AUNIP was a candidate marker for prognosis and immunology in pan-cancer
Abstract
Electronic supplementary material:
The online version of this article (10.1007/s13205-025-04294-6) contains supplementary material, which is available to authorized users.
Article type: Research Article
Keywords: AUNIP, Pan-cancer, Prognostic, Immunity, Hepatocellular carcinoma
Affiliations: https://ror.org/03s8txj32grid.412463.60000 0004 1762 6325Department of Pathology, The Second Affiliated Hospital of Harbin Medical University, Harbin, 150086 Heilongjiang China; https://ror.org/013xs5b60grid.24696.3f0000 0004 0369 153XDepartment of Pathology, Beijing Ditan Hospital, Capital Medical University, No. 8 Jing Shun East Street, Chaoyang District, Beijing, 100015 China; https://ror.org/02s7c9e98grid.411491.8Department of Pathology, The Fourth Affiliated Hospital of Harbin Medical University, Harbin, 150081 Heilongjiang China; Cancer Center, Department of Pathology, Zhejiang Provincial People’s Hospital, Affiliated People’s Hospital, Hangzhou Medical College, Hangzhou, Zhejiang China
License: © The Author(s) 2025 CC BY 4.0 Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
Article links: DOI: 10.1007/s13205-025-04294-6 | PubMed: 40386627 | PMC: PMC12085456
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (4.8 MB)
Introduction
Cancer is a serious disease that becomes a threat to mankind’s health. The number of deaths from cancer is increasing every year. Pan-cancer research has been prevalent in recent years, with the aim of integrating TCGA data based on different tumor types and platforms, while analyzing and interpreting these data. Our research relies on multi-omics database to explore differences between tumors, guiding tumor diagnosis, prognosis, and treatment selection (Zhang and Wang ref. 2015; Yang et al. ref. 2018).
AUNIP (Aurora kinase A and Ninein-interacting protein) is a centrosomal protein that interacts to promote the maintenance of Aurora-A and Ninein centrosome structures and the formation of spindles (Zhang and Wang ref. 2015). AUNIP regulates the mitotic entry and mitotic spindle assembly by activating of Plk1 and Aurora-A. Yang et al. used bioinformatics to investigate the high expression of AUNIP in oral squamous cell carcinoma (OSCC), which is associated with tumor microenvironment, human papillomavirus infection, and cell cycle. Inhibition of AUNIP can inhibit OSCC cells’ proliferation, resulting in the G0/G1 phase arrest of OSCC cells. AUNIP overexpression predicts bad prognosis in OSCC patients (Yang et al. ref. 2019). However, there are few reports on pan-cancer research in AUNIP.
Our work used bioinformatics aspect to discuss the expression, prognosis, clinicopathological features, mutation, tumor mutation load (TMB), microsatellite instability (MSI), immune characteristics, and drug sensitivity of AUNIP from the viewpoint of pan-cancer, and comprehensively analyzed the characteristics and mechanism of AUNIP, providing new ideas for tumor treatment and prognosis.
Materials and methods
Differential expression of AUNIP mRNA for cancers and normal samples
TIMER2 database studied immune cells infiltration in different tumors, as well as the differential expression of 33 kinds of tumors and normal tissues from TCGA database (Li et al. ref. 2020). Owing to the absence of normal samples in several tumors in TIMER database, we merged TCGA and GTEx to discuss the differential expression of AUNIP in 33 tumors and normal tissues.
Prognosis and diagnostic value of AUNIP
GEPIA2 database is an online platform in which survival significance maps of genes in pan-cancer can be obtained. According to the median value of AUNIP expression, AUNIP was divided into low-expression group and high-expression group and Kaplan–Meier was used to show prognostic differences of both groups (Tang et al. ref. 2019). In addition, a receiver-operating characteristic (ROC) curve estimated the diagnose value for AUNIP using pROC in R.
Clinicopathological features
We acquired the expression data and clinicopathological parameters of 33 cancers from the TCGA database and utilized Wilcoxon test to investigate its correlation with clinicopathology, including T stage, N stage, and pathological stage.
Immunohistochemical staining
Forty-four cases of liver cancer and paracancerous tissue were collected from the Department of Pathology of the Zhejiang Provincial People’s Hospital. The study was authorized by the ethics committee of Zhejiang Provincial People’s Hospital (batch number: QT2025083), and all patients received written informed consent before surgery. Patients with a pathologic diagnosis of hepatocellular carcinoma who had not received any preoperative chemotherapy or radiotherapy were included in the study. Patients with other diagnosed malignancies were not included in the study. The slices were cut into 4 um thick. The sections were dewaxed, hydrated, and repaired by high-pressure antigen. The endogenous catalase activity was inactivated by 3% H2O2 at room temperature for 10 min. The non-specific antigen was blocked by 10% sheep serum after rinsed with PBS at 37 ℃ for 10 min, and rabbit anti-AUNIP polyclonal antibody (bs-15019R, 1:200, Bioss Company) was added at 4 ℃ overnight. The next day, the secondary antibody (Goat anti-Rabbit IgG, PV-6000, Beijing Zhongshan Jinqiao Biotechnology Co., Ltd.) was added, and then developed color with DAB. Finally, the slices were observed in the microscope. The standard of expression strength is: 0 points without staining; light yellow is 1 point; light brown is 2 points; dark brown is 3 points. The scoring criteria for positive cells were: 0 points for ≤ 5%; 6% ~ 25% is 1 score; 26% ~ 50% is 2 points; 51% ~ 75% is 3 points; > 76% is 4 points. AUNIP expression is interpreted by the percentage of positive cells multiplied by the staining intensity. The degree of positive staining was defined: ≤ 7 is classified as low expression, and > 7 is classified as high expression.
Mutational analysis of AUNIP
cBioPortal studied the frequency of AUNIP gene change in various tumors (Cerami et al. ref. 2012).
Correlative analysis of AUNIP expression with TMB and MSI
TMB is the total number of genetic coding errors, base substitution, gene insertion, or deletion errors detected in somatic cells from millions of bases, and it can effectively evaluate tumor mutation and neoantigen load and is related to immunotherapy response (Zhang et al. ref. 2019; Chan et al. ref. 2019). MSI is due to mismatch repair gene defects. Tumors with MSl molecular characteristics increase tumor antigen load due to high-frequency gene mutation, inducing killer T lymphocyte infiltration and corresponding immunosuppressive molecule high expression, and respond well to corresponding immunotherapy (Dudley and Le ref. 2016). The interrelation of AUNIP expression with TMB and MSI in 33 tumors was discussed by Spearman analysis.
Association of AUNIP with immune cell infiltration and immune checkpoints
Cells and molecules of the tumor microenvironment (TME) can influence the efficiency of immunotherapy, so research on TME is of great significance in immunotherapy. Tumor immune cell infiltration is closely linked to tumor progression in the TME. The relationship of AUNIP with 23 types of immune cell infiltration in different cancers was applied using ssGSEA algorithm in R language. In addition, the stromal score, immune score, and estimate score for different tumors were investigated using ESTIMATE algorithm. Immunotherapy with immune checkpoint inhibitors has initiated a new era of tumor treatment, and finding predictable biomarkers is a necessary pathway for achieving precise tumor immunotherapy. At present, the eight commonest immune checkpoints are PD-1, PD-L1, CTLA-4, PDCD1LG2, TIGIT, HAVCR2, SIGLEC15, and LAG3. We discussed the association of AUNIP with immune checkpoints through Spearman analysis.
Correlative analysis between AUNIP expression and drug sensitivity
GSCALite is an integrated platform for genomic, pharmacogenomic, and immunogenomic gene set cancer analysis. The CTRP dataset from the GSCALite database (http://bioinfo.life.hust.edu.cn/web/GSCALite/) was employed to explore the relationship between gene expression and drug sensitivity (Liu et al. ref. 2022).
Gene set enrichment analysis of AUNIP
We conducted GSEA analysis between high AUNIP expression and low AUNIP expression according to the KEGG dataset in the MSigDB database.
Statistical analysis
Wilcoxon test was employed to study the difference expression between tumors and normal tissues. The relationship of AUNIP expression with clinicopathological features was applied by Chi-square test. We investigated the association of AUNIP with TME and immune checkpoints using Spearman correlation. P < 0.05 was considered statistically significant.
Results
Overexpression of AUNIP mRNA in various tumors
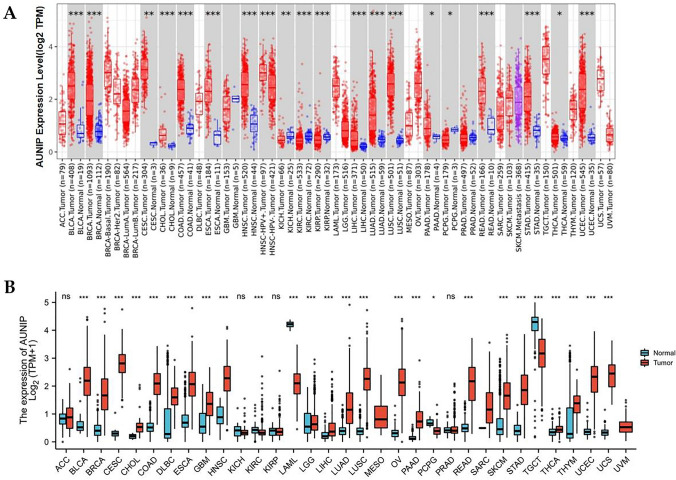
The TIMER database showed that compared with normal tissue, AUNIP had high expression in BLCA, BRCA, CESC, CHOL, COAD, ESCA, HNSC, LIHC, LUAD, LUSC, PAAD, READ, STAD, THCA, and UCEC, and low expression in KICH, KIRC, KIRP, and PCPG (Fig. 1A). Because of no normal tissues in some tumors in the TIMER database, TCGA was combined with GTEx to explore AUNIP expression in tumors and corresponding normal samples. We discovered that AUNIP expression in BLCA, BRCA, CESC, CHOL, COAD, DLBC, ESCA, GBM, HNSC, LGG, LIHC, LUAD, LUSC, OV, PAAD, READ, SKCM, STAD, THCA, THYM, UCEC, and UCS was higher than in normal tissues, but lower in KIRC, LAML, PCPG, and TGCT (Fig. 1B).
Prognosis and diagnostic value of AUNIP in pan-cancer
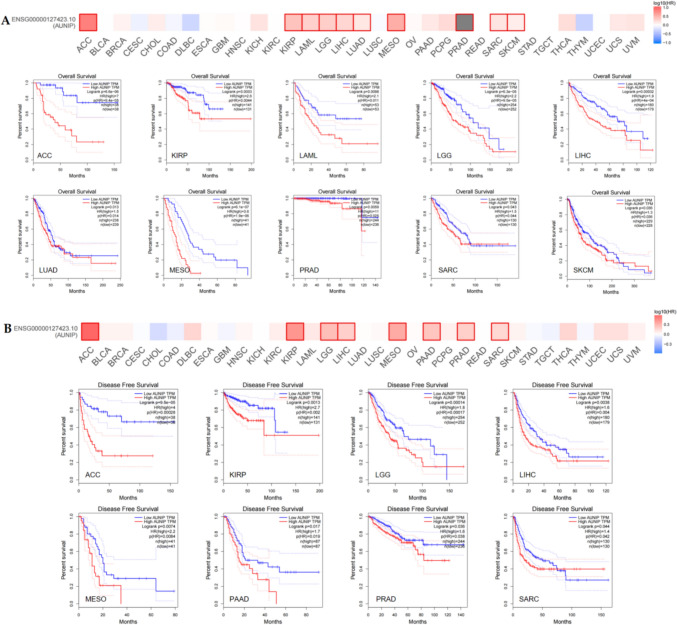
We analyzed the impact of AUNIP expression on survival in 33 tumors. Prognostic index included overall survival (OS) and disease-free survival (DFS). The findings demonstrated that the OS of low-expression AUNIP was better than that of high-expression AUNIP for ACC, KIRP, LAML, LGG, LIHC, LUAD, MESO, PRAD, SARC, and SKCM, (Fig. 2A). Regarding DFS, the DFS for low-expression AUNIP in ACC, KIRP, LGG, LIHC, MESO, PAAD, PRAD, and SARC was better than that of high-expression AUNIP (Fig. 2B). Among them, the OS and DFS with low AUNIP expression for ACC, KIRP, LGG, LIHC, MESO, PRAD, and SARC were higher than those with high AUNIP expression. There was no significant difference of AUNIP expression in KIRP and PRAD, compared to their corresponding normal tissues. Therefore, AUNIP was related to OS and DFS in ACC, LGG, LIHC, MESO, and SARC. Furthermore, we evaluated the diagnostic value of AUNIP in tumors. The area under the receiver-operating characteristic curve (AUC) > 0.7 is considered certain accuracy, and AUC > 0.9 is considered higher accuracy (Mishra et al. ref. 2023a). The results demonstrated that AUNIP has high diagnostic value in BLCA, BRCA, CESC, CHOL, COAD, ESCA, HNSC, LUAD, LUSC, PCPG, READ, STAD, UCEC, and certain diagnostic values for KICH, KIRC, LIHC, PAAD, and SARC (Figure S1).
AUNIP was associated with clinicopathological features in some tumors
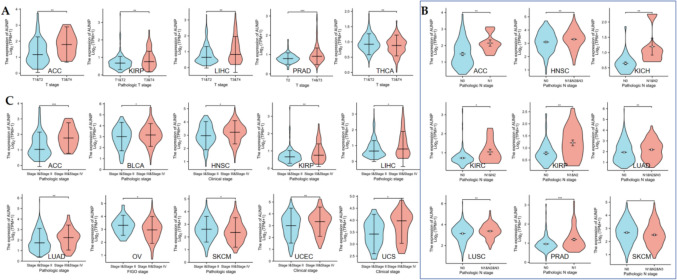
We downloaded clinicopathological data for 33 tumors. Regarding T staging, AUNIP expression was higher at T3 + T4 than at T1 + T2 in ACC, KIRP, LIHC, PRAD, and lower at T3+T4 in THCA (Fig. 3A). In ACC, HNSC, KICH, KIRC, KIRP, LUAD, LUSC, and PRAD, AUNIP was higher expressed in patients with N1&N2&N3 than in patients with N0, while in SKCM, AUNIP was higher expressed in patients with N0(Fig. 3B). In ACC, BLCA, HNSC, KIRP, LIHC, LUAD, UCEC, and UCS, AUNIP expression increased with the increase of pathological staging, while in OV and SKCM, AUNIP expression decreased with the increase of pathological staging (Fig. 3C).
Gene variation of AUNIP in pan-cancer
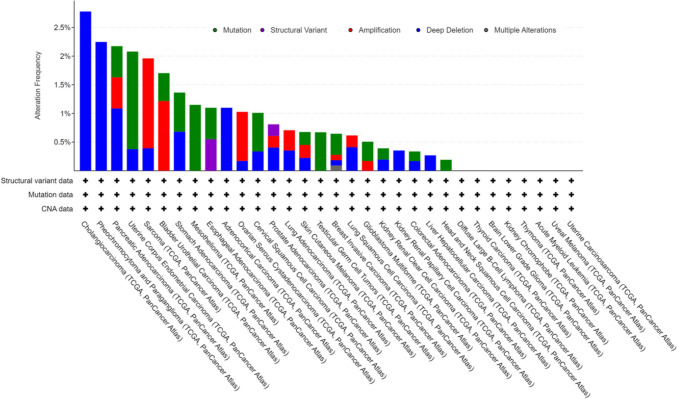
Genetic alterations are a form of epigenetics. Genetic alterations of AUNIP in different tumors appear in the form of mutation, structural variant, amplification, deep deletion, and multiple alterations. Among them, CHOL has the highest frequency of gene change, which is manifested as deep deletion, followed by PCPG, which is mainly manifested as deep deletion. The third genetic alteration is PAAD (mutation, amplification and deep deletion). No genetic alteration of AUNIP was observed in DLBC, THCA, LGG, KICH, THYM, LAML, UVM, and UCS (Fig. 4).
AUNIP expression was related to TMB and MSI in some cancers
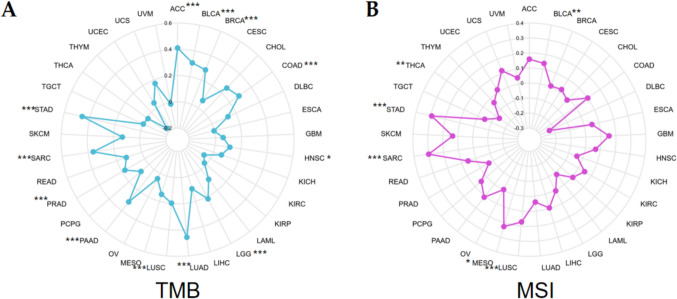
TMB and MSI are two common potential indicators for tumor immunotherapy response. The findings suggested that AUNIP expression had positive correlation with TMB in ACC, BLCA, BRCA, COAD, HNSC, LGG, LUAD, LUSC, PAAD, PRAD, SARC, and STAD, and the correlation coefficient with STAD is the highest (Fig. 5A). AUNIP had positive interrelation with MSI in BLCA, LUSC, MESO, SARC, and STAD, and negative association with MSI in THCA (Fig. 5B).
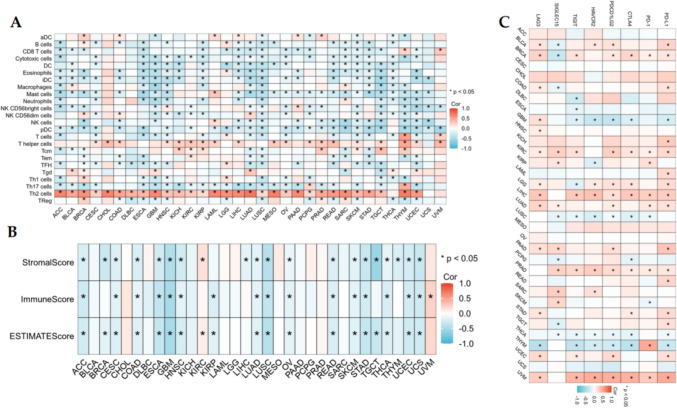
Association of AUNIP with immune cell infiltration and immune checkpoints in pan-cancer
We applied ssGSEA to investigate the interrelation between AUNIP and 23 kinds of immune cell infiltration in pan-cancer and found that AUNIP was positively linked with Th2 cells in 30 kinds of tumors, and the positive correlation coefficient was the highest. Among the remaining immune cells, AUNIP had a correlation with one or more immune cells in different cancers (Fig. 6A). For the ESTIMATE algorithm, in ACC, CESC, COAD, ESCA, GBM, HNSC, LUAD, LUSC, OV, READ, SKCM, STAD, THCA, UCEC, and UCS, AUNIP expression had negative association with stromalscore, immunescore, and estimatescore. In BLCA, CHOL, DLBC, KICH, LAML, LGG, MESO, PAAD, PCPG, PRAD, and SARC, AUNIP was not associated with stromalscore, immune score, and estimatescore, and AUNIP was related to stromalscore, immunescore, and estimate score in the remaining 7 tumors(Fig. 6B). Immune checkpoint is the target of immunotherapy at this stage, and our results showed that AUNIP was positively or negatively associated with an immune checkpoint in all tumors except ACC, CESC, CHOL, MESO, OV, and UCS (Fig. 6C).
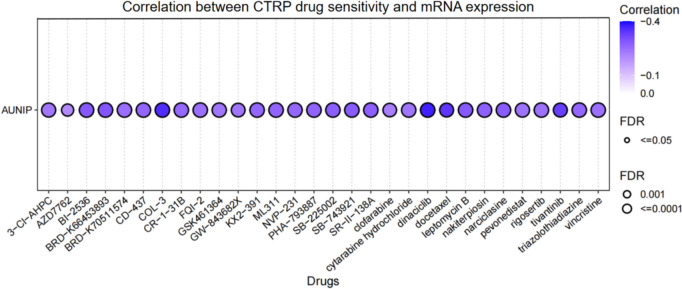
Drug sensitivity analysis
We found that AUNIP expression was negatively linked with 50% inhibitory concentration (IC50) values of 30 drugs based on the results of the CTRP dataset in GSCA. There was a strong negative correlation with IC50 of COL-3, dinaciclib, and docetaxel (Fig. 7). These findings indicated that AUNIP was significantly associated with different drug sensitivities in various tumor cell lines and may be a latent target for cancer therapy.
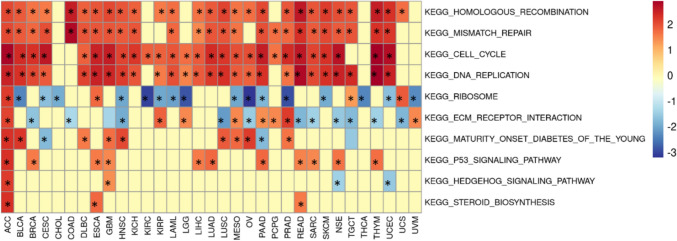
Gene functional enrichment of AUNIP in pan-cancer
The results of GSEA demonstrated that AUNIP was primarily participated in cell cycle, DNA replication, mismatch repair, and homologous recombination in most tumors (Fig. 8). In these tumors, AUNIP was mainly involved in the development of tumors through the above pathways.
Overexpression AUNIP was correlated with clinical information in LIHC and an independent prognostic gene for LIHC
IHC analysis demonstrated that AUNIP was overexpressed, compared to normal liver tissues (Fig. 9A,B). AUNIP expression was linked with histologic grade, not correlated with age, gender, and pathologic stage (Table 1). Kaplan–Meier analysis suggested that the patients with high-expression group had worse prognosis (Fig. 9C).
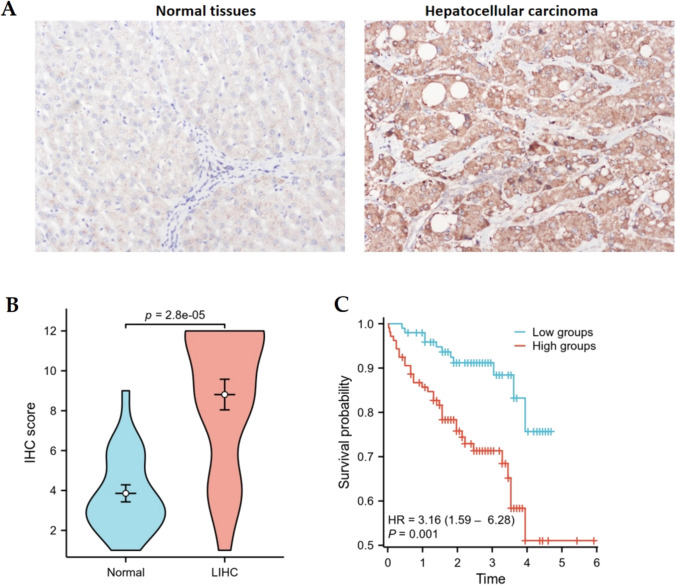
Table 1: The correlation of AUNIP expression with clinical information in LIHC
| Characteristics | AUNIP | χ2 | P | |
|---|---|---|---|---|
| Low expression | High expression | |||
| Age | 0.25 | 0.72 | ||
| ≤ 55 | 14 | 20 | ||
| > 55 | 5 | 5 | ||
| Gender | 0.24 | 0.63 | ||
| Female | 10 | 15 | ||
| Male | 9 | 10 | ||
| Pathologic stage | 0.09 | 0.76 | ||
| Stage I + Stage II | 9 | 13 | ||
| Stage III + Stage IV | 10 | 12 | ||
| Histologic grade | 4.54 | 0.03 | ||
| G1 + G2 | 13 | 9 | ||
| G3 | 6 | 16 | ||
Discussion
The global mortality burden is predominantly attributed to malignant neoplasms resulting from dysregulated cellular proliferation. Recent decades have witnessed remarkable progress in early disease detection methodologies, encompassing the conventional approaches including radiotherapy, surgical procedures, tailored therapeutic regimens, and chemotherapeutic interventions (Mishra et al. ref. 2023a, ref. 2023b, ref. 2023c). However, cancer remains a major threat to human health. Therefore, it is important to find an effective biomarker to predict the development and prognosis of cancer. DNA double-strand breakage damage is the most severe form of damage, if not repaired in time or abnormal repair occurs, it will lead to a series of changes in the cell genome, directly lead to deactivation of tumor suppressor genes or overexpression of oncogenes, and eventually lead to cell cancer (Burma et al. ref. 2006). The most critical factor affecting the selection of DNA double-strand break repair pathways is the state of the cut end of DNA, and AUNIP is a key factor regulating the state of DNA cleavage ends. AUNIP, a binding protein of protein kinase A and Ninein proteins, also known as AIBP, is a structurally specific DNA-binding protein that is localized to the 135 open-reading framework of chromosome 1. According to reports, it is highly expressed in various tumors (Ma et al. ref. 2020). AUNIP is highly expressed in astrocytoma and other brain tumors, suggesting that AUNIP may play a role as oncogenic genes in the development of brain tumors (Lieu et al. ref. 2010).
Our study analyzed the expression, clinical significance, prognosis, mutation, and immunity of AUNIP from the perspective of pan-cancer using a multi-omics system. It was found that AUNIP expression was increased significantly in most tumors compared to normal tissues, suggesting that AUNIP may be a key gene in cancer development. We conducted IHC analysis to confirm the higher expression of AUNIP in LIHC, which was consistent with TCGA database. In addition, AUNIP with high expression in ACC, LGG, LIHC, MESO, and SARC had poorer OS and DFS than those of AUNIP with low expression, suggesting that the high expression of AUNIP in some tumors influenced patients’ prognosis. Furthermore, AUNIP expression was related to the T stage, N stage, and clinicopathological stage in some cancers, indicating that AUNIP may be a promising valuable diagnostic and prognostic marker in multiple tumors. IHC analysis indicated that AUNIP was linked with histologic grade in LIHC. Moreover, AUNIP expression was an independent prognostic index by univariate and multivariate regression in LIHC. We used cBioportal to study the frequency of AUNIP gene alteration in tumors. In CHOL, the frequency of genetic changes was the highest, with all deep deletions, followed by PCPG, with all deep deletions.
Immune cell infiltration is closely linked to cancer progression (Marcas and Walzer ref. 2018). Recent studies have suggested that tumor progression is caused by an imbalance between the tumor’s immune state and the host’s immune response (Nabbi et al. ref. 2019). We studied the relationship between AUNIP and immunocyte infiltration and observed that AUNIP expression was positively related to Th2 for most tumors, indicating that with the increase of AUNIP expression, Th2 concentration was up-regulated. Th2 cells are not conducive to the anti-tumor effect of cellular immunity. Th1/Th2 drift will protect the tumor from immune surveillance and immune attack, thus promoting the development and progression of tumors (Sharma et al. ref. 2007). Furthermore, we applied the ESTIMATE algorithm to discuss the correlation between AUNIP and stromalscore, immunescore, and estimatescore in different tumors. In most tumors, AUNIP was negatively correlated with these three scores. The application in immune checkpoint inhibitors has elevated immunotherapy to a new level. Immunotherapy has been considered as an effective therapy for various advanced and invasive cancers (Morse et al. ref. 2005; Zhou and Zhong ref. 2004). At present, immunotherapy has been applied to a variety of tumors. Common immune checkpoints include PD-1, PD-L1, CTLA-4, PDCD1LG2, TIGIT, HAVCR2, SIGLEC15, and LAG3. In some tumors, AUNIP expression was positively related to immune checkpoint expression, suggesting that these patients with high AUNIP expression may benefit from immunotherapy. TMB and MSI are effective markers to predict the effect of immunotherapy. MSI-H’s tumor gene repair system is abnormal, and there may be more gene mutations, which are easily recognized by T cells and may respond better to immunotherapy (Bateman ref. 2021). The higher the TMB, the greater the probability that neoantigens expressed by the tumor will be identified by the immune system. Therefore, tumors with high TMB are more sensitive to immune therapy (Liu et al. ref. 2019). Our study demonstrated that AUNIP had positive association with TMB and MSI in BLCA, SARC, and STAD, and patients with high AUNIP expression in these three types of tumors were more susceptible to immunotherapy.
The results of gene enrichment analysis showed that AUNIP caused tumors progression through the cell cycle, DNA replication, mismatch repair, and homologous recombination in most tumors. This is consistent with the literature reports (Lou et al. ref. 2017) In addition, we also performed a correlative analysis between AUNIP and drug sensitivity, and we used a public database to predict several candidate targeted small-molecule drugs. We found that AUNIP was negatively related to IC50 values of 30 drugs, indicating that these drugs stop the progression of the tumor. This provides a novel insight into expanding the therapeutic selection of these targeted small-molecule drugs and developing new drugs specifically targeting AUNIP.
We performed IHC analysis to discuss AUNIP expression in LIHC and the findings demonstrated that AUNIP expression was up-regulated in LIHC. The patients with high-expression group had unfavorable prognosis. These findings suggested that overexpression AUNIP was correlated with the progress of LIHC development and prognosis.
In our work, the expression, prognosis, and characteristics of AUNIP were elucidated by pan-cancer analysis. However, there are some shortcomings in this study. The characteristics of AUNIP were analyzed through bioinformatics and only conducted IHC to verify the overexpression of AUNIP in LIHC. However, there was no biological experiment to verify it. Therefore, in the following studies, it needs more experiment to further validate the mechanism of effect of AUNIP in inducing tumors.
Supplementary Materials
References
- AC Bateman. DNA mismatch repair proteins: scientific update and practical guide[J]. J Clin Pathol, 2021. [DOI | PubMed]
- S Burma, BP Chen, DJ Chen. Role of non-homologous and joining (NHEJ) in maintaining genomic integrity. DNA Repair (Amst), 2006. [DOI | PubMed]
- E Cerami, J Gao, U Dogrusoz, BE Gross, SO Sumer, BA Aksoy, A Jacobsen, CJ Byrne, ML Heuer, E Larsson. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov, 2012. [DOI | PubMed]
- TA Chan, M Yarchoan, E Jaffee. Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic[J]. Ann Oncol, 2019. [DOI | PubMed]
- Chou CH, Loh JK, Yang MC, Lin CC, Hong MC, Cho CL, et al. AIBp regulates mitotic entry and mitotic spindle assembly by controlling activation of both Aurora-A and Plk1
- JC Dudley, DT Le. Microsatellite instability as a biomarker for PD-1 blockade[J]. Clin Cancer Res, 2016. [DOI | PubMed]
- Li T, Fu J, Zeng Z, Cohen D, Li J, Chen Q, Li B and Shirley Liu X (2020) TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res
- A Lieu, T Cheng, C Chou. Functional characterization of AIBp, a novel Aurora-A binding protein in centrosome structure and spindle formation. Int J Oncol, 2010. [PubMed]
- L Liu, X Bai, J Wang. Combination of TMB and CNA stratifies prognostic and predictive responses to immunotherapy across metastatic cancer [J]. Clin Cancer Res, 2019. [DOI | PubMed]
- Liu C-J, Hu F-F, Xie G-Y, Miao Y_R, Li X-W, Zeng Y, Guo A-Y (2022) GSCA: an integrated platform for gene set cancer analysis at genomic, pharmacogenomic, and immunogenomic levels. Briefings Bioinform bbac558
- J Lou, H Chen, J Han, H He, MSY Huen, X-H Feng, T Liu, J Huang. AUNIP/C1orf135 directs DNA double-strand breaks towards the homologous recombination repair pathway. Nat Commun, 2017. [DOI | PubMed]
- C Ma, W Kang, L Yu. AUNIP expression is correlated with immune infiltration and is a candidate diagnostic and prognostic biomarker for hepatocellular carcinoma and lung adenocarcinoma. Front Oncol, 2020. [DOI | PubMed]
- A Marcas, T Walzer. An immunosuppressive pathway for tumor progression [J]. Nat Med, 2018. [DOI | PubMed]
- D Mishra, A Mishra, SN Rai, E Vamanu, MP Singh. Identification of prognostic biomarkers for suppressing tumorigenesis and metastasis of hepatocellular carcinoma through transcriptome analysis. Diagnostics, 2023. [DOI | PubMed]
- D Mishra, A Mishra, SN Rai, E Vamanu, MP Singh. Demystifying the role of prognostic biomarkers in breast cancer through integrated transcriptome and pathway enrichment analyses. Diagnostics, 2023. [DOI | PubMed]
- D Mishra, A Mishra, S Rai, S Singh, E Vamanu, MP Singh. In Silico Insight to Identify Potential Inhibitors of BUB1B from Mushroom Bioactive Compounds to Prevent Breast Cancer Metastasis. Front. Biosci. Landmark Ed, 2023. [DOI | PubMed]
- MA Morse, J Garst, T Osada. A phase I study of dexosome immunotherapy in patients with advanced non-small cell lung cancer [J]. J Transl Med, 2005. [DOI | PubMed]
- A Nabbi, N Jager, S Sudhaman. Landscape of infiltrating immune repertoire in pediatric solid tumors[J]. Proc Natl Acad Sci, 2019. [PubMed]
- A Sharma, M Rajappa, A Saxena. Cytokine profile in Indian women with cervical intraepithelial neoplasia and cancer cervix[J]. Int J Gynecol Cancer, 2007. [DOI]
- Z Tang. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res, 2019. [DOI | PubMed]
- L Yang, S Yang, X Yuan. Analyzing pan-cancer DNA methylation patterns via clustering[J]. J Xidian Univ, 2018
- Z Yang, X Liang, Y Fu. Identification of AUNIP as a candidate diagnostic and prognostic biomarker for oral squamous cell carcinoma. EBioMedicine, 2019. [DOI | PubMed]
- K Zhang, H Wang. Cancer genome atlas pan-cancer analysis project[J]. Zhongguo Fei Ai Za Zhi, 2015. [PubMed]
- C Zhang, Z Li, F Qi. Exploration of the relationships between tumor mutation burden with immune infiltrates in clear cell renal cell carcinoma[J]. Ann Transl Med, 2019. [DOI | PubMed]
- J Zhou, Y Zhong. Breast cancer immunotherapy [J]. Cell Mol Immunol, 2004. [PubMed]
