A Small Cellulose-Binding-Domain Protein (CBD1) in Phytophthora is Highly Variable in the Non-binding Amino Terminus
Abstract
Electronic supplementary material:
The online version of this article (doi:10.1007/s00284-017-1315-x) contains supplementary material, which is available to authorized users.
Affiliations: 0004 0404 0958grid.463419.dGenetic Improvement of Fruits and Vegetables Laboratory, USDA-ARS, 10300 Baltimore Avenue, Beltsville, MD 20705 USA
License: © The Author(s) 2017 CC BY 4.0 Open AccessThis article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
Article links: DOI: 10.1007/s00284-017-1315-x | PubMed: 28748272 | PMC: PMC5640731
Relevance: Relevant: mentioned in keywords or abstract
Full text: PDF (56 KB)
Introduction
Cellulose-binding domains (CBD), found within the larger class of protein structural patterns known as carbohydrate binding modules (CBM), facilitate binding to target substrates. Fungal saprophyte-encoded CBDs are commonly found attached, by a serine, threonine-rich linker, to the amino or carboxy-terminus of extracellular, cellulolytic, and xylanolytic enzymes [ref. 3, ref. 22, ref. 24, ref. 26]. Pathogenic fungi generally do not have CBDs associated with their cellulolytic enzymes unless the enzymes are deployed during a necrotrophic phase [ref. 40–ref. 42]. In addition to aiding the binding of enzymes, the CBDs can have the ability to dissociate cellulose microfibrils independently, as shown with CBDs produced in protein expression systems [ref. 9, ref. 15, ref. 34].
Phytophthora species are hemibiotrophic plant pathogens that also generally lack a CBD on their cellulolytic enzymes [ref. 6, ref. 7, ref. 32]; however, they are one of the few organisms that produce CBDs as discrete proteins [ref. 12, ref. 18, ref. 28]. The Phytophthora CBD-containing proteins generally harbor the binding domains at the carboxy terminal region of the protein, while the function of the amino half of the proteins remains undefined. Most likely, these proteins are involved in the development of the cellulose-containing cell walls of Phytophthora, and the undefined regions may interact with other carbohydrates or with other proteins in the cell wall. There is currently evidence for expression of three types of proteins, harboring a single CBD (CBD1), two CBDs (CBD4), and two CBDs plus a lectin-binding domain (CBD5 or CBEL) while other genomic sequences harboring CBD motifs have not yet been shown to be expressed [ref. 18]. The CBD1 protein from P. infestans has been shown to be cell-wall-bound, and it has been shown to bind to plant cell walls when expressed in transgenic potato plants [ref. 19]. In transgenic potatoes, the binding alters the expansion of ovary cell walls, allowing for fertility restoration. The degree of divergence in CBD sequence that would still allow or further enhance this response remains to be studied.
While many extracellular proteins from Phytophthora are extensively duplicated [ref. 6, ref. 7, ref. 32], the CBD-encoding genes are single copy. The single copy feature, along with providing a critical function in cell walls, suggests limited variation; however, there have not been any studies addressing the degree of conservation in amino acid sequence of CBDs within and between species of Phytophthora. We have focused on CBD1 and compared conservation of sequence in a worldwide collection of P. infestans (USDA-ARS, Beltsville, MD). Conservation of sequence was also assessed for a divergent set of Phytophthora species, and the related biotrophic downy mildew Plasmopara halstedii. Application of this information was made in development of CBD1-specific primers for Phytophthora detection.
Materials and Methods
Isolation and Identification of Sequences Encoding CBD 1
Total genomic DNA was isolated from liquid grown cultures of Phytophthora infestans and P. mirabilis using a Qiagen Plant DNeasy kit. Preliminary screening indicated that the CBD1-encoding gene (EU179903) could be amplified from different isolates of P. infestans using the primer pair CBD1f (ATGACCTCGTTGCGACTCCTGG) and CBD1r (CTAGAGCTCCAGTCGAATGAC), amplifying the full coding region. The PCR reaction mix consisted of 20 pmol primer, 50 ng DNA, and GoTaq polymerase (Promega) in a total volume of 40 microliters. The PCR amplification consisted of one cycle of 96 C for 2 min, followed by 30 cycles of 52 C for 15 s and 72 C for 45 s, then a final cycle of 72 C for 5 min.
The amplified products were cloned using a TOPO TA cloning kit (Invitrogen). Sequencing was performed commercially (Macrogen USA) using M13 forward and reverse primers. Overall, single colonies were used for sequencing since the purpose of the study was to show general variations in CBD 1, using a large number of individual isolates (Online Resource 1). Allelic variation was not determined for each isolate where CBD1 was cloned from, and allelic variation could not be determined when using GenBank sequences.
For in silico identification of CBD1 proteins from other Phytophthora species and Plasmopara, we conducted tblastn searches of NCBI databases using the specific CBD1 Phytophthora cellulose-binding domain sequence (GVRAWAQCGGLYYLGKTKCQQHTFCKQLSEFISVC).
Alignments and Phylogeny
Nucleotide sequences were translated using WebMap (pga.mgh.harvard.edu) and signal peptides were identified using SignalP 4.1 (cbs.dtu.dk/services) [ref. 33]. Nucleotide and amino acid sequences were compared after removal of the signal peptide sequence. Alignment was performed using Clustal Omega 1.2.2 (ebi.ac.uk/Tools/msa/clustalo) [ref. 37] and amino acid-based phylogeny was assessed using Geneious 8.1.7 (geneious.com) [ref. 20]. Glycosylation patterns were assessed using NetNGlyc 1.0 and DictyOGlyc 1.1 (cbs.dtu.dk/services) [ref. 13]. Amino acid-based phylogeny was generated using MEGA6.0 [ref. 10, ref. 35, ref. 38].
Application of CBD 1 Primers for Species Detection and Identification
For PCR detection and species discrimination, primers used included P. infestans forward (PiF TCTAACCTCCGGAACGGCGAC), P. infestans reverse (PiR TAGAGCTCCAGTCGAATGACT), P. sojae forward (PsF TCCAACCTCCGAAACTCGATCCATC), P. sojae reverse (PsR TACAGCTCGAGCCGGATGACC), and CBD reverse-common (CBD-Rc GTGTGCTGCTGGCACTTGGTCT). Primer specificity was tested using 100 ng of target DNA and 10 pg of each primer. Species discrimination was assessed using P. infestans specific primers with P. sojae DNA and P. sojae primers with P. infestans DNA. The PCR amplification consisted of one cycle of 96 C for 2 min, followed by 30 cycles of 54 C (PiF plus PiR or PiF plus CBD-Rc), or 60 C (PsF plus PsR or PsF plus CBD-Rc) for 15 s and 72 C for 45 s, then a final cycle of 72 C for 5 min, using a Techne T-312 heated lid thermal cycler. Levels of detectable DNA were assessed by diluting P. infestans DNA (300 ng per microliter) to concentrations as low as 1 pg. An initial amplification used the primer pair PiF and PiR in 40 µl reactions. One microliter of the first amplification was used in a 40 µl second round amplification using the primer pair PiF and CBD-Rc. Samples were separated in 1% agarose gels. A comparison between CBD1 primers and Ypt1 primers was made using 10 pg of DNA from P. sojae and P. infestans.
Results
Isolation and Identification of Sequences Encoding CBD 1
Nucleotide alignments reveal the mature protein-coding regions that can be used in designing primers for detection of P. infestans using CBD1 (Fig. 1).

Alignments and Phylogeny
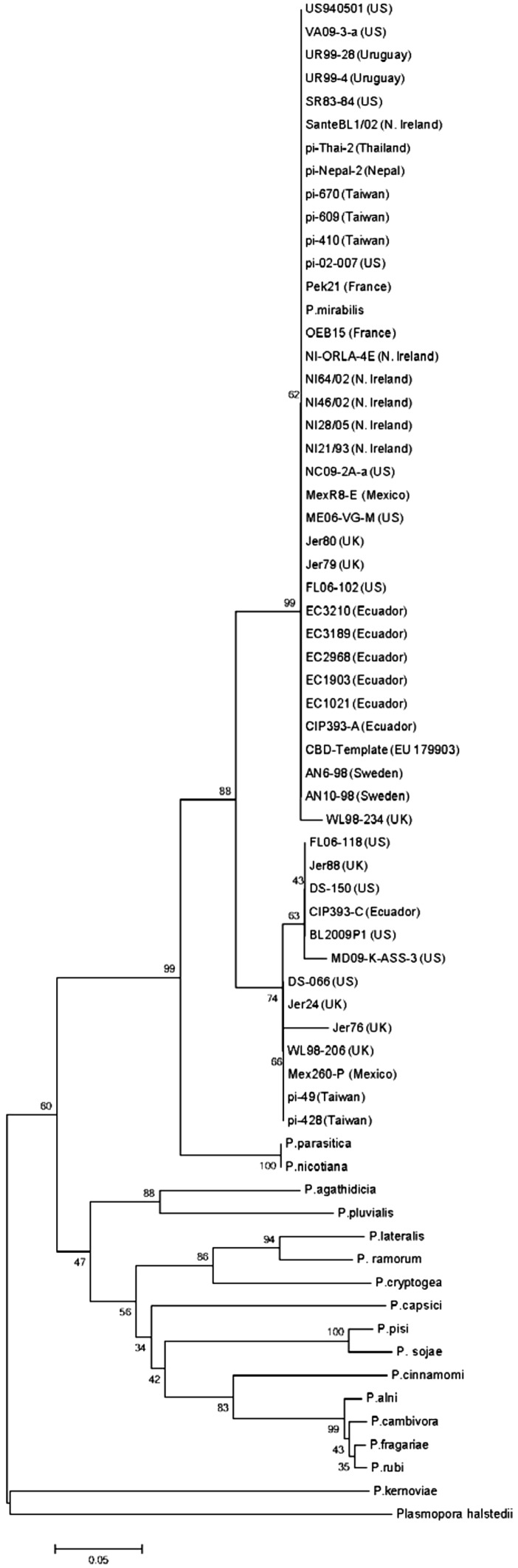
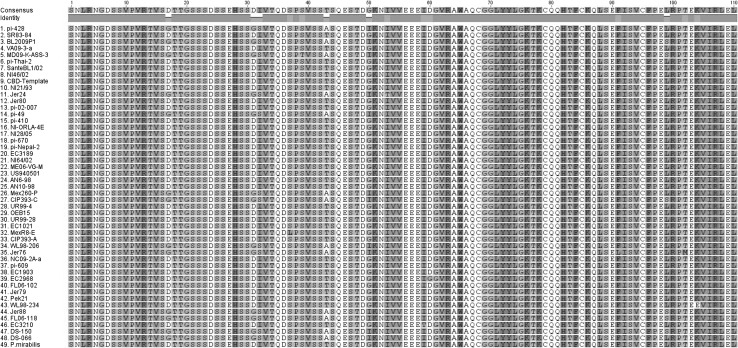
While the P. infestans CBD1 amino acid sequence was highly conserved at the cellulose-binding-domain-containing carboxy half of the protein, there were a limited number of substitutions in the amino half of the secreted protein (Fig. 2), but these were mostly synonymous. Clear differences were seen in the amino terminus of the different Phytophthora species as well as Plasmopara (Fig. 3). Interestingly the carboxy terminal half was conserved up to the last amino acid in the protein. Distinct groups of isolates within P. infestans, and between Phytophthora species could be seen when assessing phylogeny based on amino acid sequences (Fig. 4).


Application of CBD 1 Primers for Species Detection and Identification
Primer sets based on CBD1 sequence were able to discriminate between P. infestans and P. sojae. Primers targeting the full length sequence of the CBD-encoding gene, as well as a reverse primer within the conserved cellulose-binding domain allowed for species identification. Application of the terminal sequence primers followed by a nested reaction with the same forward primer and the cellulose-binding domain reverse primer allowed for the detection of P. infestans DNA concentrations of one picogram (Online Resource 2). When comparing CBD1 primers with Ypt1, the Ypt1 primers and the PiF CBD-Rc combination were able to detect 10 pg of DNA in a single reaction (Online Resource 3). The Ypt1 primers produced a stronger signal, but did not discriminate between P. infestans and P. sojae. Nested primers would be required to detect CBD1 as efficiently as Ypt1 using the primers from this study.
Discussion
The highest degree of similarity within the Phytophthora infestans CBD 1 amino acid sequence was centered on the carboxyl half of the protein where the specific amino acids involved in cellulose binding are located. There were also a very limited number of differences on the amino terminal half of the protein, suggesting that region of the protein plays an important, but as yet undefined role. Variations in the amino terminal regions were random, except for a small group of P. infestans isolates that had an elevated level of amino acid substitution (5 aa changes). This group of isolates (FL06-118-US, Jer88-UK, DS150-US, CIP393C-Ecuador, BL2009P1-US, MD09KASS3-US, DS066-US, Jer24-UK, WL98206-UK, Mex260P-Mexico, PI49-Taiwan, and PI428-Taiwan) had an identical set of substitutions (D17/G, D31/G, I32/S, T43/A, and G50/A), suggesting that an original isolate was distributed to other countries, possibly through infected plant material. Sequence variations in CBD1 may prove a simple, useful tool to track the origins of other Phytophthora isolates.
At the species level the cellulose-binding domain continued to be conserved at the carboxy half of the protein, while the amino terminus was subject to larger variations occurring through insertions, deletions, and substitutions. If the amino terminal region interacts with other protein or carbohydrate components of the cell wall, then it is possible that CBD1 variations reflect the need to interact with the differing wall compositions found in different species [ref. 39, ref. 43]. A large deletion (22 aa) is seen in Plasmopara, which differs from Phytophthora in being a strict biotroph. The presence of CBD1 in the downy mildew Plasmopara is the first evidence for this protein in downy mildews. The amino terminal region differs between Phytophthora and Plasmopara in potential for glycosylation. There are two predicted N-glycosylation sites (N8 and N13) in the Plasmopara CBD1-secreted protein and none in the Phytophthora-secreted CBD1, while there is one predicted O-glycosylation site (S8) in Phytophthora and none in Plasmopara. Glycosylation can allow for additional sites of carbohydrate interactions [ref. 14].
There are a wide range of gene targets for detecting and identification of Phytophthora species, including ITS regions, SSRs, and gene-specific primers [ref. 17, ref. 21]. Primers spanning the introns of the vesicle transport-encoding Ypt1 gene were shown to be a reliable and efficient way to detect P. nicotianae, and these primers could also be used to distinguish some species of Phytophthora [ref. 5, ref. 8, ref. 30]. We were able to apply CBD1 nucleotide sequence information to design primers for the detection of P. infestans and P. sojae. One primer set was used to amplify the region of the CBD1 gene encoding the full secreted protein product. The second set that was tested amplified the genomic region from the beginning of the secreted protein to the highly conserved cellulose-binding domain region. Since this region is highly conserved, we used a common reverse primer that could be used for many different species of Phytophthora. The fact that Phytophthora contains at least three CBD proteins with a total of five cellulose-binding domains did not interfere with the use of the common domain primer sequence used from CBD1. While Phytophthora CBD-encoding genes are single copy, they are highly expressed during infection [ref. 1, ref. 2]. Detection protocols based on RT-PCR could utilize CBD1-targeted primers and take advantage of the high transcript levels present in infected tissues.
In addition to their widespread occurrence in stramenopiles, cellulose-binding domains are found in non-enzymatic proteins of other organisms including yeast, slime molds, diatoms, and algae [ref. 18, ref. 27]. In the case of Phytophthora, there is no evidence that the CBD-containing proteins move beyond the cell wall. They are not an abundant protein in cell wall proteomic analyses [ref. 29] yet they are readily detected by immunolocalization [ref. 18], suggesting that proteomic analysis may be missing tightly bound wall proteins. Cellulose-binding domains are of interest in biotechnological applications that include cellulose bioprocessing [ref. 4, ref. 36, ref. 44], protein purification tags [ref. 25, ref. 31], and as modulators of plant growth [ref. 19, ref. 23]. Information on the native variations within the cellulose-binding domain can provide insight into the degree of substitutions that can be generated without compromising function [ref. 16]. In a random selection of clones from five isolates we did not find allelic variation in CBD1. It is possible that allelic variants occur in CBD1, and this information may also add to our knowledge on the range of variation allowed without compromising function [ref. 11].
Supplementary Materials
- Online Resource 1. Identification of Phytophthora infestans isolate origins, and GenBank accession numbers for Phytophthora species and Plasmopara gene identification. Supplementary material 1 (PDF 37 kb) (PDF)
- Online Resource 2. PCR amplification of P. infestans and P. sojae DNA using CBD1 gene primers. Lane 1. Primers PiF and PiR plus 100 pg P. infestans DNA. Lane2. Primers PiF and CBD-Rc plus 100 pg P. infestans DNA. Lane 3. Primers PiF and PiR plus 100 pg P. sojae DNA. Lane 4. Primers PiF and CBD-Rc plus 100 pg P. sojae DNA. Lane 5. Primers PsF and PsR plus 100 pg P. sojae DNA. Lane 6. Primers PsF and CBD-Rc plus 100 pg P. sojae DNA Lane 7. Primers PsF and PsR plus 100 pg P. infestans DNA. Lane 8. Primers PsF and CBD-Rc plus 100 pg P. infestans DNA. Supplementary material 2 (PDF 56 kb) (PDF)
- Online Resource 3. Comparison of CBD1 and Ypt1 primers for detection of 10 pg of P. infestans (Pi) or P. sojae (Ps) DNA using PCR amplification. Primers used for CBD1 were PiF (TCTAACCTCCGGCGAC), PiR (TAGAGCTCCAGTCGAATGACT) PsF (TCCAACCTCCGAAACTCGATCCATC), PsR (TACAGCTCGAGCCGGATGACC) and CBD-Rc (GTGTGCTGCTGGCACTTGGTCT), which is a conserved region common across species of Phytophthora. The ras-related protein gene was targeted using Ypt1F (GACACGTACACGGAGAGCTACATCTCGACCAT) and Ypt1R (GTGCGGAAACGCTCCTGGCCGGC), based on common sequence from P. infestans (GenBank U30474) and P. sojae (GenBank XM_009517214). Lane 1. PiF and CBD-Rc. Lane 2. YptF and YptR. Lane 3. PsF and CBD-Rc. Lane 4. YptF and YptR. Lane 5. PiF and PiR. Lane 6. PsF and PsR. Supplementary material 3 (PDF 94 kb) (PDF)
References
- A Attard, E Evangelisti, N KebdanMinet, F Panabieres, E Deleury, C Maggio, M Ponchet, M Gourgues. Transcriptome dynamics of Arabidopsis thaliana root penetration by the oomycete pathogen Phytophthora parasitica. BMC Genom, 2014. [DOI]
- LM Blackman, DP Cullerne, P Torreña, J Taylor, AR Hardham. RNA-Seq analysis of the expression of genes encoding cell wall degrading enzymes during infection of lupin (Lupinus angustifolius) by Phytophthora parasitica. PLoS ONE, 2015. [DOI | PubMed]
- AB Boraston, DN Bolam, HJ Gilbert, GJ Davies. Carbohydrate-binding modules: fine tuning polysaccharide recognition. Biochem J, 2004. [DOI | PubMed]
- G Carrard, A Koivula, H Soderlund, P Beguin. Cellulose-binding domains promote hydrolysis of different sites on crystalline cellulose. Proc Natl Acad Sci, 2000. [DOI | PubMed]
- Y Chen, R Roxby. Characterization of a Phytophthora infestans gene involved in vesicle transport. Gene, 1996. [DOI | PubMed]
- S Costanzo, MD Ospina-Giraldo, K Deahl, C Baker, RW Jones. Gene duplication event in family 12 glycosyl hydrolase from Phytophthora spp. Fungal Genet Biol, 2006. [DOI | PubMed]
- S Costanzo, MD Ospina-Giraldo, KL Deahl, CJ Baker, RW Jones. Alternate intron processing of family 5 endoglucanase transcripts from the genus Phytophthora. Curr Genet, 2007. [DOI | PubMed]
- AK Das, SS Bawage, SG Nerkar, A Kumar. Detection of Phytophthora nicotianae in water used for irrigating citrus trees by Ypt1 gene based nested PCR. Indian Phytopathol, 2013
- N Din, NR Gilkes, B Tekant, RC Miller, AJ Warren, DG Kilburn. Non-hydrolytic disruption of cellulose fibers by the binding domain of a bacterial cellulase. Bio-Technol, 1991. [DOI]
- J Felsenstein. Confidence limits on phylogenies: an approach using the bootstrap. Evolution, 1985. [DOI | PubMed]
- M-C Gagnon, M-J Bergeron, RC Hamelin, NJ Grunwald, GJ Bilodeau. Real-time PCR assay to distinguish Phytophthora ramorum lineages using the cellulose binding elicitor lectin (CBEL) locus. Can J Plant Pathol, 2014. [DOI]
- E Gaulin. cellulose-binding domains of a Phytophthora cell wall protein are novel pathogen-associated molecular patterns. Plant Cell, 2006. [DOI | PubMed]
- R Gupta, E Jung, AA Gooley, KL Williams, S Brunak, J Hansen. Scanning the available Dictyostelium discoideum proteome for O-linked GlcNAc glycosylation sites using neural networks. Glycobiology, 1999. [DOI | PubMed]
- R Happs, X Guan, M Resch, M Davis, G Beckham, Z Tan, M Crowley. O-glycosylation effects family 1 carbohydrate-binding module solution structures. FEBS J, 2015. [DOI | PubMed]
- C Herve, A Rogowski, AW Blake, SE Marcus, HJ Gilbert, JP Knox. Carbohydrate-binding modules promote the enzymatic deconstruction of intact plant cell walls by targeting and proximity effects. Proc Natl Acad Sci, 2010. [DOI | PubMed]
- A-M Hoffren, T Teeri, O Teleman. Molecular dynamics simulation of fungal cellulose-binding domains: differences in molecular rigidity but a preserved cellulose binding surface. Protein Eng, 1995. [DOI | PubMed]
- R Ioos, L Laugustin, N Schenk, S Rose, C Husson, P Frey. Usefulness of single copy genes containing introns in Phytophthora for the development of detection tools for the regulated species P. ramorum and P. fragariae. Eur J Plant Pathol, 2006. [DOI]
- RW Jones, MD Ospina-Giraldo. Novel cellulose-binding-domain protein in Phytophthora is cell wall localized. PLoS ONE, 2011. [DOI | PubMed]
- RW Jones, FG Perez. A cellulose-binding domain protein restores female fertility when expressed in transgenic Bintje potato. BMC Res Notes, 2016. [DOI | PubMed]
- M Kearse, R Moir, A Wilson, S Stones-Havas, M Cheung, S Sturrock, S Buxton, A Cooper, S Markowitz, C Duran, T Thierer, B Ashton, P Mentjies, A Drummond. Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics, 2012. [DOI | PubMed]
- LPN Kroon, FT Bakker, GBM van den Bosch, PJM Bonants, W Flier. Phylogenetic analysis of Phytophthora species based on mitochondrial and nuclear DNA sequences. Fungal Genet Biol, 2004. [DOI | PubMed]
- J Lehtio, J Sugiyama, M Gustavsson, L Fransson, M Linder, TT Teeri. The binding specificity and affinity determinants of family 1 and family 3 cellulose binding modules. Proc Natl Acad Sci, 2003. [DOI | PubMed]
- I Levy, O Shoseyov. Cellulose-binding domains: biotechnological applications. Biotechnol Adv, 2002. [DOI | PubMed]
- M Linder. Identification of functionally important amino acids in the cellulose-binding domain of Trichoderma reesei cellobiohydrolase I. Protein Sci, 1995. [DOI | PubMed]
- M Linder, T Nevanen, L Soderholm, O Bengs, TT Teeri. Improved immobilization of fusion proteins via cellulose-binding domains. Biotechnol Bioeng, 1998. [DOI | PubMed]
- M Linder, I Salovuori, L Ruohonen, TT Teeri. Characterization of a double cellulose-binding domain. J Biol Chem, 1996. [DOI]
- Q-Y Liu, N Ross, P Lanthier, M Reith. A gametophyte cell wall protein of the red alga Porphyra purpurea contains four apparent polysaccharide-binding domains. J Phycol, 1996. [DOI]
- FV Mateos, M Rickauer, M-T Esquerre-Tugaye. Cloning and characterization of a cDNA encoding an elicitor of Phytophthora parasitica var. nicotianae that shows cellulose-binding and lectin-like activities. Mol Plant Microb Interact, 1997. [DOI]
- H Meijer, F Mancuso, G Espadas, M Seidl, C Chiva, F Govers, E Sabido. Profiling the secretome and extracellular proteome of the potato late blight pathogen Phytophthora infestans. Mol Cell Proteom, 2014. [DOI]
- J Meng, Y Wang. Rapid detection of Phytophthora nicotianae in infected tobacco tissues and soil samples based on its Ypt1 gene. J Phytopathol, 2010. [DOI]
- C Oliveira, V Carvalho, L Domingues, F Gama. Recombinant CBM-fusion technology-applications overview. Biotechnol Adv, 2015. [DOI | PubMed]
- M Ospina-Giraldo, J Griffith, E Laird, C Mingora. The CAZyome of Phytophthora spp.: a comprehensive analysis of the gene complement coding for carbohydrate-active enzymes in species of the genus Phytophthora. BMC Genom, 2010. [DOI]
- TN Petersen, S Brunak, G von Heijne, H Nielsen. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods, 2011. [DOI | PubMed]
- R Pinto, AL Amaral, EC Ferreira, M Mota, M Vilanova, K Ruel, M Gama. Quantification of the CBD-FITC conjugates surface coating on cellulose fibres. BMC Biotechnol, 2008. [DOI | PubMed]
- N Saitou, M Nei. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol, 1987
- O Shoseyov, Z Shani, I Levy. Carbohydrate binding modules: biochemical properties and novel applications. Microbiol Mol Biol Rev, 2006. [DOI | PubMed]
- F Sievers, A Wilm, DG Dineen, TJ Gibson, K Karplus, W Li, R Lopez, H McWilliam, M Remmert, J Söding, JD Thompson, DG Higgins. Fast, scalable generation of high-quality protein multiple sequence alignments using clustal omega. Mol Syst Biol, 2011. [DOI | PubMed]
- K Tamura, G Stecher, D Peterson, A Filipski, S Kumar. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol, 2013. [DOI | PubMed]
- J Tokunaga, S Bartnicki-Garcia. Structure and differentiation of the cell wall of Phytophthora palmivora: cysts, hyphae and sporangia. Arch Microbiol, 1971
- H Wang, RW Jones. Cloning, characterization and functional expression of an endoglucanase-encoding gene from the phytopathogenic fungus Macrophomina phaseolina. Gene, 1995. [DOI | PubMed]
- H Wang, RW Jones. Analyzing the role of a cellulose-binding domain in fungal-plant interactions. Phytopathol, 1996
- Y Wei, J Shih, J Li, PH Goodwin. Two pectin lyase genes, pnl-1 and pnl-2, from Colletotrichum gloeosporioides f.sp. malvae differ in a cellulose-binding domain and in their expression during infection of Malva pusilla. Microbiology, 2002. [DOI | PubMed]
- LPTM Zevenhuizen, S Bartnicki-Garcia. Structure of the insoluble hyphal wall glucan of Phytophthora cinnamomi. Biochemistry, 1969. [DOI | PubMed]
- L Zhao, Q Pang, J Xie, J Pei, F Wang, S Fan. Enzymatic properties of Thermoanaerobacterium thermosaccharolyticum β-glucosidase fused to Clostridium cellulovorans cellulose-binding domain and its application in hydrolysis of microcrystalline cellulose. BMC Biotechnol, 2013
