Current and future perspectives on the regulation and functions of miR-545 in cancer development
Abstract
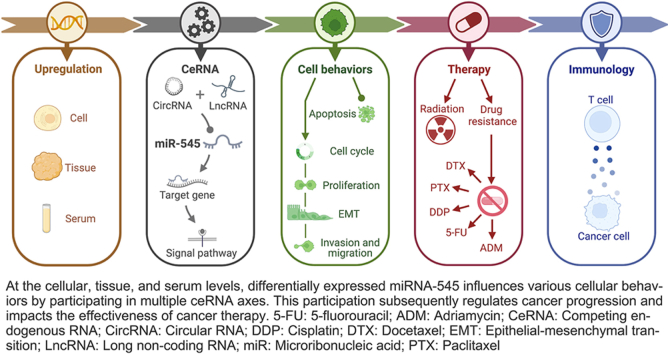
Micro ribonucleic acids (miRNAs) are a highly conserved class of single-stranded non-coding RNAs. Within the miR-545/374a cluster, miR-545 resides in the intron of the long non-coding RNA (lncRNA) FTX on Xq13.2. The precursor form, pre-miR-545, is cleaved to generate two mature miRNAs, miR-545-3p and miR-545-5p. Remarkably, these two miRNAs exhibit distinct aberrant expression patterns in different cancers; however, their expression in colorectal cancer remains controversial. Notably, miR-545-3p is affected by 15 circular RNAs (circRNAs) and 10 long non-coding RNAs (lncRNAs), and it targets 27 protein-coding genes (PCGs) that participate in the regulation of four signaling pathways. In contrast, miR-545-5p is regulated by one circRNA and five lncRNAs, it targets six PCGs and contributes to the regulation of one signaling pathway. Both miR-545-3p and miR-545-5p affect crucial cellular behaviors, including cell cycle, proliferation, apoptosis, epithelial-mesenchymal transition, invasion, and migration. Although low miR-545-3p expression is associated with poor prognosis in three cancer types, studies on miR-545-5p are yet to be reported. miR-545-3p operates within a diverse range of regulatory networks, thereby augmenting the efficacy of cancer chemotherapy, radiotherapy, and immunotherapy. Conversely, miR-545-5p enhances immunotherapy efficacy by inhibiting T-cell immunoglobulin and mucin-domain containing-3 (TIM-3) expression. In summary, miR-545 holds immense potential as a cancer biomarker and therapeutic target. The aberrant expression and regulatory mechanisms of miR-545 in cancer warrant further investigation.
Article type: Review Article
Keywords: miR-545, Competing endogenous RNA, Cell behavior, Biomarker, Therapeutic drug, Immunotherapy
License: © 2023 The Author(s) CC BY 4.0 This is an open access article under the CC BY license (http://creativecommons.org/licenses/by/4.0/).
Article links: DOI: 10.1016/j.cpt.2023.09.001 | PubMed: 39027151 | PMC: PMC11252520
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (3.5 MB)
Introduction
Micro ribonucleic acids (miRNAs) are endogenous single-stranded non-coding RNAs that are approximately 22 nucleotides long and are highly conserved in evolution.ref. bib1 They weaken or eliminate the function of downstream protein-coding genes (PCGs) by binding to the 3′-untranslated regions (3′-UTR) of target gene mRNA and play a role in post-transcriptional regulation.ref. bib2 The competing endogenous RNA (ceRNA) regulatory network connects long non-coding RNAs (lncRNAs), circular RNAs (circRNAs), miRNAs, and PCGs. miRNAs regulate cancer progression by participating in the ceRNA regulatory axis and affecting downstream PCG expression and signaling pathway activation.ref. bib3
miR-545 is located at Xq13.2, within intron 1 of the lncRNA FTX (FTX Transcript, XIST Regulator), and it has been associated with cancers in multiple systems, including the nervous,ref. bib4,ref. bib5 respiratory,ref. bib6,ref. bib7 digestiveref. bib8 and motorref. bib9 systems. miR-545 is intricately involved in an extensive ceRNA regulatory network, and it is competitively repressed by 15 circRNAs and 12 lncRNAs. It plays a crucial role in regulating multiple signaling pathways, including the catenin,ref. bib10 phosphoinositide-3-kinase (PI3K)/protein kinase B (AKT),ref. bib8,ref. bib11 cell cycle,ref. 12, ref. 13, ref. 14 and p38 pathways.ref. bib15 Via its participation in these pathways, miR-545 exerts control over critical cellular processes, such as cell cycle progression, proliferation, apoptosis, epithelial-mesenchymal transition (EMT), invasion, and migration.
This study examines miR-545 dysregulation in various human cancers and summarized the genes and pathways involved in its regulation and their roles in vivo and in vitro. Our work also provides a systematic summary of the association between miR-545 and patient prognosis as well as the relationship of miR-545 with various therapeutic approaches. Finally, we discuss the current research progress and limitations of miR-545 to provide directions for future translational medicine research.
miR-545 expression dysregulation in human cancers
Pre-miR-545 is processed into two mature miRNA products: miR-545-3p and miR-545-5p. Both of these miRNAs are expressed in various human cell types. Research has shown that both miR-545-3p and miR-545-5p levels are abnormal in 14 different cancer types Supplementary Table 1. Specifically, miR-545-3p levels were lower than normal in 11 cancers and higher than normal in two cancers. In contrast, miR-545-5p levels were lower than normal in seven types of cancer.
As shown in Supplementary Table 2 and Supplementary Figure 1, based on miRNA data from The Cancer Genome Atlas (TCGA) database, we explored the abnormal expression of miR-545 in over 30 types of cancer and paracancerous tissues and its association with the host gene FTX. Our results showed that pre-miR-545 expression was markedly negatively correlated with FTX in three tumor types: kidney chromophobe (KICH), mesothelioma (MESO), and uterine carcinosarcoma (UCS). miR-545-3p expression was upregulated in uterine corpus endometrial carcinoma (UCEC) and markedly negatively correlated with FTX in three tumor types: MESO, rectum adenocarcinoma (READ), and thymoma (THYM). miR-545-5p expression was upregulated in four tumors, head and neck squamous cell carcinoma, lung adenocarcinoma (LUAD), lung squamous cell carcinoma (LUSC), and UCEC, and substantially negatively correlated with FTX in four tumors: colon adenocarcinoma (COAD), KICH, LUAD, and prostate adenocarcinoma (PRAD) (see Supplementary Material for details).
Conflicting results have been reported regarding the expression patterns of miR-545-3p in colorectal cancer (CRC). Several studies utilizing quantitative reverse transcription polymerase chain reaction (RT-qPCR) have demonstrated that miR-545-3p expression is reduced in both CRC cell lines and tissues compared to that in normal colonic epithelium and non-tumor tissues, respectively.ref. bib11,ref. bib16,ref. bib17 However, another study using microarray profiling found that miR-545-3p expression was higher in the serum exosomes of CRC patients than that in healthy individuals.ref. bib18 The inconsistencies in miR-545-3p expression patterns in CRC may be attributed to differences in cancer tissues or measurement methods used in different studies.
In summary, variations exist in the expression of miR-545 across different contexts, and its discrepant expression in experimental data compared to that in the TCGA database has been debated. This disparity could potentially be attributed to the limited number of normal control samples in the TCGA database, leading to an imbalanced sample size in the cancer group. For instance, TCGA included 218 CESC tumor samples but only two control samples. Additionally, the reduced number of reads matching the gene in certain chips or algorithms may result from the lower accuracy of transcriptome expression results for low-expression genes as well as the short fragments and lower abundance of miR-545. Consequently, this can contribute to false results. Moreover, errors in measurement outcomes can stem from the biological properties of miRNAs. For instance, if miR-545 expression undergoes subtle changes in the early stages of cancer but either remains relatively stable during cancer progression or is subject to passive regulation by other genes within the gene regulatory network, it can lead to abnormal fluctuations in its expression levels. Therefore, further studies with larger sample sizes and diverse sample types are necessary to gain a more comprehensive understanding of the aberrant expression patterns of miR-545-3p and miR-545-5p in cancer.
miR-545 and its host gene FTX
FTX is a well-studied lncRNA that plays a crucial role in X chromosome inactivation. Located upstream of the XIST gene in the X chromosome inactivation center (XIC), FTX can enhance XIST expression and trigger X chromosome inactivation.ref. bib19 Additionally, FTX interacted with DExH-box helicase 9 (DHX9) and endoribonuclease dicer (DICER) to positively regulate both A-to-I RNA editing and miRNA expression.ref. bib11
miR-374b/421 and miR-545/374a clusters are located within the FTX intron.ref. bib20 Both FTX and miR-545-3p are upregulated and exhibit tumor-promoting functions in CRC.ref. bib11 However, FTX suppressed miR-545-5p expression in monocytes from patients with liver cirrhosis.ref. bib21 These findings suggest that regulation of miR-545-3p/5p expression by the lncRNA FTX may be tissue-specific and dependent on the host FTX transcript.
We analyzed TCGA data and found that pre-miR-545 was substantially negatively correlated with FTX in three tumor types (KICH, MESO, and UCS) but was substantially positively correlated with FTX in skin cutaneous melanoma. miR-545-5p expression was substantially negatively correlated with FTX levels in COAD, KICH, LUAD, and PRAD. miR-545-3p was substantially negatively correlated with FTX levels in the MESO, READ, and THYM groups. Additionally, the expression levels of pre-mir-374a, pre-mir-374b, pre-miR-374c, pre-mir-421, and pre-mir-545 in the miR-374b/421 and miR-545/374a clusters were strongly positively correlated Supplementary Material. This suggests that the regulation of the expression of miR-545 and that of its neighboring genes is related and may be tissue-specific.
As the host gene of miR-545, FTX is located on the X chromosome, it is worth paying attention to sex differences in its expression changes. TCGA data showed that sex differences in miR-545 expression were not significant in most cancers, as only four (pheochromocytoma and paraganglioma, thyroid carcinoma [THCA], MESO, and READ) showed significant differences (see Supplementary Material for details). Previous studies have not considered the sex of the cell line source. More in-depth research is needed to verify whether sex-related differences exist in the regulation of miR-545 expression.
miR-545 and its competing endogenous ribonucleic acids (ceRNAs)
CeRNAs competitively bind to miRNA, weakening its inhibitory effect on target mRNA and regulating cellular activities at the post-transcriptional level.ref. bib22 As shown in Supplementary Table 3 and Figure 1, the ceRNA regulatory network of miR-545 involves circRNAs and lncRNAs that play regulatory roles in cell biology.
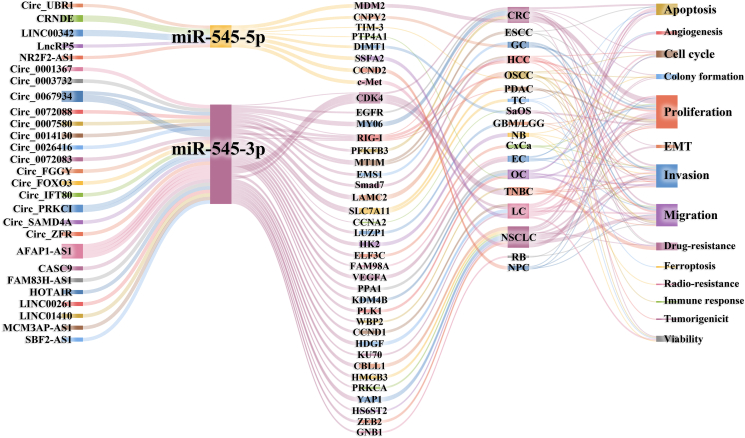
Micro ribonucleic acid-545–related circular ribonucleic acids (CircRNAs)
CircRNAs are non-coding RNA with closed circular structures that can bind to miRNAs to regulate downstream gene expression.ref. bib23 As shown in Supplementary Table 3 and Figure 1, the inhibitory effects of miR-545 on target genes were competitively inhibited by 15 circRNAs: Circ_PRKCI, Circ_0067934, Circ_0072083, Circ_0026416, Circ_0067835, Circ_0003732, Circ_0001367, Circ_FOXO3, Circ_UBR1, Circ_0007580, Circ_0014130, Circ_0072008, Circ_FGGY, Circ_ZFR, and Circ_SAMD4A.
Nineteen circRNA/miR-545-3p axes promote cancer progression, including Circ_PRKCI/miR-545-3p/E2F7 in glioblastoma (GBM)/low-grade glioma (LGG)ref. bib24; Circ_0072083/miR-545-3p/CBLL1 in lung cancer (LC)ref. bib7; Circ_0007580/miR-545-3p/PRKCA,ref. bib15 Circ_FOXO3/miR-545-3p/HMGB3,ref. bib25 Circ_0014130/miR-545-3p/YAP1,ref. bib26 Circ_PRKCI/miR-545-3p/E2F7,ref. bib27 and Circ_PRKCI/miR-545-3p in gastric cancer (GC)ref. bib8; Circ_FGGY/miR-545-3p/SMAD7 in HCCref. bib28; Circ_0026416/miR-545-3p/MYO6 in CRCref. bib16; Circ_SAMD4A/miR-545-3p/PFKFB3ref. bib29 and Circ_0003732/miR-545-3p/CCNA2 in osteosarcoma (SaOS)ref. bib30; Circ_0072008/miR-545-3p/SLC7A11 in pancreatic ductal adenocarcinoma (PDAC)ref. bib60; Circ_IFT80 (Circ_0067835)/miR-545-3p/FAM98A in endometrial cancer (EC)ref. bib31; Circ_0067934/miR-545-3p/PPA1 in ovarian cancer (OC)ref. bib32; Circ_PRKCI/miR-545-3p/WBP2ref. bib33 and Circ_0067934/miR-545-3p/EIF3C in cervical carcinoma (CxCa)ref. bib34; Circ_ZFR/miR-545-3p/WMT5A in bladder cancer; and Circ_0067934/miR-545-3p/SLC7A11 axis in THCA.ref. bib35 Additionally, the Circ_UBR1/miR-545-5p/SSFA2 axis promoted the malignant behavior of cancer cells in LC.ref. bib36 However, the Circ_0001367/miR-545-3p/LUZP1 axis suppressed cancer progression in GBM/LGG.ref. bib4
miR-545–related long non-coding ribonucleic acids (lncRNAs)
LncRNAs are long non-coding RNAs longer than 200 nt.ref. bib37 As shown in Supplementary Table 3 and Figure 1, 12 lncRNAs competitively inhibit miR-545: AFAP1-AS1, CRNDE, HOTAIR, LINC00342, SBF2-AS1, CASC9, LINC00261, MCM3AP-AS1, LINC01410, LncRP5, FAM83H-AS1, and NR2F2-AS1.
Nine lncRNA/miR-545-3p/PCG axes promote cancer progression, including AFAP1-AS1/miR-545-3p/HDGFref. bib6 and FAM83H-AS1/miR-545-3p/HS6ST2ref. bib38 in LC; CASC9/miR-545-3p/LAMC2 in oral squamous cell carcinoma (OSCC)ref. bib39; SBF2-AS1/miR-545-3p/EMS1 in GCref. bib40; HOTAIR/miR-545-3p/EGFRref. bib17 and MCM3AP-AS1/miR-545-3p/CDK4ref. bib12 in CRC; AFAP1-AS1/miR-545-3p/VEGFA in ECref. bib41; AFAP1-AS1/miR-545-3p/CDK4 in triple-negative breast cancer (TNBC),ref. bib13 and AFAP1-AS1/miR-545-3p/GNB1 in retinoblastoma (RB).ref. bib42 Two lncRNA/miR-545-3p/PCG axes can suppress cancer progression, including the LINC01410/miR-545-3p/HK2 axis in neuroblastoma (NB) to suppress tumorigenesis and tumor radioresistanceref. bib5 and the LINC00261/miR-545-3p/MT1M axis to suppress cisplatin resistance in esophageal squamous cell carcinoma (ESCC) cell lines.ref. bib43
Five lncRNA/miR-545-5p/PCG axes promote cancer progression, including CRNDE/miR-545-5p/CCND2 in nasopharyngeal carcinoma (NPC),ref. bib44 NR2F2-AS1/miR-545-5p/MET in LC,ref. bib45 LncRP5/miR-545-5p/PTP4A1 in OC,ref. bib46 CRNDE/miR-545-5p/TIM-3 in OSCC,ref. bib47 and LINC00342/miR-545-5p/MDM2 in CRC.ref. bib48
miR-545 regulates cell behaviors
Because miRNAs and their target gene sequences may not be fully complementary, miRNAs have a wide range of regulatory functions that affect cancer progression by regulating various biological behaviors.ref. bib49 As shown in Supplementary Table 4 and Figure 2, multiple ceRNA/miR-545/PCG axes can regulate cell cycle, proliferation, apoptosis, EMT, invasion, and migration.
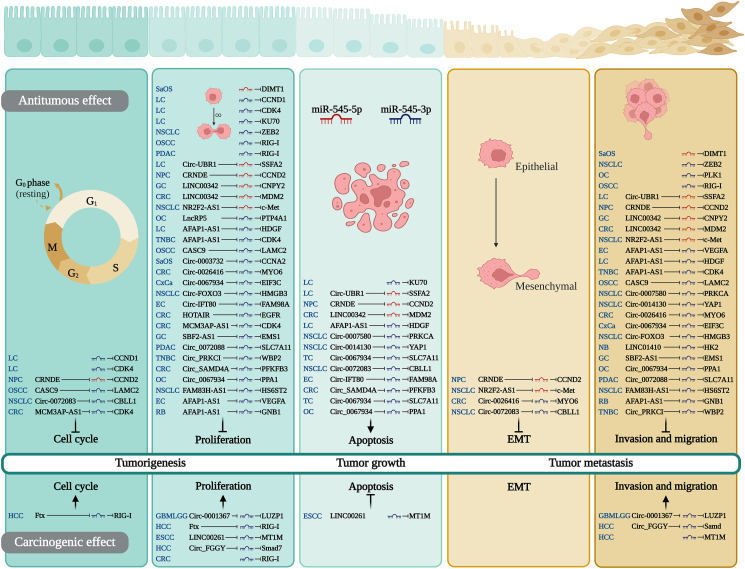
miR-545 regulates the cell cycle
The cell cycle is closely related to cell proliferation and requires appropriate mitotic signals and an appropriate environment for normal cell proliferation.ref. bib50 miR-545 can inhibit cell cycle progression by targeting four target genes (CCND1, CCND2, CDK4, and CBLL1), and it promotes cell cycle progression by targeting RIG-I. miR-545-3p induces G0/G1 phase arrest in cancer cell lines by targeting CCND1 and CDK4 in LC,ref. bib14 CBLL1 in non-small cell lung cancer (NSCLC),ref. bib7 and LAMC2 in OSCC.ref. bib39 In CRC, miR-545-3p arrests the cell cycle at the G1 phase by targeting CDK4.ref. bib12 miR-545-5p targets CCND2ref. bib44 in NPC, causing G0/G1 phase arrest in the cell line.
miR-545 regulates cell proliferation
The degree of cell proliferation is a key indicator of the pathways and mechanisms governing cell survival and death.ref. bib51 Disruptions in the inhibitory pathways associated with proliferation can result in abnormal cell growth, a hallmark of malignancy.ref. bib50 As shown in Supplementary Table 4, miR-545-3p inhibits cell proliferation by targeting 18 PCGs. In GBM/LGG and HCC cells, miR-545-3p promotes cancer cell proliferation by targeting LUZP1ref. bib4 and SMAD7,ref. bib28 respectively. miR-545-3p targets multiple genes and regulates tumor growth in xenograft animals. For example, it promotes tumor growth via four targets in LC (CCND1ref. bib14 and CDK4ref. bib14) and OC (PLK1ref. bib52 and KDM4Bref. bib52) but inhibits tumor growth in HCC via two targets (RIG-Iref. bib8 and MT1Mref. bib53). Additionally, miR-545-5p promotes tumor growth in animal models of NSCLC and SaOS by targeting c-Metref. bib45 and DIMT1,ref. bib9 respectively.
The miR-545-3p/RIG-I axis inhibits the proliferation of OSCC (HSC4)ref. bib54 and PDAC (HEK293, PANC1, and SW1990)ref. bib55 cell lines while promoting the proliferation CRC DLD-1/HCT116ref. bib11 cell lines. In CRC xenografts using LOVO cells in BALB/c nude mice, miR-545-3p promotes tumor growthref. bib17 but inhibits it in HT29/HCT116 cells.ref. bib18 The impact of miR-545-3p on cell proliferation differs between various cancers owing to variations in gene regulatory networks. Further research is required to elucidate the underlying mechanisms.
miR-545 regulates apoptosis
Apoptosis is a type of programmed cell death that can be initiated by intracellular or extracellular signals, which plays a crucial role in antitumor processes.ref. bib56 miR-545-3p and miR-545-5p promote apoptosis by targeting ten and two genes, respectively. Additionally, miR-545-3p inhibits apoptosis by downregulating MT1M expression.
miR-545-3p promotes apoptosis by targeting 10 downstream PCGs, including CCND2 in NPCref. bib57; KU70ref. bib57 and HDGFref. bib6 in LC; CBLL1,ref. bib7 PRKCA,ref. bib15 and YAP1 in NSCLCref. bib26; FAM98A in EC32; SLC7A11 in THCAref. bib35; PFKFB3 in CRCref. bib29; and PPA1 in OC.ref. bib32 miR-545-5p promotes apoptosis by targeting SSFA2 in LCref. bib36 and MDM2 in CRC.ref. bib48
Notably, miR-545-3p overexpression inhibits apoptosis by targeting MT1M in two ESCC cell lines (TE-1 and ECA109).ref. bib43 The effects of miR-545-3p on apoptosis vary between different cancers, as they act as intermediate signal transducers and relay different upstream signals. Further research is required to fully understand these underlying mechanisms.
miR-545 regulates epithelial-mesenchymal transition
EMT is a key process in tumor metastasis that enables tumor cells to invade blood or lymphatic vessels and generate circulating tumor cells.ref. bib58,ref. bib59 miR-545-3p inhibits EMT in NSCLC and CRC cells by targeting CBLL1ref. bib7 and MYO6,ref. bib16 which suppress the neural-like transformation of E-cadherin and vimentin expression. MiR-545-5p inhibits EMT in NPC cells by targeting CCND2, which prevents the reduction of E-cadherin and the replacement of the cytokeratin cytoskeleton with vimentin.ref. bib44 Additionally, miR-545-5p downregulates biliverdin reductase by targeting c-Met, thereby inhibiting EMT in NSCLC cells.ref. bib45
miR-545 regulates cell invasion and migration
Invasion and migration are critical steps in tumor metastasis and major contributors to cancer mortality. Studies have shown that miR-545-3p inhibits cancer cell invasion and migration by targeting 21 genes, including HK2, HDGF, PRKCA, HMGB3, YAP1, ZEB2, HS6ST2, EMS1, MYO6, VEGFA, PLK1, KDM4B, PPA1, CDK4, EIF3C, SLC7A11, WBP2, LAMC2, RIG-I, and GNB1. Conversely, miR-545-3p promotes cancer cell invasion and migration by targeting three genes, LUZP1, SMAD7, and MT1M, and miR-545-5p promotes them by targeting six genes, including CCND2, SSFA2, MET, CNPY2, MDM2, and DIMT1. miR-545-3p inhibits cancer cell invasion and migration by targeting 20 genes including HK2 in NBref. bib5; HDGF in LCref. bib6; PRKCA,ref. bib15 HMGB,ref. bib25 YAP1,ref. bib26 ZEB2,ref. bib10 and HS6ST2ref. bib38 in NSCLC; EMS1ref. bib40 in GC; MYO6ref. bib16 in CRC; VEGFA in ECref. bib41; PLK1,ref. bib52 KDM4B,ref. bib52 and PPA1ref. bib32 in OC; CDK4 in TNBCref. bib13; EIF3C in CxCaref. bib34; SLC7A11 in THCAref. bib35 and PDAC61; WBP2 in TNBCref. bib33; LAMC2 and RIG-I in OSCCref. bib39,ref. bib54; and GNB1 in RB.ref. bib42 miR-545-3p can inhibit tumor metastasis in nude mice by targeting KDM4B and PLK1 in BALB/c nude mice transplanted with OC cells.ref. bib52
In contrast, research has shown that miR-545-3p can promote cancer cell invasion and migration by targeting three genes: LUZP1 in GBM/LGG,ref. bib4 SMAD7 in HCC,ref. bib28 and MT1M in HCC.ref. bib53 Additionally, miR-545-5p promotes cancer cell invasion and migration by targeting six genes: CCND2 in NPC,ref. bib44 SSFA2 in LC,ref. bib36 c-MET in NSCLC,ref. bib45 CNPY2 in GC,ref. bib61 MDM2 in CRC,ref. bib48 and DIMT1 in SaOS.ref. bib9
miR-545 regulates ferroptosis
Ferroptosis is a type of programmed cell death that differs from apoptosis. It occurs because of iron-dependent lipid peroxidation, and it is independent of caspase and RIPK1 activities. Ferroptosis plays a significant role in tumor suppression.ref. bib62
The xc− transport system is an amino acid antiporter that facilitates GSH synthesis and maintains redox homeostasis.ref. bib63 SLC7A11 is a membrane-localized light-chain subunit that forms an xc− transport system with the heavy-chain subunit SLC3A2.ref. bib64 SLC7A11 overexpression can lead to tumorigenesis and ferroptosis resistance.ref. bib65
In THCA, miR-545-3p targets SLC7A11 to promote ferroptosis in FTC133 and TPC-1 cells. In CRC, miR-545-3p targets transferrin (TF) to render HT-29 or HCT-116 cell-based xenograft mouse models resistant to the ferroptosis inducer erastin. This inhibits abnormal ferroptosis signaling and promotes cancer progression.ref. bib18
miR-545 is involved in cancer signaling
As shown in Figure 3, miR-545-3p plays a role in several cancer-related signaling pathways. These include the protein wnt (Wnt)/β-catenin,ref. bib10 PI3K/AKTref. bib11 cell cycle,ref. 12, ref. 13, ref. 14 and p38 pathways.ref. bib15 Additionally, miR-545-5p participates in the p53 pathway by targeting MDM2.ref. bib48
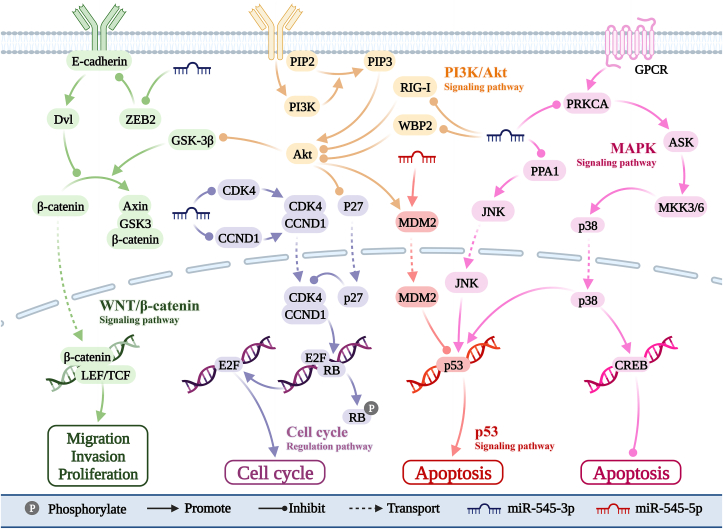
Wnt/β-catenin signaling pathway
The Wnt/β-catenin pathway is crucial for stem cells.ref. bib66 Abnormal activation of this pathway promotes tumor stem cell renewal, proliferation, and differentiation.ref. bib67 ZEB2 is a transcription factor that functions alone or in combination with other proteins.ref. bib68 In NSCLC, miR-545-3p inhibits ZEB2 expression and reduces free β-catenin protein levels in the cytoplasm. This prevents β-catenin from entering the nucleus and blocks the Wnt/β-catenin pathway. Consequently, cancer cell proliferation, invasion, and metastasis are inhibited.ref. bib10,ref. bib69
PI3K/AKT signaling pathway
The PI3K/AKT pathway regulates cell phenotypes, such as proliferation, invasion, autophagy, and senescence. It is often abnormally activated in human cancers.ref. bib70,ref. bib71 RIG-I is an innate immune sensor that acts as a tumor suppressor.ref. bib72 miR-545-3p targets and inhibits RIG-I expression in CRC and HCC. This activates the PI3K/AKT pathway and promotes cell proliferation.ref. bib11 However, miR-545-3p inhibits cancer progression by targeting RIG-I in PDAC and OSCC.ref. bib54,ref. bib55 WBP2 is an oncoprotein that promotes AKT phosphorylation.ref. bib73 miR-545-3p targets and inhibits WBP2 expression in TNBC. This inhibits PI3K/AKT pathway activation and slows cancer progression.ref. bib33
Mitogen-activated protein kinase signaling pathway
The mitogen-activated protein kinase (MAPK) pathway is important for cancer development and is frequently activated in various cancers.ref. bib74 JNKref. bib75 and p38ref. bib76 signals are key components of this pathway that affect cell proliferation, differentiation, and migration. PRKCA encodes PKCα, which promotes p38 phosphorylation and activates the p38 pathway via MKK3/6.ref. bib77,ref. bib78 In NSCLC, miR-545-3p inhibits p38 pathway activation by targeting PRKCA.ref. bib15 It also inhibits cancer cell invasion and promotes apoptosis. PPA1 encodes an inorganic pyrophosphatase dysregulated in several cancers.ref. bib79 PPA1 activates the JNK pathway and promotes NSCLC progression in a TP53-dependent manner.ref. bib80 In OC, miR-545-3p suppresses tumor malignancy by targeting PPA1 to inhibit JNK pathway activation.ref. bib32
Cellular tumor antigen p53 signaling pathway
TP53 is a crucial tumor suppressor gene that encodes p53. This protein is deleted or mutated in more than half of all cancers.ref. bib81,ref. bib82 p53 regulates cell cycle arrest and apoptosis.ref. bib83,ref. bib84 MDM2 is an E3 ubiquitin ligase and a major antagonist of p53.ref. bib85 It cooperates with MDMX to ubiquitinate p53, leading to its degradation and nuclear export. This inhibits p53 transcriptional activity.ref. bib86 In CRC, miR-545-5p inhibits MDM2 expression, increases p53 transcriptional activity, and inhibits tumor growth.ref. bib48
Cell cycle regulation pathway
The cell cycle in cancer metabolism is complex, and its dysregulation is a key feature of cancer progression.ref. bib87 The cell cycle pathway regulates cell proliferation and affects cancer progression. This is an important target in cancer therapy.ref. bib88 Various cyclin/CDK complexes play crucial roles in regulating the cell cycle pathway.ref. bib89,ref. bib90 The CCND1/CDK4 complex phosphorylates RB1 and promotes E2F transcriptional activity. This promotes the G1/S transition of cells.ref. bib91,ref. bib92 The cell cycle is regulated by upstream pathways, such as PI3K/AKTref. bib93 and p53 signaling.ref. bib94 In CRC and TNBC, miR-545-3p targets CDK4 to inhibit E2F transcriptional activity and causes G1 arrest in cancer cells.ref. bib12,ref. bib13 In LC, miR-545-3p inhibits cancer cell proliferation by targeting CCND1 and CDK4.ref. bib14
miR-545 has prognostic value in cancer
As shown in Table 1, abnormal expression of miR-545 is substantially associated with the prognosis and clinicopathological features of cancer patients. Thus, it can serve as a biomarker for cancer prognosis. In NSCLC, PDAC, and OC, the low miR-545 expression indicates poor OS, progression-free survival (PFS), and DFS.
Table 1: Prognostic value of miR-545-3p.
| Origin of mature miR-545 | Cancer | Sample size, n | miR-545 expression | Clinicopathological characteristics | Prognostic value | Reference |
|---|---|---|---|---|---|---|
| 3p | NSCLC | 84 | Down-regulation | Associated with lymph node metastasis↑, and tumor stage↑ | Shorter OS | ref. 10 |
| PDAC | 78 | Down-regulation | – | Shorter OS and DFS | ref. 55 | |
| OC | 60 | Down-regulation | Associated with pTNM stage↑ | Shorter OS | ref. 52 | |
| 218 | Down-regulation | Associated with response status↓ | Shorter OS and PFS | ref. 95 |
DFS: Disease-free survival; HCC: Hepatocellular carcinoma; NSCLC: Non-small-cell lung cancer; OC: Ovarian cancer; OS: Overall survival; PDAC: Pancreatic ductal adenocarcinoma; PFS: Progression-free survival; pTNM: Pathological tumor node metastasis; TNM: Tumor node metastasis.
In NSCLC, low miR-545-3p expression is associated with more severe lymph node metastasis and a higher tumor stage. This predicts a shorter OS.ref. bib10 In PDAC, patients with low miR-545-3p expression have poor OS and DFS.ref. bib55 In OC, low miR-545-3p expression is associated with poor OS and PFS. It is also strongly associated with a higher pTNM stage and stronger platinum drug resistance.ref. bib52,ref. bib95
Current research on the clinical value of miR-545 is limited, and it has mostly focused on miR-545-3p. Further studies with larger sample sizes are required to understand the association between miR-545 expression and the prognosis of cancer patients. The potential differences between miR-545-3p and miR-545-5p expression in association with patient prognosis need to be determined.
miR-545 and cancer therapy
miRNAs can influence cancer therapy by participating in extensive regulatory networks.ref. bib96 miR-545 mediates the effects of chemical factors, such as formononetin,ref. bib13 and physical factors, such as ionizing radiation,ref. bib97 on cancer cells. It can target downstream factors, such as YAP1, HK2, PFKFB3, KU70, and C–C motif chemokine ligand 22 (CCL22), to inhibit cell resistance to chemotherapy and radiotherapy. MiR-545 can also promote resistance to chemotherapy and radiotherapy via MT1M. Additionally, miR-545 enhances immune cell proliferation and infiltration by inhibiting downstream factors, such as TIM-3 and CCL22.
miR-545 and chemotherapy
Chemotherapy is a supplementary treatment for advanced cancers that prevents tumor cell growth.ref. bib98 Figure 4A shows that formononetin indirectly increases miR-545 levels and improves cell sensitivity to four chemotherapeutic drugs (adriamycin, docetaxel DTX, paclitaxel, and cisplatin DDP). By targeting YAP1, miR-545 increases cell sensitivity to DTX. Targeting HT1M and PPA1 increases cell resistance and sensitivity to DDP. Targeting PFKFB3 increases cell sensitivity to 5-fluorouracil (5-FU).
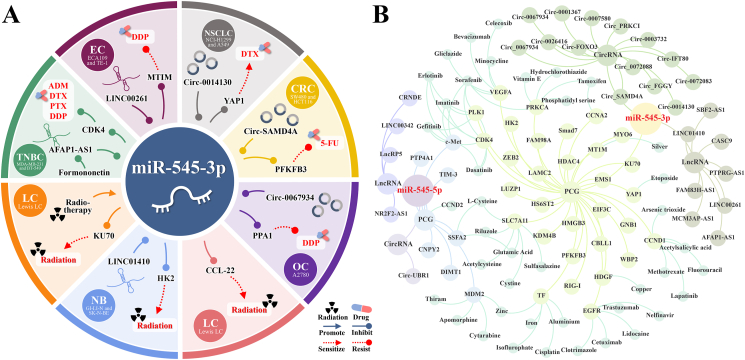
Formononetin is a 7-hydroxyisoflavone with a methoxy group that can be found in red clovers and the Chinese herb Astragalus membraneceus.ref. bib99 It can regulate cell death and cell cycle processes, and it shows promise in preventing and treating cancer.ref. bib100 Formononetin inhibits the growth, invasion, and movement of two TNBC cell lines (MDA-MB-231 and BT-549) by decreasing AFAP1-AS1 expression, increasing miR-545-3p levels, and improving sensitivity to the four drugs mentioned above.ref. bib13
DTX is a chemotherapeutic drug in the taxane family commonly used to treat NSCLC patients.ref. bib101 In NSCLC, DTX-resistant NCI–H1299 and A549 cell lines exhibit lower miR-545-3p expression levels than non-resistant NCI–H1299 and A549 cell lines.ref. bib26 High miR-545-3p expression in the circ _0014130/miR-545-3p/YAP1 axis reduces DTX resistance in two NSCLC cell lines (NCI–H1299 and A549).ref. bib26
DDP is a stable platinum coordination compound at normal temperature and pressure that can be used as a standard treatment for ESCC.ref. bib102 In ESCC, miR-545-3p is a part of the LINC00261/miR-545-3p/MT1M axis that increases DDP resistance in two ESCC cell lines (TE-1 and ESCC109).ref. bib43 In OC, miR-545-3p involvement in the Circ_0067934/miR-545/PPA1 pathway reduces DDP resistance in the A2780/DDP cell line.ref. bib32
In addition, 5-FU, a naturally occurring uracil analog,ref. bib103 is often used as a nanoagent to treat different types of cancers, including CRC and breast cancer.ref. bib104 Highly expressed miR-545-3p participated in the Circ_SAMD4A/miR-545/PFKFB3 axis in two CRC cell lines (SW480/5-FU and HCT-116/5-FU) and inhibited 5-FU resistance.ref. bib29
miR-545 and radiotherapy
Radiotherapy uses ionizing radiation to treat cancer by targeting cells and molecules.ref. bib105 It increases miR-545 expression, which inhibits KU70, HK2, and CCL22 expression and enhances tumor radiosensitivity Figure 4A. In Lewis lung carcinoma cells, radiotherapy increases miR-545-3p expression in irradiated areas and inhibits tumor progression by reducing CCL22 expression.ref. bib106 High miR-545-3p levels also enhance radiosensitivity in C57BL/6 mice by reducing KU70 expression.ref. bib57 In the NB cell lines GI-LI-N and SK-N-BE(2), high levels of miR-545-3p increase radioresistance by inhibiting glycolysis via the LINC01410/miR-545-3p/HK2 axis.ref. bib5
miR-545 and immunotherapy
Cancer cells can evade or suppress the immune system by altering their antigensref. bib107 and modifying the tumor microenvironment.ref. bib108,ref. bib109 Immunotherapy is a novel cancer treatment that induces antitumor immune responses.ref. bib110 As shown in Figure 5, miR-545 enhances antitumor immunity by reducing TIM-3 and CCL22 levels.
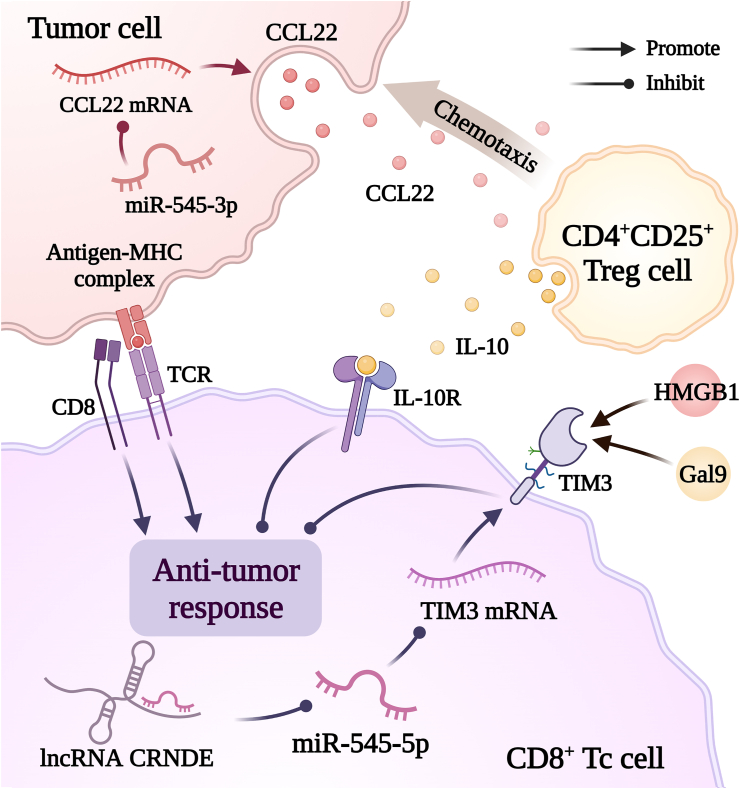
TIM-3 is an immune checkpoint that negatively regulates immunity in various cancers (CRC, CxCa, and GC) and leukemia stem cells.ref. bib111 Co-expression of PD-1 reduces the effectiveness of tumor immunotherapy and promotes tumor growth.ref. bib112,ref. bib113 In OSCC, miR-545-5p reduces TIM-3 expression to enhance immunity and inhibit tumor growth.ref. bib47
A large number of regulatory T cells can suppress immunity and worsen cancer prognosis.ref. bib114 CCL22 increases regulatory T cell (Treg) infiltration in tumors and promotes tumor growth.ref. bib115 In Lewis lung carcinoma, miR-545-3p recruits CD4+CD25+ regulatory T cells via CCL22 to inhibit tumor growth.ref. bib106
miR-545–related therapeutic drugs
As shown in Figure 4B, a search of the CADDIE database (https://exbio.wzw.tum.de/caddie/)ref. bib116 revealed that several approved drugs target miR-545 downstream of PCGs. These include apomorphine, cytarabine, zinc, and thiram targeting MDM2, as well as bevacizumab, sorafenib, celecoxib, minocycline, and gliclazide targeting VEGFA. Further research is needed to understand the interactions between miR-545 and these PCG-targeted drugs to develop new combination therapies for clinical use.
Discussion
MiR-545, located on chromosome Xq13.2, has two mature forms: miR-545-3p and miR-545-5p. Extensive evidence has demonstrated the dysregulation of miR-545 expression in various cancers, making it a promising cancer biomarker. The distinct expression patterns of miR-545 in different tumor types may be attributed to its participation in diverse regulatory networks and varying roles in different cancer contexts. However, comprehensive studies outlining the regulatory landscape of miR-545 in different cancers are lacking. Thus, it is imperative to map the regulatory mechanisms of miR-545 in diverse tumor types in future studies. Although dysregulated miR-545 expression in most tumors has been extensively explored, its differential expression in CRC remains controversial. This discrepancy may stem from factors such as cellular and tissue sample heterogeneity or differences in detection methods. Furthermore, findings from pan-cancer analyses using large databases, such as TCGA, do not align with experimental data consistently. This discrepancy could be attributed to factors such as the data volume within the database, the differential analysis algorithms employed, or the inherent heterogeneity of the experimental samples. To address these challenges, future research endeavors could increase the sample size and diversity and reduce error rates by integrating data from additional databases, such as the Gene Expression Omnibus (GEO), for joint analysis. Moreover, the expression level of miR-545 could be precisely measured using advanced chips or sequencing technologies. Concurrently, expanding the repertoire of verified cell lines and increasing their numbers would contribute to obtaining more robust and convincing results. Notably, there are discrepancies in the expression patterns of miR-545-3p and miR-545-5p, possibly stemming from insufficient data on certain tumor types in existing studies. Hence, future investigations should focus on elucidating the differential expression of miR-545 in diverse tumor subtypes, stages, clinical features, and prognoses. Leveraging multi-source data and employing multilevel analysis methods would enable a comprehensive assessment of the potential diagnostic value of miR-545 in various tumors.
MiR-545 is located within intron 1 of the lncRNA FTX. Analysis of the TCGA database revealed a significant correlation between miR-545 and FTX, as well as other miRNA clusters in certain tumors. However, this correlation was not significant in most TCGA tumors, and FTX CpG island methylation levels did not substantially influence miR-545 expression. This discrepancy may be attributed to missing data in the database. To elucidate the role and mechanisms of the lncRNA FTX and miR-545, along with their adjacent genes, in tumorigenesis, future research could enhance data availability by integrating databases or incorporating self-owned samples. Additionally, comprehensive analyses integrating genomics, transcriptomics, proteomics, metabolomics, and other data can be employed to investigate the multifaceted functions of miR-545 at various levels. Network analysis, systems biology, machine learning, and other methodologies can be applied to construct regulatory networks and functional modules associated with the lncRNA FTX and miR-545. Subsequently, a series of molecular biology experiments should be conducted to further investigate the mechanisms of their interaction. In recent years, sex disparities in tumorigenesis and disease progression have attracted increasing attention. Studies have indicated that fat accumulation or distribution exhibits varying effects on colorectal, esophageal, and liver cancers depending on sex.ref. bib117 Therefore, it is crucial to consider the unique chromosomal locations of FTX and miR-545, as well as their potential sex-specific implications in tumor diagnosis and treatment. Nevertheless, TCGA results only revealed sex differences in miR-545 expression in a few tumor types, and the current study did not focus on cell line selection based on sex. Future investigations should collect gene expression profiles from patient tissues of different sexes and incorporate clinical data from male and female cancer patients for comprehensive statistical analysis. Furthermore, the expression levels and biological functions of miR-545 were validated using in vitro-cultured cancer cell lines and animal models encompassing both males and females.
miR-545 is competitively inhibited by 15 circRNAs and 12 lncRNAs, actively participating in five tumor-associated pathways that regulate key tumor cell behaviors affecting tumor progression, including cell cycle, proliferation, apoptosis, EMT, invasion, and migration. Although the initial discovery of the miR-545 regulatory network in tumors is promising, most mechanisms identified thus far involve single ceRNA axes, highlighting the need for more systematic and comprehensive network-based studies. Additionally, given the intricate mechanism of action of miRNAs, research on miR-545 should not solely focus on the inhibition of protein translation at the post-transcriptional level but also explore the mechanisms underlying the induction of target RNA degradation. To address these challenges, future studies should integrate bioinformatic analysis with in vivo and in vitro experiments to explore the regulatory network involving miR-545 and its associated molecules more comprehensively. This will help to elucidate their potential as therapeutic targets, thereby establishing a foundation for the development of more precise and targeted therapeutic strategies.
The extensive differential expression patterns and regulatory mechanisms of miR-545 highlight its potential clinical applications. This review revealed a significant correlation between abnormal miR-545 expression and the prognostic and clinicopathological features of various tumors. However, considering the ambiguity surrounding miR-545 expression, its reliability as a stable biomarker warrants further investigation. Future studies should collect extensive clinical data, conduct cohort analyses, and perform validation studies to assess the potential of miR-545 as a reliable biomarker, either alone or in combination with other genes. Moreover, although miR-545 has been implicated in interactions with multiple tumor treatments, such as chemotherapy, radiotherapy, and immunotherapy, via at least 10 downstream factors, research on its interactions with common treatments remains limited. For instance, the role of miR-545 in cancer immunity, including its relationship with immune checkpoints and immune cell functions, requires further exploration. Future studies should expand their scope to include a variety of treatments and drugs and monitor changes in miR-545 expression pre- and post-treatment to gain a better understanding of its involvement in chemotherapy and radiotherapy resistance. RNA therapy offers advantages, such as high therapeutic efficacy, low drug toxicity, strong specificity, and broad applicability.ref. bib118 Given the involvement of miRNAs in diverse regulatory networks and their close association with normal physiological processes and cancer development, therapeutic strategies targeting miRNAs hold significant promise, and several miRNA-based therapeutic approaches already exist. For instance, rMMN nanomedicines loaded with miR-30a-5p upregulated miR-30a-5p levels both in vitro and in vivo, thereby suppressing the malignant phenotype of ocular melanoma by targeting the transcription factor E2F7.ref. bib119 Numerous therapeutic approaches targeting miRNAs and miRNA mimics have been used in clinical research.ref. bib120 Future research should elucidate the regulatory modes of miR-545, develop appropriate drug delivery vehicles and methods, and ultimately achieve the clinical translation of miR-545 with high efficiency and low side effects.
MiR-545 exhibits differential expression patterns across various tumors and plays a pivotal role in intricate regulatory networks that influence tumor initiation and progression. In the future, we anticipate increased research emphasis on miR-545 and its target interactions, aiming to unravel its molecular mechanisms, therapeutic potential, and diagnostic value. Such investigations will provide valuable insights into the clinical applications of miR-545 in cancer and establish a robust theoretical foundation for subsequent advancements in clinical diagnosis and treatment. Given its potential as a promising diagnostic and therapeutic molecule, miR-545 warrants further comprehensive investigation and in-depth studies.
Conclusions
This review systematically examines miR-545 and its dysregulated expression in cancer cells. It summarizes the regulatory role and molecular mechanisms of miR-545 in cancer and its relationship with patient prognosis and drug treatments. This review highlights the potential of miR-545 as a biomarker and therapeutic target for human cancers and identifies gaps in the current research to guide future studies. MiR-545 is an important regulatory molecule with potential practical applications in clinical cancer treatment.
Funding
This study was supported by the Qiantang Scholars Fund of the Hangzhou City University (No. 210000-581835).
Authors contribution
Jinze Shen: Conceptualization, Writing – Original Draft, Visualization; Xinming Su: Writing – Original Draft; Qurui Wang: Visualization; Yufei Ke: Visualization; Tianyu Zheng: Visualization; Yunan Mao: Visualization; Zehua Wang: Visualization; Jingyin Dong: Writing – Review & Editing; Shiwei Duan: Conceptualization, Writing – Review & Editing, Funding acquisition.
Ethics statement
None.
Data availability statement
All data generated or analyzed during this study are included in this published article and its supplementary information files.
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
References
- M. Bhaskaran, M. Mohan. MicroRNAs: history, biogenesis, and their evolving role in animal development and disease. Vet Pathol, 2014. [DOI | PubMed]
- K. Felekkis, E. Touvana, C. Stefanou, C. Deltas. MicroRNAs: a newly described class of encoded molecules that play a role in health and disease. Hippokratia, 2010. [PubMed]
- H. Li, Y. Li, D. Tian, J. Zhang, S. Duan. MiR-940 is a new biomarker with tumor diagnostic and prognostic value. Mol Ther Nucleic Acids, 2021. [DOI | PubMed]
- X. Dong, P. Zhang, L. Liu. The Circ_0001367/miR-545-3p/LUZP1 axis regulates cell proliferation, migration and invasion in glioma cells. Front Oncol, 2021. [DOI]
- L. Mou, L. Wang, S. Zhang, Q. Wang. Long noncoding RNA LINC01410 suppresses tumorigenesis and enhances radiosensitivity in neuroblastoma cells through regulating miR-545-3p/HK2 axis. OncoTargets Ther, 2021. [DOI]
- J. Sun, H. Min, L. Yu, G. Yu, Y. Shi, J. Sun. The knockdown of LncRNA AFAP1-AS1 suppressed cell proliferation, migration, and invasion, and promoted apoptosis by regulating miR-545-3p/hepatoma-derived growth factor axis in lung cancer. Anti Cancer Drugs, 2021. [DOI | PubMed]
- H. Li, F. Liu, W. Qin. Circ_0072083 interference enhances growth-inhibiting effects of cisplatin in non-small-cell lung cancer cells via miR-545-3p/CBLL1 axis. Cancer Cell Int, 2020. [DOI | PubMed]
- L. Wu, Y. Li, X.M. Xu, X. Zhu. Circular RNA circ-PRKCI promotes cell proliferation and invasion by binding to microRNA-545 in gastric cancer. Eur Rev Med Pharmacol Sci, 2019. [DOI | PubMed]
- H.Z. Zhou, B. Chen, X.J. Li. MicroRNA-545-5p regulates apoptosis, migration and invasion of osteosarcoma by targeting dimethyladenosine transferase 1. Oncol Lett, 2021. [DOI | PubMed]
- J. Cui, G. Pan, Q. He, L. Yin, R. Guo, H. Bi. MicroRNA-545 targets ZEB2 to inhibit the development of non-small cell lung cancer by inactivating Wnt/β-catenin pathway. Oncol Lett, 2019. [DOI | PubMed]
- Z.H. Kwok, B. Zhang, X.H. Chew. Systematic analysis of intronic miRNAs reveals cooperativity within the multicomponent FTX locus to promote colon cancer development. Cancer Res, 2021. [DOI | PubMed]
- X. Ma, J. Luo, Y. Zhang, D. Sun, Y. Lin. LncRNA MCM3AP-AS1 upregulates CDK4 by sponging miR-545 to suppress G1 arrest in colorectal cancer. Cancer Manag Res, 2020. [DOI | PubMed]
- J. Wu, W. Xu, L. Ma. Formononetin relieves the facilitating effect of lncRNA AFAP1-AS1-miR-195/miR-545 axis on progression and chemo-resistance of triple-negative breast cancer. Aging (Albany NY), 2021. [DOI | PubMed]
- B. Du, Z. Wang, X. Zhang. MicroRNA-545 suppresses cell proliferation by targeting cyclin D1 and CDK4 in lung cancer cells. PLoS One, 2014. [DOI]
- S. Chen, S. Lu, Y. Yao. Downregulation of hsa_circ_0007580 inhibits non-small cell lung cancer tumorigenesis by reducing miR-545-3p sponging. Aging (Albany NY), 2020. [DOI | PubMed]
- L. Zhang, R. Yu, C. Li, Y. Dang, X. Yi, L. Wang. Circ_0026416 downregulation blocks the development of colorectal cancer through depleting MYO6 expression by enriching miR-545-3p. World J Surg Oncol, 2021. [DOI | PubMed]
- X. Huang, S. Lu. MicroR-545 mediates colorectal cancer cells proliferation through up-regulating epidermal growth factor receptor expression in HOTAIR long non-coding RNA dependent. Mol Cell Biochem, 2017. [DOI | PubMed]
- S. Zheng, L. Hu, Q. Song. MiR-545 promotes colorectal cancer by inhibiting transferring in the non-normal ferroptosis signaling. Aging (Albany NY), 2021. [DOI | PubMed]
- C. Chureau, S. Chantalat, A. Romito. FTX is a non-coding RNA which affects Xist expression and chromatin structure within the X-inactivation center region. Hum Mol Genet, 2011. [DOI | PubMed]
- Q. Zhao, T. Li, J. Qi, J. Liu, C. Qin. The miR-545/374a cluster encoded in the FTX lncRNA is overexpressed in HBV-related hepatocellular carcinoma and promotes tumorigenesis and tumor progression. PLoS One, 2014. [DOI]
- X. Liu, C. Li, J. Zhu, W. Li, Q. Zhu. Dysregulation of FTX/miR-545 signaling pathway downregulates Tim-3 and is responsible for the abnormal activation of macrophage in cirrhosis. J Cell Biochem, 2019. [DOI | PubMed]
- X. Qi, D.H. Zhang, N. Wu, J.H. Xiao, X. Wang, W. Ma. ceRNA in cancer: possible functions and clinical implications. J Med Genet, 2015. [DOI | PubMed]
- A.C. Panda. Circular RNAs act as miRNA sponges. Adv Exp Med Biol, 2018. [DOI | PubMed]
- X. Zhang, H. Yang, L. Zhao, G. Li, Y. Duan. Circular RNA PRKCI promotes glioma cell progression by inhibiting microRNA-545. Cell Death Dis, 2019. [DOI | PubMed]
- M. Yang, E. Zheng, J. Ni, X. Xu, X. Jiang, G. Zhao. Circular RNA circFOXO3 facilitate non-small cell lung cancer progression through upregulating HMGB3 via sponging miR-545-3p/miR-506-3p. Tissue Cell, 2022. [DOI]
- D. Du, X. Cao, X. Duan, X. Zhang. Blocking circ_0014130 suppressed drug resistance and malignant behaviors of docetaxel resistance-acquired NSCLC cells via regulating miR-545-3p-YAP1 axis. Cytotechnology, 2021. [DOI | PubMed]
- M. Qiu, W. Xia, R. Chen. The circular RNA circPRKCI promotes tumor growth in lung adenocarcinoma. Cancer Res, 2018. [DOI | PubMed]
- K.L. Feng, N. Diao, Z.W. Zhou. CircFGGY inhibits cell growth, invasion and epithelial-mesenchymal transition of hepatocellular carcinoma via regulating the miR-545-3p/Smad7 axis. Front Cell Dev Biol, 2022. [DOI]
- Y. Gao, C. Liu, X. Xu, Y. Wang, Y. Jiang. Circular RNA sterile alpha motif domain containing 4A contributes to cell 5-fluorouracil resistance in colorectal cancer by regulating the miR-545-3p/6-phosphofructo-2-kinase/fructose-2,6-bisphosphataseisotype 3 axis. Anti Cancer Drugs, 2022. [DOI | PubMed]
- L. Li, X.A. Kong, M. Zang. Hsa_circ_0003732 promotes osteosarcoma cells proliferation via miR-545/CCNA2 axis. Biosci Rep, 2020. [DOI]
- N. Wang, Y. Guo, L. Song, T. Tong, X. Fan. Circular RNA intraflagellar transport 80 facilitates endometrial cancer progression through modulating miR-545-3p/FAM98A signaling. J Gynecol Oncol, 2022. [DOI | PubMed]
- Y. Yin, J. Li, J. Rong, B. Zhang, X. Wang, H. Han. Circ_0067934 reduces JNK phosphorylation through a microRNA-545-3p/PPA1 axis to enhance tumorigenesis and cisplatin resistance in ovarian cancer. Immunopharmacol Immunotoxicol, 2022. [DOI | PubMed]
- X. Wang, H. Song, L. Fang, T. Wu. EIF4A3-mediated circPRKCI expression promotes triple-negative breast cancer progression by regulating WBP2 and PI3K/AKT signaling pathway. Cell Death Discov, 2022. [DOI | PubMed]
- C. Hu, Y. Wang, A. Li, J. Zhang, F. Xue, L. Zhu. Overexpressed circ_0067934 acts as an oncogene to facilitate cervical cancer progression via the miR-545/EIF3C axis. J Cell Physiol, 2019. [DOI | PubMed]
- H.H. Wang, J.N. Ma, X.R. Zhan. Circular RNA Circ_0067934 attenuates ferroptosis of thyroid cancer cells by miR-545-3p/SLC7A11 signaling. Front Endocrinol, 2021. [DOI]
- P. Su, F. Mao, J. Zhang, H. Zhang, M. Wang. Circular RNA UBR1 promotes the proliferation, migration, and invasion but represses apoptosis of lung cancer cells via modulating microRNA-545-5p/SSFA2 axis. Bioengineered, 2021. [DOI | PubMed]
- D. Karagkouni, A. Karavangeli, M.D. Paraskevopoulou, A.G. Hatzigeorgiou. Characterizing miRNA-lncRNA interplay. Methods Mol Biol, 2021. [DOI | PubMed]
- Y. Zhang, Y. Yu, X. Cao, P. Chen. Role of lncRNA FAM83H antisense RNA1 (FAM83H-AS1) in the progression of non-small cell lung cancer by regulating the miR-545-3p/heparan sulfate 6-O-sulfotransferase (HS6ST2) axis. Bioengineered, 2022. [DOI | PubMed]
- B. Ning, S. Guo, Y. Mei. Long non-coding RNA CASC9 promotes tumor progression in oral squamous cell carcinoma by regulating microRNA-545-3p/laminin subunit gamma 2. Bioengineered, 2021. [DOI | PubMed]
- M. He, L. Feng, L. Qi, M. Rao, Y. Zhu. Long noncoding RNASBF2-AS1 promotes gastric cancer progression via regulating miR-545/EMS1 axis. BioMed Res Int, 2020. [DOI]
- Y. Zhong, Y. Wang, H. Dang, X. Wu. LncRNA AFAP1-AS1 contributes to the progression of endometrial carcinoma by regulating miR-545-3p/VEGFA pathway. Mol Cell Probes, 2020. [DOI]
- W. Tang, L. Zhang, J. Li, Y. Guan. AFAP1 antisense RNA 1 promotes retinoblastoma progression by sponging microRNA miR-545-3p that targets G protein subunit beta 1. Bioengineered, 2022. [DOI | PubMed]
- L. Wang, X. Wang, P. Yan, Y. Liu, X. Jiang. LINC00261 suppresses cisplatin resistance of esophageal squamous cell carcinoma through miR-545-3p/MT1M axis. Front Cell Dev Biol, 2021. [DOI]
- S. Ge, C. Jiang, M. Li, Z. Cheng, X. Feng. Long non-coding RNA CRNDE exacerbates NPC advancement mediated by the miR-545-5p/CCND2 axis. Cancer Cell Int, 2021. [DOI | PubMed]
- C. Liu, Q.G. Li, Y. Zhou. LncRNA NR2F2-AS1 induces epithelial-mesenchymal transition of non-small cell lung cancer by modulating BVR/ATF-2 pathway via regulating miR-545-5p/c-Met axis. Am J Cancer Res, 2021. [DOI | PubMed]
- P. Sun, A. Bao, X. Hua, J. Cao, Y. Ding. RP5-1148A21.3 (lncRP5) exerts oncogenic function in human ovarian carcinoma. Acta Biochim Biophys Sin, 2022. [DOI | PubMed]
- Y. Ai, S. Wu, H. Gao. Repression of CRNDE enhances the anti-tumour activity of CD8 + T cells against oral squamous cell carcinoma through regulating miR-545-5p and TIM-3. J Cell Mol Med, 2021. [DOI | PubMed]
- Z. Miao, S. Liu, X. Xiao, D. Li. LINC00342 regulates cell proliferation, apoptosis, migration and invasion in colon adenocarcinoma via miR-545-5p/MDM2 axis. Gene, 2020. [DOI]
- Y. Zhang, Z. Wang, R.A. Gemeinhart. Progress in microRNA delivery. J Control Release, 2013. [DOI | PubMed]
- G.I. Evan, K.H. Vousden. Proliferation, cell cycle and apoptosis in cancer. Nature, 2001. [DOI | PubMed]
- A. Adan, Y. Kiraz, Y. Baran. Cell proliferation and cytotoxicity assays. Curr Pharm Biotechnol, 2016. [DOI | PubMed]
- H. Zhang, K. Zhang, Z. Xu. MicroRNA-545 suppresses progression of ovarian cancer through mediating PLK1 expression by a direct binding and an indirect regulation involving KDM4B-mediated demethylation. BMC Cancer, 2021. [DOI | PubMed]
- L. Changjun, H. Feizhou, P. Dezhen, L. Zhao, M. Xianhai. MiR-545-3p/MT1M axis regulates cell proliferation, invasion and migration in hepatocellular carcinoma. Biomed Pharmacother, 2018. [DOI | PubMed]
- G. Yuan, H. Wu, Y. Du, F. He. Tumor suppressor role of microRNA-545 in oral squamous cell carcinoma. Oncol Lett, 2019. [DOI | PubMed]
- B. Song, W. Ji, S. Guo. MiR-545 inhibited pancreatic ductal adenocarcinoma growth by targeting RIG-I. FEBS Lett, 2014. [DOI | PubMed]
- G. Pistritto, D. Trisciuoglio, C. Ceci, A. Garufi, G. D’Orazi. Apoptosis as anticancer mechanism: function and dysfunction of its modulators and targeted therapeutic strategies. Aging (Albany NY), 2016. [DOI | PubMed]
- C. Liao, W. Xiao, N. Zhu. MicroR-545 enhanced radiosensitivity via suppressing Ku70 expression in Lewis lung carcinoma xenograft model. Cancer Cell Int, 2015. [DOI | PubMed]
- G. Babaei, S.G. Aziz, N.Z.Z. Jaghi. EMT, cancer stem cells and autophagy: the three main axes of metastasis. Biomed Pharmacother, 2021. [DOI]
- M. Saitoh. Involvement of partial EMT in cancer progression. J Biochem, 2018. [DOI | PubMed]
- H. Sun, F. Liu, H. Zhang. Circ_0072008, an oncogene in pancreatic ductal adenocarcinoma, contributes to tumour cell malignant progression and glycolysis by regulating miR-545-3p/SLC7A11 axis. Autoimmunity, 2022. [DOI | PubMed]
- R. Liu, X. Yang. LncRNA LINC00342 promotes gastric cancer progression by targeting the miR-545-5p/CNPY2 axis. BMC Cancer, 2021. [DOI | PubMed]
- C.M. Bebber, F. Muller, L. Prieto Clemente, J. Weber, S. von Karstedt. Ferroptosis in cancer cell biology. Cancers, 2020. [DOI | PubMed]
- P. Koppula, Y. Zhang, L. Zhuang, B. Gan. Amino acid transporter SLC7A11/xCT at the crossroads of regulating redox homeostasis and nutrient dependency of cancer. Cancer Commun, 2018. [DOI]
- S. Li, Z. Lu, R. Sun. The role of SLC7A11 in cancer: friend or foe?. Cancers, 2022. [DOI | PubMed]
- P. Koppula, L. Zhuang, B. Gan. Cystine transporter SLC7A11/xCT in cancer: ferroptosis, nutrient dependency, and cancer therapy. Protein Cell, 2021. [DOI | PubMed]
- T. Zhan, N. Rindtorff, M. Boutros. Wnt signaling in cancer. Oncogene, 2017. [DOI | PubMed]
- Y. Zhang, X. Wang. Targeting the Wnt/beta-catenin signaling pathway in cancer. J Hematol Oncol, 2020. [DOI | PubMed]
- S.V. Hegarty, A.M. Sullivan, G.W. O’Keeffe. Zeb2: a multifunctional regulator of nervous system development. Prog Neurobiol, 2015. [DOI | PubMed]
- S. Lin, Y. Zhen, Y. Guan, H. Yi. Roles of Wnt/beta-catenin signaling pathway regulatory long non-coding RNAs in the pathogenesis of non-small cell lung cancer. Cancer Manag Res, 2020. [DOI | PubMed]
- J.A. Fresno Vara, E. Casado, J. de Castro, P. Cejas, C. Belda-Iniesta, M. Gonzalez-Baron. PI3K/Akt signalling pathway and cancer. Cancer Treat Rev, 2004. [DOI | PubMed]
- M. Aoki, T. Fujishita. Oncogenic roles of the PI3K/AKT/mTOR axis. Curr Top Microbiol Immunol, 2017. [DOI | PubMed]
- J. Bragelmann, C. Lorenz, S. Borchmann. MAPK-pathway inhibition mediates inflammatory reprogramming and sensitizes tumors to targeted activation of innate immunity sensor RIG-I. Nat Commun, 2021. [DOI | PubMed]
- H. Tabatabaeian, A. Rao, A. Ramos, T. Chu, M. Sudol, Y.P. Lim. The emerging roles of WBP2 oncogene in human cancers. Oncogene, 2020. [DOI | PubMed]
- A.A. Samatar, P.I. Poulikakos. Targeting RAS-ERK signalling in cancer: promises and challenges. Nat Rev Drug Discov, 2014. [DOI | PubMed]
- E.H. Lee, H.T. Kim, S.Y. Chun. Role of the JNK pathway in bladder cancer. OncoTargets Ther, 2022. [DOI]
- T. Wada, J.M. Penninger. Mitogen-activated protein kinases in apoptosis regulation. Oncogene, 2004. [DOI | PubMed]
- D. Mul, S. Wu, R.A. de Paus. A mosaic de novo duplication of 17q21-25 is associated with GH insensitivity, disturbed in vitro CD28-mediated signaling, and decreased STAT5B, PI3K, and NF-kappaB activation. Eur J Endocrinol, 2012. [DOI | PubMed]
- Y. Cheng, Y. Zhu, W. Xu. PKCalpha in colon cancer cells promotes M1 macrophage polarization via MKK3/6-P38 MAPK pathway. Mol Carcinog, 2018. [DOI | PubMed]
- D. Xu, Y. Miao, X. Gu, J. Wang, G. Yu. Pyrophosphatase 1 expression is associated with future recurrence and overall survival in Chinese patients with intrahepatic cholangiocarcinoma. Oncol Lett, 2018. [DOI | PubMed]
- D. Luo, D. Liu, W. Shi. PPA1 promotes NSCLC progression via a JNK- and TP53-dependent manner. Oncogenesis, 2019. [DOI | PubMed]
- V.J.N. Bykov, S.E. Eriksson, J. Bianchi, K.G. Wiman. Targeting mutant p53 for efficient cancer therapy. Nat Rev Cancer, 2018. [DOI | PubMed]
- Z. Luo, R. Cui, E. Tili, C. Croce. Friend or foe: MicroRNAs in the p53 network. Cancer Lett, 2018. [DOI | PubMed]
- L.J. Hernandez Borrero, W.S. El-Deiry. Tumor suppressor p53: biology, signaling pathways, and therapeutic targeting. Biochim Biophys Acta Rev Cancer, 2021. [DOI]
- B. Hong, A.P. van den Heuvel, V.V. Prabhu, S. Zhang, W.S. El-Deiry. Targeting tumor suppressor p53 for cancer therapy: strategies, challenges and opportunities. Curr Drug Targets, 2014. [DOI | PubMed]
- U.M. Moll, O. Petrenko. The MDM2-p53 interaction. Mol Cancer Res, 2003. [PubMed]
- K. Linke, P.D. Mace, C.A. Smith, D.L. Vaux, J. Silke, C.L. Day. Structure of the MDM2/MDMX RING domain heterodimer reveals dimerization is required for their ubiquitylation in trans. Cell Death Differ, 2008. [DOI | PubMed]
- P. Icard, L. Fournel, Z. Wu, M. Alifano, H. Lincet. Interconnection between metabolism and cell cycle in cancer. Trends Biochem Sci, 2019. [DOI | PubMed]
- M. Ingham, G.K. Schwartz. Cell-cycle therapeutics come of age. J Clin Oncol, 2017. [DOI | PubMed]
- B.R. Topacio, E. Zatulovskiy, S. Cristea. Cyclin D-Cdk4,6 drives cell-cycle progression via the retinoblastoma protein’s C-terminal helix. Mol Cell, 2019. [DOI | PubMed]
- M. Malumbres, M. Barbacid. Cell cycle, CDKs and cancer: a changing paradigm. Nat Rev Cancer, 2009. [DOI | PubMed]
- E.S. Knudsen, S.C. Pruitt, P.A. Hershberger, A.K. Witkiewicz, D.W. Goodrich. Cell cycle and beyond: exploiting new RB1 controlled mechanisms for cancer therapy. Trends Cancer, 2019. [DOI | PubMed]
- P. Indovina, F. Pentimalli, N. Casini, I. Vocca, A. Giordano. RB1 dual role in proliferation and apoptosis: cell fate control and implications for cancer therapy. Oncotarget, 2015. [DOI | PubMed]
- S. Lu, C. Ren, Y. Liu, D.E. Epner. PI3K-Akt signaling is involved in the regulation of p21(WAF/CIP) expression and androgen-independent growth in prostate cancer cells. Int J Oncol, 2006. [DOI | PubMed]
- K. Engeland. Cell cycle regulation: p53-p21-RB signaling. Cell Death Differ, 2022. [DOI | PubMed]
- Y. Gu, M. Zhang, F. Peng. The BRCA1/2-directed miRNA signature predicts a good prognosis in ovarian cancer patients with wild-type BRCA1/2. Oncotarget, 2015. [DOI | PubMed]
- H. Mollaei, R. Safaralizadeh, Z. Rostami. MicroRNA replacement therapy in cancer. J Cell Physiol, 2019. [DOI | PubMed]
- M. Bugden, S. Billing, K.C. Mak, F. Norton, D. Klokov, Y. Wang. Ionizing radiation affects miRNA composition in both young and old mice. Int J Radiat Biol, 2019. [DOI | PubMed]
- E. Perez-Herrero, A. Fernandez-Medarde. Advanced targeted therapies in cancer: drug nanocarriers, the future of chemotherapy. Eur J Pharm Biopharm, 2015. [DOI | PubMed]
- K.C. Tay, L.T. Tan, C.K. Chan. Formononetin: a review of its anticancer potentials and mechanisms. Front Pharmacol, 2019. [DOI | PubMed]
- J. Machado Dutra, P.J.P. Espitia, R. Andrade Batista. Formononetin: biological effects and uses – a review. Food Chem, 2021. [DOI]
- Y. Zou, Y. Sun, B. Guo. Alpha(3)beta(1) integrin-targeting polymersomal docetaxel as an advanced nanotherapeutic for nonsmall cell lung cancer treatment. ACS Appl Mater Interfaces, 2020. [DOI | PubMed]
- O. Frontiers Editorial. Retraction: corrigendum: Tan IIA combined with DDP inhibits esophageal squamous cell carcinoma via down-regulating the NF-kB/COX-2/VEGF pathway. Front Oncol, 2021. [DOI]
- R.B. Diasio, B.E. Harris. Clinical pharmacology of 5-fluorouracil. Clin Pharmacokinet, 1989. [DOI | PubMed]
- C. Sethy, C.N. Kundu. 5-Fluorouracil (5-FU) resistance and the new strategy to enhance the sensitivity against cancer: implication of DNA repair inhibition. Biomed Pharmacother, 2021. [DOI]
- R.J. Griffin, K.M. Prise, S.J. McMahon, X. Zhang, J. Penagaricano, K.T. Butterworth. History and current perspectives on the biological effects of high-dose spatial fractionation and high dose-rate approaches: GRID, Microbeam & FLASH radiotherapy. Br J Radiol, 2020. [DOI]
- C. Liao, W. Xiao, N. Zhu. Radiotherapy suppressed tumor-specific recruitment of regulator T cells via up-regulating microR-545 in Lewis lung carcinoma cells. Int J Clin Exp Pathol, 2015. [PubMed]
- R. Upadhyay, J.A. Boiarsky, G. Pantsulaia. A critical role for Fas-mediated off-target tumor killing in T-cell immunotherapy. Cancer Discov, 2021. [DOI | PubMed]
- T. Frankel, M.P. Lanfranca, W. Zou. The role of tumor microenvironment in cancer immunotherapy. Adv Exp Med Biol, 2017. [DOI | PubMed]
- M. Abbott, Y. Ustoyev. Cancer and the immune system: the history and background of immunotherapy. Semin Oncol Nurs, 2019. [DOI]
- J. van den Bulk, E.M. Verdegaal, N.F. de Miranda. Cancer immunotherapy: broadening the scope of targetable tumours. Open Biol, 2018. [DOI]
- N. Acharya, C. Sabatos-Peyton, A.C. Anderson. Tim-3 finds its place in the cancer immunotherapy landscape. J Immunother Cancer, 2020. [DOI]
- M. Das, C. Zhu, V.K. Kuchroo. Tim-3 and its role in regulating anti-tumor immunity. Immunol Rev, 2017. [DOI | PubMed]
- R. Yang, L. Sun, C.F. Li. Galectin-9 interacts with PD-1 and TIM-3 to regulate T cell death and is a target for cancer immunotherapy. Nat Commun, 2021. [DOI | PubMed]
- A. Tanaka, S. Sakaguchi. Targeting Treg cells in cancer immunotherapy. Eur J Immunol, 2019. [DOI | PubMed]
- Q. Wang, E. Schmoeckel, B.P. Kost. Higher CCL22+ cell infiltration is associated with poor prognosis in cervical cancer patients. Cancers, 2019. [DOI | PubMed]
- M. Hartung, E. Anastasi, Z.M. Mamdouh. Cancer driver drug interaction explorer. Nucleic Acids Res, 2022. [DOI | PubMed]
- M. Rask-Andersen, E. Ivansson, J. Hoglund, W.E. Ek, T. Karlsson, A. Johansson. Adiposity and sex-specific cancer risk. Cancer Cell, 2023. [DOI | PubMed]
- M. Winkle, S.M. El-Daly, M. Fabbri, G.A. Calin. Noncoding RNA therapeutics – challenges and potential solutions. Nat Rev Drug Discov, 2021. [DOI | PubMed]
- Y. Ma, H. Lin, P. Wang. A miRNA-based gene therapy nanodrug synergistically enhances pro-inflammatory antitumor immunity against melanoma. Acta Biomater, 2023. [DOI | PubMed]
- A. Gupta, J.L. Andresen, R.S. Manan, R. Langer. Nucleic acid delivery for therapeutic applications. Adv Drug Deliv Rev, 2021. [DOI]
