Possible role of human ribonuclease dicer in the regulation of R loops
Abstract
R loops are three‐stranded nucleic acid structures that form naturally in cells under various conditions, mainly as intermediates during replication or as by‐products during transcription. R loops are involved in the regulation of many important cellular processes, including replication, transcription, centromere stabilization, protection of chromosome ends, or control of telomere length. Unscheduled R loops are linked to many diseases, including cancer, neurodegenerative, or inflammatory disorders. The list of cancer diseases linked to excessive R loop accumulation is growing rapidly. There is currently much debate about the understanding of abnormal R loop formation and its impact on genome instability and cancer development. In this review, we briefly describe the nature of R loops, their formation under physiological and pathological conditions, and the proteins involved in the regulation of R loops. In addition, we emphasize the possible role of the human ribonuclease Dicer, a multi‐tasking protein mostly known for its important role in microRNA biogenesis, in the regulation of R loops. We also discuss the involvement of R loops in cancer development and their potential use as diagnostic biomarkers. Knowledge of the molecular mechanisms underlying R loop dysregulation may significantly improve our understanding of cancer biology and provide new directions for research.
Article type: Review Article
Keywords: cancer, G‐quadruplexes, long non‐coding RNAs, R loops, ribonuclease Dicer, TERRA
Affiliations: Department of Ribonucleoprotein Biochemistry Institute of Bioorganic Chemistry Polish Academy of Sciences Poznan Poland
License: © 2025 The Author(s). FEBS Open Bio published by John Wiley & Sons Ltd on behalf of Federation of European Biochemical Societies. CC BY 4.0 This is an open access article under the terms of the http://creativecommons.org/licenses/by/4.0/ License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Article links: DOI: 10.1002/2211-5463.70026 | PubMed: 40127989 | PMC: PMC12401183
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (1.2 MB)
R loops are three‐stranded nucleic acid structures that consist of a DNA–RNA hybrid and a displaced single‐stranded DNA (ssDNA) [ref. 1]. Among the physical properties of chromatin that promote the opening of the double‐stranded DNA (dsDNA) and the formation of an R loop are negative DNA supercoiling, high GC content, and the breakage of chromosomal DNA. In addition, R loops can be stabilized by G‐quadruplex structures (Fig. 1) [ref. 2]. R loops form naturally as intermediates during replication or as by‐products during transcription. They also regulate telomere length and protect chromosome ends [ref. 3].
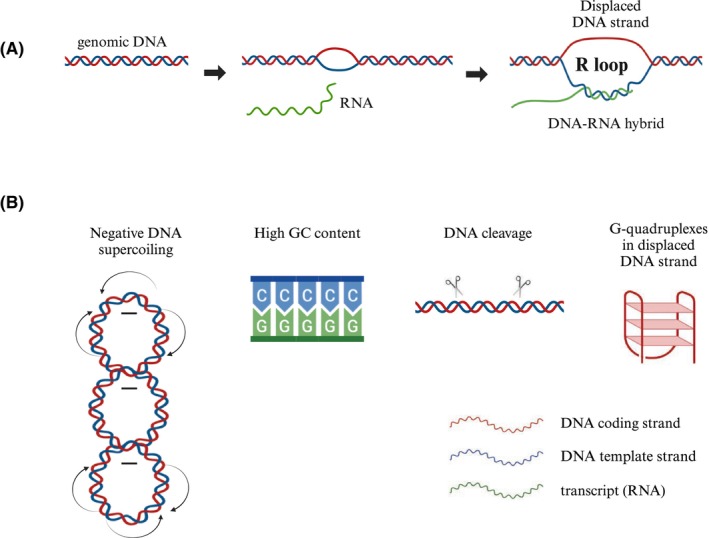
R loops as natural intermediates and regulators
R loops form during DNA replication initiation in mitochondrial DNA (mtDNA) [ref. 4], bacterial plasmids [ref. 5], and some bacteriophages [ref. 6, ref. 7]. In these cases, RNA produced by the RNA polymerase serves as a primer for replication initiation. Interestingly, R loop formation has been found to regulate the copy number of mtDNA. It is known that a somatic cell contains 1000–5000 mitochondria, each with 5–10 copies of mtDNA [ref. 8]. In the case of gametes: spermatozoa contain less than one hundred mitochondria, each with a single mtDNA, and oocytes have about 100 000–400 000 mitochondria, also with a single mtDNA copy [ref. 9, ref. 10].
R loops are also involved in immunoglobulin class‐switch recombination (CSR) [ref. 11, ref. 12]. CSR is important for the production of different classes of immunoglobulin by B lymphocytes after contact with a pathogen [ref. 13]. It is hypothesized that R loops boost mutation rates by generating long tracts of ssDNAs that serve as the substrate for activation‐induced deaminase (AID), the enzyme that co‐transcriptionally mutates ssDNA in so‐called switch recombination sequences [ref. 14]. AID‐induced mutations lead to DNA breakage and subsequent repair of two DNA sequences, which ultimately result in class‐switch recombination [ref. 15].
R loops can also be formed by long non‐coding RNAs (lncRNAs) [ref. 16]. A well‐known example of such structures is a telomeric R loop mediated by a telomeric repeat‐containing RNA (TERRA) [ref. 17, ref. 18, ref. 19]. TERRA, a lncRNA transcribed by RNA polymerase II from telomeric DNA sequences present at the ends of eukaryotic chromosomes [ref. 20, ref. 21], can invade telomeric DNA through direct base‐pairing to form R loop structures. G‐rich repeats (in humans, TTAGGG), single‐strand breaks, and G‐quadruplex structures in telomere sequences promote telomeric R loop formation. TERRA can regulate the telomeres [ref. 22, ref. 23, ref. 24]. The interaction between TERRA and G‐quadruplexes at telomeres plays a critical role in maintaining telomere stability [ref. 25]. The shelterin complex, composed of six proteins: TRF1, TRF2, POT1, RAP1, TIN2, and TPP1, serves as the primary regulator of TERRA, playing a crucial role in maintaining telomere stability and protecting chromosome ends [ref. 18, ref. 19]. TRF1, TRF2, and POT1 bind directly to telomeric DNA sequences. The remaining three (RAP1, TIN2, and TPP1) are associated with telomeres via protein–protein interactions [ref. 18, ref. 19]. The shelterin complex is also involved in the regulation of the R loops [ref. 18, ref. 19, ref. 26]. Other examples of lncRNAs that can form R loops are enhancer RNAs (eRNAs) [ref. 27, ref. 28]. eRNAs are relatively long non‐coding RNA molecules (50–2000 nucleotides) that are transcribed by RNA polymerase II from the DNA sequence of enhancer regions. eRNAs hybridize with their corresponding DNAs to form R loop structures [ref. 29]. In this way, eRNAs may modulate the activity of the matching enhancer in target genes. It has been shown that increased level of eRNAs leads to unscheduled R loop formation and consequent genomic instability at enhancer regions [ref. 30], which is causally linked to tumorigenesis.
R loop formation may also induce antisense transcription, leading to double‐stranded RNA (dsRNA) generation, followed by H3K9me2 marks and recruitment of heterochromatin protein 1γ (HP1γ) [ref. 31]. These events lead to RNA polymerase II pausing and transcription termination [ref. 31]. In addition, R loop formation may protect CpG islands within the promoter regions from DNA methyltransferases (DNMTs), which in consequence affect methylation‐dependent repression of transcription [ref. 32].
R loops as transcriptional by‐products
R loops are primarily transcriptional by‐products and are abundant in nucleolar regions, where RNA polymerase I drives the expression of ribosomal DNA (rDNA) [ref. 1]. These sites of R loop formation are particularly critical for genome stability, as RNA polymerase I accounts for more than 60% of total transcription [ref. 33].
R loop structures form also at centromeric regions [ref. 34]. Centromeres are chromosomal fragments responsible for the proper distribution of DNA during cell division. They consist of repetitive α‐satellite sequences that are intensely transcribed and form DNA–RNA hybrids. R loop structures at centromeres are recognized by and associated with BRCA1, a well‐known tumor suppressor preventing the accumulation of DNA–RNA hybrids [ref. 35]. In budding yeast, unscheduled R loops at centromeric regions have been found to contribute to defects in kinetochore biorientation and chromosomal instability [ref. 34].
One report has shown the importance of active transcription, the presence of a functional poly(A) signal, and termination G‐rich pause elements in the formation of R loops [ref. 36]. It has been observed that depletion of senataxin, a known RNA/DNA helicase, leads to the accumulation of R loops specifically downstream of the poly(A) signal, emphasizing a senataxin’s role in resolving these structures in transcription termination regions [ref. 36]. This finding suggests that while R loops may facilitate certain aspects of transcription termination, their persistence can be detrimental, highlighting the critical role of senataxin in clearing R loops to ensure proper gene expression and maintain genomic stability [ref. 36].
R loops as potentially harmful structures
R loops may impair replication fork progression and transcription; therefore, their removal is essential to preserve genome integrity [ref. 11, ref. 37]. Replication and transcription occur during the cell cycle. Since replication and transcription use the same DNA strand as a template, the cell cycle must be tightly regulated. Conflicts between these two events can result in errors in DNA replication and/or protein synthesis, both of which are essential for normal cell division [ref. 38]. R loop formation and removal is an important mechanism regulating these two processes. When replication and transcription proceed in opposite directions, RNA polymerase II cannot function due to a collision with DNA polymerase, and R loops accumulate, whereas co‐directional collision does not result in R loop accumulation [ref. 38]. R loops have been found during several stages of cell division: G1, G2, M phases, and mostly in S phase. The S phase leads to DNA synthesis, which requires DNA replication and transcription to produce multiple proteins [ref. 39]. R loops can inhibit transcription and enable DNA replication, especially during meiosis [ref. 40, ref. 41]. Thus, precise processing of R loops is crucial to maintain the balance between replication and transcription.
Displaced ssDNAs in R loop structures are susceptible to damage. Unscheduled R loops often lead to replication stress, as they can stall replication forks, potentially causing double‐strand breaks (DSBs) [ref. 42]. Moreover, unscheduled R loops may disturb the balance between replication and transcription, which may contribute to genomic instability. To counteract this, cells employ a variety of enzymes involved in R loop regulation and resolving, including topoisomerases, RNA/DNA helicases, chromatin modulators, RNA processing factors, and ribonucleases [ref. 43, ref. 44]. Examples of such proteins and their roles in the cell are summarized in Table 1.
Table 1: Exemplary enzymes involved in the regulation of R loops.
| Group | Enzyme | Role | Source |
|---|---|---|---|
| Topoisomerases | Topoisomerase I (TOP1) | Relaxes DNA supercoiling and prevents R loop formation | [ref. 118] |
| RNA/DNA helicases | DEAD‐Box Helicase 1 (DDX1) | Plays a crucial role in the formation of R loops over immunoglobulin heavy‐chain (IgH) switch regions by targeting AID (activation‐induced cytidine deaminase) through a post‐transcriptional mechanism. It binds to G‐quadruplex structures within switch transcripts and facilitates their conversion into R loops in the switch recombination sequences (the S‐regions) | [ref. 119] |
| Senataxin (SETX) | Catalyzes the unwinding of DNA–RNA hybrid in R loops, promoting their resolution | [ref. 120] | |
| Chromatin modulators | Metastasis‐associated protein 2 (MTA2) | Regulates chromatin dynamics in regions where R loops are formed, promoting genome stability by controlling access to DNA | [ref. 121] |
| DNA‐dependent ADP‐ribosyl transferase (PARP1) | Binds to R loops and initiates DNA repair processes, preventing genomic instability | [ref. 122] | |
| Breast cancer type 1 susceptibility protein (BRCA1) | Prevents accumulation of DNA–RNA hybrids | [ref. 35] | |
| RNA processing factors | The conserved THO complex (THOC) | Regulates TERRA‐associated R loops by binding to nucleoplasmic TERRA and reducing R loop accumulation at telomeres, thus maintaining telomere stability | [ref. 123] |
| Ribonucleases | RNase H1 | Degrades the RNA portion of the R loops, returning the two DNA strands to dsDNA form | [ref. 124] |
| RNase H2 | Degrades RNA in DNA–RNA hybrids, including RNA primers and ribonucleotides erroneously incorporated into DNA. The activity of RNase H2 is mainly found during cell division, in the G2/M checkpoint. This phase of the cell cycle controls the quality of replicated DNA and allows or prevents cells from dividing | [ref. 43, ref. 125] | |
| RNA Exonuclease 5 (REXO5) | Plays a key role in the physiological control of R loops using its exonuclease domain | [ref. 126] | |
| Dicer | Cleaves the RNA strand of DNA–RNA hybrids within R loop structures | [ref. 57] | |
| Recombinase | DNA repair protein (RAD51) | RAD51 interacts with TERRA and catalyzes R loop formation, a direct role in TERRA recruitment via strand invasion | [ref. 19, ref. 127] |
Ribonuclease dicer as a multi‐tasking protein
The Dicer ribonuclease is mostly known for its important role in the biogenesis of small regulatory RNAs, microRNAs (miRNAs) and small interfering RNAs (siRNAs) [ref. 45, ref. 46]. This canonical role of Dicer is associated with its cytoplasmic localization [ref. 47]. Dicer can also function in the nucleus. For example, it is already known that nuclear Dicer can participate in chromatin structure remodeling [ref. 48, ref. 49, ref. 50, ref. 51, ref. 52], restrict the deleterious accumulation of endogenous dsRNAs [ref. 53] or contribute to DNA damage response activation by generating small non‐coding RNAs, referred to as damage‐induced small RNAs (diRNAs) or DDRNAs [ref. 54, ref. 55, ref. 56]. Interestingly, a recent study has shown that ribonuclease Dicer may be involved in nuclear R loop processing [ref. 57]. Under the in vitro conditions and in the cell, human Dicer (hDicer) cleaved RNA within harmful R loops, but not within the DNA–RNA hybrid without a loop, which suggested that hDicer activity was specific for the R loop structures [ref. 57]. Silencing of the human DICER1 gene led to the accumulation of R loop structures in the nuclei, and even the overexpression of other ribonucleases involved in the R loop removal did not counteract the accumulation of R loops in the cells with downregulated DICER1 expression [ref. 57]. It has been suggested that R loop removal by hDicer is supported by its annealing activity through facilitating DNA–RNA hybrid formation [ref. 57]. The annealing activity of hDicer was first revealed [ref. 55] and extensively characterized [ref. 56, ref. 58] in our laboratory. We demonstrated that hDicer can support hybridization between complementary sequences present in nucleic acids even when both of them are trapped within secondary structures [ref. 55, ref. 56]. Besides cleaving the RNA within the R loop structures, hDicer might influence R loops by involvement in DNA damage repair. Specifically, hDicer has been implicated in the DNA damage response and repair of DSBs in mammalian cells due to replication stress [ref. 59]. As mentioned above, during DSB stress, hDicer produces diRNAs that are essential for the repair of damaged DNA [ref. 60, ref. 61]. These small RNAs correspond to the sites of DSBs and are thought to serve as templates for efficient DNA repair [ref. 60]. In addition, hDicer can potentially interplay with other R loop‐binding proteins, by direct interactions; e.g., helicase DHX9 [ref. 62, ref. 63, ref. 64], or indirectly; e.g., BRCA2/BRCA1 [ref. 65, ref. 66], BRD4 [ref. 67, ref. 68], SETX [ref. 69, ref. 70]. Furthermore, hDicer can potentially stabilize R loops by binding to G‐quadruplexes. Importantly, the results of our recent studies have indicated that hDicer binds both DNA and RNA G‐quadruplexes, including TERRA, with high affinity [ref. 71]. The potential of hDicer to interact with G‐quadruplexes and R loops, two structures with regulatory functions in the cell, strongly indicates that this ribonuclease is a multi‐tasking protein not only involved in miRNA and siRNA biogenesis, but also in many other cellular pathways (summarized in Fig. 2).
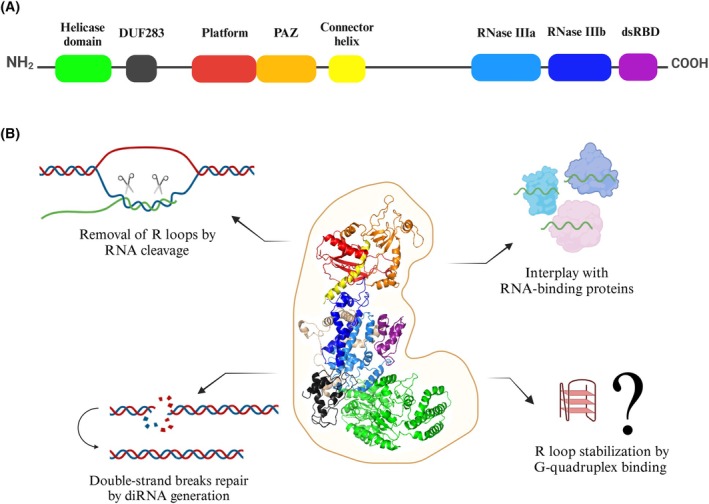
Dysregulation of hDicer in cancer
Many cases have been reported in which abnormal DICER1 expression was correlated with the development of carcinogenesis; reviewed in [ref. 46]. In the context of a specific cancer, disturbances in DICER1 expression should be considered at multiple levels, including mutations in the DICER1 gene, the level of DICER1 expression, as well as global miRNA expression. As mentioned above, disturbed levels of hDicer may also affect the R loop accumulation in cells [ref. 57]. On the one hand, DICER1 overexpression can cause excessive R loop removal, either directly (by RNA cleavage) or indirectly (by impaired miRNA biogenesis). On the other hand, decreased levels of hDicer can cause ineffective removal of R loops in cells, which threaten genome integrity [ref. 57]. Here, we focus on aberrant DICER1 expression in leukemia and thyroid cancer.
Leukemia is a hematologic malignancy originating in the bone marrow, characterized by the abnormal proliferation of blood cells. Four main types of leukemia can be distinguished: acute lymphoblastic leukemia (ALL), acute myeloid leukemia (AML), chronic lymphocytic leukemia (CLL), and chronic myeloid leukemia (CML) [ref. 72]. Disturbances in the hDicer level have been implicated in distinct leukemia subtypes. The extremely high overexpression of DICER1 was found in AML (in TCGA database referred to as LAML) [ref. 73, ref. 74, ref. 75] (Fig. 3), and reduced levels of hDicer were detected in CLL, CML, and ALL [ref. 76, ref. 77, ref. 78, ref. 79]. Interestingly, in ALL, overexpression of DICER1 has also been reported [ref. 80], which indicates that, depending on the cancer stage, the DICER1 expression may be affected differently.
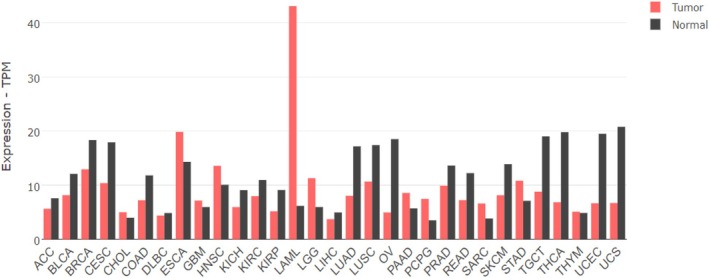
Several reports have shown the elevated levels of DICER1 expression in samples from the bone marrow of AML patients [ref. 74, ref. 75] and leukemia cell lines [ref. 74], confirmed on both mRNA [ref. 74, ref. 75] and protein [ref. 74] levels. However, since no difference in overall survival of AML patients with increased versus decreased expression of DICER1 in bone marrow was found [ref. 75], it was suggested that the influence of DICER1 expression on clinical outcomes may depend on the tissue of origin [ref. 75]. Moreover, it was shown that in AML bone marrow samples, the DICER1 gene expression was upregulated by the hematopoietic transcription factor, GATA1 [ref. 74], whose mutations are highly significant in AML patients [ref. 81]. In addition, upregulation of hDicer was found in various myelomas [ref. 82, ref. 83] and T‐cell lymphoma [ref. 84]. Abnormal expression of the DICER1 gene can affect the course of diseases [ref. 85, ref. 86, ref. 87, ref. 88].
While the precise cause of leukemia remains unclear, genetic mutations and environmental factors, such as smoking, ionizing radiation, viral infections, and exposure to harmful chemicals, are believed to play a role in its development and progression. These factors contribute to DNA damage [ref. 72]. As mentioned above, hDicer is one of the proteins involved in damaged DNA repair [ref. 89, ref. 90]. Knockdown of DICER1 has been found to reduce DSB repair [ref. 91] and increase the accumulation of unscheduled R loops [ref. 57]. It is also important to mention that loss of a single allele of DICER1 enhances tumorigenesis (e.g., by increased amounts of oncogenic versus tumor suppressive miRNAs) [ref. 46], while its complete loss leads to the accumulation of DNA damage and cell death [ref. 90]. Indeed, silencing of the DICER1 gene in AML cells inhibited cell proliferation and promoted apoptosis [ref. 74].
Reduced levels of DICER1 expression were observed in chemotherapy‐resistant CML cells [ref. 79]. In this leukemia, the rs13078 DICER1 variant (a single nucleotide polymorphism located in the 3′‐UTR of the DICER1 gene) was extensively examined [ref. 92]. Both the minor allele frequency and the minor homozygote genotype of rs13078 were significantly more prevalent in CML patients than in healthy controls. This variant is thought to alter the interaction of miRNAs or RNA‐binding proteins with DICER1 transcripts, potentially affecting the hDicer protein level and global miRNA expression, which may drive CML pathogenesis [ref. 92].
Reduced hDicer levels have also been reported in CLL [ref. 76, ref. 77]. The DICER1 expression analysis revealed statistically significant differences between ill and healthy samples (based on RNA isolated from 30 patients and 29 controls), where the DICER1 gene was downregulated in the patients with CLL compared to the controls [ref. 76]. Another study has demonstrated that reduced levels of hDicer in CLL are strongly associated with shorter overall survival and reduced treatment‐free survival [ref. 77]. In this context, it is also important to remember that the reduced hDicer level may impact DSB repair and R loop regulation.
A significantly reduced DICER1 level was also observed in thyroid cancer (THCA) [ref. 93] (Fig. 3). Thyroid cancer originates from thyroid parenchymal cells and has shown a steady rise in incidence globally, although mortality rates have remained stable in recent years [ref. 72, ref. 94]. hDicer is crucial for normal thyroid gland development, and mutations in the DICER1 gene are associated with various thyroid abnormalities [ref. 95]. A reduced level of Dicer in thyroid cancer could exacerbate the accumulation of R loops. In addition, a study from 2021 showed that thyroid hormone receptor‐associated protein 3 (Thrap3) plays a critical role in regulating R loop resolution by interacting with DEAD‐box helicase 5 (DDX5) [ref. 96]. Since both DDX5 and hDicer can regulate R loops, it would be of great importance to understand the connections among proteins involved in R loop regulation.
The abovementioned cases highlight the importance of understanding the molecular mechanisms underlying Dicer dysregulation and its contributions to cancer biology, particularly regarding R loop dynamics and their implications for genomic stability.
R loops, lncRNAs and cancer
The understanding of abnormal R loop formation and its impact on genome instability and cancer development is extensively discussed. The list of cancer diseases linked to the excessive R loop accumulation is growing rapidly [ref. 43]. Here, we focus on unscheduled R loops formed by lncRNAs.
It has been shown that a lncRNA HOTTIP (the HOXA transcript at the distal tip) regulates oncogene expression through topologically associating domains (TAD) formation via R loops, without causing DNA damage [ref. 72]. Mammalian chromosomes are organized within the nucleus into TADs, which support gene regulation, DNA replication, and repair [ref. 97]. TADs are dynamic structures formed by actively extruding loops that are restricted by TAD boundaries, enhancing intra‐domain interactions [ref. 98, ref. 99, ref. 100]. HOTTIP lncRNA is abundantly expressed in AML. The HOTTIP‐mediated TAD formation via R loops drives aberrant oncogene transcription and leukemia development [ref. 101].
Abnormal accumulation of TERRA‐mediated R loops can interfere with telomere function, potentially leading to telomere‐related dysfunction, promoting genomic instability in diseases like cancer [ref. 3]. It has also been reported that TERRA forms R loops to promote homology‐directed DNA synthesis in the alternative lengthening of telomeres (ALT) pathway [ref. 102, ref. 103, ref. 104]. For example, TERRA contributes to ALT by recruiting the lysine‐specific demethylase 1A (LSD1) to telomeres. The interaction between TERRA and LSD1 has been shown to promote R loop formation and enhance the activity of R loop regulatory molecules, supporting telomere maintenance in ALT pathways [ref. 102]. ALT can also be promoted by RAD51‐associated protein 1 (RAD51AP1) [ref. 103, ref. 104]. It has been shown that TERRA R loops mediated by RAD51AP1 regulate repressive chromatin at telomeres [ref. 104]. Moreover, it has been reported that in multiple myeloma, stabilization of TERRA G‐quadruplexes induced dissociation of telomeric repeat‐binding factor 2 (TRF2) from telomeres, leading to the activation of the DNA damage response, cell cycle arrest, proliferation block, and apoptotic death [ref. 105].
Perspectives
There are reports suggesting that R loops can serve as diagnostic biomarkers. Their unscheduled accumulation has emerged as a promising diagnostic biomarker, with the potential to stratify patients and monitor disease severity across various conditions, including immune deficiencies like Wiskott–Aldrich syndrome [ref. 106], leukemia [ref. 72], embryonal tumors with correlation to loss of DICER1 function [ref. 107], and glioblastomas [ref. 108], also in correlation with DICER1 expression [ref. 109, ref. 110, ref. 111].
Landmark studies over the past decade have shown that small extracellular vesicles (sEVs, also referred to as “exosomes” [ref. 112]) from human blood contain genomic DNA fragments reflecting the host cell genome [ref. 111]. Notably, sEV‐DNA from cancer patients accurately mirrors the mutational status of the original tumor cells, highlighting its potential as a liquid biopsy biomarker for cancer detection and monitoring metastasis [ref. 111]. In addition, recent studies have shown the existence of microvesicles containing miRNAs and lncRNAs, which may activate multiple pathways involved in tumor development when transferred to the target cells [ref. 113, ref. 114, ref. 115]. The microvesicle‐derived lncRNAs might as well contribute to unscheduled R loop formation. Elevated R loop levels are linked to genomic instability and tumor heterogeneity, and their presence offers prognostic value, guiding therapeutic approaches and predicting outcomes in disorders such as multiple myeloma [ref. 116] and uterine fibroids [ref. 117], particularly when analyzed alongside extracellular vesicle‐derived genetic materials [ref. 111]. This could lead to a breakthrough in diagnosing diseases by utilizing easily accessible biological samples, such as blood, for rapid and accurate testing.
Conflict of interest
The authors declare no conflict of interest.
Author contributions
AK‐K conceptualized the work, KW and PK wrote the draft of the manuscript. KW prepared all figures and Table 1. AK‐K revised and edited the manuscript and was responsible for its final form. All authors have read and agreed to the published version of the manuscript.
References
- R loops: new modulators of genome dynamics and function.. Nat Rev Genet, 2015. [PubMed]
- R‐loop induced G‐quadruplex in non‐template promotes transcription by successive R‐loop formation.. Nat Commun, 2020. [PubMed]
- R‐loops at chromosome ends: from formation, regulation, and cellular consequence.. Cancers (Basel), 2023. [PubMed]
- A mechanism of formation of a persistent hybrid between elongating RNA and template DNA.. Cell, 1990. [PubMed]
- R‐loop‐dependent replication and genomic instability in bacteria.. DNA Repair, 2019. [PubMed]
- Initiation of bacteriophage T4 DNA replication and replication fork dynamics: a review in the virology journal series on bacteriophage T4 and its relatives.. Virol J, 2010. [PubMed]
- The essential role of recombination in phage T4 growth.. Annu Rev Genet, 1987. [PubMed]
- Maternal inheritance of human mitochondrial DNA.. Proc Natl Acad Sci USA, 1980. [PubMed]
- The role of mitochondrial DNA copy number in mammalian fertility.. Biol Reprod, 2010. [PubMed]
- Rearranged mitochondrial genomes are present in human oocytes.. Am J Hum Genet, 1995. [PubMed]
- R loops: from transcription byproducts to threats to genome stability.. Mol Cell, 2012. [PubMed]
- R‐loops at immunoglobulin class switch regions in the chromosomes of stimulated B cells.. Nat Immunol, 2003. [PubMed]
- Class‐switch recombination: interplay of transcription, DNA deamination and DNA repair.. Nat Rev Immunol, 2004. [PubMed]
- An evolutionarily conserved target motif for immunoglobulin class‐switch recombination.. Nat Immunol, 2004. [PubMed]
- Nucleic acid structures and enzymes in the immunoglobulin class switch recombination mechanism.. DNA Repair, 2003. [PubMed]
- Author correction: gene regulation by long non‐coding RNAs and its biological functions.. Nat Rev Mol Cell Biol, 2021
- TERRA R‐loops trigger a switch in telomere maintenance towards break‐induced replication and PrimPol‐dependent repair.. 2025. [DOI]
- TERRA R‐loops connect and protect sister telomeres in mitosis.. Cell Rep, 2023. [PubMed]
- The makings of TERRA R‐loops at chromosome ends.. Cell Cycle, 2021. [PubMed]
- Telomeric repeat‐containing RNA structure in living cells.. Proc Natl Acad Sci USA, 2010. [PubMed]
- TERRA and Telomere maintenance in the yeast Saccharomyces cerevisiae .. Genes (Basel), 2023. [PubMed]
- The emerging roles of TERRA in telomere maintenance and genome stability.. Cells, 2019. [PubMed]
- Telomere‐specific regulation of TERRA and its impact on telomere stability.. Semin Cell Dev Biol, 2024. [PubMed]
- Role of TERRA in the regulation of telomere length.. Int J Biol Sci, 2015. [PubMed]
- G‐quadruplexes at telomeres: friend or foe?. Molecules, 2020. [PubMed]
- Trypanosoma brucei RAP1 maintains telomere and subtelomere integrity by suppressing TERRA and telomeric RNA:DNA hybrids.. Nucleic Acids Res, 2017. [PubMed]
- Widespread transcription at neuronal activity‐regulated enhancers.. Nature, 2010. [PubMed]
- A large fraction of extragenic RNA pol II transcription sites overlap enhancers.. PLoS Biol, 2010. [PubMed]
- Carcinogen‐induced super‐enhancer RNA promotes nasopharyngeal carcinoma metastasis through NPM1/c‐Myc/NDRG1 axis.. Am J Cancer Res, 2023. [PubMed]
- RNA exosome‐regulated long non‐coding RNA transcription controls super‐enhancer activity.. Cell, 2015. [PubMed]
- R‐loops induce repressive chromatin marks over mammalian gene terminators.. Nature, 2014. [PubMed]
- R‐loop formation is a distinctive characteristic of unmethylated human CpG Island promoters.. Mol Cell, 2012. [PubMed]
- Real‐time imaging of RNA polymerase I activity in living human cells.. J Cell Biol, 2023. [PubMed]
- R‐loops at centromeric chromatin contribute to defects in kinetochore integrity and chromosomal instability in budding yeast.. Mol Biol Cell, 2021. [PubMed]
- BRCA1 prevents R‐loop‐associated centromeric instability.. Cell Death Dis, 2021. [PubMed]
- Human senataxin resolves RNA/DNA hybrids formed at transcriptional pause sites to promote Xrn2‐dependent termination.. Mol Cell, 2011. [PubMed]
- R‐loops cause replication impairment and genome instability during meiosis.. EMBO Rep, 2012. [PubMed]
- R‐loop and diseases: the cell cycle matters.. Mol Cancer, 2024. [PubMed]
- DNA replication and progression through S phase.. Oncogene, 2005. [PubMed]
- R‐loop formation in meiosis: roles in meiotic transcription‐associated DNA damage.. Epigenomes, 2022. [PubMed]
- Dual roles of R‐loops in the formation and processing of programmed DNA double‐strand breaks during meiosis.. Cell Biosci, 2023. [PubMed]
- DNA:RNA hybrids form at DNA double‐strand breaks in transcriptionally active loci.. Cell Death Dis, 2020. [PubMed]
- R‐loops in genome instability and cancer.. Cancers (Basel), 2023. [PubMed]
- The contribution of co‐transcriptional RNA:DNA hybrid structures to DNA damage and genome instability.. DNA Repair, 2014. [PubMed]
- Genetic insight into the domain structure and functions of dicer‐type ribonucleases.. Int J Mol Sci, 2021. [PubMed]
- The many faces of dicer: the complexity of the mechanisms regulating dicer gene expression and enzyme activities.. Nucleic Acids Res, 2015. [PubMed]
- Role for a bidentate ribonuclease in the initiation step of RNA interference.. Nature, 2001. [PubMed]
- Dicer is essential for formation of the heterochromatin structure in vertebrate cells.. Nat Cell Biol, 2004. [PubMed]
- Convergent transcription induces transcriptional gene silencing in fission yeast and mammalian cells.. Nat Struct Mol Biol, 2012. [PubMed]
- RNAi‐mediated targeting of heterochromatin by the RITS complex.. Science, 2004. [PubMed]
- Regulation of heterochromatic silencing and histone H3 lysine‐9 methylation by RNAi.. Science, 2002. [PubMed]
- RITS acts in cis to promote RNA interference‐mediated transcriptional and post‐transcriptional silencing.. Nat Genet, 2004. [PubMed]
- Human nuclear dicer restricts the deleterious accumulation of endogenous double‐stranded RNA.. Nat Struct Mol Biol, 2014. [PubMed]
- Nuclear phosphorylated dicer processes double‐stranded RNA in response to DNA damage.. J Cell Biol, 2017. [PubMed]
- Revealing a new activity of the human dicer DUF283 domain in vitro.. Sci Rep, 2016. [PubMed]
- The RNA‐RNA base pairing potential of human dicer and Ago2 proteins.. Cell Mol Life Sci, 2020. [PubMed]
- DICER ribonuclease removes harmful R‐loops.. Mol Cell, 2023. [PubMed]
- The significance of the DUF283 domain for the activity of human ribonuclease dicer.. Int J Mol Sci, 2021. [PubMed]
- Dicer prevents genome instability in response to replication stress.. Oncotarget, 2019. [PubMed]
- Site‐specific DICER and DROSHA RNA products control the DNA‐damage response.. Nature, 2012. [PubMed]
- A role for small RNAs in DNA double‐strand break repair.. Cell, 2012. [PubMed]
- Human DHX9 helicase preferentially unwinds RNA‐containing displacement loops (R‐loops) and G‐quadruplexes.. DNA Repair, 2011. [PubMed]
- RNA/DNA hybrid interactome identifies DXH9 as a molecular player in transcriptional termination and R‐loop‐associated DNA damage.. Cell Rep, 2018. [PubMed]
- The biology of DHX9 and its potential as a therapeutic target.. Oncotarget, 2016. [PubMed]
- BRCA2 regulates transcription elongation by RNA polymerase II to prevent R‐loop accumulation.. Cell Rep, 2018. [PubMed]
- The coexistence of a BRCA2 germline and a DICER1 somatic variant in two first‐degree cousins suggests their potential synergic effect.. Sci Rep, 2024. [PubMed]
- BRD4 prevents R‐loop formation and transcription‐replication conflicts by ensuring efficient transcription elongation.. Cell Rep, 2020. [PubMed]
- Dicer promotes genome stability via the bromodomain transcriptional co‐activator BRD4.. Nat Commun, 2022. [PubMed]
- Senataxin plays an essential role with DNA damage response proteins in meiotic recombination and gene silencing.. PLoS Genet, 2013. [PubMed]
- Walking a tightrope: the complex balancing act of R‐loops in genome stability.. Mol Cell, 2022. [PubMed]
- RNA and DNA G‐quadruplexes bind to human dicer and inhibit its activity.. Cell Mol Life Sci, 2021. [PubMed]
- Clinical and mechanistic implications of R‐loops in human leukemias.. Int J Mol Sci, 2023. [PubMed]
- GEPIA2: an enhanced web server for large‐scale expression profiling and interactive analysis.. Nucleic Acids Res, 2019. [PubMed]
- Overexpression of DICER1 induced by the upregulation of GATA1 contributes to the proliferation and apoptosis of leukemia cells.. Int J Oncol, 2013. [PubMed]
- Dicer and outcomes in patients with acute myeloid leukemia (AML).. Leuk Res, 2009. [PubMed]
- Dicer gene expression as a prognostic factor in acute lymphoblastic leukemia and chronic lymphocytic leukemia in Fars Province.. Iran J Med Sci, 2016. [PubMed]
- Downregulated dicer expression predicts poor prognosis in chronic lymphocytic leukemia.. Cancer Sci, 2012. [PubMed]
- MicroRNA processing and human cancer.. J Clin Med, 2015. [PubMed]
- Crosstalk between kinases, phosphatases and miRNAs in cancer.. Biochimie, 2014. [PubMed]
- The impact of variations in transcription of DICER and AGO2 on exacerbation of childhood B‐cell lineage acute lymphoblastic leukaemia.. Int J Exp Pathol, 2019. [PubMed]
- GATA1 mutation analysis and molecular landscape characterization in acute myeloid leukemia with trisomy 21 in pediatric patients.. Int J Lab Hematol, 2021. [PubMed]
- Role of dicer as a prognostic predictor for survival in cancer patients: a systematic review with a meta‐analysis.. Oncotarget, 2016. [PubMed]
- Upregulation of dicer is more frequent in monoclonal gammopathies of undetermined significance than in multiple myeloma patients and is associated with longer survival in symptomatic myeloma patients.. Haematologica, 2011. [PubMed]
- Polymorphisms in microRNA‐related genes are associated with survival of patients with T‐cell lymphoma.. Oncologist, 2014. [PubMed]
- Abnormal expression of DICER1 leads to dysregulation of inflammatory effectors in human synoviocytes.. Mediat Inflamm, 2019
- DICER1 syndrome and cancer predisposition: from a rare pediatric tumor to lifetime risk.. Front Oncol, 2020. [PubMed]
- DICER1 tumor predisposition syndrome: an evolving story initiated with the pleuropulmonary blastoma.. Mod Pathol, 2022. [PubMed]
- DICER1 syndrome and its various paediatric presentations: case series and review of the literature.. EJC Paediatr Oncol, 2024
- Next‐generation sequencing reveals two populations of damage‐induced small RNAs at endogenous DNA double‐strand breaks.. Nucleic Acids Res, 2018. [PubMed]
- The paradox of dicer in cancer.. Mol Cell Oncol, 2016. [PubMed]
- Dicer1 promotes colon cancer cell invasion and migration through modulation of tRF‐20‐MEJB5Y13 expression under hypoxia.. Front Genet, 2021. [PubMed]
- Gene variants in components of the microRNA processing pathway in chronic myeloid leukemia.. Genes (Basel), 2024. [PubMed]
- Understanding the dosage‐dependent role of Dicer1 in thyroid tumorigenesis.. Int J Mol Sci, 2024. [PubMed]
- Thyroid cancer: A review.. JAMA, 2024. [PubMed]
- DICER1 gene alterations in thyroid diseases.. Cancer Cytopathol, 2020. [PubMed]
- Thrap3 promotes R‐loop resolution via interaction with methylated DDX5.. Exp Mol Med, 2021. [PubMed]
- Principles of genome folding into topologically associating domains.. Sci Adv, 2019. [PubMed]
- TADs: dynamic structures to create stable regulatory functions.. Curr Opin Struct Biol, 2023. [PubMed]
- Making sense of the linear genome, gene function and TADs.. Epigenetics Chromatin, 2022. [PubMed]
- Chromosome compartments on the inactive X guide TAD formation independently of transcription during X‐reactivation.. Nat Commun, 2021. [PubMed]
- HOTTIP‐dependent R‐loop formation regulates CTCF boundary activity and TAD integrity in leukemia.. Mol Cell, 2022. [PubMed]
- TERRA‐LSD1 phase separation promotes R‐loop formation for telomere maintenance in ALT cancer cells.. Nat Commun, 2024. [PubMed]
- TERRA and RAD51AP1 promote alternative lengthening of telomeres through an R‐ to D‐loop switch.. Mol Cell, 2022. [PubMed]
- RAD51AP1 regulates ALT‐HDR through chromatin‐directed homeostasis of TERRA.. Mol Cell, 2022. [PubMed]
- TERRA G‐quadruplex stabilization as a new therapeutic strategy for multiple myeloma.. J Exp Clin Cancer Res, 2023. [PubMed]
- ZFP281‐BRCA2 prevents R‐loop accumulation during DNA replication.. Nat Commun, 2022. [PubMed]
- The molecular landscape of ETMR at diagnosis and relapse.. Nature, 2019. [PubMed]
- Increased replication stress and R‐loop accumulation in EGFRvIII‐expressing glioblastoma present new therapeutic opportunities.. Neurooncol Adv, 2022. [PubMed]
- Dicer deficiency impairs proliferation but potentiates anti‐tumoral effect of macrophages in glioblastoma.. Oncogene, 2022. [PubMed]
- The nuclear DICER‐circular RNA complex drives the deregulation of the glioblastoma cell microRNAome.. Sci Adv, 2020. [PubMed]
- Extracellular vesicle‐associated DNA: ten years since its discovery in human blood.. Cell Death Dis, 2024. [PubMed]
- Small extracellular vesicles: A novel avenue for cancer management.. Front Oncol, 2021. [PubMed]
- Extracellular vesicles as tools and targets in therapy for diseases.. Signal Transduct Target Ther, 2024. [PubMed]
- Extracellular vesicles: new classification and tumor immunosuppression.. Biology, 2023. [PubMed]
- Exosome‐transmitted lncRNA PCGEM1 promotes invasive and metastasis in gastric cancer by maintaining the stability of SNAI1.. Clin Transl Oncol, 2021. [PubMed]
- From R‐lupus to cancer’: reviewing the role of R‐loops in innate immune responses.. DNA Repair, 2023. [PubMed]
- Aberrant R‐loop‐induced replication stress in MED12‐mutant uterine fibroids.. Sci Rep, 2022. [PubMed]
- Topoisomerase 1 prevents replication stress at R‐loop‐enriched transcription termination sites.. Nat Commun, 2020. [PubMed]
- RNA helicase DDX1 converts RNA G‐quadruplex structures into R‐loops to promote IgH class switch recombination.. Mol Cell, 2018. [PubMed]
- Senataxin and R‐loops homeostasis: multifaced implications in carcinogenesis.. Cell Death Discov, 2023. [PubMed]
- The chromatin network helps prevent cancer‐associated mutagenesis at transcription‐replication conflicts.. Nat Commun, 2023. [PubMed]
- PARP1 associates with R‐loops to promote their resolution and genome stability.. Nucleic Acids Res, 2023. [PubMed]
- The THO complex counteracts TERRA R‐loop‐mediated telomere fragility in telomerase+ cells and telomeric recombination in ALT+ cells.. Nucleic Acids Res, 2023. [PubMed]
- RNase H1, the gold standard for R‐loop detection.. Methods Mol Biol, 2022. [PubMed]
- RNase H2 degrades toxic RNA:DNA hybrids behind stalled forks to promote replication restart.. EMBO J, 2023. [PubMed]
- REXO5 promotes genomic integrity through regulating R‐loop using its exonuclease activity.. Leukemia, 2024. [PubMed]
- RAD51‐dependent recruitment of TERRA lncRNA to telomeres through R‐loops.. Nature, 2020. [PubMed]
- Cryo‐EM structure of human dicer and its complexes with a pre‐miRNA substrate.. Cell, 2018. [PubMed]
