Enhanced Krylov Methods for Molecular Hamiltonians: Reduced Memory Cost and Complexity Scaling via Tensor Hypercontraction
Abstract
We introduce an algorithm that is simultaneously memory-efficient and low-scaling for applying ab initio molecular Hamiltonians to matrix-product states (MPS) via the tensor-hypercontraction (THC) format. These gains carry over to Krylov subspace methods, which can find low-lying eigenstates and simulate quantum time evolution while avoiding local minima and maintaining high accuracy. In our approach, the molecular Hamiltonian is represented as a sum of products of four MPOs, each with a bond dimension of only 2. Iteratively applying the MPOs to the current quantum state in MPS form, summing and recompressing the MPS leads to a scheme with the same asymptotic memory cost as the bare MPS and reduces the computational cost scaling compared to the Krylov method using a conventional MPO construction. We provide a detailed theoretical derivation of these statements and conduct supporting numerical experiments to demonstrate the advantage. Our algorithm is highly parallelizable and thus lends itself to large-scale HPC simulations.
Affiliations: † Department of Computer Science, 9184Technical University of Munich, CIT, Boltzmannstraße 3, 85748 Garching, Germany; ‡ Max Planck Institute of Quantum Optics, Hans-Kopfermann-Straße 1, 85748 Garching, Germany; § Munich Center for Quantum Science and Technology, Schellingstraße 4, 80799 München, Germany; ∥ Institute for Advanced Study, Technical University of Munich, Lichtenbergstraße 2a, 85748 Garching, Germany
License: © 2025 The Authors. Published by American Chemical Society CC BY 4.0 This article is licensed under CC-BY 4.0
Article links: DOI: 10.1021/acs.jctc.5c00525 | PubMed: 40601313 | PMC: PMC12288016
Relevance: Relevant: mentioned in keywords or abstract
Full text: PDF (2.9 MB)
Introduction
We aim to simulate a molecular Hamiltonian, which is also known as electronic structure Hamiltonian, of the form (with L the number of electronic spatial orbitals)
using tensor network methods, specifically the matrix product state (MPS) formalism.ref1−ref2ref3 and a p,σ are the Fermionic creation and annihilation operators, respectively, and t pq, v pqrs are coefficients resulting from single- and two-body orbital overlap integrals. To understand molecular properties such as the electronic structure,ref4,ref5 optoelectronic properties,ref. ref6 or molecular vibrations,ref. ref7 the density matrix renormalization group (DMRG) methodref8,ref9 is widely applied to chemical systems with strong correlations, where traditional density functional theory and coupled cluster approaches face significant challenges.ref3,ref7,ref10−ref11ref12ref13ref14ref15ref16 The development of attosecond-level experimental techniquesref17−ref18ref19ref20ref21ref22ref23ref24 motivates the simulation of ultrafast electron dynamics since they determine the formation and breaking of chemical bonds.ref. ref21 The time-dependent variational principle (TDVP) is a widely used time evolution method to predict electron dynamics.ref5,ref25−ref26ref27ref28ref29 However, both of the above are variational methods in which the MPS evolves locally. It could result in the DMRG method getting trapped in local minimaref26,ref30 and might lead to an inaccurate time evolution simulation by TDVPref31,ref32 even for simple models.ref. ref33
In contrast, global Krylov subspace methods optimize all the sites globally and simultaneously,ref. ref34 offering a reliable alternative method if DMRG or TDVP run into problems. Krylov subspace methods like the Lanczos algorithmref35−ref36ref37 can compute low-energy eigenstates reliably without local minima. The Lanczos algorithm also has the favorable capability of finding multiple excited states, being less sensitive to the results of lower eigenstates.ref38,ref39 Conversely, using the DMRG algorithm, one has to explicitly project out the lower eigenstates, implying that inaccuracies propagate to the higher ones. To simulate time evolution, the global Krylov methodfn1 provides high-order error scalingref28,ref40,ref41 and works reliably. The value of the Krylov method lies in its ability to ensure accuracy while remaining robust across all models, rather than failing in a few cases like DMRG and TDVP methods.
However, the reachable system sizes and MPS bond dimensions in the Krylov methods are relatively small when using the molecular Hamiltonian in conventional matrix product operator (MPO) form. Especially when one chooses high-accuracy MPS compression methods such as singular value decomposition (SVD),ref28,ref42 the restriction results from the core step: applying the Hamiltonian to a quantum state, i.e., computing H|ψ⟩ in the tensor network formalism and compressing it. Considering a molecular Hamiltonian of the form (eq1), the maximum bond dimension D scales as when using conventional MPO constructions.ref11,ref12,ref43 One needs intensive memory to store H|ψ⟩, whose bond dimension is the product of the MPS and MPO bond dimensions. Moreover, compressing H|ψ⟩ to MPS form with smaller bond dimensions is essential for further calculations;ref34,ref41 the computational cost is also high. The difficulty arises from the nonlocality of the two-body integral tensor , which makes the molecular Hamiltonian more complicated than a Hamiltonian containing only local interactions.
In this work, we propose and study an alternative Krylov method based on the tensor hypercontraction (THC) representation of v ref44−ref45ref46
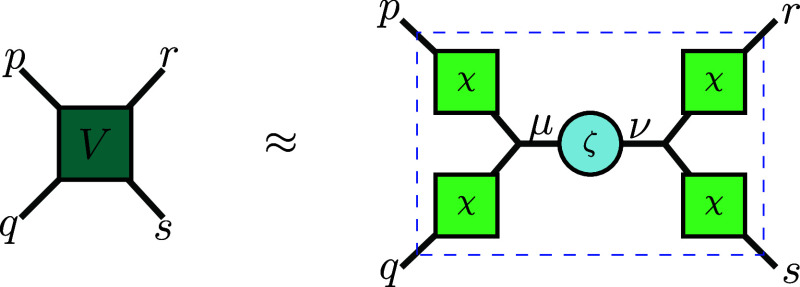
where N is the THC rank. This formulation involves only two distinct matrices χ and ζ, as illustrated in Figure . We will show that the THC representation allows us to rewrite the electronic Hamiltonian into a sum of sub-Hamiltonians, denoted THC-MPO, where each sub-Hamiltonian can be constructed as the product of four small MPOs with bond dimensions of only 2. Compared to calculations using a conventional MPO, such a small and constant bond dimension enables us to compute and compress H|ψ⟩ with significantly reduced memory requirements and better complexity scaling; both are reduced by a factor of asymptotically. We demonstrate the advantages of our THC-MPO by utilizing it for low-lying eigenstates search and time evolution simulations based on Krylov subspace methods, exemplified by the water molecule H2O, hydrogen chain with ten atoms H10, and the Ammonia molecule NH3. This allows us to track the accuracy and error sources by comparing them to results from the full configuration interaction (FCI) or exact diagonalization (ED) method. The numerical experiments show that our method enables us to calculate previously inaccessible system sizes when using Krylov methods with SVD compression; the memory advantages of our method become immediately apparent. Additionally, we will provide a general estimation of the computational complexity of larger molecules to highlight the potential. We will also illustrate why our method is well-suited for parallel computing.
Theoretical Background
Matrix Product States and Operators
In the tensor network framework, the wave function |Ψ⟩ is typically represented as a matrix product state (MPS), also called tensor trainref1,ref2,ref42,ref47
Each A[i] is a tensor of order three, as shown in Figure a. The superscript n i is a physical index enumerating the possible states at site i, and A[i]n i is a χi × χi+1 matrix for each n i. The variable χi is the i-th bond dimension. We denote the maximum MPS bond dimension by M in the following.
Alongside MPS, there is a corresponding formalism for operators, known as matrix product operators (MPOs),ref11,ref12,ref42 given by
Each element is a matrix of shape βi × βi+1, where βi is the i-th bond dimension, as shown in Figure b. We denote the maximum MPO bond dimension by D in the following. In eq eq4 , an operator Ô is represented with respect to computational basis states |m 1, …, m L⟩⟨n 1, …, n L|, and the corresponding coefficients are obtained by contracting the W matrices.
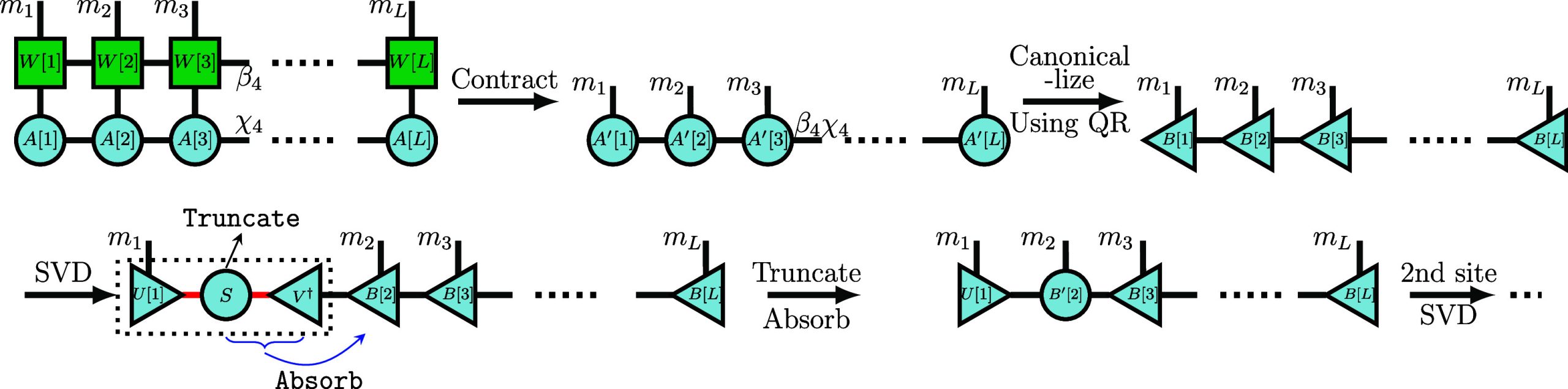
Analogous to how applying a Hamiltonian matrix to a state vector yields a state vector, applying an MPO to an MPS will result in a new MPS by contracting the local physical tensor legs. Such an operation will increase the MPS bond dimension from M to D·M. Thus, it is significant to compress this resulting MPS for further calculations, especially for iterative applications appearing in Krylov methods. The combined application and compression procedure is illustrated in Figure .
The Krylov Methods Based on Matrix Product States
The complexity of the Hamiltonian in full matrix form scales exponentially with system size; thus, exact diagonalization approaches are restricted to relatively small systems. A possibility to overcome this restriction consists in combining the Lanczos algorithmref35,ref36 with the MPS representation.ref37,ref48 Namely, the MPS ansatz offers an economical representation of quantum states, and the Lanczos algorithm confines the evolving space to the Krylov subspace spanned by {|ψ⟩, H|ψ⟩, H 2|ψ⟩, …, H K–1|ψ⟩}. In this work, we construct an orthogonal basis of the Krylov subspace. Starting from some initial state |v 0⟩ ≡|ψ⟩, we compute the next Krylov vector |v i+1⟩ by applying the Hamiltonian to |v i⟩ and orthogonalizing it w.r.t. the previous ones using the Gram-Schmidt algorithm.ref34,ref37,ref41 It is also possible to use these Krylov vectors without orthogonalization; see ref ref34,ref41 for further details.
The Hamiltonian is projected onto the Krylov subspace, and the elements of the resulting effective Hamiltonian are given by
where {|v 0⟩, |v 1⟩, …, |v K–1⟩} are orthonormalized Krylov vectors forming a basis of the subspace. Typically, we assume this effective Hamiltonian to be tridiagonal so that we only retain the entries with |i – j| ≤ 1.ref34,ref37 Assuming that the subspace dimension K is sufficiently large, the diagonalization of this effective Hamiltonian H̃ provides a reliable estimate of the low-lying eigenstates. Such a procedure is the well-known Lanczos algorithm.ref35−ref36ref37 Regarding the Hamiltonian as a linear combination of eigenspace projectors, the Krylov space can only contain components already present in the initial state. Therefore, one can also obtain excited states starting from an initial state orthogonal to the lower eigenstates.
Time evolution can also be simulated based on the Krylov subspace,ref34,ref41,ref49−ref50ref51 where the Krylov vectors {|v 0⟩, |v 1⟩, …, |v K–1⟩} are built based on the initial state at t = 0. A general quantum state in such a subspace can be written as
We represent the state as , where a i refers to the amplitude for each basis vector. With the help of such a formalism, the time-evolved state in the Krylov subspace is formulated as
where is (1,0,0,···,0)T since the initial state is just |v 0⟩. One can explicitly reconstruct the time-evolved quantum state by eq eq6 . The accuracy of this time evolution method depends on the size of the Krylov subspace, and the error is of order for a single time step δ and thus for a fixed duration.ref. ref34 The accuracy varies for different models: the upper error bounds are determined by the spectral width W, the step size δ, and the subspace size N; see Appendix A for quantitative discussion. As a practical guide, a subspace dimension of 3∼10 is typically sufficient to achieve satisfactory accuracy when selecting small time step sizes, as suggested in ref ref. ref34.
When using the MPS formalism to implement these algorithms, extra errors are introduced due to the MPS truncation, particularly the loss of orthogonality of the Krylov basis. We employ the strategy proposed in ref ref. ref37 to address this issue, and we noted that the canonical orthogonalization method might also be useful.ref. ref52 Both these techniques aim at finding a linear combination
of current Krylov vectors |v i⟩, so that the resulting vectors |ψa⟩ are well-orthogonalized. Note that we do not change the |v i⟩ vectors; instead, we only focus on solving the matrix C. Consequently, the elements of effective Hamiltonian in the |ψa⟩ basis are given by
The MPS truncation will still reduce accuracy even though the Krylov vectors are well orthogonalized; we find that restarting the Lanczos algorithm is helpful in improving the convergence. For simulating time evolution, truncation errors become significant only at very small time steps.
The Krylov method’s most expensive and memory-intensive part is obtaining the Krylov vectors whose core step is to compute H|v i⟩. Conventionally, one multiplies the Hamiltonian’s MPO with an MPS, resulting in an intermediate MPS with a large maximum bond dimension for ab initio molecular Hamiltonians. The memory cost to store such an intermediate MPS scales as for all sites, which could exhaust the available memory for even small-sized systems. Second, the high computational cost of compressing the intermediate MPS of H|v i⟩ is another bottleneck of constructing the Krylov subspace. One must compress the bond dimensions of H|v i⟩ back to smaller target bond dimensions to avoid the exponential increase in the next multiplications. To achieve fully controllable and highly accurate truncation, one typically employs the SVD method. In this approach, one first brings the intermediate MPS into canonical form using QR decomposition and then truncates the bonds by SVD, as depicted in Figure . Given that one can easily read off the Schmidt values from the mixed-canonical form, the truncation can be performed with a desired accuracy.ref42,ref53 The QR decompositions are the main contributors to the cost of compression. The computational complexity of QR decomposition for an MPS tensor with bond dimension is , leading to a total complexity across all sites as high as which makes it challenging to apply the Krylov method on large molecules.
There are also alternative MPS compression methods. The zip-up methodref. ref54 is more efficient, but since the algorithm works on a nonorthogonalized basis, the error is not fully controlled. The variational method requires a proper initial guess. Otherwise, one needs a large number of iterations and sweeps.ref. ref42 The recently proposed density matrix methodref. ref55 provides another fully controllable compression scheme that merits further study in future research. Our THC-MPO discussed in this paper can improve most of these MPS compression schemes when simulating molecular Hamiltonians, see Appendix B for more details; we focus on the traditional SVD method in this paper. We would like to emphasize that our method is compatible with all the MPS compression methods, and we can always check whether our method can be integrated into them when a newer compression scheme is proposed.
The THC Factorization
Employed widely in the simulation of molecular systems already,ref46,ref56−ref57ref58 the tensor hypercontraction (THC) proposed by Hohenstein et al.ref46,ref56,ref57 approximates the two-electron integrals v pqrs as
for all p, q, r, s ∈ {1, …, L}, as illustrated in Figure .
Currently, several relatively mature algorithms exist to obtain these tensors. The original papers proposed the PF-THCref. ref44 and LS-THCref. ref45 methods as algorithms. Subsequently, the interpolative separable density fitting (ISDF) methodref. ref59 enhanced the computational efficiency and improved the approximation accuracy. In density-fitting (DF),ref60−ref61ref62 one approximates the product of two orbitals as
where P μ for μ = 1, 2, …, N a are auxiliary basis functions. The idea of ISDF is that if we approximate ρpq by interpolation, the THC factorization can directly be obtainedref. ref59
where r k are selected grid points in the Becke scheme. The selection is implemented by interpolative decomposition, aimed at choosing a limited number of rows to approximate ρpq(r k) interpreted as a N g × L 2 matrix, where N g is the total number of Becke grid points. Since the row indices represent individual grid points, the procedure can also be interpreted as discarding less important grid points. Their importance is revealed by randomized QR decomposition with column-pivoting.ref59,ref63 We then determine fit functions F k after we obtain selected grid points. The fit functions are chosen as auxiliary basis functions P μ ref ref. ref59, but in this work, we obtained them following the strategy introduced in LS-THC,ref. ref45 as suggested in ref ref. ref56.
To improve the accuracy of the THC decomposition, we minimize the relative error
where denotes the Frobenius norm. It is essentially an optimization problem, and we carried it out using the Adam optimizer,ref. ref64 implemented in Optax.ref. ref65 Although Adam was introduced for stochastic optimization problems, its adaptive moment mechanism also converges reliably in deterministic settings. In our tests, Adam converged faster than classical optimizers such as the Broyden-Fletcher-Goldfarb-Shanno (BFGS) algorithm,ref. ref66 so we adopted it for the optimization step. It appears that there are two matrices, namely χ and ζ, to be optimized. However, since ζ can be obtained from χ by LS-THC, the number of free parameters is reduced to the entries of χ. Exemplified by the hydrogen chain, we first carry out the optimization by 1000 rounds with a learning rate of 0.001, followed by another 1000 rounds with a learning rate of 0.0005. In the numerical experiment, we reach the acceptable chemical accuracy (1.6 mHatree) with N = 4L for the water molecule H2O, N = 3L – 3 for the hydrogen chain H10, and N = 4.5L for the Ammonia molecule NH3, all in the STO-6G basis set. We ensure the accuracy reaches chemical accuracy by comparing it with the energy obtained via FCI results from PySCF. Regarding our future studies in larger systems, we noticed that previous studies have demonstrated that the THC rank N typically exhibits near-linear scaling with respect to the system size L.ref44−ref45ref46,ref56,ref59,ref63,ref67−ref68ref69 For instance, there are 76 spatial orbitals needed for the active-space model of the FeMoco system proposed by Li et al.,ref. ref70 and the THC rank of 450 is enough to reach the chemical accuracy;ref. ref67 for hydrogen chain of L atoms (L spatial orbitals in STO-6G basis) with distances of 1.4 Å, a THC rank of 3L – 3 is sufficient to achieve the accuracy of 5 × 10–5 Hartree per atom.ref. ref71 We remark that this part is a preprocessing step. If a prepared THC decomposition is available (e.g., from other work or a database), we can apply our THC-MPO method directly.
The Krylov Method Based on THC
Constructing MPOs Using THC
In this section, we first show how to use the THC factorization to construct a special representation of the molecular Hamiltonian (THC-MPO). Then, we will utilize the THC-MPO in Krylov methods and discuss its advantages. We focus on the challenging Coulomb term here. An MPO of the kinetic term T can be easily constructed following the strategy introduced in ref ref. ref11, and we will also discuss the kinetic term in Section sec3.2 .
Inserting the THC factorization eq eq10 into the Coulomb term V, one immediately arrives at
where G μσ,νσ′ is defined as
Each subterm (exemplified by ) in G μσ,νσ′ can explicitly be converted to an MPO as follows
and the first and last tensors
One can contract the W matrices sequentially to verify the correctness of the construction. It is worth noting that the bond dimension of W[s] is always only 2, independent of the system size L.
In this work, we follow the convention of treating each spatial orbital as a single site in the MPS. When implementing a corresponding MPO numerically using the Jordan-Wigner transformation,ref. ref72 we replace the Fermionic operators with their bosonic counterparts and substitute the identities in each W at position (1,1) by Pauli-Z operators (to account for Fermionic sign factors)
where I 4 denotes the identity matrix of size 4 × 4, and b s,σ′ is defined as the local annihilation operator for spin σ′ of size 4 × 4, for which we detail these in Appendix C.
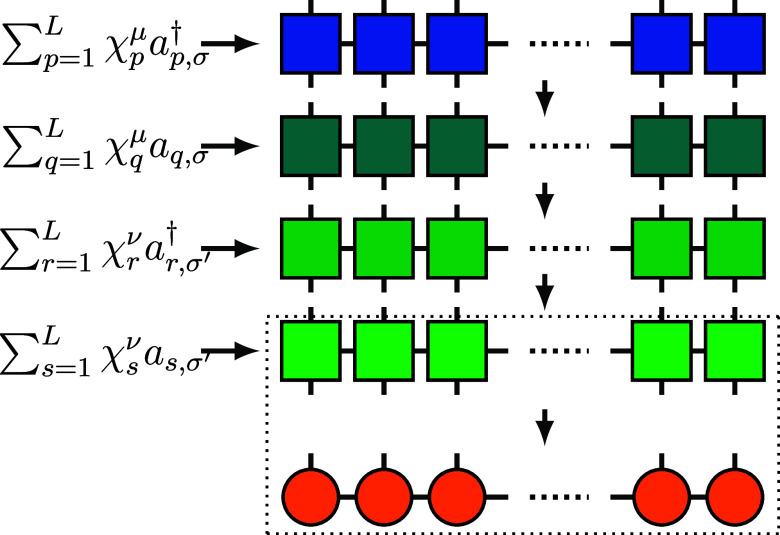
Following the strategy above, one can analogously construct MPOs for the other three subterms: , and . The entire MPO of G μσ,νσ′ is thus the product of the MPOs of these four subterms as shown in Figure , and the scalar ζμν can be absorbed into W[1] at the first site. Therefore, the MPO of G μσ,νσ′ likewise has a constant bond dimension. The whole MPO of the Coulomb term is thus the summation of MPOs of sub-Hamiltonians G μσ,νσ′, but to calculate V|ψ⟩ in compressed MPS form, we will refrain from merging them into a large MPO, see below.
Krylov Method Using THC-MPO
As discussed in Section sec2.2 , the essential step in Krylov methods is multiplying H with |ψ⟩. Here, we focus on the Coulomb term V in H = T + V. We present how to take advantage of our THC-MPO to execute the multiplication and subsequent compression. With the help of eq eq14 , we can write V|ψ⟩ as
where we apply each sub-Hamiltonian to |ψ⟩ and sum the resulting states up. Instead of manipulating large matrices, we compute V|ψ⟩ via the small MPOs of G μσ,νσ′.
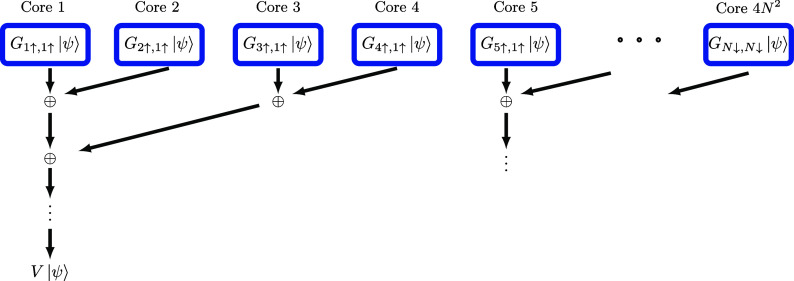
For each term G μσ,νσ′|ψ⟩, we execute multiplication and compression for each elementary MPO (layers shown in Figure ) sequentially instead of treating G μσ,νσ′ as a whole. Each compression returns the bond dimension to M so that the maximum bond dimension is only 2M during the calculation (since the MPO bond dimension for each layer is 2). Beginning with G 1↑,1↑ |ψ⟩, we add each subsequent G μσ,νσ′|ψ⟩. Such an MPS addition likewise leads to an intermediate MPS of bond dimension 2M, which is still cheap to store and compress. In summary, memory is required to store the largest intermediate MPS, which is less by a factor compared to a conventional MPO algorithm. In addition, the memory for storing G μσ,νσ′|ψ⟩ and G μσ,νσ′ is immediately released after adding G μσ,νσ′|ψ⟩ to others. Implementing eq eq18 is flexible regarding the order of additions.
Another optimization can be achieved by reusing intermediate results. We first notice that one can write G μσ,νσ′ as
where
and similarly for G μσ. It indicates that for two sub-Hamiltonians G aτ,νσ′ and G bκ,νσ′ that share the same latter two indices, the term G νσ′ can be factored out. Therefore, the intermediate state G νσ′|ψ⟩, which is obtained from compressing the two elementary MPOs (layers) in G νσ′|ψ⟩, can be reused. By applying this optimization, we reduce the computational cost by nearly half. Alg. 1 One includes all these steps and illustrates the overall algorithm as pseudocode.
In practice, we must take the kinetic term T into account as well. The conventional MPO representation of T has bond dimension , which leads to an overall memory requirement of to store T|ψ⟩ as MPS (without compression). We can improve on that situation using similar ideas as for the interaction term: We perform a spectral decomposition of (t pq) in eq eq1 and construct a sum of products of elementary MPOs with bond dimension 2. Therefore, the memory requirement can be reduced to for obtaining a compressed MPS. We explain these steps in detail in Appendix D.
Numerical Results and Resource Estimation
Ground- and Low-Lying States Finding
To benchmark the MPS-based Lanczos algorithm using our THC-MPO, we apply our method to the water molecule H2O and the hydrogen chain H10 using the STO-6G basis. The electronic integrals and FCI reference are calculated by PySCF;ref73,ref74 the tensor network calculation is implemented with PyTeNet,ref. ref75 in which Abelian quantum number conservation laws (electron number and spin) are enforced. We chose these relatively small systems because they allow for easier analysis of error sources and algorithmic behavior. But even so, we will see that the memory advantage has been fully verified. To fully explore our approach’s computational complexity capabilities, we plan to switch to high-performance computers and utilize parallel computing to benchmark them for large systems in the future, as discussed in Section sec4.5 .
We first present the results of the water molecule using the STO-6G basis, which leads to 7 spatial orbitals (14 spinor orbitals). In this case, we limit the maximum MPS bond dimension for the Krylov vectors to 30. The THC rank N for H2O is set to 28, resulting in the Frobenius norm error ∥v – v′∥ ≈ 3 × 10–11, where v′ is the Coulomb term reconstructed by THC tensors according to eq eq10 .
While the Lanczos algorithm performs well with a random initial state, selecting a proper initial state can significantly speed up convergence. In practice, we start from a state close to the target state, obtained from a heuristic guess or a low-cost algorithm. In this work, we simply use the Hartree–Fock state as the initial state for ground state finding, where paired electrons occupy the five lowest-energy molecular spatial orbitals. Additionally, we excite the highest occupied orbital in the Hartree–Fock state to serve as the initial state for finding the first excited state since the Hartree–Fock state is orthogonal to the ground state. As shown in Figure , we obtain the ground state and the first excited state within acceptable chemical accuracy (1.6 mHartree) using only 15 and 35 krylov vectors, respectively. The first excited state energy converges much slower because the gap between the first excited state and the second excited state is smaller than the one between the ground state and the first excited state. The energy error is obtained by comparison with the numerically exact value calculated by the FCI method in PySCF.
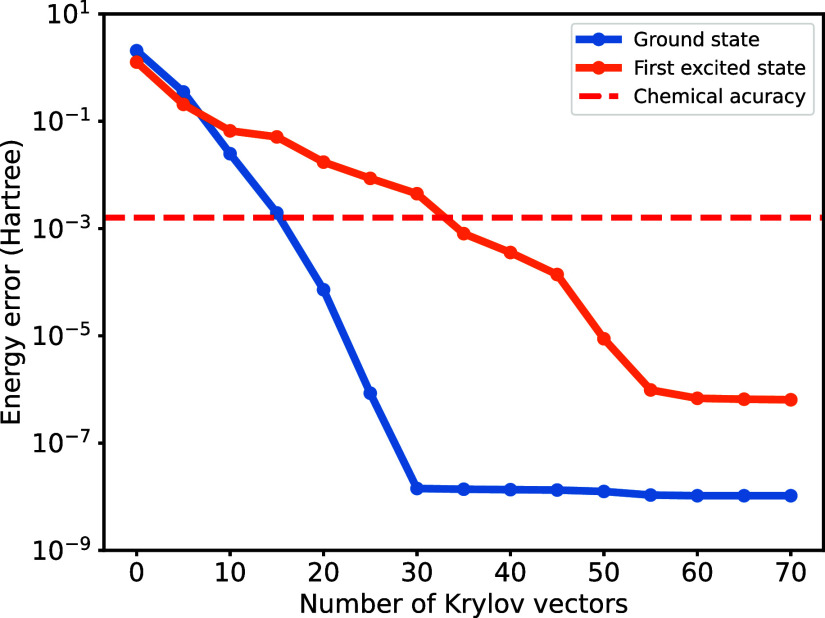
However, a high-accurate THC decomposition as above is unnecessary because, first, the total error is also bounded by MPS truncation. Second, a high THC rank will increase the computational cost. A way to reduce computational cost at the expense of accuracy is using a smaller THC rank N. To explore this possibility and quantify the resulting error, we study the hydrogen chain of ten atoms H10 with distances of 1.4 Å in the STO-6G basis, which leads to 10 spatial orbitals. Allowing a ground state energy error of 3 × 10–6 Hartree per atom, its THC rank is as low as 27.
The MPS bond dimensions for representing Krylov vectors are capped at 250. Like the previous example, we again use the Hartree–Fock ground and single-excited states as the initial states for the Krylov method. Interestingly, when using the exact Hamiltonian to calculate the energy expectation value for the approximated ground state
where |ψapprox⟩ is obtained by the THC-MPO-based Krylov method and H exact is the exact Hamiltonian, the resulting energy error is smaller than the error introduced by the THC approximation. While the THC approximation leads to an energy error of around 3 × 10–6 Hartree per atom, we can obtain the ground and first excited state with energy error ∼10–7 Hartree per atom, as illustrated in Figure . This indicates that accurate results could still be obtained using the THC-MPO, even when choosing a smaller THC rank that introduces non-negligible errors.
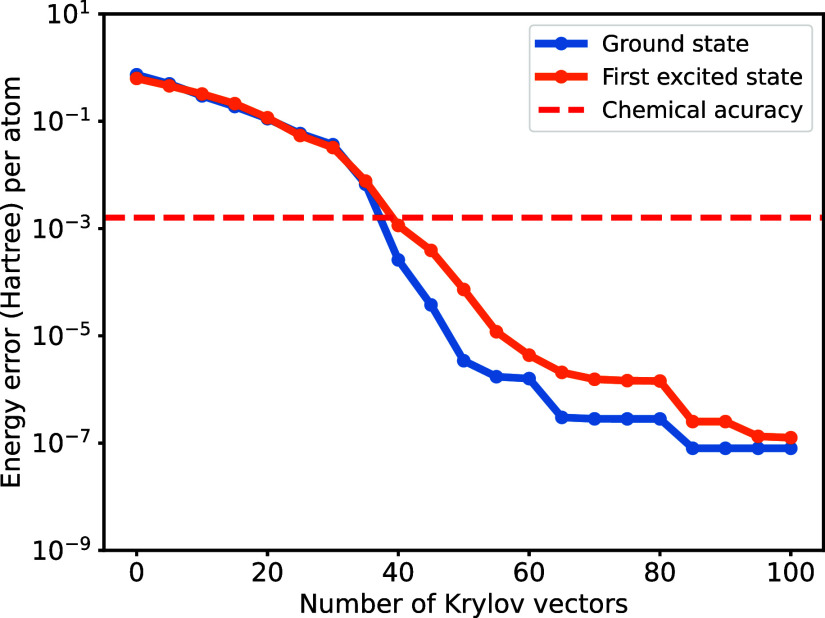
The results also suggest that although a large number of truncations is required to implement eq eq18 , the MPS truncations introduce only a minor error. Intuitively, assuming that each G μσ,νσ′|ψ⟩ term admits a relative error ϵ, the summation of them also admits a relative error ϵ, especially when allowing larger bond dimensions during reduction (and compress the final bond dimension back to M). Despite errors introduced by compressing the terms G μσ,νσ′|ψ⟩, additional errors also arise during the subsequent reduction (summation) process. Again, Assuming that pairwise addition-compression of these terms each results in a relative error ϵ′, it follows that each hierarchical reduction level similarly contributes a relative error of ϵ′. Given there are log(4N 2) = 2 log(N) + 2 reduction levels (as shown in Figure ), the cumulative error can thus be bounded by , assuming that the errors are all in different “directions”. Furthermore, since the reduction only manipulates the canonical MPS, increasing the bond dimension for this process will significantly enhance the accuracy without costing too much computational resources. Additionally, since the final MPS |ψ⟩ closely approximates the ground state (or other low-lying eigenstates), the bond dimension required to accurately represent the resulting MPO-MPS is expected to remain moderate. Therefore, many subterms should not contribute a much larger error.
Time Evolution Using Global Krylov Method
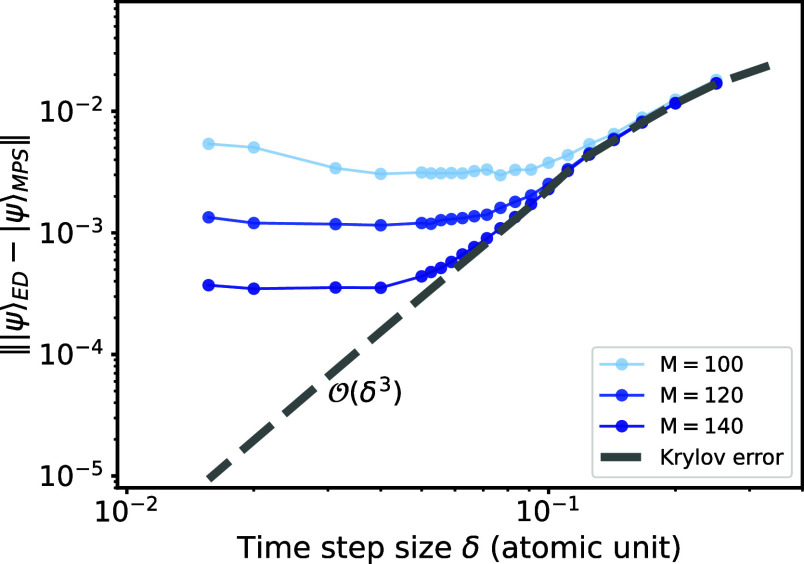
We also study the Krylov subspace time evolution based on our THC-MPO, where we set the subspace dimension to 4, leading to a single step error and total error for a fixed duration T for a single time step size δ. We apply the global Krylov method to the Ammonia molecule NH3 in the STO-6G basis, which leads to 8 spatial orbitals (16 spinor orbitals). The THC rank N for NH3 is set to 36, resulting in the Frobenius norm error ∥v – v′∥ ≈ 4 × 10–12. The initial state is defined as |ψ(t = 0)⟩ = a 3,↑ |ψ0⟩, where a spin-up electron is annihilated from the third spatial orbital of the ground state. Three factors determine the accuracy: SVD cutoff (bond dimension limitation), time step size δ, and the THC error from the THC factorization. The THC error is negligible for the NH3 molecule since the THC rank N = 4.5L results in a very accurate approximation.
As depicted in Figure , we measure the time evolution error for duration T = 1 atomic unit (a.u.) for different step sizes δ and maximum bond dimensions. The behavior of the errors can be explained well: As expected, the Krylov error dominates the overall error for larger time step sizes. Conversely, the scaling leads to small Krylov errors when the step size δ is reduced, causing the truncation error to dominate the overall error. To balance efficiency and accuracy, one can reach a sweet spot where the truncation error is comparable to the Krylov error. In Figure , it occurs where the total error curve converges with the Krylov error curve. For the case N = 4, one can observe that when setting M = 140, a step size of δ ∈ [0.05, 0.1] a.u. appears to be optimal.
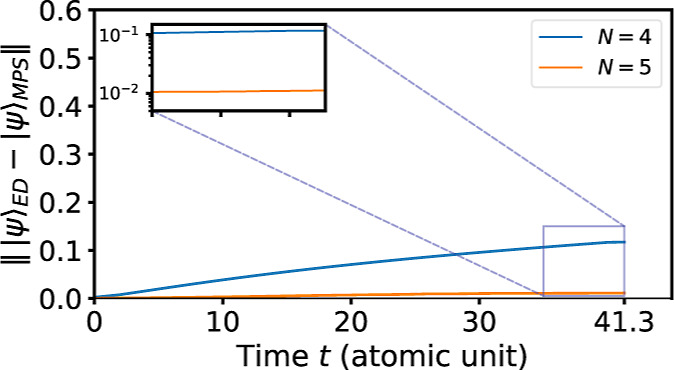
It is also meaningful to enlarge the Krylov subspace size and examine whether it would enhance the accuracy as predicted. Specifically, we calculate the time evolution for duration T = 41.3 au (1 fs) with M = 140, for both subspace size N = 4 and N = 5; the step size is set as δ = 0.1. As illustrated in Figure , the cumulative wave function error is 0.131 for N = 4 but only 0.011 for N = 5, indicating that the accuracy is enhanced by a factor of 10 when adding one more vector. For this time step size δ = 0.1, such a reduction is consistent with the expected error scaling .
Memory Consumption Comparison
One of the advantages of our THC-MPO method is the significantly reduced memory cost by a factor , We separately monitored memory consumption to store intermediate MPS in the Krylov algorithm based on the conventional MPO and the THC-MPO to test this prediction in our numerical experiments. We denote memory consumption when using conventional MPOs as P, and when using THC-MPOs as Q. Figure shows the quotient P/Q for the water molecule and hydrogen chains. By studying systems of different sizes, one clearly observes the predicted scaling difference. For example, considering the hydrogen chain of eight atoms, the memory required for storing an intermediate state H|ψ⟩ calculated with the conventional MPO amounts to 12,586 MB. In contrast, only 3.49 MB is needed when using the THC-MPO method, leading to a factor P/Q as large as 3606. We measure the memory cost by saving these intermediate MPS in HDF5 files and directly accessing their sizes. This case’s maximum bond dimension is 80, and we utilized double-precision complex numbers. Such a large memory usage is even too large to apply the Krylov methods on the H8 molecule. Therefore, in this respect, the H10 example has already proved the advantage over the original Krylov methods.
The Krylov method based on THC-MPO also outperforms the DMRG algorithm in terms of memory usage. Theoretically, the DMRG algorithm requires memory (to store the left and right environment blocks), which is times larger than the THC-MPO-based Krylov method. As shown in Figure , we numerically compare Q with the memory consumption S for the DMRG algorithm, using the same MPS bond dimensions. The results suggest that our method requires significantly less memory than the DMRG algorithm, and the observed values agree with the theoretically predicted scaling. Since the TDVP method could be implemented within a framework similar to the DMRG algorithm, our method also outperforms TDVP in terms of memory consumption when simulating time evolution. We do not continue to increase the system size to measure more cases since the memory usage for conventional MPO methods rapidly exceeds our available memory (32 GB), and the results shown in Figure are sufficient to demonstrate the memory advantage of our method. Due to memory constraints, the runtime comparison for large systems is also infeasible.
Computational Complexity Estimation
Besides memory consumption, the global Krylov methods based on our THC-MPO also perform better in terms of computational cost scaling than global Krylov methods using conventional MPOs. Here, we only present the summary; see Appendix E for a detailed derivation.
The primary contributor to the overall cost is obtaining compressed Krylov vectors. When using conventional MPO construction, renormalization is the most expensive step in compression. Since we have to handle the intermediate MPS with bond dimension , the renormalization has an overall complexity of for all sites. In contrast, for global Krylov methods utilizing THC-MPO, we only need to deal with MPS with bond dimension since the bond dimension of each layer in the subterms G μσ,νσ′ is only 2. Therefore, it costs to obtain G μσ,νσ′|ψ⟩ as compressed MPS, leading to an overall cost of for all G μσ,νσ′|ψ⟩ (when assuming that the THC rank N scales linearly with L). This computational cost has a large prefactor; it could be around 105 when taking hydrogen chains as an example. The large prefactor leads to longer run times for small molecules; for example, on a 13th-generation Intel Core i7-1355U CPU (12 cores, 1.70 GHz base frequency), the average wall-time runtime is 56 s for computing a Krylov vector for our H2O case, and 26 min is needed for a Krylov vector for our H10 case. The relatively slow performance can be attributed to our current implementation, which is not yet performance-oriented, and much of the multicore capacity is left idle. Switching to a more efficient programming language and applying further optimizations should substantially improve the runtime. We are actively developing a more efficient implementation.ref. ref76
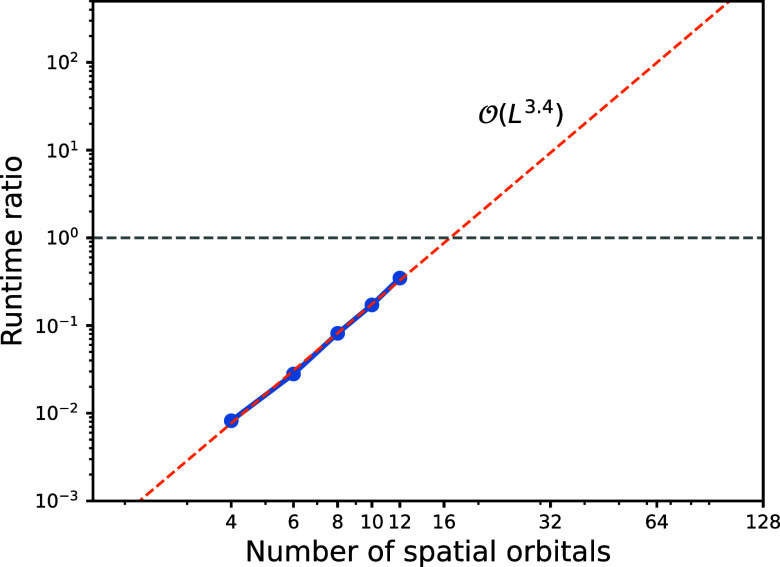
Nevertheless, we expect an advantage for medium- and large-sized molecules due to their promising scaling gap . To quantify the computational benefit of the THC-MPO method, we benchmark the wall-clock time required to compute the compressed MPS H|ψ⟩. Denote the runtime with the explicit (original) Hamiltonian by t o and that with THC-MPO by t THC, we report the runtime ratio t THC/t o, i.e., how many times faster the THC-MPO method is on hydrogen chains comprising 4–12 orbitals, as shown in Figure . Even though we only use a small bond dimension M = 30, the benchmark using conventional MPOs is still infeasible when L ≥ 14 due to the large memory requirements. The curve admits a scaling , which is not as perfect accurate as our predicts, but the obtained runtime ratio can clearly reveal the ratio trend, that is, our THC-MPO method should be more efficient when the system size goes beyond 20.
A Natural Scalable Parallelization Scheme
Parallel computing has been effectively integrated into DMRG algorithms for quantum chemistry to take advantage of high-performance computing platforms. This integration has significantly enhanced the ability to study large molecular systems; various parallel schemes were proposed,ref11,ref77−ref78ref79ref80ref81ref82 and notable open-source packages like Block2 were developed.ref. ref83 The Krylov method based on our THC-MPO can straightforwardly use the potential of parallel computing: to obtain H|ψ⟩ following eq eq18 , each of the 4N 2 subterms can be calculated and compressed independently, and the summation of these subterms can also be performed in parallel by a reduction.
More specifically, we propose a parallelism scheme as illustrated in Figure . For each core, we first assign the task of computing and compressing one (or several) subterms G μσ,νσ′|ψ⟩. The power of multiple cores can be perfectly utilized for this part. After this step, we add and compress these terms pairwise in parallel. It appears that some computational resources are idling during such a process, but the compression can utilize multiple cores for parallel computation when using packages like multithreaded LAPACK implementations.ref. ref84 Because the SVD and QR decomposition can be significantly sped up by parallel computing,ref85,ref86 the reduction part can utilize the power of parallel computing as well. Also, since each compressed term G μσ,νσ′|ψ⟩ has already been canonical form (up to a factor), the MPS addition-compression process does not contain the QR decomposition, which makes the reduction inexpensive. Another bottleneck of parallel computing is communication;ref. ref77 an extra advantage of our parallel scheme is that communication only occurs during the reduction.
As a preliminary demonstration (with more advanced systems and larger molecules planned for future work), we benchmark our parallel scheme on a 112-core node using OpenMP.ref. ref87 Each thread (core) is tasked with calculating and compressing G μσ,νσ′|ψ⟩. As displayed in Figure , we compare the runtime for computing H|ψ⟩ with a single thread versus K threads. The results indicate near-ideal speedups when multiple threads are utilized. Extending this approach to multiple nodes should also yield near-linear scaling because each node can achieve this speedup independently, and communication among nodes is only required once all nodes have completed their tasks.
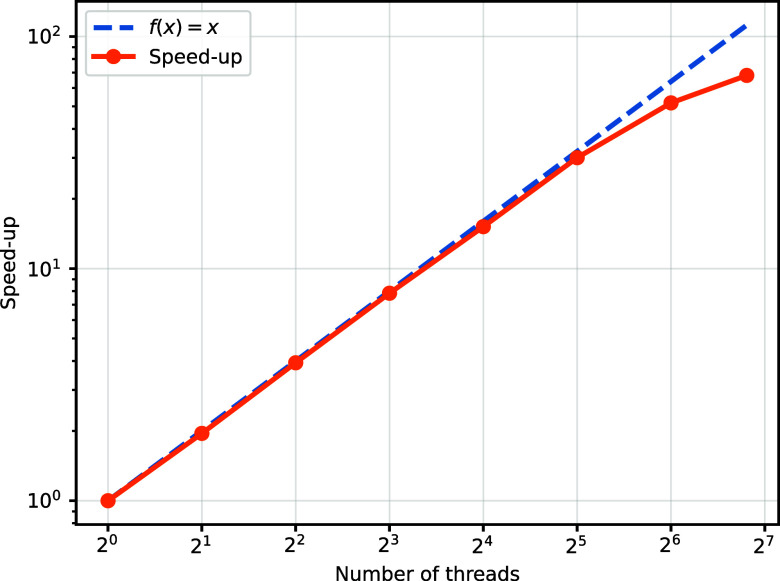
Due to such an efficient and scalable parallel scheme, the parallel runtime scales as under ideal parallelization conditions with 4N 2 available cores efficiently. Currently, advanced tensor network methods in quantum chemistry (e.g., DMRG) can utilize thousands of cores efficiently,ref77,ref82 the Krylov methods (Lanczos algorithm and Krylov time evolution method) based on THC-MPO have the potential to leverage cores scaling as 4N 2 with high efficiency, making it possible to surpass the current state of the art in CPU utilization. Note that each subtask shown in Figure can also be implemented by multiple cores (e.g., a node), thereby further increasing the number of cores we can efficiently utilize and decreasing the reduction depth.
Conclusions
The THC-MPO approach allows us to implement Krylov subspace methods, e.g., the Lanczos algorithm and the global Krylov method for time evolution, with reduced memory usage and lower computational cost scaling. When compared to the Krylov method based on the conventional MPO representation, the memory advantage of THC-MPO is apparent, even for the smallest molecules. Moreover, it outperforms popular methods like DMRG and TDVP in terms of memory consumption, suggesting that THC-MPO can potentially enable simulations of even larger systems than currently reachable by DMRG or TDVP. While the benefit of computational cost is not immediate for small molecules due to large prefactors, we expect that the improvement will become significant for moderate and large molecular systems. We emphasize that the THC-MPO is essentially a proper decomposition. A group of MPOs with a small bond dimension can also be achieved by building an MPO for each term in the molecular Hamiltonian, but the final computational complexity will be as large as if we do so.
A cornerstone of our work is the compressed THC representation of the two-body integral tensor v. A promising research direction (complementary to the present study) could be the exploitation of sparsity structures of v, for example, due to localized orbitals or wavelet-type orbitals supported on a fine grid. Also, the tensor v pqrs (which originates from overlap integrals) is symmetric with respect to interchanges of p ↔ q, r ↔ s, and (p, q) ↔ (r, s). The symmetries are passed on to the THC representation. It is worth exploring how to exploit such symmetries in our approach. We also noticed that our THC-MPO could help enable the computation of spectral functionsref52,ref88,ref89 for large-size molecular Hamiltonians when combining with the Chebyshev expansions, where multiplication-compression H|ψ⟩ also remains a major bottleneck. With our THC-MPO method and optimized parallel computing implementations, our primary plan is to explore these ideas and the reachable system sizes in future works.
References
- S. Östlund, S. Rommer. Thermodynamic Limit of Density Matrix Renormalization. Phys. Rev. Lett., 1995. [DOI | PubMed]
- S. Rommer, S. Östlund. Class of ansatz wave functions for one-dimensional spin systems and their relation to the density matrix renormalization group. Phys. Rev. B, 1997. [DOI]
- Ma, H. ; Schollwöck, U. ; Shuai, Z. Density Matrix Renormalization Group (DMRG)-Based Approaches in Computational Chemistry; Elsevier, 2022.
- G. K.-L. Chan, S. Sharma. The density matrix renormalization group in quantum chemistry. Annu. Rev. Phys. Chem., 2011. [DOI | PubMed]
- Y. Xu, Z. Xie, X. Xie, U. Schollwöck, H. Ma. Stochastic adaptive single-site time-dependent variational principle. JACS Au, 2022. [DOI | PubMed]
- J. Ren, Z. Shuai, G. Kin-Lic Chan. Time-dependent density matrix renormalization group algorithms for nearly exact absorption and fluorescence spectra of molecular aggregates at both zero and finite temperature. J. Chem. Theory Comput., 2018. [DOI | PubMed]
- A. Baiardi, C. J. Stein, V. Barone, M. Reiher. Vibrational density matrix renormalization group. J. Chem. Theory Comput., 2017. [DOI | PubMed]
- S. R. White. Density matrix formulation for quantum renormalization groups. Phys. Rev. Lett., 1992. [DOI | PubMed]
- S. R. White. Density-matrix algorithms for quantum renormalization groups. Phys. Rev. B, 1993. [DOI]
- Y. Xu, Y. Cheng, Y. Song, H. Ma. New density matrix renormalization group approaches for strongly correlated systems coupled with large environments. J. Chem. Theory Comput., 2023. [DOI | PubMed]
- G. K. Chan, A. Keselman, N. Nakatani, Z. Li, S. R. White. Matrix product operators, matrix product states, and ab initio density matrix renormalization group algorithms. J. Chem. Phys., 2016. [DOI | PubMed]
- S. Keller, M. Dolfi, M. Troyer, M. Reiher. An efficient matrix product operator representation of the quantum chemical Hamiltonian. J. Chem. Phys., 2015. [DOI | PubMed]
- S. Szalay, M. Pfeffer, V. Murg, G. Barcza, F. Verstraete, R. Schneider, O. Legeza. Tensor product methods and entanglement optimization for ab initio quantum chemistry. Int. J. Quantum Chem., 2015. [DOI]
- G. Friesecke, G. Barcza, O. ¨. Legeza. Predicting the FCI energy of large systems to chemical accuracy from restricted active space density matrix renormalization group calculations. J. Chem. Theory Comput., 2024. [DOI | PubMed]
- Y. Cheng, H. Ma. Renormalized-Residue-Based Multireference Configuration Interaction Method for Strongly Correlated Systems. J. Chem. Theory Comput., 2024. [DOI | PubMed]
- Z. Xie, Y. Song, F. Peng, J. Li, Y. Cheng, L. Zhang, Y. Ma, Y. Tian, Z. Luo, H. Ma. Kylin 1.0: An ab-initio density matrix renormalization group quantum chemistry program. J. Comput. Chem., 2023. [DOI | PubMed]
- K. Klünder, J. M. Dahlström, M. Gisselbrecht, T. Fordell, M. Swoboda, D. Guénot, P. Johnsson, J. Caillat, J. Mauritsson, A. Maquet, R. Taïeb, A. L’Huillier. Probing single-photon ionization on the attosecond time scale. Phys. Rev. Lett., 2011. [DOI | PubMed]
- F. Krausz, M. Ivanov. Attosecond physics. Rev. Mod. Phys., 2009. [DOI]
- F. Lépine, M. Y. Ivanov, M. J. Vrakking. Attosecond molecular dynamics: fact or fiction?. Nat. Photonics, 2014. [DOI]
- E. Goulielmakis, Z.-H. Loh, A. Wirth, R. Santra, N. Rohringer, V. S. Yakovlev, S. Zherebtsov, T. Pfeifer, A. M. Azzeer, M. F. Kling, S. R. Leone, F. Krausz. Real-time observation of valence electron motion. Nature, 2010. [DOI | PubMed]
- M. Nisoli, P. Decleva, F. Calegari, A. Palacios, F. Martín. Attosecond electron dynamics in molecules. Chem. Rev., 2017. [DOI | PubMed]
- A. Baltuška, T. Udem, M. Uiberacker, M. Hentschel, E. Goulielmakis, C. Gohle, R. Holzwarth, V. S. Yakovlev, A. Scrinzi, T. W. Hänsch, F. Krausz. Attosecond control of electronic processes by intense light fields. Nature, 2003. [DOI | PubMed]
- R. Borrego-Varillas, M. Lucchini, M. Nisoli. Attosecond spectroscopy for the investigation of ultrafast dynamics in atomic, molecular and solid-state physics. Rep. Prog. Phys., 2022. [DOI]
- K. Midorikawa. Progress on table-top isolated attosecond light sources. Nat. Photonics, 2022. [DOI]
- J. Haegeman, C. Lubich, I. Oseledets, B. Vandereycken, F. Verstraete. Unifying time evolution and optimization with matrix product states. Phys. Rev. B, 2016. [DOI]
- A. Baiardi, M. Reiher. The density matrix renormalization group in chemistry and molecular physics: Recent developments and new challenges. J. Chem. Phys., 2020. [DOI | PubMed]
- A. Baiardi. Electron dynamics with the time-dependent density matrix renormalization group. J. Chem. Theory Comput., 2021. [DOI | PubMed]
- J. Ren, W. Li, T. Jiang, Y. Wang, Z. Shuai. Time-dependent density matrix renormalization group method for quantum dynamics in complex systems. Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2022. [DOI]
- Y. Xu, C. Liu, H. Ma. Kylin-V: An open-source package calculating the dynamic and spectroscopic properties of large systems. J. Chem. Phys., 2024. [DOI | PubMed]
- W. Hu, G. K.-L. Chan. Excited-State Geometry Optimization with the Density Matrix Renormalization Group, as Applied to Polyenes. J. Chem. Theory Comput., 2015. [DOI | PubMed]
- B. Kloss, Y. B. Lev, D. Reichman. Time-dependent variational principle in matrix-product state manifolds: Pitfalls and potential. Phys. Rev. B, 2018. [DOI]
- S. Goto, I. Danshita. Performance of the time-dependent variational principle for matrix product states in the long-time evolution of a pure state. Phys. Rev. B, 2019. [DOI]
- M. Yang, S. R. White. Time-dependent variational principle with ancillary Krylov subspace. Phys. Rev. B, 2020. [DOI]
- S. Paeckel, T. Köhler, A. Swoboda, S. R. Manmana, U. Schollwöck, C. Hubig. Time-evolution methods for matrix-product states. Ann. Phys., 2019. [DOI]
- R. Haydock. The recursive solution of the Schrödinger equation. Solid State Phys., 1980. [DOI]
- C. Lanczos. An Iteration Method for the Solution of the Eigenvalue Problem of Linear Differential and Integral Operators. J. Res. Natl. Bur. Stand., 1950. [DOI]
- P. E. Dargel, A. Woellert, A. Honecker, I. McCulloch, U. Schollwöck, T. Pruschke. Lanczos algorithm with matrix product states for dynamical correlation functions. Phys. Rev. B, 2012. [DOI]
- Improved Lanczos Algorithm using Matrix Product States.. 2025
- Inexact subspace projection methods for low-rank tensor eigenvalue problems.. 2025
- E. Ronca, Z. Li, C. A. Jimenez-Hoyos, G. K.-L. Chan. Time-step targeting time-dependent and dynamical density matrix renormalization group algorithms with ab initio Hamiltonians. J. Chem. Theory Comput., 2017. [DOI | PubMed]
- L.-H. Frahm, D. Pfannkuche. Ultrafast ab initio quantum chemistry using matrix product states. J. Chem. Theory Comput., 2019. [DOI | PubMed]
- U. Schollwöck. The density-matrix renormalization group in the age of matrix product states. Ann. Phys., 2011. [DOI]
- J. Ren, W. Li, T. Jiang, Z. Shuai. A general automatic method for optimal construction of matrix product operators using bipartite graph theory. J. Chem. Phys., 2020. [DOI | PubMed]
- E. G. Hohenstein, R. M. Parrish, T. J. Martínez. Tensor hypercontraction density fitting. I. Quartic scaling second- and third-order Møller-Plesset perturbation theory. J. Chem. Phys., 2012. [DOI | PubMed]
- R. M. Parrish, E. G. Hohenstein, T. J. Martínez, C. D. Sherrill. Tensor hypercontraction. II. Least-squares renormalization. J. Chem. Phys., 2012. [DOI | PubMed]
- E. G. Hohenstein, R. M. Parrish, C. D. Sherrill, T. J. Martínez. Communication: Tensor hypercontraction. III. Least-squares tensor hypercontraction for the determination of correlated wavefunctions. J. Chem. Phys., 2012. [DOI | PubMed]
- J. C. Bridgeman, C. T. Chubb. Hand-waving and interpretive dance: An introductory course on tensor networks. J. Phys. A: Math. Theor., 2017. [DOI]
- P. E. Dargel, A. Honecker, R. Peters, R. M. Noack, T. Pruschke. Adaptive Lanczos-vector method for dynamic properties within the density matrix renormalization group. Phys. Rev. B, 2011. [DOI]
- T. J. Park, J. Light. Unitary quantum time evolution by iterative Lanczos reduction. J. Chem. Phys., 1986. [DOI]
- R. Kosloff. Time-dependent quantum-mechanical methods for molecular dynamics. J. Phys. Chem., 1988. [DOI]
- M. Hochbruck, C. Lubich. On Krylov subspace approximations to the matrix exponential operator. SIAM J. Numer. Anal., 1997. [DOI]
- T. Jiang, J. Ren, Z. Shuai. Chebyshev matrix product states with canonical orthogonalization for spectral functions of many-body systems. J. Phys. Chem. Lett., 2021. [DOI | PubMed]
- J. Hauschild, F. Pollmann. Efficient numerical simulations with tensor networks: Tensor Network Python (TeNPy). SciPost Phys. Lect. Notes, 2018. [DOI]
- E. M. Stoudenmire, S. R. White. Minimally entangled typical thermal state algorithms. New J. Phys., 2010. [DOI]
- L. Ma, M. Fishman, E. M. Stoudenmire, E. Solomonik. Approximate contraction of arbitrary tensor networks with a flexible and efficient density matrix algorithm. Quantum, 2024. [DOI]
- J. Lee, L. Lin, M. Head-Gordon. Systematically improvable tensor hypercontraction: Interpolative separable density-fitting for molecules applied to exact exchange, second-and third-order Møller–Plesset perturbation theory. J. Chem. Theory Comput., 2020. [DOI | PubMed]
- R. M. Parrish, C. D. Sherrill, E. G. Hohenstein, S. I. Kokkila, T. J. Martínez. Communication: Acceleration of coupled cluster singles and doubles via orbital-weighted least-squares tensor hypercontraction. J. Chem. Phys., 2014. [DOI | PubMed]
- R. Schutski, J. Zhao, T. M. Henderson, G. E. Scuseria. Tensor-structured coupled cluster theory. J. Chem. Phys., 2017. [DOI | PubMed]
- J. Lu, L. Ying. Compression of the electron repulsion integral tensor in tensor hypercontraction format with cubic scaling cost. J. Comput. Phys., 2015. [DOI]
- J. L. Whitten. Coulombic potential energy integrals and approximations. J. Chem. Phys., 1973. [DOI]
- B. I. Dunlap, J. Connolly, J. Sabin. On some approximations in applications of Xα theory. J. Chem. Phys., 1979. [DOI]
- H.-J. Werner, F. R. Manby, P. J. Knowles. Fast linear scaling second-order Møller–Plesset perturbation theory (MP2) using local and density fitting approximations. J. Chem. Phys., 2003. [DOI]
- W. Hu, L. Lin, C. Yang. Interpolative separable density fitting decomposition for accelerating hybrid density functional calculations with applications to defects in silicon. J. Chem. Theory Comput., 2017. [DOI | PubMed]
- Adam: A Method for Stochastic Optimization.. Proceedings of the 3rd International Conference on Learning Representations (ICLR)., 2015
- https://github.com/google-deepmind/optax (accessed May 2025).
- Nocedal, J. ; Wright, S. J. Numerical Optimization, 2nd ed.; Springer, 2006.
- J. Lee, D. W. Berry, C. Gidney, W. J. Huggins, J. R. McClean, N. Wiebe, R. Babbush. Even More Efficient Quantum Computations of Chemistry Through Tensor Hypercontraction. PRX Quantum, 2021. [DOI]
- D. A. Matthews. Improved grid optimization and fitting in least squares tensor hypercontraction. J. Chem. Theory Comput., 2020. [DOI | PubMed]
- K. Dong, W. Hu, L. Lin. Interpolative separable density fitting through centroidal voronoi tessellation with applications to hybrid functional electronic structure calculations. J. Chem. Theory Comput., 2018. [DOI | PubMed]
- Z. Li, J. Li, N. S. Dattani, C. Umrigar, G. K. Chan. The electronic complexity of the ground-state of the FeMo cofactor of nitrogenase as relevant to quantum simulations. J. Chem. Phys., 2019. [DOI | PubMed]
- M. Luo, J. I. Cirac. Efficient simulation of quantum chemistry problems in an enlarged basis set. PRX Quantum, 2025. [DOI]
- P. Jordan, E. Wigner. Über das Paulische Äquivalenzverbot. Z. Phys., 1928. [DOI]
- Q. Sun, T. C. Berkelbach, N. S. Blunt, G. H. Booth, S. Guo, Z. Li, J. Liu, J. D. McClain, E. R. Sayfutyarova, S. Sharma, S. Wouters, G. K.-L. Chan. PySCF: the Python-based simulations of chemistry framework. Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2018. [DOI]
- Q. Sun, X. Zhang, S. Banerjee, P. Bao, M. Barbry, N. S. Blunt, N. A. Bogdanov, G. H. Booth, J. Chen, Z.-H. Cui. Recent developments in the PySCF program package. J. Chem. Phys., 2020. [DOI | PubMed]
- C. B. Mendl. PyTeNet: A concise Python implementation of quantum tensor network algorithms. J. Open Source Softw., 2018. [DOI]
- https://github.com/qc-tum/chemtensor (accessed: May 2025).
- H. Zhai, G. K. Chan. Low communication high performance ab initio density matrix renormalization group algorithms. J. Chem. Phys., 2021. [DOI | PubMed]
- C. Xiang, W. Jia, W.-H. Fang, Z. Li. Distributed multi-GPU ab initio density matrix renormalization group algorithm with applications to the P-cluster of nitrogenase. J. Chem. Theory Comput., 2024. [DOI | PubMed]
- R. Olivares-Amaya, W. Hu, N. Nakatani, S. Sharma, J. Yang, G. K.-L. Chan. The ab-initio density matrix renormalization group in practice. J. Chem. Phys., 2015. [DOI | PubMed]
- G. K.-L. Chan. An algorithm for large scale density matrix renormalization group calculations. J. Chem. Phys., 2004. [DOI | PubMed]
- S. Wouters, D. Van Neck. The density matrix renormalization group for ab initio quantum chemistry. Eur. Phys. J. D, 2014. [DOI]
- J. Brabec, J. Brandejs, K. Kowalski, S. Xantheas, O. ¨. Legeza, L. Veis. Massively parallel quantum chemical density matrix renormalization group method. J. Comput. Chem., 2021. [DOI | PubMed]
- H. Zhai, H. R. Larsson, S. Lee, Z.-H. Cui, T. Zhu, C. Sun, L. Peng, R. Peng, K. Liao, J. Tölle, J. Yang, S. Li, G. K.-L. Chan. Block2: A comprehensive open source framework to develop and apply state-of-the-art DMRG algorithms in electronic structure and beyond. J. Chem. Phys., 2023. [DOI | PubMed]
- L. S. Blackford, J. Demmel, J. Dongarra, I. Duff, S. Hammarling, G. Henry, M. Heroux, L. Kaufman, A. Lumsdaine, A. Petitet, R. Pozo, K. Remington, R. C. Whaley. An updated set of basic linear algebra subprograms (BLAS). ACM Trans. Math. Software, 2002. [DOI]
- J. Demmel, L. Grigori, M. Hoemmen, J. Langou. Communication-optimal parallel and sequential QR and LU factorizations. SIAM J. Sci. Comput., 2012. [DOI]
- E. R. Jessup, D. C. Sorensen. A parallel algorithm for computing the singular value decomposition of a matrix. SIAM J. Matrix Anal. Appl., 1994. [DOI]
- L. Dagum, R. Menon. OpenMP: an industry standard API for shared-memory programming. IEEE Comput. Sci. Eng., 1998. [DOI]
- Y. Yang, S. Iblisdir, J. I. Cirac, M. C. Banuls. Probing thermalization through spectral analysis with matrix product operators. Phys. Rev. Lett., 2020. [DOI | PubMed]
- A. Holzner, A. Weichselbaum, I. P. McCulloch, U. Schollwöck, J. von Delft. Chebyshev matrix product state approach for spectral functions. Phys. Rev. B, 2011. [DOI]
- M. Hochbruck, C. Lubich. On Krylov Subspace Approximations to the Matrix Exponential Operator. SIAM J. Numer. Anal., 1997. [DOI]
- Golub, G. H. ; Van Loan, C. F. Matrix Computations; Johns Hopkins University Press, 2013.
- M. Gu, S. C. Eisenstat. Efficient algorithms for computing a strong rank-revealing QR factorization. SIAM J. Sci. Comput., 1996. [DOI]
- Anderson, E. ; Bai, Z. ; Bischof, C. ; Blackford, L. S. ; Demmel, J. ; Dongarra, J. ; Du Croz, J. ; Greenbaum, A. ; Hammarling, S. ; McKenney, A. ; Sorensen, D. LAPACK Users’ Guide; Society for Industrial and Applied Mathematics, 1999.
