Synthesis, Molecular Docking, and Biological Studies of New Naphthalene‐1,4‐Dione‐Linked Phenylpiperazine and Thioether
Abstract
Quinones are an important family of chemical compounds that are used in the development of anticancer drugs. Due to the anticancer properties of quinones, this study aims to investigate the antiproliferative and antimicrobial activities of newly substituted quinone derivatives synthesized through reactions with various nucleophiles (such as thiols and amines). The antiproliferative effects of the synthesized compounds on human cancer cell lines were screened at a concentration of 10 µM using the 3‐(4,5‐dimethylthiazol‐2‐yl)‐2,5‐diphenyltetrazolium bromide (MTT) assay. The cytotoxic effects of the synthesized compounds were also assessed in NIH/3T3 cells. Target molecules for the cancer cell lines A549, PC‐3, and MCF‐7 employed in this study were identified in light of the literature in order to perform molecular docking studies based on the MTT experiment results. Moreover, the antimicrobial effects of newly synthesized quinone derivatives were also investigated. Among the synthesized compounds, especially compound 10 had strong antiproliferative effects on a variety of cancer cell lines, most notably PC‐3, while having no cytotoxicity toward NIH/3T3 cells. Furthermore, compounds 5 and 8 exhibited antimicrobial activity against both bacteria and fungi. Our results indicate that the newly synthesized quinone molecules may function as therapeutic agents for the treatment of cancer and infectious diseases.
Article type: Research Article
Keywords: anticancer activity, antimicrobial activity, molecular docking, naphthoquinone, spectroscopy
Affiliations: Faculty of Engineering, Department of Chemistry, Division of Organic Chemistry Istanbul University‐Cerrahpaşa Istanbul Turkey; Faculty of Pharmacy, Department of Biochemistry Marmara University, Basibuyuk Campus Istanbul Turkey; Institute of Health Sciences, Department of Biochemistry (Pharmacy) Marmara University, Basibuyuk Campus Istanbul Turkey; Faculty of Pharmacy, Department of Pharmaceutical Microbiology Marmara University, Basibuyuk Campus Istanbul Turkey; Faculty of Pharmacy, Department of Basic Pharmaceutical Sciences Marmara University, Basibuyuk Campus Istanbul Turkey
License: © 2025 The Author(s). Chemistry & Biodiversity published by Wiley‐VHCA AG. CC BY 4.0 This is an open access article under the terms of the http://creativecommons.org/licenses/by/4.0/ License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Article links: DOI: 10.1002/cbdv.202500774 | PubMed: 40714915 | PMC: PMC12629165
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (1.0 MB)
Introduction
Cancer is one of the most prevalent and lethal diseases worldwide, contributing to a significant number of deaths annually. Because current cancer therapy regimens lack selectivity and have many adverse effects, research into alternative treatment techniques is still important. For decades, scientists have continued to investigate potential new drug candidates for cancer treatment by exploring both naturally derived compounds and synthetic pharmaceutical agents produced in laboratory settings [ref. 1, ref. 2].
Compounds with a naphthoquinone (NQ) framework are known to show strong activity against human breast, liver, cervical, and gastric cancer cells by inducing apoptosis [ref. 3, ref. 4, ref. 5]. The primary mechanism underlying the anticancer effect of quinones is thought to be the triggering of the autophagy system [ref. 6, ref. 7, ref. 8, ref. 9] and the formation of the reactive oxygen species (ROS) [ref. 10, ref. 11, ref. 12], such as hydroxyl radicals [ref. 13]. Ansamycin antibiotics, substituted‐1,4‐NQ derivatives containing amino groups, represent a significant part of antibiotics with anticancer activity [ref. 14, ref. 15]. Many NQ compounds are toxic to various cancer cell lines due to properties of redox potential [ref. 16, ref. 17, ref. 18, ref. 19]. For example, it has been shown that some alkylamino quinone derivatives have a high cytotoxic effect against human colon, brain, and pancreatic cancer cells [ref. 20]. Ourworking group has investigated a variety of quinone compounds and their uses in anticancer research on quinone derivatives replaced with heterogroups [ref. 21, ref. 22, ref. 23, ref. 24, ref. 25, ref. 26]. Furthermore, we discovered that 2‐(tert‐butylthio)‐3‐chloronaphthalene‐1,4‐dione, one of the NQ derivatives with a sulfur atom substitution, had a strong antiproliferative effect on HeLa cells with an IC50 value of 10.16 µM [ref. 27, ref. 28, ref. 29].
We planned to direct our synthesis toward the discovery of novel N‐, S‐substituted quinone analogues and investigate their antiproliferative activities against human lung, breast, and prostate cancer cells and their cytotoxic activities on mouse embryonic fibroblast cells in this study. The molecular docking approach is a simulation technique used to predict potential interactions in drug development processes. This method allows for the determination of interactions and binding affinities between drug candidate molecules and their target receptors. Drug development efforts can benefit from generating a prediction profile by analyzing the interactions between disease‐related macromolecules and potential therapeutic compounds, providing valuable insights to complement experimental studies. Given the anticancer properties of 1,4‐NQ derivatives, molecular docking studies were conducted to explore the interactions between newly synthesized naphthalene‐1,4‐dione‐linked phenylpiperazine and thioether derivatives and various cancer‐related targets in this study. Moreover, the antimicrobial activity potential of quinone derivatives was determined by the agar well diffusion method. As well, the minimal inhibitory concentrations (MICs) of quinone derivatives were determined.
Results and Discussion
Chemistry of 1,4‐NQ Derivatives
The reaction of 2,3‐dihalo(Cl)‐1,4‐NQ (1) with some nucleophiles (2a–d) gave S,S‐, N‐, and N,S‐substituted quinone analogues (Scheme 1). Both halogen atoms, such as chlorine and bromine, can be replaced by using nitrogen‐containing heterocycles [ref. 30]. The one strong nucleophile property reacts with 2,3‐dihalo‐1,4‐NQ; only one chlorine atom can be displaced due to the electronic density of the quinone system. The displacement of the second chlorine atom and its separation from the structure can be easily achieved if the quinone system contains an electron‐withdrawing group (EWG) or in the presence of ethanol [ref. 31, ref. 32, ref. 33, ref. 34, ref. 35].
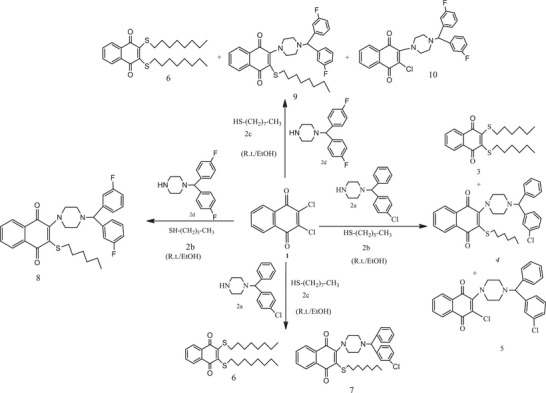
The synthesis of 1,4‐NQ derivatives (3–10) was investigated by the reactions of 2,3‐dihalo‐1,4‐NQ (1) with 1‐(4‐chlorobenzhydryl)piperazine (2a), hexanethiol (2b), octanethiol (2c), and 1‐[bis(4‐fluorophenyl)‐methyl]piperazine (2d) (Scheme 1).
In a previous study, heteroatom‐substituted NQ derivatives containing piperonylpiperazine or diphenylmethylpiperazine in Position two and chlorine atom in Position three of the 1,4‐NQ moiety showed good antiproliferative profiles and also antibacterial and antioxidant capacity. In the literature [ref. 31], the compounds of 2‐(N‐diphenylmethylpiperazin‐1‐yl)‐3‐chloro‐1,4‐NQ and 2‐[1‐piperonylpiperazin‐1‐yl]‐3‐chloro‐1,4‐NQ showed a strong cytotoxic effect on MCF‐7 cell lines and significant antibacterial activity against Micrococcus luteum. Interestingly, they also have powerful antioxidant capacity [ref. 31]. In this study, a similar compound, 2‐(4‐(bis(3‐fluorophenyl)methyl)piperazin‐1‐yl)‐3‐chloronaphthalene‐1,4‐dione, also gave similar results. The strong antimicrobial activity demonstrated by our similar compounds has encouraged us to further our studies on this subject.
Spectral Studies of Synthesized Compounds
As illustrated in Scheme 1, when starting compound (1) reacted in a single reaction vessel with an equimolar amount of various amines and thiols (2a–d). The products (4 and 7–9) were synthesized in good yields with appropriate reaction times and high purity. In the FT‐IR spectra of newly synthesized compounds, the stretching bands for the carbonyl group gave peaks in the range of 1537–1730 cm−1.
According to 13C(APT) NMR spectrum, except for synthesized quinone compounds (4, 5, and 7–10), two carbonyl signals were given between 178.05 and 182.90 ppm. It was proved that two different substituents were connected to the quinone unit. However, the individual carbonyl carbons (C═O) of compounds 3 and 6 provided single chemical shift values at δ 178.94 and 178.98 ppm, respectively. These 13C(APT) shift values have agreed well with the related literature [ref. 36, ref. 37, ref. 38, ref. 39].
The respective molecular ion peaks were observed at m/z (%) m/z = 391.28329 [M]+ (compound 3), 559.21674 [M + H]+ (compound 4), 477.11203 [M]+ (compound 5), 447.23743 [M]+ (compound 6), 587.24829 [M]+ (compound 7), 561.23718 [M]+ (compound 8), 589.26807 [M]+ (compound 9), and 479.13260 [M + H]+ (compound 10), respectively, for the synthesized compounds (3–10) by using the ESI method of high‐resolution mass spectrum (HRMS) method.
Antiproliferative Activity of Synthesized Compounds
The antiproliferative effect of the synthesized compounds (3–10) on human cancer cell lines was screened at a concentration of 10 µM using the 3‐(4,5‐dimethylthiazol‐2‐yl)‐2,5‐diphenyltetrazolium bromide (MTT) assay. The effect of the compounds on the cell viability at 24 h is shown in Figure 1. The cytotoxic effects of the synthesized compounds were also assessed in NIH/3T3 cells. According to the results, compound 10 was determined to be the most effective compound against all cancer cell lines in this study. The highest inhibitory activity of compound 2‐(4‐(Bis(3‐Fluorophenyl)Methyl)Piperazin‐1‐yl)‐3‐Chloronaphthalene‐1,4‐Dion) 10 was observed in PC‐3 cells (51.66%). Additionally, compound 10 has low toxicity toward healthy cells (10.1%), indicating that it does not significantly harm NIH/3T3 cells. Interestingly, compound 10 showed a stronger inhibition on cancer cell proliferation than the reference drug doxorubicin (DOX), which contains a 1,4‐NQ ring. On the other hand, the compounds 5 and 7 showed promising results against A549 cells (36.17% and 36.71%, respectively); however, the remaining compounds at 10 µM concentration had virtually no antiproliferative effects.
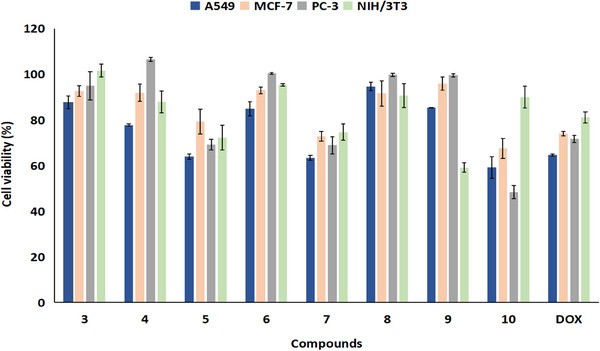
The fact that compound 10 has an effect on all three cancer cell lines is due to the fluorophenyl piperazine ring it contains. The biological effects of piperazine‐conjugated molecules might vary greatly, with the main one being anticancer action [ref. 40, ref. 41]. Consistent with our results, a very recent study by Zsoldos et al. reported that 1‐bis(4‐fluorophenyl)methylpiperazine derivatives exhibited potent anticarcinogenic activity on all cancer cell lines used in the study [ref. 42].
Molecular Docking Studies of Compounds (5, 7, and 10)
On the basis of the results of the MTT experiment, docking was carried out among the three compounds (5, 7, and 10) that gave the highest cytotoxic activity in all three cancer cell lines and the target proteins. The selection of target proteins for molecular docking studies was based on their biological roles in cancer signaling pathways as supported by the literature [ref. 43, ref. 44, ref. 45]. The cell lines utilized in this study (A549, PC‐3, and MCF‐7) serve as model systems for lung, prostate, and breast cancers, respectively. For these cell lines, proteins such as epidermal growth factor receptor (EGFR), vascular endothelial growth factor receptor 2 (VEGFR2), mammalian target of rapamycin (mTOR), Caspase‐3, focal adhesion kinase (FAK), and cyclin‐dependent kinase 6 (CDK6) were selected due to their critical roles in the biology of the respective cancer types, as highlighted in previous studies. Comparison with docetaxel, which is used as a positive control, reveals that three synthesized quinone compounds (5, 7, and 10) exhibit significantly higher binding scores, as depicted in Table 2.
TABLE 2: Inhibition zones of quinone derivatives determined by the agar well diffusion method (mm).
| Quinone derivatives | Staphylococcus aureus ATCC 25923 | Staphylococcus aureus ATCC 43300 | Staphylococcus epidermidis ATCC 12228 | Enterococcus faecalis ATCC 29212 | Pseudomonas aeruginosa ATCC 27853 | Escherichia coli ATCC 25922 | Klebsiella pneumoniae ATCC 4352 | Candida albicans ATCC 90028 |
|---|---|---|---|---|---|---|---|---|
| 3 | 11.10 ± 0.18 | 27.87 ± 0.17 | 10.92 ± 0.19 | 0 | 0 | 0 | 9.46 ± 0.17 | 17.79 ± 0.24 |
| 4 | 0 | 17.45 ± 0.15 | 23.58 ± 0.22 | 0 | 15.46 ± 0.31 | 0 | 14.77 ± 0.27 | 0 |
| 5 | 15.13 ± 0.22 | 16.35 ± 0.24 | 15.73 ± 0.23 | 29.30 ± 0.21 | 12.22 ± 0.24 | 14.31 ± 0.17 | 11.93 ± 0.25 | 21.12 ± 0.11 |
| 7 | 21.00 ± 0.25 | 16.83 ± 0.23 | 16.70 ± 0.31 | 19.27 ± 0.20 | 13.18 ± 0.27 | 0 | 16.22 ± 0.25 | 16.37 ± 0.13 |
| 8 | 29.73 ± 0.11 | 15.28 ± 0.10 | 13.84 ± 0.14 | 12.96 ± 0.23 | 10.63 ± 0.10 | 11.41 ± 0.08 | 11.13 ± 0.08 | 17.40 ± 0.18 |
| 9 | 0 | 12.28 ± 0.10 | 15.42 ± 0.25 | 0 | 18.23 ± 0.21 | 0 | 19.41 ± 0.29 | 0 |
| 10 | 9.83 ± 0.14 | 0 | 14.13 ± 0.17 | 14.20 ± 0.15 | 7.32 ± 0.09 | 10.83 ± 0.21 | 12.67 ± 0.17 | 17.82 ± 0.31 |
| M (10 µg/mL) | 31.80 ± 0.29 | 29.68 ± 0.32 | 23.49 ± 0.42 | 18.44 ± 0.22 | 29.32 ± 0.21 | 30.12 ± 0.12 | 28.56 ± 0.25 | — |
| AmB (100 µg/mL) | — | — | — | — | — | — | — | 21.14 ± 0.27 |
Note: “—”: untested.
Abbreviations: AmB, Amphotericin B; M, meropenem.
Simultaneously with the cytotoxic activity results, we observed that the molecule coded as compound 10 demonstrates excellent docking scores against all proteins. Furthermore, the highest score (−9.8 kcal/mol), as indicated in Table 1, is obtained from the docking experiment of the compound 10 molecule with the EGFR protein.
TABLE 1: Docking scores of compounds (5, 7, and 10) with proteins (kcal/mol).
| Proteins | Docetaxel | 5 | 7 | 10 |
|---|---|---|---|---|
| EGFR | −8.5 | −9.4 | −8.9 | −9.8 |
| Caspase‐3 | −7.1 | −7.6 | −6.4 | −8 |
| VEGFR2 | −6.7 | −8.4 | −9 | −9 |
| mTOR | −8 | −8.6 | −8.7 | −9.3 |
| FAK | −6.6 | −9.4 | −8.7 | −9.5 |
| CDK6 | −7.1 | −9.8 | −8.3 | −9.3 |
Abbreviations: CDK6, cyclin‐dependent kinase 6; EGFR, epidermal growth factor receptor; FAK, focal adhesion kinase; mTOR, mammalian target of rapamycin; VEGFR2, vascular endothelial growth factor receptor 2.
The overexpression or mutation of EGFR is often associated with the poor prognosis, rapid metastasis, short‐term recurrence, and short survival time of epithelium tumors, such as breast, gastrointestinal, ovarian, prostate, and cervical cancer [ref. 46]. Increasing evidence has shown that EGFR inhibitors have great potential in cancer treatment, especially for non‐small cell lung cancer, colon cancer, and hepatocellular carcinoma, which has led to an increasing number of research studies based on the design and synthesis of EGFR inhibitors [ref. 43, ref. 47]. NQs have been shown to be involved in various molecular processes that drive cancer cells to enter the apoptotic process, such as suppression of the EGFR–NF‐κB signaling pathway [ref. 48]. Furthermore, many studies have shown that 1,4‐NQ derivatives exhibit strong effects on various cancer cells as potential EGFR inhibitors [ref. 19, ref. 29, ref. 49].
Overexpression of EGFR activates pro‐oncogenic downstream signaling pathways, including PI3K/AKT/mTOR [ref. 50, ref. 51]. This signaling plays key roles in oncogenesis‐related events, such as cell survival, cell proliferation, cell differentiation, and cellular apoptosis. In vitro studies have shown that PI3K/AKT/mTOR signaling plays an important role not only in the proliferation and apoptosis of prostate cancer cells [ref. 52] but also in migration and invasion [ref. 53]. In our study, compound 10 exhibits the strongest inhibitory effect on prostate cancer cells and shows a strong binding with mTOR, suggesting that this compound may be an alternative potential therapeutic agent for prostate cancer treatment by inhibiting PI3/AKT/mTOR signaling.
Analyzing the protein–ligand interaction of the positive control and the compound 10 molecule when docked with EGFR, considering the numerical data, both molecules exhibit high‐density hydrophobic interactions with similar amino acids (Figure 2a,b). Moreover, it is noteworthy that both molecules position themselves in a comparable location within the active region of the protein (Figure 2c,d).
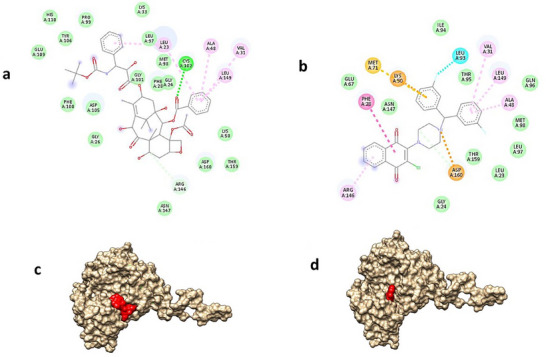
In conclusion, the docking studies were found to be in agreement with the in vitro antiproliferative data, particularly for compound 10. The observed interactions between compound 10 and EGFR suggest a possible mechanism of action, which may help guide future experimental investigations.
Antimicrobial Activity of Quinone Derivatives
In the present study, compounds 5 and 8 were effective against all of the microorganisms, both bacteria and yeast. Compound 7 was effective against all of the microorganisms except Escherichia coli ATCC 25922, and compound 10, a derivative, was effective against all of the microorganisms except Staphylococcus aureus ATCC 43300 (Table 3). Moreover, compound 3 was effective against S. aureus ATCC 25923 (Figure 3), S. aureus ATCC 43300, Staphylococcus epidermidis ATCC 11228, Klebsiella pneumoniae ATCC 4352, and Candida albicans ATCC 90028. Compounds 4 and 9 showed similar activities as being effective against S. aureus ATCC 43300, S. epidermidis ATCC 11228, Pseudomonas aeruginosa ATCC 27853, and K. pneumoniae ATCC 4352 (Table 2).
TABLE 3: The antimicrobial activity of quinone derivatives (3–10) (µM).
| Compounds | Staphylococcus aureus ATCC 25923 | Staphylococcus aureus ATCC 43300 | Staphylococcus epidermidis ATCC 12228 | Enterococcus faecalis ATCC 29212 | Pseudomonas aeruginosa ATCC 27853 | Escherichia coli ATCC 25922 | Klebsiella pneumoniae ATCC 4352 | Candida albicans ATCC 90028 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MIC | MBC | MIC | MBC | MIC | MBC | MIC | MBC | MIC | MBC | MIC | MBC | MIC | MBC | MIC | MFC | |
| 3 | 3.51 | 6.99 | 112.01 | 224.01 | 224.01 | 448.03 | >448.03 | >448.03 | >448.03 | >448.03 | >448.03 | >448.03 | 56.02 | 112.01 | 112.01 | 224.01 |
| 4 | >89.58 | >89.58 | 11.20 | 22.40 | 44.79 | 89.58 | >89.58 | >89.58 | 11.20 | 22.40 | >89.58 | >89.58 | 22.40 | 44.79 | >89.58 | >89.58 |
| 5 | 1.63 | 3.27 | 26.18 | 52.36 | 52.36 | 104.72 | 26.18 | 52.36 | 6.56 | 13.09 | 26.18 | 52.36 | 6.56 | 13.09 | 26.18 | 52.36 |
| 7 | 3.73 | 7.46 | 14.90 | 29.80 | 14.90 | 29.80 | 29.80 | 59,60 | 7.46 | 14.90 | >29.80 | >29.80 | 14.90 | 29.80 | 14.90 | 29.80 |
| 8 | 64.65 | 129.30 | 16.16 | 32.33 | 64.65 | 129.30 | 64.65 | 129.30 | 32.33 | 64.65 | 32.33 | 64.65 | 8.08 | 16.16 | 16.16 | 32.33 |
| 9 | >55.20 | >55.20 | 6.90 | 13.81 | 13.81 | 27.60 | >55.20 | >55.20 | 27.60 | 55.20 | >55.20 | >55.20 | 13.81 | 27.60 | >55.20 | >55.20 |
| 10 | 36.60 | 73.20 | >292.81 | >292.81 | 73.20 | 146.40 | 146.40 | 292.81 | 146.40 | 292.81 | 73.20 | 146.40 | 18.30 | 36.60 | 146.40 | 292.81 |
| M | 0.0052 | 0.0104 | 0.0417 | 0.0834 | 0.0007 | 0.0013 | 0.0209 | 0.0417 | 0.0013 | 0.0052 | 0.0002 | 0.0003 | 0.0013 | 0.0052 | — | — |
| AmB | — | — | — | — | — | — | — | — | — | — | — | — | — | — | 1.08 | 4.33 |
Note: “—”: untested.
Abbreviations: AmB, Amphotericin B; M, meropenem; MBC, minimal bactericidal concentration; MFC, minimal fungicidal concentration; MIC, minimal inhibition concentration.
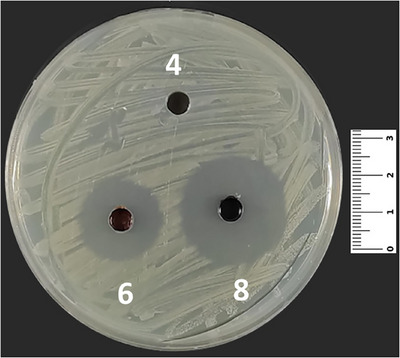
Cho et al. investigated the antimicrobial effects of 1,4‐quinone derivatives against various microorganisms [ref. 54, ref. 55, ref. 56]. In their study, the authors declared that synthesized compounds (3–5 and 7–10) showed antibacterial effects against S. aureus and P. aeruginosa. Additionally, it was stated that the five synthesized compounds (3–5, 7, and 9) in our study have shown antifungal effects (Figure 4) against C. albicans at various concentrations of MIC (Table 3).
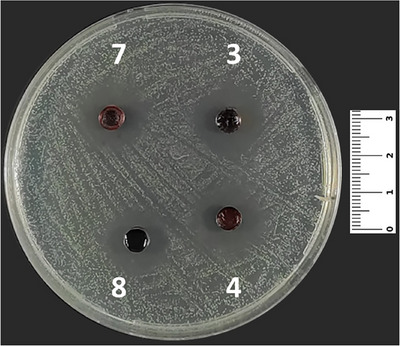
As it is known, quinone can be transformed to detrimental ROS by catalyzing redox cycles where NAP(P)H and molecular O2 take place. Moreover, some quinone derivatives have the ability to show their antiproliferative activity by impairing mitochondrial functions not only in bacteria (such as S. aureus) but also in fungi and protozoa [ref. 57].
In order to find out a solution for new agents against the antimicrobial resistance, Ulfah et al. have shown that diterpene quinone–derived compounds are safe to use. The investigators have stated that in vitro assay results of their study had shown that the antimicrobial activities of diterpene quinone–derived compounds were related to the inhibition of ATP as a function and disruption of the membrane function of S. aureus. Additionally, diterpene quinone–derived compounds exhibited good antibacterial activity. On the basis of in vitro and in silico assays, the mechanism of action of this compound is related to the disruption of bacterial membrane function, and it has the potential to inhibit the ATPase enzyme and the S. aureus gyrase [ref. 58].
In our opinion, the synthesized naphthalene‐1,4‐diones in our study (naphthalene‐1,4‐dione‐linked phenyl piperazine and thioether) might show efficacy against microorganisms by detrimental ROS by catalyzing redox cycles and impairing mitochondrial functions and bacterial membrane function and inhibiting the ATPase enzyme.
Conclusions
The new quinone analogues (3–10) were synthesized by the reaction of 1,4‐NQ (1) with heterocyclic ring‐containing nucleophiles such as thiols and piperazine derivatives in this study. The chemical structures of these newly synthesized compounds were elucidated by FT‐IR, 1H NMR, 13C(APT) NMR, and HRMS. In the past study, some synthetic NQ derivatives showed good antiproliferative profiles and also antibacterial and antioxidant capacity. The strong antimicrobial activity demonstrated by our similar compounds has encouraged us to further our studies on this subject.
The compounds were assessed for their ability to inhibit the growth of different cancer cell lines (MCF‐7, A549, PC‐3) and NIH/3T3 using the MTT assay. Compound 10 exhibited potent antiproliferative effects on various cancer cell lines, notably PC‐3, with minimal cytotoxicity toward NIH/3T3 cells. However, compounds 5 and 7 showed promising activity against A549 cells. These compounds offer the potential to treat cancer, and their different biological effects should be investigated in future studies.
Our molecular docking analysis revealed favorable binding affinities for compound 10, particularly with EGFR, in alignment with its observed in vitro cytotoxicity. Although these findings are promising, they should be interpreted as preliminary insights that require further biological validation to confirm their relevance. These findings corroborate previous studies, which also employed docking methods and reported high‐affinity binding of quinone derivatives to EGFR. The potential interaction between quinone‐based compounds and EGFR has been previously explored in silico. For instance, Nafie et al. reported that quinoline‐based thiazolidinone derivatives exhibited significant binding affinity toward EGFR, suggesting their potential as anticancer agents. Similarly, Mohanty et al. conducted computational studies on quinazoline derivatives, highlighting their favorable interactions with the EGFR active site. These findings align with our docking results, further supporting the hypothesis that quinone derivatives may serve as effective EGFR inhibitors [ref. 59, ref. 60]. Collectively, our findings indicate that quinone derivatives, especially compound 10, may have potential as anticancer agents targeting EGFR; however, further in vitro and in vivo studies are necessary to validate this hypothesis. Further studies are warranted to validate these findings in vivo and explore the molecular mechanisms underlying compound 10’s anticancer activity.
We have investigated the antimicrobial effects of various quinone derivatives, which we synthesized. The findings we have mentioned above are quite important, as the new quinone derivatives that we have synthesized have shown various antimicrobial effects against both important gram‐positive and gram‐negative bacteria, together with antifungal effect. Hence, these compounds show activity against various microorganisms that are human pathogens; considering this antimicrobial activity, our study may have potential for drug development. Accordingly, our study can be quite beneficial because it might shed light for future studies on this area.
Materials and Methods
Chemistry
Melting points were measured by using a Büchi B‐540 melting point apparatus. FT‐IR spectra (cm−1) were recorded as KBr pellets in Nujol mulls on a Shimadzu IR Prestige 21 model Diamond spectrometer (ATR method). 1H NMR and 13C(APT) NMR spectrums were obtained using a Varian Unity Inova (500 MHz) spectrometer by using TMS as the internal standard and deuterated chloroform as the solvent. HRMSs were obtained on an LC‐HRMS Thermo Q Exactive. The synthesized compounds were separated and purified by using column chromatography on silica gel (Fluka Silica gel 60, particle size 63–200 µm). Kieselgel 60 F‐254 plates were used for thin‐layer chromatography (TLC) All chemicals were of reagent grade and were used without further purification. Moisture was excluded from the glass apparatus with CaCl2 drying tubes.
General Method for the Synthesis of Compounds (3–10)
2,3‐Dichloro‐1,4‐NQ (1) was used as a starting compound, and nucleophiles were refluxed in abs. ethanol. The reaction mixture was monitored by TLC until the disappearance of starting material. The reaction mixture was extracted by using 30 mL of CHCl3, washed three times with water, and then dried using Na2SO4. Evaporator system was used to remove the extra amount of solvent. The crude dark residue was separated and purified by column chromatography and dried using a vacuum oven at the end [ref. 61, ref. 62, ref. 63].
The spectral charts are given in the Supporting Information section.
Anticancer Studies of NQ Compounds
Cell Culture
Human lung cancer cell line A549 (ATCC, CCL‐185), human breast cancer cell line MCF‐7 (ATCC, HTB‐22), human prostate cancer cell line PC‐3 (ATCC, CRL‐1435), and mouse embryonic fibroblast cells NIH/3T3 (ATCC, CRL‐1658) were used in this study (Table 1). NIH/3T3, MCF‐7, and A549 cells were cultured in Dulbecco’s Modified Eagle Medium/F12 (DMEM/F12), whereas PC‐3 cells were cultured in RPMI 1640 medium (Gibco, Rockville, MD, USA). All used media contain 10% v/v fetal bovine serum (FBS) and 1% v/v penicillin‐streptomycin (Gibco, Rockville, MD, USA). Cells were grown in a humidified incubator at 37°C with 5% CO2. In our study, DOX, which is one of the most well‐known antineoplastic drugs, was also used as a positive control in this method.
Cell Viability Assay
The MTT assay (Vybrant MTT Cell Proliferation Assay Kit, Thermo Fisher Scientific) was utilized to investigate the antiproliferative and cytotoxic effects of the compounds [ref. 64]. The cells were seeded in a 96‐well plate (1 × 104 cells per well) and incubated for 24 h. The next day, the cells were treated with a 10 µM concentration of compounds for 24 h. At the end of the period, MTT solution was added to each well at a final concentration of 0.5 mg/mL and incubated for an additional 4 h. Then, the purple formazan product was solubilized by adding 100 µL of SDS buffer. Absorption measurements were taken at wavelengths of 570 and 630 nm using a microplate reader (Biotech Instruments, Winooksi, VT, USA). Cell viability percentages were calculated with optical density.
Molecular Docking Studies of NQs
Data, Database, and Tools
To conduct molecular docking studies for the cell lines A549, PC‐3, and MCF‐7 used in this study, target molecules were first determined in light of the literature. Accordingly, two protein targets were determined for each cell line (Table 4) [ref. 65, ref. 66, ref. 67]. Crystal structures of EGFR (EGFR tyrosine kinase), Caspase‐3, VEGFR2, mTOR, FAK, and CDK6 proteins were downloaded from the Protein Data Bank (PDB) (https://www.rcsb.org/). The PDB IDs are as follows, respectively: 1M17, 2XYG, 2OH4, 4JT6, 1MP8, and 3NUP.
TABLE 4: Target proteins identified for molecular docking.
| Target protein | Cancer cell line |
|---|---|
| EGFR | A549 |
| Caspase‐3 | A549 |
| VEGFR2 | MCF‐7 |
| mTOR | MCF‐7 |
| FAK | PC‐3 |
| CDK6 | PC‐3 |
Abbreviations: CDK6, cyclin‐dependent kinase 6; EGFR, epidermal growth factor receptor; FAK, focal adhesion kinase; mTOR, mammalian target of rapamycin; VEGFR2, vascular endothelial growth factor receptor 2.
The structures of the ligands were obtained with Chimera software after converting the image files of the molecules into SMILES format. Proteins and ligands were pre‐prepared for molecular docking with USCF Chimera 1.14 software. Subsequently, docking studies were performed using AutoDock Vina 1.1.2 software to investigate molecular interactions. Biovia Discovery Studio Visualizer software was used for detailed examination and visualization of protein–ligand interactions. In the validation of our molecular docking processes, the Zhang Group web server was used to calculate the RMSD values (https://zhanggroup.org//DockRMSD/).
Molecular Docking Process
PDB was utilized to obtain the crystallographic structures of the proteins associated with cancer (lung, breast, and prostate). For the molecular docking study, the coordinates of the active sites of each protein are given in Table 5, based on the coordinates of the ligand. After obtaining SMILES formats of the ligands using 3D images, their 3D structures were obtained with the Build Structure module in Chimera 1.12 software. Then, the lowest energy conformations were obtained in mol2 format with the Docking Prepare module in Chimera software.
TABLE 5: Active site coordinates of proteins.
| Proteins | PDB ID | X | Y | Z | Grid size |
|---|---|---|---|---|---|
| EGFR | 1M17 | 21.54 | −0.26 | 51.31 | 25 × 25 × 25 |
| Caspase‐3 | 2XYG | 35.8 | 37.75 | 32.75 | 20 × 20 × 20 |
| VEGFR2 | 2OH4 | 2.98 | 33.97 | 16.03 | 26 × 26 × 26 |
| mTOR | 4JT6 | 49.51 | 1.48 | −47.14 | 20 × 20 × 20 |
| FAK | 1MP8 | 35.55 | −4.43 | 25.06 | 20 × 20 × 20 |
| CDK6 | 3NUP | 22.14 | 34.94 | −7.51 | 20 × 20 × 20 |
Abbreviations: CDK6, cyclin‐dependent kinase 6; EGFR, epidermal growth factor receptor; FAK, focal adhesion kinase; mTOR, mammalian target of rapamycin; VEGFR2, vascular endothelial growth factor receptor 2.
Molecular docking energy calculations were made according to the standard protocol of AutoDock Vina 1.1.2 software. The best energy from 10 different bindings was obtained as a result of these processes and recorded in PDB format. Biovia Discovery Studio Visualizer software was used for more detailed protein–ligand interactions on the structure with the best binding energy.
Determination of Antimicrobial Activity of Quinone Derivatives
Antimicrobial Activity Test
In order to find out the antimicrobial activity potential of quinone derivatives the agar well diffusion method was applied. Additionally, MICs were found for quinone derivatives. Later, moreover, the minimal bactericidal concentrations (MBCs) and minimal fungicidal concentrations (MFCs) of the derivatives were assessed.
Agar Well Diffusion Test
In our study, S. aureus ATCC 25923, S. aureus ATCC 43300, S. epidermidis ATCC 11228, Enterococcus faecalis ATCC 29212, P. aeruginosa ATCC 27853, E. coli ATCC 25922, and K. pneumoniae ATCC 4352 were the bacteria, and also C. albicans ATCC 90028 was the yeast used. All of the bacteria were inoculated on the surface of Mueller–Hinton agar (MHA), left for incubation at 37°C for 24 h, and Sabouraud dextrose agar (SDA) was used for C. albicans, left for incubation at 35°C for 48 h. After that the formed colonies suspensions of bacteria were adjusted to 108 cfu/mL and those of C. albicans to 106 cfu/mL in accordance with McFarland 0.5 standard turbidity by using 0.85% NaCl physiological saline solution (PSS). The prepared bacterial suspensions were spread on the surface of the MHA by using sterile swabs and yeast suspension on the surface of SDA, and then by using sterile punch, a 5 mm diameter wells were opened on the surface of the media. Then 50 µL of the extracts dissolved in their solvents were put into the wells. While as a positive control for bacteria meropenem and for the yeast amphotericin were assessed, as negative controls solvent (DMSO) and PSS were appraised. After incubation of the bacteria‐seeded plates at 37°C for 18–24 h and yeast‐seeded ones at 35°C for 24–48 h, the formed inhibition zones were measured and expressed in mm. All of the experiments were done in a triplicate average range [ref. 56, ref. 68, ref. 69].
Assessment of MIC for Bacteria
In our study, the assessment of MIC for bacteria was done in accordance with the standards of the Clinical and Laboratory Standards Institute (CLSI) [ref. 70]. Here, although meropenem was used as positive control, CAMHB and DMSO were used as negative control [ref. 70].
Assessment of MIC for Yeasts
The assessment of MIC for yeast was done in accordance with the standards of the CLSI [ref. 71]. Here, although Amphotericin B was used as positive control, RPMI and DMSO were used as negative control [ref. 71].
Assessment of MBC and MFC
In order to assess the MIC and MFC values of the quinone derivatives, inoculation to the appropriate plates was done from each well of the microplate. Then the plates were left to incubate at 37°C for 24 h for bacteria and 48 h for the yeast. Finally, MBC and MFC values were determined according to the lowest values where no growth was observed [ref. 72].
Author Contributions
Berrin Yilmaz: methodology, software, formal analysis. Nahide Gulsah Deniz: conceptualization, investigation, supervision, writing – original draft. Cigdem Sayil: conceptualization, investigation, supervision, writing – original draft. Ozlem Bingol Ozakpinar: data curation, formal analysis, software. Merve Gurboga: data curation, formal analysis, software. Turgut Sekerler: formal analysis, software. Pervin Rayaman: data curation, formal analysis, software. Erkan Rayaman: formal analysis, software. Elif Caliskan Salihi: data curation, formal analysis, software, writing – review and editing, supervision.
Conflicts of Interest
The authors declare no conflicts of interest.
Supplementary Materials
References
- Anticancer Drug Discovery From Iranian Chrysanthemum Cultivars Through System Pharmacology Exploration and Experimental Validation,”. Scientific Reports, 2021. [DOI | PubMed]
- Pentacyclic Triterpenoids With Nitrogen‐Containing Heterocyclic Moiety, Privileged Hybrids in Anticancer Drug Discovery,”. Molecules (Basel, Switzerland), 2021. [DOI | PubMed]
- Menadione: Role in Cancer Prevention and Methods of Analysis,”. World Journal of Pharmaceutical Sciences, 2014
- Vitamin K: A Novel Cancer Chemosensitizer,”. Biotechnology and Applied Biochemistry, 2022. [DOI | PubMed]
- Amygdalin as Multi‐Target Anticancer Drug Against Targets of Cell Division Cycle: Double Docking and Molecular Dynamics Simulation,”. Journal of Biomolecular Structure and Dynamics, 2021. [DOI | PubMed]
- Primary Systemic Therapy in HER2‐Positive Operable Breast Cancer Using Trastuzumab and Chemotherapy: Efficacy Data, Cardiotoxicity and Long‐Term Follow‐Up in 142 Patients Diagnosed From 2005 to 2016 at a Single Institution,”. Breast Cancer (Tokyo, Japan), 2018. [DOI]
- First‐Line Therapy for Metastatic Soft Tissue Sarcoma,”. Current Treatment Options in Oncology, 2019. [DOI]
- The Diverse Mechanisms and Anticancer Potential of Naphthoquinones,”. Cancer Cell International, 2019. [DOI | PubMed]
- Anticancer Activities of Vitamin K3 Analogues,”. Investigational New Drugs, 2020. [DOI | PubMed]
- Quinone‐Induced Oxidative Stress Elevates Glutathione and Induces Gamma‐Glutamylcysteine Synthetase Activity in Rat Lung Epithelial L2 Cells,”. Journal of Biological Chemistry, 1994. [DOI | PubMed]
- DNA Strand Scission and Free Radical Production in Menadione‐Treated Cells. Correlation With Cytotoxicity and Role of NADPH Quinone Acceptor Oxidoreductase,”. Journal of Biological Chemistry, 1992. [DOI | PubMed]
- Quinone‐Induced DNA Single Strand Breaks in a Human Colon Carcinoma Cell Line,”. Carcinogenesis, 1997. [DOI | PubMed]
- An Approach to Aminonaphthoquinone Ansamycins Using a Modified Danishefsky Diene,”. Organic Letters, 2012. [DOI | PubMed]
- Two Approaches to the Aromatic Core of the Aminonaphthoquinone Antibiotics,”. Journal of Organic Chemistry, 2013. [DOI | PubMed]
- Vitamin K: Redox‐Modulation, Prevention of Mitochondrial Dysfunction and Anticancer Effect,”. Redox Biology, 2018. [DOI | PubMed]
- 2‐Phenylaminonaphthoquinones and Related Compounds: Synthesis, Trypanocidal and Cytotoxic Activities,”. Bioorganic & Medicinal Chemistry, 2014. [DOI | PubMed]
- C–H Trifluoromethylation of 2‐Substituted/Unsubstituted Aminonaphthoquinones at Room Temperature With Bench‐Stable (CF3SO2)2Zn: Synthesis and Antiproliferative Evaluation,”. Journal of Organic Chemistry, 2017. [DOI | PubMed]
- Biological Evaluation of Donor‐Acceptor Aminonaphthoquinones as Antitumor Agents,”. European Journal of Medicinal Chemistry, 2010. [DOI | PubMed]
- Quinonoid Compounds via Reactions of Lawsone and 2‐Aminonaphthoquinone With α‐Bromonitroalkenes and Nitroallylic Acetates: Structural Diversity by C‐Ring Modification and Cytotoxic Evaluation Against Cancer Cells,”. European Journal of Medicinal Chemistry, 2018. [DOI | PubMed]
- Synthesis and Characterization of n‐Alkylamino Derivatives of Vitamin K3: Molecular Structure of 2‐Propylamino‐3‐Methyl‐1,4‐Naphthoquinone and Antibacterial Activities,”. Journal of Molecular Structure, 2015. [DOI]
- The Synthesis of Novel Mono(alkoxy)‐, Tris(thio)‐ and Tetrakis(thio)‐Substituted Quinones From the Reactions of p‐Chloranil With Various S‐Nucleophiles,”. Bulletin of the Korean Chemical Society, 2009. [DOI]
- Synthesis and Investigation of Antimicrobial and Antioxidant Activity of Anthraquinonylhydrazones,”. Monatshefte Für Chemie—Chemical Monthly, 2018. [DOI]
- Synthesis and Spectral Properties of Novel Thionaphtoquinone Dyes,”. Bulletin of the Korean Chemical Society, 2010. [DOI]
- Synthesis and Characterization of Nitrogen and Sulfur Containing 1,4‐Naphthoquinones,”. Phosphorus, Sulfur, and Silicon and Related Elements, 2013. [DOI]
- Synthesis and Cyclization Reactions of Novel Benzo[a]phenazine‐ and Phenoxazine-5‐Ones Derivatives,”. Bulgarian Chemical Communications, 2016
- Synthesis, Antimicrobial Properties, and Inhibition of Catalase Activity of 1,4‐Naphtho‐ and Benzoquinone Derivatives Containing N‐, S‐, O‐Substituted,”. Heteroatom Chemistry, 2019. [DOI]
- Synthesis and Characterization of New Naphtho‐and Tetracyclic Diazaquinone Derivatives,”. Journal of the Chemical Society of Pakistan, 2019
- New Vitamin K3 (Menadione) Analogues: Synthesis, Characterization, Antioxidant and Catalase Inhibition Activities,”. Journal of Chemical Sciences, 2020. [DOI]
- Synthesis and Antiproliferative Evaluation of Some 1,4‐Naphthoquinone Derivatives Against Human Cervical Cancer Cells,”. Open Chemistry, 2019. [DOI]
- Nucleophilic Reactions of Quinones,”. Russian Chemical Reviews, 1991. [DOI]
- Design, Synthesis, Biological Evaluation, and Antioxidant and Cytotoxic Activity of Heteroatom‐Substituted 1,4‐Naphtho‐ and Benzoquinones,”. Chemical & Pharmaceutical Bulletin, 2015. [DOI | PubMed]
- 1,1′‐Bis(Diphenylphosphino)Ferrocene]Dichloropalladium/1,1′‐Bis(Diphenylphosphino)Ferrocene Catalyzed Synthesis of 2,3‐Diamino‐1,4‐Naphthoquinones,”. Synthesis (Stuttg), 2007. [DOI]
- Facile Synthesis of 2‐Amino‐1,4‐Naphthoquinones Catalyzed by Molecular Iodine Under Ultrasonic Irradiation,”. Synthetic Communications, 2008. [DOI]
- Novel Amino‐ and Thio(Substituted)‐1,4‐Naphthoquinone (NQ) Compounds: Synthesis and Characterization,”. Phosphorus, Sulfur, and Silicon and Related Elements, 2020. [DOI]
- Synthesis, Characterization of N‐, S‐, O‐Substituted Naphtho‐ and Benzoquinones and a Structural Study,”. Journal of Chemical Sciences, 2012. [DOI]
- Methoxy‐Enriched Cationic Stilbenes as Anticancer Therapeutics,”. Bioorganic Chemistry, 2020. [DOI | PubMed]
- Synthesis and Biological Evaluation of Naphthalene‐1,4‐Dione Derivatives as Potent Antimycobacterial Agents,”. Medicinal Chemistry (Los Angeles), 2008. [DOI]
- Rapid Colorimetric Assay for Cellular Growth and Survival: Application to Proliferation and Cytotoxicity Assays,”. Journal of Immunological Methods, 1983. [DOI | PubMed]
- Viscum album L. Extracts Protects HeLa Cells Against Nuclear and Mitochondrial DNA Damage,”. Evidence‐Based Complementary and Alternative Medicine, 2012. [DOI | PubMed]
- Green Chemistry Approach Towards Piperazine: Anticancer Agents,”. Journal of Molecular Structure, 2023. [DOI]
- N‐Substituted Piperazinopyridylsteroid Derivatives as Abiraterone Analogues Inhibit Growth and Induce Pro‐Apoptosis in Human Hormone‐Independent Prostate Cancer Cell Lines,”. Chemical Biology & Drug Design, 2013. [DOI | PubMed]
- Novel Piperazine Derivatives of Vindoline as Anticancer Agents,”. International Journal of Molecular Sciences, 2024. [DOI | PubMed]
- Epidermal Growth Factor Receptor Dual‐Target Inhibitors as a Novel Therapy for Cancer: A Review,”. International Journal of Biological Macromolecules, 2023. [DOI | PubMed]
- To Investigate Growth Factor Receptor Targets and Generate Cancer Targeting Inhibitors,”. Current Topics in Medicinal Chemistry, 2023. [DOI | PubMed]
- Targeting a Key FAK‐Tor: The Therapeutic Potential of Combining Focal Adhesion Kinase (FAK) Inhibitors and Chemotherapy for Chemoresistant Non‐Small Cell Lung Cancer,”. Expert Opinion on Investigational Drugs, 2024. [DOI | PubMed]
- EGFR Over‐Expression in Non‐Small Cell Lung Cancers Harboring EGFR Mutations Is Associated With Marked Down‐Regulation of CD82,”. Biochimica et Biophysica Acta, 2015. [DOI | PubMed]
- Design and Synthesis of Some New 2,4,6‐Trisubstituted Quinazoline EGFR Inhibitors as Targeted Anticancer Agents,”. Bioorganic Chemistry, 2020. [DOI | PubMed]
- The Diverse Mechanisms and Anticancer Potential of Naphthoquinones,”. Cancer Cell International, 2019. [DOI | PubMed]
- Discovery of Anilino‐1,4‐Naphthoquinones as Potent EGFR Tyrosine Kinase Inhibitors: Synthesis, Biological Evaluation, and Comprehensive Molecular Modeling,”. ACS Omega, 2022. [DOI | PubMed]
- Epidermal Growth Factor Receptor (EGFR) Signaling in Cancer,”. Gene, 2006. [DOI | PubMed]
- Targeting EGFR/PI3K/AKT/mTOR Signaling in Lung and Colon Cancers: Synthesis, Antitumor Evaluation of New 1,2,4‐Oxdiazoles Tethered 1,2,3‐Triazoles,”. RSC Advances, 2024. [DOI | PubMed]
- Role of PI3K/AKT/mTOR Signaling in the Cell Cycle Progression of Human Prostate Cancer,”. Biochemical and Biophysical Research Communications, 2003. [DOI | PubMed]
- TGF‐β Effects on Prostate Cancer Cell Migration and Invasion Are Mediated by PGE2 Through Activation of PI3K/AKT/mTOR Pathway,”. Endocrinology, 2013. [DOI | PubMed]
- Anti‐Acne Activities of Pulsaquinone, Hydropulsaquinone, and Structurally Related 1,4‐Quinone Derivatives,”. Archives of Pharmaceutical Research, 2009. [DOI]
- Antifungal Susceptibility Testing: Current Approaches,”. Clinical Microbiology Reviews, 2020. [DOI | PubMed]
- Methods for In Vitro Evaluating Antimicrobial Activity: A Review,”. Journal of Pharmaceutical Analysis, 2016. [DOI | PubMed]
- Lawsone, Juglone, and β‐Lapachone Derivatives With Enhanced Mitochondrial‐Based Toxicity,”. ACS Chemical Biology, 2018. [DOI | PubMed]
- The Mechanism of Action Underlying Antibacterial Activity of a Diterpene Quinone Derivative Against Staphylococcus aureus Through the In Vitro and In Silico Assays,”. Pharmacy Education, 2024. [DOI]
- Deciphering Quinazoline Derivatives’ Interactions With EGFR: A Computational Quest for Advanced Cancer Therapy Through 3D‐QSAR, Virtual Screening, and MD Simulations,”. Frontiers in Pharmacology, 2024. [DOI | PubMed]
- Quinoline‐Based Thiazolidinone Derivatives as Potent Cytotoxic and Apoptosis‐Inducing Agents Through EGFR Inhibition,”. Chemical Biology & Drug Design, 2022. [DOI | PubMed]
- Synthesis and Biological Evaluation of Alkyl/Arylamino Derivatives of Naphthalene‐1,4‐Dione as Antimycobacterial Agents,”. Der Pharma Chemica, 2010
- Light‐Driven Carbon–Carbon Coupling of α‐sp3 –CH of Aliphatic Alcohols With sp2 –CH Bond of 1,4‐Naphthoquinones,”. Organic Letters, 2022. [DOI | PubMed]
- The Preparation of Alkylthio Substituted Naphthoquinones‐1,4 and Benzoquinones‐1,4,”. Recueil Des Travaux Chimiques Des Pays‐Bas, 1952. [DOI]
- Stereochemistry‐Dependent Cytotoxicity of Some Artemisinin Derivatives,”. Journal of Natural Products, 1997. [DOI | PubMed]
- Molecular Docking as a Tool for the Discovery of Molecular Targets of Nutraceuticals in Diseases Management,”. Scientific Reports, 2023. [DOI | PubMed]
- CDK6 Is Upregulated and May be a Potential Therapeutic Target in Enzalutamide‐Resistant Castration‐Resistant Prostate Cancer,”. European Journal of Medical Research, 2022. [DOI | PubMed]
- Synthesis, ADMET Predictions, Molecular Docking Studies, and In‐Vitro Anticancer Activity of Some Benzoxazines Against A549 Human Lung Cancer Cells,”. Journal of Basic and Clinical Physiology and Pharmacology, 2021. [DOI | PubMed]
- An Antibiotic Assay by Agar Well Diffusion Method,”. Acta Biologiae Et Medicinae Experimentalis, 1990
- Antifungal Susceptibility Testing,”. Clinical Microbiology Reviews, 1993. [DOI]
- 70 Clinical and Laboratory Standard Institute (CLSI) , Performance Standards for Anti‐Microbial Susceptibility Testing. 30th Ed., CLSI supplement M100, Clinical and Laboratory Standard Institute, 2020.
- 71 Clinical and Laboratory Standard Institute (CLSI) , M27M44S Performance Standards for Antifungal Susceptibility Testing of Yeasts, 3rd Ed., CLSI supplement M27M44S, Clinical and Laboratory Standard Institute (CLSI), 2022.
- Antibacterial Activity of Doxycycline‐Loaded Nanoparticles,”. Methods in Enzymology, 2012. [DOI | PubMed]
