Neuromedin U receptor 1 deletion leads to impaired immunotherapy response and high malignancy in colorectal cancer
Abstract
Colorectal cancer (CRC) exhibits significant heterogeneity, impacting immunotherapy efficacy, particularly in immune desert subtypes. Neuromedin U receptor 1 (NMUR1) has been reported to perform a vital function in immunity and inflammation. Through comprehensive multi-omics analyses, we have systematically characterized NMUR1 across various tumors, assessing expression patterns, genetic alterations, prognostic significance, immune infiltration, and pathway associations at both the bulk sequencing and single-cell scales. Our findings demonstrate a positive correlation between NMUR1 and CD8+ T cell infiltration, with elevated NMUR1 levels in CD8+ T cells linked to improved immunotherapy outcomes in patients with CRC. Further, we have validated the NMUR1 expression signature in CRC cell lines and patient-derived tissues, revealing its interaction with key immune checkpoints, including lymphocyte activation gene 3 and cytotoxic T-lymphocyte-associated protein 4. Additionally, NMUR1 suppression enhances CRC cell proliferation and invasiveness. Our integrated analyses and experiments open new avenues for personalized immunotherapy strategies in CRC treatment.
Article type: Research Article
Keywords: Bioinformatics, Cancer, Microenvironment, Molecular biology, Therapeutics
Affiliations: Department of Oncology, Xiangya Hospital, Central South University, Changsha, Hunan 410008, China; Department of Microbiology, Immunology & Molecular Genetics, University of Texas Long School of Medicine, UT Health Science Center, San Antonio, TX 78229, USA; National Clinical Research Center for Geriatric Disorders, Xiangya Hospital, Central South University, Changsha, Hunan 410008, China; Department of Emergency, Hainan General Hospital, Hainan Affiliated Hospital of Hainan Medical University, Haikou, Hainan 570311, China; Department of Oncology, Yueyang People’s Hospital, Yueyang Hospital Affiliated to Hunan Normal University, Yueyang, Hunan 414000, China; Department of Hematology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou, Henan 450052, China
License: © 2024 The Authors CC BY 4.0 This is an open access article under the CC BY license (http://creativecommons.org/licenses/by/4.0/).
Article links: DOI: 10.1016/j.isci.2024.110318 | PubMed: 39055918 | PMC: PMC11269305
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (4.3 MB)
Introduction
Neuromedin U (NMU), a highly conserved neuropeptide, is known to regulate smooth muscle contraction, intestinal motility, and blood flow.ref. bib1,ref. bib2 NMU receptors (NMURs), belonging to the G-protein-coupled receptor (GPCR) family, exhibit distinct expression patternsref. bib3,ref. bib4; NMUR1 is primarily peripheral, while NMUR2 is central nervous system-oriented.ref. bib5,ref. bib6 Besides physiological function, accumulated studies have identified that NMU has a significant impact on tumorigenesis and metastasis in colorectal cancer (CRC), lung cancer, renal cancer, breast cancer, hepatocellular cancer, endometrial cancer, and myeloid leukemia.ref. bib7,ref. bib8,ref. bib9,ref. bib10,ref. bib11,ref. bib12,ref. bib13 And NMUR1 is expressed on various cell types, including macrophages and endothelial cells, known to be active modulators of the tumor microenvironment.ref. bib6 The interaction between NMU and Group 2 innate lymphoid cells (ILC2s), which are implicated in cytokine secretion and express NMUR1, underscores the importance of NMU-NMUR1 signaling in immunity and inflammation.ref. bib14,ref. bib15,ref. bib16,ref. bib17,ref. bib18 Furthermore, through an NMUR1-dependent manner, NMU was found to regulate anti-tumor activity of CD8+ T cells, which are the most powerful effectors in the anticancer immune response and form the backbone of current successful cancer immunotherapies.ref. bib19,ref. bib20 NMU has also been shown to be a candidate biomarker and therapeutic target in drug resistance.ref. bib21,ref. bib22 Although NMUR1 is widely expressed on many immune cells, such as macrophages, endothelial cells, ILC2, and CD8+ T cells, especially in the gastrointestinal tract, and is known to interact with NMU in the tumor microenvironment and drug resistance, its role in the CRC microenvironment and treatment remains unclear.
Tumor microenvironment (TME) is a dynamic and complicated component consisting of stromal cells, immune cells, and signaling molecules, except for tumor cells.ref. bib23,ref. bib24 Accumulating researches suggest that the ratio of tumor-infiltrating immune cells, stromal component, and cytokines have a significant impact on the regulation of tumor progression, evasion, metastasis, and evolution.ref. bib24,ref. bib25,ref. bib26,ref. bib27,ref. bib28,ref. bib29,ref. bib30 The complicated correlation between tumor and immune system influences the efficiency of traditional anticancer therapy strategies, such as surgery, radiotherapy, and chemotherapy. This interplay accelerates advances in tumor therapeutic methods that function by overcoming tumor evasion and optimizing the clinical benefit.ref. bib31,ref. bib32,ref. bib33,ref. bib34,ref. bib35 Immunotherapy is a novel anticancer therapy that suppresses cancer development by the activation of the immune system.ref. bib36,ref. bib37 As a promising therapeutic technique in immunotherapy, immune checkpoint inhibitors (ICIs) have been acknowledged to be involved in conquering tumor microenvironment immunosuppression and enhancing the immunological anti-cancer responses by blocking inhibitory immune checkpoints,ref. bib38,ref. bib39,ref. bib40,ref. bib41 such as cytotoxic T lymphocyte antigen-4 (CTLA4)ref. bib42 and programmed cell death protein 1 (PD-1),ref. bib43 which have shown survival benefits in clinical trials. Among emerging treatment strategies for patients with cancer, few have progressed as rapidly as neoadjuvant immune checkpoint blockade (ICB). Combining neoadjuvant chemotherapy with PD-1 blockade has shown increased expression of CD8+ T cell activation markers and reduced expression of markers related to immunosuppression.ref. bib44 Neoadjuvant ICB can induce a diverse range of inflammatory infiltrates, including CD8+ tumor-infiltrating lymphocytes (TIL), neutrophils, and plasma cells. It also has the potential to upregulate certain immune checkpoint pathways and remodel cancer-associated fibroblasts in the tumor bed.ref. bib45,ref. bib46 Despite these promising developments, there’s a pressing need to identify new biomarkers that can stratify patients based on their sensitivity to ICB and predict therapy response.ref. bib47
Based on the high expression of NMUR1 in the gastrointestinal tractref. bib1 and its close association with immune cells,ref. bib17 we hypothesized that NMUR1 played a crucial role in CRC. To identify the potential of NMUR1 as a biomarker in CRC, our study first conducted a comprehensive analysis of the function of NMUR1 in pan-cancer, including NMUR1 expression profiles, epigenetic alteration, the impact of survival, correlation with immune infiltration and associated pathways in 33 types of cancer. Given the important role of NMUR1 in pan-cancer, we aimed to investigate immune signatures and the function of NMUR1 in CRC cell lines and patient tissues by performing single-cell RNA sequencing and functional experiments. And we also revealed the prognosis value and immune signature of NMUR1 in the CRC patient cohort. In summary, this study uncovered that NMUR1 is a promising biomarker for immunotherapy response and function as a suppressor in CRC.
Results
Neuromedin U receptor 1 expression in human pan-cancer
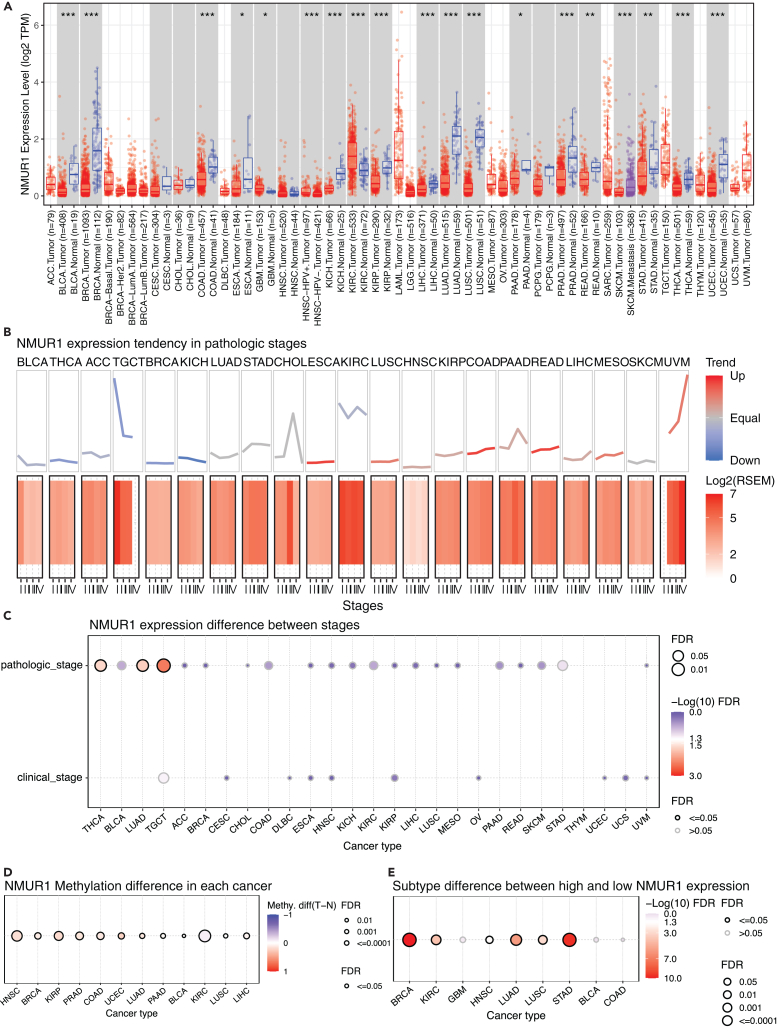
In our investigation into NMUR1’s role in oncogenesis, we explored the TIMER2 database to examine its expression levels across a spectrum of malignancies. Our analysis identified a pervasive pattern of NMUR1 downregulation in a variety of cancer types, including Bladder urothelial carcinoma (BLCA), Breast invasive carcinoma (BRCA), Colon adenocarcinoma (COAD), Esophageal carcinoma (ESCA), Glioblastoma multiforme (GBM), Head and Neck squamous cell carcinoma-HPVneg (HNSC-HPV-), Kidney chromophobe (KICH), Kidney renal clear cell carcinoma (KIRC), Kidney renal papillary cell carcinoma (KIRP), Liver hepatocellular carcinoma (LIHC), Lung adenocarcinoma (LUAD), Lung squamous cell carcinoma (LUSC), Pancreatic adenocarcinoma (PAAD), Prostate adenocarcinoma (PRAD), Rectum adenocarcinoma (READ), Skin Cutaneous Melanoma (SKCM), Stomach adenocarcinoma (STAD), Thyroid carcinoma (THCA), and Uterine Corpus Endometrial Carcinoma (UCEC) (Figure 1A). Further examination revealed that NMUR1 expression consistently varied with cancer progression, with discernible trends across different pathological stages (Figure 1B). Significantly, NMUR1’s expression was found to correlate with both pathological and clinical stages in THCA, LUAD, TGCT, and COAD, as corroborated by data from GSCA and TISIDB databases (Figures 1C and S1A). In consonance with its expression patterns in most cancers, NMUR1 demonstrated heightened methylation levels in HNSC, BRCA, KIRP, PRAD, COAD, and UCEC (Figure 1D). Further exploration of NMUR1’s correlation with molecular subtypes revealed marked subtype-specific signatures, particularly in BRCA, KIRC, LUAD, LUSC, and STAD according to the GSCA database (Figure 1E), and in BRCA, COAD, ESCA, LUSC, OV, PCPG, STAD, and UCEC based on the TISIDB database (Figure S1B). Therefore, the abnormal NMUR1 expression across pan-cancer in comparison to normal tissues implies its potential role in tumor initiation.
Prognostic analysis of neuromedin U receptor 1 expression in human pan-cancer
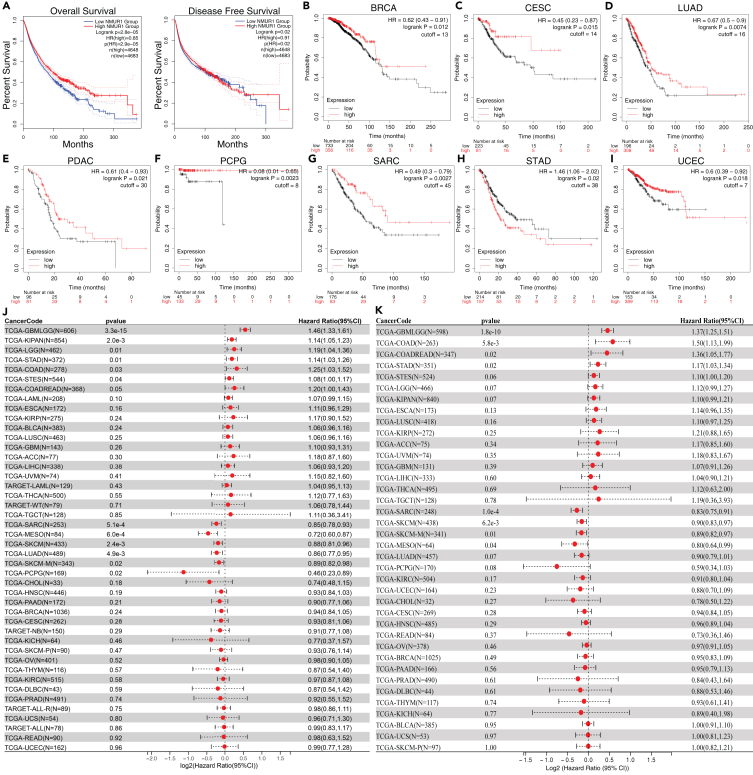
Subsequent investigations included a thorough assessment of overall survival (OS) and disease-free survival (DFS) across pan-cancer cohorts to explore the potential of NMUR1 as a prognostic biomarker. The aggregate data from 33 cancer types in the TCGA dataset compellingly indicated that higher NMUR1 expression is associated with improved OS and DFS (Figure 2A). Kaplan-Meier plotter analysesref. bib48 further reinforced the link between NMUR1 overexpression and enhanced OS for cancers, such as BRCA, Cervical squamous cell carcinoma (CESC), LUAD, Pancreatic ductal adenocarcinoma (PDAC), Pheochromocytoma and paraganglioma (PCPG), Sarcoma (SARC), and UCEC (Figures 2B–2I and S2). Moreover, Cox proportional hazards analyses revealed that lower NMUR1 expression is significantly related to poor patient prognoses in SARC (p = 2.2e-4), SKCM (p = 5.3e-4), Mesothelioma (MESO) (p = 6.0E-4), Metastatic Skin Cutaneous Melanoma (SKCM-M) (p = 9.5e-3), LUAD (p = 0.01), and PCPG (p = 0.02) (Figure 2J). A parallel trend was observed in disease-specific survival (DSS) metrics, with reduced NMUR1 expression aligning with adverse outcomes, notably in SARC (p = 1.0e-4), SKCM (p = 6.2e-3), and MESO (p = 0.04) (Figure 2K). Thus, NMUR1 emerges as a prognostic biomarker with potential implications for various malignancies.
Relationship between neuromedin U receptor 1 expression and immune infiltration in tumor microenvironment
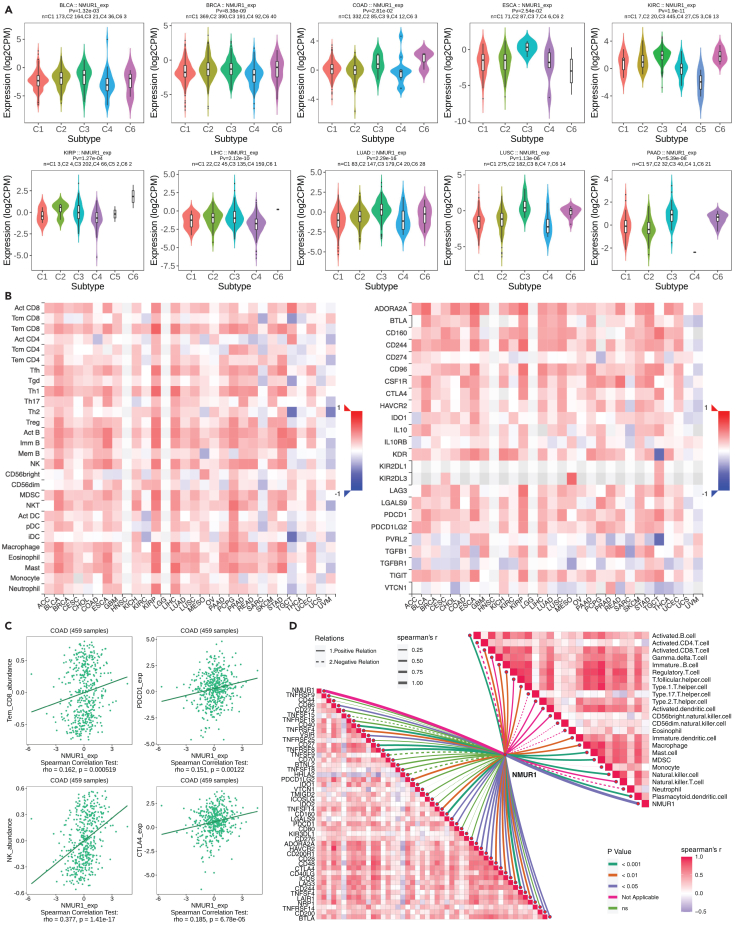
After establishing the expression and survival landscape of NMUR1 in pan-cancer, we explored the immune infiltration of NMUR1. Initial investigations revealed a pronounced positive correlation between NMUR1 and several immune subtypes (C1: wound healing; C2: IFN-gamma dominant; C3: inflammatory; C4: lymphocyte depleted; C5: immunologically quiet; C6: TGF-β dominant) in pan-cancer contexts, including BLCA (p = 1.32e-03), BRCA (p = 8.38e-09), COAD (p = 2.81e-02), ESCA (p = 2.54e-02), KIRC (p = 1.9e-11), KIRP (p = 1.27e-04), LIHC (p = 2.12e-10), LUAD (p = 2.29e-16), LUSC (p = 1.13e-06), and PAAD (p = 5.39e-06) (Figures 3A and S3). Further, NMUR1 was positively associated with a spectrum of immune cells, notably CD8+ T cells and NK cells, and immune inhibitors such as PDCD1 and CTLA4, across diverse cancers (Figure 3B). These associations were robustly supported by analyses using TIMER and xCELL databases (Figures S4A and S4B). These findings suggest that NMUR1 might play a significant role in immune response across a wide range of cancer types. Given the high expression of NMUR1 in the gastrointestinal tract,ref. bib1 we decided to focus our study on CRC. To substantiate our findings, we conducted a Spearman correlation analysis on the TCGA-COAD dataset, confirming a significant link between NMUR1 and memory CD8 T (Tem_CD8) cells (p = 0.000519), Natural killer (NK) cells (p = 1.41e-17), PDCD1 (p = 0.00122), and CTLA4 (p = 6.78e-05) (Figure 3C). Further exploration of TCGA-COAD data unveiled heightened NMUR1 expression in most immune cells (Figure S4C). Intriguingly, elevated NMUR1 expression corresponded with higher stromal, immune, and estimate scores (Figures S4D and S4E). Through meticulous analysis involving the ssGSEA algorithm applied to the TCGA-COAD dataset, we deciphered the intricate relationship between NMUR1 expression and various cancer related pathways. Notably, our investigation unveiled a strikingly positive association between NMUR1 and pivotal pathways implicated in tumor inflammation, Epithelial-Mesenchymal Transition (EMT), Extracellular Matrix (ECM) dynamics including degradation, angiogenesis, apoptosis, PI3K/AKT signaling, Transforming Growth Factor β (TGFB) signaling, and collagen formation. Conversely, a contrasting negative correlation was identified in pathways governing cellular response to G2M checkpoint, hypoxia, tumor proliferation, MYC targets, DNA repair, and DNA replication (Figure S5). These observations provide compelling insights into the multifaceted impact of NMUR1 on diverse molecular pathways within the context of COAD. Finally, RNA-Seq analysis within the Xiangya CRC validation cohort corroborated these findings, revealing consistent positive correlations between NMUR1 and various immune cell subtypes, such as CD8+ T cells and NK cells, as well as immune checkpoints such as CD244, LAG3, CD40, and PDCD1 (Figure 3D). Collectively, our data indicate that NMUR1 is involved in shaping the tumor microenvironment and modulating immune infiltration, establishing it as a promising biomarker for predicting the efficacy of immunotherapy.
Neuromedin U receptor 1 expression landscape in colorectal cancer single cell sequencing datasets
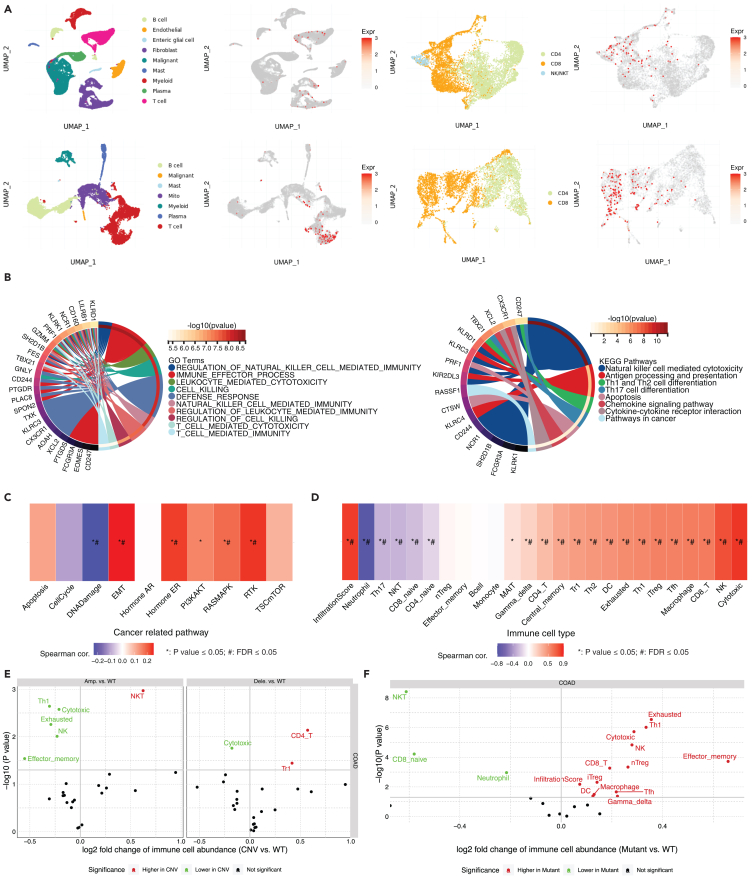
Building upon the established correlation between NMUR1 expression and the immune landscape of CRC, we further investigated NMUR1’s specific role within immune cells. Utilizing publicly available single-cell sequencing datasets, we observed a marked enrichment of NMUR1 within CD8+ T cells, aligning with our preliminary findings in Figures 3B and 3D (Figure 4A). This enrichment was corroborated by an extensive validation across eight CRC single-cell datasets within the TISIH database, which confirmed the heightened expression of NMUR1 in both CD8+ T cells and NK cells (Figure S6A). Subsequently, we delved into the top 50 genes exhibiting correlation across two single-cell datasets (CRC_GSE108989 and CRC_GSE146771_Smartseq2), depicted as an informative heatmap (Figure S6B). Through Gene Set Enrichment Analysis (GSEA), we determined that these genes were negatively enriched in COAD samples (Figure S6C). The application of Gene Set Variation Analysis (GSVA) enabled us to quantify the collective expression of these NMUR1-correlated genes, revealing that higher GSVA scores were indicative of a decreased mortality risk (Figure S6D) and were associated with earlier pathologic stages of COAD (Figure S6E). These scores were also found to be elevated in normal tissues (Figure S6F), earlier stages of COAD (Figure S6G), and the microsatellite instability-high (MSI-H) subtype (Figure S6H). Further examination through Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses highlighted a significant enrichment of NMUR1-correlated gene sets in biological processes and pathways integral to NK cell-mediated and T cell-mediated immunity (Figure 4B). Positive correlations were also identified between these gene sets and pivotal cancer-related pathways, including EMT, hormone ER, RAS/MAPK, and RTK signaling pathways (Figure 4C), as well as immune infiltration scores (Figure 4D). Notably, we observed a compelling link between the levels of copy number variations (CNVs) and single nucleotide variants (SNVs) in NMUR1-correlated gene sets and the extent of immune infiltration (Figures 4E and 4F). Together, these comprehensive data shed light on the profound role of NMUR1 within the CRC immune microenvironment and underscore its functional significance.
High neuromedin U receptor 1 expression in CD8+ T cell correlated with a better immunotherapy response in patients with colorectal cancer patients
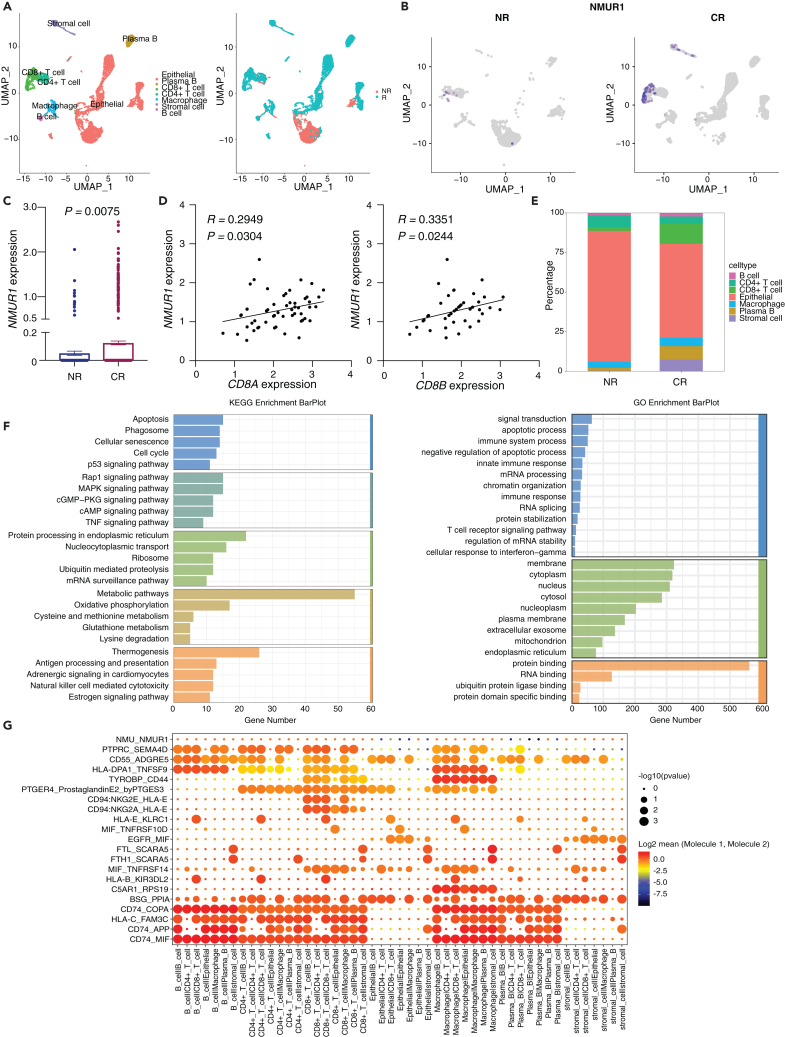
In probing the involvement of NMUR1 in the COAD immune microenvironment, our study made a significant advance by analyzing single-cell sequencing data from two patients with CRC post-immunotherapy—one non-responder (NR) and one complete responder (CR). Utilizing the SciBet platform for precise cell type identification,ref. bib49 we were able to categorize the diverse cell populations present in the samples (Figure 5A). Notably, NMUR1 expression was significantly upregulated in the sample from the responding patient, underscoring its potential role in mediating immunotherapy efficacy (Figures 5B and 5C). Additionally, NMUR1 showed a positive correlation with CD8+ T cell markers CD8A and CD8B (Figure 5D). Furthermore, a striking enrichment of CD8+ T cells, which are known to express NMUR1, was observed in the responding patient’s sample (Figures 5E and S7). In a deeper investigation into the functional implications of NMUR1 within T cells from NR and CR samples, we conducted a thorough GO enrichment and KEGG pathway analysis, revealing significant insights (Figures 5E and 5F). Additionally, regarding the role of NMUR1 as a receptor for NMU, we examined cellular communication at the single-cell level. Here, the NMU-NMUR1 interaction appears as a weak communicative pathway among immune cells within the CRC immune microenvironment, possibly due to the low secretion of NMU in immune cells (Figure 5G). Our finding was further supported by validation in two additional public single-cell RNA datasets, GSE205506ref. bib50 (Figure S8) and GSE222300ref. bib51 (Figure S9), where CD8+ T cell infiltration and NMUR1 enrichment were consistently observed in patients with CR. Collectively, these analyses across three independent datasets solidify the concept of NMUR1 as a determinant of immunotherapy response in CRC, enriching our understanding of its role in the immune landscape of cancer therapy.
Neuromedin U receptor 1 as a prospective immunotherapy biomarker for colorectal cancer patient stratification
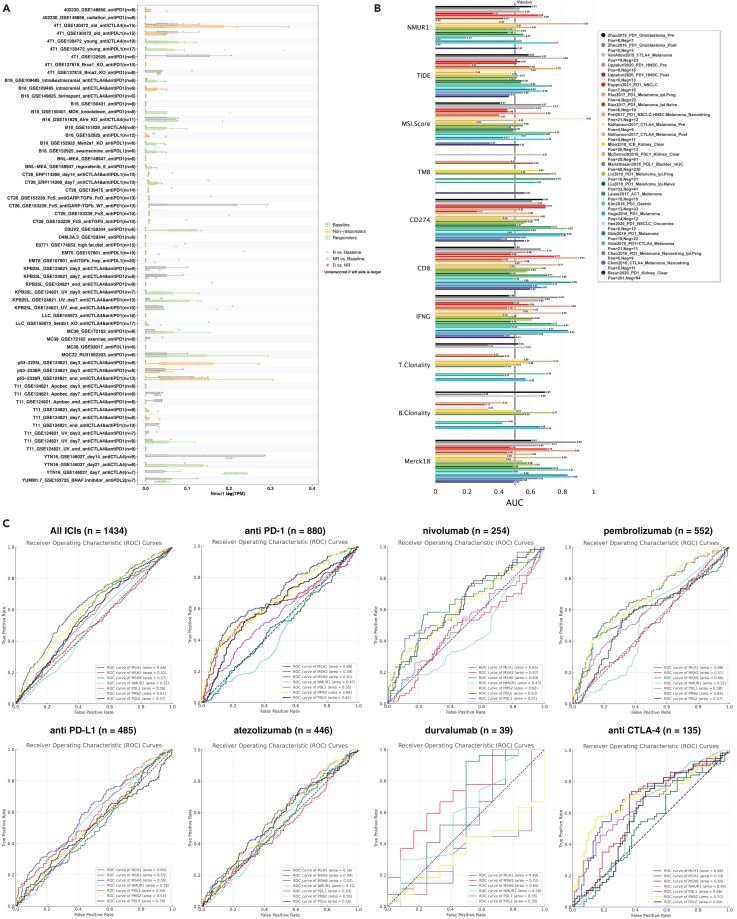
Given our insightful exploration into NMUR1’s enrichment in patients with CRC immunotherapy responder, we extended our analysis to assess its biomarker potential in publicly available immunotherapy cohorts. Utilizing the TISMO database, we conducted a comprehensive analysis of gene expression, pathways, and immune cell infiltration associated with ICB treatment. This analysis yielded a predictive model for immunotherapeutic response, revealing that NMUR1 expression is modulated by distinct ICB treatments across different cancer models (Figure 6A). The Receiver Operating Characteristic (ROC) curve, which plots the true positive rate against the false positive rate at various threshold settings, was employed as a key metric. The Area Under the ROC Curve (AUC) is particularly telling, serving as a measure of the predictive accuracy. Our results are promising, showing that NMUR1 alone achieved an AUC greater than 0.7 in two immunotherapy cohorts (Figure 6B), a performance comparable to well-established biomarkers such as TIDE, MSI score, Tumor Mutational Burden (TMB), and T and B Cell Clonality. When juxtaposed with biomarkers such as CD8, Interferon Gamma (IFNG), and the T cell-inflamed signature (Merck 18), which displayed AUCs greater than 0.7 in multiple cohorts, NMUR1 presented a modest predictive capacity. This could be attributed to the limited dataset sizes available for CRC cohorts. PD-L1, POLE, and MSI-H status are well recognized biomarkers for the guidance of immunotherapy.ref. bib47 We then analyzed the immunotherapy cohorts from pan-cancer to compare NMUR1 with those biomarkers in different ICB treatments (Figure 6C). Furthermore, validation in the GSE235919 dataset, comprising 23 responders and 11 non-responders to CRC immunotherapy, demonstrated that higher NMUR1 expression was significantly associated with increased CD8+ T cell infiltration as determined by the xCELL algorithm (Figure S10A). Moreover, NMUR1 expression was consistently elevated in responders, even though without statistical significance (Figure S10B). In a comparative evaluation of biomarker potential using AUC, NMUR1 (AUC = 0.626) outperformed PD-L1 (AUC = 0.555), underscoring its viability as a biomarker in CRC immunotherapy (Figure S10C–I). In summary, these results collectively underscore the potential of NMUR1 as a significant biomarker in the realm of CRC immunotherapy.
Deletion of neuromedin U receptor 1 potentiates the proliferation and invasion of colorectal cancer cells
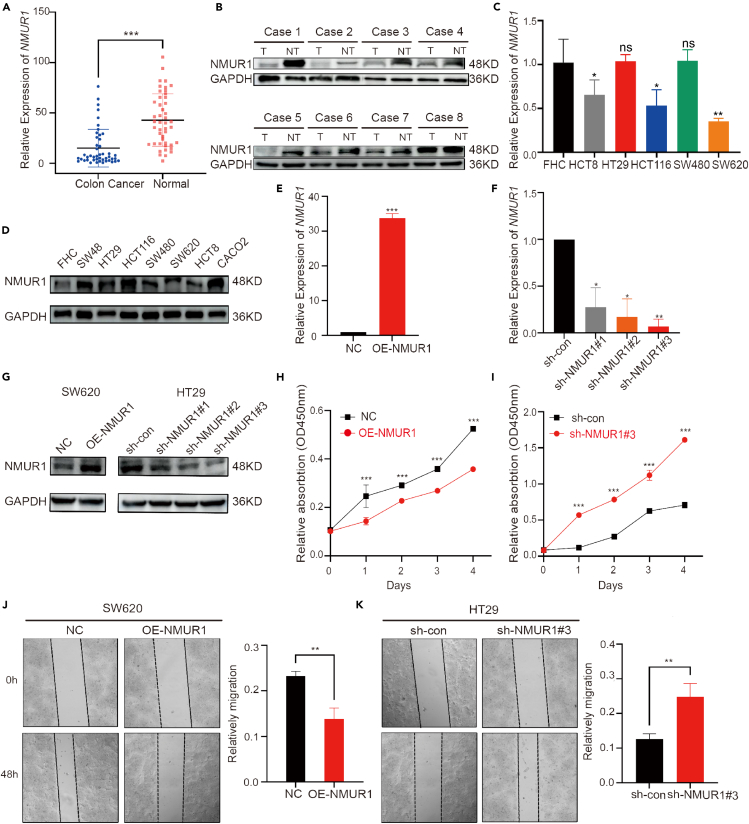
Analysis of total RNA from 46 CRC patient samples using RT-qPCR revealed a significant reduction of NMUR1 mRNA expression in tumor tissues compared to adjacent non-tumor tissues (Figure 7A). Further validation was performed using protein extracts from 8 pairs of CRC tissues and adjacent non-tumor tissues. Western blot analysis confirmed the reduced expression of NMUR1 in CRC tissues, aligning with our mRNA expression findings (Figure 7B). The disparity in NMUR1 expression suggests its potential involvement in the pathobiology of CRC cells. Subsequently, we examined the NMUR1 expression signature in five CRC cell lines alongside a normal intestinal mucosal epithelial cell line through qRT-PCR and western blotting. Most CRC cell lines demonstrated lower NMUR1 expression compared to the normal cell line (Figures 7C and 7D). HT29 cells, which exhibited relatively higher levels of NMUR1, were transfected with knockdown plasmids, while SW620 cells, with lower NMUR1 levels, were transfected with overexpression plasmids. The efficiency of these transfections was confirmed by RT-qPCR and western blot (Figures 7E–7G). To discern the impact of NMUR1 on CRC cell proliferation and metastatic potential, we employed the CCK8 assay, revealing that NMUR1 overexpression markedly decreased SW620 cell viability, whereas NMUR1 knockdown increased HT29 cell viability (Figures 7H and 7I). A wound healing assay indicated that NMUR1 overexpression significantly impeded the invasive capability of SW620 cells, whereas NMUR1 knockdown had the converse effect in HT29 cells. The differences in invasion between the overexpression and knockdown groups were statistically significant (Figures 7J and 7K). In conclusion, our investigations provide substantial evidence that NMUR1 may act as a tumor suppressor in CRC.
Neuromedin U receptor 1 prognostic value in Xiangya colorectal cancer cohort
In line with previous observations, NMUR1 expression levels in tissue samples were markedly higher in tumor tissues than in adjacent non-tumor tissues. To confirm NMUR1’s expression characteristics and prognostic value, we analyzed two independent real-world cohorts (Xiangya training cohort and Xiangya validation cohort), both of which echoed the findings from our bioinformatics analysis. Tissue microarrays from 64 patients with CRC in the Xiangya training cohort were used for immunohistochemical analysis to investigate NMUR1’s prognostic significance. Patients were stratified into high (36 patients) and low (28 patients) NMUR1 expression groups, with representative staining patterns depicted in Figure 8A. Log rank test results revealed that patients with lower NMUR1 expression had a significantly poorer overall survival (OS) compared to their higher-expression counterparts, corroborating our database analysis (Figure 8B). The prognostic accuracy, as indicated by AUC for survival rate, was 0.6756 (Figure 8C). Further, multivariate Cox regression analysis identified NMUR1 expression, tumor size, and clinical stage as independent predictors of OS (Figure 8D; Table 1). Additionally, qRT-PCR of 46 CRC patient tissues from the Xiangya validation cohort further supported these findings. Kaplan-Meier analysis showed that patients with high NMUR1 expression had a better OS compared to those with low expression (Figure 8E), with an impressive AUC of 0.72 for the survival rate (Figure 8F).
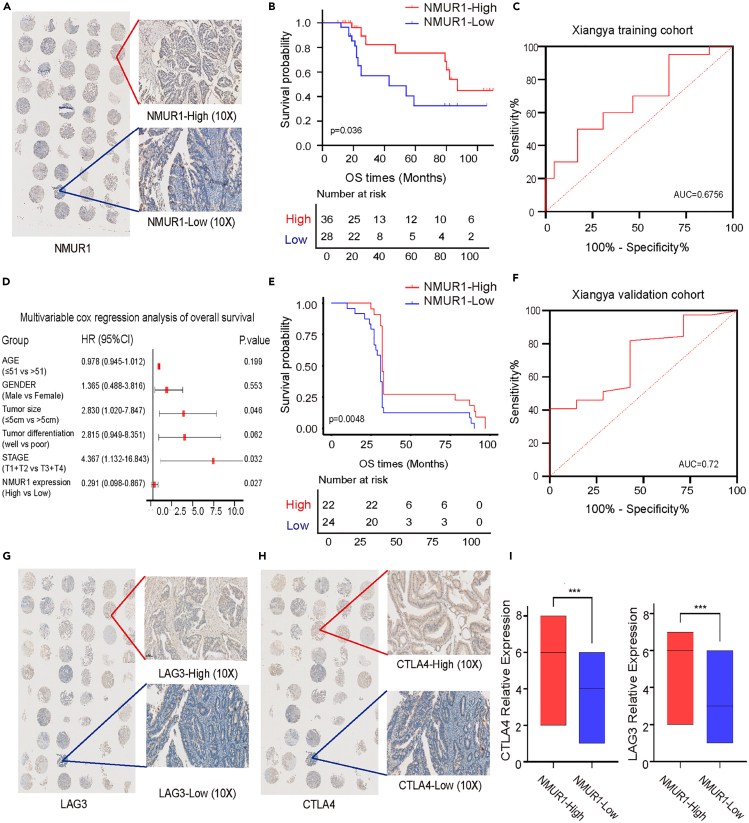
Table 1: Cox regression analysis of overall survival in the Xiangya CRC training cohort
| Variables | subgroup | Patients | Univariate analysis | Multivariate analysis | ||||
|---|---|---|---|---|---|---|---|---|
| HR | 95% CI | p Value | HR | 95% CI | p Value | |||
| Age | ≤50 | 11 | 0.986 | (0.957–1.015) | 0.343 | 0.978 | (0.945–1.012) | 0.199 |
| >50 | 53 | 1.000 | 1.000 | |||||
| Gender | Male | 40 | 1.209 | (0.493–2.963) | 0.678 | 1.365 | (0.488–3.816) | 0.553 |
| Female | 24 | 1.000 | 1.000 | |||||
| Tumor size | ≤5cm | 37 | 2.451 | (0.999–6.011) | 0.050 | 2.830 | (1.020–7.847) | 0.046 |
| >5cm | 27 | 1.000 | 1.000 | |||||
| Tumor differentiation | well | 40 | 3.018 | (1.152–7.904) | 0.025 | 2.815 | (0.949–8.351) | 0.062 |
| poor | 24 | 1.000 | 1.000 | |||||
| Stage | I + II | 37 | 6.488 | (1.894–22.218) | 0.003 | 4.367 | (1.132–16.843) | 0.032 |
| III+IV | 27 | 1.000 | 1.000 | |||||
| NMUR1 expression | High | 36 | 0.393 | (0.159–0.975) | 0.044 | 0.291 | (0.098–0.867) | 0.027 |
| Low | 28 | 1.000 | 1.000 | |||||
We also investigated NMUR1’s predictive value regarding immunotherapy response. Immunohistochemistry was used to validate the association between NMUR1 expression and immune checkpoints LAG3 and CTLA4, which were selected based on database analysis and clinical relevance (Figures 8G and 8H). We observed a significant positive correlation between the expression levels of NMUR1, LAG3, and CTLA4 (Figure 8I). These findings suggest that patients with higher NMUR1 expression might derive greater benefit from immunotherapy, which could account for their improved prognosis. Therefore, NMUR1 shows promise as a predictive biomarker for immunotherapy response.
Discussion
As a major receptor of NMU which was already considered as a tumor growth and/or progression marker in endometrial, renal, and breast cancers, NMUR1 could interact with NMU leading to autocrine tumor-promoting pathway activation, including intracellular Caflux, the phosphorylation of ERK1/2 kinases and hypoxia-inducible factors activation.ref. bib9,ref. bib10,ref. bib11 Considering that NMUR1 was mainly expressed in peripheral tissues especially in gut and lung,ref. bib1,ref. bib5,ref. bib52 NMUR1 was widely reported to be involved in airway inflammation and immune response,ref. bib15,ref. bib53 while few literatures revealed the association between NMUR1 and gastrointestinal immune response, especially in gastrointestinal cancer. Our study identified NMUR1 as a potential biomarker for CRC prognosis and immunotherapy response by combining multi-omics analyses and in vitro functional experiments. We proved that high NMUR1 expression contributed to the inhibition of CRC cells proliferation and invasion, and was positively correlated with immune checkpoints in patients with CRC’ tissues.
In this study, firstly, we explored the expression level of NMUR1 at pan-cancer level using TCGA and GEO databases. The results obtained reveal that NMUR1 had a significantly lower expression level in COAD, LUAD, and so forth, which is consistent with recent publications in CRC and lung cancer.ref. bib7,ref. bib11 NMUR1 expression level was also found to be a positive correlation with the different stages and molecular subtypes of certain cancers such as COAD, TGCT, BRCA, and so forth. For the NMUR1 prognostic values, we performed the analysis based on GEPIA2 and Kaplan-Meier Plotter. Similar to previous research, decreased NMUR1 expression correlated with a poor prognosis in certain cancers, including STAD, COAD, GBM, LGG, and BRCA, as well as in pan-cancer. Taken together, these pan-cancer data offered us a perspective that NMUR1 played a crucial role in the initiation and progression of tumors.
The TME has gained significant attention for its dynamic role in tumor promotion or suppression. Recent literature has categorized tumors as "cold" or "hot" based on the heterogeneity of TILs, stromal cells, and extracellular matrix components.ref. bib54 Insight distinction and alteration of hot tumor is beneficial to the intervention of immunotherapy strategies. We conducted a bioinformatics analysis which revealed a huge difference proportion of NMUR1 in different cancers. Notably, high expression of NMUR1 in TCGA-COAD was found in the C3 (Inflammatory) subtype which presented increased expression level of Th17 and Th1 and low cancer cells proliferation when compared with lower expression in C1 (Wound Healing) and C2 (IFN-γ Dominant) subtypes which herald boost cancer cells proliferation and worse prognosis. These results further indicated that NMUR1 might be an important molecular and biomarker in different subtypes of cancer.
TILs are diverse within the TME, affecting tumor metastasis, angiogenesis, and immunotherapy resistance.ref. bib55 CD4+ T cells, multiple subtypes including T helper type 1 (Th1), Th2, Th17, Th9, Treg and T follicular helper cells, contribute to diverse functions that are opposite in antitumor immunity leading to an opposite outcome.ref. bib56 CD8+ T cells which were also called cytotoxic T lymphocytes (CTLs) serve as a durable and critical antitumor immune effect via binding to membrane-expressed T cell receptors (TCR) leading to the activation of apoptosis.ref. bib57,ref. bib58 NK cells, a member of the innate lymphoid cell (ILC) family, have a significant impact on anti-tumor immunity which presents direct cytolytic activity interacted with target cells, up-regulated expression of TNF and death-inducing ligands, and a capacity to secrete multiple cytokines and chemokines.ref. bib59 Tumor-associated macrophages (TAM), macrophages infiltrated in tumor types, exerting antitumor or protumor function according to the different status, M1 (antitumor role) and M2 (protumor role). In this study, we initially discovered an association between NMUR1 expression and the proportion of immune cell infiltration in pan-cancer using TIMER2 and xCELL databases. In line with our hypothesis, NMUR1 was enriched in CD4+ T cells, CD8+ T cells, and macrophage cells across different cancers and positively connected with StromalScores, ImmuneScores, and ESTIMATE scores in pan-cancer. Based on the above results, NMUR1 showed a robust correlation with multi-types of tumor-associated immune infiltration cells.
TISCH database was selected to explore how NMUR1 affects CRC TME in single-cell insight. The results obtained show different immune cell distributions. Similar to previous results, higher NMUR1 expression was observed in CD4+ T cells and CD8+ T cells in five independent CRC datasets. Notably, Guozhong Xiao et al. found that CD4+ T cells in overweight/obese CRC exhibited higher expression of immune checkpoint molecules and immunosuppressive microenvironment.ref. bib60 NMU-NMUR1 signaling has also been investigated to show a connection with the metabolism and regulation of anti-tumor activity of CD8+ T cells and glycolysis of tumor cells.ref. bib19,ref. bib61,ref. bib62,ref. bib63 Our findings indicated a pronounced correlation between increased NMUR1 levels in CD8+ T cells and enhanced immunotherapy outcomes, suggesting that NMU-NMUR1 signaling could potentially alter CD8+ T cell metabolism in the CRC microenvironment. In addition, the ratio of NK cells exhibited a positive association with NMUR1 expression in the CRC_GSE146771_10X dataset. Therefore, the patients with high expression of NMUR1 might present a tumor suppressive TME and benefit more from immunotherapy strategies. Besides, a positive correlation with chemokines and related receptors was noticed in the high expression of NMUR1. Surprisingly, the remarkable results reported that NMU-NMUR1 signaling performs a vital function in immunity and allergic inflammation by the activation of ILC2s causing increased secretion of IL-5, IL-9, and IL-13.ref. bib16,ref. bib17,ref. bib18 Consistently, high expression of NMUR1 was found to correlate with the upregulated proportion of NK cell which is a member of the ILC family and secrets diverse chemokine.
TMB and MSI have been proven to increase mutation frequency which might lead to immunogenic neoantigens transcription and translation because of mismatch repair (MMR) deficiency. They serve as the predicting factors for immunotherapy efficacy.ref. bib64 A hot tumor feature of a high T cell infiltration degree was found to be effective in ICI therapy. For instance, CTLA4, a classic inhibitory receptor expressed in exhausted or dysfunctional TILs, inhibits the early activation and differentiation of T cells. Strategies used to inhibit CTLA4 have now been approved to be effective in the treatment of advanced-stage melanoma,ref. bib65 RCC,ref. bib66 and NSCLC.ref. bib67 We performed IHC to detect the relative protein expression of classic ICP genes, CTLA4, and LAG3 in the Xiangya cohort. TMB and MSI are predictors of immunotherapy efficacy, and our results suggest NMUR1 could be added to this repertoire, especially considering its positive correlation with immune checkpoint protein LAG3 and CTLA4, indicative of a prognostic and therapeutic biomarker for CRC.
NMU has been shown to play an important role in cancer through NMUR1 or NMUR2 dependent pathway. Przygodzka et al. had tested NMU mRNA expression and protein level in 6 CRC cell lines (Caco-2, HT29, SW620, HCT15, HCT116, and SW480).ref. bib11 Even though all the tested CRC cells expressed NMU mRNA, the NMU protein was only detectable in the lysates and cell-conditioned media of HCT15, HCT116, and SW480 cells. And NMU induced an invasive phenotype of CRC cells in a NMUR2 dependent way instead of NMUR1. Notably, CRC cells exhibited higher expression of NMUR2 and lower expression of NMUR1. And cognate residues for both NMUR1 and NMUR2 are completely conserved, thus making highly similar interactions with peptide.ref. bib68 Given these findings, we hypothesize that overexpression of NMUR1 might compete with NMUR2 for NMU binding, leading to a suppressive effect in CRC cells by interfering with NMUR2-dependent signaling. It is worthwhile to investigate if there is a competition between NMUR2 and NMUR1 for NMU in the future experiment.
Collectively, our study showed that downregulated expression of NMUR1 was correlated with worse prognosis and immune infiltration in CRC. NMUR1 has emerged as an effective predictor of CRC patient prognosis and immunotherapy response, highlighting its significant role in inhibiting CRC proliferation and invasion. Thus, NMUR1 represents a promising biomarker for prognosis and immunotherapy response in CRC.
Limitations of the study
There are still some limitations need to be considered in our study. Even though we thoroughly searched the public database, there are still extremely few cohorts for CRC immunotherapy. In the future, we need to increase the sample size and demographic diversity. The association between the expression of NMUR1 and CTLA4 is conflicting between IHC experiment results in the Xiangya cohort and the bioinformatics analysis. This may be due to the limited bioinformatics database and the inaccurate assessment of the complex and heterogeneous TME using tissue microarray. The information from databases exhibited conflict and specificity deficiency and the verification of the NMUR1 expression signature was only performed in CRC tissue samples and CRC cell lines. The definite molecular mechanisms and exact pathways of how NMUR1 participates in tumor progression and invasion, and immune regulation remain are still required to be investigated in mice model and in vitro experiments. It has been proven that NMU may be an important resistance-enhancing factor and display an anorexigenic effect in animal models,ref. bib21,ref. bib61 deeper insight into NMUR1 as a cancer cachexia regulator and drug target would be critical in clinical significance.
STAR★Methods
Key resources tables
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| Anti-hu-NMUR1 | Invitrogen | Cat. # PA5-30376; RRID: AB_2547850 |
| Anti-GAPDH | Abcam | Cat. # ab181602; RRID: AB_2630358 |
| Anti-hu-LAG3 | CST | Cat. # 15372; RRID: AB_2798739 |
| Anti-hu-CTLA4 | Abcam | Cat. # ab237712; RRID: AB_2905652 |
| Biological samples | ||
| Human CRC and adjacent non-tumor tissues | Xiangya Hospital of Central South University | N/A |
| Chemicals, peptides, and recombinant proteins | ||
| Trizol | Invitrogen | Cat. # 10296010 |
| RPMI-1640 medium | HyClone | Cat. # SH30027.FS |
| L-15 medium | Gibco | Cat. # 21-083-027 |
| Fetal Bovine Serum | Gibco | Cat. # 10437028 |
| Immobilon Western HRP substrate | Millipore | Cat. # WBKLS0500 |
| Critical commercial assays | ||
| SYBR Green fluorescent-based assay | TaKaRa Bio Inc | Cat. # 638320 |
| PrimeScript RT-PCR Kit | TaKaRa Bio Inc | Cat. # RR014B |
| Lipofectamine 3000 | Invitrogen | Cat. #L3000008 |
| Cell Counting Kit 8 | Dojindo | Cat. # CK04 |
| Deposited data | ||
| Single cell RNA-seq data | This study | GSA-Human: HRA006722 & HRA007564 |
| Bulk sequencing data | This study | GSA-Human: HRA007645 |
| Public single cell RNA-seq data | GEO | GSE108989GSE136394GSE139555GSE146771GSE166555GSE179784GSE205506GSE222300GSE235919 |
| Experimental models: cell lines | ||
| Human CRC cell lines and normal intestinal epithelial cells | Institutes of Biomedical Sciences | N/A |
| Oligonucleotides | ||
| NMUR1 | 5′-GCCGGAGACAAGTGACCAAGA-3′ | 5′-TGACACGACGCTCCACATG-3′ |
| GAPDH | 5′-CTGGGCTACACTGAGCACC-3′ | 5′-AAGTGGTCGTTGAGGGCAATG-3′ |
| Software and algorithms | ||
| R studio Version 1.2.1335 (R version4.1.1) | RStudio, Inc. | https://www.rstudio.com/ |
| GraphPadPrism7.0 | GraphPad | www.graphpad.com |
| Adobe Illustrator 2023 | Adobe | http://aotucad2.xmjfg.com/pg/230.html |
| ImageJ | ImageJ | https://imagej.nih.gov/ij/ |
| ImageLabsoftware | Bio-Rad | www.bio-rad.com |
| OmicStudio | LC-Bio Technology Co | https://www.omicstudio.cn/tool |
| SPSS20.0 | IBM | https://spss.en.softonic.com/ |
Resource availability
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the lead contact Dr. Shan Zeng (zengshan2000@csu.edu.cn).
Materials availability
This study did not generate unique reagents.
Data and code availability
- All the public datasets can be downloaded in the Cancer Genome Atlas (TCGA) (https://cancergenome.nih.gov/), Gene Expression Omnibus (GEO) (https://www.ncbi.nlm.nih.gov/geo/) database, and UCSC Xena (https://xena.ucsc.edu/) database. Bulk sequencing data can be found in supplementary table and are available on GSA human database (https://ngdc.cncb.ac.cn/gsa-human/) under accession number HRA007645. Single cell sequencing data are available on GSA human database under accession number HRA006722 & HRA007564. As per local regulations, human sequencing data cannot be made publicly accessible. Access can be requested by following the guidance from GSA Human database (https://ngdc.cncb.ac.cn/gsa-human/document/GSA-Human_Request_Guide_for_Users_us.pdf).ref. bib91,ref. bib92
- This paper does not report original code.
- Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
Experimental model and study participant details
Patients and follow-up
From October 2011 to November 2019, clinical samples were gathered from patients diagnosed with CRC at Xiangya Hospital of Central South University. In the Xiangya cohort, a total of 110 patients were recruited for the retrospective study. Follow-up was completed in November 2021. Detailed information of CRC patients was acquired from the Xiangya digital medical records system. Overall survival (OS) was chosen as the endpoint of this study. We separated 110 patients into a training cohort (64 patients) and a validation cohort (46 patients). This study was reviewed and approved by the Xiangya Hospital Medical Ethics Committee of Central South University (No.201905131), and got the consent from all participates. Sex/gender as a variable factor did not show significant difference.
Cell lines and cell culture
Human CRC cell lines SW480, SW620, HCT8, HT29, HCT116 and human normal intestinal epithelial cell FHC were supplied by the Institutes of Biomedical Sciences (IBS, Shanghai, China). Cell lines were cultured in RPMI-1640 medium (HyClone, Cat. # SH30027.FS, UT) or L-15 medium (Gibco, Grand Island, NY) with 10% FBS (Gibco, Grand Island, NY) and were cultured in a humidified 37°C and 5% CO2 atmosphere.
Method details
Data collection
The Cancer Genome Atlas (TCGA) and Genotype-tissue expression (GTEx) profiles and clinical information were collected from the UCSC Xena (https://xena.ucsc.edu/) database for this investigation.ref. bib69 R software (version 4.1.1) and related website databases were used to assist in the analysis of the data.
Differential expression of NMUR1 analysis
The expression differences of NMUR1 in pan-cancer were investigated using TCGA and GTEx. TISIDB (http://cis.hku.hk/TISIDB/index.php) was chosen to examine NMUR1 expression in TCGA tumor samples from various immunological subtypes and molecular subtypes.ref. bib70 To identify expression differences between tumor and normal tissues, we employed the R "limma" package.ref. bib71 Boxplots were generated using the R "ggplot2″ package.ref. bib72 The methylation analysis was performed by GSCA tool, which performed correlation analyses for each gene to identify the methylation site most negatively correlated with the gene’s expression.ref. bib73
Prognostic value analysis
GEPIA2 (http://gepia.cancer-pku.cn/) was elected to detect OS and DFS survival data for NMUR1 in all tumor types. Kaplan–Meier plotter (http://kmplot.com/analysis/) was utilized to evaluate the prognosis of NMUR1 expression in various cancers,ref. bib74 and the best performing cutoff was determined as stated in Nagy et al.ref. bib48 By using SangerBox online database,ref. bib75 we investigated the impact of NMUR1 on patients’ survival in pan-cancer according to univariate Cox regression.
Assessment of immune infiltration levels
Immunedeconv, an R package that combines with TIMER, xCELL, MCP-counter, CIBERSORT, EPIC and quanTIseq algorithms, was considered to explore immune scores.ref. bib76 To determine the Stromal, Immune and ESTIMATE scores in each sample based on NMUR1 expression, we chose the R "ESTIMATE" package.ref. bib77 GSVA packageref. bib78 was chosen to estimate NMUR1 immune cell infiltration level. The markers for 24 immune cells were obtained from an article in Immunity.ref. bib79 Besides, we downloaded the pan-cancer dataset from the UCSC (https://xenabrowser.net/) database, and further extracted NMUR1 expression and immune-related genes including chemokine, receptor, MHC, Immunoinhibitor and Immunostimulator in each sample.
NMUR1 related gene functional and pathway enrichment investigation
We collected the gene sets contained in the relevant pathwaysref. bib80 and calculated the enrichment score of NMUR1 on each pathway according to the single sample Gene Set Enrichment Analysis (ssGSEA) algorithm, so as to obtain the relationship between NMUR1 and gene ontology (GO) biological function. By calculating the correlation between gene expression and pathway score, we can get the gene-pathway relationship. The R "GSVA" package was used to perform a Spearman correlation analysis on the relationship between NMUR1 and pathway scores.ref. bib78,ref. bib80
Single cell sequencing data analysis
CancerSEA (http://biocc.hrbmu.edu.cn/CancerSEA/home.jsp) is an appropriative online database utilized to investigate distinct functional states across various cancers at a single-cell level.ref. bib81 Based on single-cell sequencing profiling, we studied the NMUR1 function characteristics in 14 different states and drew a heatmap to explore the differences in pan-cancer. TISCH (https://doi.org/10.1093/nar/gkaa1020) is a scRNA-seq atlas that provides single-cell transcriptomic profiles data collected from GEOref. bib82 and ArrayExpress,ref. bib83 which was chosen for a comprehensive exploration of the TME components and its correlation of NMUR1 expression at the single-cell level.ref. bib84 We utilized the TIGER database (http://tiger.canceromics.org/#/home) to show NMUR1 expression in two CRC single cell sequencing.ref. bib85,ref. bib86 In addition, to assess the correlation between NMUR1 and immune infiltration in colon cancer, we applied the GSCA R package to calculate the NMUR1 score, and further explored the correlation of the score with multiple immune cells, CNV, and SNV.
Biomarker analysis
The data comparing gene expression levels across different tumor models and ICB treatments, between pre- and post-ICB treatment and responders and non-responders were acquired from the Tumor Immune Syngeneic MOuse (TISMO, http://tismo.cistrome.org/). TIDE platform can perform the comparison between the custom biomarker and other published biomarkers based on their predictive power of response outcome and overall survival.ref. bib87,ref. bib88 ROC plotter is a platform to investigate biomarkers of immunotherapy response in a large cohort of solid tumor samples.ref. bib89
Western blot
We carried out western blot to analyze the relative protein expression under standard technique. Briefly, the CRC cell (5×106 cells) and CC tissue samples (40mg) were lysed by RIPA cell lysis (NCM Biotech, Suzhou, China) with protease inhibitor and phosphatase inhibitor. The proteins samples (20ug) were then loaded in a 12.5% SDS PAGE and transferred to PVDF membrane under constant 300 mA at 90 min. The membranes were incubated in NMUR1(1:1000) and GAPDH (1:10000) Rabbit Polyclonal antibody at 4°C overnight (NMUR1, #PA5-30376, Invitrogen, USA; GAPDH, ab181602, Abcam, USA). The membranes were incubated in HRP-conjugated IgG for 2 h at 4°C after being washed by TBST. The chemiluminescence substrate Immobilon (Merck Millipore, Cat#WBKLS0500, Germany) was employed for signal detection according to the manufacturer’s instructions. The ChemiDoc XRS+ System was used to automatically detect the signals (Bio-Rad, Hercules, CA). GAPDH protein expression was chosen as internal control.
RNA extraction and real-time quantitative PCR (RT-qPCR)
The relative mRNA levels were assessed by reverse transcription PCR (RT-PCR). Total RNA was isolated from the CRC cells and CRC tissue samples by Trizol reagent (Invitrogen, Cat. #10296010, CA), and the PrimeScript RT-PCR Kit (TaKaRa Bio Inc., Cat. # RR014B, Japan) was performed for cDNA synthesis. RT-qPCR was performed on a ViiATM7 RT-PCR system (Applied Biosystems, Carlsbad, CA) using SYBR Green fluorescent-based assay (TaKaRa Bio Inc., Cat. #638320). After normalizing to GAPDH expression level, the relative expression level of NMUR1 was determined using the 2−(ΔΔCt) method. Sequences of primers:
NMUR1: forward 5′-GCCGGAGACAAGTGACCAAGA-3′; reverse 5′-TGACACGACGCTCCACATG-3′.
GAPDH: forward:5′-CTGGGCTACACTGAGCACC-3′; reverse 5′-AAGTGGTCGTTGAGGGCAATG-3′.
Construction of plasmid and transfection
SW620 and HT29 human colon cancer cells were selected according to their expression levels of NMUR1 and were transfected with lentivirus expressing shRNAs targeting NMUR1 and lentivirus selectively expressing NMUR1 (GeneChem Inc, Shanghai, China) by using Lipofectamine 3000 (Invitrogen, Carlsbad, CA). The target sequences of shRNAs were as follows: sh-NMUR1 #1 (5′-ccGGAGACAAGTGACCAAGAT-3′), sh-NMUR1 #2 (5′-cgCTACTGTTTGAGATGGTCT-3′), and sh-NMUR1 #3 (5′-gcGCACGCCTACCAACTACTA-3′). Cells were cultured for 72 h following transfection and selected in puromycin (3 μg/mL). Effective knockdown and overexpression were evaluated by fluorescence microscopy. RT-qPCR and western blot were conducted to verify the transcriptional and translational transfection efficiency.
Cell Counting Kit 8 (CCK8)
A CCK8 Kit (Dojindo, Tokyo, Japan) was used for determining cell viability. 2.0 × 103/well cells were seeded and 10 μL/well CCK8 was added into 96-well plates. The plates were incubated at 37°C for 2 h and were then measured using OD 490 nm absorbance values each day.
Wound healing assay
Wound-healing experiments were used to determine cell invasion abilities. Briefly, HT29 and SW620 cells (1×106/well) were seeded into 6-well plates. A wound was generated in the plate scratched by a 10 μL pipette tip. The floating cells were gently washed by PBS and cultured in a serum-free RPMI-1640 or L-15 medium. The wound healing status was observed under a microscope and representative fields pictures were captured at 0 and 48 h.
Immunohistochemistry staining (IHC)
The tissue microarray (TMA) was established from tissue samples of 64 CRC patients after follow-up. CRC tissue sample was immersed in paraformaldehyde and embedded in paraffin to constitute tissue microarray. After slicing into 4 μm slides, dimethyl benzene was used for dewaxing and ethanol was used for hydration, EDTA buffer was chosen for antigen retrieval at 95°C for 20 min 10% goat serum was used to block charged groups for 30 min. Then the slides were incubated with NMUR1 (1:100), LAG3 (1:200) and CTLA4 (1:500) Rabbit Polyclonal antibody at 4°C overnight (NMUR1, #PA5-30376, Invitrogen, USA; LAG3, #15372, CST, USA; CTLA4, ab237712, Abcam, USA). The secondary antibody was incubated for 30 min. After DAB staining, the slides were dehydrated, transparent, sealed with gumdrops and scanned by a Multispectral imaging system. IHC staining score was evaluated by 3 separate pathologists according to the percentage of positive staining cells (0–25%, 1; 26%–50%, 2; 51%–75%, 3; 76–100%, 4) and staining intensity (no staining, 0; Yellow, 1; Light brown, 2; Brown, 3). The results were sorted into two groups based on the sum of the percentage of positive cells and staining intensity scores: high expression (total score ≥4) and low expression (total score<4).
Single cell RNA sequencing and bulk RNA sequencing of real-world cohort
2 fresh tumor samples were collected from CRC patients received immunotherapy of Xiangya Hospital of Central South University. One patient showed no response (NR) and the other showed good response. Sequencing was performed according to the 10x Single Cell 3′ Reagent Kit. The furthering analysis was based on R software. 55 fresh tumor samples were collected from CRC patients of Xiangya Hospital of Central South University and were stored in liquid nitrogen. After TRIzol (Invitrogen, Cat. # 10296010, CA, USA) treated, total RNA was extracted from tissues samples for sequencing. Raw reads were processed using the Illumina Hiseq. Trimmomatic was utilized for quality control. Reads per kilobase per million reads (RPKM) was used for calculating gene expression levels. Bioinformatic analysis was performed using the OmicStudio tools at https://www.omicstudio.cn/tool.ref. bib90
Quantification and statistical analysis
Data processing, statistical analysis, and visualization were performed in R software (version 4.1.1), SPSS 23, and GraphPad Prism 9 software. All results of this study were estimated for statistical significance using Wilcoxon test, Kruskal-Wallis test, Spearman coefficients, Student t-test, and ANOVA appropriately. The Kaplan-Meier method and log rank test were selected to perform survival analysis. Univariate and multivariable Cox regressions were utilized to calculate HRs and explore the predictive biomarkers that independently affected CRC prognosis. Statistical significance was defined as p values or adjusted p values less than 0.05.
References
- A.D. Howard, R. Wang, S.S. Pong, T.N. Mellin, A. Strack, X.M. Guan, Z. Zeng, D.L. Williams, S.D. Feighner, C.N. Nunes. Identification of receptors for neuromedin U and its role in feeding. Nature, 2000. [PubMed]
- P.J. Brighton, P.G. Szekeres, G.B. Willars. Neuromedin U and its receptors: structure, function, and physiological roles. Pharmacol. Rev., 2004. [PubMed]
- L. Shan, X. Qiao, J.H. Crona, J. Behan, S. Wang, T. Laz, M. Bayne, E.L. Gustafson, F.J. Monsma, J.A. Hedrick. Identification of a novel neuromedin U receptor subtype expressed in the central nervous system. J. Biol. Chem., 2000. [PubMed]
- L.K. Malendowicz, M. Rucinski. Neuromedins NMU and NMS: An Updated Overview of Their Functions. Front. Endocrinol., 2021. [DOI]
- R. Raddatz, A.E. Wilson, R. Artymyshyn, J.A. Bonini, B. Borowsky, L.W. Boteju, S. Zhou, E.V. Kouranova, R. Nagorny, M.S. Guevarra. Identification and characterization of two neuromedin U receptors differentially expressed in peripheral tissues and the central nervous system. J. Biol. Chem., 2000. [PubMed]
- P. Przygodzka, K. Soboska, E. Sochacka, J. Boncela. Neuromedin U: A Small Peptide in the Big World of Cancer. Cancers, 2019. [DOI]
- K. Takahashi, C. Furukawa, A. Takano, N. Ishikawa, T. Kato, S. Hayama, C. Suzuki, W. Yasui, K. Inai, S. Sone. The neuromedin U-growth hormone secretagogue receptor 1b/neurotensin receptor 1 oncogenic signaling pathway as a therapeutic target for lung cancer. Cancer Res., 2006. [PubMed]
- Q. Li, L. Han, S. Ruan, S. Shen, Q. Cao, X. Cai, Y. Yan, B. Peng, Y. Hua. The prognostic value of neuromedin U in patients with hepatocellular carcinoma. BMC Cancer, 2020. [DOI | PubMed]
- S.K. Harten, M.A. Esteban, D. Shukla, M. Ashcroft, P.H. Maxwell. Inactivation of the von Hippel-Lindau tumour suppressor gene induces Neuromedin U expression in renal cancer cells. Mol. Cancer, 2011. [DOI | PubMed]
- S.E. Shetzline, R. Rallapalli, K.J. Dowd, S. Zou, Y. Nakata, C.R. Swider, A. Kalota, J.K. Choi, A.M. Gewirtz. Neuromedin U: a Myb-regulated autocrine growth factor for human myeloid leukemias. Blood, 2004. [PubMed]
- P. Przygodzka, E. Sochacka, K. Soboska, M. Pacholczyk, I. Papiewska-Pająk, T. Przygodzki, P. Płociński, S. Ballet, A. De Prins, J. Boncela. Neuromedin U induces an invasive phenotype in CRC cells expressing the NMUR2 receptor. J. Exp. Clin. Cancer Res., 2021. [DOI | PubMed]
- V.G. Martinez, S. O’Neill, J. Salimu, S. Breslin, A. Clayton, J. Crown, L. O’Driscoll. Resistance to HER2-targeted anti-cancer drugs is associated with immune evasion in cancer cells and their derived extracellular vesicles. OncoImmunology, 2017. [DOI]
- X. Yang, C.C. Wang, W.Y.W. Lee, J. Trovik, T.K.H. Chung, J. Kwong. Long non-coding RNA HAND2-AS1 inhibits invasion and metastasis in endometrioid endometrial carcinoma through inactivating neuromedin U. Cancer Lett., 2018. [DOI | PubMed]
- C.S.N. Klose, T. Mahlakõiv, J.B. Moeller, L.C. Rankin, A.-L. Flamar, H. Kabata, L.A. Monticelli, S. Moriyama, G.G. Putzel, N. Rakhilin. The neuropeptide neuromedin U stimulates innate lymphoid cells and type 2 inflammation. Nature, 2017. [DOI | PubMed]
- A. Wallrapp, S.J. Riesenfeld, P.R. Burkett, R.-E.E. Abdulnour, J. Nyman, D. Dionne, M. Hofree, M.S. Cuoco, C. Rodman, D. Farouq. The neuropeptide NMU amplifies ILC2-driven allergic lung inflammation. Nature, 2017. [DOI | PubMed]
- V. Cardoso, J. Chesné, H. Ribeiro, B. García-Cassani, T. Carvalho, T. Bouchery, K. Shah, N.L. Barbosa-Morais, N. Harris, H. Veiga-Fernandes. Neuronal regulation of type 2 innate lymphoid cells via neuromedin U. Nature, 2017. [DOI | PubMed]
- Y. Ye, Z. Liang, L. Xue. Neuromedin U: potential roles in immunity and inflammation. Immunology, 2021. [DOI | PubMed]
- P. Wibisono, S. Wibisono, J. Watteyne, C.-H. Chen, D. Sellegounder, I. Beets, Y. Liu, J. Sun. Neuronal GPCR NMUR-1 regulates distinct immune responses to different pathogens. Cell Rep., 2022. [DOI]
- R. Zheng, S. Wang, J. Wang, M. Zhou, Q. Shi, B. Liu. Neuromedin U regulates the anti-tumor activity of CD8+ T cells and glycolysis of tumor cells in the tumor microenvironment of pancreatic ductal adenocarcinoma in an NMUR1-dependent manner. Cancer Sci., 2024. [DOI | PubMed]
- M. Philip, A. Schietinger. CD8+ T cell differentiation and dysfunction in cancer. Nat. Rev. Immunol., 2022. [DOI | PubMed]
- S. Rani, C. Corcoran, L. Shiels, S. Germano, S. Breslin, S. Madden, M.S. McDermott, B.C. Browne, N. O’Donovan, J. Crown. Neuromedin U: A Candidate Biomarker and Therapeutic Target to Predict and Overcome Resistance to HER-Tyrosine Kinase Inhibitors. Cancer Res., 2014. [DOI | PubMed]
- S. You, L. Gao. Identification of NMU as a potential gene conferring alectinib resistance in non-small cell lung cancer based on bioinformatics analyses. Gene, 2018. [DOI | PubMed]
- J.A. Joyce, D.T. Fearon. T cell exclusion, immune privilege, and the tumor microenvironment. Science (New York, N.Y.), 2015. [DOI | PubMed]
- H. Zhang, Y. Zhou, B. Cui, Z. Liu, H. Shen. Novel insights into astrocyte-mediated signaling of proliferation, invasion and tumor immune microenvironment in glioblastoma. Biomed. Pharmacotherapy, 2020. [DOI]
- Y. Lin, J. Xu, H. Lan. Tumor-associated macrophages in tumor metastasis: biological roles and clinical therapeutic applications. J. Hematol. Oncol., 2019. [DOI | PubMed]
- M. Binnewies, E.W. Roberts, K. Kersten, V. Chan, D.F. Fearon, M. Merad, L.M. Coussens, D.I. Gabrilovich, S. Ostrand-Rosenberg, C.C. Hedrick. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat. Med., 2018. [DOI | PubMed]
- K.V. Woan, J.S. Miller. Harnessing Natural Killer Cell Antitumor Immunity: From the Bench to Bedside. Cancer Immunol. Res., 2019. [DOI | PubMed]
- P. Berraondo, M.F. Sanmamed, M.C. Ochoa, I. Etxeberria, M.A. Aznar, J.L. Pérez-Gracia, M.E. Rodríguez-Ruiz, M. Ponz-Sarvise, E. Castañón, I. Melero. Cytokines in clinical cancer immunotherapy. Br. J. Cancer, 2019. [DOI | PubMed]
- C. Cai, Y. Peng, E. Shen, R. Wan, L. Gao, Y. Gao, Y. Zhou, Q. Huang, Y. Chen, P. Liu. Identification of tumour immune infiltration-associated snoRNAs (TIIsno) for predicting prognosis and immune landscape in patients with colon cancer via a TIIsno score model. EBioMedicine, 2022. [DOI]
- Y. Gao, Y. Zhou, L. Wei, Z. Feng, Y. Chen, P. Liu, Y. Peng, Q. Huang, L. Gao, Y. Liu. Hsa_Circ_0066351 Acts as a Prognostic and Immunotherapeutic Biomarker in Colorectal Cancer. Front. Immunol., 2022. [DOI]
- M.W.L. Teng, S.F. Ngiow, A. Ribas, M.J. Smyth. Classifying Cancers Based on T-cell Infiltration and PD-L1. Cancer Res., 2015. [DOI | PubMed]
- A. Facciabene, G.T. Motz, G. Coukos. T-regulatory cells: key players in tumor immune escape and angiogenesis. Cancer Res., 2012. [DOI | PubMed]
- G. Xie, H. Dong, Y. Liang, J.D. Ham, R. Rizwan, J. Chen. CAR-NK cells: A promising cellular immunotherapy for cancer. EBioMedicine, 2020. [DOI]
- H. Zhang, B. Cui, Y. Zhou, X. Wang, W. Wu, Z. Wang, Z. Dai, Q. Cheng, K. Yang. B2M overexpression correlates with malignancy and immune signatures in human gliomas. Sci. Rep., 2021. [DOI | PubMed]
- H. Zhang, Y. Zhou, Q. Cheng, Z. Dai, Z. Wang, F. Liu, F. Fan, B. Cui, H. Cao. PDIA3 correlates with clinical malignant features and immune signature in human gliomas. Aging (Albany NY), 2020. [DOI | PubMed]
- A.D. Waldman, J.M. Fritz, M.J. Lenardo. A guide to cancer immunotherapy: from T cell basic science to clinical practice. Nat. Rev. Immunol., 2020. [DOI | PubMed]
- K. Yang, Z. Wu, H. Zhang, N. Zhang, W. Wu, Z. Wang, Z. Dai, X. Zhang, L. Zhang, Y. Peng. Glioma targeted therapy: insight into future of molecular approaches. Mol. Cancer, 2022. [DOI | PubMed]
- S. Sadreddini, B. Baradaran, A. Aghebati-Maleki, S. Sadreddini, D. Shanehbandi, A. Fotouhi, L. Aghebati-Maleki. Immune checkpoint blockade opens a new way to cancer immunotherapy. J. Cell. Physiol., 2019. [DOI | PubMed]
- S.L. Topalian, C.G. Drake, D.M. Pardoll. Immune checkpoint blockade: a common denominator approach to cancer therapy. Cancer Cell, 2015. [DOI | PubMed]
- S.L. Topalian. Targeting Immune Checkpoints in Cancer Therapy. JAMA, 2017. [DOI | PubMed]
- P. Sharma, J.P. Allison. The future of immune checkpoint therapy. Science (New York, N.Y.), 2015. [DOI | PubMed]
- A. Derakhshani, S. Hashemzadeh, Z. Asadzadeh, M.A. Shadbad, F. Rasibonab, H. Safarpour, V. Jafarlou, A.G. Solimando, V. Racanelli, P.K. Singh. Cytotoxic T-Lymphocyte Antigen-4 in Colorectal Cancer: Another Therapeutic Side of Capecitabine. Cancers, 2021. [DOI]
- H. Zhang, Z. Dai, W. Wu, Z. Wang, N. Zhang, L. Zhang, W.-J. Zeng, Z. Liu, Q. Cheng. Regulatory mechanisms of immune checkpoints PD-L1 and CTLA-4 in cancer. J. Exp. Clin. Cancer Res., 2021. [DOI | PubMed]
- T. Cascone, C.H. Leung, A. Weissferdt, A. Pataer, B.W. Carter, M.C.B. Godoy, H. Feldman, W.N. William, Y. Xi, S. Basu. Neoadjuvant chemotherapy plus nivolumab with or without ipilimumab in operable non-small cell lung cancer: the phase 2 platform NEOSTAR trial. Nat. Med., 2023. [DOI | PubMed]
- P. Qin, H. Chen, Y. Wang, L. Huang, K. Huang, G. Xiao, C. Han, J. Hu, D. Lin, X. Wan. Cancer-associated fibroblasts undergoing neoadjuvant chemotherapy suppress rectal cancer revealed by single-cell and spatial transcriptomics. Cell Rep. Med., 2023. [DOI]
- S.L. Topalian, P.M. Forde, L.A. Emens, M. Yarchoan, K.N. Smith, D.M. Pardoll. Neoadjuvant immune checkpoint blockade: A window of opportunity to advance cancer immunotherapy. Cancer Cell, 2023. [DOI | PubMed]
- A. Sveen, S. Kopetz, R.A. Lothe. Biomarker-guided therapy for colorectal cancer: strength in complexity. Nat. Rev. Clin. Oncol., 2020. [DOI | PubMed]
- Á. Nagy, G. Munkácsy, B. Győrffy. Pancancer survival analysis of cancer hallmark genes. Sci. Rep., 2021. [DOI | PubMed]
- C. Li, B. Liu, B. Kang, Z. Liu, Y. Liu, C. Chen, X. Ren, Z. Zhang. SciBet as a portable and fast single cell type identifier. Nat. Commun., 2020. [DOI | PubMed]
- J. Li, C. Wu, H. Hu, G. Qin, X. Wu, F. Bai, J. Zhang, Y. Cai, Y. Huang, C. Wang. Remodeling of the immune and stromal cell compartment by PD-1 blockade in mismatch repair-deficient colorectal cancer. Cancer Cell, 2023. [DOI | PubMed]
- L. Guo, Y. Wang, W. Yang, C. Wang, T. Guo, J. Yang, Z. Shao, G. Cai, S. Cai, L. Zhang. Molecular Profiling Provides Clinical Insights Into Targeted and Immunotherapies as Well as Colorectal Cancer Prognosis. Gastroenterology, 2023. [DOI | PubMed]
- T.D. Westfall, G.P. McCafferty, M. Pullen, S. Gruver, A.C. Sulpizio, V.N. Aiyar, J. Disa, L.C. Contino, I.J. Mannan, J.P. Hieble. Characterization of neuromedin U effects in canine smooth muscle. J. Pharmacol. Exp. Ther., 2002. [PubMed]
- Y. Ye, J. Luo, N. Zeng, S. Jiang, W. Chen, R.D. Hoyle, P. Klenerman, I.D. Pavord, L. Xue. Neuromedin U promotes human type 2 immune responses. Mucosal Immunol., 2022. [DOI | PubMed]
- J. Galon, D. Bruni. Approaches to treat immune hot, altered and cold tumours with combination immunotherapies. Nat. Rev. Drug Discov., 2019. [DOI | PubMed]
- E. Pérez-Ruiz, I. Melero, J. Kopecka, A.B. Sarmento-Ribeiro, M. García-Aranda, J. De Las Rivas. Cancer immunotherapy resistance based on immune checkpoints inhibitors: Targets, biomarkers, and remedies. Drug Resist. Updat., 2020. [DOI]
- M. Ben Khelil, Y. Godet, S. Abdeljaoued, C. Borg, O. Adotévi, R. Loyon. Harnessing Antitumor CD4+ T Cells for Cancer Immunotherapy. Cancers, 2022. [DOI]
- B. Farhood, M. Najafi, K. Mortezaee. CD8+ cytotoxic T lymphocytes in cancer immunotherapy: A review. J. Cell. Physiol., 2019. [DOI | PubMed]
- C. Hao, X. Wu, R. Zhou, H. Zhang, Y. Zhou, X. Wang, Y. Feng, L. Mei, C. He, X. Cai, L. Wu. Downregulation of p66Shc can reduce oxidative stress and apoptosis in oxidative stress model of marginal cells of stria vascularis in Sprague Dawley rats. Drug Des. Devel. Ther., 2019. [DOI]
- F. Souza-Fonseca-Guimaraes, J. Cursons, N.D. Huntington. The Emergence of Natural Killer Cells as a Major Target in Cancer Immunotherapy. Trends Immunol., 2019. [DOI | PubMed]
- G. Xiao, Y. Zheng, H. Chen, M. Luo, C. Yang, D. Ren, P. Qin, H. Zhang, H. Lin. Single-cell transcriptome analysis reveals immunosuppressive landscape in overweight and obese colorectal cancer. J. Transl. Med., 2024. [DOI | PubMed]
- R. Hanada, H. Teranishi, J.T. Pearson, M. Kurokawa, H. Hosoda, N. Fukushima, Y. Fukue, R. Serino, H. Fujihara, Y. Ueta. Neuromedin U has a novel anorexigenic effect independent of the leptin signaling pathway. Nat. Med., 2004. [DOI | PubMed]
- A.M. Peier, K. Desai, J. Hubert, X. Du, L. Yang, Y. Qian, J.R. Kosinski, J.M. Metzger, A. Pocai, A.R. Nawrocki. Effects of peripherally administered neuromedin U on energy and glucose homeostasis. Endocrinology, 2011. [DOI | PubMed]
- H. Teranishi, R. Hanada. Neuromedin U, a Key Molecule in Metabolic Disorders. Int. J. Mol. Sci., 2021. [DOI]
- D.T. Le, J.N. Uram, H. Wang, B.R. Bartlett, H. Kemberling, A.D. Eyring, A.D. Skora, B.S. Luber, N.S. Azad, D. Laheru. PD-1 Blockade in Tumors with Mismatch-Repair Deficiency. N. Engl. J. Med., 2015. [DOI | PubMed]
- J.D. Wolchok, V. Chiarion-Sileni, R. Gonzalez, P. Rutkowski, J.-J. Grob, C.L. Cowey, C.D. Lao, J. Wagstaff, D. Schadendorf, P.F. Ferrucci. Overall Survival with Combined Nivolumab and Ipilimumab in Advanced Melanoma. N. Engl. J. Med., 2017. [DOI | PubMed]
- R.J. Motzer, N.M. Tannir, D.F. McDermott, O. Arén Frontera, B. Melichar, T.K. Choueiri, E.R. Plimack, P. Barthélémy, C. Porta, S. George. Nivolumab plus Ipilimumab versus Sunitinib in Advanced Renal-Cell Carcinoma. N. Engl. J. Med., 2018. [DOI | PubMed]
- M.D. Hellmann, T.-E. Ciuleanu, A. Pluzanski, J.S. Lee, G.A. Otterson, C. Audigier-Valette, E. Minenza, H. Linardou, S. Burgers, P. Salman. Nivolumab plus Ipilimumab in Lung Cancer with a High Tumor Mutational Burden. N. Engl. J. Med., 2018. [DOI | PubMed]
- C. You, Y. Zhang, P. Xu, S. Huang, W. Yin, H. Eric Xu, Y. Jiang. Structural insights into the peptide selectivity and activation of human neuromedin U receptors. Nat. Commun., 2022. [DOI | PubMed]
- T. Chen, X. Chen, S. Zhang, J. Zhu, B. Tang, A. Wang, L. Dong, Z. Zhang, C. Yu, Y. Sun. The genome sequence archive family: toward explosive data growth and diverse data types. Dev. Reprod. Biol., 2021. [DOI]
- Database Resources of the National Genomics Data Center, China National Center for Bioinformation in 2024. Nucleic Acids Res., 2024. [DOI | PubMed]
- J. Vivian, A.A. Rao, F.A. Nothaft, C. Ketchum, J. Armstrong, A. Novak, J. Pfeil, J. Narkizian, A.D. Deran, A. Musselman-Brown. Toil enables reproducible, open source, big biomedical data analyses. Nat. Biotechnol., 2017. [DOI | PubMed]
- B. Ru, C.N. Wong, Y. Tong, J.Y. Zhong, S.S.W. Zhong, W.C. Wu, K.C. Chu, C.Y. Wong, C.Y. Lau, I. Chen. TISIDB: an integrated repository portal for tumor-immune system interactions. Bioinformatics, 2019. [DOI | PubMed]
- M.E. Ritchie, B. Phipson, D. Wu, Y. Hu, C.W. Law, W. Shi, G.K. Smyth. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res., 2015. [DOI | PubMed]
- V. Gómez-Rubio. ggplot2 – Elegant Graphics for Data Analysis (2nd Edition). J. Statist. Softw., 2017. [DOI]
- C.-J. Liu, F.-F. Hu, G.-Y. Xie, Y.-R. Miao, X.-W. Li, Y. Zeng, A.-Y. Guo. GSCA: an integrated platform for gene set cancer analysis at genomic, pharmacogenomic and immunogenomic levels. Brief. Bioinform., 2023. [DOI]
- A. Lánczky, B. Győrffy. Web-Based Survival Analysis Tool Tailored for Medical Research (KMplot): Development and Implementation. J. Med. Internet Res., 2021. [DOI]
- W. Shen, Z. Song, X. Zhong, M. Huang, D. Shen, P. Gao, X. Qian, M. Wang, X. He, T. Wang. Sangerbox: A comprehensive, interaction-friendly clinical bioinformatics analysis platform. iMeta, 2022. [DOI | PubMed]
- G. Sturm, F. Finotello, M. List. Immunedeconv: An R Package for Unified Access to Computational Methods for Estimating Immune Cell Fractions from Bulk RNA-Sequencing Data. Methods Mol. Biol., 2020. [DOI | PubMed]
- K. Yoshihara, M. Shahmoradgoli, E. Martínez, R. Vegesna, H. Kim, W. Torres-Garcia, V. Treviño, H. Shen, P.W. Laird, D.A. Levine. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun., 2013. [DOI | PubMed]
- S. Hänzelmann, R. Castelo, J. Guinney. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinf., 2013. [DOI]
- G. Bindea, B. Mlecnik, M. Tosolini, A. Kirilovsky, M. Waldner, A.C. Obenauf, H. Angell, T. Fredriksen, L. Lafontaine, A. Berger. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity, 2013. [DOI | PubMed]
- J. Wei, K. Huang, Z. Chen, M. Hu, Y. Bai, S. Lin, H. Du. Characterization of Glycolysis-Associated Molecules in the Tumor Microenvironment Revealed by Pan-Cancer Tissues and Lung Cancer Single Cell Data. Cancers, 2020. [DOI]
- H. Yuan, M. Yan, G. Zhang, W. Liu, C. Deng, G. Liao, L. Xu, T. Luo, H. Yan, Z. Long. CancerSEA: a cancer single-cell state atlas. Nucleic Acids Res., 2019. [DOI | PubMed]
- T. Barrett, S.E. Wilhite, P. Ledoux, C. Evangelista, I.F. Kim, M. Tomashevsky, K.A. Marshall, K.H. Phillippy, P.M. Sherman, M. Holko. NCBI GEO: archive for functional genomics data sets–update. Nucleic Acids Res., 2013. [DOI | PubMed]
- A. Athar, A. Füllgrabe, N. George, H. Iqbal, L. Huerta, A. Ali, C. Snow, N.A. Fonseca, R. Petryszak, I. Papatheodorou. ArrayExpress update – from bulk to single-cell expression data. Nucleic Acids Res., 2019. [DOI | PubMed]
- D. Sun, J. Wang, Y. Han, X. Dong, J. Ge, R. Zheng, X. Shi, B. Wang, Z. Li, P. Ren. TISCH: a comprehensive web resource enabling interactive single-cell transcriptome visualization of tumor microenvironment. Nucleic Acids Res., 2021. [DOI | PubMed]
- J. Qian, S. Olbrecht, B. Boeckx, H. Vos, D. Laoui, E. Etlioglu, E. Wauters, V. Pomella, S. Verbandt, P. Busschaert. A pan-cancer blueprint of the heterogeneous tumor microenvironment revealed by single-cell profiling. Cell Res., 2020. [DOI | PubMed]
- T.D. Wu, S. Madireddi, P.E. de Almeida, R. Banchereau, Y.J.J. Chen, A.S. Chitre, E.Y. Chiang, H. Iftikhar, W.E. O’Gorman, A. Au-Yeung. Peripheral T cell expansion predicts tumour infiltration and clinical response. Nature, 2020. [DOI | PubMed]
- J. Fu, K. Li, W. Zhang, C. Wan, J. Zhang, P. Jiang, X.S. Liu. Large-scale public data reuse to model immunotherapy response and resistance. Genome Med., 2020. [DOI | PubMed]
- P. Jiang, S. Gu, D. Pan, J. Fu, A. Sahu, X. Hu, Z. Li, N. Traugh, X. Bu, B. Li. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med., 2018. [DOI | PubMed]
- S.A. Kovács, J.T. Fekete, B. Győrffy. Predictive biomarkers of immunotherapy response with pharmacological applications in solid tumors. Acta Pharmacol. Sin., 2023. [DOI | PubMed]
- F. Lyu, F. Han, C. Ge, W. Mao, L. Chen, H. Hu, G. Chen, Q. Lang, C. Fang. OmicStudio: A composable bioinformatics cloud platform with real-time feedback that can generate high-quality graphs for publication. iMeta, 2023. [DOI | PubMed]
