Identification and Expression Analysis of the MLO Gene Family Under Salt Stress in Cotton (Gossypium hirsutum L.)
Abstract
MLO (Mildew Resistance Locus O) genes encode seven-transmembrane proteins that function as critical regulators of powdery mildew resistance and abiotic stress responses. Despite their established importance, the MLO gene family in Gossypium hirsutum L. has not been systematically investigated under salt stress conditions. Here, we performed genome-wide identification of 46 GhMLO members using Hidden Markov Model and BLAST searches based on the latest cotton genome assembly. Phylogenetic analysis classified these genes into four distinct subfamilies. Transmembrane topology and conserved domain analyses revealed that all GhMLO proteins contain typical MLO domains and transmembrane structures, maintaining high structural similarity with dicotyledonous model plants. Synteny analysis demonstrated that the expansion of the GhMLO family was primarily driven by segmental and tandem duplications. Integration of transcriptomic data from the COTTONOMICS database revealed tissue-specific expression patterns, with higher transcript abundance in receptacles, stems, and roots, but lower levels in stamens and petals. Salt, drought, and cold stress treatments induced upregulation of GhMLO family members, with most genes showing increased expression over time. RT-qPCR analysis validated that five candidate GhMLO genes were significantly upregulated under salt stress. In summary, this study provides a comprehensive genome-wide characterization of the GhMLO gene family, elucidating their phylogenetic relationships and expression dynamics, which establishes a theoretical basis for identifying key regulatory genes involved in abiotic stress responses and offers novel genetic resources for improving stress tolerance in cotton molecular breeding.
Article type: Research Article
Keywords: transcriptome analysis, abiotic stress, gene expression
Affiliations: Institute of Cotton, Hebei Academy of Agricultural and Forestry Sciences, Key Laboratory of Biology and Genetic Improvement of Cotton in Huanghuaihai Semiarid Area, Ministry of Agriculture and Rural Affairs, Hebei Key Laboratory of Cotton Bio-Breeding and Cultivation Physiology, Shijiazhuang 050051, China; fch19942024@163.com (C.-H.F.); zjb210@126.com (J.Z.); spp2016@126.com (L.L.); limengzhe0131@126.com (M.L.); jmm18270464849@163.com (M.J.); Genetics Laboratory, College of Life Science, Hebei University, Baoding 071002, China
License: © 2026 by the authors. CC BY 4.0 Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Article links: DOI: 10.3390/life16030476 | PubMed: 41900994 | PMC: PMC13028013
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (8.0 MB)
1. Introduction
Gossypium hirsutum, the world’s primary source of natural fiber, holds dual strategic value for ecological restoration and rural revitalization in the saline-alkali regions of northern and northwestern China [ref. 1,ref. 2]. Salt stress significantly suppresses cotton growth, causing stunted development, leaf chlorosis, impaired root systems, and reductions in both yield and fiber quality [ref. 3,ref. 4]. Therefore, the discovery, identification, and utilization of salt-tolerant genes to enhance cotton’s salinity tolerance and improve the comprehensive utilization efficiency of saline-alkali lands holds profound implications [ref. 5,ref. 6].
A gene family comprises a set of homologous genes derived from a common ancestral gene through duplication and divergence, often encoding functionally similar or synergistic proteins that constitute a functional module within an organism [ref. 7,ref. 8,ref. 9]. The MLO (Mildew Resistance Locus O) gene family encodes specialized seven-transmembrane proteins initially identified as powdery mildew susceptibility factors in barley [ref. 10]. MLO proteins mediate broad spectrum resistance against fungal, bacterial, and viral pathogens by modulating apoplastic Ca2+ channel activity and ROS homeostasis [ref. 11]. Additionally, they participate in signal transduction pathways under abiotic stresses including salinity, drought, cold, and heavy metal toxicity and maintain cell wall integrity [ref. 12,ref. 13]. In Arabidopsis, mutations in AtMLO2/6/12 confer complete resistance to powdery mildew, whereas loss of function mutants of OsMLO12 exhibit male gametophyte sterility, with impaired pollen hydration and germination both in vitro and in vivo, preventing generation of homozygous progeny [ref. 14,ref. 15,ref. 16]. Although initially recognized for their role in regulating powdery mildew resistance, recent studies have demonstrated that MLO genes also play significant roles in abiotic stresses such as salt, drought, extreme temperature stress, etc. [ref. 14,ref. 17,ref. 18]. The function of MLO in abiotic stress can be summarized as “negatively regulating osmotic/oxidative stress tolerance and participating in stress-immune cross-signaling [ref. 19]. At present, the MLO protein family has been defined and explored in more than 40 plant species, including Arabidopsis thaliana, Malus pumila, Citrullus lanatus and Gosspium [ref. 20,ref. 21,ref. 22]. However, the role of MLO genes in responding to salt stress in cotton lacks systematic investigation.
A bHLH family member, designated GhbHLH (GenBank: KJ605396), from a normalized cDNA library of G. hirsutum cultivar JiMian 228. This gene encodes a 199 amino acid protein with a 600 bp open reading frame, shares 52% homology with Arabidopsis AtbHLH149, and contains a conserved basic helix-loop-helix (bHLH) domain. Expression of GhbHLH is induced by abiotic stresses such as high salinity and drought, showing maximal upregulation in roots that peaks at 2 h post treatment and remains significantly higher than in stems and leaves. Agrobacterium-mediated transformation generated GhbHLH149-like overexpression lines, and comparative transcriptomic analysis against recipient control JIN668 revealed significant upregulation of GhMLO43 in transgenic lines.
Previously, we generated GhbHLH149-like overexpressing cotton lines via Agrobacterium-mediated transformation. Comparative transcriptome analysis against the recipient control JIN668 revealed significant upregulation of GhMLO43 in transgenic lines, a finding further corroborated by RT-qPCR demonstrating consistent upregulation at multiple time points under salt stress. In this study, based on the latest genomic data, we identified 46 GhMLO members through a combination of hidden Markov models and iterative BLAST searches, clustering them into four subfamilies. To characterize the structural features of GhMLO proteins, transmembrane topology and conserved domain analyses were performed, and the results indicated that all GhMLO members harbor the conserved MLO domain as well as the typical transmembrane architectures that are characteristic of the MLO protein family. Comparative structural analysis revealed a high degree of similarity between GhMLO proteins and their homologs from dicotyledonous model plant species, suggesting evolutionary conservation of this gene family across dicot lineages. Collinearity assessment was also conducted to explore the evolutionary mechanisms underlying the expansion of the GhMLO gene family in Gossypium hirsutum, and the findings demonstrated that segmental duplication and tandem duplication are the primary driving forces responsible for the family’s expansion. Transcriptomic datasets obtained from the COTTONOMICS database were integrated and analyzed, and the results showed that GhMLO genes display obvious tissue-specific expression patterns: notably high expression levels were detected in vegetative tissues including leaves, stems, and roots, whereas expression was relatively weak in reproductive tissues such as stamens and petals. Additionally, abiotic stress treatments (salt, drought, and low temperature) were applied to explore the stress-responsive characteristics of GhMLO genes, and the results showed that numerous GhMLO family members were induced and up-regulated under these stress conditions, with the majority of these genes showing a gradual increase in expression level as the stress treatment duration prolonged. RT-qPCR results confirmed that all five selected candidate GhMLO genes exhibited significant up-regulation in response to salt stress. Collectively, the systematic identification and comprehensive analysis of the GhMLO gene family in G. hirsutum not only offer novel perspectives and potential targets for the screening of candidate GhMLO genes but also establish a solid basis for further deciphering their biological roles in cotton growth and development processes, as well as their regulatory mechanisms in mediating responses to abiotic stresses. In conclusion, this study provides crucial theoretical support for the identification of key regulatory genes involved in cotton abiotic stress adaptation and developmental regulation, which is expected to provide valuable guidance for subsequent research on cotton stress resistance breeding and genetic improvement.
2. Results
2.1. Identification, Phylogenetic Analysis and Sequence Alignment of the GhMLOs Gene Family
To systematically investigate the function of GhMLO gene family in G. hirsutum, we performed genome-wide identification and curation of family members. Following elimination of redundancies and integration of NCBI chromosome mapping information, we identified a total of 46 GhMLO genes (Supplementary Table S1). Physicochemical characterization revealed that the encoded proteins ranged from 85 to 584 amino acids (average 484.7 aa), with 85% exhibiting molecular weights exceeding 50 kDa. Predicted isoelectric points (pI) varied from 6.30 to 9.86, with three proteins classified as acidic and 44 as basic. 22 members displayed instability indices below 40, indicating high stability. 21 proteins were predicted to be hydrophilic, whereas the remainder were hydrophobic. Subcellular localization prediction revealed plasma membrane localization for 45 GhMLO proteins.
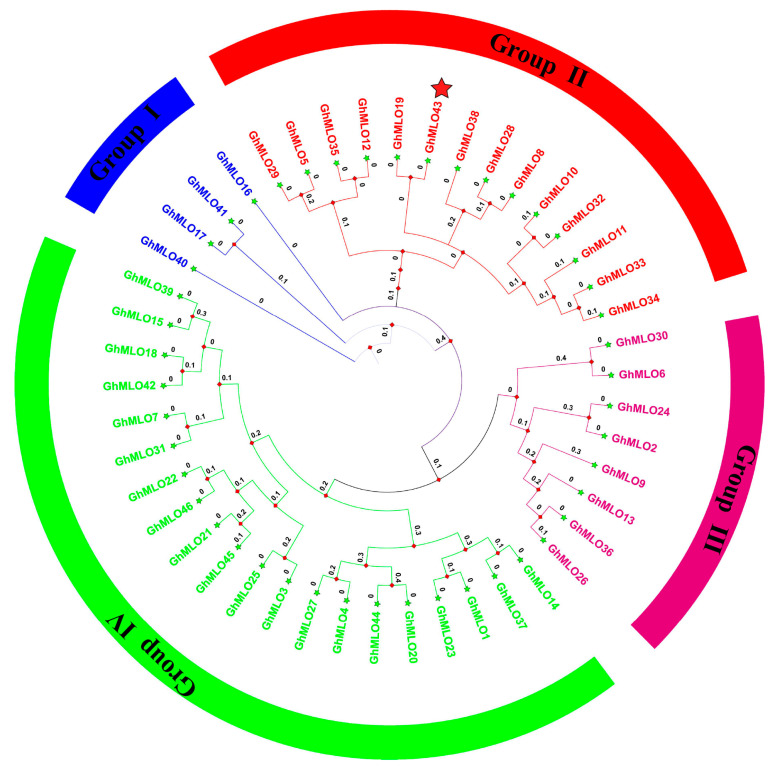
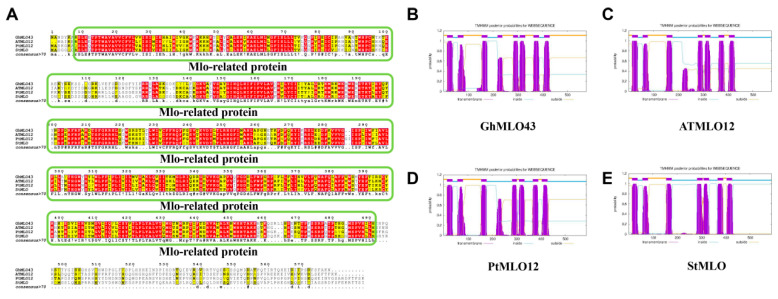
Based on 46 G. hirsutum GhMLO amino acid sequences, a phylogenetic tree was constructed using the MEGA-X maximum likelihood method (ML). Phylogenetic analysis divided the GhMLO family into four distinct branches (Figure 1), each with different numbers of members, but sequences within the same branch were highly homologous (Figure 1). To further verify conservation, CLUSTALX multi-sequence alignment was performed on MLO proteins from four representative species (G. hirsutum, Arabidopsis thaliana, Populus trichocarpa, and Solanum tuberosum), and the results showed that all MLO proteins contained typical MLO conserved domains and seven transmembrane helical regions (Figure 2A–E). To sum up, the GhMLO gene family is highly conserved during evolution, and its structural and functional characteristics are common between G. hirsutum and other species.
2.2. Chromosomal Distribution and Collinearity Analysis of the GhMLOs Gene Family
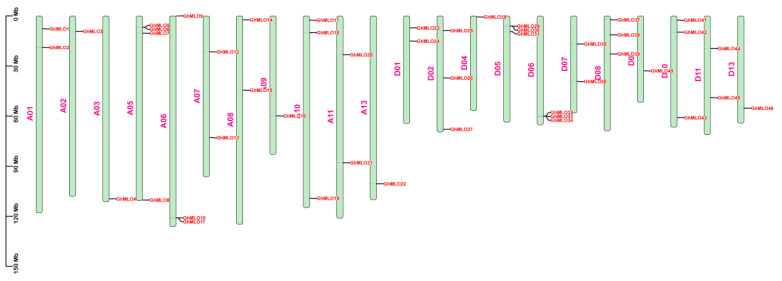
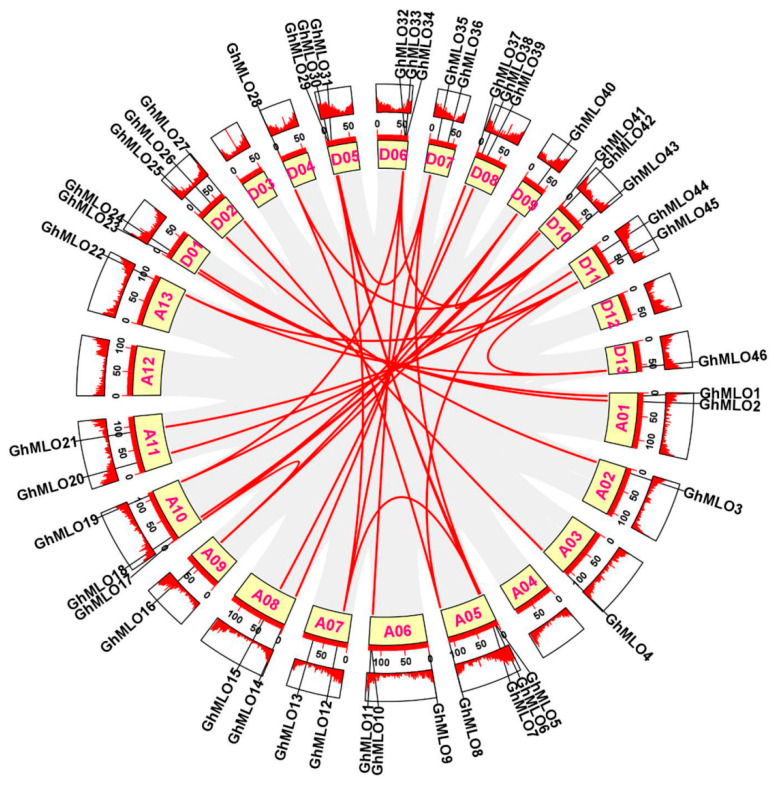
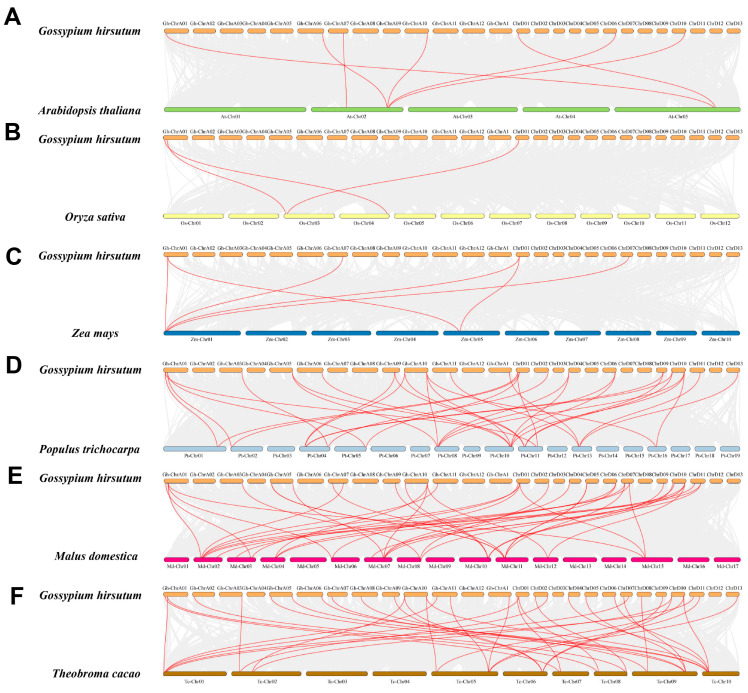
Chromosome localization results showed that 46 GhMLO genes were distributed across 22 chromosomes of G. hirsutum (Figure 3). Among them, chromosome A05 carries the largest number of members (4), while the rest of the chromosomes contain 1–3 genes. 4 GhMLO tandem repeat clusters (GhMLO5/6, GhMLO10/11, GhMLO29/30 and GhMLO32/33/34) were detected on chromosomes A05, A06, D05 and D06. Intraspecific collinearity analysis further identified 33 pairs of GhMLO fragment duplication events (Figure 4). To analyze cross-species evolutionary relationships, collinearity comparisons of G. hirsutum were conducted with Arabidopsis thaliana, rice, corn, G. hirsutum, apple, and cocoa, resulting in 7, 3, 6, 35, 36, and 37 homologous gene pairs (Figure 5A–F). The results suggest that the GhMLO family has a closer genetic relationship with woody plants in terms of evolution.
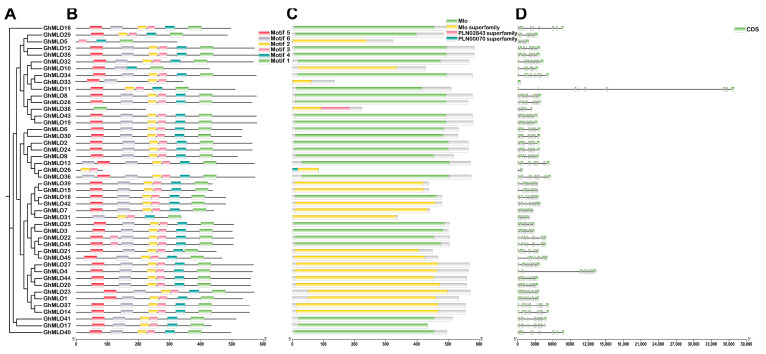
2.3. Analysis of the Gene Structure and Protein Conserved Motifs of the GhMLOs Gene Family
To clarify the evolutionary relationships of the GhMLO gene family, we integrated phylogenetic trees, conserved motifs, protein domains and gene structures for multi-dimensional analysis (Figure 6A–D). Six conserved motifs were identified from 46 GhMLO protein sequences. The number and distribution of motifs varied, but the family as a whole was highly conserved: all members contained at least two motifs, with Motif-3 present in all 46 proteins, forming a characteristic marker of the MLO domain. Domain analysis revealed that all GhMLO proteins contained the MLO core domain. Notably, GhMLO38 and GhMLO26 also carry two additional domains, PLN02843 and PLN00070. In terms of genetic structure, except for GhMLO33 which is a single exon gene, the rest of the members have introns, with the number of exons ranging from 1 to 16.
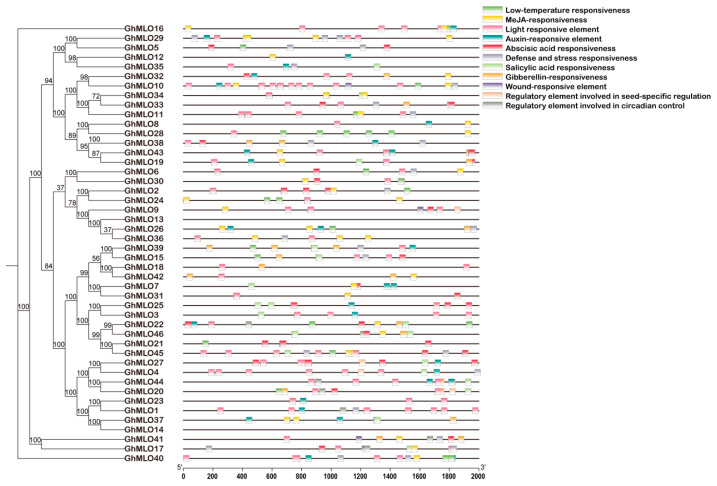
2.4. Analysis of Cis-Acting Elements in the GhMLOs Gene Family Promoter
To analyze the expression regulatory characteristics of the GhMLOs gene family, the cis-acting elements in its promoter region were systematically predicted using the PlantCARE database. The results showed that all GhMLOs promoters contained photoresponsive elements, and most members carried hormone regulatory elements in response to methyl jasmonate (MeJA), abatinic acid (ABA), gibberellin (GA), salicylic acid (SA), and auxin (IAA). In addition, elements related to the low-temperature response, circadian rhythm, defense and stress response were also widely present (Figure 7). There are significant differences in the types and quantities of cis-type elements among different members, suggesting that the GhMLOs family has diverse and specific regulatory functions in G. hirsutum growth and development and stress response.
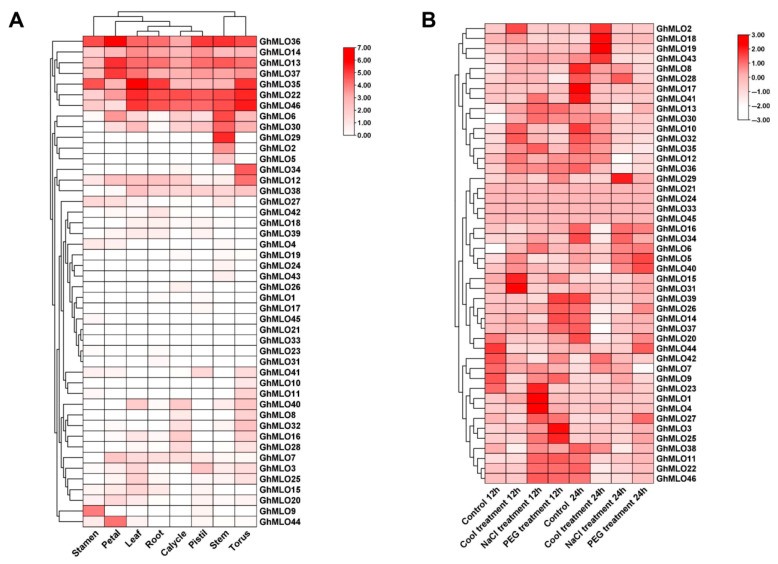
2.5. Expression Patterns of the GhMLOs Gene Family
To further clarify the role of the GhMLOs gene family in cotton growth and development and abiotic stress, this study obtained expression profiles of eight representative tissues and three abiotic stresses (cold, simulated drought, salt) from the COTTONOMICS transcriptome database (Figure 8A,B). The tissue expression profiles showed that the vast majority of GhMLOs were highly expressed in receptacles, such as GhMLO22, GhMLO35 and GhMLO46; GhMLO36 was maintained at high levels in all eight tissues, suggesting that it may play an important regulatory role in the overall growth and development of cotton. Stress expression profiles indicated that GhMLO1, GhMLO4, GhMLO23, etc., were significantly upregulated under salt stress, while GhMLO15, GhMLO31, GhMLO43, etc., were enhanced under cold stress, revealing that related members of the gene family were respectively involved in the response processes of salt stress and cold stress.
2.6. Expression of GhMLOs Under Salt Stress
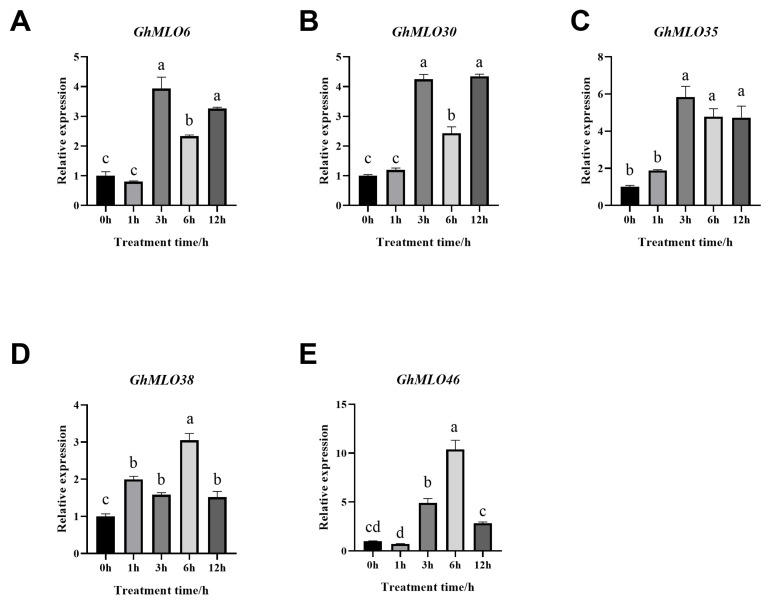
The GhMLO gene family encodes transmembrane proteins whose expression is regulated by ABA, ROS, and temperature signaling networks, making it an important node for studying stress-immune cross-responses. The expression levels of five GhMLO genes at different time points under normal conditions and salt stress were detected by RT-qPCR to verify the role of these genes under salt stress. The results showed that all five GhMLO genes were upregulated under salt stress treatment, but the response times were different (Figure 9A–E). The results showed that the GhMLO gene families in G. hirsutum responded differently to salt stress treatment.
3. Discussion
A total of 46 GhMLO genes were identified in this study. Phylogenetic and motif analyses revealed that all members contained seven transmembrane domains and calmodulin binding domains (CaMBD) at the C-end, showing typical basic isoelectric point characteristics. Subcellular localization predictions indicated that they were mainly located on the plasma membrane (Figure 6B,C, Supplementary Table S1). The CaMBD conserved domain gives GhMLO protein the ability to interact dynamically with calmodulin (CaM), thereby mediating calcium signal transduction in response to external stress [ref. 23,ref. 24]. It is notable that the GhMLO gene family typically contains multiple introns, and given that intron-free genes usually have higher transcriptional efficiency, it is speculated that the presence of introns may give GhMLO transcriptional dynamic characteristics of delayed regulation or fine regulation under adverse conditions (Figure 6D) [ref. 25]. GhMLO genes in different plants are highly homologous, and comparisons of their amino acid sequences suggest that they are evolutionarily conserved. The mechanisms of gene family growth and genome evolution are largely dependent on gene duplication events, including whole genome duplication (WGD), segmental duplication (SD), and tandem duplications (TD), etc. [ref. 26]. In this study, 46 GhMLO genes were distributed across 22 chromosomes, including 33 SD gene pairs and 4 TD gene clusters (Figure 3 and Figure 4). Gene pairs of gene replication events may have similar functions and expression patterns. For example, the highly homologous gene pairs GhMLO5/6 and GhMLO10/11 formed through series replication have similar transcriptional levels under different stress conditions and in different tissues, respectively (Figure 8A,B).
Although the functions of the MLO protein family have been identified in many species, there are certain differences in the functions among different species [ref. 17]. The MLO protein inhibits the deposition of antibacterial substances such as callose by directly interacting with SNARE protein ROR2 and blocking the directional transport of disease-resistant vesicles to the infection site [ref. 27,ref. 28]. At the same time, the interaction also downregulates the transcriptional levels of the glycoside hydrolase PEN2 and the ABC transporter PEN3, weakening the plant’s ability to degrade and excrete mycotoxins [ref. 29]. CML45 (calmodulin protein), ZTP29 (zinc transporter), LSD1 (zinc finger protein), and MYB in Goji berries may specifically interact with TC-rich repeats to regulate, thereby coordinating the expression of the LbMLO2 gene to enhance goji berries’ resistance to powdery mildew [ref. 30,ref. 31,ref. 32].
It participates in abiotic stress responses in a variety of species such as melons, rice and peppers. The analysis in this study showed that the vast majority of GhMLO genes were significantly upregulated under simulated conditions of drought, low temperature and salt stress (Figure 8B). We further conducted a systematic analysis of the GhMLO gene family promoter sequences and found that they were enriched with a variety of cation-acting elements associated with abiotic and biological stresses, such as ABRE and ERE (Figure 7) [ref. 33]. At 6 h of salt stress and 12 h of drought stress in bitter gourd, the relative expression of McMLO11 in bitter gourd leaves reached its peak, 56.7 and 9.07 times that of the control, respectively, suggesting that the McMLO11 gene can respond positively to drought and salt induction [ref. 34]. We treated the HX1 strain with 200 mM salt stress and found that all 5 GhMLO genes were upregulated under salt stress and may play an important role in resisting salt stress (Figure 9). bHLH (basic helix-loop-helix) transcription factor is one of the core nodes in the plant salt tolerance regulatory network, and bHLH can enhance plant salt tolerance by eliminating reactive oxygen species (ROS), maintaining ionic homeostasis, and hormone signaling crossover. The GhbHLH149-like gene in G. hirsutum plays a significant role in salt tolerance stress, and overexpression can enhance salt tolerance in cotton. The expression of GhMLO43 in overexpressed GhbHLH149-like cotton plants was significantly higher than that in wild-type cotton, and the expression of GhMLO43 was also significantly upregulated under salt stress, indicating that GhMLO43 functions under salt stress and may be regulated by GhbHLH149-like expression (Figure S1A,B).
Soil salinization is a major limiting factor for global agricultural production. The core mechanism by which plants respond to salt stress depends on the SOS pathway [ref. 3]. Salt stress induces an increase in cytoplasmic Ca2+, the calcium sensor CBL4/CBL10 binds and activates the kinase CIPK24 (SOS2), and then phosphorylates the Na+/H+ reverse transporter SOS1 for Na+ efflorescence or vacuole fragmentation [ref. 35,ref. 36]. Enhanced salt tolerance. MLO (such as AtMLO1/5/9/15) may be a type of calcium channel specific to plants that mediates extracellular Ca2+ influx and serves as a core hub for Ca2+ signaling in plant polar growth [ref. 37]. However, the mechanism by which the GhMLO gene regulates plant salt tolerance through the Ca2+ signaling pathway is still understudied. It can be speculated that clarifying the function of the GhMLO gene in the salt tolerance response of G. hirsutum may be of great significance. This study systematically identified members of the GhMLO gene family using bioinformatics methods and screened out multiple candidate proteins involved in the salt stress response. Compared with previous studies [ref. 22], we identified more GhMLO genes by utilizing updated genome data, and performed comprehensive analyses on their gene structure, protein conserved motifs, cis-acting elements, as well as the expression levels of some members at different response time after salt stress treatment. In summary, the GhbHLH149-like—GhMLO43 signaling module may play a key role in salt tolerance of G. hirsutum by regulating calcium ion signal transduction. GhMLO43 is currently the only member with identified regulatory context, and future studies will explore the functional roles of other candidates.
4. Materials and Methods
4.1. Cotton Test Materials
The cotton materials used in this study were the JIN668 and HX1 lines. Among these, HX1 is a multi-year selected line developed by our laboratory, exhibiting excellent comprehensive traits, strong drought resistance, and broad adaptability. JIN668 was provided by Professor Jin Shuangxia of Huazhong Agricultural University and serves as a commonly used model recipient in cotton genetic transformation research. The GhbHLH149-like overexpression vector was used to transform JIN668 by Agrobacterium tumefaciens-mediated transformation to obtain GhbHLH149-like overexpressing cotton lines. Cotton seeds were soaked in water for 12–18 h before being inoculated into nutrient soil (nutrient soil:vermiculite 1:1). After two weeks, uniformly growing seedlings were selected for salt stress treatment (200 mM NaCl solutions were supplemented with nutrient solution). The treatment was performed with three biological replicates. Five plants were mixed for RNA extraction per sample, which was subsequently used for qRT-PCR data analysis [ref. 2].
4.2. Identification of the GhMLOs Gene Family in G. hirsutum and Analysis of Their Protein Properties
Using the HMMER 3.0 software, candidate gene family members were identified by performing homology searches in the G. hirsutum reference genome using the MLO protein hidden Markov model (PF03094) provided by the Pfam 35.0 database. The GhMLO gene family was confirmed using the protein BLAST database (https://blast.ncbi.nlm.nih.gov/Blast.cgi (accessed on 4 August 2025)). The G. hirsutum TM-1 reference genome (v2.1), transcripts, amino acids, and 2000 base pairs upstream of the ATG promoter region were extracted from the COTTONOMICS database (http://cotton.zju.edu.cn/ (accessed on 4 August 2025)) [ref. 38]. GhMLOs were named according to their chromosomal positions (GhMLO1–GhMLO46). GhMLO gene positions and chromosome sizes were obtained from the COTTONOMICS database and visualized using TBtools -II (South China Agricultural University, Guangzhou, China) [ref. 39]. Protein physicochemical property predictions were performed using the ProtParam website (https://web.expasy.org/protparam/ (accessed on 4 August 2025)). Subcellular localization predictions for the GhMLO gene family were conducted via the Plant-mPLoc website (http://www.csbio.sjtu.edu.cn/bioinf/plant-multi/ (accessed on 4 August 2025)).
4.3. Sequence Alignment and Phylogenetic Analysis of the GhMLOs Gene Family
Using MEGA7.0 software, a phylogenetic tree of the GhMLOs gene family was constructed via the Neighbor-Joining (NJ) method. Branch reliability was assessed through 1000 bootstrap resampling runs [ref. 40]. The resulting tree file was further imported into the iTOL online platform (https://itol.embl.de/ (accessed on 20 August 2025)) for advanced visualization, clearly illustrating its evolutionary relationships.
The amino acid sequences of the ATML012, PtMLO12, and StMLO were obtained from the Phytozome14 database (https://phytozome-next.jgi.doe.gov/ (accessed on 20 August 2025)). Sequence comparisons and result visualization were performed using the ClustalW website (https://www.genome.jp/tools-bin/clustalw (accessed on 20 August 2025)) and ENDscript/ESPrip (https://espript.ibcp.fr/ESPript/ESPript/index.php (accessed on 20 August 2025)). Additionally, the protein structures of these four genes were predicted using the TMHMM-2.0 website (https://services.healthtech.dtu.dk/services/TMHMM-2.0/ (accessed on 20 August 2025)) [ref. 41].
4.4. Analysis of the GhMLOs Gene Family: Conserved Motifs, Conserved Domains, and Gene Structure
To analyze the structural characteristics of the GhMLOs gene family, we first imported a file containing all GhMLOs gene ID and gff3 annotation files into TBtools for visualizing their exon-intron layouts. Subsequently, GhMLO protein sequences were submitted to MEME Suite (https://meme-suite.org/meme/index.html (accessed on 25 October 2025)). Under default parameters, the motif count was set to 6 to generate corresponding MEME results. The obtained protein sequences were simultaneously uploaded to the NCBI protein database to retrieve Hitdata files. Finally, the three datasets were integrated within the TBtools platform to jointly construct a phylogenetic tree, a conserved motif map, and a gene structure diagram for the GhMLOs family. This enabled a comprehensive analysis of the family at the sequence, evolutionary, and structural levels [ref. 42].
4.5. Collinearity Analysis of the GhMLOs Gene Family
Using TBtools with default parameters, we performed MCScanX analysis on the G. hirsutum genome’s FAST and gff3 files. We further assessed collinearity between G. hirsutum and other species. We employed the default parameters of TBtools’ One Step MCScanX function to identify potential homologous genes. We compared the genomic FAST and gff3 files of G. hirsutum with those of other species, obtaining three key files: the genomic gff file, the CTL file (control file), and the Collinearity file. First, we manually removed unnamed chloroplast and mitochondrial sequences from the CTL file, then reordered the data, and finally visualized it using TBtools [ref. 43].
4.6. Analysis of Promoter Cis-Acting Elements in the GhMLOs Gene Family
The cis-acting elements within the 2000 bp upstream region of the ATG promoter for the GhMLOs gene family were predicted using the PlantCARE website (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/ (accessed on 25 October 2025)), and the results were visualized with TBtools.
4.7. Transcriptome Analysis of the GhMLOs Gene Family
To systematically analyze the expression characteristics of the GhMLOs gene family, this study downloaded the corresponding transcriptome dataset from the COTTONOMICS public database (http://cotton.zju.edu.cn/index.htm (accessed on 30 October 2025)). This dataset encompasses FPKM expression matrices from eight different tissues (root, stem, leaf, etc.) of G. hirsutum and three abiotic stress treatments (salt, simulated drought, cold). Subsequently, the “HeatMap” function module in TBtools software was used to generate an expression heatmap. Row scale normalization was applied with all other parameters set to default values to visually illustrate the differential expression patterns of GhMLOs across different tissues and stress conditions.
4.8. RT-qPCR Analysis of the GhMLOs Gene Family
RNA extraction was performed using the Nuoto® AutoExtracter-32 Nucleic Acid Extractor with the 5fz PCR DNA/RNA AutoPurification Kit from Kangma-Healthcode (Shanghai, China). Subsequently, first-strand cDNA was synthesized using the AT311 Reverse Transcription Kit (TransGen, Beijing, China). Specific primers were designed using the NCBI Primer-BLAST online tool (Supplementary Table S2). RT-qPCR was performed on a CFX96 Touch™ real-time fluorescent quantitative PCR instrument (Bio-Rad, Co. Ltd., Hercules, CA,USA) using the SYBR Green detection system. Reaction program: 95 °C pre-denaturation for 15 min; 45 cycles of 95 °C denaturation for 10 s, 60 °C annealing for 30 s, and 72 °C extension for 30 s [ref. 44]. Relative expression levels were calculated using the 2−ΔΔCT method, with GhUBQ7 (Ghir_D12G021700) as the reference gene [ref. 2], and four technical replicates to ensure accuracy.
4.9. Statistical Analyses
We analyzed RT-qPCR experimental data using GraphPad Prism 8(GraphPad Software, Inc., Boston, Massachusetts, USA). For relative gene expression levels, we performed one-way analysis of variance (ANOVA) with LSD multiple comparison tests (p < 0.05). Prior to ANOVA, data were tested for normality and homogeneity of variance [ref. 45].
5. Conclusions
The MLO gene family is a plant-specific gene family with seven transmembrane domains and carboxyl-terminal calmodulin binding domains. The plant GhMLO gene is integrated and regulated by ABA, ROS, and temperature signaling networks, making it an important node for studying stress-immune cross-responses. By genomic and bioinformatics methods, 46 GhMLO genes in the MLO (Mildew resistance Locus O) gene family of G. hirsutum were classified and identified in this study. Chromosomal localization, evolutionary tree, gene structure, gene doubling, promoter cis-element, and expression profile of GhMLO were analyzed. Members of the GhMLO family could be significantly induced to expression by drought, low-temperature stress and salt stress, suggesting that GhMLO may play an important role in the resistance of G. hirsutum to abiotic stress. This study represents a preliminary investigation into the GhMLO gene family in G. hirsutum. The results provide candidate genes and a basis for future studies exploring their possible roles in abiotic stress adaptation, which could ultimately contribute to the development of cotton varieties with enhanced tolerance to salt, drought, and temperature stresses. Expression levels of the GhMLO43 gene in GhbHLH149-like overexpression lines and under salt stress.
References
- Y. Chen, Z. Yang, Y. Xiao, P. Wang, Y. Wang, X. Ge, C. Zhang, X. Zhang, F. Li. Genome-Wide Analysis of the NF-YB Gene Family in Gossypium hirsutum L. and Characterization of the Role of GhDNF-YB22 in Embryogenesis. Int. J. Mol. Sci., 2018. [DOI | PubMed]
- M. Jiang, C. Feng, J. Zhen, L. Liu, D. Liu, S. Zhang, J. Chi. Genome-Wide Identification and Characterization of the VOZ Gene Family in Gossypium hirsutum L. and Functional Characterization in Abiotic Stress and Somatic Embryogenesis. Int. J. Mol. Sci., 2025. [DOI | PubMed]
- J.-F. Cao, J.-Q. Huang, X. Liu, C.-C. Huang, Z.-S. Zheng, X.-F. Zhang, X.-X. Shangguan, L.-J. Wang, Y.-G. Zhang, J.F. Wendel. Genome-wide characterization of the GRF family and their roles in response to salt stress in Gossypium. BMC Genom., 2020. [DOI]
- P. Yang, J. Jin, J. Zhang, D. Wang, X. Bai, W. Xie, T. Hu, X. Zhao, T. Mao, T. Qin. MDP25 mediates the fine-tuning of microtubule organization in response to salt stress. J. Integr. Plant Biol., 2022. [DOI | PubMed]
- M. Zhang, X. Liang, L. Wang, Y. Cao, W. Song, J. Shi, J. Lai, C. Jiang. A HAK family Na+ transporter confers natural variation of salt tolerance in maize. Nat. Plants, 2019. [DOI | PubMed]
- W. Xu, Y. Wang, J. Xie, S. Tan, H. Wang, Y. Zhao, Q. Liu, Y.A. El-Kassaby, D. Zhang. Growth-regulating factor 15-mediated gene regulatory network enhances salt tolerance in poplar. Plant Physiol., 2023. [DOI | PubMed]
- Z. Chan, P.B. Cao, S. Azar, H. SanClemente, F. Mounet, C. Dunand, G. Marque, C. Marque, C. Teulières. Genome-Wide Analysis of the AP2/ERF Family in Eucalyptus grandis: An Intriguing Over-Representation of Stress-Responsive DREB1/CBF Genes. PLoS ONE, 2015. [DOI | PubMed]
- X.-M. Han, Z.-L. Yang, Y.-J. Liu, H.-L. Yang, Q.-Y. Zeng. Genome-wide profiling of expression and biochemical functions of the Medicago glutathione S-transferase gene family. Plant Physiol. Biochem., 2018. [DOI | PubMed]
- W. Hu, H. Yan, S. Luo, F. Pan, Y. Wang, Y. Xiang. Genome-wide analysis of poplar SAUR gene family and expression profiles under cold, polyethylene glycol and indole-3-acetic acid treatments. Plant Physiol. Biochem., 2018. [DOI | PubMed]
- P. Iovieno, G. Andolfo, A. Schiavulli, D. Catalano, L. Ricciardi, L. Frusciante, M.R. Ercolano, S. Pavan. Structure, evolution and functional inference on the Mildew Locus O (MLO) gene family in three cultivated Cucurbitaceae spp.. BMC Genom., 2015. [DOI]
- N. Rispail, D. Rubiales. Genome-wide identification and comparison of legume MLO gene family. Sci. Rep., 2016. [DOI | PubMed]
- J. Liu, Y. Wu, X. Zhang, R.A. Gill, M. Hu, Z. Bai, C. Zhao, Y. Zhang, Y. Liu, Q. Hu. Functional and evolutionary study of MLO gene family in the regulation of Sclerotinia stem rot resistance in Brassica napus L.. Biotechnol. Biofuels Bioprod., 2023. [DOI | PubMed]
- R.R. Tapia, C.R. Barbey, S. Chandra, K.M. Folta, V.M. Whitaker, S. Lee. Evolution of the MLO gene families in octoploid strawberry (Fragaria × ananassa) and progenitor diploid species identified potential genes for strawberry powdery mildew resistance. Hortic. Res., 2021. [DOI | PubMed]
- J. Acevedo-Garcia, S. Kusch, R. Panstruga. Magical mystery tour: MLO proteins in plant immunity and beyond. New Phytol., 2014. [DOI | PubMed]
- Z. Chen, H.A. Hartmann, M.-J. Wu, E.J. Friedman, J.-G. Chen, M. Pulley, P. Schulze-Lefert, R. Panstruga, A.M. Jones. Expression analysis of the AtMLO Gene Family Encoding Plant-Specific Seven-Transmembrane Domain Proteins. Plant Mol. Biol., 2006. [DOI | PubMed]
- H. Cheng, W. Kong, D. Hou, J. Lv, X. Tao. Isolation, characterization, and expression analysis of CmMLO2 in muskmelon. Mol. Biol. Rep., 2012. [DOI | PubMed]
- M. Appiano, D. Catalano, M. Santillán Martínez, C. Lotti, Z. Zheng, R.G.F. Visser, L. Ricciardi, Y. Bai, S. Pavan. Monocot and dicot MLO powdery mildew susceptibility factors are functionally conserved in spite of the evolution of class-specific molecular features. BMC Plant Biol., 2015. [DOI | PubMed]
- Z. Zheng, M. Appiano, S. Pavan, V. Bracuto, L. Ricciardi, R.G.F. Visser, A.-M.A. Wolters, Y. Bai. Genome-Wide Study of the Tomato SlMLO Gene Family and Its Functional Characterization in Response to the Powdery Mildew Fungus Oidium neolycopersici. Front. Plant Sci., 2016. [DOI | PubMed]
- B.R. Acharya, R.N. Day, K.H. Day, F.M. Pavalko. Direct visualization by FRET-FLIM of a putative mechanosome complex involving Src, Pyk2 and MBD2 in living MLO-Y4 cells. PLoS ONE, 2021. [DOI | PubMed]
- N. Pépin, F.O. Hebert, D.L. Joly. Genome-Wide Characterization of the MLO Gene Family in Cannabis sativa Reveals Two Genes as Strong Candidates for Powdery Mildew Susceptibility. Front. Plant Sci., 2021. [DOI | PubMed]
- T. Zhang, N. Xu, S. Amanullah, P. Gao. Genome-wide identification, evolution, and expression analysis of MLO gene family in melon (Cucumis melo L.). Front. Plant Sci., 2023. [DOI | PubMed]
- X. Wang, Q. Ma, L. Dou, Z. Liu, R. Peng, S. Yu. Genome-wide characterization and comparative analysis of the MLO gene family in cotton. Plant Physiol. Biochem., 2016. [DOI | PubMed]
- S. Han, J.M. Smith, Y. Du, A.F. Bent. Soybean transporter AAT Rhg1 abundance increases along the nematode migration path and impacts vesiculation and ROS. Plant Physiol., 2023. [DOI | PubMed]
- R. Oelmüller, Y.-H. Tseng, A. Gandhi. Signals and Their Perception for Remodelling, Adjustment and Repair of the Plant Cell Wall. Int. J. Mol. Sci., 2023. [DOI | PubMed]
- M. Wouters, H. Bastiaanse, S. Rombauts, L. de Vries, T. De Pooter, M. Strazisar, G. Neutelings, R. Vanholme, W. Boerjan. Suppression of the Arabidopsis cinnamoyl-CoA reductase 1-6 intronic T-DNA mutation by epigenetic modification. Plant Physiol., 2023. [DOI | PubMed]
- H. Jian, F. Wei, P. Chen, T. Hu, X. Lv, B. Wang, H. Wang, X. Guo, L. Ma, J. Lu. Genome-wide analysis of SET domain genes and the function of GhSDG51 during salt stress in upland cotton (Gossypium hirsutum L.). BMC Plant Biol., 2023. [DOI | PubMed]
- K. Morioka, C. Tanikawa, K. Ochi, Y. Daigo, T. Katagiri, H. Kawano, H. Kawaguchi, A. Myoui, H. Yoshikawa, N. Naka. Orphan receptor tyrosine kinase ROR2 as a potential therapeutic target for osteosarcoma. Cancer Sci., 2009. [DOI | PubMed]
- G. Rebagay, S. Yan, C. Liu, N.-K. Cheung. ROR1 and ROR2 in Human Malignancies: Potentials for Targeted Therapy. Front. Oncol., 2012. [DOI | PubMed]
- J.B. Blackmon, T. Salverson. Analytical Advances in Reciprocating Feed System Trade Study Code. Proceedings of the AIAA Propulsion and Energy 2020 Forum
- T. Ma, X. Tian, B. Zhang, M. Li, Y. Wang, C. Yang, J. Wu, X. Wei, Q. Qu, Y. Yu. Low-dose metformin targets the lysosomal AMPK pathway through PEN2. Nature, 2022. [DOI | PubMed]
- M. Wang, Q. Xu, J. Yu, M. Yuan. The putative Arabidopsis zinc transporter ZTP29 is involved in the response to salt stress. Plant Mol. Biol., 2010. [DOI | PubMed]
- F. Yi, Y. Li, A. Song, X. Shi, S. Hu, S. Wu, L. Shao, Z. Chu, K. Xu, L. Li. Positive roles of the Ca2+ sensors GbCML45 and GbCML50 in improving cotton Verticillium wilt resistance. Mol. Plant Pathol., 2024. [DOI | PubMed]
- M. Beltramino, J.M. Debernardi, A. Ferela, J.F. Palatnik. ARF2 represses expression of plant GRF transcription factors in a complementary mechanism to microRNA miR396. Plant Physiol., 2021. [DOI | PubMed]
- X. Zhang, J. Cui, C. Zhang, X. Yang, H. Pan, H. Du, A. Ahmad, T. Wu, C. Yao. Genome-wide identification and analysis of the MLO gene family for candidate powdery mildew susceptibility factors in Momordica charantia. Sci. Hortic., 2021. [DOI]
- W.-Z. Liu, M. Deng, L. Li, B. Yang, H. Li, H. Deng, Y.-Q. Jiang. Rapeseed calcineurin B-like protein CBL4, interacting with CBL-interacting protein kinase CIPK24, modulates salt tolerance in plants. Biochem. Biophys. Res. Commun., 2015. [DOI | PubMed]
- Y. Yang, C. Zhang, R.-J. Tang, H.-X. Xu, W.-Z. Lan, F. Zhao, S. Luan. Calcineurin B-Like Proteins CBL4 and CBL10 Mediate Two Independent Salt Tolerance Pathways in Arabidopsis. Int. J. Mol. Sci., 2019. [DOI | PubMed]
- Q. Zhang, C. Hou, Y. Tian, M. Tang, C. Feng, Z. Ren, J. Song, X. Wang, T. Li, M. Li. Interaction Between AtCML9 and AtMLO10 Regulates Pollen Tube Development and Seed Setting. Front. Plant Sci., 2020. [DOI | PubMed]
- F. Dai, J. Chen, Z. Zhang, F. Liu, J. Li, T. Zhao, Y. Hu, T. Zhang, L. Fang. COTTONOMICS: A comprehensive cotton multi-omics database. Database, 2022. [DOI | PubMed]
- C. Chen, Y. Wu, J. Li, X. Wang, Z. Zeng, J. Xu, Y. Liu, J. Feng, H. Chen, Y. He. TBtools-II: A “one for all, all for one” bioinformatics platform for biological big-data mining. Mol. Plant, 2023. [DOI | PubMed]
- B.G. Hall. Building Phylogenetic Trees from Molecular Data with MEGA. Mol. Biol. Evol., 2013. [DOI | PubMed]
- C.-H. Feng, M.-X. Niu, X. Liu, Y. Bao, S. Liu, M. Liu, F. He, S. Han, C. Liu, H.-L. Wang. Genome-Wide Analysis of the FBA Subfamily of the Poplar F-Box Gene Family and Its Role under Drought Stress. Int. J. Mol. Sci., 2023. [DOI | PubMed]
- F. He, Y.-J. Shi, Q. Chen, J.-L. Li, M.-X. Niu, C.-H. Feng, M.-M. Lu, F.-F. Tian, F. Zhang, T.-T. Lin. Genome-Wide Investigation of the PtrCHLP Family Reveals That PtrCHLP3 Actively Mediates Poplar Growth and Development by Regulating Photosynthesis. Front. Plant Sci., 2022. [DOI | PubMed]
- F. He, Y.-J. Shi, J.-L. Li, T.-T. Lin, K.-J. Zhao, L.-H. Chen, J.-X. Mi, F. Zhang, Y. Zhong, M.-M. Lu. Genome-wide analysis and expression profiling of Cation/H+ exchanger (CAX) family genes reveal likely functions in cadmium stress responses in poplar. Int. J. Biol. Macromol., 2022. [DOI | PubMed]
- H.L. Wang, J. Chen, Q. Tian, S. Wang, X. Xia, W. Yin. Identification and validation of reference genes for Populus euphratica gene expression analysis during abiotic stresses by quantitative real-time PCR. Physiol. Plant., 2014. [DOI | PubMed]
- F. He, H.G. Li, J.J. Wang, Y. Su, H.L. Wang, C.H. Feng, Y. Yang, M.X. Niu, C. Liu, W. Yin. PeSTZ1, a C2H2-type zinc finger transcription factor from Populus euphratica, enhances freezing tolerance through modulation of ROS scavenging by directly regulating PeAPX2. Plant Biotechnol. J., 2019. [DOI | PubMed]
