Exploring Emergent Properties in Enzymatic Reaction Networks: Design and Control of Dynamic Functional Systems
Abstract
The intricate and complex features of enzymatic reaction networks (ERNs) play a key role in the emergence and sustenance of life. Constructing such networks in vitro enables stepwise build up in complexity and introduces the opportunity to control enzymatic activity using physicochemical stimuli. Rational design and modulation of network motifs enable the engineering of artificial systems with emergent functionalities. Such functional systems are useful for a variety of reasons such as creating new-to-nature dynamic materials, producing value-added chemicals, constructing metabolic modules for synthetic cells, and even enabling molecular computation. In this review, we offer insights into the chemical characteristics of ERNs while also delving into their potential applications and associated challenges.
License: © 2024 The Authors. Published by American Chemical Society CC BY 4.0 Permits the broadest form of re-use including for commercial purposes, provided that author attribution and integrity are maintained (https://creativecommons.org/licenses/by/4.0/).
Article links: DOI: 10.1021/acs.chemrev.3c00681 | PubMed: 38476077 | PMC: PMC10941194
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (15.6 MB)
Introduction
All key functions of living systems, such as metabolism, reproduction, sensing the environment, adaptation, and homeostasis, are enabled by enzymatic reaction networks (ERNs). In metabolic networks, the activities of many enzymatic reactions are finely tuned to provide responsiveness and controlled behavior. Highly complex emergent properties such as chemotaxis,1,2 cell division, and the cell cycle2 are all orchestrated by enzymatic networks. Signaling pathways consisting of cascades of kinases and phosphatases interlock to form complex networks that allow cells to process information from the environment and activate suitable genetic programs.3−5
In the field of synthetic biology, enormous progress has been made in appropriating and harnessing some of the enzymatic networks in living cells to create, for example, artificial proteolysis signaling pathways and control intracellular self-assembly.6−8 Progress in synthetic biology has been reviewed earlier.9,10 Here, we wish to focus on synthetic enzymatic reaction networks outside a cellular context. These in vitro systems offer the key advantage of precise control over all components. This control enables deep and detailed chemical understanding of the dynamics of ERNs, which is a prerequisite for designing systems with emergent properties. The bottom-up construction of functional ERNs could then be used to create life-like materials, where the emergent dynamics of the networks can be coupled to material properties. Ultimately, ever more complex artificial ERNs may be used in the construction of synthetic cells, composed of multiple networks at the genetic and metabolic level, all compartmentalized within a synthetic compartment.
To achieve these long-term ambitions, the field has started to develop robust methods to construct ERNs and control their temporal and spatial dynamics. In this review, we will provide an overview of the progress to date, discuss challenges ahead (Figure ), and finish with an overview table listing all enzymes used in enzymatic reaction networks (Table 1). The structure of our review is as follows: we start in section 2 with an overview of the structure of networks by discussing topology, motifs, and kinetics, highlighting approaches to modeling the dynamic properties of ERNs as well as opportunities for computational design of functional ERNs. In section 3, we provide examples of artificial ERNs with nonlinear dynamic behavior. In section 4, we discuss various means to control enzymatic activity via external stimuli such as pH, light, magnetic fields, and sound. Combined, these design and control principles have led to progress in enzymatically powered dynamic materials with life-like properties, which are reviewed in section 5. Having established the key aspects of designing and controlling ERNs, we devote a final set of sections on potential areas of application. In section 6, we review how ERNs can be used in the cell-free synthesis of valuable compounds. In section 7, we summarize the ambitious efforts to build a synthetic cell from the bottom up and highlight new developments where the information processing typically reserved for living systems is harnessed in synthetic ERNs. We conclude the review with a brief summary of the key goals and potential bottlenecks for future research.
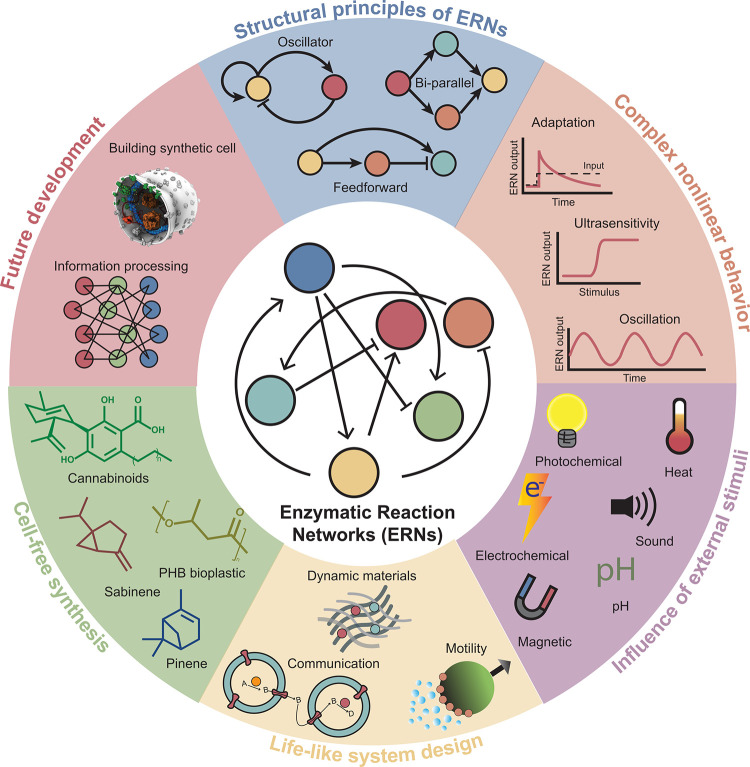
Table 1: Overview of Enzymes Mentioned in the Review
| enzymes | Enzyme Commission number | class | mode of action | section | ref |
|---|---|---|---|---|---|
| alcohol dehydrogenase (ADH) | 1.1.1.1 | oxidoreductase | ADH converts primary alcohols to aldehydes | 5.2 | (ref. 165) |
| l-lactate dehydrogenase (LDH) | 1.1.1.27 | oxidoreductase | LDH converts pyruvate to lactate and vice versa in glycolysis | 5.2 | (ref. 164) |
| l-malate NADP+ oxidoreductase (ME) | 1.1.1.40 | oxidoreductase | ME catalyzes oxidative decarboxylation of l-malate to form pyruvate (reversible) | 4.3 | (ref. 105) |
| hydroxymethylglutaryl-CoA reductase (HMGR) | 1.1.1.88 | oxidoreductase | HMGR converts 3-hydroxy-3-methylglutaryl-CoA (HMG-CoA) to mevalonic acid | 6.1 | (ref. 208) |
| glucose-6-phosphate dehydrogenase (G6PDH) | 1.1.1.49 | oxidoreductase | G6PDH converts glucose-6-phosphate to 6-phosphogluconolactone in pentose-phosphate pathway | 5.2, 5.3, and 6.1 | (ref. 164), (ref. 179), (ref. 202) |
| hydroxybutyryl-CoA dehydrogenase (Hbd) | 1.1.1.157 | oxidoreductase | Hbd catalyzes conversion of acetoacetyl-CoA to hydroxybutyryl-CoA and vice versa | 6.1 | (ref. 205) |
| l-lactate oxidase (LO) | 1.1.3.2 | oxidoreductase | LO is a FMN-containing enzyme that catalyzes conversion of lactate to pyruvate | 5.2 | (ref. 164) |
| glucose oxidase (GOx) | 1.1.3.4 | oxidoreductase | GOx catalyzes oxidation of d-glucose to d-gluconic acid by oxygen | 3.1, 3.3, 4.2, 4.5, and 5.1 | (ref. 60), (ref. 61), (ref. 78), (ref. 96), (ref. 99), (ref. 110), (ref. 139), (ref. 147), (ref. 185) |
| choline oxidase (COx) | 1.1.3.17 | oxidoreductase | COx is a flavoprotein, which catalyzes formation of betaine from choline | 5.1 | (ref. 147) |
| glyceraldehyde-3-phosphate dehydrogenase (Gap) | 1.2.1.12 | oxidoreductase | Gap converts glyceraldehyde 3-phosphate to 1,3-bisphosphoglycerate during glycolysis | 6.1 | (ref. 200) |
| pyruvate dehydrogenase (PDH) | 1.2.4.1 | oxidoreductase | PDH catalyzes the reaction of pyruvate and a lipoamide to give dihydrolipoamide and CO2 and is involved many metabolic pathways like glycolysis and TCA cycle | 6.1 | (ref. 200), (ref. 201), (ref. 203) |
| pyruvate:ferredoxin oxidoreductase (PFOR) | 1.2.7.1 | oxidoreductase | PFOR converts acetyl-CoA to pyruvate in many metabolic cycles including pyruvate metabolism, propanoate metabolism, and butanoate metabolism | 6.2 | (ref. 219) |
| d-amino acid oxidase (DAAO) | 1.4.3.3 | oxidoreductase | DAAO converts d-amino acids to 2-oxo carboxylates and is involved in d-amino acid metabolism | 4.2 | (ref. 95) |
| sarcosine oxidase (SOx) | 1.5.3.1 | oxidoreductase | SOx catalyzes formation of glycine from sarcosine by oxidative demethylation | 5.1 and 5.2 | (ref. 147), (ref. 165) |
| NADH oxidase (NoxE) | 1.6.3.4 | oxidoreductase | water-forming NoxE is a flavoprotein that specifically oxidizes NADH, not NADP | 6.1 | (ref. 199−ref. 202) |
| urate oxidase (UOx) | 1.7.3.3 | oxidoreductase | UOx is involved in the allantoin pathway and converts uric acid to 5-hydroxyisourate | 5.1 | (ref. 147) |
| catalase (Cat) | 1.11.1.6 | oxidoreductase | Cat is a peroxidase and is involved in biosynthesis of tryptophan and secondary metabolites | 5.2 | (ref. 161−ref. 163), (ref. 165), (ref. 168−ref. 170), (ref. 184), (ref. 185), (ref. 187) |
| horseradish peroxidase (HRP) | 1.11.1.7 | oxidoreductase | catalyzes oxidation of various organic substrates by hydrogen peroxide; heme containing glycoprotein with many isoforms | 3.3, 4.2, 4.5, 5.1, and 5.3 | (ref. 74−ref. 76), (ref. 97), (ref. 110), (ref. 111), (ref. 140), (ref. 178), (ref. 183), (ref. 184) |
| NiFe hydrogenase | 1.12.2.1 | oxidoreductase | NiFe hydrogenase oxidizes H2 to H+ (reversible), present in prokaryotes | 3.3 and 4.3 | (ref. 79), (ref. 106) |
| renillaluciferase (RLuc) | 1.13.12.5 | oxidoreductase | RLuc converts coelenterazine to excited coelenteramide and emits blue light | 5.3 | (ref. 176) |
| ferredoxin NADP+ reductase (FNR) | 1.18.1.2 | oxidoreductase | FNR is a flavoprotein involved in photosynthesis | 4.3 | (ref. 105) |
| Mo-dependent nitrogenase | 1.18.6.1 | oxidoreductase | Mo-dependent nitrogenase involved in nitrogen fixation, catalyzes ammonia formation from nitrogen | 4.3 | (ref. 106) |
| acetyl-CoA acetyltransferase (ACAT) | 2.3.1.9 | transferase | ACAT is present in many metabolic pathways, where it catalyzes formation of acetoacetyl-CoA from acetyl-CoA | 6.1 | (ref. 208) |
| PHB synthase (PhaC) | 2.3.1.304 | transferase | Phac catalyzed formation of Bioplastic PHB from 3-hydroxybutyryl-CoA | 6.1 | (ref. 202) |
| hydroxymethylglutaryl-CoA synthase (HMGS) | 2.3.3.10 | transferase | HMGS catalyzes formation of HMG-CoA from acetyl-CoA and acetoacetyl-CoA in the mevalonate pathway | 6.1 | (ref. 208) |
| glycogen phosphorylase b (GPb) | 2.4.1.1 | transferase | GPb breaks down glycogen to form glucose-1-phosphate and is involved in starch and sucrose metabolism | 5.3 | (ref. 179) |
| geranyl diphosphate synthase (GPPS) | 2.5.1.1 | transferase | GPPS forms geranyl diphosphate from the condensation of dimethylallyl diphosphate (DMAPP) and isopentenyl diphosphate (IPP) | 6.1 | (ref. 204), (ref. 208) |
| hexokinase (HK) | 2.7.1.1 | transferase | hexokinase phosphorylate d-hexose sugars in the presence of ATP, thus playing a very important role in glycolysis | 4.3 and 5.2 | (ref. 104), (ref. 163) |
| mevalonate kinase (MK) | 2.7.1.36 | transferase | MK phosphorylates mevalonate to mevalonate-6-phosphate | 6.1 | (ref. 208) |
| protein kinase A (PKA) | 2.7.1.37 | transferase | PKA initiates phosphorylation of serine residues present in the peptide chain | 5.1 | (ref. 145) |
| pyruvate kinase (PK) | 2.7.1.40 | transferase | PK catalyzes the last step of glycolysis by transferring the phosphate group from PEP to ADP | 5.2 | (ref. 163) |
| phosphomevalonate kinase (PMK) | 2.7.4.2 | transferase | PMK catalyzes the phosphorylation of mevalonate-6-phosphate to form diphosphomevalonate in the mevalonate pathway | 6.1 | (ref. 208) |
| esterase (Est, PLE for pig liver esterase) | 3.1.1.1 | hydrolase | Est catalyzes hydrolysis of the ester bonds of carboxyl esters | 3.1, 4.1, and 4.5 | (ref. 59), (ref. 60), (ref. 65), (ref. 93), (ref. 110) |
| phospholipase A2 (PLA2) | 3.1.1.4 | hydrolase | PLA2 catalyzes cleavage of phospholipids | 5.3 | (ref. 176) |
| acetylcholine esterase (AchE) | 3.1.1.7 | hydrolase | AchE breaks down the ester bond of neurotransmitter acetylcholine to form choline and acetic acid | 4.2 and 5.1 | (ref. 99), (ref. 147) |
| phosphatase | 3.1.3.1/3.1.3.2 | hydrolase | phosphatases dephosphorylate phosphate esters | 5.1 | (ref. 139), (ref. 141) |
| amyloglucosidase (AMG) | 3.2.1.3 | hydrolase | AMG breaks down starch to glucose, hence involved in starch metabolism | 4.5 | (ref. 110) |
| lactase | 3.2.1.108 | hydrolase | lactase catalyzes the conversion of lactose to galactose and glucose | 5.3 | (ref. 183) |
| aminopeptidase (Ap) | 3.4.11.2 | hydrolase | catalyzes cleavage of N-terminal amino acids from peptides and is involved in glutathione metabolism | 3.2 | (ref. 67) |
| chymotrypsin (Cr) | 3.4.21.1 | hydrolase | Cr is a serine protease that cleaves peptides on the C terminal of phenylalanine, tyrosine, tryptophan, and leucine amino acids | 3.2 and 5.1 | (ref. 70), (ref. 73), (ref. 146) |
| trypsin (Tr) | 3.4.21.4 | hydrolase | Tr is a serine protease that cleaves peptides on the C terminal of arginine and lysine amino acids | 3.2, 4.1, 4.4, and 5.2 | (ref. 42), (ref. 67−ref. 73), (ref. 86), (ref. 89), (ref. 146) |
| elastase (Els) | 3.4.21.36 | hydrolase | elastase breaks down elastin (responsible for the elasticity of connective tissue) and cleaves after glycine, alanine, and valine amino acids | 3.2 | (ref. 73) |
| proteinase K | 3.4.21.64 | hydrolase | proteinase K is a serine protease with a broad spectrum of cleavage site preferences | 5.3 | (ref. 175) |
| matrix metalloproteinase 2 (MMP 2) | 3.4.24.24 | hydrolase | MMP 2 is an endopeptidase and cleaves collagens type IV, V, VII, and X; also known as gelatinase A | 5.1 | (ref. 145) |
| matrix metalloproteinase 9 (MMP 9) | 3.4.24.35 | hydrolase | similar to MMP 2; also known as gelatinase B | 5.1 | (ref. 17) |
| urease (Ur) | 3.5.1.5 | hydrolase | catalyzes the hydrolysis of urea to carbon dioxide and ammonia, which basifies the solution | 3.1, 4.1, 4.2, 5.1, and 5.2 | (ref. 51−ref. 54), (ref. 57−ref. 59), (ref. 61), (ref. 93), (ref. 97−ref. 99), (ref. 110), (ref. 157−ref. 160), (ref. 171), (ref. 185), (ref. 187), (ref. 189) |
| apyrase | 3.6.1.5 | hydrolase | apyrase hydrolyses di- and triphosphate nucleotides to monophosphate nucleotides | 5.3 | (ref. 179) |
| oxaloacetate acetylhydrolase (OAH) | 3.7.1.1 | hydrolase | OAH breaks down oxaloacetate to form oxalate and acetate | 6.2 | (ref. 219) |
| pyrophosphomevalonate decarboxylase (PMD) | 4.1.1.33 | lyase | PMD catalyzes the last step of the mevalonate pathway, where it converts diphosphomevalonate to isopentenyl diphosphate | 6.1 | (ref. 208) |
| carbonic anhydrase (CA) | 4.2.1.1 | lyase | CA catalyzes the equilibrium between carbon dioxide and carbonic acid | 4.3 | (ref. 105) |
| fumarase (FumC) | 4.2.1.2 | lyase | FumC converts malate to fumarate (reversible) and is involved in the TCA cycle and pyruvate metabolism | 4.3 | (ref. 105) |
| limonene synthase (LS) | 4.2.3.16 | lyase | limonene synthase catalyzes limonene formation from geranyl diphosphate | 6.1 | (ref. 200), (ref. 208) |
| pinene synthase | 4.2.3.121 | lyase | pinene synthase catalyzes the conversion of geranyl diphosphate to pinene | 6.1 | (ref. 200) |
| l-aspartate ammonia-lyase (AspA) | 4.3.1.1 | lyase | AspA converts l-aspartate to fumarate (reversible) and is involved in amino acid metabolism | 4.3 | (ref. 105) |
| isopentenyl pyrophosphate isomerase (IDI) | 5.3.3.2 | isomerase | IDI catalyzes the conversion of IPP to DMAPP | 6.1 | (ref. 208) |
| phosphoglucomutase (PMG) | 5.4.2.2 | isomerase | PMG catalyzes the isomerization of glucose-1-phosphate to glucose-6-phosphate and is involved in many metabolic pathways including glycolysis and the pentose phosphate pathway | 5.3 | (ref. 179) |
| acetate-CoA ligase (ACS) | 6.2.1.1 | ligase | ACS catalyzes acetyl-CoA formation from acetate and CoA in the presence of ATP | 6.2 | (ref. 219) |
| pyruvate carboxylase (PYC) | 6.4.1.1 | ligase | PYC catalyzes the carboxylation of pyruvate to form oxaloacetate in the TCA cycle | 6.2 | (ref. 219) |
| cytochrome C oxidase | 7.1.1.9 | translocase | cytochrome C oxidase catalyzes the translocation of hydrons and is involved in oxidative phosphorylation pathways | 4.2 | (ref. 95) |
| F-type ATP synthase | 7.1.2.2 | translocase | F-type ATP synthase forms ATP from ADP and phosphate (Pi) | 7.1 | (ref. 227), (ref. 228) |
Structural Principles of Enzymatic Networks
Despite a huge variety in enzymes and enzymatic reactions, many of the enzymatic reaction networks (ERNs) found in living cells feature recurring topologies and motifs. By focusing on these patterns, it becomes clear how certain functionalities arise from specific groupings of enzymes, and similar or alternative networks can be identified.
Network Topology
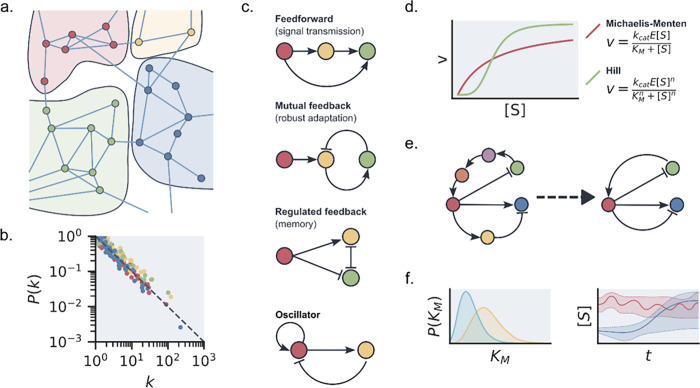
At first sight, the enzymatic networks encountered in living systems show an overwhelming complexity. However, statistical network analysis shows that these networks possess a strongly structured topology, where network topology refers to the large-scale logical structure of a network and the statistical properties of its connections.11 For example, the full metabolism is divided into strongly connected modules and pathways that each have their own function (a schematic example is shown in Figure a).12,13 This modularity is reflected in the statistical description of interactions in ERNs. Generally, these are found to follow scaling relationships (Figure b).14−16 The distribution of reactions per enzyme follows an exponential scaling distribution. This means that many types of enzymes only promote one or a few reactions, while a few enzymes promote many reactions. This characteristic distribution makes the network more robust to failure while remaining efficient. Identifying the most common network topologies for specific functionalities and the most and least prominent enzymes in those networks can lead to insights into plasticity and redundancy features of specific network topologies,17 which can be exploited by ERNs to function in a range of different environments.18 A downside to this statistical approach to ERNs is that it only gives a large-scale picture, neglecting any heterogeneity introduced by different types of enzyme reactions and dynamics, and does not distinguish between substrates and effector molecules. Additionally, it does not translate well to the analysis of artificial ERNs, as these networks are often too small to warrant a proper statistical treatment.
Network Motifs
While ERN topology analysis can provide us with information on the dynamics of large and complex systems, zooming in to smaller subunits can help us gain a better understanding of how certain properties arise from the combination of just a few enzymes. Motifs are the smallest functional units, meaning that an individual motif is capable of showcasing complex behavior such as oscillations, adaptation and memory, amplification, and filtering.19 A classic example is the emergence of ultrasensitivity in a simple network containing a forward and backward reaction catalyzed by two different enzymes and the emergence of adaptation by inhibitory feedback between two different enzymes converting the same substrate, both key motifs in signaling networks. In their classic paper, Goldbeter and Koshland analyzed the kinetics of such systems and explored the control parameters which would show sensitivity.20,21 In 1997, Barkai and Leibler showed how robust adaptation results directly from network connectivity.22 Milo and co-workers generalized the concept of these small subnetworks and identified network motifs across a range of different networks—repeated patterns of interactions occurring in large and complex networks at a higher rate than a random network with equivalent topology.23 Examples of these include simple cascades, positive or negative feedback loops, or bifans (see Figure c). Interestingly, network motifs can vary dramatically between different types of networks but may remain largely the same when different networks have the same type of functionality. For example, gene regulation, neurons, and logic circuits often contain feed-forward loops and bifans to enable rapid switch-like behavior, which are different from those found in food webs, electronic circuits used for arithmetic, and social networks. Similarly, in ERNs, different networks with similar behavior are found to often have the same motifs.24 This can be clearly seen in the design of enzymatic oscillators, which almost exclusively involve delayed negative feedback loops, although more complex motifs may be involved as well.25 More recently, researchers have shown how different large network topologies can lead to the same effective network motifs capable of, for example, homeostasis and how descriptions of larger networks can be reduced to smaller functional motifs.26 Network motifs can be used as a starting point for the design of specific behavior, inspired by either behavior shown in already existing ERNs27,28 or recreating network motifs not found in ERNs but in other types of networks.
Enzyme Kinetics
Topology and motifs constrain what types of behavior are possible in ERNs, but the kinetics of enzymatic reactions ultimately determine what will happen and how fast. Many enzymatic reactions are traditionally characterized by Michaelis–Menten (MM) kinetics. Michaelis–Menten kinetics assume a two-step mechanism, where the binding of the enzyme to the substrate is reversible and the release of product is not. While a number of assumptions underlie MM kinetics, it is found to be generally applicable even when some of those assumptions do not hold. In the case of cooperative binding, the Michaelis–Menten equation can be extended to the Hill equation (Figure d). For multisubstrate enzymes and inhibitory effects, further extensions of the MM equation are possible.
In recent years, further improvements and replacements to MM kinetics have been developed. For example, Piephoff et al. established a generalized form of the Michaelis–Menten equation by explicitly incorporating nonequilibrium effects.29 They took into account how different enzyme conformations impact the speed of the reaction and automatically corrected for different types of substrate binding and allosteric effects. Alternatively, Rowher et al. have shown that the reversible Hill equation is a good general approximation for a large variety of mechanisms found in enzyme kinetics.30
However, kinetics data for individual enzymes or complete ERNs are often limited. Consequently, the data used to obtain kinetic parameters can be insufficient to obtain correct estimates. To warrant against overfitting of data, care should be taken to choose equations that do not contain an excess of parameters.31 This can be checked for by performing sensitivity analyses on each of the parameters in the kinetic model if a large network proves to be partially unidentifiable; a viable reduced-form network description might still be obtained. By determining which interactions and variables are sloppy and subsequently removing those from the network model, a reduced description shows only those enzymes and interactions that influence the network behavior (schematically shown in Figure e).32 This method has been used effectively to obtain efficient descriptions of cell-signaling networks33 but has yet to be used for artificial ERNs.
Alternatively, so-called Bayesian inference techniques can be used for parameter estimation.34 Here, one can use prior knowledge, such as the physically allowed range or probabilistic estimates obtained from previous measurements, to constrain parameter estimations and obtain probabilistic uncertainty intervals and predictions (Figure f).35 Data with non-normal, or unknown, noise distributions can also be incorporated with relative ease by including additional measurement noise terms in the likelihood function. Bayesian inference methods allow for much more accurate quantification of the uncertainties remaining in a model fit.36 Hierarchical Bayesian models can be used to correctly combine data from different experiments and measurement techniques, taking into account different accuracies, noise profiles, and different subsets of observable parameters, and can even be used to compare the likelihood of different reaction mechanisms.37
More recently, techniques to check the identifiability of parameters in enzymatic networks directly from equations have become available. These require the construction of so-called sensitivity matrices directly from the kinetic models in combination with experimental data.38 When parameters are unidentifiable from a single-input experiment alone, further analysis is required to check if changing the experimental conditions can improve parameter identifiability.39 More recently, a new approach has been developed to calculate optimal experimental designs for parameter identification.40 This method uses pulse patterns for input substrates in flow to decorrelate kinetic parameters and improve identifiability.246 Alternatively, work has been done to decompose a full ERN into separate submodules which are individually identifiable.41
Choosing the right analysis method to obtain an accurate kinetic model depends on both the nature and the complexity of the system and the goal behind it. If the goal is optimizing for specific reaction products, it requires a different amount of knowledge than when the goal is to achieve a fundamental understanding of all interactions in an ERN. As shown by the recent developments discussed above, structural identifiability analysis and (Bayesian) parameter estimation highlight the importance of proper uncertainty quantification in increasingly large and complex ERNs.
Complex Nonlinear Behavior in Artificial ERNs
As discussed above, nonlinear dynamics introduced by positive (autocatalytic) or negative (inhibitory) feedback loops are key characteristics of networks exhibiting complex properties. In contrast to the large metabolic and signaling networks found in living systems, synthetic enzymatic networks with complex behavior are typically based on one or just a few feedback reactions. Figure a illustrates the general principle behind the design of artificial ERNs with complex nonlinear dynamic behavior: a nonlinear core reaction motif is chosen and combined with additional processes; the actual behavior of this combination is then dependent on the type of reactor that is used. Although the additional processes are typically dependent on the enzymes chosen for the core motif and thus the two are often intertwined, we explicitly separate the two to aid the reader in appreciating the network design and analyzing their dynamics.
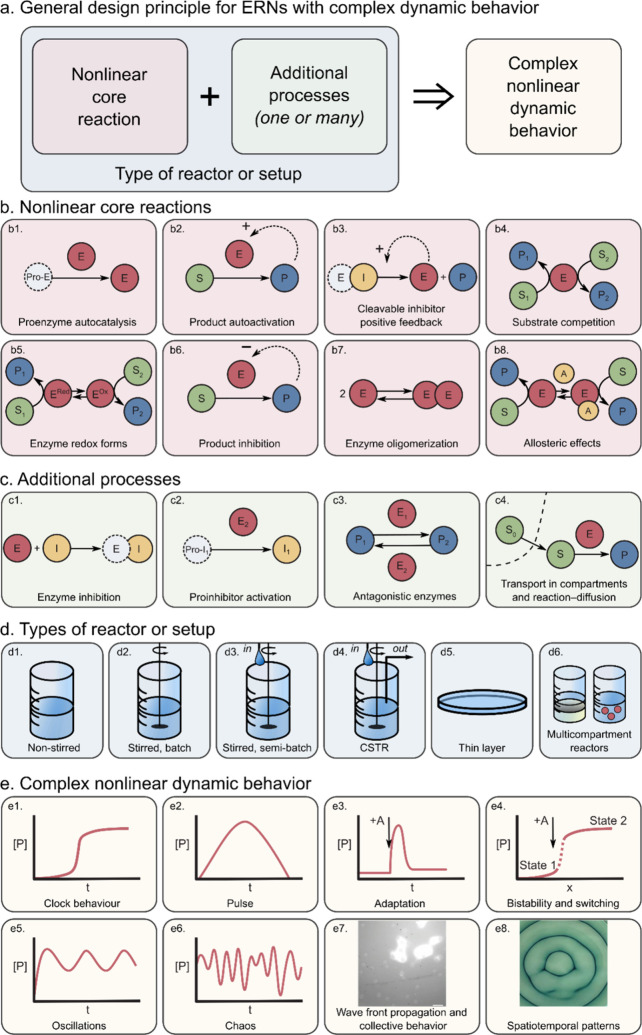
The full toolbox of nonlinear core enzymatic reactions described in the literature to date is presented in Figure b. The use of autocatalytic core reactions is prominent not only in enzymatic networks but also in organic and DNA networks,42 and the design principles of autocatalytic networks were recently reviewed by Semenov and Plasson et al..43,45 The exploitation of feedback in the engineering of ERNs was reviewed by Bánsági and Taylor.44,45 The three general ways to autocatalysis in ERNs are shown in Figure b1–b3. Figure b1 shows autocatalysis by self-activation of an inactive zymogen, with the trypsinogen–trypsin (Tg–Tr) pair as a well-known example discussed in detail in section 3.2. Product autoactivation (Figure b2) is another very general mechanism with examples mostly involving urease (section 3.1). The cleavable inhibitor positive feedback is relevant for proteolytic networks (see section 3.2) and shown in Figure b3. As we discuss later, individual autocatalytic steps are not a general prerequisite to complex behavior, and autocatalysis can also emerge from closed cycles where the separate steps are linear.242 The alternative nonlinear cores, namely, those built on substrate competition (Figure b4) and switching between enzyme redox forms (Figure b5), also deliver nonlinear dynamics, as discussed in section 3.3 using the example of oxidoreductases. Additionally, there are three types of nonlinear cores that are somewhat underexploited. Product inhibition (Figure b6), an example of which is the autoinhibitory catalysis of esterase, can be a nonlinear core but has only been found as an additional process in networks comprising a urease core (see section 3.1). The enzyme oligomerization (Figure b7) and allosteric effects (Figure b8) are well-known ways to achieve nonlinearity, but they have only been demonstrated in theoretical studies and in in vivo networks.20,21
To achieve controlled complex behavior, a core motif is supplemented and altered by what we call additional processes (Figure c). These processes may also serve to connect different network nodes. Inhibition (Figure c1) slows down the reaction rate of a certain enzyme, which is an exceptionally useful feedback loop in the case when an inhibitor is produced by another enzyme (Figure c2). Effectively, this connects two enzymatic activities that are otherwise independent (see section 3.2 on the trypsin oscillator). Another way to connect enzymatic activities is via the use of antagonistic enzymes (Figure c3), with examples mostly on pH-dependent urease networks (section 3.1). Finally, throughout sections 3.1–3.3 we discuss how transport processes enrich modes of dynamic behavior. Transport across membranes, compartment borders, between phases, and simple diffusion (Figure c4) add complexity and provide important types of nonlinear dynamics relevant to nature (collective behavior and patterns, Figure e7 and 3e8 correspondingly). The influence of transport processes has been recently studied theoretically, highlighting how spatial localization and transport phenomena in nonstirred systems can yield complex nonlinear responses even in fairly simple networks.46,47 The transport processes are only relevant in nonstirred systems (Figure d1 and 3d5) or in compartmentalized systems (Figure d6). Other reactor features presented in Figure d can supply reagents and remove products, maintaining the network out of equilibrium (Figure d3 and 3d4).
Combinations of the core reactions and additional processes yield a range of possible complex behaviors as shown in Figure e. While some types are more common and relatively straightforward to obtain (for instance, clock behavior (Figure e1) or pulse-like response (Figure e2)), some other types are rare (chaos as shown in Figure e6 is, to the best of our knowledge, only obtained in peroxidase–oxidase reaction (section 3.3)).
In order to provide the reader with a readily accessible overview, we grouped the synthetic ERNs with complex nonlinear behavior according to the central enzymatic reaction around which the network is built. For all of the discussed networks, we refer to the motifs they contain and behavior types they produce, as per Figure .
Networks Based on the Urea–Urease Reaction
In general, enzymes producing acid or base are a fruitful ground for designing systems with complex dynamics due to the nonlinear kinetics intrinsic to pH-reactive systems.48−50 A simple and well-studied enzymatic network that produces nonlinear behavior is the urea–urease network. The enzyme urease (Ur) is found in many bacteria and plants. It catalyzes the hydrolysis of urea into carbon dioxide and ammonia, resulting in an increase in pH. The enzyme has a standard bell-shaped activity–pH profile with an optimum around 7.5.51 When starting from a lower pH (usually buffers of pH 3.5–5.0 with low buffer capacity), the enzyme basifies the solution, thereby overcoming the buffer and increasing its own activity, up to pH 7.5. As the reaction proceeds further to even higher pH values, the Ur activity drops again and the final pH of about 8.5 is that of ammonia/ammonium buffer. This process creates an autocatalytic sigmoidal signature in pH vs time, and the time of the steep transition from low to high pH state is addressed in the literature as “clock time” (Figure e1).51,52 The increase in Ur activity due to the Ur reaction is an example of product autoactivation (Figure b2).
Compartmentalization and coupling to diffusion enrich the range of complex behaviors obtained in Ur networks. Effectively, the simple combination of the Ur nonlinear core (Figure b2) with transport phenomena (Figure c4) yields five types of complex behavior (Figure e1–e7), particularly because diffusion coefficients for H+ and other species are very different. Ur-loaded millimeter-sized particles exhibit quorum-sensing-like autoactivation (similarly to Figure e7), which is a consequence of autocatalysis and diffusion front propagation.53 Miele et al. demonstrated the collective synchronized behavior of Ur-loaded microvesicles.52 Muzika et al. showed sustained oscillations in a flow reactor with differential influx of urea and H+,54 building on earlier work on bistability,55 and noise-induced irregular oscillatory switching.56 Oscillations were also obtained in a system with Ur-loaded lipid vesicles by combining the clock reaction inside of the vesicle with the influx of urea and H+ from the bulk phase.57 Finally, oscillations in packed arrays of Ur-loaded beads with influx of urea and acid from the bulk solution were predicted theoretically but not yet experimentally proven.58
An alternative approach to obtain richer dynamic behavior from the urea–urease network is combining it with antagonistic acidifying enzymes (Figure c3). Heinen et al. showed that a urea–urease/ester–esterase network produces pulse-like pH responses under batch conditions with, first, a Ur activity-related increase followed by an esterase (Est) activity-induced decrease in the pH (see Figure d for network topology). The kinetic profile of the response can be finely tuned by changing the buffer and substrate composition.59 Compartmentalization of Ur and Est units in a two-layered gel aided in programming of transient acidic pH flips.60 This latter work also addressed the lack of nonlinearity of the Est response (as compared to Ur), which originates in the shape of the Est pH–activity profile and its autoinhibitory nature (Figure b6). An alternative acidifying enzyme is glucose oxidase (GOx), converting glucose to δ-gluconolactone, hydrolyzing spontaneously into gluconic acid. Wang achieved dynamic pH switching in response to the addition of urea and glucose in a system with Ur and GOx incorporated in pH-responsive polymersomes.61 While Ur is very fast, all of the acidifying enzymes explored thus far exhibited certain limitations on rates. We hypothesize that coupling the urea–urease network to faster acidifying counterplayers would yield more robust dynamic behavior and possibly oscillating systems. The pH transition caused by the Ur-based ERNs can be coupled to a wide range of chemical and physical processes,62 such as polymerization and sol–gel transitions,59,63 gel growth,64,65 and precipitation of CaCO3,66 that all can serve as handles for functionalities as discussed in section 5.
Networks Based on Proteases
A general route to autocatalysis is to exploit the formation of certain proteases from their inactive precursors (zymogens or proenzymes), as shown in Figure b1. Semenov et al. constructed a bienzymatic oscillatory network with a core reaction of trypsin (Tr)-mediated trypsinogen (Tg) activation modified with a delayed negative feedback loop via an aminopeptidase-activated trypsin proinhibitor (combination of motifs Figure b1, 3c1, and 3c2; full network topology is depicted in Figure a).67 Further investigations of this trypsin oscillator included studies on the boundary conditions of the oscillatory regimes using a range of slightly different inhibitors in the negative feedback loop68 and synchronization with external temperature oscillations.69 Exploiting protease zymogen activation as a core step, Helwig et al. demonstrated adaptation to an external stimulus in an incoherent type 1 feed-forward loop network motif, comprising the chymotrypsinogen–chymotrypsin (Cg–Cr) pair and Tr.70 The network responded with a peak to a persistent external stimulus (influx of Tr) and adapted over time, bringing the response Cr activity close to the original baseline, a behavior type in Figure e3. The possibility to use autocatalytic trypsinogen activation for signal enhancing was demonstrated in a diffusion-reaction system with trypsin and its inhibitor in a gel.71 Finally, Kriukov et al. showed in a Tr–Tg network with an additional inhibition step that the combination of autocatalytic activation and flow leads to hysteresis, and the inhibition can be used to control the network dynamics in a history-dependent manner.72
A different and novel type of autoactivation in enzymatic systems was introduced by Pogodaev et al. using slow substrates (cleavable inhibitors) for proteinases Tr, Cr, and elastase (Els), as shown in Figure b3.73 The reactions started in highly inhibited states, where the slow substrates occupied the active sites of the enzymes. With time, the peptides were cleaved into low-inhibitory fragments, thereby liberating more of the free enzymes, leading to an autocatalytic increase in enzyme activity. It was also demonstrated that in mixtures with Cr, Tr, and their corresponding slow substrates, crosstalk occurred, while Cr and Els remained orthogonal under these conditions, which opens roads to modular construction of modified networks. By converting a cleavable peptide inhibitor into a phosphate-modified proinhibitor, the behavior of the network was coupled to the activity of alkaline phosphatase, which acted as an initiator of the flip response in Cr activity. This is another example of using the motif of proinhibitor activation (Figure c1 and 3c2)) to connect behaviors of two enzymes.73
Networks Based on Oxidoreductases
In oxidoreductase networks, the nonlinearity often originates from the switching between redox and coordination forms of the enzymes (Figure b5).
The earliest example of an in vitro enzymatic oscillatory reaction is the peroxidase–oxidase (PO) reaction, first described by Yokota and Yamazaki in 1965.74 Oscillations occur during the horseradish peroxidase (HRP)-catalyzed oxidation of NADH (reduced nicotinamide adenine dinucleotide) by oxygen in the presence of additives such as methylene blue and 2,4-dichlorophenol in a semibatch stirred reactor with an influx of NADH and diffusion of oxygen from a gas mixture and no outlet as shown in Figure d3. The PO reaction has a complex mechanism comprising multiple redox and coordination forms of the heme cofactor of HRP and very rich dynamics that include normal, period-doubled, and mixed-mode oscillations, quasi-periodicity, and chaos, shown in Figure e5 and 3e6.75,76 It is hypothesized that many more new dynamic states of the PO reaction are still to be discovered using experimental parameter combinations that go beyond the current regimes, with the pH being an especially important control parameter.76 We refer the reader to more specialized literature for further details.75,77
Recently, HRP was combined with GOx to produce tunable pulse responses in a stirred reactor (Figure e2) and form spatiotemporal patterns (Figure e8) in a nonstirred thin layer experiment. Remarkably, this example of complex dynamics is not comprised of individual steps of autocatalytic activation or inhibition, with the nonlinearity originating only in delayed feedback loops and substrate competition (Figure b4).78 A unique aspect of the behavior of this network is the spontaneous formation of patterns of convection flows when not stirred.
Another enzyme exhibiting autocatalytic kinetics is hydrogenase, a metalloenzyme catalyzing the reaction H2 = 2H+ + 2e–. The mechanism of autocatalysis differs from the simple acid–base activation/deactivation shown for urease and is thought to be connected to redox forms.79 The hydrogenase reaction has been observed to produce reaction fronts and oscillations in a thin layer reaction–diffusion experiment via a mechanism that is not yet fully understood.80
Summary
In summary, synthetic in vitro ERNs with complex behavior have exploited a limited set of core reactions but demonstrated a large range of types of complex behavior. As the recent finding with hydrogenase demonstrates, identifying novel experimental realizations of theoretical network motifs is still a fruitful area of research. Expanding the types of enzymes and combining multiple enzymes into more complex networks are necessary to broaden the scope of accessible network motifs and to realize the theoretical motifs not found in real networks yet. Currently, only two out of seven EC classes of enzymes (oxidoreductases and hydrolases) are used as core reactions in complex ERNs.
An important open question is the design of an ERN which would sustain oscillations under batch conditions (stirred or nonstirred), an enzymatic analogue of the Belousov–Zhabotinsky (BZ) reaction.247 The oscillators described in the sections 3.1–3.3 rely on fluxes of reagents (by means of flow reactors or transport processes between compartments). In contrast to the BZ reaction, all enzymatic networks reported thus far consume all “fuel” in a single reaction cycle under well-stirred conditions, thus making it impossible to sustain oscillations in batch. In general and as highlighted by the nonautocatalytic HRP/GOx network example, it is still an open fundamental question what network motifs are required to yield a certain desired type of nonlinear behavior. A strong coupling between experimental and theoretical studies is needed in this regard, especially when incorporating diffusion and other transport phenomena.
Influence of External Stimuli on ERNs
It is difficult to engineer the properties of complex enzymatic reaction networks as their dynamic output requires all reaction rates to be adjusted to each other, while the formation of byproducts or the emergence of hidden interactions could prevent or disrupt the dynamic behavior of the system.81 Fine-tuning rates by controlling the activity of individual enzyme activities is therefore desired. Multiple methods were developed to gain control over enzyme activity by applying an external stimulus such as light, pH, redox potential, heat, magnetic field, and sound.81−83 In this section, we will focus on the use of external stimuli to control enzymatic reaction networks that consist of two or more enzymes.
Light
Light irradiation in combination with photocleavable or photoswitchable molecules allows for selective, rapid, and spatiotemporal control over enzyme activities.81,84,85
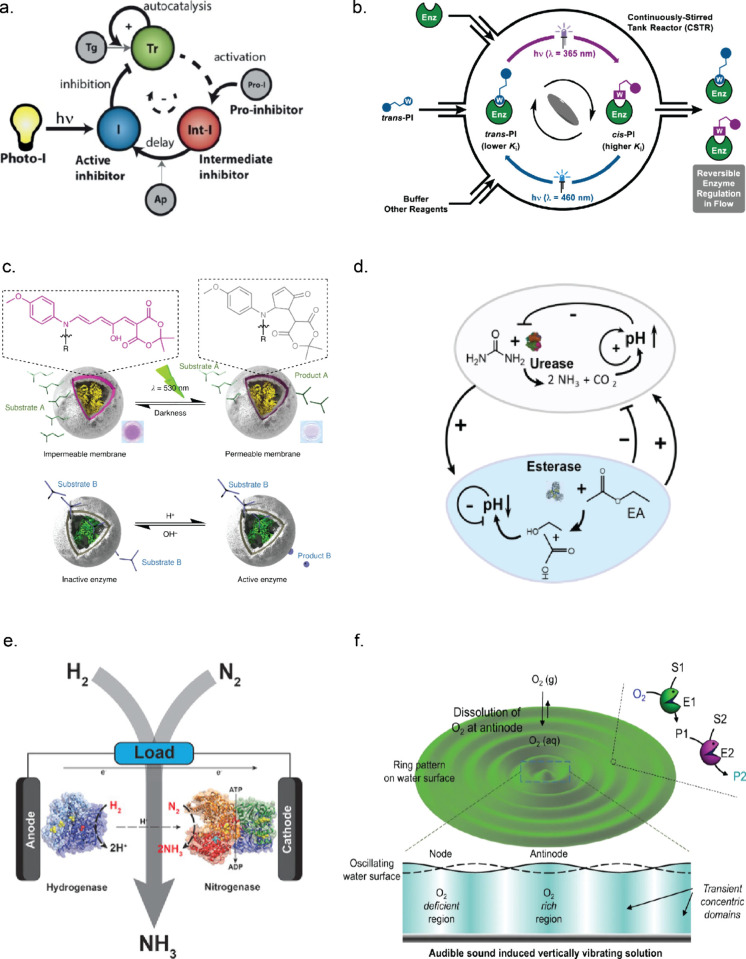
Pogodaev et al. used ultraviolet (UV) irradiation of a photocaged irreversible inhibitor to obtain control over the oscillations of the Tr oscillator discussed in section 3.2 and presented in Figure a.86 The oscillations were dependent on the duration and timing of UV light, demonstrating that external stimuli can introduce extra complexity without disrupting the rest of the system. However, for many enzymatic systems it is desirable to obtain reversible control over the enzymatic activity. The groups of Feringa, König, and Szymanski have reported many examples where photoresponsive molecules are used to reversibly inhibit specific enzymes upon light irradiation.83,85,87 For this, photoswitchable molecules such as azobenzenes, spiropyrans, diarylethenes, and donor–acceptor Stenhouse adducts (DASAs) could be used.85
The method to use photoswitchable molecules in combination with light is known as the chromophore-warhead strategy, where one of the two conformations acts as a stronger inhibitor. The warhead, a functional group that is known to inhibit the enzyme, can be covalently attached to a chromophore that can change their 3D structure upon light irradiation.85 Using this strategy, Teders et al. recently designed an azobenzene-based inhibitor for Tr and Cr.88,89 The activity of these enzymes could be reversibly adjusted under flow conditions upon different light intensity, duration, and wavelength of light, see Figure b. These systems exhibited an ultrasensitive response, illustrating potential for using light as an input for controlling complex dynamics in ERNs.
In addition to directly photoswitching molecules that interact with the enzyme (via the active site or allosteric interactions), one could alter the kinetics by controlling the accessibility of the substrate toward the enzyme, as demonstrated by encapsulating enzymes in vesicles or polymersomes and controlling the transport of substrate using light.90−93 The latter example, presented in Figure c, was based on polymersomes containing a DASA dye in the polymer shell, allowing switching between semipermeable states.93 These DASA polymersomes were filled with Est and then mixed with Ur containing polymersomes that were permeable and light insensitive. Upon green light irradiation, the Est–DASA polymersomes became semipermeable, which resulted in the conversion of ethyl acetate into acetic acid by Est. A concomitant pH decrease from pH ≈ 8 to pH ≈ 7 activated urea hydrolysis by Ur, which resulted in a pH rise. The pH increase by Ur resulted in the deprotonation of a pigment that absorbs green light in the protonated form. Therefore, the absorption of green light stopped, and the Est activity could be reactivated.
pH
All enzymes have a specific pH value at which their activity is maximized.94,95 In pH-dependent networks, the pH can be altered by addition or generation of acid or base by another enzyme. Often used combinations contain Ur, Est, GOx, or HRP as sources of acid or base.48,59,96 As described by Che et al., the addition of chemical fuel can influence the biocatalysis of pH-sensitive polymersomes loaded with HRP or Ur.97 The polymersomes shrank in a high-pH buffer due to deprotonation, whereby they became impermeable and thereby inactivate the enzyme. Addition of hydrochloric acid (HCl) and urea (fuel) resulted in a decrease of pH, swelling of the polymersomes, and activation of the enzymes. The depletion of urea resulted in shrinkage of the polymersomes and a decrease in the enzymatic activity. The activation cycle could be repeated multiple times by addition of acid and fuel to create a pH-dependent system. Maity et al. constructed an Ur/Est loop by encapsulating Ur and Est in different hydrogel beads that formed pH fronts.65 As described in section 3.1, the conversion of urea by Ur results in a pH increase which is counteracted by the conversion of ethyl acetate into acetic acid by Est, see Figure d. The same pH-dependent enzymatic reaction network was used to create enzymatic logic gates.98
Wang et al. developed a DNA-based hydrogel that contained GOx, acetylcholine esterase (AchE), and Ur to alter the pH.99 GOx and AchE activity resulted in a decrease in pH, while the hydrolysis of urea by Ur increased the pH. Different enzyme compositions allowed control over the pH and thereby the stiffness of the hydrogel. Control over enzymatic batch processes was obtained with acid-producing enzymes and poly(methacrylic acid) (PMAA)-functionalized gold particles. First, these particles were dispersed in the reactor, and reaction-induced pH changes were allowed to take place. When the buffer capacity was exceeded, protonation of the PMAA led to aggregation of the functionalized gold nanoparticles, which could be resuspended upon addition of fresh buffer and substrate.100 Instead of changing the pH of the bulk solution, Zhang and co-workers showed that the enzyme kinetics of immobilized enzymes could also be influenced by pH changes in the microenvironment.95 The local pH around cytochrome C, which is most active under acidic conditions, could be lowered by immobilization on negatively charged high-density polyelectrolytes that can attract ions of opposite charge. The throughput of a cascade with immobilized cytochrome C and d-amino acid oxidase was increased 10-fold by lowering the local pH of cytochrome C in an alkaline environment where d-amino acid oxidase is most active.
Electrochemical Control
The redox potential can be used as an external stimulus to control the activity of enzymes.101−103 Mallawarachchi et al. showed that the accessibility of the substrate toward the active site of hexokinase (HK) entrapped in an electroresponsive hydrogel was altered upon application of an electrochemical potential due to the reversible contraction and expansion of the hydrogel.104 The reaction kinetics of enzymes can also be modified by trapping enzymatic cascades in a porous conducting metal oxide electrode material. This was done by Morello et al. to reversibly recycle a nicotinamide cofactor that was generated with an enzymatic cascade consisting of ferredoxin NADP+ reductase, l-malate NADP+ oxidoreductase, fumarase, l-aspartate ammonia-lyase, and carbonic anhydrase.105 It was shown that the reaction direction and rate of ferrodoxin NADP+ reductase could be regulated based on the electrochemical potential which allowed control over the synthesis of aspartic acid from pyruvic acid or the reverse reaction. Furthermore, Milton et al. used a combination of nitrogenase and hydrogenase enzymes to electrify the production of NH3 from N2 and H2, which is depicted in Figure e. In this work, methyl viologen was used as an electron transfer agent between the enzymes and the electrodes to produce NH3 and an electrical current at the same time.106
Heat
Heat can be used as a parameter to influence the dynamics of ERNs since the rate constants of enzymes are dependent on temperature.107,108 As mentioned in section 3.2, Maguire et al. studied the influence of temperature on the Tr oscillator. They investigated the effect of temperature perturbations under conditions close to the so-called tipping point, the boundary between the oscillatory and the steady state regime.68 Close to the boundaries of the stable oscillatory regime (between 15 and 38 °C), sensitivity to short temperature perturbations increased, causing the recovery time to stable oscillations to increase as well. In another study, they investigated the use of small temperature oscillations to regulate the periodicity of the Tr oscillator.69 For temperature oscillations with an amplitude of just 3 °C, the periodicity of stable oscillations could be shifted to match the externally induced periodicity through a process known as phase locking or synchronization. Outside the phase-locking regime, quasi-periodic behavior was observed. Since the temperature can have a significant influence on the dynamic behavior of a network, a method to locally adjust the temperature was developed by Zhang et al. This involved the embedding of enzymes on platinum nanoparticles decorated with thermoresponsive copolymers, allowing local heating by irradiation with near-infrared light. The polymer–enzyme hybrids formed aggregates in solution below a certain temperature and disassembled above that temperature, thereby adjusting the accessibility of the substrate toward the active site of the enzyme. In general, the formation of aggregates resulted in a decreased enzymatic activity compared to the disassembled state of the particles.108 Thermoresponsive polymers were also used by Gobbo and co-workers to reversibly regulate enzyme activity by contractions of a polymer–protein-based protocell upon temperature changes.109
Other Control Factors
The Katz group presented an enzymatic system where a magnetic field was used to induce local pH changes, yielding reversible control over the enzyme activity.110 Here, two types of enzyme-modified magnetic nanoparticles that contained amyloglucosidase (AMG) and Ur or Est were used. Upon applying a magnetic field, the nanoparticles with AMG and Ur displayed aggregation. Due to the aggregation, the enzymes were in closer proximity to each other and therefore showed higher enzyme activity. The AMG activity was monitored by a cascade reaction with GOx and HRP.
Furthermore, sound can be used as an external stimulus to control the dynamics of enzymatic cascades. Gradients and patterns of vibrations in an aqueous medium form upon application of sound, see Figure f. Dhasaiyan et al. found that at the minima of the vibration, gas dissolution was lower than that at the maxima of the vibration. This was exploited in an enzymatic cascade dependent on dissolved oxygen. First, GOx-FAD was reduced to GOx-FADH2 by glucose consumption. The dissolved oxygen was used to produce H2O2 that can be used by HRP to oxidize ABTS (2,2′-azino-bis(3-ethylbenzothiazoline-6-sulfonic acid). ABTS is a colorful compound that was used to visualize gradients in the enzyme activity of the cascade, see Figure f.111 Please note, the compartmentalization of enzymes or the immobilization of enzymes on different types of support can enhance the reaction rate and therefore the total output of an enzymatic cascade. For more information about compartmentalization,120−122 in DNA nanocages,123−128 immobilization in metal organic frameworks (MOFs),129−133 or immobilization of enzymes on other supports,112−119 we would refer to other reviews.
Designing “Life-Like” Systems Using ERNs
In previous sections, we discussed how ERNs can exhibit intricate, complex features such as feedback loops, bistability, ultrasensitivity, or adaptive behavior. Employing these complex features of enzyme-catalyzed reactions to trigger morphological transformations and modulate the physical properties could lead to the development of so-called life-like systems. In this section, we will delve into specific examples that highlight the design of complex systems utilizing enzymatic networks.
Dynamic Materials
Enzyme-catalyzed reactions typically occur under mild conditions and show high substrate specificity, making enzymes an attractive choice for the design of new dynamic materials. To make the link to materials, researchers studied the influence of enzymatic reactions on polymer chains in order to induce self-assembly or disassembly or other morphological reorganizations.134,135 For example, Amir et al. reported the use of acid phosphatase to cleave phosphate moieties of hydrophilic monomers, resulting in the formation of an amphiphilic diblock copolymer of polyethylene glycol (PEG) and poly(4-hydroxystyrene) that assembled into spherical micelles.136 In another example, the autocatalytic nature of urea-Ur was harnessed to control base-catalyzed thiol-Michael addition reaction for time-lapse hydrogelation.63 Enzyme-catalyzed reactions have been utilized to initiate polymerization of monomeric building blocks.137,138 Mao et al. used acid phosphatase and GOx to make hydrogels out of supramolecular and polymeric networks.139 A similar strategy was later implemented to design printable hydrogels using GOx and HRP.140 In this study, an acrylic acid-modified hydrogelator (NapFFK-acrylic acid) was used as a monomer, undergoing self-assembly to form hydrogels (gel-I, Figure a). GOx converted glucose to gluconic acid and subsequently reduced O2 to H2O2. Next, HRP utilized H2O2 to catalyze the formation of acetylacetone (AcAc) radicals via oxidation of AcAc. These radicals initiated polymerization of poly(ethylene glycol) methacrylate (PEGMA) with acrylic-modified hydrogelators, resulting in a cross-linked hydrogel (gel-II, Figure a). Gel-II showed very good mechanical properties with an approximately 16 times higher storage modulus compared to gel-I (without cross-linking). Klemperer et al. utilized a similar strategy to design cytocompatible bioinks with interpenetrating polymer network (IPN) formed under mild and aerobic conditions.138
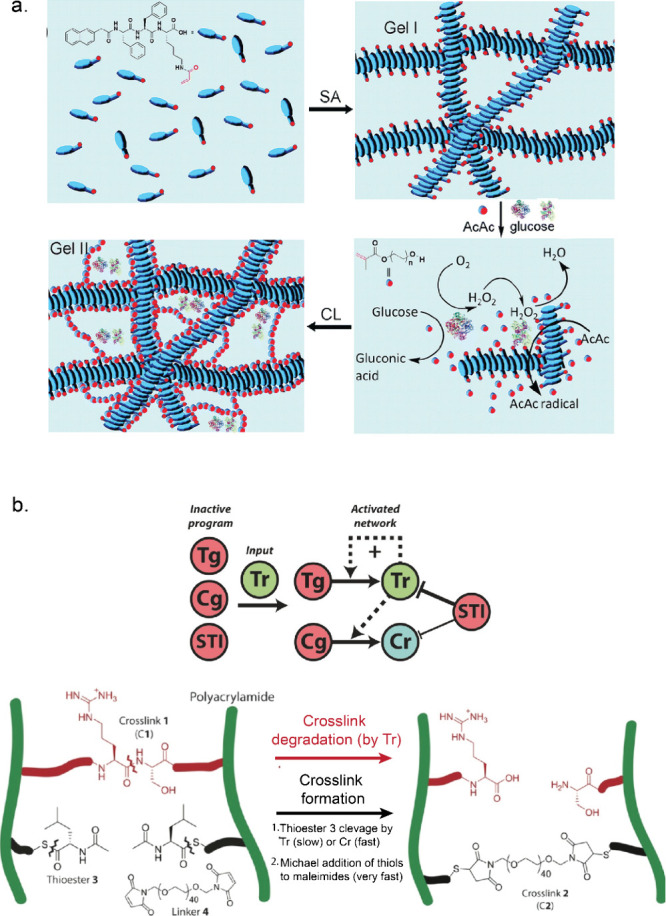
Moreover, enzyme-mediated de-crosslinking or cleavage has been exploited to install stimuli responsiveness in the presence of enzymes. Hydrogels containing enzyme-cleavable groups were used for designing responsive materials susceptible to enzymes like proteases, lipase, Est, and phosphatase activities.141−144 Yang et al. demonstrated for the first time an enzyme-regulated reversible gel–sol transformation. They utilized kinase–phosphatase switch to regulate assembly of peptide-based hydrogelator (Nap-FFGEY). The addition of kinase and adenosine 5′-triphosphate (ATP) led to phosphorylation of hydrogelator, disrupting self-assembly, while phosphatase reversed this process, restoring the self-assembly property.141 Interestingly, micelles formed from polymer–peptide block copolymers modified with substrates for different enzymes (protein kinase A (PKA), protein phosphatase-1 (PP1), and matrix-metalloproteinases MMP-2 and MMP-9) showed different morphologies and aggregation behavior upon enzymatic reactions.145 In addition, Postma et al. reported a systematic approach in designing adaptive matter by integrating reaction networks with materials.146 They designed a polyacrylamide (PAAm)-based hydrogel containing two orthogonal types of cross-links, which could either be enzymatically degraded or formed (Figure b). Tr rapidly cleaved the first cross-links, thus forming a liquid phase. Simultaneously, Tr slowly cleaved a copolymerized thioester (cross-link precursor), and these newly exposed thiol groups reacted rapidly with linker (poly(ethylene glycol)-bis-maleimide) to form a new gel. Upon addition of Tg and Cg, Tr was formed autocatalytically from Tg and thus speeding up the degradation and activation processes of the first and second cross-links, respectively. Tr also rapidly converted Cg into Cr, which induced even faster cleavage of copolymerized thioester. Finally, to set a threshold for activating the entire network, soybean trypsin inhibitor (STI) protein was introduced, which deactivated the enzymatic network by inhibiting Tr activity (Figure b).
Enzyme-catalyzed reaction products can also be used as triggering stimuli which influence polymeric assemblies to exhibit dynamic functions. For example, oxidases like GOx, sarcosine oxidase (SOx), choline oxidase (COx), and urate oxidase (UOx) generate H2O2 upon oxidation of their substrates. Hydrogels containing both hydrolases (AchE) and oxidases displayed a gel–sol transition in the presence of hydrolase substrate (acetylcholine) by the cascade generation of H2O2.147 In earlier sections, we have discussed how enzymes like Ur, GOx, and Est are capable of controlling the pH of the system environment. Walther and co-workers have demonstrated how enzyme-mediated pH feedback can be introduced to design stimuli-responsive smart materials. For example, Heuser et al. integrated Ur-mediated pH feedback to a pH-responsive photonic gel composed of polystyrene-b-poly(2-vinylpyridine) (PS-b-P2VP) diblock copolymers to control swelling of the photonic hydrogel, which in turn controls transient optical reflection.148 Moreover, enzyme-mediated pH change was used to design temporal materials where gelation is controlled by a change in pH via enzymatic reactions.59,65 To demonstrate pH-dependent transient gelation, Heinen et al. used a hybrid hydrogel made of DNA-containing polyacrylamide copolymer with pH-responsive DNA cross-linker (i-motif). This pH-responsive unit formed a tetraplex intramolecularly at low pH and as a result did not take part in cross-linking (sol state). With increasing pH, the unreactive tetraplexes unfold and induce hybridization with the copolymerized strand, resulting in a gel state. A sol–gel–sol transition was observed by the antagonistic effect of Ur-mediated pH increase and pH decrease by Est.59 In short, enzyme-catalyzed chemical reactions have a diverse range of applications, from enabling cross-linking polymerization under mild conditions to directly controlling material properties to install responsiveness.
Enzyme-Powered Motile Systems
The inspiration for designing motile systems stems from various biological motor proteins which operate via conversion of chemical energy into mechanical motion by the hydrolysis of ATP. Besides these motor proteins, enzymes have shown increased diffusion during substrate turnover and can impart forces (ca. 10 pN) quite comparable with biological motor proteins.149,150 In addition, an asymmetric distribution of reactant and product molecules during enzyme catalysis can create local gradients of concentration or an electrical field, which can lead to enhanced diffusion of enzymes.151 However, active movement of enzymes in the presence of substrates is part of an ongoing debate, and we refer to other literature sources for a more thorough discussion.152,153 A number of research groups have worked on endowing small micro/nanosized particles with enzymes in order to create motile systems.154−156 These particles achieved motility by enzyme-catalyzed chemical reactions. For example, Ur can show ionic self-diffusiophoresis due to the generation of a local electric field from diffusivity differences between the oppositely charged ions (NH4+ and CO32–) formed from urea.157 Ur has been extensively used as a catalytic engine to design self-propelling motors.158−160 On the other hand, oxygen bubble generation from H2O2 by Cat or a pair of enzymes like GOx/Cat can propel microswimmers opposite to bubble formation.161,162 One promising system is based on stomatocytes, bowl-shaped particles formed by deformation of polymersomes by osmotic pressure. During the deformation process, enzymes can be encapsulated in the nanocavity that is formed when the shell folds in on itself, and the enzymatic activity of the encapsulated enzymes provides a force that can propel these stomatocytes.163 For example, a metabolic network of six enzymes was compartmentalized in stomatocytes, which was responsible for converting glucose (fuel) into motion (Figure a). This network started with an ATP-mediated activation module containing HK and pyruvate kinase with phosphoenolpyruvate (PEP, phosphate donor) and glucose (energy source). Pyruvate (the product of the first cycle) triggered the pyruvate–l-lactate cycle where l-lactate dehydrogenase (LDH) consumed pyruvate. But, l-lactate oxidase (LO) catalyzed the reverse reaction. NADH was regenerated by the conversion of glucose-6-phosphate by glucose-6-phosphate dehydrogenase (G6PDH) into 6-phosphogluconolactone. This resulted in net H2O2 production, which was converted to molecular oxygen by Cat.164 Here, ATP determined the concentration of NADH, thus regulating the entire network. In another example, amyloid microphases loaded with alcohol dehydrogenase (ADH), SOx, and Cat exhibited microscopic motility. These self-assembled structures contained imidazole moieties, which hydrolyzed the 6-methoxy naphthyl alcohol ester of N-methyl glycine (starting substrate) to 6-methoxynaphthyl alcohol and N-methyl glycine. Subsequently, ADH converted 6-methoxynaphthyl alcohol to 6-methoxynaphthaldehyde (a fluorescent compound), while the cascade of SOx and Cat utilized N-methyl glycine to generate oxygen bubbles, responsible for the observed motility.165 Increased diffusive movement of free enzymes and enzyme-coated particles toward higher substrate concentration has been reported and extensively studied.150,166,167 The Wilson group designed Cat-powered PLGA (poly(lactic-co-glycolic acid) micromotors as chemotactic drug delivery vehicles, which can follow the H2O2 gradient produced by macrophage cells.168 Joseph et al. demonstrated how coupling enzyme cascades (GOx and Cat) with asymmetric polymeric vehicles could be used to design potential chemotactic drug delivery vehicles for crossing the blood–brain barrier.169
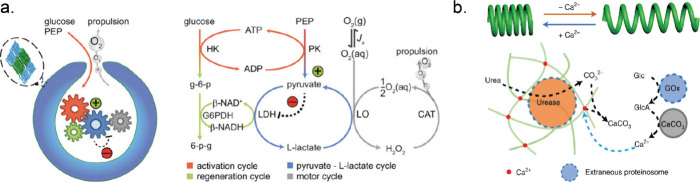
Aside from enzyme-powered self-propulsion, oscillatory movement was achieved in self-assembled organoclay/DNA semipermeable microcapsules containing Cat and GOx. In the presence of H2O2, Cat produced oxygen bubbles which were taken up by GOx in the presence of glucose. The generation and consumption of oxygen bubbles was responsible for sustained oscillatory movement. The system was able to achieve multiple oscillations under a continuous flow of glucose and H2O2.170 In other recent work by the same group, the interaction between protocells and their environment was explored by designing microactuators capable of chemically induced spring-like compression and relaxation.171 Helical filaments of calcium alginate containing Ur-loaded proteinosomes were fabricated using microfluidics. When these coiled hydrogels were exposed to urea, carbonate ions were produced and started chelating the calcium ions (Ca2+) from the hydrogel cross-links. This Ur-mediated removal of Ca2+ ions led to the release of the stored elastic potential energy, which in turn induced extension of the helical filaments. To contract these extended filaments, they were exposed to proteinosomes containing GOx. In the presence of glucose, gluconic acid was produced, which dissolves CaCO3 and releases Ca2+. This protocell-mediated Ca2+ flux in turn restored cross-links in the calcium alginate matrix which led to a slow retraction of the extended filaments (Figure b). These examples highlight current endeavors to utilize enzymes to control motility of artificial systems, which can be useful in designing intelligent sensors and novel therapeutic vehicles.189
Communication in Compartmentalized Bioreactors
Compartmentalization is an effective strategy to segregate enzymes catalyzing sequential reactions and can be helpful to increase productivity by concentrating substrates and avoiding unwanted side reactions.120,172,173 This spatial segregation, coupled with interactions between compartments with distinct chemistries, can further lead to the emergence of life-like properties in artificial systems like predator–prey behavior,174−176 chemical signaling,177−181 and chemostructural feedbacks.171,182 In this context, Elani et al. designed a multicompartment vesicle-based platform to spatially segregate reaction processes where chemical signals can transverse from one compartment to another. A three step cascade by lactase, GOx and HRP was conducted where each enzyme catalyzed step was isolated in distinct compartment.183 Wang et al. developed methodologies to design microarrays of giant unilamellar lipid vesicles (GUVs) containing GOx and HRP to facilitate controlled chemical signaling in the presence of melittin, a pore-forming peptide.178 Buddingh’ et al. demonstrated chemical communication between giant vesicles through allosteric signal amplification. They used glycogen phosphorylase b (GPb) which switched to a high-activity state upon binding adenosine 5′-monophosphate (AMP). The sender vesicle contained apyrase which produced AMP in the presence of ATP (Figure a). The produced AMP diffused to the receiver vesicle, where a small ERN comprising phosphoglucomutase (PMG), G6PDH, GPb, and glycogen was compartmentalized. As AMP acts as allosteric activator of GPb, it induced conversion of glycogen to glucose-1-phosphate, followed by the conversion of glucose-1-phosphate to glucose-6-phosphate by PMG, and subsequent generation of NADH by G6PDH (Figure a). This allosteric amplification of the initial weak signal from the sender vesicles, achieved through GPb activation, resulted in a robust response (high NADH output) in the receiver vesicles. Remarkably, the strong signal amplification engineered into this pathway allowed long distance (5 mm) communication between senders and receiver vesicles.179
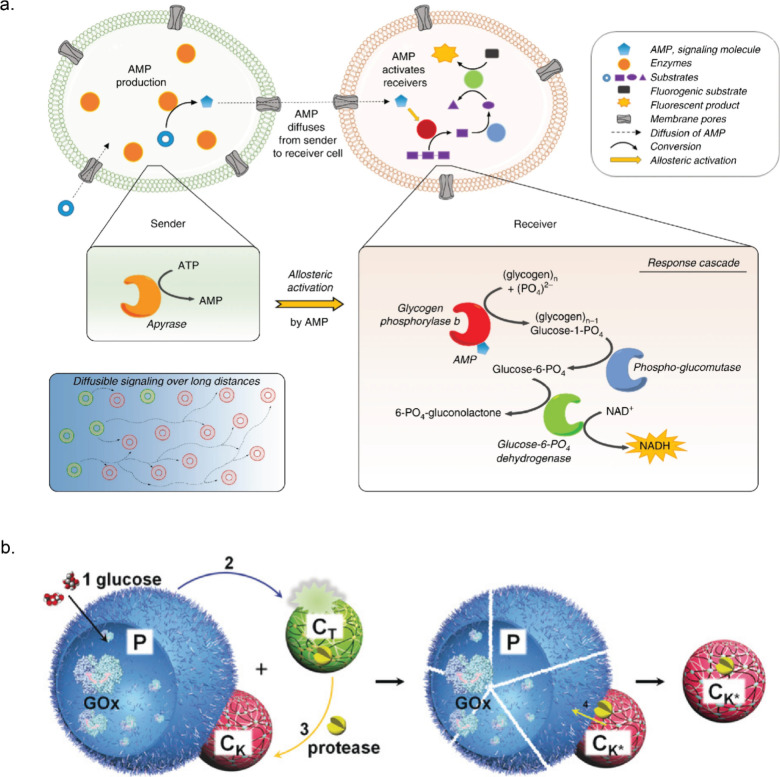
Qiao et al. achieved artificial response–retaliation behavior using ternary protocell populations. These populations included proteinase K-sensitive GOx-containing proteinosomes (P) that released H+ in the presence of glucose.175 These also consisted of small pH-sensitive proteinase K-containing polypeptide (poly-d-lysine)/adenosine 5′-diphosphate (ADP) coacervates (CT) and pH-resistant positively charged polymer (poly(diallyldimethylammonium chloride), PDDA)/polysaccharide (dextransulfate, DS) coacervate droplets (CK) that adhered to negatively charged proteinosomes via electrostatic interactions (Figure b). Proteinase K was initially sequestered in CT coacervates, but the addition of glucose led to gluconic acid release from P, lowering the pH and causing disassembly of the coacervates CT. The released proteinase K was then sequestered by the proteinosome-adherent coacervates CK. Finally, proteinosomes were destroyed by proteinase K, and only CK remained. Expanding on the idea of communication between compartments, Chakraborty et al. demonstrated how a bioluminescent signal triggered prey–predator behavior in GUVs. Renillaluciferase (RLuc) containing GUVs (sender) produced blue light in the presence of coelenterazine, activating iLID and Nano proteins in the outer membranes of both sender and receiver GUVs. The interaction between iLID and Nano under blue light mediated the adhesion between the GUVs. GUV–GUV adhesion allowed transfer of Ca2+ from the sender to the receiver through unblocked α-hemolysin (α-HL) pores. Ca2+ activation of phospholipase A2 (PLA2) in prey GUVs led to phospholipid cleavage and collapse of receiver GUVs.176 A recent study from the same group showcased bidirectional communication in GUVs through similar chemiluminescence-triggered adhesion, enabling exchange of H2O2 from the sender and Ca2+ from the receiver. Interestingly, GUVs separated when the signaling molecule production ceased.180 Liu and Zhang et al. designed a three-layer tubular prototissue comprising concentrically arranged agarose hydrogel layers containing GOx, HRP, and CAT containing coacervates as the outer, middle, and inner layers, respectively. Glucose and hydroxyurea were added specifically to the exterior side of the model prototissue. Through inward diffusion, glucose was first processed by GOx containing a hydrogel layer, leading to H2O2 production. HRP in the middle layer converted hydroxyurea in the presence of H2O2 into nitric oxide (NO). Excess H2O2 was then consumed by Cat in the final layer, such that NO became the main output, which was further used to inhibit blood coagulation in samples located within the device’s internal lumen.184
Interestingly, enzymes fixed in a microchamber can act as “chemical pumps” in the presence of specific substrates. Density differences between the substrates and the enzyme-catalyzed products give rise to solutal buoyancy, in turn generating convective flows of the enclosed fluids.185,186 This system can be understood as follows: when the density of the enzyme-catalyzed products exceeds that of the reactants, the fluid becomes denser, sliding down and away from the patch like an outward pump. Conversely, if the product density is lower, the fluid rises up, falls back down due to convective flows, and moves toward the enzyme patch, forming an inward pump. The fluid flow velocity is dependent on the enzymatic reaction rates. These micropumps have been utilized as proof-of-concept delivery vehicles to transport small molecules like insulin in the presence of a glucose stimulus or act as sensors to detect toxins that hamper enzymatic reactions.185,187 Furthermore, simulations on flexible sheets with two enzyme patches have revealed complex motility behaviors. The first patch produces substrates for the second, while the products from the second patch inhibit the enzymes in the first patch. The spatial separation between these two enzyme patches led to the time delay required for chemical oscillations in the system. The resulting fluid flow induced oscillatory mechanical deformation, displaying diverse motility behaviors.188 These strategies utilizing ERNs to enable communication between spatially segregated bioreactors are crucial for designing systems with diverse life-like applications.
Cell-Free Synthesis
Cell-free biosynthesis is a rapidly growing field in which complex biochemical pathways are assembled in vitro. The aim of this field is to better understand how such pathways function and how they can be manipulated. The reconstruction of these pathways in vitro also provides a route to the synthesis of valuable organic molecules using environmentally friendly processes and without the need for culturing organisms. Reaction networks carried out in cell-free environments can, in principle, be much better controlled by optimizing the pH, temperature, and enzyme loading. Furthermore, competition with other enzymatic pathways is excluded, and enzymes produced in various organisms can be combined into a single pathway.190 In this section, we will focus on the biochemical pathways that consist of multiple enzymes. For the two-step enzyme cascades, we refer the reader to the reviews in refs (ref. 172) and (ref. 191−ref. 193) and specifically for enzymatic cofactor regeneration to the reviews in refs (ref. 194) and (ref. 195). Importantly, only cell-free synthesis of small molecules is covered in this review; cell-free protein synthesis has been the topic of several reviews.196−198
Sugars as Substrates
Bowie’s group demonstrated how complex enzymatic chemical reaction networks can be used to synthesize valuable molecules starting from simple substrates such as glucose or phosphoenolpyruvate (PEP) (Figure a). All reactions were performed using purified enzymes in batch conditions. One of the first complex enzymatic reaction pathways included isoprene synthesis, which was synthesized from PEP in a combined glycolysis and mevalonate pathway containing in total 12 enzymes.199 ADP, NADPH, and coenzyme A (CoA) regeneration cycles were employed to reuse the cofactors and to avoid CoA build up, which was causing inhibition of the forward direction of the mevalonate pathway. After this, the same group showed how a 27-enzyme cell-free system in batch converts glucose into different monoterpenes using glycolysis and mevalonate.200 The choice for the final enzyme in the pathway, a limonene synthase, pinene synthase, or limonene synthase-N345A mutant, determined which monoterpene (limonene, pinene, or sabinene) was produced. The whole pathway also included a molecular purge valve (Figure b).201 Molecular purge valves maintain cofactor balance in the system without the need to perfectly match stoichiometric cofactor formation and carbon consumption. Once there is a NADPH cofactor buildup, the NADP+-dependent reductase enzyme is starved of oxidized cofactor and the pathway shuts down. In the meantime, the purge valve is turned on and NAD+-dependent reductase and NADH-specific oxidase are activated. In this case, it was used to control the balance of NAD(P)H/NAD(P)+ using three enzymes: glyceraldehyde-3-phosphate dehydrogenase (Gap), mutant glyceraldehyde-3-phosphate dehydrogenase (mGap), and NADH oxidase (NoxE). This cascade regenerates NADPH, purges the excess of NADH, and continues generating carbon building blocks for the glycolysis pathway. It was demonstrated that excluding at least one enzyme of the molecular purge valve reduced limonene production drastically, while a system without the NoxE enzyme did not convert any glucose to limonene. Researchers managed to produce all three terpenes starting from 500 mM glucose over 7 days at high titers.
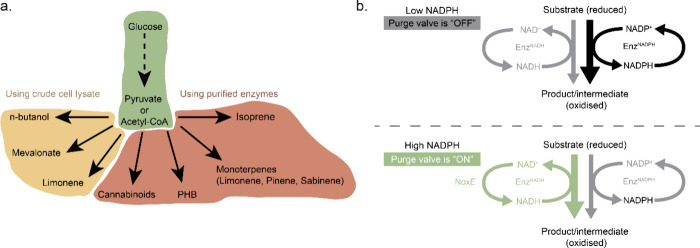
Another example of synthetic pathways producing valuable organics includes the so-called PBG (pentose–bifido–glycolysis) cycle in which the final product is polyhydroxybutyrate (PHB), a bioplastic.202 The whole cycle consists of 20 enzymes from three partial pathways: the pentose phosphate pathway, the bifidobacterium shunt, and the glycolysis pathway. This ERN also contained two purge valves to regulate NAD(P)H concentrations as well as a metabolite salvage pathway which allowed erythrose-4-phosphate to re-enter the cycle. Two new enzymes, a G6PDH and a 6-phosphogluconate dehydrogenase (Gnd) mutant, had to be engineered for the purge valve to favor NAD+ instead of NADP+, as their wild forms do. A batch reaction containing all enzymes was productive for up to 55 h, and only the last enzyme of the cycle—PHB synthase (PhaC)—had to be added again after each 10 h cycle. This was done because the enzyme was covalently linked to the growing end of the PHB product, which meant the enzyme would be removed every time during sampling of the bioplastic out of the reaction vessel to quantitatively measure produced PHB. Valliere et al. constructed an even more complex pathway consisting of 23 enzymes, which was used to synthesize a variety of prenylated molecules, including cannabinoids, from glucose.203 In their work, a modified glycolysis module was developed which included a purge valve to balance NADPH concentration and carbon flux. This module was connected to acetyl coenzyme A (acetyl-CoA) and mevalonate modules to form geranyl-pyrophosphate (GPP). From there, different substrates and their complementary enzymes were added to yield various prenylated compounds. Performing this pathway using only purified enzymes is also advantageous as GPP is toxic to cells in medium concentrations.204 The original system containing pyruvate dehydrogenase (PDH) had to be modified as it was found that PDH was inhibited by 1,6-dihydroxynaphthalene (1,6-DHN), which is the preferred substrate for wild-type prenyltransferase (NphB) enzyme. Therefore, a PDH bypass system consisting of two additional enzymes to convert pyruvate via acetyl phosphate to acetyl-CoA was introduced. After it was proved that various prenylated aromatic compounds can be generated using prenyl transferases, the authors focused on optimizing cannabinoid production. The results showed that the production rate of the cannabinoid precursor cannabigerolic acid (CBGA) using the primary system reached a not particularly high titer. However, a new mutant CBGA synthase (M23) was designed to produce CBGA using olivetolic acid (OA) without side products, which helped to reach a higher titer. In combination with in situ product removal using a flow system, a final CBGA titer was increased even more.
All of these examples demonstrated complex cell-free purified enzymatic reaction networks and possibilities for future industrial applications. Obviously, some important aspects of any industrial application include optimization, cost reduction, and yield increase. The authors proposed employing stable enzymes that could be used for longer periods of time in order to reduce costs, introducing cofactor (ATP/NAD(P)H) regeneration cycles to increase yields,204 recycling enzymes, and introducing cheaper purification methods.202 Another potential improvement could come from enzyme immobilization, which allows for easier purification and optimization at a cost of potentially reduced enzyme activity upon immobilization.
Instead of using purified enzymes, in some networks it would be desirable to construct ERNs in crude cell lysate. Karim et al. established a cell-free protein synthesis-driven metabolic engineering (CFPS-ME) framework for fast enzyme selection and pathway construction.205 As a model system, they chose a CoA-dependent n-butanol pathway starting from glucose including 17 steps in vitro. For this, the authors combined cell-free metabolic engineering (CFME), where heterologous enzymes are selectively overexpressed, and cell-free protein synthesis (CFPS). First, it was shown that mixing five crude lysates each of them with different selectively overexpressed enzymes via CFME can activate the whole pathway and produce n-butanol. Next, CFPS was modularly integrated in CFME to speed up the biosynthesis. Hydroxybutyryl-CoA dehydrogenase from Clostridia beijerinckii (Hbd2) lysate was excluded from the mixture of overexpressed enzymes, and instead, the CFPS was performed for Hbd2. As the downstream enzymes could not be activated without their substrates, the pathway would stay inactive until Hbd2 was synthesized. Therefore, first, CFPS of Hbd2 was performed for 3 h, and glucose was added to initiate the n-butanol synthesis pathway. Later, a similar experiment was repeated by excluding every other one of the five n-butanol pathway enzymes from overexpressed enzyme lysate mix and synthesizing them using CFPS for 3 h. After adding glucose, NAD, and coenzyme-A (CoA) to initiate the CFPS–ME pathway reactions, all variants produced n-butanol. This study demonstrated how the CFPS–ME method can be used for fast enzyme screening for ERNs. The same group also demonstrated the synthesis of mevalonate from glucose using enzyme-enriched lysate mixtures206 and investigated a pathway from mevalonate to limonene without any optimization.207 This research laid the groundwork for a combined study involving a full pathway from glucose to limonene using enzyme-enriched Escherichia coli (E. coli) lysates requiring in total 20 metabolic steps.208 To begin with, the endogenous metabolism in E. coli crude cell lysate already consisted of all of the glycolytic enzymes which transformed glucose to acetyl-CoA. This part was then connected to a single extract enriched in three enzymes—acetyl-CoA acetyltransferase (ACAT), hydroxymethylglutaryl-CoA synthase (HMGS), and hydroxymethylglutaryl-CoA reductase (HMGR)—converting acetyl-CoA to mevalonate. Last, it was coupled to a module of six extracts each enriched in a single enzyme using heterologous overexpression in vivo—mevalonate kinase (MK), phosphomevalonate kinase (PMK), pyrophosphomevalonate decarboxylase (PMD), isopentenyl pyrophosphate isomerase (IDI), geranyl diphosphate synthase (GPPS), and limonene synthase (LS). After additional experiments and optimization, the full pathway produced final molecule limonene.
Cell-free biosynthesis points out attractive possibilities of performing enzymatic synthesis and creating complex ERNs without any cell membrane constraints and avoiding cell toxicity. This growing field offers the opportunity of producing valuable compounds using sustainable biosynthesis. In the ethanol industry, commonly accepted thresholds required for industrial production are yields of 90%, productivity of 1 g/L/h, and titers of 40 g/L.202 Some of the discussed network final product titers are 12.5 ± 0.3, 14.9 ± 0.6, and 15.9 ± 0.4 g/L and yields of 88.3 ± 4.6%, 103.9 ± 8.1%, and 94.5 ± 4.7% for limonene, pinene, and sabinene, respectively.200 These final product values are close to the threshold values used in the ethanol industry, and it gives high hopes that cell-free biosynthesis will be applied for industrial purposes.
CO2 as Substrate
In addition to glucose, CO2 is also a versatile starting material for the sustainable synthesis of valuable compounds. Many inorganic and metallic catalysts, like SnO2, TiO2, Cu, and transition metal complexes, have been successfully developed as catalysts to reduce CO2.209 These inorganic catalysts in general show low selectivity; hence, the products are mainly limited to C1 and C2 compounds, including methanol and ethanol. Nature has evolved several metabolic networks to reduce CO2 into higher carbon compounds. However, many of the key carboxylase enzymes that couple CO2 to organic substrates have low efficiency.210−212 A new-to-nature carboxylase, glycolyl-CoA carboxylase, was recently developed and used in the successful in vitro implementation of the previously hypothesized tartronyl-CoA (TaCo) pathway, which had been proposed as a direct route for the assimilation of glycolate into central carbon metabolism and expected to outperform all naturally evolved glycolate assimilation routes.213 In contrast, in vitro construction of artificial CO2 fixation ERNs by freely combining different enzymes from various biological sources is another promising approach. The Erb group assembled the so-called crotonyl–coenzyme A (CoA)/ethylmalonyl-CoA/hydroxybutyryl-CoA (CETCH) cycle with 17 enzymes from nine different organisms to convert CO2 into malate at a rate of 5 nmol of CO2 per minute per milligram of protein, providing a seventh synthetic alternative to the six naturally evolved CO2 fixation pathways (Figure a).214 This is comparable to the CO2 fixation rate (1–3 nmol of CO2 per minute per milligram of protein) of the Calvin–Benson–Bassham (CBB) cycle, which fixes more than 90% of carbon in nature.215 Moreover, this CETCH cycle could be solar powered through the incorporation of chloroplast extracts.216 Subsequently, the output glyoxylate from the CETCH cycle can be applied to produce acetyl-CoA, the central precursor for many natural compounds. Hence, a series of value-added compounds, including monoterpenes, sesquiterpenes, polyketides, and 6-deoxyerythronolide B, has been successfully produced from CO2 in one pot based on the CETCH cycle.217,218 In addition, a small ERN with four enzymes (pyruvate carboxylase–oxaloacetate acetylhydrolase–acetate-CoA ligase–pyruvate ferredoxin oxidoreductase), named the POAP cycle, was constructed to couple two CO2 into one oxalate at the expense of two ATP and one NAD(P)H, reaching a rate of 8 nmol/min/mg CO2-fixing enzymes (Figure b).219 For a more detailed overview, the enzymatic conversion of CO2 has been discussed in a recent review on the topic.220
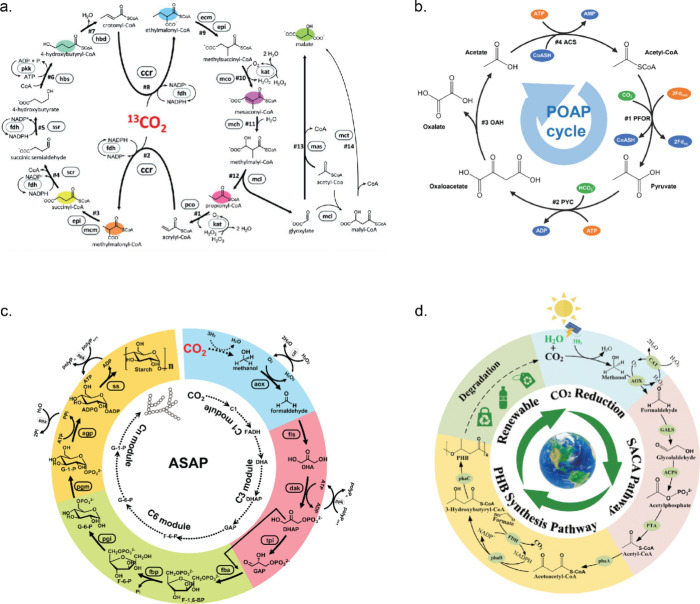
Other Substrates
In addition to using sugar and CO2 as substrates, progress has been made on fabricating ERNs to upgrade sustainable C1 and C2 stocks, like methanol, ethanol, and acetate, to compounds with longer chains and hence higher value. Cai et al. designed and constructed an 11-enzyme ERN to convert C1 methanol to polymeric starch (Figure c).221 Zhou et al. designed several ERNs to convert methanol to ethylene glycol, glycolic acid, and d-erythrose.222 In addition, the Bioplastic PHB has been produced from methanol and acetate (Figure d).223,224 Furthermore, Liu et al. managed to use ethanol to produce acetyl-CoA and regenerate ATP using the isoprenol production pathway.225
Future Development
The ERNs discussed so far exhibit a diverse range of functionalities and emergent properties. Yet, they remain relatively small based on several reactions occurring at once. However, as the sections on control over enzyme reactivity and the construction of elaborate reaction cascades for the synthesis of complex organic molecules have demonstrated, the field is now in a strong position to develop ERNs with much more advanced functionalities. In this section, we will discuss two such possible functions (without the ambition of providing complete overviews of those fields): synthetic cells and molecular computers.
Toward Building a Synthetic Cell from the Bottom Up
Building a synthetic cell from the bottom up could address the fundamental questions on how life emerged and evolved from nonliving components and at the same time generate a wide range of applications. Any functioning synthetic cell will inevitably require the construction of (minimal) metabolic ERNs for the continuous supply of building blocks and energy to support other cellular processes and also response to external environmental changes.226 ATP fuels many cellular processes and also serves as a building block for gene transcription and translation. Lee et al. successfully designed and built a photosynthetic artificial organelle consisting of an ATP synthase and two photoconverters in a giant vesicle. This synthetic organelle was capable of harvesting light to produce ATP and to drive endergonic reactions, like pyruvate carboxylase-mediated carbon fixation and actin polymerization into filaments (Figure a).227 Similarly, Berhanu et al. designed a light-harvesting artificial organelle, which was composed of two kinds of membrane proteins—bacteriorhodopsin and F-type ATP synthase. In this work, the organelle, reconstituted inside a GUV together with a cell-free protein synthesis system, generated ATP required in transcription and at the same time powered the synthesis of GTP and translation (Figure b).228 Additionally, Bailoni et al. reconstituted the l-arginine breakdown pathway of three cytosolic enzymes to phosphorylate ADP into ATP to fuel the sustainable formation of phospholipid headgroups in synthetic cells.229 Blanken et al. managed to encode seven phospholipid-producing enzymes in a synthetic minigenome, which were then expressed within cell-like liposomes. This de novo-constructed ERN can make use of fatty acyl coenzyme A and glycerol-3-phosphate as precursors to produce two phospholipids phosphatidylethanolamine (PE) and phosphatidylglycerol (PG) simultaneously (Figure c). The balance of PE and PG is transcriptionally regulated by the activity of specific genes together with a metabolic feedback mechanism.230 In addition, Partipilo et al. constructed a small ERN with three enzymes in liposomes to control the redox status of NADH and NADPH cofactors.231
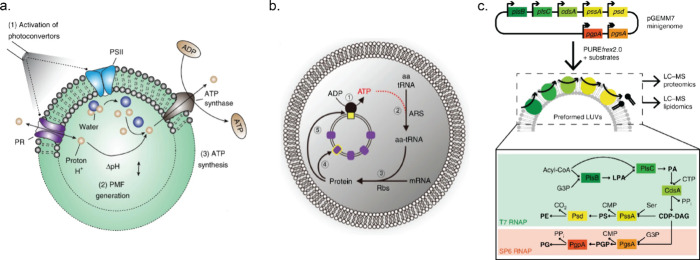
Living cells maintain physicochemical homeostasis (including pH, ionic strength, osmotic pressure) to enable the internal components to function near their optimum. This homeostasis state was successfully achieved inside a cell-like system by Pols et al., who co-reconstituted the arginine-breakdown pathway and an ionic strength-gated ATP-driven osmolyte transporter inside vesicles. The cell-like vesicles shrank under increased medium osmolality, causing the internal ionic strength and pH to increase, leading to the inactivation of enzymes. Once the internal ionic strength reached a critical value, the transporter was activated and glycine betaine was pumped in with the accompaniment of the influx of water into the vesicles. Hence, the volume of vesicles increased, and the internal ionic strength and pH decreased. As a result, homeostasis of the ATP/ADP ratio, internal ionic strength, and pH was achieved.232 All of these examples clearly demonstrate how enzymatic modules are crucial for mimicking complex metabolic functions of living systems in the field of designing synthetic cells.
Information Processing and Computation
An emerging direction for the research on and application of ERNs is special-purpose information processing and computation. So far, the development of Boolean gates and logic circuits has seen a long tradition in enzymatic network research and has been extensively reviewed several times (examples are shown in Figure a).233,234 However, the development of systems based on digital paradigms has been slowing down, as the principles of Boolean logic and von Neumann computer architectures are challenging to adapt in biochemical systems, which are intrinsically of a nonbinary and dynamic nature. Instead, the potential application of analog and neuromorphic computation principles in biochemistry is increasingly appreciated.235 While these developments are still in their infancy, here we focus on recent progress in this direction.
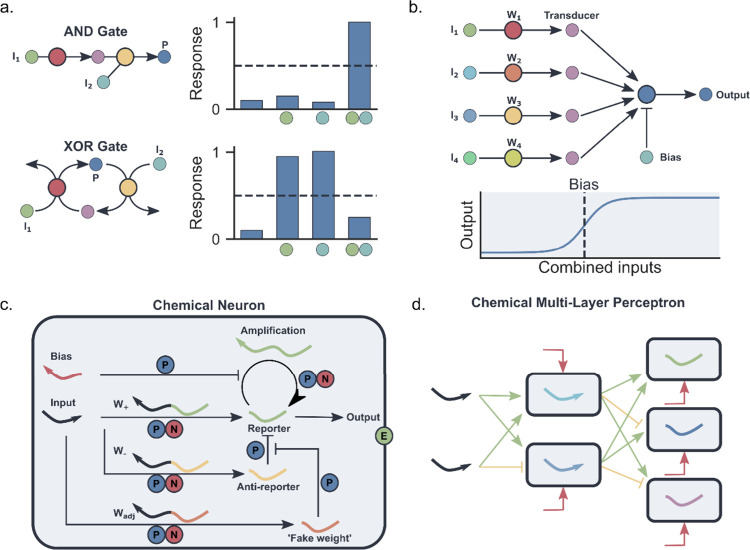
Rather than recreating digital infrastructure directly, recent work on small ERNs has aimed to establish modules and motifs capable of analogue computation. One example of this shows the development of small ERNs that are capable of arithmetic operations.236 The key insight here is that specific enzymatic interactions, such as cascading pathways, inhibitors, and parallel product conversion, can be interpreted as arithmetic functions if operated in the right parameter regime. Similarly, specific forms of substrate competition between different enzymes can lead to nonlinear Boolean logic as the theoretical study by Genot and co-workers shows.237 This work essentially presents network motifs that act like a nonlinear and nonadditive switch of the starting substrates. This type of research showcases the intrinsic computational power of ERNs, circumventing the requirements necessary for developing explicit digital circuitry.
Recently, this approach has been expanded to enzymatic systems capable of more generalized types of computation. Inspired by concepts from machine learning and neuromorphic computation, Pandi et al. created in vitro metabolic perceptrons.238 They employed computational retrosynthesis tools to design enzymatic models that can act as transducers (converting different incoming metabolite signals into one substrate) and nonlinear actuators (to achieve sigmoidal responses) and when combined together can act as analogue adders (to perform a weighted sum of incoming signals, e.g., a perceptron). A schematic overview of this is shown in Figure b. These perceptrons are essentially based on ultrasensitivity network motifs to create nonlinear weighted adders (sigmoidal sums) of multiple metabolites but do so in a generalizable and extensible manner. Modifying the transducer reactions changes how the incoming signals are weighted, allowing the perceptron to be controlled and trained similar to an artificial neural network.
A different approach is investigated in the work of Okumura et al., where neural networks capable of nonlinear classification have been created by assembling various components of the polymerase–exonuclease–nickase (PEN) toolbox, as shown in Figure c.239 These enzymes, respectively, polymerize, degrade, and cut DNA strands. Neurons are constructed by supplying specific DNA template strands that convert DNA (or RNA) inputs either into a “reporter” DNA strand or into an “antireporter” DNA strand. The reporter strand can be transformed into a fluorescent signal by another DNA template strand. The antireporter strand polymerizes with the reporter strand, inhibiting its output. Thus, modifying the concentration of reporter and antireporter templates impacts fluorescence, respectively, in a positive or negative way, establishing an effective weight on input conversion. These effective weights can be further modified by incorporating so-called “fake” weight templates, which competitively inhibit the antireporter–reporter interaction. The reporter signal is further amplified by an amplification template, resulting in an ultrasensitive (sigmoidal) response. Finally, the location of the ultrasensitive regime can be adjusted by inclusion of a “bias” template, which operates similarly to the antireporter strand but is not input dependent. Polymerase and nickase drive these DNA reactions, while exonuclease can be used to degrade the fluorescent output over time. The interaction of all enzymes and DNA template strands together establishes a perceptron where all weights and biases can be fine tuned by controlling the concentration of the reporter, antireporter, fake, and bias template strands. By exchanging the reporter templates for strands that can serve as input to another perceptron (Figure d), a multilayer neural network was created that could accurately classify input concentration regimes. Importantly, the computational power in this work originates from the highly heterogeneous substrates used in the three-enzyme ERN, in contrast to the previous example of a metabolic network where it originates from the design of the ERN itself.
The above examples show the significant progress that has been made in enzymatic information processing by directly exploiting similarities between enzymatic interactions and analogous principles instead of recreating small Boolean operations and assembling those in a circuit. By either rationally fine tuning the parameter regime in small existing ERNs, by computationally designing larger ERNs to convert many different substrates into the same products, or by intelligently choosing which substrates (and specifically which templates) to exploit using just a small ERN, impressive feats of computation can be achieved. However, further developments have been mostly theoretical up until now, including work showcasing different enzymatic network motifs capable of noise filtering,240 pattern recognition and pulse counting,241 and systems that can convert information into work and vice versa.242 It remains to be seen if these proposals can be realized experimentally, but they highlight the potential for ERNs in future information processing tasks.
Conclusion
We have presented this comprehensive review of the field of in vitro enzymatic reaction networks in the hope to inspire a broad group of researchers to participate in expanding the field. Key design principles, basic emergent properties, and the need for an integrated approach comprising experiments, computational approaches, and analytical methods have been highlighted. Several key challenges for future research were identified: there are currently no “blueprints” for the design of ERNs with functions that go beyond “simple” temporal behavior. For example: we know how to create oscillatory networks, but we cannot design networks that design with a specific frequency and amplitude. Most ERNs reported to date have relatively simple topologies. To significantly expand current designs, we need to establish new strategies for the reverse design of functional ERNs and establish high-throughput methods for the rapid evaluation of networks consisting of multiple feedback loops. Instead of current “trial-and-error” experiments to optimize network properties, iterative “design–build–test–learn” cycles need to be established, and we are hopeful that advances in machine learning and artificial intelligence could be leveraged to identify novel network topologies with hitherto “undesignable” properties. We note that many experimental designs (especially if they are aimed at constructing materials with life-like properties) can only be realized within a relatively narrow scope of kinetic values. Simulations can significantly accelerate discoveries of the most relevant experimental regime, but this will need to be coupled to methods to control or expand the kinetic parameters of enzymes. Although current networks are incomparable in complexity to ERNs found in extant life, our overview on possibilities to control enzymatic activity together with the many examples of feedback loops gives us confidence significantly more elaborate reaction networks will be within reach in the next decade. Indeed, the successful attempts to construct complex reaction cascades and cycles to synthesize complex organic molecules are testament to the potential of more complex ERNs.
We foresee many new developments in the construction of life-like materials, where properties such as homeostasis, sensing the environment, and motility are all carried out by coupled enzymatic reaction networks. The design of intelligent materials by incorporating ERNs, which are able to learn from their environment to adopt desired features, could open up innovative avenues in material science. In section 4, we have explored the influence of external stimuli to regulate ERN activity, serving as physical learning rules for enhanced task performance. Interestingly, these materials potentially can exhibit task adaptiveness through training with different stimuli.243 Another important direction for designing life-like materials would be to incorporate some type of fuel regeneration and methods to maintain such materials out of equilibrium as many of the properties associated with living systems (motility, homeostasis, regeneration) require dissipative systems and continuous input of energy. In section 5, we have introduced different strategies like energy production, compartmentalization, and communication using ERNs, which facilitate crosstalk between multiscale processes to design truly autonomous systems.244 Such materials could then find applications as delivery vehicles in medicine, in soft robotics, or as interfaces between living systems and electronic systems. In section 6, we demonstrated the progress in cell-free biosynthesis, showcasing the in vitro design of metabolic modules for unraveling complex metabolic machinery and producing value-added chemicals. Looking ahead, we anticipate the development of ever more complex ERNs for designing novel catalytic networks with promising industrial applications. Significantly, this can augment the design of novel synthetic cells as chemical factories. Finally, enzymes are key in processing information in biology, but the potential of synthetic ERNs to achieve similar capabilities remains largely unrealized.245 Building on the multifarious uses of enzymes in complex (and not so complex) reaction networks, there is much potential for designing novel computing paradigms in which enzymes are used to process multimodal and multiplexed information at the molecular level.
References
- R. Colin, V. Sourjik. Emergent properties of bacterial chemotaxis pathway.. Curr. Opin. Microbiol., 2017. [DOI | PubMed]
- A. Typas, V. Sourjik. Bacterial protein networks: properties and functions.. Nat. Rev. Microbiol., 2015. [DOI | PubMed]
- T. Hunter. Protein Kinases and Phosphatases: The yin and yang of protein phosphorylation andsignaling.. Cell, 1995. [DOI | PubMed]
- M. R. Junttila, S. P. Li, J. Westermarck. Phosphatase-mediated crosstalk between MAPK signaling pathways in the regulation of cell survival.. FASEB J., 2008. [DOI | PubMed]
- J. Saravia, J. L. Raynor, N. M. Chapman, S. A. Lim, H. Chi. Signaling networks in immunometabolism.. Cell Res., 2020. [DOI | PubMed]
- T. Fink, J. Lonzaric, A. Praznik, T. Plaper, E. Merljak, K. Leben, N. Jerala, T. Lebar, Z. Strmsek, F. Lapenta. Design of fast proteolysis-based signaling and logic circuits in mammalian cells.. Nat. Chem. Biol., 2019. [DOI | PubMed]
- Z. Feng, H. Wang, F. Wang, Y. Oh, C. Berciu, Q. Cui, E. H. Egelman, B. Xu. Artificial intracellular filaments.. Cell Rep. Phys. Sci., 2020. [DOI | PubMed]
- S. Kohyama, A. Merino-Salomon, P. Schwille. In vitro assembly, positioning and contraction of a division ring in minimal cells.. Nat. Commun., 2022. [DOI | PubMed]
- A. K. Fisher, B. G. Freedman, D. R. Bevan, R. S. Senger. A review of metabolic and enzymatic engineering strategies for designing and optimizing performance of microbial cell factories.. Comput. Struct. Biotechnol. J., 2014. [DOI | PubMed]
- S. Hirschi, T. R. Ward, W. P. Meier, D. J. Muller, D. Fotiadis. Synthetic biology: bottom-up assembly of molecular systems.. Chem. Rev., 2022. [DOI | PubMed]
- M. Boguñá, I. Bonamassa, M. De Domenico, S. Havlin, D. Krioukov, M. Á. Serrano. Network geometry.. Nat. Rev. Phys., 2021. [DOI]
- E. Ravasz, A. L. Somera, D. A. Mongru, Z. N. Oltvai, A.-L. Barabási. Hierarchical organization of modularity in metabolic networks.. Science, 2002. [DOI | PubMed]
- A.-L. Barabási, Z. N. Oltvai. Network biology: understanding the cell’s functional organization.. Nat. Rev. Genet., 2004. [DOI | PubMed]
- M. A. Serrano, M. Boguñá, F. Sagués. Uncovering the hidden geometry behind metabolic networks.. Mol. Biosyst., 2012. [DOI | PubMed]
- H. Kim, H. B. Smith, C. Mathis, J. Raymond, S. Walker. Universal scaling across biochemical networks on Earth.. Sci. Adv., 2019. [DOI | PubMed]
- D. C. Gagler, B. Karas, C. P. Kempes, J. Malloy, V. Mierzejewski, A. D. Goldman, H. Kim, S. I. Walker. Scaling laws in enzyme function reveal a new kind of biochemical universality.. Proc. Natl. Acad. Sci. U. S. A., 2022. [DOI | PubMed]
- O. Guell, F. Sagues, M. A. Serrano. Essential plasticity and redundancy of metabolism unveiled by synthetic lethality analysis.. PLoS Comput. Biol., 2014. [DOI | PubMed]
- G. Sambamoorthy, K. Raman. Understanding the evolution of functional redundancy in metabolic networks.. Bioinformatics, 2018. [DOI | PubMed]
- U. Alon. Network motifs: theory and experimental approaches.. Nat. Rev. Genet., 2007. [DOI | PubMed]
- A. Goldbeter, D. E. Koshland. An amplified sensitivity arising from covalent modification in biological systems.. Proc. Natl. Acad. Sci. U. S. A., 1981. [DOI | PubMed]
- D. E. Koshland, A. Goldbeter, J. B. Stock. Amplification and adaptation in regulatory and sensory systems.. Science, 1982. [DOI | PubMed]
- N. Barkai, S. Leibler. Robustness in simple biochemical networks.. Nature, 1997. [DOI | PubMed]
- R. Milo, S. Shen-Orr, S. Itzkovitz, N. Kashtan, D. Chklovskii, U. Alon. Network motifs: simple building blocks of complex networks.. Science., 2002. [DOI | PubMed]
- J. J. Tyson, B. Novak. Functional motifs in biochemical reaction networks.. Annu. Rev. Phys. Chem., 2010. [DOI | PubMed]
- B. Novak, J. J. Tyson. Design principles of biochemical oscillators.. Nat. Rev. Mol. Cell Biol., 2008. [DOI | PubMed]
- R. P. Araujo, L. A. Liotta. Universal structures for adaptation in biochemical reaction networks.. Nat. Commun., 2023. [DOI | PubMed]
- E. R. Shellman, C. F. Burant, S. Schnell. Network motifs provide signatures that characterize metabolism.. Mol. Biosyst., 2013. [DOI | PubMed]
- M. E. Beber, C. Fretter, S. Jain, N. Sonnenschein, M. Muller-Hannemann, M. T. Hutt. Artefacts in statistical analyses of network motifs: general framework and application to metabolic networks.. J. R. Soc. Interface, 2012. [DOI | PubMed]
- D. E. Piephoff, J. Wu, J. Cao. Conformational nonequilibrium enzyme kinetics: generalized Michaelis-Menten equation.. J. Phys. Chem. Lett., 2017. [DOI | PubMed]
- J. M. Rohwer, A. J. Hanekom, C. Crous, J. L. Snoep, J. H. Hofmeyr. Evaluation of a simplified generic bi-substrate rate equation for computational systems biology.. Syst. Biol. (Stevenage), 2006. [DOI | PubMed]
- M. K. Transtrum, B. B. Machta, K. S. Brown, B. C. Daniels, C. R. Myers, J. P. Sethna. Perspective: sloppiness and emergent theories in physics, biology, and beyond.. J. Chem. Phys., 2015. [DOI | PubMed]
- R. N. Gutenkunst, J. J. Waterfall, F. P. Casey, K. S. Brown, C. R. Myers, J. P. Sethna. Universally sloppy parameter sensitivities in systems biology models.. PLoS Comput. Biol., 2007. [DOI | PubMed]
- M. K. Transtrum, P. Qiu. Model reduction by manifold boundaries.. Phys. Rev. Lett., 2014. [DOI | PubMed]
- R. van de Schoot, S. Depaoli, R. King, B. Kramer, K. Märtens, M. G. Tadesse, M. Vannucci, A. Gelman, D. Veen, J. Willemsen, C. Yau. Bayesian statistics and modelling.. Nat. Rev. Methods Primers, 2021. [DOI]
- B. Choi, G. A. Rempala, J. K. Kim. Beyond the Michaelis-Menten equation: accurate and efficient estimation of enzyme kinetic parameters.. Sci. Rep., 2017. [DOI | PubMed]
- N. J. Linden, B. Kramer, P. Rangamani. Bayesian parameter estimation for dynamical models in systems biology.. PLoS Comput. Biol., 2022. [DOI | PubMed]
- M. G. Baltussen, J. van de Wiel, C. L. Fernandez Regueiro, M. Jakstaite, W. T. S. Huck. A Bayesian Approach to Extracting kinetic information from artificial enzymatic networks.. Anal. Chem., 2022. [DOI | PubMed]
- A. Gábor, A. F. Villaverde, J. R. Banga. Parameter identifiability analysis and visualization in large-scale kinetic models of biosystems.. BMC Syst. Biol., 2017. [DOI | PubMed]
- A. F. Villaverde, N. D. Evans, M. J. Chappell, J. R. Banga. Input-dependent structural identifiability of nonlinear systems.. IEEE Control Systems Letters, 2019. [DOI]
- B. van Sluijs, R. J. M. Maas, A. J. van der Linden, T. F. A. de Greef, W. T. S. Huck. A microfluidic optimal experimental design platform for forward design of cell-free genetic networks.. Nat. Commun., 2022. [DOI | PubMed]
- C. Hold, S. Billerbeck, S. Panke. Forward design of a complex enzyme cascade reaction.. Nat. Commun., 2016. [DOI | PubMed]
- A. S. Y. Wong, W. T. S. Huck. Grip on complexity in chemical reaction networks.. Beilstein J. Org. Chem., 2017. [DOI | PubMed]
- A. I. Hanopolskyi, V. A. Smaliak, A. I. Novichkov, S. N. Semenov. Autocatalysis: Kinetics, mechanisms and design.. ChemSystemsChem., 2021. [DOI]
- T. Bánsági, A. F. Taylor. Exploitation of feedback in enzyme-catalysed reactions.. Isr. J. Chem., 2018. [DOI]
- R. Plasson, A. Brandenburg, L. Jullien, H. Bersini. Autocatalyses.. J. Phys. Chem. A, 2011. [DOI | PubMed]
- M. A. Budroni, F. Rossi, L. Rongy. From transport phenomena to systems chemistry: chemohydrodynamic oscillations in A+B⃗C systems.. ChemSystemsChem., 2022. [DOI]
- G. Menon, J. Krishnan. Spatial localisation meets biomolecular networks.. Nat. Commun., 2021. [DOI | PubMed]
- C. Sharma, I. Maity, A. Walther. pH-feedback systems to program autonomous self-assembly and material lifecycles.. Chem. Commun., 2023. [DOI]
- B. Dúzs, I. Molnár, I. Lagzi, I. Szalai. Reaction–diffusion dynamics of pH oscillators in oscillatory forced open spatial reactors.. ACS Omega, 2021. [DOI | PubMed]
- B. Duzs, I. Lagzi, I. Szalai. Functional rhythmic chemical systems governed by pH-driven kinetic feedback.. ChemSystemsChem, 2023. [DOI]
- I. N. Bubanja, T. Bánsági, A. F. Taylor. Kinetics of the urea–urease clock reaction with urease immobilized in hydrogel beads.. Reac. Kinet. Mech. Catal., 2018. [DOI]
- Y. Miele, S. J. Jones, F. Rossi, P. A. Beales, A. F. Taylor. Collective behavior of urease pH clocks in nano- and microvesicles controlled by fast ammonia transport.. J. Phys. Chem. Lett., 2022. [DOI | PubMed]
- V. M. Markovic, T. Bánsági, D. McKenzie, A. Mai, J. A. Pojman, A. F. Taylor. Influence of reaction-induced convection on quorum sensing in enzyme-loaded agarose beads.. Chaos, 2019. [DOI | PubMed]
- F. Muzika, M. RuZicka, L. Schreiberova, I. Schreiber. Oscillations of pH in the urea-urease system in a membrane reactor.. Phys. Chem. Chem. Phys., 2019. [DOI | PubMed]
- F. Muzika, T. Bánsági, I. Schreiber, L. Schreiberova, A. F. Taylor. A bistable switch in pH in urease-loaded alginate beads.. Chem. Commun., 2014. [DOI]
- G. Hu, J. A. Pojman, S. K. Scott, M. M. Wrobel, A. F. Taylor. Base-catalyzed feedback in the urea-urease reaction.. J. Phys. Chem. B, 2010. [DOI | PubMed]
- A. V. Straube, S. Winkelmann, C. Schütte, F. Höfling. Stochastic pH oscillations in a model of the urea-urease reaction confined to lipid vesicles.. J. Phys. Chem. Lett., 2021. [DOI | PubMed]
- T. Bánsági, A. F. Taylor. Switches induced by quorum sensing in a model of enzyme-loaded microparticles.. J. R. Soc. Interface., 2018. [DOI]
- L. Heinen, T. Heuser, A. Steinschulte, A. Walther. Antagonistic enzymes in a biocatalytic pH feedback system program autonomous DNA hydrogel life cycles.. Nano Lett., 2017. [DOI | PubMed]
- X. Fan, A. Walther. Autonomous transient pH flips shaped by layered compartmentalization of antagonistic enzymatic reactions.. Angew. Chem., Int. Ed., 2021. [DOI]
- X. Wang, S. Moreno, S. Boye, P. Wen, K. Zhang, P. Formanek, A. Lederer, B. Voit, D. Appelhans. Feedback-induced and oscillating pH regulation of a binary enzyme–polymersomes.. System. Chem. Mater., 2021. [DOI]
- B. Dúzs, I. Lagzi, I. Szalai. Functional rhythmic chemical systems governed by pH-Driven kinetic feedback.. ChemSystemsChem, 2023. [DOI]
- E. Jee, T. Bánsági, A. F. Taylor, J. A. Pojman. Temporal control of gelation and polymerization fronts driven by an autocatalytic enzyme reaction.. Angew. Chem., Int. Ed., 2016. [DOI]
- A. Q. Mai, T. Bánsági, A. F. Taylor, J. A. Pojman. Reaction-diffusion hydrogels from urease enzyme particles for patterned coatings.. Commun. Chem., 2021. [DOI | PubMed]
- I. Maity, C. Sharma, F. Lossada, A. Walther. Feedback and communication in active hydrogel spheres with pH fronts: facile approaches to grow soft hydrogel structures.. Angew. Chem., Int. Ed., 2021. [DOI]
- N. Rauner, L. Buenger, S. Schuller, J. C. Tiller. Post-polymerization of urease-induced calcified, polymer hydrogels.. Macromol. Rapid Commun., 2015. [DOI | PubMed]
- S. N. Semenov, A. S. Wong, R. M. van der Made, S. G. Postma, J. Groen, H. W. van Roekel, T. F. de Greef, W. T. Huck. Rational design of functional and tunable oscillating enzymatic networks.. Nat. Chem., 2015. [DOI | PubMed]
- O. R. Maguire, A. S. Y. Wong, J. H. Westerdiep, W. T. S. Huck. Early warning signals in chemical reaction networks.. Chem. Commun., 2020. [DOI]
- O. R. Maguire, A. S. Y. Wong, M. G. Baltussen, P. van Duppen, A. A. Pogodaev, W. T. S. Huck. Dynamic environments as a tool to preserve desired output in a chemical reaction network.. Chem. Eur. J., 2020. [DOI | PubMed]
- B. Helwig, B. van Sluijs, A. A. Pogodaev, S. G. J. Postma, W. T. S. Huck. Bottom-up construction of an adaptive enzymatic reaction network.. Angew. Chem., Int. Ed., 2018. [DOI]
- S. N. Semenov, A. J. Markvoort, T. F. de Greef, W. T. Huck. Threshold sensing through a synthetic enzymatic reaction-diffusion network.. Angew. Chem., Int. Ed., 2014. [DOI]
- D. V. Kriukov, A. H. Koyuncu, A. S. Y. Wong. History dependence in a chemical reaction network enables dynamic switching.. Small, 2022. [DOI]
- A. A. Pogodaev, C. L. Fernández Regueiro, M. Jakštaitė, M. J. Hollander, W. T. S. Huck. Modular design of small enzymatic reaction networks based on reversible and cleavable inhibitors.. Angew. Chem., Int. Ed., 2019. [DOI]
- I. Yamazaki, K. Yokota, R. Nakajima. Oscillatory oxidations of reduced pyridine nucleotide by peroxidase.. Biochem. Biophys. Res. Commun., 1965. [DOI | PubMed]
- L. Folke Olsen. Complex dynamics in an unexplored simple model of the peroxidase-oxidase reaction.. Chaos, 2023. [DOI | PubMed]
- L. F. Olsen, A. Lunding. Chaos in the peroxidase-oxidase oscillator.. Chaos, 2021. [DOI | PubMed]
- A. Scheeline, D. L. Olson, E. P. Williksen, G. A. Horras, M. L. Klein, R. Larter. The peroxidase–oxidase oscillator and its constituent chemistries.. Chem. Rev., 1997. [DOI | PubMed]
- Y. Zhang, S. Tsitkov, H. Hess. Complex dynamics in a two-enzyme reaction network with substrate competition.. Nat. Catal., 2018. [DOI]
- L. Gyevi-Nagy, E. Lantos, T. Gehér-Herczegh, A. Tóth, C. Bagyinka, D. Horváth. Reaction fronts of the autocatalytic hydrogenase reaction.. J. Chem. Phys., 2018. [DOI | PubMed]
- C. Bagyinka, G. Pankotai-Bodó, R. M. M. Branca, M. Debreczeny. Oscillating hydrogenase reaction.. Int. J. Hydrogen Energy, 2014. [DOI]
- C. Claaßen, T. Gerlach, D. Rother. Stimulus-responsive regulation of enzyme activity for one-step and multi-step syntheses.. Adv. Synth. Catal., 2019. [DOI | PubMed]
- Z. Chen, Y. Zhao, Y. Liu. Advanced strategies of enzyme activity regulation for biomedical applications.. ChemBioChem, 2022. [DOI | PubMed]
- M. W. H. Hoorens, W. Szymanski. Reversible, spatial and temporal control over protein activity using light.. Trends Biochem. Sci., 2018. [DOI | PubMed]
- A. C. Kneuttinger. A guide to designing photocontrol in proteins: methods, strategies and applications.. Biol. Chem., 2022. [DOI | PubMed]
- J. Volarić, W. Szymanski, N. A. Simeth, B. L. Feringa. Molecular photoswitches in aqueous environments.. Chem. Soc. Rev., 2021. [DOI | PubMed]
- A. A. Pogodaev, T. T. Lap, W. T. S. Huck. The dynamics of an oscillating enzymatic reaction network is crucially determined by side reactions.. ChemSystemsChem, 2021. [DOI]
- S. Crespi, N. A. Simeth, B. König. Heteroaryl azo dyes as molecular photoswitches.. Nat. Rev. Chem., 2019. [DOI]
- M. Teders, A. A. Pogodaev, G. Bojanov, W. T. S. Huck. Reversible photoswitchable inhibitors generate ultrasensitivity in out-of-equilibrium enzymatic reactions.. J. Am. Chem. Soc., 2021. [DOI | PubMed]
- M. Teders, N. R. Murray, W. T. S. Huck. Reversible photoswitchable inhibitors enable wavelength-selective regulation of out-of-equilibrium bi-enzymatic systems.. ChemSystemsChem, 2021. [DOI]
- K. Li, M. D. Liu, Q. X. Huang, C. J. Liu, X. Z. Zhang. Nanoplatforms with donor-acceptor Stenhouse adduct molecular switch for enzymatic reactions remotely controlled with near-infrared light.. Sci. China Mater., 2023. [DOI]
- J. W. Hindley, Y. Elani, C. M. McGilvery, S. Ali, C. L. Bevan, R. V. Law, O. Ces. Light-triggered enzymatic reactions in nested vesicle reactors.. Nat. Commun., 2018. [DOI | PubMed]
- O. Babii, S. Afonin, C. Diel, M. Huhn, J. Dommermuth, T. Schober, S. Koniev, A. Hrebonkin, A. Nesterov-Mueller, I. V. Komarov. Diarylethene-based photoswitchable inhibitors of serine proteases.. Angew. Chem., Int. Ed., 2021. [DOI]
- O. Rifaie-Graham, J. Yeow, A. Najer, R. Wang, R. Sun, K. Zhou, T. N. Dell, C. Adrianus, C. Thanapongpibul, M. Chami. Photoswitchable gating of non-equilibrium enzymatic feedback in chemically communicating polymersome nanoreactors.. Nat. Chem., 2023. [DOI | PubMed]
- H. Bisswanger. Enzyme assays.. Perspectives in Science, 2014. [DOI]
- Y. Zhang, Q. Wang, H. Hess. Increasing enzyme cascade throughput by pH-engineering the microenvironment of individual enzymes.. ACS Catal., 2017. [DOI]
- X. Fan, A. Walther. pH feedback lifecycles programmed by enzymatic logic gates using common foods as fuels.. Angew. Chem., Int. Ed., 2021. [DOI]
- H. Che, S. Cao, J. C. M. van Hest. Feedback-induced temporal control of ″breathing″ polymersomes to create self-adaptive nanoreactors.. J. Am. Chem. Soc., 2018. [DOI | PubMed]
- P. K. Wells, O. Smutok, A. Melman, E. Katz. Switchable biocatalytic reactions controlled by interfacial pH changes produced by orthogonal biocatalytic processes.. ACS Appl. Mater. Interfaces, 2021. [DOI | PubMed]
- C. Wang, A. Fischer, A. Ehrlich, Y. Nahmias, I. Willner. Biocatalytic reversible control of the stiffness of DNA-modified responsive hydrogels: applications in shape-memory, self-healing and autonomous controlled release of insulin.. Chem. Sci., 2020. [DOI | PubMed]
- Y. Zhang, N. Nie, H. Wang, Z. Tong, H. Xing, Y. Zhang. Smart enzyme catalysts capable of self-separation by sensing the reaction extent.. Biosens. Bioelectron., 2023. [DOI | PubMed]
- M. Jain, B. J. Ravoo. Fuel-driven and enzyme-regulated redox-responsive supramolecular hydrogels.. Angew. Chem., Int. Ed., 2021. [DOI]
- T. Eixelsberger, B. Nidetzky. Enzymatic redox cascade for one-pot synthesis of uridine 5′-diphosphate xylose from uridine 5′-diphosphate glucose.. Adv. Synth. Catal., 2014. [DOI | PubMed]
- S. K. Kuk, R. K. Singh, D. H. Nam, R. Singh, J. K. Lee, C. B. Park. Photoelectrochemical reduction of carbon dioxide to methanol through a highly efficient enzyme cascade.. Angew. Chem., Int. Ed., 2017. [DOI]
- S. Mallawarachchi, V. Gejji, L. S. Sierra, H. Wang, S. Fernando. Electrical field reversibly modulates enzyme kinetics of hexokinase entrapped in an electro-responsive hydrogel.. ACS Appl. Bio Mater., 2019. [DOI]
- G. Morello, C. F. Megarity, F. A. Armstrong. The power of electrified nanoconfinement for energising, controlling and observing long enzyme cascades.. Nat. Commun., 2021. [DOI | PubMed]
- R. D. Milton, R. Cai, S. Abdellaoui, D. Leech, A. L. De Lacey, M. Pita, S. D. Minteer. Bioelectrochemical Haber-Bosch process: an ammonia-producing H2/N2 fuel cell.. Angew. Chem., Int. Ed., 2017. [DOI]
- Y. Cao, Y. Wang. Temperature-mediated regulation of enzymatic activity.. ChemCatChem., 2016. [DOI]
- S. Zhang, C. Wang, H. Chang, Q. Zhang, Y. Cheng. Off-on switching of enzyme activity by near-infrared light-induced photothermal phase transition of nanohybrids.. Sci. Adv., 2019. [DOI | PubMed]
- P. Gobbo, A. J. Patil, M. Li, R. Harniman, W. H. Briscoe, S. Mann. Programmed assembly of synthetic protocells into thermoresponsive prototissues.. Nat. Mater., 2018. [DOI | PubMed]
- K. Szekeres, P. Bollella, Y. Kim, S. Minko, A. Melman, E. Katz. Magneto-controlled enzyme activity with locally produced pH changes.. J. Phys. Chem. Lett., 2021. [DOI | PubMed]
- P. Dhasaiyan, T. Ghosh, H. G. Lee, Y. Lee, I. Hwang, R. D. Mukhopadhyay, K. M. Park, S. Shin, I. S. Kang, K. Kim. Cascade reaction networks within audible sound induced transient domains in a solution.. Nat. Commun., 2022. [DOI | PubMed]
- A. Küchler, M. Yoshimoto, S. Luginbühl, F. Mavelli, P. Walde. Enzymatic reactions in confined environments.. Nat. Nanotechnol., 2016. [DOI | PubMed]
- E. T. Hwang, S. Lee. Multienzymatic Cascade Reactions Enzyme complex by immobilization.. ACS Catal., 2019. [DOI]
- Q. Ji, B. Wang, J. Tan, L. Zhu, L. Li. Immobilized multienzymatic systems for catalysis of cascade reactions.. Process Biochemistry, 2016. [DOI]
- F. Kazenwadel, M. Franzreb, B. E. Rapp. Synthetic enzyme supercomplexes: co-immobilization of enzyme cascades.. Anal. Methods, 2015. [DOI]
- K. Xu, X. Chen, R. Zheng, Y. Zheng. Immobilization of multi-enzymes on support materials for efficient biocatalysis.. Front. Bioeng. Biotechnol., 2020. [DOI | PubMed]
- J. Bié, B. Sepodes, P. C. B. Fernandes, M. H. L. Ribeiro. Enzyme immobilization and co-immobilization: main framework, advances and some applications.. Processes, 2022. [DOI]
- A. I. Benitez-Mateos, D. Roura Padrosa, F. Paradisi. Multistep enzyme cascades as a route towards green and sustainable pharmaceutical syntheses.. Nat. Chem., 2022. [DOI | PubMed]
- J. Liang, K. Liang. Multi-enzyme cascade reactions in metal-organic frameworks.. Chem. Rec., 2020. [DOI | PubMed]
- S. Tsitkov, H. Hess. Design principles for a compartmentalized enzyme cascade reaction.. ACS Catal., 2019. [DOI]
- W. Oh, D. Jeong, J.-W. Park. An Artificial compartmentalized biocatalytic cascade system constructed with enzyme-caged reticulate nanoporous membranes.. Adv. Mater. Interfaces, 2023. [DOI]
- E. Diamanti, D. Andrés-Sanz, A. H. Orrego, S. Carregal-Romero, F. López-Gallego. Surpassing substrate–enzyme competition by compartmentalization.. ACS Catal., 2023. [DOI]
- W. M. Aumiller, M. Uchida, T. Douglas. Protein cage assembly across multiple length scales.. Chem. Soc. Rev., 2018. [DOI | PubMed]
- A. Jaekel, P. Stegemann, B. Saccà. Manipulating enzymes properties with DNA nanostructures.. Molecules, 2019. [DOI | PubMed]
- Y. Jiao, Y. Shang, N. Li, B. Ding. DNA-based enzymatic systems and their applications.. iScience, 2022. [DOI | PubMed]
- W. Engelen, B. M. G. Janssen, M. Merkx. DNA-based control of protein activity.. Chem. Commun., 2016. [DOI]
- J. Fu, Z. Wang, X. H. Liang, S. W. Oh, E. St. Iago-McRae, T. Zhang. DNA-scaffolded proximity assembly and confinement of multienzyme reactions.. Top. Curr. Chem., 2020. [DOI]
- S. Kröll, C. M. Niemeyer. Nucleic acid-based enzyme cascades—current trends and future perspectives.. Angew. Chem., Int. Ed., 2024. [DOI]
- J. Liang, K. Liang. Multi-enzyme cascade reactions in metal-organic frameworks.. Chem. Rec., 2020. [DOI | PubMed]
- A. Dhakshinamoorthy, A. M. Asiri, H. Garcia. Integration of metal organic frameworks with enzymes as multifunctional solids for cascade catalysis.. Dalton Trans., 2020. [DOI | PubMed]
- X. Wang, P. C. Lan, S. Ma. Metal–organic frameworks for enzyme immobilization: beyond host matrix materials.. ACS Cent. Sci., 2020. [DOI | PubMed]
- X. Lian, Y. Fang, E. Joseph, Q. Wang, J. Li, S. Banerjee, C. Lollar, X. Wang, H.-C. Zhou. Enzyme–MOF (metal–organic framework) composites.. Chem. Soc. Rev., 2017. [DOI | PubMed]
- W. Liang, P. Wied, F. Carraro, C. J. Sumby, B. Nidetzky, C.-K. Tsung, P. Falcaro, C. J. Doonan. Metal–organic framework-based enzyme biocomposites.. Chem. Rev., 2021. [DOI | PubMed]
- J. Hu, G. Zhang, S. Liu. Enzyme-responsive polymeric assemblies, nanoparticles and hydrogels.. Chem. Soc. Rev., 2012. [DOI | PubMed]
- P. Li, Y. Zhong, X. Wang, J. Hao. Enzyme-regulated healable polymeric hydrogels.. ACS Cent. Sci., 2020. [DOI | PubMed]
- R. J. Amir, S. Zhong, D. J. Pochan, C. J. Hawker. Enzymatically triggered self-assembly of block copolymers.. J. Am. Chem. Soc., 2009. [DOI | PubMed]
- S. Kobayashi, H. Uyama, S. Kimura. Enzymatic polymerization.. Chem. Rev., 2001. [DOI | PubMed]
- R. G. Klemperer, M. R. Shannon, J. L. Ross Anderson, A. W. Perriman. Bienzymatic generation of interpenetrating polymer networked engineered living materials with shape changing Pproperties.. Adv. Mater. Technol., 2023. [DOI]
- Y. Mao, T. Su, Q. Wu, C. Liao, Q. Wang. Dual enzymatic formation of hybrid hydrogels with supramolecular-polymeric networks.. Chem. Commun., 2014. [DOI]
- Q. Wei, M. Xu, C. Liao, Q. Wu, M. Liu, Y. Zhang, C. Wu, L. Cheng, Q. Wang. Printable hybrid hydrogel by dual enzymatic polymerization with superactivity.. Chem. Sci., 2016. [DOI | PubMed]
- Z. Yang, G. Liang, L. Wang, B. Xu. Using a kinase/phosphatase switch to regulate a supramolecular hydrogel and forming the supramolecular hydrogel in vivo.. J. Am. Chem. Soc., 2006. [DOI | PubMed]
- J. M. Knipe, F. Chen, N. A. Peppas. Enzymatic biodegradation of hydrogels for protein delivery targeted to the small intestine.. Biomacromolecules, 2015. [DOI | PubMed]
- C. G. Pappas, I. R. Sasselli, R. V. Ulijn. Biocatalytic pathway selection in transient tripeptide nanostructures.. Angew. Chem., Int. Ed., 2015. [DOI]
- A. B. Cook, P. Decuzzi. Harnessing endogenous stimuli for responsive materials in theranostics.. ACS Nano, 2021. [DOI | PubMed]
- T. H. Ku, M. P. Chien, M. P. Thompson, R. S. Sinkovits, N. H. Olson, T. S. Baker, N. C. Gianneschi. Controlling and switching the morphology of micellar nanoparticles with enzymes.. J. Am. Chem. Soc., 2011. [DOI | PubMed]
- S. G. Postma, I. N. Vialshin, C. Y. Gerritsen, M. Bao, W. T. Huck. Preprogramming complex hydrogel responses using enzymatic reaction networks.. Angew. Chem., Int. Ed., 2017. [DOI]
- M. Ikeda, T. Tanida, T. Yoshii, K. Kurotani, S. Onogi, K. Urayama, I. Hamachi. Installing logic-gate responses to a variety of biological substances in supramolecular hydrogel-enzyme hybrids.. Nat. Chem., 2014. [DOI | PubMed]
- T. Heuser, R. Merindol, S. Loescher, A. Klaus, A. Walther. Photonic devices out of equilibrium: transient memory, signal propagation, and sensing.. Adv. Mater., 2017. [DOI]
- Y. Hong, D. Velegol, N. Chaturvedi, A. Sen. Biomimetic behavior of synthetic particles: from microscopic randomness to macroscopic control.. Phys. Chem. Chem. Phys., 2010. [DOI | PubMed]
- S. Sengupta, K. K. Dey, H. S. Muddana, T. Tabouillot, M. E. Ibele, P. J. Butler, A. Sen. Enzyme molecules as nanomotors.. J. Am. Chem. Soc., 2013. [DOI | PubMed]
- X. Zhao, K. Gentile, F. Mohajerani, A. Sen. Powering motion with enzymes.. Acc. Chem. Res., 2018. [DOI | PubMed]
- Y. Zhang, H. Hess. Enhanced diffusion of catalytically active enzymes.. ACS Cen. Sci., 2019. [DOI]
- Y. Zhang, H. Hess. Chemically-powered swimming and diffusion in the microscopic world.. Nat. Rev. Chem., 2021. [DOI | PubMed]
- T. Patiño, X. Arqué, R. Mestre, L. Palacios, S. Sánchez. Fundamental aspects of enzyme-powered micro- and nanoswimmers.. Acc. Chem. Res., 2018. [DOI | PubMed]
- L. Wang, S. Song, J. van Hest, L. Abdelmohsen, X. Huang, S. Sánchez. Biomimicry of cellular motility and communication based on synthetic soft-architectures.. Small, 2020. [DOI]
- S. Hermanová, M. Pumera. Biocatalytic micro- and nanomotors.. Chem. Eur. J., 2020. [DOI]
- H. S. Muddana, S. Sengupta, T. E. Mallouk, A. Sen, P. J. Butler. Substrate catalysis enhances single-enzyme diffusion.. J. Am. Chem. Soc., 2010. [DOI | PubMed]
- A. C. Hortelão, S. García-Jimeno, M. Cano-Sarabia, T. Patiño, D. Maspoch, S. Sanchez. LipoBots: Using liposomal vesicles as protective shell of urease-based nanomotors.. Adv. Funct. Mater., 2020. [DOI]
- H. Choi, S. H. Cho, S. K. Hahn. Urease-powered polydopamine nanomotors for intravesical therapy of bladder diseases.. ACS Nano, 2020. [DOI | PubMed]
- Z. Yang, L. Wang, Z. Gao, X. Hao, M. Luo, Z. Yu, J. Guan. Ultrasmall enzyme-powered janus nanomotor working in blood circulation system.. ACS Nano, 2023. [DOI | PubMed]
- B. J. Toebes, F. Cao, D. A. Wilson. Spatial control over catalyst positioning on biodegradable polymeric nanomotors.. Nat. Commun., 2019. [DOI | PubMed]
- B. Qiu, L. Xie, J. Zeng, T. Liu, M. Yan, S. Zhou, Q. Liang, J. Tang, K. Liang, B. Kong. Interfacially super-assembled asymmetric and H2O2 sensitive multilayer-sandwich magnetic mesoporous silica nanomotors for detecting and removing heavy metal ions.. Adv. Funct. Mater., 2021. [DOI]
- L. K. Abdelmohsen, M. Nijemeisland, G. M. Pawar, G. J. Janssen, R. J. Nolte, J. C. van Hest, D. A. Wilson. Dynamic loading and unloading of proteins in polymeric stomatocytes: formation of an enzyme-loaded supramolecular nanomotor.. ACS Nano, 2016. [DOI | PubMed]
- M. Nijemeisland, L. K. Abdelmohsen, W. T. Huck, D. A. Wilson, J. C. van Hest. A compartmentalized out-of-equilibrium enzymatic reaction network for sustained autonomous movement.. ACS Cent. Sci., 2016. [DOI | PubMed]
- A. Chatterjee, S. Ghosh, C. Ghosh, D. Das. Fluorescent microswimmers based on cross-beta amyloid nanotubes and divergent cascade networks.. Angew. Chem., Int. Ed., 2022. [DOI]
- K. K. Dey, X. Zhao, B. M. Tansi, W. J. Mendez-Ortiz, U. M. Cordova-Figueroa, R. Golestanian, A. Sen. Micromotors powered by enzyme catalysis.. Nano Lett., 2015. [DOI | PubMed]
- X. Zhao, H. Palacci, V. Yadav, M. M. Spiering, M. K. Gilson, P. J. Butler, H. Hess, S. J. Benkovic, A. Sen. Substrate-driven chemotactic assembly in an enzyme cascade.. Nat. Chem., 2018. [DOI | PubMed]
- J. Wang, B. J. Toebes, A. S. Plachokova, Q. Liu, D. Deng, J. A. Jansen, F. Yang, D. A. Wilson. Self-propelled PLGA micromotor with chemotactic response to inflammation.. Adv. Healthcare Mater., 2020. [DOI]
- A. Joseph, C. Contini, D. Cecchin, S. Nyberg, L. Ruiz-Perez, J. Gaitzsch, G. Fullstone, X. Tian, J. Azizi, J. Preston, G. Volpe, G. Battaglia. Chemotactic synthetic vesicles: design and applications in blood-brain barrier crossing.. Sci. Adv., 2017. [DOI | PubMed]
- B. Kumar, A. J. Patil, S. Mann. Enzyme-powered motility in buoyant organoclay/DNA protocells.. Nat. Chem., 2018. [DOI | PubMed]
- N. Gao, M. Li, L. Tian, A. J. Patil, B. Pavan Kumar, S. Mann. Chemical-mediated translocation in protocell-based microactuators.. Nat. Chem., 2021. [DOI | PubMed]
- I. Wheeldon, S. D. Minteer, S. Banta, S. C. Barton, P. Atanassov, M. Sigman. Substrate channelling as an approach to cascade reactions.. Nat. Chem., 2016. [DOI | PubMed]
- F. Hinzpeter, U. Gerland, F. Tostevin. Optimal compartmentalization strategies for metabolic microcompartments.. Biophys. J., 2017. [DOI | PubMed]
- Y. Qiao, M. Li, R. Booth, S. Mann. Predatory behaviour in synthetic protocell communities.. Nat. Chem., 2017. [DOI | PubMed]
- Y. Qiao, M. Li, D. Qiu, S. Mann. Response-retaliation behavior in synthetic protocell communities.. Angew. Chem., Int. Ed., 2019. [DOI]
- T. Chakraborty, S. V. Wegner. Cell to cell signaling through light in artificial cell communities: glowing predator lures prey.. ACS Nano, 2021. [DOI | PubMed]
- A. F. Mason, B. C. Buddingh, D. S. Williams, J. C. M. van Hest. Hierarchical self-assembly of a copolymer-stabilized coacervate protocell.. J. Am. Chem. Soc., 2017. [DOI | PubMed]
- X. Wang, L. Tian, H. Du, M. Li, W. Mu, B. W. Drinkwater, X. Han, S. Mann. Chemical communication in spatially organized protocell colonies and protocell/living cell micro-arrays.. Chem. Sci., 2019. [DOI | PubMed]
- B. C. Buddingh’, J. Elzinga, J. C. M. van Hest. Intercellular communication between artificial cells by allosteric amplification of a molecular signal.. Nat. Commun., 2020. [DOI | PubMed]
- Y. Ji, T. Chakraborty, S. V. Wegner. Self-regulated and bidirectional communication in synthetic cell communities.. ACS Nano, 2023. [DOI | PubMed]
- H. Taylor, N. Gao, S. Mann. Chemical communication and protocell-matrix dynamics in segregated colloidosome micro-colonies.. Angew. Chem., Int. Ed., 2023. [DOI]
- G. Fusi, D. Del Giudice, O. Skarsetz, S. Di Stefano, A. Walther. Autonomous soft robots empowered by chemical reaction networks.. Adv. Mater., 2023. [DOI | PubMed]
- Y. Elani, R. V. Law, O. Ces. Vesicle-based artificial cells as chemical microreactors with spatially segregated reaction pathways.. Nat. Commun., 2014. [DOI | PubMed]
- S. Liu, Y. Zhang, X. He, M. Li, J. Huang, X. Yang, K. Wang, S. Mann, J. Liu. Signal processing and generation of bioactive nitric oxide in a model prototissue.. Nat. Commun., 2022. [DOI | PubMed]
- S. Sengupta, D. Patra, I. Ortiz-Rivera, A. Agrawal, S. Shklyaev, K. K. Dey, U. Cordova-Figueroa, T. E. Mallouk, A. Sen. Self-powered enzyme micropumps.. Nat. Chem., 2014. [DOI | PubMed]
- S. Maiti, O. E. Shklyaev, A. C. Balazs, A. Sen. Self-organization of fluids in a multienzymatic pump system.. Langmuir, 2019. [DOI | PubMed]
- I. Ortiz-Rivera, T. M. Courtney, A. Sen. Enzyme micropump-based inhibitor assays.. Adv. Funct. Mater., 2016. [DOI]
- R. K. Manna, O. E. Shklyaev, A. C. Balazs. Chemically driven multimodal locomotion of active, flexible sheets.. Langmuir, 2023. [DOI | PubMed]
- C. Simo, M. Serra-Casablancas, A. C. Hortelao, V. Di Carlo, S. Guallar-Garrido, S. Plaza-Garcia, R. M. Rabanal, P. Ramos-Cabrer, B. Yague, L. Aguado, L. Bardia, S. Tosi, V. Gomez-Vallejo, A. Martin, T. Patino, E. Julian, J. Colombelli, J. Llop, S. Sanchez. Urease-powered nanobots for radionuclide bladder cancer therapy.. Nat. Nanotechnol., 2024. [DOI]
- J. A. Rollin, T. K. Tam, Y. H. P. Zhang. New biotechnology paradigm: cell-free biosystems for biomanufacturing.. Green Chem., 2013. [DOI]
- Y. Zhang, H. Hess. Toward rational design of high-efficiency enzyme cascades.. ACS Catal., 2017. [DOI]
- J. Shi, Y. Wu, S. Zhang, Y. Tian, D. Yang, Z. Jiang. Bioinspired construction of multi-enzyme catalytic systems.. Chem. Soc. Rev., 2018. [DOI | PubMed]
- Z. Wang, B. Sundara Sekar, Z. Li. Recent advances in artificial enzyme cascades for the production of value-added chemicals.. Bioresour. Technol., 2021. [DOI | PubMed]
- S. Mordhorst, J. N. Andexer. Round, round we go – strategies for enzymatic cofactor regeneration.. Nat. Prod. Rep., 2020. [DOI | PubMed]
- K. Bachosz, J. Zdarta, M. Bilal, A. S. Meyer, T. Jesionowski. Enzymatic cofactor regeneration systems: A new perspective on efficiency assessment.. Sci. Total Environ., 2023. [DOI | PubMed]
- C. Monck, Y. Elani, F. Ceroni. Cell-free protein synthesis: biomedical applications and future perspectives.. Chem. Eng. Res. Des., 2022. [DOI]
- J. G. Perez, J. C. Stark, M. C. Jewett. Cell-free synthetic biology: engineering beyond the cell.. Cold Spring Harb. Perspect. Biol., 2016. [DOI | PubMed]
- A. D. Silverman, A. S. Karim, M. C. Jewett. Cell-free gene expression: an expanded repertoire of applications.. Nat. Rev. Genet., 2020. [DOI | PubMed]
- T. P. Korman, B. Sahachartsiri, D. Li, J. M. Vinokur, D. Eisenberg, J. U. Bowie. A synthetic biochemistry system for the in vitro production of isoprene from glycolysis intermediates.. Protein Sci., 2014. [DOI | PubMed]
- T. P. Korman, P. H. Opgenorth, J. U. Bowie. A synthetic biochemistry platform for cell free production of monoterpenes from glucose.. Nat. Commun., 2017. [DOI | PubMed]
- P. H. Opgenorth, T. P. Korman, J. U. Bowie. A synthetic biochemistry molecular purge valve module that maintains redox balance.. Nat. Commun., 2014. [DOI | PubMed]
- P. H. Opgenorth, T. P. Korman, J. U. Bowie. A synthetic biochemistry module for production of bio-based chemicals from glucose.. Nat. Chem. Biol., 2016. [DOI | PubMed]
- M. A. Valliere, T. P. Korman, N. B. Woodall, G. A. Khitrov, R. E. Taylor, D. Baker, J. U. Bowie. A cell-free platform for the prenylation of natural products and application to cannabinoid production.. Nat. Commun., 2019. [DOI | PubMed]
- S. Sarria, B. Wong, H. G. Martin, J. D. Keasling, P. Peralta-Yahya. Microbial synthesis of pinene.. ACS Synth. Biol., 2014. [DOI | PubMed]
- A. S. Karim, M. C. Jewett. A cell-free framework for rapid biosynthetic pathway prototyping and enzyme discovery.. Metab. Eng., 2016. [DOI | PubMed]
- Q. M. Dudley, K. C. Anderson, M. C. Jewett. Cell-free mixing of Escherichia coli crude extracts to prototype and rationally engineer high-titer mevalonate synthesis.. ACS Synth. Biol., 2016. [DOI | PubMed]
- A. Casini, F. Y. Chang, R. Eluere, A. M. King, E. M. Young, Q. M. Dudley, A. Karim, K. Pratt, C. Bristol, A. Forget. A pressure test to make 10 molecules in 90 days: external evaluation of methods to engineer biology.. J. Am. Chem. Soc., 2018. [DOI | PubMed]
- Q. M. Dudley, C. J. Nash, M. C. Jewett. Cell-free biosynthesis of limonene using enzyme-enriched Escherichia coli lysates.. Synthetic Biology, 2019. [DOI | PubMed]
- D. U. Nielsen, X.-M. Hu, K. Daasbjerg, T. Skrydstrup. Chemically and electrochemically catalysed conversion of CO2 to CO with follow-up utilization to value-added chemicals.. Nat. Catal., 2018. [DOI]
- S. P. Long, E. A. Ainsworth, A. Rogers, D. R. Ort. Rising atmospheric carbon dioxide: plants FACE the future.. Annu. Rev. Plant Biol., 2004. [DOI | PubMed]
- R. J. Spreitzer, M. E. Salvucci. Rubisco: structure, regulatory interactions, and possibilities for a better enzyme.. Annu. Rev. Plant Biol., 2002. [DOI | PubMed]
- M. Könneke, D. M. Schubert, P. C. Brown, M. Hugler, S. Standfest, T. Schwander, L. Schada von Borzyskowski, T. J. Erb, D. A. Stahl, I. A. Berg. Ammonia-oxidizing archaea use the most energy-efficient aerobic pathway for CO2 fixation.. Proc. Natl. Acad. Sci. U. S. A., 2014. [DOI | PubMed]
- M. Scheffen, D. G. Marchal, T. Beneyton, S. K. Schuller, M. Klose, C. Diehl, J. Lehmann, P. Pfister, M. Carrillo, H. He. A new-to-nature carboxylation module to improve natural and synthetic CO2 fixation.. Nat. Catal., 2021. [DOI]
- T. Schwander, L. Schada von Borzyskowski, S. Burgener, N. S. Cortina, T. J. Erb. A synthetic pathway for the fixation of carbon dioxide in vitro.. Science, 2016. [DOI | PubMed]
- N. L. Gale, J. V. Beck. Evidence for the Calvin cycle and hexose monophosphate pathway in Thiobacillus ferrooxidans.. J. Bacteriol., 1967. [DOI | PubMed]
- T. E. Miller, T. Beneyton, T. Schwander, C. Diehl, M. Girault, R. McLean, T. Chotel, P. Claus, N. S. Cortina, J.-C. Baret, T. J. Erb. Light-powered CO2 fixation in a chloroplast mimic with natural and synthetic parts.. Science, 2020. [DOI | PubMed]
- S. Sundaram, C. Diehl, N. S. Cortina, J. Bamberger, N. Paczia, T. J. Erb. A modular in vitro platform for the production of terpenes and polyketides from CO2.. Angew. Chem., Int. Ed., 2021. [DOI]
- C. Diehl, P. D. Gerlinger, N. Paczia, T. J. Erb. Synthetic anaplerotic modules for the direct synthesis of complex molecules from CO2.. Nat. Chem. Biol., 2023. [DOI | PubMed]
- L. Xiao, G. Liu, F. Gong, H. Zhu, Y. Zhang, Z. Cai, Y. Li. A minimized synthetic carbon fixation cycle.. ACS Catal., 2022. [DOI]
- S. Bierbaumer, M. Nattermann, L. Schulz, R. Zschoche, T. J. Erb, C. K. Winkler, M. Tinzl, S. M. Glueck. Enzymatic conversion of CO2: from natural to artificial utilization.. Chem. Rev., 2023. [DOI | PubMed]
- T. Cai, H. Sun, J. Qiao, L. Zhu, F. Zhang, J. Zhang, Z. Tang, X. Wei, J. Yang, Q. Yuan. Cell-free chemo-enzymatic starch synthesis from carbon dioxide.. Science, 2021. [DOI | PubMed]
- J. Zhou, X. Tian, Q. Yang, Z. Zhang, C. Chen, Z. Cui, Y. Ji, U. Schwaneberg, B. Chen, T. Tan. Three multi-enzyme cascade pathways for conversion of C1 to C2/C4 compounds.. Chem. Catal., 2022. [DOI]
- J. Zhang, D. Liu, Y. Liu, H. Chu, J. Bai, J. Cheng, H. Zhao, S. Fu, H. Liu, Y. Fu. Hybrid synthesis of polyhydroxybutyrate bioplastics from carbon dioxide.. Green Chem., 2023. [DOI]
- F. Li, X. Wei, L. Zhang, C. Liu, C. You, Z. Zhu. Installing a green engine to drive an enzyme cascade: a light-powered in vitro biosystem for poly(3-hydroxybutyrate) synthesis.. Angew. Chem., Int. Ed., 2022. [DOI]
- H. Liu, M. A. Arbing, J. U. Bowie. Expanding the use of ethanol as a feedstock for cell-free synthetic biochemistry by implementing acetyl-CoA and ATP generating pathways.. Sci. Rep., 2022. [DOI | PubMed]
- E. Bailoni, M. Partipilo, J. Coenradij, D. A. J. Grundel, D. J. Slotboom, B. Poolman. Minimal out-of-equilibrium metabolism for synthetic cells: a membrane perspective.. ACS Synth. Biol., 2023. [DOI | PubMed]
- K. Y. Lee, S. J. Park, K. A. Lee, S. H. Kim, H. Kim, Y. Meroz, L. Mahadevan, K. H. Jung, T. K. Ahn, K. K. Parker. Photosynthetic artificial organelles sustain and control ATP-dependent reactions in a protocellular system.. Nat. Biotechnol., 2018. [DOI | PubMed]
- S. Berhanu, T. Ueda, Y. Kuruma. Artificial photosynthetic cell producing energy for protein synthesis.. Nat. Commun., 2019. [DOI | PubMed]
- E. Bailoni, B. Poolman. ATP recycling fuels sustainable glycerol 3-phosphate formation in synthetic cells fed by dynamic dialysis.. ACS Synth. Biol., 2022. [DOI | PubMed]
- D. Blanken, D. Foschepoth, A. C. Serrao, C. Danelon. Genetically controlled membrane synthesis in liposomes.. Nat. Commun., 2020. [DOI | PubMed]
- M. Partipilo, E. J. Ewins, J. Frallicciardi, T. Robinson, B. Poolman, D. J. Slotboom. Minimal pathway for the regeneration of redox cofactors.. JACS Au, 2021. [DOI | PubMed]
- T. Pols, H. R. Sikkema, B. F. Gaastra, J. Frallicciardi, W. M. Smigiel, S. Singh, B. Poolman. A synthetic metabolic network for physicochemical homeostasis.. Nat. Commun., 2019. [DOI | PubMed]
- E. Katz, V. Privman. Enzyme-based logic systems for information processing.. Chem. Soc. Rev., 2010. [DOI | PubMed]
- Y. Benenson. Biomolecular computing systems: principles, progress and potential.. Nat. Rev. Genet., 2012. [DOI | PubMed]
- L. Grozinger, M. Amos, T. E. Gorochowski, P. Carbonell, D. A. Oyarzun, R. Stoof, H. Fellermann, P. Zuliani, H. Tas, A. Goni-Moreno. Pathways to cellular supremacy in biocomputing.. Nat. Commun., 2019. [DOI | PubMed]
- N. M. Ivanov, M. G. Baltussen, C. L. F. Regueiro, M. Derks, W. T. S. Huck. Computing arithmetic functions using immobilised enzymatic reaction networks.. Angew. Chem., Int. Ed., 2023. [DOI]
- A. J. Genot, T. Fujii, Y. Rondelez. Computing with competition in biochemical networks.. Phys. Rev. Lett., 2012. [DOI | PubMed]
- A. Pandi, M. Koch, P. L. Voyvodic, P. Soudier, J. Bonnet, M. Kushwaha, J. L. Faulon. Metabolic perceptrons for neural computing in biological systems.. Nat. Commun., 2019. [DOI | PubMed]
- S. Okumura, G. Gines, N. Lobato-Dauzier, A. Baccouche, R. Deteix, T. Fujii, Y. Rondelez, A. J. Genot. Nonlinear decision-making with enzymatic neural networks.. Nature, 2022. [DOI | PubMed]
- D. Hathcock, J. Sheehy, C. Weisenberger, E. Ilker, M. Hinczewski. Noise filtering and prediction in biological signaling networks.. IEEE Transactions on Molecular, Biological and Multi-Scale Communications, 2016. [DOI]
- J. O’Brien, A. Murugan. Temporal pattern recognition through analog molecular computation.. ACS Synth. Biol., 2019. [DOI | PubMed]
- T. McGrath, N. S. Jones, P. R. Ten Wolde, T. E. Ouldridge. Biochemical machines for the interconversion of mutual information and work.. Phys. Rev. Lett., 2017. [DOI | PubMed]
- M. Stern, A. Murugan. Learning without neurons in physical systems.. Annu. Rev. Condens. Matter Phys., 2023. [DOI]
- O. E. Shklyaev, A. C. Balazs. Interlinking spatial dimensions and kinetic processes in dissipative materials to create synthetic systems with lifelike functionality.. Nat. Nanotechnol., 2024. [DOI | PubMed]
- D. Bray. Protein molecules as computational elements in living cells.. Nature, 1995. [DOI | PubMed]
- B. van Sluijs, T. Zhou, B. Helwig, M. G. Baltussen, F. H. T. Nelissen, H. A. Heus, W. T. S. Huck. Inverse design of enzymatic reaction network states.. Nat. Commun., 2024. [DOI | PubMed]
- A. N. Zaikin, A. M. Zhabotinsky. Concentration wave propagation in two-dimensional liquid-phase self-oscillating system.. Nature, 1970. [DOI]
