Engineering Plant Cell Fates and Functions for Agriculture and Industry
Abstract
Many plant species are grown to enable access to specific organs or tissues, such as seeds, fruits, or stems. In some cases, a value is associated with a molecule that accumulates in a single type of cell. Domestication and subsequent breeding have often increased the yields of these target products by increasing the size, number, and quality of harvested organs and tissues but also via changes to overall plant growth architecture to suit large-scale cultivation. Many of the mutations that underlie these changes have been identified in key regulators of cellular identity and function. As key determinants of yield, these regulators are key targets for synthetic biology approaches to engineer new forms and functions. However, our understanding of many plant developmental programs and cell-type specific functions is still incomplete. In this Perspective, we discuss how advances in cellular genomics together with synthetic biology tools such as biosensors and DNA-recording devices are advancing our understanding of cell-specific programs and cell fates. We then discuss advances and emerging opportunities for cell-type-specific engineering to optimize plant morphology, responses to the environment, and the production of valuable compounds.
Affiliations: †Engineering Biology, Earlham Institute, Norwich Research Park, Norwich, NR4 7UZ United Kingdom; ‡Department of Plant Sciences, University of Cambridge, Downing Street, Cambridge, CB2 3EA United Kingdom
License: © 2024 The Authors. Published by American Chemical Society CC BY 4.0 Permits the broadest form of re-use including for commercial purposes, provided that author attribution and integrity are maintained (https://creativecommons.org/licenses/by/4.0/).
Article links: DOI: 10.1021/acssynbio.4c00047 | PubMed: 38573786 | PMC: PMC11036505
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (2.0 MB)
Introduction
Plant cultivation is considered to be one of the most important cultural transitions in human history. Today, 40% of land mass is dedicated to growing plants for food as well as for products such as wood, textiles, dyes, and medicines.1 For many species, the value is associated with a specific organ, tissue, or cell-type (Figure ). In food production, this is exemplified by seeds harvested from grain crops and roots from Daucus carota subsp. Sativus (carrot), tubers from Solanum tuberosum (potato), stems from Apium graveolens (celery), and the fruit of numerous species, including Solanum lycopersicum (tomato). Industrial products include seed hairs from Gossypium hirsutum (cotton), extraxylary fibers from Linum usitatissimum (flax), and natural products from numerous species that often accumulate in specialized cell types. Although recent efforts have sought to maximize the use of plant biomass, for example by producing biofuels from Zea mays (corn) leaves and chippings for playgrounds and gardens from timber waste, efforts are still primarily focused on maximizing yields of target organs and products.
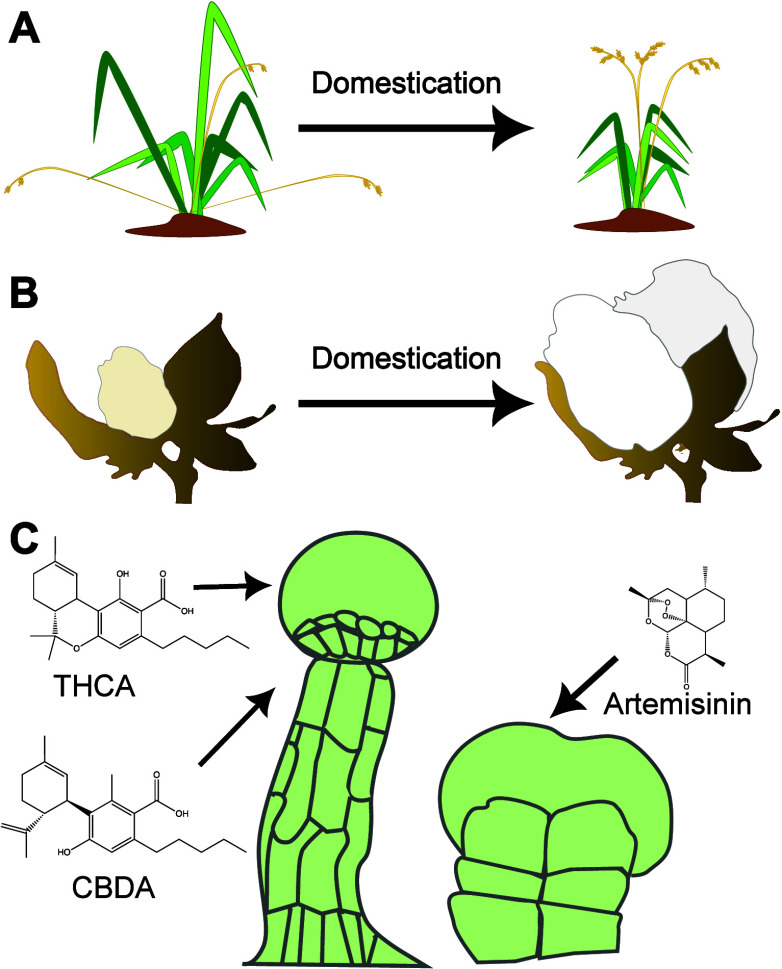
Selective breeding has improved crop yields by increasing the size, number, and quality of valuable harvested organs or tissues and by modifying plant architecture, optimizing it for large-scale cultivation (Figure A). For example, domestication of Oryza sativa (rice) fixed semidwarfism and increased inflorescence branching to prevent lodging and increase grain-yield, respectively.2 Similarly, tomato breeding fixed a more compact morphology with more fruit per stem,3 while cotton breeding selected for longer and more plentiful seed coat hairs to produce larger bolls4 (Figure B). Many of the mutations that underlie these phenotypes have been identified to be key regulators of cellular identity. For example the compact morphology of tomato is attributed to a mutation in SELF-PRUNING (SELF), which regulates the development of terminal flowers instead of further branching stems.5 Fruit yield per stem is attributed to a mutation in WUSCHEL HOMEOBOX9 (WOX9), a homeobox gene involved in maintaining the shoot’s capacity for growth.6 The cell-type-specific expression of such regulators therefore controls the shape, size, arrangement, and abundance of plant organs.
In species cultivated for compounds used in health care and industry, the location of the target product is often even more limited, accumulating in rare and specialized cell types. Some cell types, such as trichomes (epidermal outgrowths), are relatively easy to isolate and characterize, and many natural products have been found to be produced by these cells. For example, the antimalarial artemisinin, from Artemisia annua (sweet wormwood) and tetrahydrocannabinolic acid (THCA) from Cannabis sativa (cannabis) accumulate in leaf and floral glandular trichomes, respectively7,8 (Figure C). Other specialized plant cells are relatively poorly studied as they are embedded within complex tissues, but recent advances in single-cell technologies are producing novel insights. For example, in Catharanthus roseus (Madagascar periwinkle), the biosynthesis of vindoline, a precursor of the anticancer drugs vinblastine and vincristine, was located to laticifer and idioblast cells.9 However, relatively little is known about the differentiation and development of these cell types.10
Synthetic biology provides the potential to reprogram plant cell fates to engineer novel architectures and manipulate cellular identities. It could be applied to further optimize the architecture of cultivated species to, for example, engineer forms better suited to the water and nutrient stress events that are predicted to be more frequent and severe in a changing climate.11 Synthetic biology is already revolutionizing access to plant natural products via pathway reconstruction in heterologous hosts.12 However, extraction from field-grown crops or cultured tissue continues to be the most economical route to many compounds. The ability to engineer cell identities and conduct cell-type-specific metabolic engineering in the native host provides the potential to increase the yield of high-value compounds. Achieving these aims requires detailed information about plant cell-identities and of the cellular events that underlie cell fate. In this Perspective, we first discuss how synthetic biology can contribute to deepening our understanding of plant cell-types and cell-specific programs. We then discuss opportunities for engineering cell fates and functions for the benefit of agriculture and industry.
Synthetic Biology Approaches for Elucidating Plant Cellular Functions and Lineages
In the past, many key regulators of cellular identity were discovered either in mutation screens or using microarrays and RNA-sequencing to identify genes that change expression in specific organs or tissues. This was followed by functional characterization of candidates by overexpression/silencing and promoter-fusions to glucuronidase, luciferase, or fluorescent reporters (recently reviewed in refs (ref. 13 and ref. 14)). Later, the identification of genes expressed in specific cells was enabled by microdissection, isolation of marked nuclei, and fluorescence-activated cell sorting (FACS) of protoplasts isolated from reporter lines.15 In recent years, single-cell omics of protoplasts and isolated nuclei has led to an explosion in data. However, the identification and comparison of cell types across the diversity of plant lineages from gene-expression signatures remains a significant challenge.16 The advent of spatial genomics, which allows high-throughput fluorescent in situ hybridization (FISH) is beginning to address this, though to date it has only been applied to a few plant species.17,18 To understand how metabolic pathways are organized across cell-types, single cell transcriptomics and metabolomics have been integrated to link gene expression with the presence of specific compounds.19
Omics techniques provide a snapshot of the molecular state of cells but are destructive, making it challenging to observe dynamics. In contrast biosensors enable the high-resolution monitoring and quantification of metabolites, nutrients, hormones, small molecules, and gene expression within individual cells while preserving spatial information.20 In recent years, the ability to design novel biosensors and tune their function has been revolutionized by synthetic biology, resulting in an expansion of the biosensors for plants.
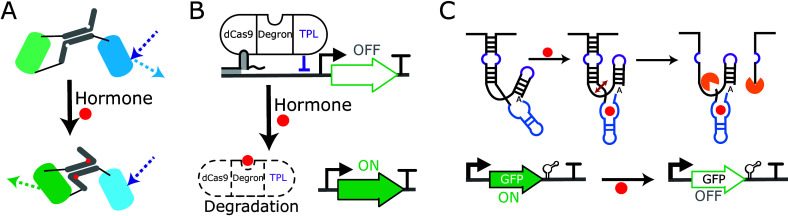
Direct biosensors, consisting of a single multifunctional module, are typically based on fluorescent proteins with optical properties dependent on the signal (e.g., pH, Ca2+), or are a fusion of two Forster Resonance Energy Transfer (FRET)-compatible fluorophores and a ligand-binding domain. In plants, direct biosensors have been designed to detect phytohormones including auxin21 (Figure A) and ABA,22 nutrients such as inorganic phosphate,23 and stress, e.g., via pH.24 Indirect biosensors have separate sensing, processing, and output components, which decouples sensing from output. Examples deployed in plants include those based on transcriptional regulation, e.g., a copper sensor fused with a Gal4 activator;25 on post-translational modification, e.g., a dCas9-based biosensor with a degron for the detection of plant hormones26 (Figure B); and on translation regulation, e.g., riboswitch-based biosensors for the detection of thiamine27 and theophylline28 (Figure C). Modularity of detection and output responses was demonstrated using a plant hormone receptor (PYR1) as a reprogrammable scaffold for detecting a range of molecules including cannabinoids and organophosphates, combined with various ligand-responsive outputs.29
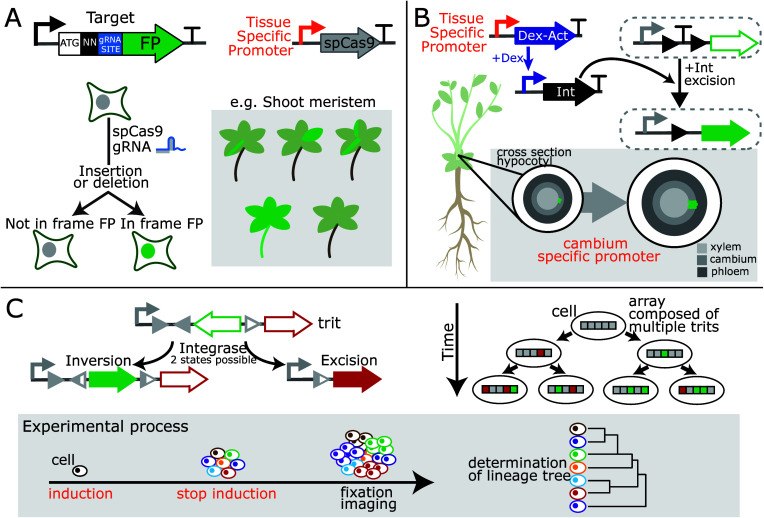
In general, live imaging is required to detect signals from in vivo biosensors. However, live imaging of large species is challenging, and the autofluorescence of many plant tissues can be difficult to overcome. An alternative is to record biosensor signals into DNA. This enables the presence of even transient signals to be detected after their presence either by DNA-sequencing or by imaging a fluorescent reporter, the expression of which is activated/repressed by biosensor-induced genetic changes. An advantage of these so-called DNA-recording devices is that the memory of detection is heritable, allowing cell lines to be traced. For this reason, they have been applied to understanding cell fates, cell lineages, and organ development. For example, CRISPR-Cas9-based DNA-recording was used to track cellular lineages in Arabidopsis thaliana (Arabidopsis) and Marchantia polymorpha(ref. 30) (Figure A). Integrase-based DNA-recorders were implemented in Arabidopsis to detect the expression of transcription factors that control lateral root development,31 and for analysis of cambium stem cells32 (Figure B). As plant cells are not generally mobile, live imaging has been used to track some cell lineages. Nevertheless, these new techniques provide an opportunity to understand the molecular mechanism of cellular transitions and can be used to differentiate lineages over different time periods while retaining spatial information. They also provide new opportunities for tissues and species for which live-imaging has presented technical challenges. In animals, these approaches are more advanced and have been used to obtain complete maps of cell ancestry for tissues and organs by multiplexing DNA editing sites with different probabilities forming molecular barcodes33,34 (Figure C). The barcodes can be either read using single-cell sequencing to reconstitute the cell-lineage tree or visualized by single molecule fluorescence in situ hybridization (smFISH). The application of such methods to plants might enable a complete map of cell ancestry and could be combined with omics methods to link cell lineages to cell fates and molecular states of cells.
Emerging Technologies for Controlling Plant Development and Performance
The architecture of plants is most often defined as the three-dimensional organization of the major organs. This can influence organ quantity, location, and size, which are significant contributors to the yields of target products. In aerial tissues, architecture includes the branching pattern of the stem(s) as well as the size, shape, and position of leaves and inflorescences. Fewer side branches are often beneficial in field systems, as they allow planting at higher density and facilitate mechanized harvesting. Compact architectures suited to high-density cultivation are also desirable for vertical farms, which require less water and can increase yield per unit area of land.35 Synthetic biology can be used to engineer these traits. For example, in some species, auxin prevents the development of axillary buds via a complex mechanism dependent on the strength of repression of auxin transporter PIN-FORMED1 (PIN1). Khakhar and colleagues engineered an auxin-activated HACR (hormone-activated Cas9-based repressors) to decrease the activation of expression of PIN1 by auxin and reduce feedback, leading to fewer side branches26 (Figure A).
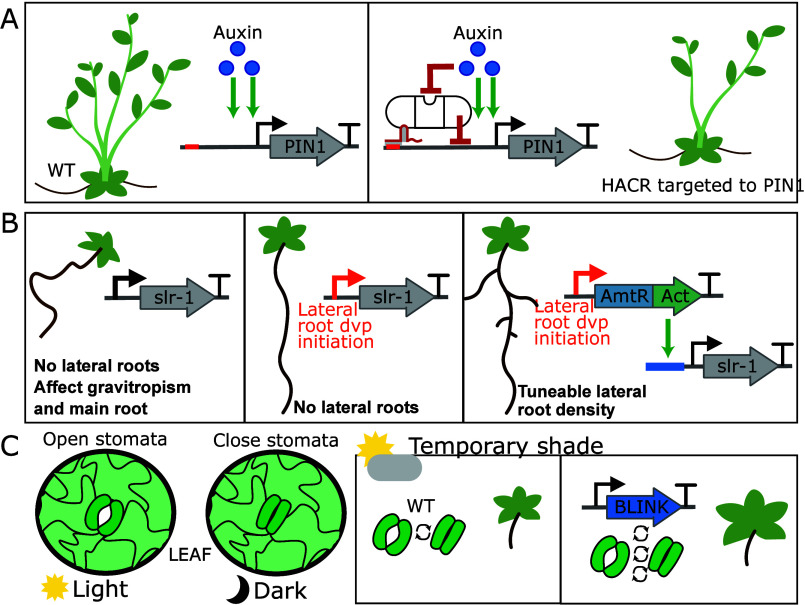
In root systems, the number, position, and density of lateral roots are essential parameters for optimum nutrient and water absorption.36 Synthetic biology approaches have been used to tune root architecture, including the combination of cell-type-specific expression with synthetic signal processing. When expressed via the native promoter, a mutant allele of a developmental regulator, solitary root (slr-1), affects root branching, gravitropism, root hair development, and primary root growth. Brophy and colleagues used buffer gates to tune the location and level of slr-1 in order to control root branching and limit unwanted effects37 (Figure B).
Plants also need to adapt to changes in growth conditions, including fluctuating light conditions. Stomata, pores in the leaf epidermis that regulate gas exchange, are generally open in light to enable CO2 absorption but close in the dark to limit water loss via transpiration. To accelerate stomatal kinetics, reducing the time in which photosynthesis is repressed, researchers engineered a optogenetic system consisting of a synthetic potassium channel and a blue-light sensor (BLINK1) under the control of guard cell-specific promoter38 (Figure C). Implementation resulted in increased growth under fluctuating white light without increasing water consumption.
Finally, certain cell types, architectures, and organs are present only in specific plant lineages. For example, symbiotic nitrogen fixation in the nodules of legumes is beneficial for plant growth. Many researchers are exploring engineering strategies to introduce nitrogen-fixation or symbiont-signaling symbiosis into nonlegumes (for a review, see ref (ref. 39)), but it remains challenging. Similarly, efficient photosynthesis at high temperatures is enabled by the specific leaf cellular architecture and photosynthetic biochemistry of C4 species, such as corn. Engineering the internal structure of the leaves of C3 plants such as rice is being explored to improve photosynthetic capacity (for a review, see ref (ref. 40)). In both of these cases, the development of strategies to engineer cellular identities and tissue architectures is likely to be beneficial.
Toward Cell-Type-Specific Metabolic Engineering
As noted above, many valuable bioactive and industrially relevant compounds are found in plants, often accumulating in specific tissues and cells. These include trichomes and laticifer cells in vascular plants7,8,41 and oil bodies in liverworts.42 This may be linked to specific functions, or because biosynthesis presents metabolic challenges to other cells such as toxicity and competition for precursors.43,44 Increasing the number of specialized cell-types represents a strategy to increase product yields. As several trichome-associated compounds have been associated with defense, this could also provide a route to improved resistance to pests and pathogens. Cannabis breeding has manipulated the number and type of trichomes to improve yields of THCA.45 In sweet wormwood, glandular trichome density was increased by manipulating the levels of developmental regulators, improving artemisinin yields (for a review, see ref (ref. 46)) (Figure A). Equivalent approaches might be used to increase the population of idioblast or laticifer cells, which are also known to accumulate valuable natural products in other species, including Madagascar periwinkle.9 However, relatively little is known about the developmental programs that lead to the differentiation of these cell-types.47
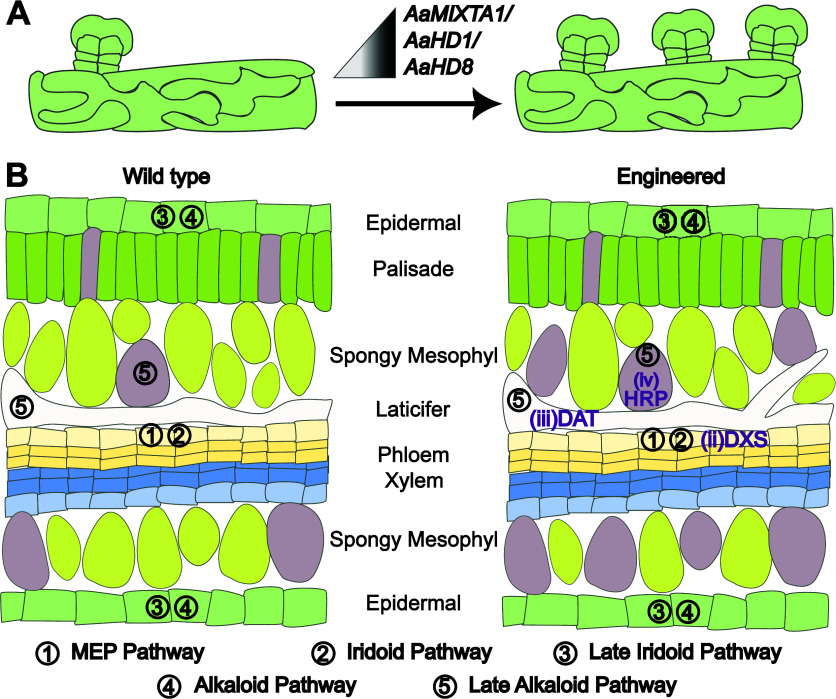
A second strategy is to upregulate production by cell-type-specific metabolic engineering. Strong, constitutive promoters can negatively affect plant growth and development.48 Further, increasing precursor availability in a cell-type-dependent manner rather than constitutively would reduce the transcriptional burden of the synthetic circuits, the impacts of which have been noted in other systems.49,50 Previously, it was found that overexpression of acetyl-CoA:4-O-deacetylvindoline 4-O-acetyltransferase (DAT), a rate-limiting enzyme in terpene-indole alkaloid biosynthesis in Madagascar periwinkle, led to increased vindoline biosynthesis.51 Recent single-cell data suggest that, as its precursor is mainly available in idioblast and laticifer cells, upregulation in these cell types may be sufficient (Figure B). Cell-type-specific engineering is likely to be even more beneficial when engineering precursor availability for the first committed steps. Enabling this in specific cells would avoid metabolic burdens in cells that lack the downstream pathway. In Madagascar periwinkle, precursor availability could be specifically engineered in leaf epidermal cells (Figure B). Similarly, it has been proposed that the rate-limiting steps for production of vinblastine is the oxidative coupling of catharanthine and vindoline, which may be improved by the expression of a peroxidase in idioblast cells19 (Figure B).
An alternative route to accessing natural products is heterologous biosynthesis. Although microbial production chassis are widely used, pathway reconstruction in plants, particularly species of Nicotiana including tobacco (N. tabacum) and an Australian relative, N. benthamiana, have been successful.52 To date, most studies have expressed pathways using strong constitutive promoters. This is generally successful when expression is transient, achieved by the leaf-infiltration of multiple strains of the Agrobacterium shuttle chassis each carrying individual genes.53 However, as this strategy requires containment glasshouses and bacterial cultivation, it may be economically viable only for high-value products. In contrast, field production of recombinant products in stable transgenics, e.g., N. tabacum, is low-cost,54 but requires the stable integration of synthetic pathways into the nuclear or plastid genome. Here, strong, constitutive promoters can lead to detrimental phenotypes, including impaired growth.48 Engineering strategies that limit expression to specific organs such as fruits, specific cell types, or tightly inducible expression are less likely to impact growth. A novel application for metabolically engineered plants is as living dispensers of volatile chemicals for pest control.55 In this case, production in trichomes and increases in trichome density are expected to be beneficial for product release.
Other plant species proposed for photosynthesis-driven bioproduction include the moss Physcomitrium patens(ref. 56) and the liverwort M. polymorpha.57 The latter produces complex oil bodies, specialized organelles that accumulate unique compounds including bisbibenzyls and several isoprenoid-derived compounds.58 It has been demonstrated that overexpression of the transcription factor, MpERF13, increases the number of oil body cells, which could improve its utility as a production chassis, but has deleterious effects on growth.59 These negative impacts might be avoided by precise spatiotemporal control of the expression.
Conclusions
Synthetic biology approaches have been applied to engineer a number of model plant species, demonstrating their utility for elucidating and engineering developmental programs and for metabolic engineering. While such proof-of-concepts are important, there is now a need to accelerate the engineering of crop plants for agriculture. The recent, rapid uptake of single-cell omics by the plant community is already producing a wealth of information about cellular functions in a wide range of agriculturally and industrially important species.16 Advances in synthetic biosensors and DNA-recording devices (Figure and 3) are also likely to provide novel insights into the molecular events that lead to cell differentiation and cell-type-specific functions. Currently, the plant synthetic biology community experiences bottlenecks imposed by low-efficiency and laborious plant transformation protocols. Methods to study cellular programs might also be applied to better understand and engineer improvements in plant regeneration. Ultimately, these advances will inform cell-type-specific engineering strategies to optimize plant body plans and accelerate the production of plant varieties for a rapidly changing climate and for the sustainable production of compounds for health and industry.
References
- A. Chandrasekhar, D. Dunne, G. Viglione. UN land report: Five key takeaways for climate change, food systems and nature loss.. Carbon Brief. April 27,, 2022
- R. A. Wing, M. D. Purugganan, Q. Zhang. The Rice Genome Revolution: From an Ancient Grain to Green Super Rice.. Nat. Rev. Genet., 2018. [DOI | PubMed]
- T. Lin, G. Zhu, J. Zhang, X. Xu, Q. Yu, Z. Zheng, Z. Zhang, Y. Lun, S. Li, X. Wang, Z. Huang, J. Li, C. Zhang, T. Wang, Y. Zhang, A. Wang, Y. Zhang, K. Lin, C. Li, G. Xiong, Y. Xue, A. Mazzucato, M. Causse, Z. Fei, J. J. Giovannoni, R. T. Chetelat, D. Zamir, T. Städler, J. Li, Z. Ye, Y. Du, S. Huang. Genomic Analyses Provide Insights into the History of Tomato Breeding.. Nat. Genet., 2014. [DOI | PubMed]
- R. A. Rapp, C. H. Haigler, L. Flagel, R. H. Hovav, J. A. Udall, J. F. Wendel. Gene Expression in Developing Fibres of Upland Cotton (Gossypium Hirsutum L.) Was Massively Altered by Domestication.. BMC Biol., 2010. [DOI | PubMed]
- L. Pnueli, L. Carmel-Goren, D. Hareven, T. Gutfinger, J. Alvarez, M. Ganal, D. Zamir, E. Lifschitz. The SELF-PRUNING Gene of Tomato Regulates Vegetative to Reproductive Switching of Sympodial Meristems and Is the Ortholog of CEN and TFL1.. Development, 1998. [DOI | PubMed]
- A. Hendelman, S. Zebell, D. Rodriguez-Leal, N. Dukler, G. Robitaille, X. Wu, J. Kostyun, L. Tal, P. Wang, M. E. Bartlett, Y. Eshed, I. Efroni, Z. B. Lippman. Conserved Pleiotropy of an Ancient Plant Homeobox Gene Uncovered by Cis-Regulatory Dissection.. Cell, 2021. [DOI | PubMed]
- M. Chen, T. Yan, Q. Shen, X. Lu, Q. Pan, Y. Huang, Y. Tang, X. Fu, M. Liu, W. Jiang, Z. Lv, P. Shi, Y.-N. Ma, X. Hao, L. Zhang, L. Li, K. Tang. GLANDULAR TRICHOME-SPECIFIC WRKY 1 Promotes Artemisinin Biosynthesis in Artemisia Annua.. New Phytol., 2017. [DOI | PubMed]
- C. A. S. Tanney, R. Backer, A. Geitmann, D. L. Smith. Cannabis Glandular Trichomes: A Cellular Metabolite Factory.. Front. Plant Sci., 2021. [DOI | PubMed]
- K. Yamamoto, K. Takahashi, L. Caputi, H. Mizuno, C. E. Rodriguez-Lopez, T. Iwasaki, K. Ishizaki, H. Fukaki, M. Ohnishi, M. Yamazaki, T. Masujima, S. E. O’Connor, T. Mimura. The Complexity of Intercellular Localisation of Alkaloids Revealed by Single-Cell Metabolomics.. New Phytol., 2019. [DOI | PubMed]
- A. R. Johnson, G. D. Moghe, M. H. Frank. Growing a Glue Factory: Open Questions in Laticifer Development.. Curr. Opin. Plant Biol., 2021. [DOI | PubMed]
- R. Anderson, P. E. Bayer, D. Edwards. Climate Change and the Need for Agricultural Adaptation.. Curr. Opin. Plant Biol., 2020. [DOI | PubMed]
- A. Cravens, J. Payne, C. D. Smolke. Synthetic Biology Strategies for Microbial Biosynthesis of Plant Natural Products.. Nat. Commun., 2019. [DOI | PubMed]
- N. M. Adamski, P. Borrill, J. Brinton, S. A. Harrington, C. Marchal, A. R. Bentley, W. D. Bovill, L. Cattivelli, J. Cockram, B. Contreras-Moreira, B. Ford, S. Ghosh, W. Harwood, K. Hassani-Pak, S. Hayta, L. T. Hickey, K. Kanyuka, J. King, M. Maccaferrri, G. Naamati, C. J. Pozniak, R. H. Ramirez-Gonzalez, C. Sansaloni, B. Trevaskis, L. U. Wingen, B. B. Wulff, C. Uauy. A Roadmap for Gene Functional Characterisation in Crops with Large Genomes: Lessons from Polyploid Wheat.. Elife, 2020. [DOI | PubMed]
- L. Villao-Uzho, T. Chávez-Navarrete, R. Pacheco-Coello, E. Sánchez-Timm, E. Santos-Ordóñez. Plant Promoters: Their Identification, Characterization, and Role in Gene Regulation.. Genes, 2023. [DOI | PubMed]
- R. Shaw, X. Tian, J. Xu. Single-Cell Transcriptome Analysis in Plants: Advances and Challenges.. Mol. Plant, 2021. [DOI | PubMed]
- K. D. Birnbaum, M. S. Otegui, J. Bailey-Serres, S. Y. Rhee. The Plant Cell Atlas: Focusing New Technologies on the Kingdom That Nourishes the Planet.. Plant Physiol., 2022. [DOI | PubMed]
- T. Nobori, M. Oliva, R. Lister, J. R. Ecker. Multiplexed Single-Cell 3D Spatial Gene Expression Analysis in Plant Tissue Using PHYTOMap.. Nat. Plants, 2023. [DOI | PubMed]
- L. Zhao, A. Fonseca, A. Meschichi, A. Sicard, S. Rosa. Whole-Mount smFISH Allows Combining RNA and Protein Quantification at Cellular and Subcellular Resolution.. Nat. Plants, 2023. [DOI | PubMed]
- C. Li, J. C. Wood, A. H. Vu, J. P. Hamilton, C. E. Rodriguez Lopez, R. M. E. Payne, D. A. Serna Guerrero, K. Gase, K. Yamamoto, B. Vaillancourt, L. Caputi, S. E. O’Connor, C. Robin Buell. Single-Cell Multi-Omics in the Medicinal Plant Catharanthus Roseus.. Nat. Chem. Biol., 2023. [DOI | PubMed]
- C. E. French, K. de Mora, N. Joshi, A. Elfick, J. Haseloff, J. Ajioka. SYNTHETIC BIOLOGY AND THE ART OF BIOSENSOR DESIGN;, 2011
- O. Herud-Sikimić, A. C. Stiel, M. Kolb, S. Shanmugaratnam, K. W. Berendzen, C. Feldhaus, B. Höcker, G. Jürgens. A Biosensor for the Direct Visualization of Auxin.. Nature, 2021. [DOI | PubMed]
- J. Rowe, M. Grangé-Guermente, M. Exposito-Rodriguez, R. Wimalasekera, M. O. Lenz, K. N. Shetty, S. R. Cutler, A. M. Jones. Next-Generation ABACUS Biosensors Reveal Cellular ABA Dynamics Driving Root Growth at Low Aerial Humidity.. Nat. Plants, 2023. [DOI | PubMed]
- P. Mukherjee, S. Banerjee, A. Wheeler, L. A. Ratliff, S. Irigoyen, L. R. Garcia, S. W. Lockless, W. K. Versaw. Live Imaging of Inorganic Phosphate in Plants with Cellular and Subcellular Resolution.. Plant Physiol., 2015. [DOI | PubMed]
- K. S. K. Gjetting, C. K. Ytting, A. Schulz, A. T. Fuglsang. Live Imaging of Intra- and Extracellular pH in Plants Using pHusion, a Novel Genetically Encoded Biosensor.. J. Exp. Bot., 2012. [DOI | PubMed]
- E. Garcia-Perez, B. Diego-Martin, A. Quijano-Rubio, E. Moreno-Giménez, S. Selma, D. Orzaez, M. Vazquez-Vilar. A Copper Switch for Inducing CRISPR/Cas9-Based Transcriptional Activation Tightly Regulates Gene Expression in Nicotiana Benthamiana.. BMC Biotechnol., 2022. [DOI | PubMed]
- A. Khakhar, A. R. Leydon, A. C. Lemmex, E. Klavins, J. L. Nemhauser. Synthetic Hormone-Responsive Transcription Factors Can Monitor and Re-Program Plant Development.. Elife, 2018. [DOI | PubMed]
- A. Subki, C. L. Ho, N. F. N. Ismail, A. A. Zainal Abidin, Z. N. Balia Yusof. Identification and Characterisation of Thiamine Pyrophosphate (TPP) Riboswitch in Elaeis Guineensis.. PLoS One, 2020. [DOI | PubMed]
- N. Shanidze, F. Lenkeit, J. S. Hartig, D. Funck. A Theophylline-Responsive Riboswitch Regulates Expression of Nuclear-Encoded Genes.. Plant Physiol., 2020. [DOI | PubMed]
- J. Beltrán, P. J. Steiner, M. Bedewitz, S. Wei, F. C. Peterson, Z. Li, B. E. Hughes, Z. Hartley, N. R. Robertson, A. V. Medina-Cucurella, Z. T. Baumer, A. C. Leonard, S.-Y. Park, B. F. Volkman, D. A. Nusinow, W. Zhong, I. Wheeldon, S. R. Cutler, T. A. Whitehead. Rapid Biosensor Development Using Plant Hormone Receptors as Reprogrammable Scaffolds.. Nat. Biotechnol., 2022. [DOI | PubMed]
- M. Donà, G. Bradamante, Z. Bogojevic, R. Gutzat, S. Streubel, M. Mosiolek, L. Dolan, O. Mittelsten Scheid. A Versatile CRISPR-Based System for Lineage Tracing in Living Plants.. Plant J., 2023. [DOI | PubMed]
- S. Guiziou, C. J. Maranas, J. C. Chu, J. L. Nemhauser. An Integrase Toolbox to Record Gene-Expression during Plant Development.. Nat. Commun., 2023. [DOI | PubMed]
- D. Shi, I. Lebovka, V. López-Salmerón, P. Sanchez, T. Greb. Bifacial Cambium Stem Cells Generate Xylem and Phloem during Radial Plant Growth.. Development, 2019. [DOI | PubMed]
- K.-H. K. Chow, M. W. Budde, A. A. Granados, M. Cabrera, S. Yoon, S. Cho, T.-H. Huang, N. Koulena, K. L. Frieda, L. Cai, C. Lois, M. B. Elowitz. Imaging Cell Lineage with a Synthetic Digital Recording System.. Science, 2021. [DOI | PubMed]
- A. McKenna, J. A. Gagnon. Recording Development with Single Cell Dynamic Lineage Tracing.. Development, 2019. [DOI | PubMed]
- C. A. O’Sullivan, C. L. McIntyre, I. B. Dry, S. M. Hani, Z. Hochman, G. D. Bonnett. Vertical Farms Bear Fruit.. Nat. Biotechnol., 2020. [DOI | PubMed]
- J. A. Postma, A. Dathe, J. P. Lynch. The Optimal Lateral Root Branching Density for Maize Depends on Nitrogen and Phosphorus Availability.. Plant Physiol., 2014. [DOI | PubMed]
- J. A. N. Brophy, K. J. Magallon, L. Duan, V. Zhong, P. Ramachandran, K. Kniazev, J. R. Dinneny. Synthetic Genetic Circuits as a Means of Reprogramming Plant Roots.. Science, 2022. [DOI | PubMed]
- M. Papanatsiou, J. Petersen, L. Henderson, Y. Wang, J. M. Christie, M. R. Blatt. Optogenetic Manipulation of Stomatal Kinetics Improves Carbon Assimilation, Water Use, and Growth.. Science, 2019. [DOI | PubMed]
- R. Huisman, R. Geurts. A Roadmap toward Engineered Nitrogen-Fixing Nodule Symbiosis.. Plant Commun., 2020. [DOI | PubMed]
- M. Ermakova, F. R. Danila, R. T. Furbank, S. von Caemmerer. On the Road to C4 Rice: Advances and Perspectives.. Plant J., 2020. [DOI | PubMed]
- S. Sun, X. Shen, Y. Li, Y. Li, S. Wang, R. Li, H. Zhang, G. Shen, B. Guo, J. Wei, J. Xu, B. St-Pierre, S. Chen, C. Sun. Single-Cell RNA Sequencing Provides a High-Resolution Roadmap for Understanding the Multicellular Compartmentation of Specialized Metabolism.. Nat. Plants, 2023. [DOI | PubMed]
- M. Tanaka, T. Esaki, H. Kenmoku, T. Koeduka, Y. Kiyoyama, T. Masujima, Y. Asakawa, K. Matsui. Direct Evidence of Specific Localization of Sesquiterpenes and Marchantin A in Oil Body Cells of Marchantia Polymorpha L.. Phytochemistry, 2016. [DOI | PubMed]
- F. Taura, M. Iijima, F. Kurosaki. Daurichromenic Acid and Grifolic Acid: Phytotoxic Meroterpenoids That Induce Cell Death in Cell Culture of Their Producer Rhododendron Dauricum.. Plant Signal. Behav., 2018. [DOI | PubMed]
- K. Malhotra, M. Subramaniyan, K. Rawat, M. Kalamuddin, M. I. Qureshi, P. Malhotra, A. Mohmmed, K. Cornish, H. Daniell, S. Kumar. Compartmentalized Metabolic Engineering for Artemisinin Biosynthesis and Effective Malaria Treatment by Oral Delivery of Plant Cells.. Mol. Plant, 2016. [DOI | PubMed]
- J. L. Parsons, S. L. Martin, T. James, G. Golenia, E. A. Boudko, S. R. Hepworth. Polyploidization for the Genetic Improvement of Cannabis Sativa.. Front. Plant Sci., 2019. [DOI | PubMed]
- C. Chalvin, S. Drevensek, M. Dron, A. Bendahmane, A. Boualem. Genetic Control of Glandular Trichome Development.. Trends Plant Sci., 2020. [DOI | PubMed]
- W. F. Pickard. Laticifers and Secretory Ducts: Two Other Tube Systems in Plants.. New Phytol., 2008. [DOI | PubMed]
- R. Mateos-Fernández, E. Moreno-Giménez, S. Gianoglio, A. Quijano-Rubio, J. Gavaldá-García, L. Estellés, A. Rubert, J. L. Rambla, M. Vazquez-Vilar, E. Huet, A. Fernández-Del-Carmen, A. Espinosa-Ruiz, M. Juteršek, S. Vacas, I. Navarro, V. Navarro-Llopis, J. Primo, D. Orzáez. Production of Volatile Moth Sex Pheromones in Transgenic Nicotiana Benthamiana Plants.. Biodes Res., 2021. [DOI | PubMed]
- F. Ceroni, R. Algar, G.-B. Stan, T. Ellis. Quantifying Cellular Capacity Identifies Gene Expression Designs with Reduced Burden.. Nat. Methods, 2015. [DOI | PubMed]
- T. Frei, F. Cella, F. Tedeschi, J. Gutiérrez, G.-B. Stan, M. Khammash, V. Siciliano. Characterization and Mitigation of Gene Expression Burden in Mammalian Cells.. Nat. Commun., 2020. [DOI | PubMed]
- Q. Wang, S. Xing, Q. Pan, F. Yuan, J. Zhao, Y. Tian, Y. Chen, G. Wang, K. Tang. Development of Efficient Catharanthus Roseus Regeneration and Transformation System Using Agrobacterium Tumefaciens and Hypocotyls as Explants.. BMC Biotechnol., 2012. [DOI | PubMed]
- M. J. Stephenson, J. Reed, N. J. Patron, L. P. George, A. Osbourn, B. Hung-Wen, T. P. B. Liu. 6.11 – Engineering Tobacco for Plant Natural Product Production. In. Comprehensive Natural Products III;, 2020. [DOI]
- Q. Chen, H. Lai, J. Hurtado, J. Stahnke, K. Leuzinger, M. Dent. Agroinfiltration as an Effective and Scalable Strategy of Gene Delivery for Production of Pharmaceutical Proteins.. Adv. Tech Biol. Med., 2013. [DOI | PubMed]
- J. F. Buyel. Plant Molecular Farming – Integration and Exploitation of Side Streams to Achieve Sustainable Biomanufacturing.. Front. Plant Sci., 2019. [DOI | PubMed]
- R. Mateos Fernández, M. Petek, I. Gerasymenko, M. Juteršek, Š. Baebler, K. Kallam, E. Moreno Giménez, J. Gondolf, A. Nordmann, K. Gruden, D. Orzaez, N. J. Patron. Insect Pest Management in the Age of Synthetic Biology.. Plant Biotechnol. J., 2022. [DOI | PubMed]
- R. Reski, H. Bae, H. T. Simonsen. Physcomitrella Patens, a Versatile Synthetic Biology Chassis.. Plant Cell Rep., 2018. [DOI | PubMed]
- C. R. Boehm, B. Pollak, N. Purswani, N. Patron, J. Haseloff. Synthetic Botany.. Cold Spring Harb. Perspect. Biol., 2017. [DOI | PubMed]
- Y. Asakawa, A. Ludwiczuk, F. Nagashima. Phytochemical and Biological Studies of Bryophytes.. Phytochemistry, 2013. [DOI | PubMed]
- T. Kanazawa, H. Morinaka, K. Ebine, T. L. Shimada, S. Ishida, N. Minamino, K. Yamaguchi, S. Shigenobu, T. Kohchi, A. Nakano, T. Ueda. The Liverwort Oil Body Is Formed by Redirection of the Secretory Pathway.. Nat. Commun., 2020. [DOI | PubMed]
