nature of cancer biology. Their association with distinct cancer subtypes, disease progression stages, and immune cell dynamics not only sheds new light on the underlying mechanisms of cancer but also paves the way for the development of innovative diagnostic tools and prognostic indicators. Furthermore, the intriguing link between MGAM and alpha‐glucosidase inhibitors opens up exciting avenues for repurposing existing drugs in oncology, thereby offering hope for more effective and less toxic treatments. As our understanding deepens through continuous research and the identification of specific genetic variants like rs2960746, the potential of MGAM and MGAM2 in revolutionizing cancer care becomes even more apparent. However, conducting functional studies and updating databases will comprehensively complete our understanding of these genes’ roles in cancer. This commitment will be crucial in leveraging the full potential of MGAM and MGAM2 in advancing personalized medicine and ultimately enhancing patient outcomes.
In conclusion, our investigation provides robust evidence supporting the critical function of the MGAM and MGAM2 genes in cancer biology. This discovery opens up promising directions for further exploration into their utility in prognostication, diagnosis, and treatment strategies. To the best of our knowledge, this study marks the inaugural comprehensive examination of MGAM across a spectrum of cancer types. The implications of our findings highlight the potential of MGAM as a significant biomarker for assessing cancer prognosis. Furthermore, they offer valuable insights into the possible pathways through which MGAM might influence tumor development.
AUTHOR CONTRIBUTIONS
Conceptualization: Hanieh Azari; Methodology: Farzaneh Alizadeh, Rawaa Chasib Mezher, Hamid Fiuji and Hanieh Azari; Software: Reza Khayami and Hanieh Azari; Validation: Ibrahim Saeed Gataa; Formal analysis: Hamid Fiuji; Investigation: Hanieh Azari, Farzaneh Alizadeh, Majid Rajabian; Resources: Ladan Goshayeshi; Data curation: Reza Khayami; Writing—original draft preparation: Rawaa Chasib Mezher and Hanieh Azari; Writing—review and editing: Majid Rajabian; Visualization: Reza Khayami and Hanieh Azari; Supervisions: Seyed Mahdi Hassanian and Amir Avan; Project administration: Amir Avan: Funding acquisition: Amir Avan. All authors have read and agreed to the published version of the manuscript.
CONFLICT OF INTEREST STATEMENT
The authors declare no conflicts of interest.
ETHICS STATEMENT
Ethical approval for this study was obtained from the Mashhad University of Medical Sciences Review Board (IR.MUMS.MEDICAL.REC.1402.389).
Supplementary Materials

References
- Drug Repurposing and Prediction of Multiple Interaction Types via Graph Embedding.”. BMC Bioinformatics, 2023. [DOI | PubMed]
- Global Cancer Statistics 2022: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries.”. CA: A Cancer Journal for Clinicians, 2024. [DOI | PubMed]
- Drug Repurposing for Cancer Therapy.”. Signal Transduction and Targeted Therapy, 2024. [DOI | PubMed]
- Drug Repurposing for Cancer Therapy in the Era of Precision Medicine.”. Current Molecular Pharmacology, 2022. [DOI | PubMed]
- 5 Aggarwal, S. , S. S. Verma , S. Aggarwal and S. C. Gupta . 2021. “Drug Repurposing for Breast Cancer Therapy: Old Weapon for New Battle.”. Seminars in Cancer Biology. Elsevier.
- Computational Approaches to Drug Repurposing: Methods, Challenges, and Opportunities.”. Annual Review of Biomedical Data Science, 2024. [DOI]
- Drug Repositioning or Target Repositioning: a Structural Perspective of drug‐target‐indication Relationship for Available Repurposed Drugs.”. Computational and Structural Biotechnology Journal, 2020. [DOI | PubMed]
- Drug Repurposing: an Overview.”. Asian Journal of Pharmaceutical Research and Development, 2020
- Repurposing Existing Therapeutics, Its Importance in Oncology Drug Development: Kinases as a Potential Target.”. British Journal of Clinical Pharmacology, 2022. [DOI | PubMed]
- Drug Repurposing in Dermatology: Molecular Biology and Omics Approach: Intechopen.. 2020
- Oncogenic Signaling Pathways in the Cancer Genome Atlas.”. Cell, 2018. [DOI | PubMed]
- 12 Issa, N. T. , V. Stathias , S. Schürer and S. Dakshanamurthy . 2021. “Machine and Deep Learning Approaches for Cancer Drug Repurposing.”. Seminars in Cancer Biology. Elsevier.
- A Systematic Review of Computational Approaches to Understand Cancer Biology for Informed Drug Repurposing.”. Journal of Biomedical Informatics, 2023. [DOI | PubMed]
- Biomarkers of Human Colonic Cell Growth Are Influenced Differently by a History of Colonic Neoplasia and the Consumption of Acarbose.”. The Journal of nutrition, 2000. [DOI | PubMed]
- Suppression of Lysosomal Acid Alpha‐Glucosidase Impacts the Modulation of Transcription Factor EB Translocation in Pancreatic Cancer.”. Cancer Science, 2021. [DOI | PubMed]
- Study of the Inhibition of Two Human maltase‐glucoamylases Catalytic Domains by Different α‐glucosidase Inhibitors.”. Carbohydrate Research, 2011. [DOI | PubMed]
- Use of an α‐glucosidase Inhibitor and the Risk of Colorectal Cancer in Patients with Diabetes: a Nationwide, Population‐based Cohort Study.”. Diabetes Care, 2015. [DOI | PubMed]
- Cancer Cells Change Their Glucose Metabolism to Overcome Increased ROS: One Step from Cancer Cell to Cancer Stem Cell?”. Biomedicine & Pharmacotherapy, 2019. [DOI | PubMed]
- In‐vitro, in‐silico Studies on the Proposed Penasulfate A and Its Derivatives as Dual Inhibitors of α‐glucosidases and Human Cancer Cells.”. Journal of Molecular Structure, 2024. [DOI]
- Proteins with Evolutionarily Hypervariable Domains Are Associated with Immune Response and Better Survival of Basal‐like Breast Cancer Patients.”. Computational and Structural Biotechnology Journal, 2019. [DOI | PubMed]
- Genome‐Wide Association Study of 1, 5‐anhydroglucitol Identifies Novel Genetic Loci Linked to Glucose Metabolism.”. Scientific Reports, 2017. [DOI | PubMed]
- Characterization of the SGLT2 Interaction Network and Its Regulation by SGLT2 Inhibitors: a Bioinformatic Analysis.”. Frontiers in Pharmacology, 2022. [DOI | PubMed]
- Human Intestinal maltase–glucoamylase: Crystal Structure of the N‐terminal Catalytic Subunit and Basis of Inhibition and Substrate Specificity.”. Journal of Molecular Biology, 2008. [DOI | PubMed]
- Gene Amplifications at Chromosome 7 of the Human Gastric Cancer Genome.”. International Journal of Molecular Medicine, 2007. [DOI | PubMed]
- G‐Protein Signaling Modulator 2 as a Potential Biomarker in Colorectal Cancer: Integrative Analysis Using Genetic Profiling and Pan‐Cancer Studies.”. Genes, 2024. [DOI | PubMed]
- Firebrowser: an R Client to the Broad Institute’s Firehose Pipeline.”. Database, 2017. [DOI | PubMed]
- GSCA: an Integrated Platform for Gene Set Cancer Analysis at Genomic, Pharmacogenomic and Immunogenomic Levels.”. Briefings in Bioinformatics, 2023. [DOI | PubMed]
- The STRING Database in 2021: Customizable protein–protein Networks, and Functional Characterization of user‐uploaded gene/measurement Sets.”. Nucleic Acids Research, 2020. [DOI]
- Dgidb 5.0: Rebuilding the drug–gene Interaction Database for Precision Medicine and Drug Discovery Platforms.”. Nucleic Acids Research, 2024. [DOI | PubMed]
- SIFT Missense Predictions for Genomes.”. Nature Protocols, 2016. [DOI | PubMed]
- Predicting Functional Effect of Human Missense Mutations Using PolyPhen‐2.”. Current protocols in human genetics, 2013. [DOI]
- Variant Selection and Interpretation: an Example of Modified Varsome Classifier of ACMG Guidelines in the Diagnostic Setting.”. Genes, 2021. [DOI | PubMed]
- COSMIC: Exploring the World’s Knowledge of Somatic Mutations in Human Cancer.”. Nucleic Acids Research, 2015. [DOI | PubMed]
- Repurposing of Chronically Used Drugs in Cancer Therapy: a Chance to Grasp.”. Cancers, 2023. [DOI | PubMed]
- The Diagnostic and Prognostic Value of C1Orf174 in Colorectal Cancer.”. BioImpacts, 2024. [DOI | PubMed]
- Integrated Analysis of multi‐omics Data for the Discovery of Biomarkers and Therapeutic Targets for Colorectal Cancer.”. Computers in Biology and Medicine, 2023. [DOI | PubMed]
- The Prognostic Value of ASPHD1 and ZBTB12 in Colorectal Cancer: a Machine Learning‐based Integrated Bioinformatics Approach.”. Cancers, 2023. [DOI | PubMed]
- Drug Repurposing in Oncology: Compounds, Pathways, Phenotypes and Computational Approaches for Colorectal Cancer.”. Biochimica et Biophysica Acta (BBA)‐Reviews on Cancer, 2019. [DOI | PubMed]
- Omics‐Driven Drug Repurposing as a Source of Innovative Therapies in Rare Cancers.”. Expert Opinion on Orphan Drugs, 2018. [DOI]
- Pathway‐Based drug‐repurposing Schemes in Cancer: the Role of Translational Bioinformatics.”. Frontiers in Oncology, 2021. [DOI | PubMed]
- Repurposing Drugs Targeting Metabolic Diseases for Cancer Therapeutics.”. Drug Discovery Today, 2023. [DOI | PubMed]
- Drug Repositioning in Drug Discovery of T2DM and Repositioning Potential of Antidiabetic Agents.”. Computational and Structural Biotechnology Journal, 2022. [DOI | PubMed]
- The Effect of Metformin on Mortality Following Cancer Among Patients with Diabetes.”. Cancer Epidemiology, Biomarkers & Prevention, 2014. [DOI]
- Pharmacologic Therapy of Diabetes and Overall Cancer Risk and Mortality: a meta‐analysis of 265 Studies.”. Scientific Reports, 2015. [DOI | PubMed]
- The Effects of 6 Common Antidiabetic Drugs on Anti‐PD1 Immune Checkpoint Inhibitor in Tumor Treatment.”. Journal of Immunology Research, 2022. [DOI | PubMed]
- Increased Tumor Glycolysis Characterizes Immune Resistance to Adoptive T Cell Therapy.”. Cell Metabolism, 2018. [DOI]
- Metabolic Barriers to T Cell Function in Tumors.”. The Journal of Immunology, 2018. [DOI | PubMed]
- Targeting Metabolism for Cancer Therapy.”. Cell Chemical Biology, 2017. [DOI | PubMed]
- Immune Cells: Plastic Players Along Colorectal Cancer Progression.”. Journal of Cellular and Molecular Medicine, 2013. [DOI | PubMed]
- Comprehensive Outline of Whole Exome Sequencing Data Analysis Tools Available in Clinical Oncology.”. Cancers, 2019. [DOI | PubMed]
- Explore the Mechanism and Substance Basis of Mahuang Fuzixixin Decoction for the Treatment of Lung Cancer Based on Network Pharmacology and Molecular Docking.”. Computers in Biology and Medicine, 2022. [DOI | PubMed]
- Distinct Patterns of Somatic Genome Alterations in Lung Adenocarcinomas and Squamous Cell Carcinomas.”. Nature Genetics, 2016. [DOI | PubMed]
- Co‐mutations of Epidermal Growth Factor Receptor and BRAF in Chinese Non‐small Cell Lung Cancer Patients.”. Annals of Translational Medicine, 2021. [DOI | PubMed]
- Association of Survival and immune‐related Biomarkers with Immunotherapy in Patients with Non–small Cell Lung Cancer: a meta‐analysis and Individual patient–level Analysis.”. JAMA Network Open, 2019. [DOI | PubMed]
- Urinary Glycoproteomic Profiling of Non‐muscle Invasive and Muscle Invasive Bladder Carcinoma Patients Reveals Distinct N‐glycosylation Pattern of CD44, MGAM, and GINM1.”. Oncotarget, 2020. [DOI | PubMed]
- Candidate Serum Biomarkers for Early Intestinal Cancer Using 15N Metabolic Labeling and Quantitative Proteomics in the ApcMin/+ Mouse.”. Journal of Proteome Research, 2013. [DOI | PubMed]
- Comparing Urinary Glycoproteins Among Three Urogenital Cancers and Identifying Prostate Cancer‐specific Glycoproteins.”. ACS Omega, 2022. [DOI | PubMed]
- A seven‐gene Prognosis Model to Predict Biochemical Recurrence for Prostate Cancer Based on the TCGA Database.”. Frontiers in Surgery, 2022. [DOI | PubMed]
- Molecular Profiling of Peripheral Blood Is Associated with Circulating Tumor Cells Content and Poor Survival in Metastatic castration‐resistant Prostate Cancer.”. Oncotarget, 2015. [DOI | PubMed]
- Identification of Subgroups Along the glycolysis‐cholesterol Synthesis Axis and the Development of an Associated Prognostic Risk Model.”. Human Genomics, 2021. [DOI | PubMed]
- Integrated Profiling Revealed Potential Regulatory Networks Among Long Noncoding Rnas and Mrnas in Mucosal Gastric Cancer.”. Technology in Cancer Research and Treatment, 2020. [DOI | PubMed]
- Genome‐Wide Analysis of Gene Expression in Intestinal‐type Gastric Cancers Using a Complementary DNA Microarray Representing 23,040 Genes.”. Cancer Research, 2002. [PubMed]
- Genome Wide Analysis of Chromosomal Alterations in Oral Squamous Cell Carcinomas Revealed over Expression of MGAM and ADAM9.”. PLoS One, 2013. [DOI | PubMed]
- Integrative Comparison of Mrna Expression Patterns in Breast Cancers from Caucasian and Asian Americans with Implications for Precision Medicine.”. Cancer Research, 2017. [DOI | PubMed]
- Neoadjuvant PD‐1 Blockade Combined with Chemotherapy Followed by Concurrent Immunoradiotherapy in Locally Advanced Anal Canal Squamous Cell Carcinoma Patients: Antitumor Efficacy, Safety and Biomarker Analysis.”. Frontiers in Immunology, 2022. [DOI | PubMed]
- Analysis of Genomic Pathogenesis According to the Revised Bethesda Guidelines and Additional Criteria.”. Journal of Cancer Research and Clinical Oncology, 2021. [DOI | PubMed]
- Genetic Variants in Carbohydrate Digestive Enzyme and Transport Genes Associated with Risk of Irritable Bowel Syndrome.”. medRxiv:, 2023. [DOI]
