Computational pathology: A survey review and the way forward☆
Abstract
Computational Pathology (CPath) is an interdisciplinary science that augments developments of computational approaches to analyze and model medical histopathology images. The main objective for CPath is to develop infrastructure and workflows of digital diagnostics as an assistive CAD system for clinical pathology, facilitating transformational changes in the diagnosis and treatment of cancer that are mainly address by CPath tools. With evergrowing developments in deep learning and computer vision algorithms, and the ease of the data flow from digital pathology, currently CPath is witnessing a paradigm shift. Despite the sheer volume of engineering and scientific works being introduced for cancer image analysis, there is still a considerable gap of adopting and integrating these algorithms in clinical practice. This raises a significant question regarding the direction and trends that are undertaken in CPath. In this article we provide a comprehensive review of more than 800 papers to address the challenges faced in problem design all-the-way to the application and implementation viewpoints. We have catalogued each paper into a model-card by examining the key works and challenges faced to layout the current landscape in CPath. We hope this helps the community to locate relevant works and facilitate understanding of the field’s future directions. In a nutshell, we oversee the CPath developments in cycle of stages which are required to be cohesively linked together to address the challenges associated with such multidisciplinary science. We overview this cycle from different perspectives of data-centric, model-centric, and application-centric problems. We finally sketch remaining challenges and provide directions for future technical developments and clinical integration of CPath. For updated information on this survey review paper and accessing to the original model cards repository, please refer to GitHub. Updated version of this draft can also be found from arXiv.
Article type: Review Article
Keywords: Digital pathology, Whole slide image (WSI), Deep learning, Computer aided diagnosis (CAD), Clinical pathology, Survey
Affiliations: Department of Computer Science and Software Engineering (CSSE), Concordia Univeristy, Montreal, QC H3H 2R9, Canada; Qualcomm AI Research, Qualcomm Technologies Netherlands B.V., Amsterdam, The Netherlands; Institute for Research in Immunology and Cancer of the University of Montreal, Montreal, QC H3T 1J4, Canada; The Edward S. Rogers Sr. Department of Electrical & Computer Engineering (ECE), University of Toronto, Toronto, ON M5S 3G4, Canada; Huron Digitial Pathology, St. Jacobs, ON N0B 2N0, Canada; Department of Electrical and Computer Engineering, University of New Brunswick, Fredericton, NB E3B 5A3, Canada; University of Montreal Hospital Center, Montreal, QC H2X 0C2, Canada; Department of Computer Science, Stony Brook University, Stony Brook, NY 11794, United States
License: © 2024 The Author(s) CC BY 4.0 This is an open access article under the CC BY license (http://creativecommons.org/licenses/by/4.0/).
Article links: DOI: 10.1016/j.jpi.2023.100357 | PubMed: 38420608 | PMC: PMC10900832
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (1.9 MB)
Introduction
April 2017 marked a turning point for digital pathology when the Philips IntelliSite digital scanner received the US Food & Drugs Administration (FDA) approval (with limited use case) for diagnostic applications in clinical pathology.ref. bb0005,ref. bb0010 A subsequent validation guideline was created to help ensure the produced Whole Slide Image (WSI) scans could be used in clinical settings without compromising patient care, while maintaining similar results to the current gold standard of optical microscopy.ref. 3, ref. 4, ref. 5, ref. 6 The use of WSIs offers significant advantages to the pathologist’s workflow: digitally captured images, compared to tissue slides, are immune from accidental physical damage and maintain their quality over time.ref. bb0035,ref. bb0040 Clinics and practices can share and store these high-resolution images digitally enabling asynchronous viewing/collaboration worldwide.ref. bb0045,ref. bb0050 The development of digital pathology shows great promise as a framework to improve work efficiency in the practice of pathology.ref. bb0050,ref. bb0055 Adopting a digital workflow also opens immense opportunities for using computational methods to augment and expedite their workflow–the field of Computational Pathology (CPath) is dedicated to researching and developing these methods.ref. 12, ref. 13, ref. 14, ref. 15, ref. 16, ref. 17
However, despite the aforementioned advantages, the adoption of digital pathology, and hence computational pathology, has been slow. Some pathologists consider the analysis of WSIs as opposed to glass slides as an unnecessary change in their workflowref. bb0045,ref. 18, ref. 19, ref. 20 and recent surveys indicate that the switch to digital pathology does not provide enough financial incentive.ref. bb0040,ref. 21, ref. 22, ref. 23, ref. 24, ref. 25 This is where advances from CPath can address or overpower many of the concerns in adopting a digital workflow. For example, CPath models to identify morphological features that correlate with breast cancerref. bb0130 provide substantial benefits to clinical accuracy. Further, CPath models that identify lymph node metastases with better sensitivity while reducing diagnostic timeref. bb0135 can streamline workflows to increase pathologist throughput and generate more revenue.ref. bb0140,ref. bb0145
Similar to digital pathology, the adoption of CPath methods has also lagged despite the many benefits it offers to improve efficiency and accuracy in pathology.ref. bb0010,ref. 30, ref. 31, ref. 32 This lack of adoption and integration into clinical practice raises a significant question regarding the direction and trends of current work in CPath. This survey looks to review the field of CPath in a systematic fashion by breaking down the various steps involved in a CPath workflow and categorizing CPath works to both determine trends in the field and provide a resource for the community to reference when creating new works.
Existing survey papers in the field of CPath can be clustered into a few groups. The first focuses on the design and applications of smart diagnosis tools.ref. 15, ref. 16, ref. 17,ref. 33, ref. 34, ref. 35, ref. 36, ref. 37, ref. 38, ref. 39, ref. 40, ref. 41, ref. 42, ref. 43 These works focus on designing novel architectures for artificial intelligence (AI) models with regards to specific clinical tasks, although they may briefly discuss clinical challenges and limitations. A second group of works focus on clinical barriers for AI integration discussing specific certifications and regulations required for the development of medical devices under clinical settings.ref. 44, ref. 45, ref. 46, ref. 47, ref. 48, ref. 49 Lastly, the final group focuses on both the design and the integration of AI tools with clinical applications.ref. 12, ref. 13, ref. 14,ref. bb0145,ref. 50, ref. 51, ref. 52, ref. 53, ref. 54, ref. 55, ref. 56 These works speak to both the computer vision and pathology communities in developing machine learning (ML) models that can satisfy clinical use cases.
Our work is situated in this final group as we breakdown the end-to-end CPath workflow into stages and systematically review works related to and addressing those stages. We oversee this as a workflow for CPath research that breaks down the process of problem definition, data collection, model creation, and clinical validation into a cycle of stages. A visual representation of this cycle is provided in Fig. 1. We review over 700 papers from all areas of the CPath field to examine key works and challenges faced. By reviewing the field so comprehensively, our goal is to layout the current landscape of key developments to allow computer scientists and pathologists alike to situate their work in the overall CPath workflow, locate relevant works, and facilitate an understanding of the field’s future directions. We also adopt the idea of generating model cards fromref. bb0285 and designed a card format specifically tailored for CPath. Each paper we reviewed was catalogued as a model card that concisely describes (1) the organ of application, (2) the compiled dataset, (3) the machine learning model, and (4) the target task. The complete model card categorization of the reviewed publications is provided in Appendix A.12 for the reader’s use.
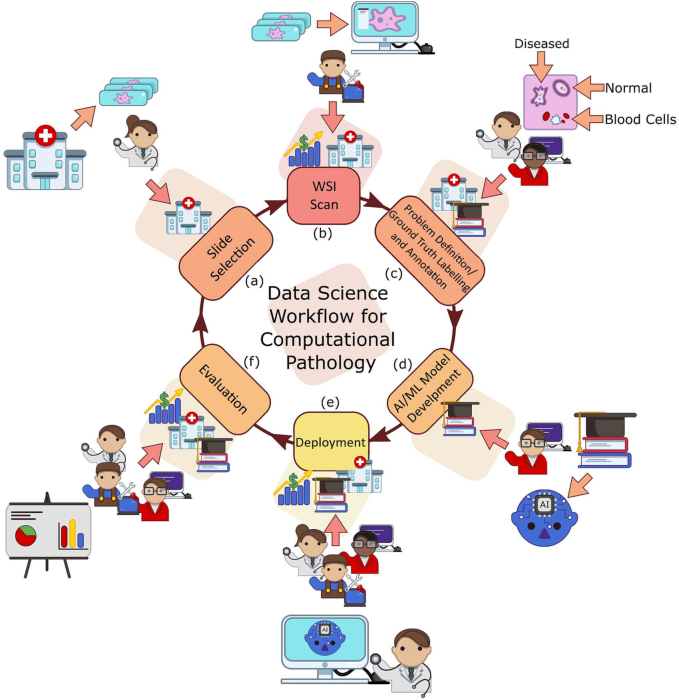
In our review of the CPath field, we find that two main approaches emerge in works: 1) a data-centric approach and 2) a model-centric approach. Considering a given application area, such as specific cancers, e.g. breast ductal carcinoma in-situ (DCIS), or a specific task, e.g. segmentation of benign and malignant regions of tissue, researchers in the CPath field focus generally on either improving the data or innovating on the model used.
Works with data-centric approaches focus on collecting pathology data and compiling datasets to train models on certain tasks based on the premise that the transfer of domain expert knowledge to models is captured by the process of collecting and labeling high-quality data.ref. bb0255,ref. bb0290,ref. bb0295 The motivation behind this approach in CPath is driven by the need to 1) address the lack of labeled WSI data representing both histology and histopathology cases due to the laborious annotation processref. bb0120 and 2) capture a predefined pathology ontology provided by domain expert pathologists for the class definitions and relations in tissue samples. Regarding the lack of labeled WSI data our analysis reveals that there are a larger number of datasets with granular labels, but there is a larger total amount of data available for a given organ and disease application that have weakly supervised labels at the Slide or Patient-level. Although some tasks, such as segmentation and detection, require WSI data to have more granular labels at the region-of-interest (ROI) or image mosaic/tiles (known as patch) levels, to capture more precise information for training models, there is a potential gap to leverage the large amount of weakly-supervised data to train models that can be later used downstream on smaller strongly-supervised datasets for those tasks. When considering the ontology of pathology as compared to the field of computer vision, we note that pathology datasets have far fewer classes than computer vision (e.g. ImageNet-20K contains 20,000 class categories for natural imagesref. bb0300 whereas CAMELYON17 has four annotated classes for breast cancer metastasesref. bb0305), but has much more variation within each of these classes in terms of representations and fuzzy boundaries around the grade of cancers which subdivides each class into many more in reality. There are also very rare classes in the form of rare diseases and cancers, as presented in Fig. 12 and discussed in Section 2, which present a class imbalance challenge when compiling data or training models. If one considers the complexities involved in representational learning of related tissues and diseases, it raises the question of whether there is a clear understanding and consensus in the field of how an efficient dataset should be compiled for model development. Our survey analyzes the availability of CPath datasets along with what area of application they address and their annotation level in detail in Section 3.3, and the complete table of datasets we have covered is available in Appendix A.9. Section 4 goes into more depth about the various levels of annotation, the annotation process, and selecting the appropriate annotation level for a task.
The model-centric approach, by contrast, is favoured by computer scientists and engineers, who design algorithmic approaches based on the available pathology data. Selection of a modelling approach, such as self-supervised, weakly-supervised, or strongly-supervised learning, is dictated directly by the amount of data available for a given annotation level and task. Currently, many models are developed on datasets with strongly-supervised labels at the ROI, Patch, or Pixel-levels to address tasks such as tissue type classification or disease detection. However, a recent trend is developing to apply self-supervised and weakly-supervised learning methods to leverage the large amount of data with Slide and Patient-level annotations.ref. bb0310 Models are trained in a self or weakly supervised manner to learn representations on a wider range of pathology data across organs and diseases, which can be leveraged for other tasks requiring more supervision but without the need for massive labeled datasets.ref. 63, ref. 64, ref. 65 This trend points to the future direction of CPath models following a similar trend to that in computer vision, where large-scale models are being pre-trained using self-supervised techniques to achieve state-of-the-art performance in downstream tasks.ref. bb0330,ref. bb0335
Although data and model centric approaches are both important in advancing the performance of models and tools in CPath, we note a need for much more application centric work in CPath. We define a study to be application centric if the primary focus is on addressing a particularly impactful task or need in the clinical workflow, ideally including clinical validation of the method or tool. To this end, Section 2 details the clinical pathology workflow from specimen collection to report generation, major task categories in CPath, and specific applications per organ. Particularly, we find that very few works focus on the pre or post-analytical phases of the pathology workflow where many errors can occur, instead focusing on the analytical phase where interpretation tasks take place. Additionally, certain types of cancer with deadly survival rates are underrepresented in CPath datasets and works. Very few CPath models and tools have been validated in a clinical setting by pathologists, suggesting that there may still be massive barriers to actually using CPath tools in practice. All of this points to a severe oversight by the CPath community towards considering the actual application and implementation of tools in a clinical setting. We suspect this to be a major reason as to why there is a slow uptake in adopting CPath tools by pathology labs.
The contributions of this survey include the provision of an end-to-end workflow for developing CPath work which outlines the various stages involved and is reflected within the survey sections. Further, we propose and provide a comprehensive conceptual model card framework for CPath that clearly categorizes works by their application of interest, dataset usage, and model, enabling consistent and easy comparison and retrieval of papers in relevant areas. Based on our analysis of the field, we highlight several challenges and trends, including the availability of datasets, focus on models leveraging existing data, disregard of impactful application areas, and lack of clinical validation. Finally, we give suggestions for addressing these aforementioned challenges and provide directions for future work in the hopes of aiding the adoption and implementation of CPath tools in clinical settings.
The structure of this survey closely follows the CPath data workflow illustrated in Fig. 1. Section 2 begins by outlining the clinical pathology workflow and covers the various task domains in CPath, along with organ specific tasks and diseases. The next step of the workflow involves the processes and methods of histopathology data collection, which is outlined in Section 3. Following data collection, Section 4 details the corresponding annotation and labeling methodology and considerations. Section 5 covers deep learning designs and methodologies for CPath applications. Section 6 focuses on regulatory measures and clinical validation of CPath tools. Section 7 explores emerging trends in recent CPath research. Finally, we provide our perceived challenges and future outlook of CPath in Section 8.
Clinical applications for CPath
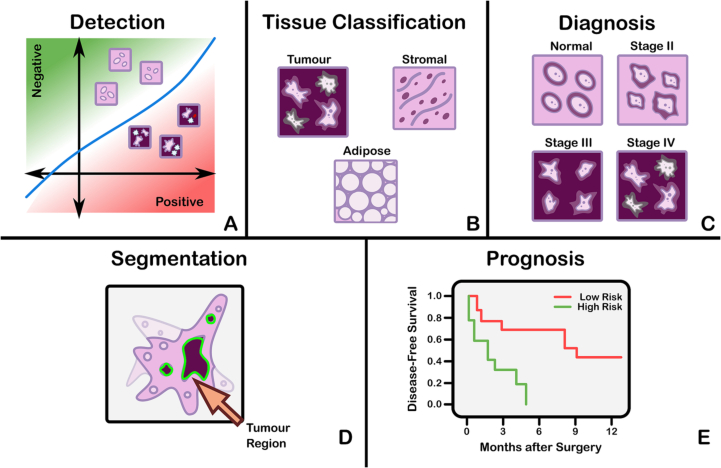
The field of CPath is dedicated to the creation of tools that address and aid steps in the clinical pathology workflow. Thus, a grounded understanding of the clinical workflow is of paramount importance before development of any CPath tool. The outcomes of clinical pathology are diagnostics, prognostics, and predictions of therapy response. Computational pathology systems that focus on diagnostic tasks aim to assist the pathologists in tasks such as tumour detection, tumour grading, quantification of cell numbers, etc. Prognostic systems aim to predict survival for individual patients while therapy response predictive models aid personalized treatment decisions based on histopathology images. Fig. 3 visualizes the goals pertaining to these tasks. In this section, we provide detail on the clinical pathology workflow, the major application areas in diagnostics, prognostics, and therapy response, and finally detail the cancers and CPath applications in specific organs. The goal is to outline the tasks and areas of application in pathology where CPath tools and systems can be developed and implemented.
Clinical pathology workflow
This subsection provides a general overview of the clinical workflow in pathology covering the collection of a tissue sample, its subsequent processing into a slide, inspection by a pathologist, and compilation of the analysis and diagnosis into a pathology. Fig. 2 summarizes these steps at a high level and provides suggestions for corresponding CPath applications. The steps are organized under the conventional pathology phases for samples: pre-analytical, analytical, and post-analytical. These phases were developed to categorize quality control measures, as each phase has its own set of potential sources of errors,ref. bb0340 and thus potential sources of corrections during which CPath and healthcare artificial intelligence tools could prove useful. For details about each step of the workflow, please refer to the Appendix A.1.
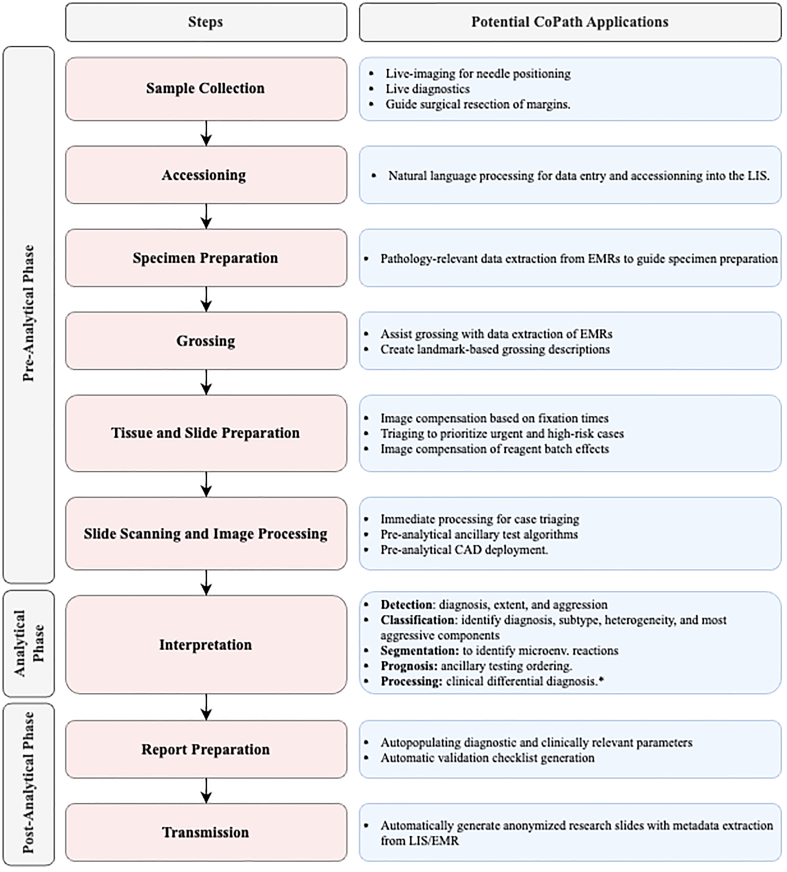
Pre Analytical Phase The first step of the pre-analytical phase is a biopsy performed to collect a tissue sample, where the biopsy method is dependent on the type of sample required and the tissue characteristics. Sample collection is followed by accessioning of the sample which involves entering of the patient and specimen information into a Laboratory Information System (LIS) and linking to the Electronic Medical Records (EMR) and potentially a Slide Tracking System (STS). After accessioning, smaller specimens that have not already been preserved by fixation in formalin are fixated. Once the basic specimen preparation has occurred, the tissue is analyzed by the pathology team without the use of a microscope; a step called grossing. Grossing involves cross-referencing the clinical findings and the EMR reports, with the operator localizing the disease, locating the pathological landmarks, describing these landmarks, and measuring disease extent. Specific sampling of these landmarks is performed, and these samples are then put into cassettes for the final fixation. Subsequently, the samples are then sliced using a microtome, stained using the relevant stains for diagnosis, and covered with a glass slide.
Analytical Phase After a slide is processed and prepared, a pathologist views the slide to analyze and interpret the sample. The approach to interpretation varies depending on the specimen type. Interpretation of smaller specimens is focused on diagnosis of any disease. Analysis is performed in a decision-tree style approach to add diagnosis-specific parameters, e.g. esophagus biopsy type of sampled mucosa presence of folveolar-type mucosa identify Barrett’s metaplasia identify degree of dysplasia. Once the main diagnosis has been identified and characterized, the pathologist sweeps the remaining tissue for secondary diagnoses which can also be characterized depending on their nature. Larger specimens are more complex and usually focus on characterizing the tissue and identifying unexpected diagnoses beyond the prior diagnosis from a small specimen biopsy. Microscopic interpretation of large specimens is highly dependent on the quality of the grossing and the appropriate detection and sampling of landmarks. Each landmark (e.g., tumor surface, tumor at deepest point, surgical margins, lymph node in mesenteric fat) is characterized either according to guidelines, if available, or according to the pathologist’s judgment. After the initial microscopic interpretation additional deeper cuts (“levels”), special stains, immunohistochemistry (IHC), and/or molecular testing may be performed to hone the diagnosis by generating new material or slides from the original tissue block.
Post-Analytical Phase The pathologist synthesizes a diagnosis by aggregating their findings from grossing and microscopic examination in combination with the patient’s clinical information, all of which are included in a final pathology report. The classic sections of a pathology report are patient information, a list of specimens included, clinical findings, grossing report, microscopic description, final diagnosis, and comment. The length and degree of complexity of the report again depends on the specimen type. Small specimen reports are often succinct, clearly and unambiguously listing relevant findings which guide treatment and follow-up. Large specimen reports depend on the disease, for example, in cancer resection specimens the grossing landmarks are specifically targeted at elements that will guide subsequent treatment.
In the past, pathology reports had no standardized format, usually taking a narrative-free text form. Free text reports can omit necessary data, include irrelevant information, and contain inconsistent descriptions.ref. bb0345 To combat this, synoptic reporting was introduced to provide a structured and standardized reporting format specific to each organ and cancer of interest.ref. bb0345,ref. bb0350 Over the last 15 years, synoptic reporting has enabled pathologists to communicate information to surgeons, oncologists, patients, and researchers in a consistent manner across institutions and even countries. The College of American Pathologists (CAP) and the International Collaboration on Cancer Reporting (ICCR) are the two major institutions publishing synoptic reporting protocols. The parameters included in these protocols are determined and updated by CAP and ICCR respectively to remain up-to-date and relevant for diagnosis of each cancer type. For the field of computational pathology, synoptic reporting provides a significant advantage in dataset and model creation, as a pre-normalized set of labels exist across a variety of cases and slides in the form of the synoptic parameters filled out in each report. Additionally, suggestion or prediction of synoptic report values are a possible CPath application area.
Diagnostic tasks
Computational pathology systems that focus on diagnostic tasks can broadly be categorized as: (1) disease detection, (2) tissue subtype classification, (3) disease diagnosis, and (4) segmentation. These tasks are visually depicted in Fig. 3. Note how the detection tasks all involve visual analysis of the tissue in WSI format. Thus computer vision approach is primarily adopted towards tackling diagnostic tasks in computer aided diagnosis (CAD). For additional detail on some previous works on these diagnostic tasks, we refer the reader to Appendix A.2
Detection We define the detection task as a binary classification problem where inputs are labeled as positive or negative, indicating the presence or absence of a certain feature. There may be variations in the level of annotation required, e.g. slide-level, patch-level, pixel-level detection depending on the feature in question. Although detection tasks may not provide an immediate disease diagnosis, it is a highly relevant task in many pathology workflows as pathologists incorporate the presence or absence of various histological features into synoptic reports that lead to diagnosis. Broadly, detection tasks fall into two main categories: (1) screening the presence of cancers and (2) detecting histopathological features specific to certain diagnoses.
Cancer detection algorithms can assist the pathologists by filtering obviously normal WSIs and directing pathologist’s focus to metastatic regions.ref. bb0355 Although pathologists have to review all the slides to check for multiple conditions regardless of the clinical diagnosis, an accurate cancer detection CAD would expedite the workflow by pinpointing the ROIs and summarizing results into synoptic reports, ultimately leading to a reduces time per slide. Due to this potential impact, cancer detection tasks have been explored in a broad set of organs. Additionally, the simple labeling in binary detection tasks allows for deep learning methods to generalize across different organs where similar cancers form.ref. 72, ref. 73, ref. 74
Tissue Subtype Classification Treatment and patient prognosis can vary widely depending on the stage of cancer, and finely classifying specific tissue structures associated with a specific disease type provides essential diagnostic and prognostic information.ref. bb0375 Accordingly, accurately classifying tissue subtypes is a crucial component of the disease diagnosis process. As an example, discriminating between two forms of glioma (a type of brain cancer), glioblastoma multiforme and lower grade glioma, is critical as they differ by over in patient survival rates.ref. bb0380 Additionally, accurate classification is key in colorectal cancer (CRC) diagnosis, as high morphological variation in tumor cellsref. bb0385 makes certain forms of CRC difficult to diagnose by pathologists.ref. bb0390 We define this classification of histological features as tissue subtype classification.
Disease Diagnosis The most frequently explored design of deep learning in digital pathology involves emulating pathologist diagnosis. We define this multi-class diagnosis problem as a disease diagnosis task. Note the similarity with detection–disease diagnosis can be considered a fine-grained classification problem which subdivides the general positive disease class into finer disease-specific labels based on the organ and patient context.
Segmentation The segmentation task moves one step beyond classification by adding an element of spatial localization to the predicted label(s). In semantic segmentation, objects of interest are delineated in an image by assigning class labels to every pixel. These class labels can be discrete or non-discrete, the latter being a more difficult task.ref. bb0395 Another variant of the segmentation task is instance segmentation, which aims to achieve both pixel-level segmentation accuracy as well as clearly defined object (instance) boundaries. Segmentation approaches can accurately capture many morphological statisticsref. bb0400 and textural features,ref. bb0405 both of which are relevant for cancer diagnosis and prognosis. Most frequently, segmentation is used to capture characteristics of individual glands, nuclei, and tumor regions in WSIs. For instance, glandular structure is a critical indicator of the severity of colorectal carcinoma,ref. bb0410 thus accurate segmentation could highlight particularly abnormal glands to the pathologist as demonstrated in.ref. 82, ref. 83, ref. 84 Overall, segmentation provides localization and classification of cancer-specific tumors and of specific histological features that can be meaningful for the pathologist’s clinical interpretation.
Prognosis
Prognosis involves predicting the likely development of a disease based on given patient features. For accurate survival prediction, models must learn to both identify and infer the effects of histological features on patient risk. Prognosis represents a merging of the diagnosis classification task and the disease-survivability regression task.
Training a model for prognosis requires a comprehensive set of both histopathology slides and patient survival data (i.e. a variant of multi-modal representation learning). Despite the complexity of the input data, ML models are still capable of extracting novel histological patterns for disease-specific survivability.ref. 85, ref. 86, ref. 87 Furthermore, strong models can discover novel prognostically-relevant histological features from WSI analysis.ref. bb0440,ref. bb0445 As the quality and comprehensiveness of data improves, additional clinical factors could be incorporated into deep learning analysis to improve prognosis.
Prediction of treatment response
With the recent advances in targeted therapy for cancer treatment, clinicians are able to use treatment options that precisely identify and attack certain types of cancer cells. While the number of options for targeted therapy are constantly increasing, it becomes increasingly important to identify patients who are potential therapy responders to a specific therapy option and avoid treating non-responding patients who may experience severe side effects. Deep learning can be used to detect structures and transformations in tumour tissue that could be used as predictive markers of a positive treatment response. Training such deep learning models usually requires large cohorts of patient data for whom the specific type of treatment option and the corresponding response is known.
Organs and diseases
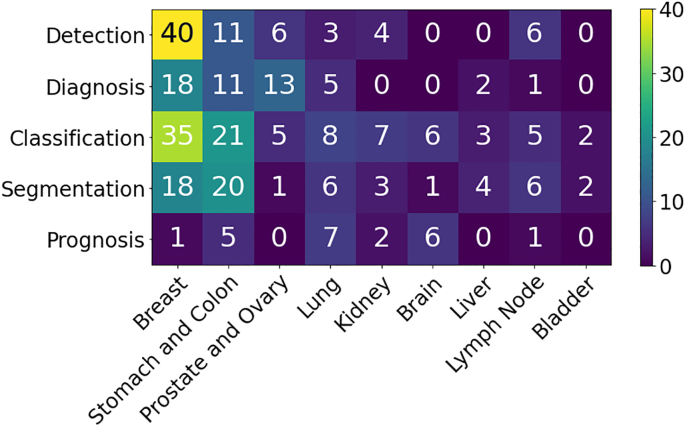
This section presents an overview of the various anatomical application areas for computational pathology grouped by the targeted organ. Each organ section gives a brief overview of the types of cancers typically found and the content of the pathology report as noted from the corresponding CAP synoptic reporting outline (discussed at 2.1). Fig. 4 highlights the intersection between the major diagnostic tasks and the anatomical focuses in state-of-the-art research. The majority of papers are dedicated to the four most common cancer sites: breast, colon, prostate, and lung.ref. bb0450 Additionally, a significant amount of research is also done on cancer types with highest mortality, brain and liver.ref. bb0450 Note that details of some additional works that may be of interest for each organ type can be found in Appendix A.7 (see Fig. 5).
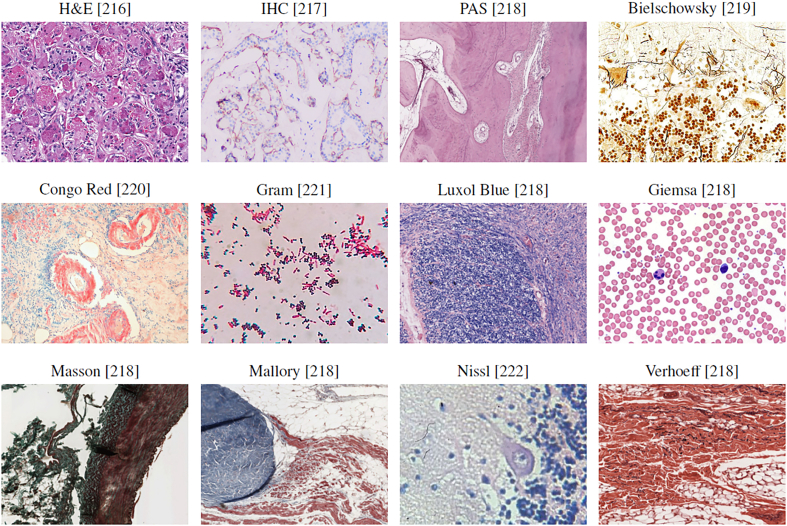
Breast Breast cancers can start from different parts of the breast and majorly consist of 1) Lobular cancers that start from lobular glands, 2) Ductal cancers, 3) Paget cancer which involves the nipple, 4) Phyllodes tumor that stems from fat and connective tissue surrounding the ducts and lobules, and 5) Angiosarcoma which starts in the lining of the blood and lymph vessels. In addition, based on whether the cancer has spread or not, breast cancers can be categorized into in situ or invasive/infiltrating forms. DCIS is a precancerous state and is still confined to the ducts. Once the cancerous cells grow out of the ducts, the carcinoma is now considered invasive or infiltrative and can metastasize.ref. bb0455
Synoptic reports for breast cancer diagnosis are divided based on the type of cancers mentioned above. For DCIS and invasive breast cancers, synoptic reports focus on the histologic type and grade, along with the nuclear grade, evidence of necrosis, margin, involvement of regional lymph nodes, and biomarker status. Notably, architectural patterns are no longer a valuable predictive tool compared to nuclear grade and necrosis to determine a relative ordering of diagnostic importance for DCIS.ref. bb0460 In contrast to DCIS and invasive cancers, Phyllodes tumours vary due to their differing origin in the fat and connective tissue, focusing on analyzing the stroma characteristics, existence of heterologous elements, mitotic rate, along with the involvement of lymph nodes. Finally, to determine therapy response and treatments, biomarker tests for Estrogen, Progesteroneref. bb0465 and HER-2ref. bb0470 receptors are recommended, along with occasional tests for Ki67 antigens.ref. bb0475,ref. bb0480
Most breast cancer-focused works in CPath propose various solutions for carcinoma detection and metastasis detection, an important step for assessing cancer stage and morbidity. Metastasis detection using deep learning methods was shown to outperform pathologists’ exhaustive diagnosis by free-response receiver operating characteristic (FROC) in.ref. bb0485
Prostate Prostate cancer is the second most prevalent cancer among the total population and the most common cancer among men (both excluding non-melanoma skin cancers). However, most prostate cancers are not lethal. Prostate cancer can occur in any of the three prostate zones: Central (CZ), Peripheral (PZ), and Transition (TZ), in increasing order of aggressiveness. Prostate cancers are almost always adenocarcinomas, which develop from the gland cells that make prostate fluid. The other types of prostate cancers are small cell carcinomas, neuroendocrine tumors, transitional cell carcinomas, isolated intraductal carcinoma, and sarcomas (which are very rare). Other than cancers, there are multiple conditions that are important to identify or diagnose as precursors to cancer or not. Prostatic intraepithelial neoplasia (PIN) is diagnosed as either low-grade PIN or high-grade PIN. Men with high-grade PIN need closely monitored follow-up sessions to screen for prostate cancer. Similarly, atypical small acinar proliferation (ASAP) is another precancerous condition requiring follow-up biopsies.ref. bb0490
To grade and score tumours, pathologists use a Tumor, Nodes, Metastasis (TNM) framework. In the synoptic report, pathologists identify and report the histologic type and grades, and involvement of regional lymph nodes to help grade and provide prognosis for any tumours. Specifically for prostate analysis, tumour size and volume are both important factors in prognosis according to multiple studies.ref. 99, ref. 100, ref. 101, ref. 102 Similarly, location is important to note for both prognosis and therapy response.ref. bb0515 Invasion to nearby (except perineural invasion) tissues is noted and can correlate to TMN classification.ref. bb0520 Additionally, margin analysis is especially important in prostate cancers as the presence of a positive margin increases the risk of cancer recurrence and metastasis.ref. bb0525 Finally, intraductal carcinoma (IDC) must be identified and distinguished from PIN and PIA; as it is strongly associated with a high Gleason score, a high-volume tumor, and metastatic disease.ref. 106, ref. 107, ref. 108, ref. 109, ref. 110
After a prostate cancer diagnosis is established, pathologists assign a Gleason Score to determine the cancer’s grade: a grade from 1 to 5 is assigned to the two most common areas and those two grades are summed to make a final Gleason Score.ref. bb0555 For Gleason scores of 7, where survival and clinical outcomes demonstrate large variance, the identification of Cribriform glands is key in helping to narrow possible outcomes.ref. bb0560,ref. bb0565
Ovary Ovarian cancer is the deadliest gynecologic malignancy and accounts for more than deaths each year.ref. bb0570 Ovarian cancer manifests in three types: 1) epithelial cell tumors that start from the epithelial cells covering the outer surface of the ovary, 2) germ cell tumors which start from the cells that produce eggs, and 3) stromal tumors which start from cells that hold the ovary together and produce the hormones estrogen and progesterone. Each of these cancer types can be classified into benign, intermediate and malignant categories. Overall, epithelial cell tumors are the most common ovarian cancer and have the worst prognosis.ref. bb0575
When compiling a synoptic report for ovarian cancer diagnosis, pathologists focus on histologic type and grade, extra-organ involvement, regional lymph nodes, T53 gene mutations, and serous tubal intraeptithelial carconma (STIC). Varying histologic tissue types are vital to determine the pathology characteristics and determining eventual prognosis. For example, generally endometrioid, mucinous, and clear cell carcinomas have better outcomes than serous carcinomas.ref. bb0580 Additionally, lymph node involvement and metastasis in both regional and distant nodes has a direct correlation to patient survival, grading, and treatment. Determining the presence of STICs correlates directly to the presence of ovarian cancer, as of ovarian cancer patients will also have an associated STIC.ref. bb0570 Finally, T53 gene mutations are the most common in epithelial ovarian cancer; which has the worst prognosis among ovarian cancers, so determining their presence is critical to patient cancer risk and therapy response.ref. bb0585,ref. bb0590 There are not a large number of works dedicated to the ovary specifically, but most works on ovary focus on classification of its five most common cancer subtypes: high-grade serous (HGSC), low-grade serous (LGSC), endometriod (ENC), clear cell (CCC), and mucinous (MUC).ref. bb0595,ref. bb0600
Lung Lung cancer is the third most common cancer, next to breast and prostate cancer.ref. bb0605 Lung cancers mostly start in the bronchi, bronchioles, or alveoli and are divided into two major types, non-small cell lung carcinomas (NSCLC) () and small cell lung carcinomas (SCLC) (). Although NSCLS cancers are different in terms of origin, they are grouped because they have similar outcomes and treatment plans. Common NSCLS cancers are 1) adenocarcinoma, 2) squamous cell carcinoma 3) large cell carcinoma, and some other uncommon subtypes.ref. bb0610
For reporting, histologic type helps determine NSCLC vs SCLC and the subtype of NSCLC. Although NSCLC generally has favourable survival rates and prognosis as compared to SCLC, certain subtypes of NSCLC can have lower survival rates due to co-factors.ref. bb0615 Histologic patterns are applicable in adenocarcinomas, consisting of favourable types: lepidic, intermediate: acinar and papillary, and unfavourable: micropapillary and solid.ref. bb0620 Grading each histologic type aids in categorization but is differentiated based on each type, and thus is out of scope for this paper. Importantly for lung cancers, tumour size is an independent prognostic factor for early cancer stages, lymph node positivity, and locally invasive disease. Additionally, the size of the invasive portion is an important factor for prognosis of nonmucinous adenocarcinoma with lepidic pattern.ref. bb0615,ref. 125, ref. 126, ref. 127, ref. 128, ref. 129 Other important lung specific features are visceral pleural invasion, which is associated with worse prognosis in early-stage lung cancer even with tumors <3cm,ref. bb0650 and lymphatic invasion which indicates an unfavourable prognostic finding.ref. bb0625,ref. bb0655
Colon and Rectum Colorectal cancers are two of the five most common cancer types.ref. bb0450 Cancer cells usually start to develop in the innermost layer of the colon and rectum walls, known as the mucosa, and continue their way up to other layers. In other layers, there are lymph and blood vessels that can be used by cancer cells to travel to nearby lymph nodes or other organs.ref. bb0660 Colorectal cancers usually start with the creation of different types of polyps, each possessing a unique risk of developing into cancer. Most colorectal cancers are adenocarcinomas, which are split into three well-studied subtypes: classic adenocarcinoma (AC), signet ring cell carcinoma (SRCC), and mucinous adenocarcinoma (MAC). In most cases, AC has a better prognosis than MAC or SRCC. Other types, albeit uncommon, of colorectal cancers are: carcinoid tumors, gastrointestinal stromal tumors (GISTs), lymphomas, and sarcomas.ref. bb0665
As in other cancers, histologic grade is the most important factor in cancer prognosis along with regional lymph node status and metastasis. The tumor site is also important in determining survival rates and prognosis.ref. bb0670 Vascular invasion of both small and large vessels are important factors in adverse outcomes and metastasis,ref. 135, ref. 136, ref. 137 and perineural invasion has been shown in multiple studies to be an indicator of poor prognosis.ref. 137, ref. 138, ref. 139 Additionally, microsatellite instability (MSI) is shown to be a good indicator of prognosis and is divided into three stages in decreasing adversity of Stable (MSI-S), Low (MSI-L), and High (MSI-H).ref. bb0700 Finally, some studies have indicated the usefulness of biomarkers in colorectal cancer treatment, with biomarkers such as BRAF mutations, KRAS mutations, MSI, APC, Micro-RNA, and PIK3CA.ref. bb0705
Works are relatively well-distributed among various tasks including disease diagnosis, segmentation, and detection. Expanding on colorectal cancer detection, work fromref. bb0710 used feature analysis for colorectal and mucinous adenocarcinomas using heatmap visualizations. They discovered that adenocarcinoma is often detected by ill-shaped epithelial cells and that misclassification can occur due to lumen regions that resemble the malformed epithelial cells. Similarly for mucinous carcinoma, the model again recognizes the dense epithelium, but this time ignores the primary characteristic of the carcinoma (abundance of extracellular mucin). These findings suggest that a thorough analysis of class activation maps can be helpful for improving the classifier’s accuracy and intuitiveness.
Bladder There are several layers within the bladder wall with most cancers starting in the internal layer, called the urothelium or transitional epithelium. Cancers remaining in the inner layer are non-invasive or carcinoma in situ (CIS) or stage 0. If they grow into other layers such as the muscle or fatty layer, the cancer is now invasive. Nearly all bladder cancers are urothelial carcinomas or transitional cell carcinomas (TCC). However, there are other types of cancer such as squamous cell carcinomas, adenocarcinomas, small cell carcinomas, and sarcomas which all are very rare. In the early stages, all types of bladder cancers are treated similarly but as their stage progresses, and chemotherapy is needed, different drugs might be used based on the type of the cancer.ref. bb0715 As with other organs, histologic type and grade also play a role in prognosis and treatment,ref. bb0720 and lymphovascular invasion is independently associated with poor prognosis and recurrence.ref. bb0725
Works focusing on the bladder display promising results that could lead to rapid clinical application. For example, a prediction method for four molecular subtypes (basal, luminal, luminal p53, and double negative) of muscle-invasive bladder cancer was proposed in,ref. bb0730 outperforming pathologists by in classification accuracy when restricted to a tissue morphology-based assessment. Further improvements in accuracy could help expedite diagnosis by complementing traditional molecular testing methods.
Kidney Each kidney is made up of thousands of glomeruli which feed into the renal tubules. Kidney cancer can occur in the cells that line the tubules (renal cell carcinoma (RCC)), blood vessels and connective tissue (sarcomas), or urothelial cells (Urothelial carcinoma). RCC accounts for about of kidney cancers and comes in two types: 1) clear cell renal carcinoma, which are most common and 2) non-clear cell renal carcinoma consisting of papillary, chromophobe and some very rare subtypes.ref. bb0735 The CAP’s cancer protocol template for the kidney is solely focused on RCCs,ref. bb0740 likely due to their high probability. Tumour size is directly associated with malignancy rates, with cm size increase resulting in increase in malignancy chance.ref. bb0745 Additionally, the RCC histologic type is correlated with metastasis, with clear cell, capillary, collecting ducts (Bellini), and medullary being the most aggressive ones.ref. bb0750
Many works are focused on glomeruli segmentation, as the number of glomeruli and glomerulosclerosis constitute standard components of a renal pathology report.ref. bb0755 In addition to glomeruli detection, some works have also detected other relevant features such as tubules, Bowman’s capsules, and arteries.ref. bb0760 The results display strong performance on PAS-stained nephrectomy samples and tissue transplant biopsies, and there seems to be a strong correlation between the visual elements identified by the network and those identified by renal pathologists.
Brain There are two main types of brain tumors: malignant and non-malignant. Malignant tumors can be classified as primary tumors (originating from the brain) or secondary (metastatic).ref. bb0765,ref. bb0770 The most common type of brain cancers is gliomas, occurring of the time, and are classified into four grades.ref. bb0775 In the synoptic reporting, tumour location is noted as it has some impact on the prognosis, with parietal tumours showing better prognosis compared to other locations.ref. bb0765 Additionally, focality of glioblastomas (a subtype of gliomas) is important to determine as multifocal glioblastoma is far more aggressive and resistant to chemotherapy as compared to unifocal.ref. bb0770 A recent summary of the World Health Organization’s (WHO) classification of tumors of the central nervous system has indicated that biomarkers as both ancillary and diagnostic predictive tools.ref. bb0780 Additionally, in a recent WHO edition of classification of tumours of the central nervous system, molecular information is now integrated with histologic information into tumor diagnosis for cases such as diffuse gliomas and embryonal tumors.ref. bb0785
Accordingly, most works focus on gliomas and more specifically glioblastoma, the most aggressive and invasive form of glioma. Due to glioblastoma’s extremely low survival rate of after 5 years, compared to a low grade glioma’s survival rate of over after 5 years,ref. bb0380,ref. bb0790 it is critical to distinguish the two forms for improved patient care and prognosis.
Liver Liver cancer is one of the most common causes of cancer death.ref. bb0795 In particular, hepatocellular carcinoma (HCC) is the most common type of primary liver cancer and has various subtypes, but they generally have little impact on treatment.ref. bb0800 Histogolical grade is divided into nuclear features and differentiation, which directly correlate to tumour size, presentation, and metastatic rate.ref. bb0805,ref. bb0810 Notably, high-grade dysplastic nodules are included in synoptic reports for HCC but are difficult to assess and have high inter-observer disagreement,ref. bb0815 and thus is an area where CAD systems could be leveraged to normalize assessments. Current grading of this cancer suffers from an unsatisfactory level of standardization,ref. bb0820 likely due to the diversity and complexity of the tissue. This could explain why relatively low number of works are dedicated to liver disease diagnosis and prognosis. Instead, most works focus on the segmentation of cancerous tissues.
Lymph Nodes There are hundreds of lymph nodes in the human body that contain immune cells capable of fighting infections. Cancer manifests in lymph nodes in two ways: 1) cancer that originates in the lymph node itself known as lymphoma and 2) cancer cells from different origins that invade lymph nodes.ref. bb0825 As mentioned in the prior organ sections, lymphocytic infiltration is correlated with cancer recurrence on multiple organs and lymph nodes are the most common site for metastasis. The generalizable impact to multiple organs and importance of detecting lymphocytic infiltration is why many works focused on lymph nodes address metastasis detection.ref. bb0830
Organ Agnostic The remaining papers focus on segmentation, diagnosis, and prognosis tasks that attempt to generalize to multiple organs, or target organ agnostic applications. An interesting approach to increase the generalization capability of deep learning in histopathology is proposed in.ref. bb0835 Currently, publicly available datasets with thorough histological tissue type annotations are organ specific or disease specific and thus constrain the generalizability of CPath research. To fill this gap, a novel dataset called Atlas of Digital Pathology (ADP) is proposed.ref. bb0835 This dataset contains multi-label patch-level annotations of Histological Tissue Types (HTTs) arranged in a hierarchical taxonomy. Through supervised training on ADP, high performance on multiple tasks is achieved even on unseen tissue types.
Data collection for CPath
One of the first steps in the workflow for any CPath research is the collection of a representative dataset. This procedure often requires large volumes of data that should be annotated with ground-truth labels for further analysis.ref. bb0310,ref. bb0320,ref. bb0840 However, creating a meaningful dataset with corresponding annotations is a significant challenge faced in the CPath community.ref. bb0310,ref. bb0320,ref. 168, ref. 169, ref. 170
This section outlines the entire process of the data-centric design approach in CPath, including tissue slide preparation and WSI scanning–the first two stages in the proposed workflow shown in Fig. 1. Additionally, the trend in dataset compilation across the 700 papers surveyed is discussed regarding dataset sizes, public availability, and annotation types; see Table 9.11 in the Supplementary Material for information regarding the derivations and investigation of said trends.
Tissue slide preparation
For the application development stages in CPath, the creation of a new WSI dataset must begin with selection of relevant glass slides. High quality WSIs are required for effective analysis, however, considerations must be made for potential slide artifacts and variations inherently present. As described in Section 2.1, pathology samples are categorized as either biopsy or resection samples, with most samples being prepared as permanent samples and some intra-operative resection samples being prepared as frozen samples.
Variations and Irregularities Throughout the slide sectioning process, artifacts and irregularities can occur which reduce the slide quality, including: uncovered portions, air bubbles in between the glass seal, tissue chatter artifacts, tissue folding and tears, ink markings present on the slide, and dirt, debris, microorganisms, or cross-contamination of slides by unrelated tissue from other organs.ref. 171, ref. 172, ref. 173 Frozen sections can present unique irregularities and variations, such as freezing artifacts, cracks in the tissue specimen block, or delay of fixation causing drying artifacts.ref. bb0870,ref. bb0875 Beyond these irregularities, glass slides may vary in stain colouring, occurring due to differences in slide thickness, tissue thickness, fixation, tissue processing schedule, patient variation, stain variation, and lab variation.ref. bb0870,ref. 176, ref. 177, ref. 178, ref. 179, ref. 180
All such defects and variations are important to keep in mind when selecting glass slides for the development and application process in CPath, as they can both reduce the quality of the WSI as well as impact the performance of developed CAD tools trained with these WSIs.ref. bb0855,ref. bb0860,ref. bb0885 A more detailed discussion on the surveyed works in CPath which seek to identify and correct issues in slide artifacts and colour variation in WSIs is found in Section 3.2. However, prior to digitization, artifacts, and irregularities can be kept at a minimum by following good pathology practices. While an in-depth discussion of this topic is outside the scope of this paper, some research provides an extensive list of recommendations for reducing such errors in slide sectioning.ref. bb0865
Whole slide imaging (WSI)
WSI Scan Once a glass slide is prepared, it must be digitized into a WSI. The digitization and processing workflow for WSIs can be summarized as a four-step processref. bb0905: (1) Image acquisition via scanning; (2) Image storage; (3) Image editing and annotation; (4) Image display.ref. bb0910 As the first two steps of the digitization workflow are the most relevant for WSI collection and with regards to the CPath workflow, they are discussed to a greater extent below.
Slide scanning is carried out through a dedicated slide scanner device. A plethora of such devices currently exist or are in development; see Appendix Table 1 for a collection of commercially available WSI scanners. Additionally, some research has investigated and compared the capabilities and performances of various WSI scanners.ref. 183, ref. 184, ref. 185, ref. 186
In order to produce a WSI that is in focus, which is especially important for CPath works, appropriate focal points must be chosen across the slide either using a depth map or by selecting arbitrarily spaced tiles in a subset.ref. bb0935 Once focal points are chosen, the image is scanned by capturing tiles or linear scans of the image, which are stitched together to form the full image.ref. bb0900,ref. bb0935 Slides can be scanned at various magnification levels depending on the downstream task and analysis required, with the vast majority being scanned at (/pixel) or (/pixel) magnification.ref. bb0900
WSI Storage and Standards WSIs are in giga-pixel dimension format.ref. bb0150,ref. bb0940 For instance a tissue in size scanned /pixel resolution can produce a GB image (uncompressed) with pixels. Due to this large size, hardware constraints may not support viewing entire WSIs at full resolution, thus WSIs are most often stored in a tiled format so only the viewed portion of the image (tile) is loaded into memory and rendered.ref. bb0945 When building CAD tools for CPath, this large WSI dimensionality must be taken into account in determining how much compute is required to analyze a WSI. Alongside the WSI, metadata regarding patient, tissue specimen, scanner, and WSI information is stored for reference.ref. bb0150,ref. bb0940,ref. bb0950 Due to their clinical use, it is important to develop effective storage solutions for WSI images and metadata, allowing for robust data management, querying of WSIs, and efficient data retrieval.ref. bb0955,ref. bb0960 Further details on WSI image formats and storage methods are discussed in Appendix A.6.
To develop CPath CAD tools in a widespread and general manner, a standardized format for WSIs and their corresponding metadata is essential.ref. bb0940 However, there is a general lack of standardization for WSI formats outputted by various scanners, as shown in Table 1, especially regarding metadata storage. The Digital Imaging and Communications in Medicine (DICOM) standard provides a format for CPath image formatting and data management through Supplement 145,ref. bb0950,ref. bb0965 and has been shown in research to allow for efficient access and interoperability of data between varying medical centers and devices.ref. bb0940 However, few scanners are DICOM-compliant and thus there are challenges to using different models of scanners, thus different image formats and metadata structures, in the context of dataset aggregation and processing.
Table 1: The following table lists commercially available WSI Scanners grouped by manufacturing company and their respective available compression slide formats.
| Company: Scanner Model (Slide Format) |
| Leica Biosystems: Aperio AT2 / CS2 / GT450 (TIFF (SVS)) |
| Hamamatsu: Nanozoomer SQ / S60 / S360 / S210 (JPEG) |
| F. Hoffmann-La Roche AG: Ventana DP200 / iScan HT / iScan Coreo (BIF, TIFF, JPG2000, DICOM) |
| Huron Digital Pathology: TissueScope IQ / LE / LE120 (BigTIFF, DICOM compliant) |
| Philips: Ultra-Fast Scanner(iSyntax Philips proprietary file) |
| 3DHistech: Pannoramic Series (MRXS, JPG, JPG2000) |
| Mikroscan Technologies: SL5 (TIFF) |
| Olympus: SL5 (JPEG, vsi, TIFF) |
| Somagen Diagnostics: Sakura VisionTek (BigTIFF, TIFF, JPG2000) |
| Akoya Biosciences: Vectra Polaris (JPEG, single-layer TIFF, BMP, or PNG) |
| Meyer Instruments: EASYSCAN PRO 6 (SVS, MDS, JPEG, JPEG2000) |
| Kfbio: KF-PRO (JPEG, JPEG2000, BMP, TIFF) |
| Motic: EasyScan Pro (JPEG, JPEG2000, Aperio Compatible) |
| Precipoint: PreciPoint O8 (GTIF) |
| Zeiss: Zeiss Axio (Not specified) |
| Objective Imaging: Glissando (SVS, BigTIFF) |
| Microvisioneer: manualWSI (Not specified) |
Apart from storage format, a general framework for storing and distributing WSIs is also an important pillar for CPath. In other medical imaging fields such as radiology, images are often stored in a picture archiving and communications systems (PACS) in a standardized DICOM format, with DICOM storage and retrieval protocols to interface with other systems.ref. bb0945 The need for standardization persists in pathology for WSI storage solutions; few works have proposed solutions to incorporate DICOM-based WSIs in a PACS, although some research has successfully implemented a WSI PACS consistent using the DICOM standard using a web-based service for viewing and image querying.ref. bb0945
WSI Defects and Variations Certain aspects of the slide scanning process can introduce unfavorable irregularities and variations.ref. bb0970 A major source of defects is out-of-focus regions in a generated WSI; often caused by glass slide artifacts, such as air bubbles and tissue folds, which interfere with selection of focus points for a slide.ref. bb0855,ref. bb0975 Out-of-focus regions degrade WSI quality and are detrimental to the performance of CAD tools developed with these WSIs, presenting concerns for performance with studies showing high false-positive errors.ref. bb0980,ref. bb0860 Additionally, as WSIs are scanned in strips or tiles, any misalignment between sections can introduce striping/stitching errors in the final image.ref. bb0985 Another source of error may appear during tissue-background segmentation where the scanner may misidentify some tissue regions as background, potentially missing crucial tissue areas on the glass slide from being digitized.ref. bb0990
Variations in staining refers to differences in colour and contrast of the tissue structures in the final WSI occurring due to differences in the staining process, staining chemicals, and tissue state. Variations in colour can lead to difficulty in generalizing CAD tools to WSIs from different labs, institutions, and settings.ref. bb0995,ref. bb1000 Even identical staining techniques can yield different WSIs due to scanner differences in sensor design, light source and calibration,ref. bb0900,ref. bb1005 creating challenges for cross-laboratory dataset generation. These additional sources of variation add layers of complexity to the WSI processing workflow, and must be kept in mind during slide selection and dataset curation for CAD tool development and deployment.
Addressing Irregularities and Variations Much work has gone into identifying areas of irregularities within WSIs, most notably blur and tissue fold detection.ref. bb0975,ref. bb0980 Some research has explored automated deep learning tools to identify these irregularities at a more efficient pace than manual inspection.ref. bb0975,ref. bb0980 Developing techniques for addressing staining variation has also been a significant research arearef. bb0885,ref. 202, ref. 203, ref. 204, ref. 205, ref. 206, ref. 207 as the use of techniques addressing stain variation is important for all future works. We list some computational approaches proposed to address these issues: An example method proposed inref. bb1010 uses a stain normalization technique, attempting to map the original WSI onto a target color profile. In this technique, a color deconvolution matrix is estimated to convert each image to a target hematoxylin and eosin (HE) color space and each image is normalized to a target image colour profile through spline interpolation.ref. bb1010 A second approach applies color normalization using the H channel with a threshold on the H channel on a Lymphocyte Detection dataset.ref. bb1025 Recent studies have shown promise in having deep neural networks accomplish the stain normalization in contrast to the previous classical approaches,ref. bb1015,ref. 208, ref. 209, ref. 210 commonly applying generative models such as generative adversarial networks (GANs) to stain normalization. Furthermore, Histogram Equalization (HE) technique for contrast enhancement is used in,ref. bb1055 where novel preprocessing technique is proposed to select and enhance a portion of the images instead of the whole dataset, resulting in improved performance and computational efficiency.
An alternative approach to address the impact of stain variation on training CAD tools is data augmentation. Such methods augment the data with duplicates of the original data, containing adjustments to the color channels of the image, creating images of varying stain coloration, and training train models that are accustomed to stain variations.ref. bb1000 This method has been frequently used as a pre-processing step in the development of training datasets for deep learning.ref. 212, ref. 213, ref. 214 A form of medically-irrelevant data augmentation based on random style transfer, called STRAP, was proposed by researchers and outperformed stain normalization.ref. bb1030 Similar to style transfer,ref. bb1075 proposes stain transfer which allows one to virtually generate multiple types of staining from a single stained WSI.
Cohort selection, scale, and challenges
The data used to create/train CPath CAD tools can greatly impact the performance and success of the tool. Curating the ideal dataset, and thus selecting the ideal set of WSIs for the development of a CAD tool is a nontrivial task. Several works suggest that datasets for deep learning in CPath should include a large quantity of data with a degree of variation and artifacts in the WSIs.ref. bb0310,ref. bb0860 Some works also recommend the inclusion of difficult or rarely diagnosed cases; other works indicate that inclusion of extremely difficult cases may decrease the performance of advanced models.ref. bb0860,ref. bb1115
A study highlighting the results of the 2016 Dataset Session at the first annual Conference on Machine Intelligence in Medical Imaging outlines several key attributes to create an ideal medical imaging dataset,ref. bb1120 including: having a large amount of data to achieve high performance on the desired task, quality ground truth annotations, and being reusable for further efforts in the field. While the scope of this conference did not include CPath, many of the points made regarding medical imaging datasets are also relevant to the development of CPath datasets. The session also outlined the impact that class imbalances can have on ML models, an issue also prevalent in CPath as healthy or benign regions often outnumber diseased regions by a significant margin.ref. bb1125
Our survey of past works in the literature reveals some trends in CPath datasets. Currently, the majority of datasets presented in the literature for CAD tool development are small-scale datasets,ref. bb0310 using a small number of images, and/or images from a small number of pathology laboratories. Examples of these smaller datasets include a dataset with 596 WSIs (401 training, 195 testing) from four centres for breast cancer detectionref. bb1130 and the Breast Cancer Histology (BACH2018) dataset, which has 500 ROI images (400 training, 100 testing) and 40 WSIs (30 training, 10 testing).ref. bb1135 Although curating a dataset from fewer pathology laboratories may be simpler, these smaller scale datasets may not be able to effectively generalize to data from other pathology centres.ref. bb0995,ref. bb0600 An example of this can be seen in which data from different pathology centres are clustered disjointly in a t-distributed stochastic neighbor embedding (t-SNE) representation demonstrated in.ref. bb0860 Another alternative was proposed inref. bb1140: using a swarm learning technique multiple AI models were trained on different small data sets separately and then unified into one central model.
Additionally, stain variations, slide artifacts, and variation of disease prevalence may sufficiently shift the feature space such that a deep learning model may not sustain high performance on unseen data in new settings.ref. bb0600,ref. bb1145 As artifacts in WSIs are inevitable, with some artifacts, such as ink mark-up on glass slides, being an important part of the pathology workflow,ref. bb1150 the ability of CAD tools to become robust to these artifacts through exposure to a diverse set of images is an important consideration.
Compared to the number of studies conducted on small-scale datasets, relatively fewer studies have been performed using large-scale, multi-centre datasets.ref. bb0310,ref. bb1155,ref. bb0860 One study uses over 44,715 WSIs from three organ types, with very little curation of the WSIs for multi-instance learning detailed in.ref. bb0310 Stomach and colon epithelial tumors were classified using 8,164 WSIs in.ref. bb1155 A similar study uses 13,537 WSIs from three laboratories to test a machine learning model trained on 5,070 WSIs and achieves high performance.ref. bb0310
Despite some advancements, there exist major barriers to using such large, multi-centre datasets in CAD development. Notably, for strongly supervised methods of learning, an immense amount of time is needed to acquire granular ground truth annotations on a large amount of data.ref. bb1155 To combat this, some researchers have implemented weakly-supervised learning by harvesting existing slide level annotations to forego the need for further annotation.ref. bb0310 Additionally, it may be difficult to aggregate data from multiple pathology centres due to regulatory, privacy, and attribution concerns, despite the improvements that diverse datasets offer. Section 5 discusses model architectures and training techniques that harness curated datasets of various annotation levels.
Dataset Availability In general computer vision, progress can be tracked by the increasing size and availability of datasets used to train models, e.g. ImageNet grew from 3.2 million images and 5000 classes in 2009 to 14 million images and 21,000 classes in 2021.ref. bb1160 We infer a similar trend in dataset growth and availability indicates progress in CPath. In our survey of over 700 CPath papers, we determine the current landscape by noting the dataset(s) used in work, along with dataset details such as the organ(s) of interest, annotation level, and stain type, tabulating the results into Table 9.11 of the supplementary materials, with summarized findings from Table 9.11 are shown in Fig. 6.
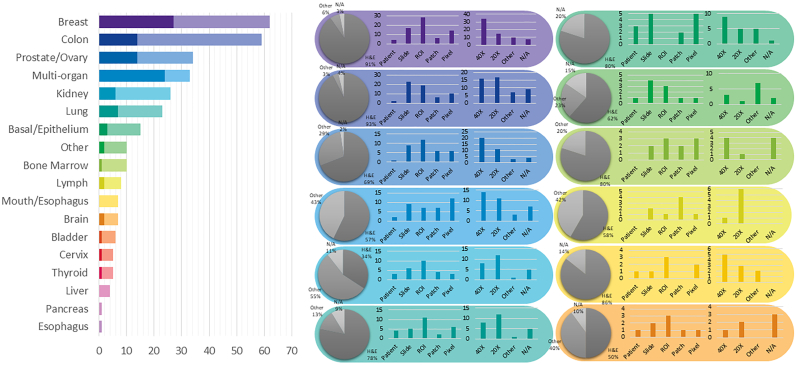
From Fig. 6 we can clearly see that the majority of datasets used for research developments in computational pathology are privately sourced or require additional registration/request. With organs represented in a small number of datasets, such as the liver, thyroid, brain, etc, having a smaller proportion of freely accessible datasets as compared to the Breast, Colon, or Porstate. This can be problematic when trying to create CAD tools for cancers in these organs due to a lack of accessible data. We additionally note that although data sets requiring registration/request for access can be easily accessible, as in the case of Breast Cancer Histopathological Database (BreakHis)ref. bb1165 being used in multiple works,ref. 227, ref. 228, ref. 229 the need for registration presents a barrier to access as requests may go unanswered or take much time to review.
In our categorization of CPath datasets, we find that a few prominent datasets have been released publicly for use by the research community. Many such datasets are made available through grand challenges in computational pathology,ref. bb1185 such as the CAMELYON16 and CAMELYON17 challenges for breast lymph node metastases detection,ref. bb0305,ref. bb1190,ref. bb1195 and the Gland Segmentation in Colon Histology Images Challenge (GLaS) competition for colon gland segmentation in conjunction with Medical Image Computing and Computer Assisted Intervention (MICCAI) 2015.ref. bb1200,ref. bb1205 Notable amongst publicly available data repositories is the cancer genome atlas (TCGA),ref. bb1210 a very large-scale repository of WSI-data containing many organs and diseases, along with covering a variety of stain types, magnification levels, and scanners. Data collected from TCGA has been used in a large number of works in the literature for the development of CAD tools.ref. bb1000,ref. bb1215,ref. bb1220 As such, TCGA represents an essential repository for the development of computational pathology. While patient confidentiality is a general concern when compiling and releasing a CPath dataset, large-scale databases such as TCGA prove that it is possible to provide relatively unrestricted data access without compromising patient confidentiality. Further evaluating public source datasets, it seems that the majority of them use data extracted from large data repositories, such as TCGA, without specifying the IDs of the images used, which provides a challenge in comparing datasets or CAD tool performance across works. However, there are a few datasets that are exceptions to that phenomenon.ref. bb0320,ref. 238, ref. 239, ref. 240
Fig. 6 also provides some insights on the dataset breakdown by organ, stain type, and annotation level. Per organ, it can be seen that the breast, colon, prostate/ovary, and lung tissue datasets are amongst the most common, understandably since cancer occurrence in these regions is the most frequentref. bb0450–complying with cancer statistics findings in 9.5. Multi-organ datasets are the other most common type, where we have designated a dataset to be multi-organ if it compiles WSIs from several different organs. To note, multi-organ datasets are especially useful for the development of generalized image analysis tools in computational pathology. The annotation level provided in the datasets did not indicate any pattern across most organs.
Dataset Bias It is also important to note the potential for bias in datasets that may influence the ability of any deep learning algorithm to generalize on unseen data.ref. bb1240,ref. bb1245 This problem is a prevalent issue in general machine learning applications,ref. 243, ref. 244, ref. 245, ref. 246 and CPath is not immune to it. The survey review inref. bb1270 reviews a large number of other examples in machine learning that exhibit such bias, both from a dataset-standpoint and an algorithm-standpoint.
Such a lack of generalizability in CPath can impact the ability of machine learning models trained on biased data to meet the needs of patients. As noted in,ref. bb1240 minority groups may be disproportionately negatively impacted if care is not taken in curating a diverse dataset that adequately reflects the relevant demographics for the problem to be solved.
Several works have delved into the issue of dataset bias in CPath specifically.ref. bb1275,ref. bb1280 A notable example is in,ref. bb1280 where the study was able to demonstrate that deep learning models trained on WSIs from TCGA were able to infer the organization that contributed the slide sample. Notably, some features, such as genetic ancestry, patient prognosis, and several key genomic markers were significantly correlated with the site the WSI was provided from. As the vast majority of data in TCGA is acquired from 24 origin centers,ref. bb1275 such site-specific factors may impact the ability of a DL model to perform well on patient data from different sites.
As discussed previously, having a large set of diverse data may help to mitigate generalization issues.ref. bb0600,ref. bb0995,ref. bb1240 Additionally, the studyref. bb1280 makes the suggestion that training data should be from separate sites than validation data, and that per-site performance of a model should be reported when validating a model. In doing this, the robustness of the model to site-specific variation, including both stain and demographic related variation, can be evaluated.
Domain expert knowledge annotation
A primary goal of CPath is to capture and distill domain expert knowledge, in this case the expertise of pathologists, into increasingly efficient and accurate CAD tools to aid pathologists everywhere. Much of the domain knowledge transfer is encompassed within the process of human experts, in this case pathologists, generating diagnostically-relevant annotations and labels for WSIs. It must be emphasized, that without some level of label, a WSI dataset is not directly usable to train a model for most CAD tasks that involve the generation of diagnoses, prognoses, or suggestions for pathologists. Thus, the process of obtaining and/or using annotations at the appropriate granularity and quality is paramount in the field. This section focuses on describing various types of ground-truth annotation to cover the spectrum of weak to strong supervision of labels, discussing the practicality of labeling across this supervision spectrum, and how a labeling workflow can be potentially designed to optimize related annotation tasks.
Supervised annotation
In contrast to general computer vision, computer scientists do not have expert-level knowledge of histopathology and thus they are not as efficient at generating annotations or labels of pathology images. Further, labels cannot be easily obtained by outsourcing the task to the general public. As a result, pathologists must be leveraged to obtain labels at some stage of the data collection and curation process, and in many annotation pipelines the first step involves recruiting the help of pathologists for their expertise in labelling.
Obtaining Expert Domain Knowledge The knowledge of pathologists is essential in the development of accurate ground truth annotations–a process most commonly completed by encircling ROI.ref. bb1130 However, there are studied instances of inter-observer variance between pathologists when determining a diagnosis.ref. 250, ref. 251, ref. 252 As obtaining the most correct label is essential when training a model for CAD, this issue must be addressed and a review of the data by several pathologists can result in higher quality ground truth data as compared to that of a single pathologist. As a result, most datasets are curated by involving a group of pathologists in the annotation process. If there exists a disagreement between the expert pathologists on the annotation of a ground truth, one of several methods is usually employed to rectify the discrepancy. A consensus can be reached on the annotation label through discussion amongst pathologists, as is done in the triple negative breast cancer (TNBC-CI) dataset,ref. bb1300 the Breast Cancer Surveillance Consortium (BCSC) datasetref. bb1305 and the minimalist histopathology image analysis dataset (MHIST) dataset.ref. bb1310 Alternatively, images, where disagreements occur, can be discarded, as is done in some works.ref. bb1315,ref. bb1320 Further, the disagreement between annotators can be recorded to determine the difficulty level of the images, as is done in the MHIST dataset.ref. bb1115 This extra metadata aids in the development of CAD tools for analysis.
Pathologists can also be involved indirectly in dataset annotation. Both the Multi-organ nuclear segmentation dataset (MoNuSeg)ref. bb1325 and ADPref. bb0835 have non-expert labelers annotate their respective datasets. A board-certified pathologist is then tasked with reviewing the annotations for correctness. Alternatively, some researchers have employed a pathologist in performing quality control on WSIs for curating a high-quality dataset with minimal artifacts.ref. bb0435,ref. bb1330 To enable the large scale collection of accurate annotated data, Lizardref. bb1335 was developed using a multi-stage pipeline with several significant “pathologist-in-the-loop” refinement steps.
Existing pathological reports, along with the metadata that comes from public large-scale databases like TCGA, can also be leveraged as additional sources of task-dependent annotations without the use of further annotation. For example, TCGA metadata was used to identify desirable slides in,ref. bb0130 while pathological diagnostic reports were used for breast ductal carcinoma in-situ grading in.ref. bb1340
To note, there are some tasks where manual annotation by pathologists can be bypassed altogether. For instance, IHC was applied to generate mitosis figure labels using a Phospho-Histone H3 (PHH3) slide-restaining approach in,ref. bb1345 while immunofluorescence staining was used as annotations to identify nuclei associated with pancreatic adenocarcinoma.ref. bb1350 These works parallel the techniques that pathologists often use in clinical practice, such as the use of IHC staining as a supplement to HE stained slides for difficult to diagnose cases.ref. bb1355 They demonstrate high performance on their respective tasks wherein the top-performing models on the Tumor Proliferation Assessment Challenge 2016 (TUPAC16)ref. bb1360 dataset were achieved.ref. bb1345 Importantly, these techniques still utilize supervision, albeit weakly, by leveraging lab techniques that have been developed and refined to identify the desired regions visually.
Ground-Truth Diagnostic Information Understanding different annotation levels and their impact on the procedural development of ML pipelines is an important step in solving tasks within CPath. There are five possible levels of annotation, in order of increasing granularity (from weakly-supervised to fully-supervised): patient, slide, ROI, patch, and pixel. Fig. 7 overviews the benefits and limitations of each level. For additional information regarding each annotation level please refer to Appendix A.8.
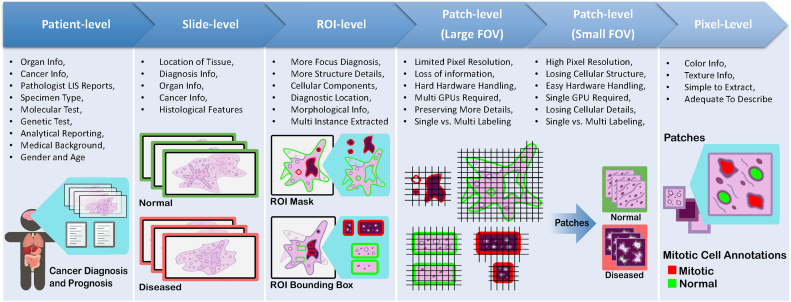
Picking the Annotation Level Selecting an annotation level depends largely on the specific CPath task being addressed, as shown in Fig. 8. For example, segmentation tasks tend to favor pixel-level annotations as they require precise delineation of a nucleus or tissue ROI. Conversely, disease diagnosis tends to favor datasets with ROI-level annotations, as diagnosis tasks are predominantly associated with the classification of diseased tissue, the higher-level annotations may provide a sufficient level of detail and context for this task.ref. bb1365
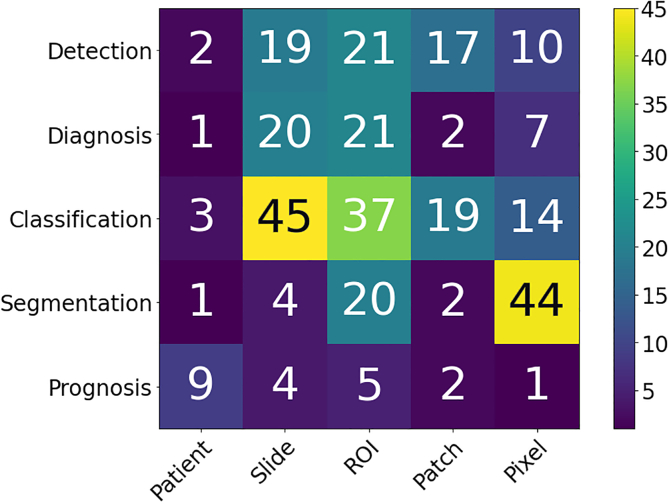
Fig. 8 shows that tasks that use stronger supervision are more likely to be used in CAD tool model development. However, due to the high cost of pixel-level annotation, fully supervised annotations are challenging to develop. Even patch-based annotations often require the division and analysis of a WSI into many small individual sub-images resulting in a similar problem to pixel-based annotations.ref. bb0315,ref. bb1060 In contrast, WSI data is most often available with an accompanying slide-level pathology report regarding diagnosis thus making such weakly labeled information at the WSI level significantly more abundant than ROI, patches, or pixel-level data.ref. bb1370,ref. bb1375 Different levels of annotation can be leveraged together, as demonstrated by a framework to use both pixel and slide level annotations to generate pseudo labels in.ref. bb1380 Additionally, it is common in CPath to further annotate the slide-level WSIs on an ROI or patch level structure.ref. bb0160,ref. bb0425,ref. bb1385,ref. bb1390
Active Learning Tools Active learning annotation tools bridge the gap between the need for highly supervised labels and the current abundance of less informative annotations. Such works seek to ease the annotation process by using computational approaches to assist the human annotator. For example, in,ref. bb0840 a platform was developed for creating nuclei and gland segmentation ground truth labels quickly and efficiently. A convolutional neural network (CNN), trained on similar cohort data, was used to segment nuclei and glands with different mouse actions.ref. bb0840 Alternatively, Awan et al.ref. bb0845 presented the HistoMaprTM platform to assist in diagnosis and ground truth data collection. Through this tool, a pathologist selects one of several proposed classes for each given ROI, thus mitigating the need for hand-drawn annotations or manual textual input.ref. bb0845 Similarly, an active learning model called the Human-Augmenting Labeling System (HALS)ref. bb1395 was developed to increase data efficiency by guiding annotators to more informative slide sections. Quick Annotator (QA)ref. bb1400 is another tool which provides an easy-to-use online interface for annotating ROIs and was designed to create a significant improvement in the annotation efficiency of histological structures by several orders of magnitude. There are other active learning annotation tools proposed for different applications in computer vision that can be investigated for use in the pathology datasets. Such examples include methods to produce object segmentation masks for still imagesref. bb1405,ref. bb1410 as well as video.ref. bb1415 One notable example is DatasetGANref. bb1410; the model is proposed as a training data creator, and it is shown that the model can produce segmentation masks with a small number of labelled images in the training data. While these systems are for general computer vision, they may be adoptable in computational pathology, and would facilitate the necessary relationship between pathologists and computer scientists in the development of CAD tools. As such, they may prove to be a valuable contributor to the CAD system development workflow.
Tissue-Class and Disease Complexity Much of the current CPath research operates under the umbrella of supervised learning tasks, and correspondingly uses labeled data to develop automated CAD tools. We refer to supervised learning to include a diverse spectrum of annotation i.e. weak-supervision (e.g. patient-level) all the way to strong-supervision (e.g. pixel-level). Classes within a dataset can be task-dependent, for example as shown in Table 9.11 of the supplementary material, datasets primarily used for segmentation such as MoNuSegref. bb1325 and CPM-17ref. bb1420 have classes for each annotated pixel indicating the presence or absence of nuclei. However, classes need not be task-dependent; datasets such as CAMELYON16ref. bb1190 outline metastases present in WSIs that can be used for a variety of applications, including disease detectionref. bb1190 and segmentation tasks.ref. bb1425
The current paradigm for dataset compilation in computational pathology, particularly for disease detection and diagnosis, treats different disease tissue types as separate independent classes. For example, BreakHis divides all data into benign/malignant breast tumours.ref. bb1165 At the ROI level, GLaS divides colon tissue into five classes: healthy, adenomatous, moderately differentiated, moderately-to-poorly differentiated, and poorly differentiated.ref. bb1200 So far, this approach to class categorization has resulted in high-performing CAD tools.ref. bb0400,ref. bb0415,ref. bb0420,ref. bb1175,ref. bb1180,ref. bb1430 However, the treatment of different disease tissue types as an independent class is perceived differently in computer vision domain where the representation learning of normal objects is done differently compared to anomalies. A similar synergy can be found by differentiating healthy tissue classes from diseased ones and one should be mindful about defining meaningful tissue ontology for annotation and labeling.
Optimum labeling workflow design
This section focuses on the steps required for compilation of a CPath dataset which is broken into three main sub-tasks: Data Acquisition, Data Annotation, and Data Management, as per Fig. 9. Each sub-task is discussed below with reference to its individual components in the hierarchical structure in Fig. 9.
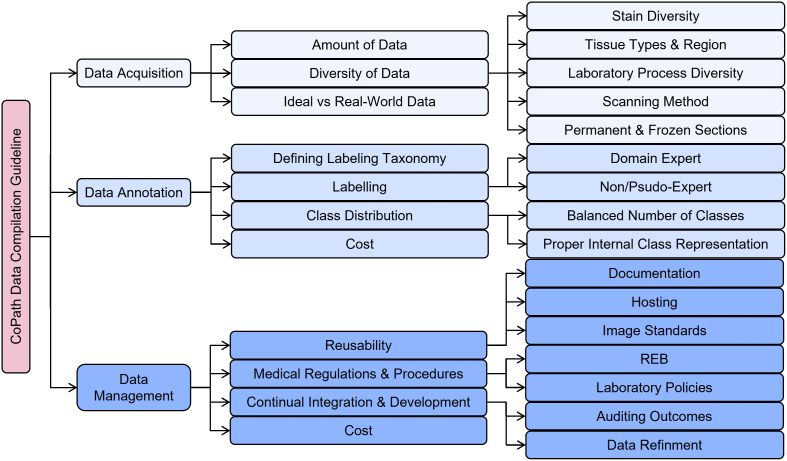
Data Acquisition Database compilation starts from data acquisition. When collecting data, it is vital that there are large amounts of data,ref. bb1435 along with having sufficient diversity.ref. bb0310,ref. bb0860 Specifically, diversity in CPath data occurs in multiple ways, such as staining methods, tissue types and regions, laboratory processes, and digital scanners. We advise that CPath researchers consult expert pathologists on the diversity of data required for various tasks. Ideally, all the data acquired in pathology should be perfect without any irregularity and artifacts. However, some level of artifacts and irregularity are unavoidable and introducing realistic artifacts that are representative of real-world scenarios into the data increases the robustness and generalizability of CAD tools.
Data Annotation After collecting sufficient data, the next task is annotation of the data. Data annotation is a costly process in both time and money, thus a budget and schedule should always be established when generating labels. There are often various approaches for annotating different structures,ref. bb1440 so a specific labelling taxonomy should be defined a priori. As mentioned previously, annotation should involve expert pathologists due to the domain knowledge requirement and importance of label correctness. A table of commonly used commercially available annotation software for annotating different slide formats are show in Table 2, along with their compatible image formats which is important to note when trying to build compatible and accessible datasets.
Table 2: Commercially available annotation software along with their manufacturing company and available input slide formats.
| Company: Annotation Tool (Input Format) |
| Leica Biosystems: Aperio eSlide Manage (JFIF, JPEG2000, PMM) |
| Pathcore: Sedeen Viewer (Aperio SVS, Leica SVN, TIFF, JPEG2000) |
| Indica: Halo (TIFF/SVS) |
| Objective Pathology: MyObjective (Scanner-wide compatibility) |
| ASAP: ASAP (Multiple formats through OpenSlide) |
| SiliconLotus: SiliconLotus (Not specified) |
| Augmentiqs: Annotation Software Suite (Not specified) |
| QuPath: QuPath (Multiple formats, Bio-formats and OpenSlide) |
| Proscia: Concentriq (Not specified) |
| Visiopharm A/S: VisioPharm (Not specified) |
| Hamamatsu: NDP (JPEG) |
| Roche: Ventana Companion Image Analysis (BIF, TIFF, JPG2000, DICOM compliant) |
| Huron: HuronViewer (BigTIFF, FlatTIFF, DICOM compliant) |
| Philips: Intellisite (iSyntax Philips proprietary file) |
| 3DHistech: CaseViewer (JPG, PNG, BMP, TIFF) |
| AnnotatorJref. bb1380: AnnotatorJ (JPG, PNG, TIFF) |
| NuClickref. bb1770: NuClick (Not specified) |
Once the ontology of class-definitions is defined (in collaboration with expert pathologists), there will be two ways to generate labels or annotations in general: domain expert labelling or non-expert labelling. The domain expert labelling refers to having pathologists annotate data that they are specialized at, which is labor-extensive. On the other hand, non-expert labelling can use crowdsourcing techniques to generate weak labels or have non-experts, such as junior pathologists or students, label the data. This process is cheaper and quicker, but it may be harder to maintain the same level of quality as domain expert labeling.ref. bb1440 Regardless of the labelling methodology used, labels generated from both should be validated. Finally, to determine the sufficiency of label quantity, one should consider the balance between the number of classes, representation size of each class, and complexity of class representation. Techniques from active-learning could be also leveraged to compensate for lack of resource management as well as maintain the quality of labeling as discussed above.
Data Management Data management is an important aspect of any dataset creation process, and is the one that is most likely to be overlooked. Proper data management should have considerations for reusability, medical regulations/procedures, and continuous integration and development.
Reusability can be broken down into detailed documentation of the data, accessible and robust hosting of data, and consideration for image standards. Poor cross-organizational documentation can lead to missing metadata, ultimately resulting in discarding entire datasets.ref. bb1445 Adherence to an established image standard, such as DICOM, can help resolve some of these issues in reusability. Medical regulations/procedures can be broken down into the construction of a Research Ethics Board (REB) and proper consideration for whom is curating the data. Through incentives for data excellence for medical practitioners, the issue of misaligned priorities between data scientists, domain experts, and field partners can be resolved.ref. bb1445 To ensure that models used on actual patients remain relevant and hidden errors do not propagate, continuous integration/development (CI/CD) must be implemented. These systems must include at least two components, a method to audit predictions from the model, as well as a way to refine the training data accounting for discrepancies found through auditing. Several algorithms deployed in high-risk environments, including medical diagnosis, proved to only work well when data was updated after initial deployment.ref. bb1450,ref. bb1445 Throughout the data management process, consultation with domain experts is a vital step in ensuring the success of data compilations.ref. bb1455
Model learning for CPath
Once an application domain and corresponding dataset have been chosen, the next step of developing a CPath tool involves designing of an appropriate model and representation learning paradigm. Representation learning refers to a set of algorithmic techniques dedicated to learning feature representations of a certain data domain that can be used in downstream tasks.ref. bb1460 In CPath, the amount of data available for a given annotation level and task are the key determinants to designing a model and learning technique. The last decade has shown neural network architectures to become the dominant method in many machine-learning domains because they are rich enough to avoid handcrafted features and offer superior performance.ref. bb1465 The annotation level of the data pertaining to the task corresponds to the level of supervision for the learning technique applied. This relationship between data annotation level and learning supervision level is surveyed in Fig. 11.
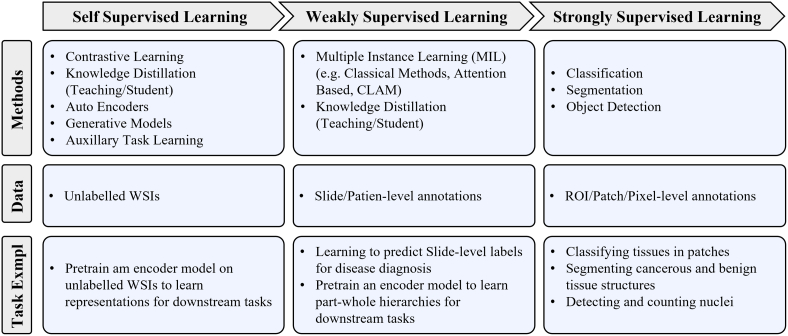
This section details the various types of models and learning techniques, along with the tasks they have been applied to in CPath. Fig. 10 highlights the most common backbone architectures used for feature encoding in SOTA research, based on the corresponding tasks. More details are provided in Table 9.11 from the supplementary materials. The selection of architectures is then compared to draw useful insights into accuracy, computational complexity, and limitations. Lastly, existing challenges in model design are investigated.
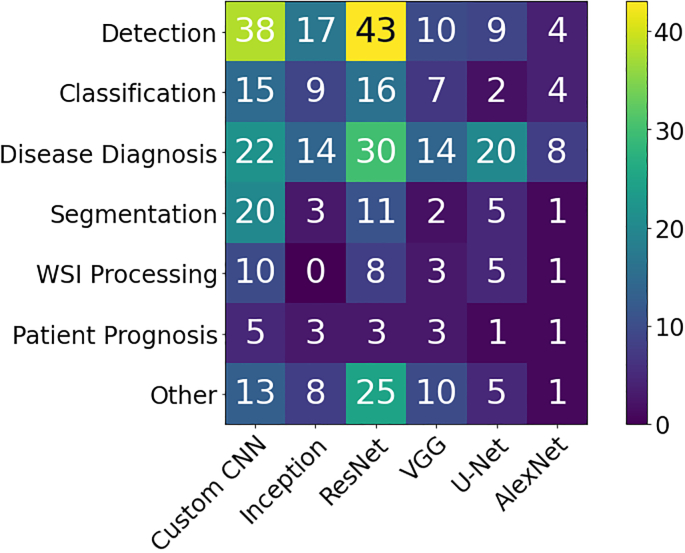
Classification architectures
In CPath, general classification architectures are the most prevalent due to their straightforward applicability to a wide range of tasks including tissue subtype classification, disease diagnosis and detection (more details in Section 2 and Fig. 4). Architectures commonly used for natural images, in particular CNNs, are widely adopted for CPath. To maximize model performance, it is a common approach to pre-train the model on large datasets like ImageNet before subsequent fine-tuning them to perform well for the specific CPath task, a task known as transfer learning.ref. bb0320,ref. bb0430,ref. bb0435,ref. bb1030,ref. bb1285,ref. bb1340,ref. bb1370,ref. 287, ref. 288, ref. 289, ref. 290, ref. 291, ref. 292, ref. 293, ref. 294, ref. 295, ref. 296, ref. 297, ref. 298, ref. 299, ref. 300, ref. 301, ref. 302, ref. 303, ref. 304, ref. 305, ref. 306, ref. 307, ref. 308, ref. 309, ref. 310, ref. 311, ref. 312 Transfer learning for CPath allows for: 1) improved generalizability, particularly for tasks with limited amount of data; and 2) improved ease in finetuning a model compared to training from scratch.ref. bb1600
Graph Convolutional Neural Networks (GCN)ref. bb1605,ref. bb1610 is an alternative architecture that can be used to improve the learning of context-aware features across the WSI. GCNs typically consist of nodes representing elements and edges defining relationships between nodes. In,ref. bb1615 a GCN was defined on a WSI, where the nodes represent patches and edges represent the connections among patches; this work obtained remarkable results on cancer prognosis task outperforming the SOTA in four out of five cancer types.ref. bb1615
Vision Transformers (ViT),ref. bb1620 have recently emerged as a direct application of Transformer modelsref. bb1625 to the image domain. In ViT, images are sub-patched and flattened into a 1D embedding along with a positional encoding which is then classified by an MLP head. By using the positional encoding, the model’s attention mechanism can focus computation on the most relevant areas of the image. ViT models have been applied with great success to CPath tasks, especially in conjunction with pre-trained CNN models.ref. bb1630 We refer the reader to a comprehensive survey of transformer methods in medical image analysis for more details.ref. bb1635
General classification architectures are also commonly used as a foundation for novel architectural designs. For example, Squeeze-and-Excitation (SE) modules were introduced to reduce the number of parameters in ResNet and DenseNet blocks while maintaining high accuracy.ref. bb1640,ref. bb1180 A fully-connected conditional random field (CRF) was incorporated on top of a CNN encoder to improve performance while maintaining the same level of computational complexity.ref. bb1645 Lastly, patch sampling and pooling were used with AlexNet to perform slide-level disease diagnosis and segmentation.ref. bb0710
Finally, in order to achieve superior performance, many researchers often rely on ensemble or multi-stage techniques which combine the predictive power or feature extraction abilities of multiple models to form a final output. These approaches have shown performance improvements compared to traditional single model classifiers.ref. 323, ref. 324, ref. 325, ref. 326, ref. 327, ref. 328 However, this often comes at the expense of higher computational requirements.
Segmentation architectures
Segmentation is widely used in CPath, as shown in Fig. 4, and enable localizing the area of interest at the pixel level.ref. bb1680 U-Net was initially developed for neuronal structure segmentation in electron microscopy image stacks,ref. bb1680 but has become one of the most common architectures for segmentation in CPath.ref. bb0760,ref. bb1025,ref. bb1035,ref. bb1220,ref. bb1400,ref. bb1425,ref. bb1555,ref. bb1650,ref. 330, ref. 331, ref. 332, ref. 333, ref. 334, ref. 335, ref. 336, ref. 337, ref. 338, ref. 339, ref. 340, ref. 341, ref. 342, ref. 343, ref. 344, ref. 345, ref. 346, ref. 347, ref. 348, ref. 349 U-Net has an encoder-decoder structure: an encoder to contract features spatially and a decoder to expand them again to capture semantically related context and generate pixel-level predictions.ref. bb1680 The U-Net model has been used to segment nuclei for creating a novel dataset with unsupervised learning,ref. bb1220 but it should be noted that this process also relies on the Mask R-CNN framework and pathologists for quality-checking purposes.
Another common approach for segmentation is to use fully convolutional networks (FCNs),ref. bb0400,ref. bb0420,ref. bb0990,ref. bb1325,ref. bb1385,ref. 349, ref. 350, ref. 351, ref. 352, ref. 353, ref. 354, ref. 355 customized architectures constructed by combining multiple components of various architectures, or introducing new components to pre-existing architectures.ref. bb0420,ref. bb0840,ref. bb1205,ref. bb1330,ref. bb1365,ref. bb1380,ref. bb1535,ref. bb1580,ref. bb1750,ref. bb1755,ref. 356, ref. 357, ref. 358, ref. 359, ref. 360, ref. 361, ref. 362, ref. 363, ref. 364, ref. 365, ref. 366, ref. 367, ref. 368, ref. 369 For example, one work used a custom CNN to predict whether each pixel was benign or malignant, while a second CNN was used to refine the initial prediction through probability fusion.ref. bb0420
Object detection architectures
In this section, we specifically focus on architectures that are used for object detection in CPath, where bounding boxes are predicted around regions of interest. A major CPath application for object detection is mitosis detection with the primary goal for counting the number of mitosis instances. To this end, a large number of studies has been dedicated to this application.ref. bb0995,ref. bb1000,ref. bb1125,ref. bb1345,ref. bb1720,ref. bb1750,ref. 370, ref. 371, ref. 372, ref. 373, ref. 374, ref. 375, ref. 376, ref. 377, ref. 378, ref. 379 Object detection has been additionally applied for nuclei,ref. bb0385,ref. 380, ref. 381, ref. 382, ref. 383, ref. 384, ref. 385 colorectal glandref. bb0400,ref. bb1740,ref. bb1965 and glomeruli detectionref. 387, ref. 388, ref. 389; however, it can also be applied to the detection of a variety of histopathological objects including tumor-infiltrating lymphocytesref. bb1985 or keratin pearls.ref. bb1855
In CPath, object detection employs a combination of pre-existing off-the-shelf architectures and customized neural networks to perform object detection tasks, as shown in Fig. 10. A model called CircleNet, which uses a deep layer aggregation network as a backbone, was proposed to detect round objects.ref. bb1975 Their approach involves using an anchor-free “center point localization” framework in order to output a heatmap with center points followed by a conversion into a bounding circle for the detection of kidney glomeruli. A multi-stage deep learning detection model based on Fast R-CNN was proposed in.ref. bb1895 First, a modified Fast R-CNN generated region proposals, then a ResNet-50 model eliminated false positives, and a Feature Pyramid Network detected mitosis in sparsely annotated images using a ResNet backbone.ref. bb1925
Multi-task learning
Multi-task models are individual models predicting for multiple tasks at once (e.g. classification and segmentation), as defined in Section 2. Multi-task learning (MTL) can be beneficial over independent task learning because sharing representations between related tasks can create more generalizable representations and encourage the task heads to make logically consistent predictions. This type of model, however, is uncommon in CPath, as it requires annotating multiple tasks for each image.ref. bb0160,ref. bb0410,ref. bb0600,ref. bb1920,ref. 391, ref. 392, ref. 393 We discuss some of these papers in further detail below.
In one work, a ResNet-50 backbone followed by independent decoders (a pyramid scene parsing network for segmentation and a fully-connected layer for classification) was used to solve 11 different tasks (4 segmentation based and 7 classification based).ref. bb0600 With significantly less computation, the MTL model achieved comparable or better results to single task learning in classification, but comparatively worse results in segmentation. Similarly, in,ref. bb0410 a ResNet-50 with two parallel branches to perform segmentation and classification, was able to achieve comparable results on both tasks through an MTL approach.
While the results are impressive, there is still work to be done in this field. One work found that model performance may be sensitive to the number and type of tasks used during training.ref. bb1990 If the tasks are unrelated, this could deteriorate the performance compared to a single-task setting. How to weigh different task objectives and select optimal tasks to be trained together is yet an active area of research.ref. bb2005,ref. bb2010 MTL represents an interesting field of research in CPath as it may reduce the necessity to train multiple deep neural networks to perform different tasks.
Multi-modal learning
As opposed to multi-task networks where multiple tasks are learned simultaneously, the multi-modal approach involves using network input features from multiple domains/modalities at once.ref. bb2015 In the case of CPath, modalities can be represented as pathologists’ reports, gene expression data, or even WSI images. Most commonly, immunohistochemistry (IHC) stains alongside the H&E stain to better visualize specific proteins.ref. 397, ref. 398, ref. 399 As a result, models can learn better unified/shared latent representations which capture correlations from multiple indicators, since some information may not be captured by individual indicators.ref. bb2035 This approach can be viewed as adding hand-crafted features to boost performance. While the use of deep learning normally implies using learned features to replace hand-crafted ones, using hand-crafted features can nonetheless improve performance compared to strictly deep learning approaches when data is limited.ref. bb2040 Indeed, many works have obtained best performance by combining manual and learned features.ref. bb1890,ref. bb1950,ref. bb2045 This was demonstrated in the case of mitotic cell classification when an ensembled classifier model using hand-crafted features set a new record for the MITOS-ATYPIA 2014 challenge with an F-score of .ref. bb2050 However, where data is plentiful, CNNs alone can outperform all other hand-crafted features. In the same MITOS-ATYPIA 2014 challenge, the previous record was broken this way with a new F-score of .ref. bb2055 Although one cannot compare these two works directly as they use different classifier heads and dataset balancing methods, one can argue that the optimal choice of approaches from deep learning, classical ML, and different modalities should depend on the situation. Multi-modal approaches are gaining traction in CPath for specific problems, especially where useful additional data is available.ref. bb2060,ref. bb2065 For example, gene expression data and WSI images are often combined to improve cancer prognosis prediction.ref. bb2070,ref. bb2075
Vision-language models
Following its successful use in the natural image domain, vision-language data (consisting of histopathology images paired with relevant natural language text) is becoming increasingly prominent in CPath. Whether it be the development of foundational modelsref. bb2080 extending to CPath, or fine tuning state-of-art large large models for use in downstream tasks,ref. 410, ref. 411, ref. 412 leveraging the semantic information embedded in the natural language data is becoming more evidently beneficial. It was only recently that foundational language models advanced enough to become useful in CPath, and this has triggered an explosion of interest into building models at the intersection of visual and language information. At the moment, language data is primarily used to address Multi-Instance Learning, although this is still an extremely new field and we anticipate that future works will surely address more advanced tasks (see Section 7.4 for further discussion).
Sequential models
Recurrent Neural Networks (RNNs) are typically used in tasks with temporally-correlated sequential data, such as speech or time series.ref. bb2100 Since RNNs consider the past through the hidden state, they are suited for handling contextual information. While images are the default data format in CPath (and hence poorly-suited for RNNs), some works opt to combine RNNs with CNNs as a feature extractor,ref. bb0310,ref. bb0440,ref. bb2105,ref. bb1065,ref. bb1155,ref. bb1320,ref. bb1330,ref. 415, ref. 416, ref. 417, ref. 418, ref. 419, ref. 420 most commonly by aggregating patches or processing feature sequences.ref. bb0310,ref. bb1155,ref. bb1330,ref. bb2125,ref. bb2140 Another application of RNNs is to consider spatial relations between patches, which can be lost after extracting from the slide.ref. bb1065,ref. bb1320
A particularly exciting use of RNNs is in deciding which region within an image should be examined next.ref. bb2120,ref. bb2145 In the “Look, Investigate, and Classify” 3-stage model, an long short-term memory (LSTM) was used to classify the ROI cropped from the current patch and predict the next region to be analyzed, and achieved good performance while only using of pixels from the original image.ref. bb2120 Similarly, an LSTM network was used to better predict ROIs by treating state features similar to time-series data, thus identifying only relevant examples to use for training.ref. bb2145 And an LSTM transformer with “Feature Aware Normalization” (FAN) units for stain normalization was used in parallel with a VGG-19 network.ref. bb1625 More recently, transformers using attention mechanisms have been used to allow parallelization and better sequence translation compared to older RNNs or LSTM networks.ref. bb2150
Synthetic data and generative models
With annotated data difficult to obtain in CPath, especially with granular labels (see Section 3.3 for more discussion), this is problematic for training generalizable models. Hence, generating synthetic data from a controlled environment (either via simulation or a trained model) for augmenting the available training set of annotated data shows much promise. Originally developed for visual style transfer in general computer vision, generative models learn to create novel instances of samples from a given data distribution – they form the dominant approach in CPath.
Initial works primarily utilized Generative Adversarial Networks (GANs) for patch synthesis,ref. bb1685,ref. 424, ref. 425, ref. 426 stain normalization,ref. bb1030,ref. 208, ref. 209, ref. 210,ref. bb2170,ref. bb2175, style transfer,ref. bb1030,ref. 429, ref. 430, ref. 431 and various other tasks.ref. bb1060 One unsupervised pipeline relied on a non-GAN model to create an initial patch that was refined by a GAN.ref. bb1685 In another work, one CycleGAN generated tumor images and another non-tumor, in order to train a classification network.ref. bb2195 One work used neural image compression to learn the optimal encoder to map image patches to spatially consistent feature vectors.ref. bb1060 Another work first classified bone marrow cell representations and then used an unsupervised GAN to generate more instances from each cluster.ref. bb2200 A self-supervised CycleGAN was also used for stain normalization, and shown to improve model performance in subsequent detection and segmentation tasks.ref. bb1050 Similarly, a CycleGAN pipeline was applied to perform artificial IHC re-staining.ref. bb2205 Recent works in GANs attempt to model spatial awareness of tissues and improve the realism of the generated samples.ref. bb2210
Lately, diffusion models have become the SOTA in general computer vision and now produce far more semantically plausible and noise-free images than GANs. These improvements promise to make synthetic data finally accepted by pathologists and the broader CPath community as reliable training data and significantly improve the generalizability of models trained on them.ref. bb2215
Multi-instance learning (MIL) models
Multi-instance learning (MIL) involves training from data that is labelled as high-level bags consisting of numerous unlabelled instances. In the context of CPath, these labelled bags often represent annotated slides of far more unlabelled patch instances.ref. bb2220 As labels at the WSI level are much easier to obtain (and hence more prevalent) than patch-level annotations, MIL has been applied to CPath by a significant number of papers.ref. 62, ref. 63, ref. 64,ref. bb1370,ref. bb1375,ref. bb1575,ref. bb1615,ref. bb2125,ref. 437, ref. 438, ref. 439, ref. 440, ref. 441, ref. 442, ref. 443, ref. 444, ref. 445, ref. 446, ref. 447, ref. 448, ref. 449, ref. 450, ref. 451, ref. 452, ref. 453, ref. 454, ref. 455, ref. 456 Since both utilize coarser annotations for training on massive images, MIL is similar to weakly-supervised learning. However, weak supervision predicts at a finer level (e.g. pixel segmentation from labelled patches) than the provided annotation while MIL prediction is typically at the same level.
One notable work used a two stage approach to first encode patches with a CNN from a slide into feature vectors and then pass the most cancer-likely ones to an slide-level classification RNN. A similar work first detected abnormal regions in the WSI before adaptively fusing the instance-level features with an importance coefficient.ref. bb2230 Adding additional instance-specific attributes tends to improve MIL performance. One work applied a nuclei grading network to provide a cell-level prediction for each patch, and demonstrate this out-performs hand-crafted cell features for overall slide classification.ref. bb1630 Recent works explore the morphological and spatial relationships between instances, which conforms with pathologist diagnostic intuitions and have demonstrably improved performance, especially with unbalanced data.ref. bb2315
As not all instances are equally relevant to the bag label, many works focus on building attention mechanisms to adaptively focus on more relevant instances. One work used such a mechanism to highlight regions of interest and improve localization relative to other SOTA CNNs.ref. bb2220 MIL models can be improved by considering multi-scale information: one work notably used embeddings from different magnification levels and self-supervised contrastive learning to learn WSI classifiers.ref. bb1375 Some works explicitly encode the patient-slide-patch hierarchy in the attention mechanism,ref. bb2320,ref. bb2325 with one work using a cellular graph for top-down attention.ref. bb2330 Graph Neural Networks (GNNs) have been explored to leverage intra- and inter-cell relationships, enabling cancer grading,ref. bb2335 classification,ref. bb2340 and survival prediction.ref. bb2345,ref. bb2350 These hierarchy- and morphology-aware models are the current SOTA and pave the way for future improvements.
One persistent challenge with using MIL in CPath, compared to natural image computer vision, is the lack of large-scale WSI datasets.ref. bb2355 One recent work addressed issues related to small sample cohorts by splitting up large bags (and their labels) into smaller ones through pseudo-bags.ref. bb2360
Contrastive self-supervised learning for few-shot generalization
The idea of using contrastive learning (CL) for self-supervised learning (SSL) dates back to 2005, yet only recently gained momentum in CPath.ref. bb1375,ref. bb1565,ref. 466, ref. 467, ref. 468, ref. 469, ref. 470, ref. 471 By using a contrastive loss, a feature embedding is learned to ensure similar (positive) examples are close in vector space, while dissimilar (negative) examples are distant.ref. bb2365,ref. bb2370 Contrastive learning is an attractive approach for CPath because when used as self-supervision for few-shot learning,ref. bb1375,ref. bb1565,ref. bb2375 it does not require labelling the massive self-supervision image set but only labelling the small subset used for training on the downstream task, an approach that has recently achieved SOTA performance in a wide array of tasks in CPath.ref. bb1565 SimCLR was originally proposed to learn representations invariant to different augmentation transforms (such as crop, noise) for natural images,ref. bb0330 and when applied to CPath, was found to match or outperform SOTA supervised techniques.ref. bb1565 Self-supervised pre-training has been shown to perform best against fully-supervised pre-training when applied to small but visually-diverse datasets.ref. bb1565 Recent works have focused on transferring the self-supervised representations to the downstream task more intelligently: through latent space transfers,ref. bb2395 with an awareness of the patient-slide-patch hierarchy,ref. bb2400,ref. bb2405 or with semi-supervised pseudo-label guidance.ref. bb2410
Novel CPath architectures
In this section, we discuss papers that made significant changes to the model design or completely designed an architecture from scratch for CPath tasks.ref. bb0320,ref. bb1420,ref. bb1820,ref. bb1840,ref. bb1945,ref. 476, ref. 477, ref. 478, ref. 479 Typically, model architectures are adapted from the natural image domain and minor changes applied for CPath tasks, rather than being designed from scratch for CPath directly. Unfortunately, general computer vision architectures typically require large computational resources not necessarily available in clinical settings and are prone to overfitting on smaller CPath training sets.ref. bb2415
More importantly, CPath tasks often comprise of multiple specialized sub-tasks not addressed by common architectures – in such cases, CPath-specific architectures perform better. “PlexusNet” achieved SOTA performance with significantly fewer parametersref. bb2415 and “Hover-Net” used a three-branched architecture for nuclei classification and instance segmentation.ref. bb1420 Path R-CNN similarly used one branch to generate epithelial region proposals and another to segment tumours.ref. bb1945
In other cases, custom architectures are designed to obtain better performance with respect to certain metricsref. bb0425,ref. bb1510,ref. bb1725,ref. bb1920,ref. bb1975 or improve computational efficiency and speed,ref. bb0980 where since model inference can be a bottleneck for WSI processing. To automate architecture design, neural architecture search (NAS) is often used. This is an umbrella term covering evolutionary algorithms (EA), deep learning (specifically Reinforcement Learning), and gradient-based NAS searches. There are two approaches to EA: (1) Neuroevolution, which more generally optimizes at the neuron-level to find optimal weights, and (2) Evolutionary-Algorithms based NAS (EANAS), which searches for optimal combinations of mid-sized neural network blocks and conducts training after this architectural search.ref. bb2435,ref. bb2440 In CPath, reinforcement learning-based NAS has designed models for cancer prediction which were found to train faster, have fewer parameters, and perform comparably with manually designed models.ref. bb2445 Another work demonstrated that a significantly smaller model can outperform existing SOTA models on a variety of CPath tasks using an adaptive optimization strategy.ref. bb2430
We hypothesize that NAS has yet to be explored significantly in CPath due to the lack of annotated data (see Section 3.3) and its relative recency as a research area. According to the “no free lunch theorems”,ref. bb2450,ref. bb2455 no single model can perform best on all tasks. However, computationally efficient but performant models are crucial for CPath applications, and NAS is the most promising approach to computationally design such architectures without manual engineering.
Model comparison
The various model architectures and types discussed above can and should be compared on common benchmarks to determine the best models for a given task.ref. bb2460 Numerous papers have conducted such benchmarking work on CNNs. One work comparing GoogLeNet, AlexNet, VGG16, and FaceNet on breast cancer metastasis classification found that deeper networks (i.e. GoogLeNet) predictably performed better.ref. bb0995 Another work found that using ResNet-34 with a custom gradient descent performed best.ref. bb1970 Finally, VGG-19 performed best in colorectal tissue subtype clasification, showing that deeper SOTA networks do not necessarily perform better universally. Which CNN performs best depends on the task, the nature of the data, the metrics used, training time, hyperparameters, and/or hardware constraints.
Likewise, third parties have organized “grand-challenges” to facilitate the fair comparison of different techniques on a common CPath task and dataset. In some cases, SOTA CNNs achieve the best results, such as the adapted GoogLeNet that obtained the highest AUCref. bb1190 and the AlexNet that achieved highest accuracyref. bb2465 for breast cancer detection in the CAMELYON16 challenge. Likewise, SqueezeNet, which is an existing SOTA network performs best in colorectal tissue subtype classification.ref. bb2470 On the contrary, the best performing models for mitosis detection in the TUPAC16ref. bb0995 and MITOS12ref. bb1725 challenges both relied on custom CNN architectures. For breast cancer diagnosis, a novel Hybrid CNN achieved the best results in the BACH18 (ICIAR18) datasetref. bb2475 while the two teams achieving the best classification accuracy in the BreakHis dataset used differing approaches: one directly used ResNet-50ref. bb1490 and the other used an ensemble of VGG networks.ref. bb2480 For nuclei segmentation on the Kumar-TCGA dataset, a novel framework using ResNet and another existing model achieved the highest F1-score.ref. bb1820 Lastly, a custom CNN achieved the best results for gladn segmentation on the GLaS dataset.ref. bb0840
However, as mentioned in Section 3.3, many grand challenges use private datasets or even extract data from larger public repositories without referencing the original WSIs used. Furthermore, benchmark datasets address different tasks and lack standardization. As models that are hyper-optimized to for specific sets of data continue to be released, the lack of more standardized benchmark datasets and model comparison studies make it impossible to systematically compare new models against existing ones or assess their robustness in clinical settings, thus impeding model development in CPath.
Evaluation and regulations
Clinical validation
Within the domain of CPath, clinical validation is essential for substantiating the decisions produced by deep learning models so that they are more readily accepted by the medical community. Generally, acceptable clinical criteria are determined by authoritative professional guidelines, consensus, or evidence-based sources. However, in CPath, prediction results are generated by the computer scientists and engineers who build the model, and may not be completely aware of where their work fits into the clinical pathology workflow–the clinical implications of this arrangement are often unknown.ref. bb0845 By incorporating pathologist expertise, clinical validation can better align the technical work with clinical objectives.
Despite the importance of this step for real-world deployment, very few works have performed clinical validation with expert pathologists. We identify three prominent types of clinical validation in the CPath literature: (1) direct performance comparison of CAD tools with pathologists on a similar task, (2) impact of CAD tool assistance on pathologist performance, and (3) pathologist validation of CAD tool outputs. Each topic is further discussed in the sections alongside notable results.
Direct Performance Comparison with Pathologists To validate the benefits of deep learning methods, it is desirable that they equal or surpass the performance humans to gain the trust of pathologists in their decisions and their willingness to use them as a second opinion.ref. bb2485 With this in mind, many papers directly compared their models with pathologists in tasks such as prognosis and diagnosis.
One study on cancer detection found that the top computational models from the CAMELYON16 challenge out-performed the 11 pathologists with a two-hour time constraint and performed similarly to the expert pathologist without a time constraint.ref. bb1190 This suggests that deep learning models could be particularly useful in clinical scenarios with excessive numbers of time-critical cases to diagnose. Similarly, for tissue subtype classification, another study performed similarly to, or slightly better than individual pathologists. The proposed model agreed with all pathologists of the time and agreed with two-thirds of pathologists of the time.ref. bb0375 An additional study claimed their deep learning model outperformed pathologists without gynecology-specific training in ovarian carcinoma classification.ref. bb0595 This pushes the idea that CAD predictions can be used as a second opinion due to the potential for human error by individual pathologists.
One paper on diagnosisref. bb1315 demonstrated that deep learning models can correctly classify images that even individual pathologists failed to correctly identify. However, another paper found that of the examples misclassified by their model were also misclassified by at least one pathologist.ref. bb2490 This suggests that deep learning models can aid pathologists in decision-making, but as they tend to achieve a specificity and sensitivity similar to pathologists, they must be applied cautiously to avoid reinforcing the biases or errors of individual pathologists.
Deep learning models for prognosis have been shown to achieve performance similar to or better than experts as well.ref. bb1225,ref. bb1480,ref. bb1940 In one study, the best model for renal clear cell carcinoma classification achieved accuracy, outperforming the inter-pathologist accuracy of .ref. bb1480 This shows that deep learning models and pathologists may perform similarly on patient prognosis.
Overall, AI approaches are not perfect but have approached expert-level ability in a variety of tasks. Deep learning could play an important role as a second opinion and in democratizing the knowledge distilled from many pathologists to other pathology centres. Specifically, deep learning models appear to be best used as a tool to enhance the pathologist workflow, and could provide aid in making quick decisions with high accuracy.ref. bb0070
Impact of CAD Tool Assistance Much of CPath research is conducted under the assumption that the resulting AI tools will be intuitive, usable, and beneficial to pathologists and patients. However, CAD tools that are developed without feedback from pathologists could fail to integrate into a realistic pathologist workflow or impact the most significant diagnostic tasks. Thus, a valuable validation experiment is to compare and comprehend the performance of expert pathologists in clinical tasks before and after being given the assistance of a CAD tool.
In one study, a CAD system called Paige Prostate Alpha leveraged a weakly-supervised algorithm to highlight patches in a WSI with the highest probability of cancer.ref. bb0310 When used by pathologists, the model significantly improved sensitivity, average review time, and accuracy over unaided diagnosis. Likewise, another study using the LYNA algorithm examined the performance of six pathologists on breast cancer tumor classification before and after being able to see the LYNA-predicted patch heatmaps. The results indicate using LYNA substantially improved sensitivity, average review time, and the subjective “obviousness” score for all breast cancer types.ref. bb0360,ref. bb0135
These studies suggest that integrating CAD tools into the clinical workflow will greatly improve pathologist efficiency. However, there is a general lack of research on the impact of CAD tools on pathology efficiency. Such studies would shed more light on the impact of CAD tools and identify approaches for implementation in clinical settings.
FDA regulations
Despite the ongoing development of CAD tools in CPath and its potential for triaging cases and providing second opinions, the regulations regarding this technology pose an obstacle to the testing and deployment of these devices. The FDA currently provides three levels of clearance on AI/ML-based medical technologies: 510(k) clearance, premarket approval, and the De Novo pathway. While one source lists 64 AI/ML-based medical solutions that are currently FDA-approved or cleared, none of these are in the field of CPath.ref. bb2495 A few companies, such as Paige AI, hold the 510(k) clearance for their digital pathology image viewer; however, an automated diagnostic system has yet to be approved. This may indicate a reluctance to change, and the lack of clarity in the process of FDA approval has prevented numerous impactful technologies from being deployed. There is a need for collaboration between researchers, doctors, and governmental bodies to establish a clear pathway for these novel technologies to be validated and implemented in clinical settings.
Emerging trends in CPath research
Computational pathology research has seen a sudden shift of focus in the past year of 2023. Driven by recent technological advances in the field of computer vision for natural images and the release of capable foundational models in natural language processing, formerly difficult research problems in CPath have been solved, opening up exciting new avenues of research, especially the difficulty of training models on adequate annotated data. We will discuss the main research trends below in further detail and make simple predictions of where the field is headed.
Contrastive self-supervised learning becomes mainstream
Data annotation for CPath is a persistent problem – it is easy to collect large amounts of visual data but much harder to annotate them. Transfer learning can help but it is difficult to transfer a model trained on one dataset to generalize to another. Whereas past efforts focused on carefully engineered methods, the recent development of contrastive self-supervised learningref. bb0330 means that it has become the mainstream approach in CPath.ref. bb1375,ref. bb1565,ref. bb2375 Not only does it utilize the massive amounts of unlabelled images typically available in CPath, but it also as a result only requires finetuning on a small set for the downstream task. We anticipate that this will lead to the development of general-use foundational models in the future to perform the most common CPath tasks, as more pathology images are collected and models become more advanced.
Prediction becoming increasingly high-level
We noticed that recent research works are increasingly addressing higher level prediction tasks than before. Whereas patch classificationref. bb1370,ref. 287, ref. 288, ref. 289, ref. 290 or pixel segmentationref. bb1220,ref. 329, ref. 330, ref. 331, ref. 332, ref. 333 was formerly mainstream, these problems appear to have been largely solved, and now there is far more research into higher-level problems dominate, such as multiple-instance learning.ref. bb0310,ref. bb2220,ref. 457, ref. 458, ref. 459 As computational methods continue to improve, it is natural that they are applied not merely as attention aids for pathologists (i.e. at the pixel or patch level), but furthermore are used to make intelligent slide- and patient-level decisions on their own. Indeed, they promise to vastly improve pathologist efficiency when used with human pathologists in the loop to validate the automated decisions, especially when paired with modern natural language capabilities.
Spatial and hierarchical relationships receiving attention
Inspired by the approach taken for natural image computer vision, the mainstream approach in CPath currently requires breaking up large WSIs into smaller patches and perceiving them independently (see Fig. 7). However, this ignores the spatial relationships between cells and tissues or between the patches and their parent slides in histopathology images, which are often relevant or even crucial when making decisions. Many works have recently found success in explicitly encoding an awareness of these inter-cell relationshipsref. bb2335,ref. bb2345,ref. bb2350 and the patch-slide-patient hierarchy,ref. 457, ref. 458, ref. 459 especially using Graph neural networks, but these suffer from higher latency than conventional CNNs. We anticipate future works will seek to speed up GNNs for tasks where spatial and hierarchical relationships are important and continue developing hierarchy-aware attention for MIL techniques.
Vision-language models for explainable predictions
One persistent problem in CPath has been developing models that can explain their decisions for human validation. One obvious route is to develop models that produce natural language output (and even converse with the human user to explain their decisions), but until recently, this would have required collecting massive amounts of pathology text paired with images. With foundational vision-language models widely available and able to generalize to great effect in the natural image domain,ref. 493, ref. 494, ref. 495 recent works have shown that they perform excellently when applied with minimal re-training to CPath images.ref. 410, ref. 411, ref. 412 Further advances require collecting more pathology-specific data, but we anticipate that crowd sourcing of public pathology annotations will become mainstream and this will lead to the development of foundational vision-language models. As natural language capabilities continue improving, we also anticipate that synoptic report automation will become feasible and reinforcement learning from human feedback (RLHF)ref. bb2515 will become common for improving CPath language models.
Synthetic data now realistic enough
Whereas one way to combat the difficulty of annotating CPath data is to develop models that require fewer annotations, another trend is to generate more annotated data for training. Whereas concerns were previously raised about their realism, new advances in generative image models have now been leveraged to produce realistic histopathology images and pixel-accurate annotations simultaneously. However, current works are limited by specific tissues, organs, diseases,ref. bb1060,ref. bb1685,ref. bb2195,ref. bb2200 or stainsref. bb1050,ref. bb2205 and are limited by their inability to easily expand to other histopathology content. We note that generating synthetic data via game engines and 3D model assets is a recent trend in the natural image domain,ref. 497, ref. 498, ref. 499 but visual modelling of histopathology entities is little explored. We anticipate that future works will attempt to improve synthetic histopathology image generation by: (1) attempting to create generative models that can generalize to a broad variety of histopathology images and (2) create simulation software to generate realistic histopathology images without learned models.
Existing challenges and future opportunities
CPath as anomaly detection
Typically in computer vision, the various classes represent distinct normative entities, such as airplanes or bears.ref. bb2535,ref. bb2540 There exist abundant “normal” samples and potentially few “anomalous” samples, which are considered data points significantly dissimilar to the majority within a given class.ref. bb2545 These anomalies are not only out of distribution from the samples in a dataset, there is also a lack of consensus on understanding anomalous representations as effectively identifying anomalies requires ML models to learn a feature space encompassing all “normal” samples within each class.ref. bb2545
In other words, in general computer vision, each class cannot simply be considered as an anomalous version of any other class. However, in CPath, since each class is often considered a different disease state on a single tissue type, diseased classes are essentially extensions of the “normal” healthy class into the “anomalous” zone. From a pathologist’s perspective, similar to the general computer vision approach, the curriculum learning process of a resident pathologist first involves training on histology and gaining a mastery of normal tissue identification, and then train on diseased tissues, so they are able to flag the sample as anomalous and follow up with possible diagnosis.
In light of this, it may be illuminating to approach the problem from an anomaly detection viewpoint: provided a model has sufficient variety of healthy tissue, any anomalies must then be diseased. The output of such an anomaly detection algorithm is dependent on the task at hand. One source describes several meaningful output types that may be producedref. bb2545: an anomaly score which describes how anomalous a sample is and a binary label indicating whether a sample is normal or anomalous. If only identifying anomalous samples is enough, a binary classification procedure may be sufficient. However, if it is necessary to identify the particular stage of progression of a disease type, then a more granular approach in assigning some anomaly score may be more appropriate as explored in a previous work.ref. bb2550 This work found that the confidence score in tissue classification was inversely correlated with disease progression, thus the confidence score may act as a proxy for an anomaly score. Theoretically, such approaches may better replicate the behaviour of pathologists. While several works have used an anomaly detection approach on medical image data outside of CPath,ref. 504, ref. 505, ref. 506 few works tackle the problem for WSI data in CPath.
Leveraging existing datasets
As mentioned in Section 3.3 of this paper, a minority of datasets in CPath are available to be freely used by the public. Additionally, the level of annotations varies for each dataset. However as can be noted in Table 9.11 of the supplementary material, for prominent public datasets such as CAMELYON16, CAMELYON17, GlaS, BreakHis, and TCGA, there is far more available data annotated at the slide level as opposed to more granular predictions. For example, considering breast datasets, there are 399 WSIs annotated at the Slide and ROI levels in CAMELYON16ref. bb2570 and 1399 WSIs annotated at the Patient, Slide, and ROI levels in CAMELYON17,ref. bb0305 in contrast, the TCGA-BRCA dataset contains diagnostic slides and tissue slides that are accompanied with labels at the Patient and Slide levels and diagnostic reports with labels for tissue features and tumor grades.ref. bb1210
The lack of publicly available datasets with granular annotations is a major challenge in CPath. To address this lack some training data, techniques have been proposed to efficiently obtain labels, such as an active deep learning frameworks that use a small amount of labelled data to suggest the most pertinent unlabelled samples to be annotated.ref. bb2575 Alternatively, other works propose models to synthetically create WSI patches, usually with the use of GANs. For example, Hou et al.ref. bb1685 introduced an unsupervised pipeline that was capable of synthesizing annotated data at a large scale, noting that even pathologists had difficulty distinguishing between real and synthesized patches. However, despite these promising results, the issue of acquiring accurate and large datasets remains a prevalent issue within CPath.
Generally, tasks such as tissue classification or gland segmentation require labels at the ROI, Patch, or Pixel levels. However, existing data annotated at the patient and slide levels can be used for these tasks by leveraging weakly supervised techniques such as MIL,ref. bb0320,ref. bb2220 or by learning rich representations using self-supervised techniques such as DINOref. bb0325,ref. bb2580 and contrastive learningref. bb1565 that can be used in downstream tasks. Specifically, work is being done to develop training methodologies and architectures that are more data efficient for patient- and slide-level annotations, such as CLAM, which is a MIL technique that is used to train a performant CPath model with as little as of the training data to get over AUC.ref. bb0320 Another recent work used self-supervised learning on WSIs without labels to train a hierarchical vision transformer and used the learned representations to fine-tune for cancer subtyping tasks. This finetuned model outperformed other SOTA methods that used supervised learning methods on both the full training set and when all models used only of the training set. These examples demonstrate a recent trend in the application of weakly and self-supervised learning techniques to leverage pre-existing and available data with weak labels, showcasing that a large amount of granular labels are not necessarily required for achieving SOTA performance. We urge researchers in the CPath field to follow this trend and focus on how to leverage existing weakly labelled datasets, especially to learn rich representations as a pre-training step for learning on smaller strongly labelled datasets.
Creating new datasets
Although we mention the availability of many datasets and comment on how to leverage this existing data, there is still a need for new CPath datasets that address overlooked clinical and diagnostic areas. Therefore, creation of new CPath datasets should focus on addressing two main goals: (1) tasks that are not addressed adequately by existing datasets and (2) accumulating as large a dataset as possible with maximal variety.
Regarding the first goal, there are still organs, diseases, and pathology tasks without freely available data or sufficient annotations to develop CAD tools. For example, in Fig. 6, we see that whereas breast tissue datasets are abundant, there are few public datasets for the brain and none for the liver. Collecting and releasing datasets for these organs would have significant impact in enabling further works focusing on these applications. Further, analysis of specific organ synoptic reports can guide CPath researchers to build CAD tools to identify or discriminate the most impactful diagnostic parameters. In the case of the prostate, which is discussed in Section 2.5, the synoptic report requires distinguishing IDC from PIN and PIA as it correlates to high Gleason scores. This is important, as high-grade PIN is a cancer precursor requiring follow-up sessions for screenings. These parameters are identified and noted in the report by the pathologist and factor into the final diagnosis and grading. Thus, collecting annotated datasets for such parameters can be crucial to developing CAD tools that are relevant to clinical workflows and can enrich learned representations.
The second goal is concerned with the scaling laws of deep learning models with respect to the amount of data available and their application to diverse clinical settings. As seen in the general computer vision domain, larger datasets tend to improve model performance, especially when used to learn rich model representations through pre-training that can be used for downstream tasks such as classification and semantic segmentation.ref. bb2585 Additionally, ensuring that datasets capture the underlying data distribution and thus sufficiently encompass the test distribution has been shown to be especially important in the medical domain.ref. bb2590 For CPath, this means ensuring a dataset captures the expected variations in tissue structure, disease progression, staining, preparation artifacts, scanner types, and image processing. Collecting a sufficiently large dataset continues to be problematic, however, so recent works have focused on using crowd sourcing to annotate histopathology data posted publicly on Twitter and YouTube,ref. bb2080,ref. bb2090 a practice that is similar to that commonly used for natural images.
Pre- and post-analytical CAD tools
In recent years, advances in image analysis, object detection, and segmentation have motivated new approaches to support the analytical phase of the clinical workflow, especially in the two steps where CAD tools could significantly increase efficiency and accuracy: (1) specimen registration and (2) pathology reports. This need is highlighted by a study determining that the pre-analytical and post-analytical phases (as shown in Fig. 2 account for up to of medical errors in pathology.ref. bb2595 Likewise, Meier et al. classify of medical errors as diagnostic errors, with an even smaller proportion being misinterpreted diagnoses in their study.ref. bb2600 Other authors attribute approximately of diagnostic errors to slide interpretation.ref. 514, ref. 515, ref. 516, ref. 517, ref. 518 These results reinforce the need for CPath applications that address more than just the analytical phase.ref. bb2630 Considering post-analytical step of compiling a pathology report, a few natural language processing efforts have been used to analyze completed pathology reports,ref. 520, ref. 521, ref. 522 extract primary site codes from reports,ref. bb2650 and generate of captions or descriptive texts for WSI patches.ref. bb2110 However, to the best of our knowledge, there are no works that reliably extract clinical data from service requests and electronic medical records to automatically generate synoptic or text reports. Developing such a tool that could explicitly identify the most significant parameters for its decisions would directly improve clinical workflow and increase the interpretability of the results at the same time. We encourage the field of CPath to expand its efforts in creating tools for the pre- and post-analytical steps in order to reduce the large proportion of clinical errors attributed to those phases, and suggest some potential applications in Fig. 2.
Multi domain learning
Despite being particularly well-suited for CPath, multi-domain learning (MDL) is still a relatively unexplored topic. MDL aims to train a unified architecture that can solve many tasks (e.g. lesion classification, tumour grading) for data coming from different domains (e.g. breast, prostate, liver). During inference, the model receives an input image and the corresponding domain indicator and is able to solve the corresponding task for the given domain. There are two reasons that make MDL attractive for CPath. The first is that the additional information from a source domain (coming from a related organ such as the stomach) can be informative for improving performance in the target domain (e.g. colon). By sharing representations between related domains, the model is enabled to generalize to other domains. The second motivation is to alleviate the data sparsity problem where one domain has a limited number of labeled data. Through MDL, the domain with limited data can benefit from the features that are jointly learned with other related tasks/domains.ref. bb2655,ref. bb2660
Federated learning for multi-central CPath
Data-driven models require a large amount of data to yield strong performance. In CPath, this requires incorporating diverse datasets with varying tissue slide preparations, staining quality, and scanners. An obvious solution to train such models is to accumulate the data from multiple medical centers into a centralized repository. However, in practice, data privacy regulations may not permit such data sharing between medical institutions, especially between countries. A possible solution lies in privacy-preserving training algorithms, such as federated learning,ref. bb2665,ref. bb2670 which can make use of decentralized data from multiple institutions while maintaining data privacy. In federated learning, training starts with a generic machine learning model in a centrally located server. But instead of transferring data to a centralized server for training, copies of the model are sent to individual institutions for training on their local data. The learning updates are encrypted and sent to the central server and then aggregated across the institutions. Ming Lu et al.ref. bb2270 demonstrated the feasibility and effectiveness of applying federated, attention-based weakly supervised learning for general-purpose classification and survival prediction on WSIs using data from different sites. Using such algorithms for CPath can facilitate cross-institutional collaborations and can be a viable solution for future commercial solutions that need to continuously augment and improve their ML models using decentralized data.
CPath-specific architecture designs
Many deep learning architectures are not designed for CPath specifically, which raises a serious question about the optimality of using “borrowed” architectures from general computer vision. For instance,ref. bb2415 notes that traditional CV architectures may not be well suited for CPath due to a large number of parameters that risk overfitting. Additionally, the field of pathology has much domain-specific knowledge that should be taken into account before choosing an ML model. For example, under varying magnifications different morphological patterns are captured, from cellular-level details to tissue architecture features.ref. bb1675 Naively applying an architecture without considering such details could discard key visual information and lead to deteriorated performance.
Unlike natural images, WSIs exhibit translational, rotational, and reflective symmetryref. bb2675 and CNNs for general vision applications do not exploit this symmetry. The conventional approach to overcome this issue is to train the model with augmented rotations and reflections, but this increases training time and does not explicitly restrict CNN kernels to exploit those symmetries. Rotation-equivariant CNNs, which are inherently equivariant to rotations and reflections were introduced for digital pathology,ref. bb2675 significantly improving over a comparable CNN on slide level classification and tumor localization. Similarly, Lafarge et al.ref. bb1915 designed a group convolution layer leveraging the rotational symmetry of pathology data to yield superior performance in mitosis detection, nuclei segmentation, and tumor classification tasks. These results motivate the application and further research of rotation-equivariant models for CPath.
In general, we note that the SOTA computer vision architectures used in computational pathology have tended to lag behind those used for natural images by a couple of years. This delay in knowledge propagation from the mainline computer vision research in natural images may be due to the data-centric nature of the CPath field. As data labelling is specialized and expensive to conduct, annotating more data or clever training tweaks to finetune established architectures is more attractive than developing advanced, specialized architectures. However, we recommend that CPath researchers should still use the most powerful relevant models available for the simple reason that they tend to perform best given the computational resources available. While computational efficiency is generally not as important during training, it is imperative at inference time if models are to be run in real-time on medical devices with limited computational resources.
Digital and computational pathology adoption
Despite the numerous advantages to the clinical workflow and applications offered by using digital pathology and CPath, the adoption of digital pathology remains the first barrier to clinical use. A major reason for adoption hesitancy is the common opinion that digital slide analysis is an unnecessary step in a pathologist’s workflow which has been refined over decades to produce reproducible and robust diagnoses without digitization.ref. bb0045,ref. 18, ref. 19, ref. 20 In terms of clinical efficiency, studies have shown mixed results, with two finding that digitization actually decreased efficiency (by increasing turnaround time by ).ref. bb2680,ref. bb2685 However, another study demonstrated a clear increase in productivity and reduction of turnaround-time.ref. bb0120 One of the co-authors (B.N.) has implemented digital pathology at a public tertiary institution, which began as a pilot study over three years including three experienced academic pathologists and showed that digitization reduced turnaround time by for biopsies and for resections, and increased case output by . These trial results led to all pathologists not retiring within two years to transition to a digital pathology workflow in 2019. Due to the varied nature of results and outcomes in studies analyzing the effectiveness of digital pathology there is more work to be done to have a multi-institution and lab analysis for more general and concrete results.
A major factor in the adoption of digital or computational pathology practices is the source of funding and the pay structure of pathologists. A few cost-analysis studies show that the transition to digital pathology becomes financially advantageous in 2 years, with savings projected to be up to $5M after 5 years in a sizeable tertiary center.ref. bb0045,ref. 531, ref. 532, ref. 533 The financial impact will also be viewed differently in public vs private healthcare settings. Public healthcare is primarily limited by funding and universal access to healthcare whereas for private lab networks improvements in processes and services are directly linked to the prospects of obtaining additional contracts and increased profitability. However, studies considering multiple institutions and funding settings are still required to fully characterize the financial impact compared to clinical benefit. Additionally, on an individual pathologist level, compensation structures can affect buy in for implementation. For example, at our co-authors’ B.N. and V.Q.T institution, a fee-for-service structure is used to compensate pathologists thus an increase in throughput and productivity has a direct correlation to increased pay. We propose that this fee-for-service model contributes to the widespread embracement of DP at this institution. In contrast, pathologists in a salary-based environment are paid based based on a combined package of services which includes diagnostics, research, teaching, administration, quality control, etc. An increase in clinical productivity would technically not benefit them directly, as it would translate to a high number of rendered diagnostics over the same amount of time.
Integration CPath into the clinical workflow is relatively understudied as few papers have actually deployed, or performed clinical validation of their results. Works in this area have either proposed methods to deploy their models in the clinic or developed tools to enable the use of their research in the clinic.ref. bb1470,ref. bb2485,ref. bb2705 However, as a primary goal of CPath is the use of CAD tools in clinical settings, more works should consider how to integrate models and tools into the clinical workflow, especially in conjunction with expert clinicians.
Institutional challenges
Several institutional challenges may affect the implementation of CPath tools, and similar challenges in implementing digital pathology workflows at medical institutions have been well-described by many studies.ref. 535, ref. 536, ref. 537, ref. 538, ref. 539 As noted by multiple studies considering the digital transition of pathology laboratories,ref. 535, ref. 536, ref. 537, ref. 538, ref. 539 the importance of a common shared goal and frequent communication between the involved parties is necessary to successfully deploy a digital system. These lessons are likely extendable in the context of CPath and CAD development as well. Specifically, Cheng et al.ref. bb2725 reported on their experiences and lessons learned as a 7-point-based system to efficiently deploy a digital pathology system in a large academic center. We believe similar systematic approaches will need to be developed to implement CPath applications in a clinical setting.
Another institutional challenge concerns the regulatory oversight at the departmental, institutional, accrediting agencies, pathology association, state/provincial, and federal levels. Regulatory measures underlying WSI scanners are well established, as well as the technical and clinical validation of their use.ref. 540, ref. 541, ref. 542 On the other hand, patient confidentiality, ethics, medical data storage regulations, and data encryption laws are equally, if not more, time-consuming and intensive to comply with. These issues can be mitigated by deploying a standardized digital pathology system throughout multiple institutions at the state/provincial level. For example, our co-author (B.N.) has obtained governmental approval and funding to distribute a set of digital pathology systems throughout the province’s public anatomical pathology laboratories. Similarly, a unified set of standards for processing and digitizing slides, along with unifying storage and access to WSIs for research use in collaborative efforts is paramount in moving forward in both the development and implementation of CAD systems.
Clinical alignment of CPath tasks
Researchers in the CPath field must ensure that the CAD tools they create are clinically relevant and applicable to pathology so that effort and resources are not allocated towards extraneous or clinically irrelevant tasks. For example, certain CADs have been proposed to facilitate case triaging and reduce turnaround time for critical diagnoses.ref. 72, ref. 73, ref. 74,ref. bb2750,ref. bb2755 However, several regulatory agencies in pathology aim for of cases to be completed within a timeframe of 72 hours for signing-out resection specimens and up to 48 hours for biopsies.ref. bb2760,ref. bb2765 In this context, triaging becomes extraneous, as signing out cases faster than 48-72 hours has no clinical impact. However, in the context of an institution operating at longer turnaround times or struggling to keep up with the caseload, this method could be lifesaving. Alternatively, identifying mitotic figures and counting positive Ki-67 nuclei are appreciated tools already in use in multiple digital pathology settings, despite these tools being seldom applied to the large caseload proportion of most practicing pathologists.
As noted previously, the overall number of pathologists in the USA has decreased from 2007 to 2017 and caseloads have increased by .ref. bb2770 This trend places further emphasis of developing CAD tools towards specific challenges encountered by pathologists and where sub-specialists may not be readily available. For example, a large consortium generated a prostate cancer CAD that achieved a concordance with expert genitourinary pathologists,ref. bb2775 a significant breakthrough for healthcare settings where prostate biopsies are not signed out by sub-specialists. Additionally, targeting specific diagnoses with high rates of medical errors and inter-observer variance, notably in dermatological, gynecological, and gastrointestinal pathology, should be prioritized and integrated into practice quickly to support patient care.ref. bb2780 Finally, advanced CAD tools capable of diagnosing features out of reach by conventional pathology could have a great impact. For example, identifying the origin of metastases from morphological cues on the WSIs without added IHCref. bb2275 or CADs capable of calculating the exact involvement of cancer on a biopsy core for prognostic purposes.ref. bb2775
Concluding remarks
Bringing pathologists and computer scientists together and initiating meaningful collaborations with shared gains between all parties is likely the most efficient path forward for CPath and CAD integration. Opportunities to facilitate collaborations should be promoted by parties such as the Pathology Innovation Collaborative Community and the Digital Pathology Association. Furthermore, we encourage involved pathologists and computer scientists to communicate and collaborate on studies towards the common goal of providing patients with fast, reproducible, and high-quality care.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
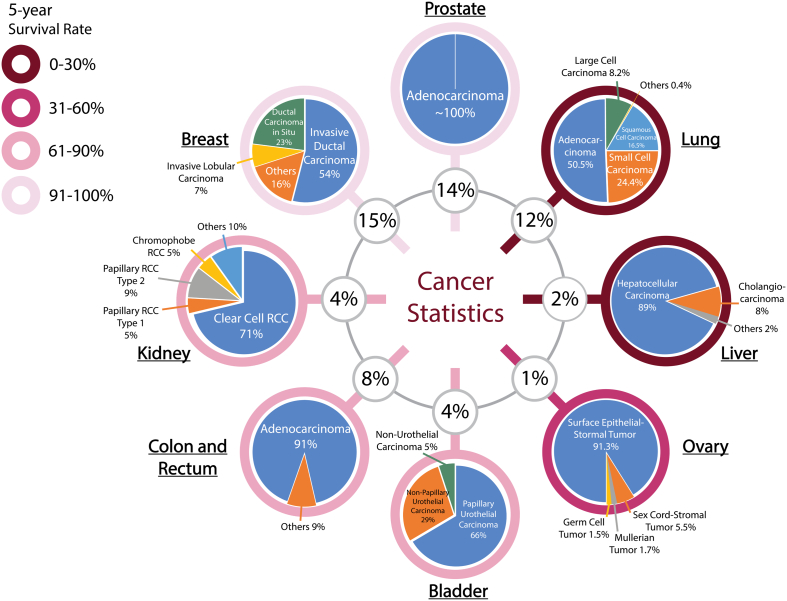
Compilation of all the datasets carefully studied in this survey with its respective information (see Table Creation Details).
| Dataset Name | References | Availability | Stain Type | Size | Res(μm)/ Mag | Annotation | Label | Class | CB |
|---|---|---|---|---|---|---|---|---|---|
| Basal/Epithelium | |||||||||
| NKI-VGH | ref. bb3070 | Link | H&E | 158 ROIs | N/A | Pixel | S | 2 | U |
| AJ-Epi-Seg | ref. bb2750 | Link | H&E | 42 ROIs | 20× | Pixel | S | 2 | U |
| TCGA-Phil | ref. bb1385 | TCGA | H&E | 50 WSIs | 40× | Pixel | S | 4 | I |
| MOIC | ref. bb3025 | N/A | N/A | 6 610 MOIs | 10× | Slide | S | 2 | U |
| MOIS | ref. bb3025 | N/A | N/A | 1 436 MOIs | 10× | Pixel | S | 2 | U |
| Jiang et al. | ref. bb3025 | N/A | N/A | 128 WSIs | 40× | Pixel | S | 2 | U |
| MIP | ref. bb3075 | N/A | H&E | 108 Patients | 40× | Patient | S | 2 | I |
| YSM | ref. bb3075 | N/A | H&E | 104 Patients | 40× | Patient | S | 2 | I |
| GHS | ref. bb3075 | N/A | H&E | 51 Patients | 40× | Patient | S | 2 | I |
| DKI | ref. bb1520 | N/A | H&E | 695 WSIs | 40× | Slide | S | 2 | I |
| Y/CSUXH-TCGA | ref. bb3080 | N/A | H&E | 2 241 WSIs | 0.275/40×, 0.5/20×, 1/10×, 5/4× | Slide | S | 4 | U |
| BE-Hart | ref. bb3085 | N/A | H&E | 300 WSIs | 40× | Patch | S | 2 | I |
| BE-Cruz-Roa | ref. bb3090 | N/A | H&E | 308 ROI, 1 417 Patches | 10× | Patch | S | 2 | U |
| DLCS | ref. bb0860 | N/A | H&E | 5 070 WSIs | 0.25/40× | Slide | S | 4 | U |
| BE-TF-Florida-MC | ref. bb0860 | N/A | H&E | 13 537 WSIs | 0.24/20×, 0.5/20×, 0.55/20× | Slide | S | 4 | U |
| Bladder | |||||||||
| TCGA+UFHSH | ref. bb2110 | By Req. | H&E | 913 WSIs | 40× | Slide, ROI | S | 2 | I |
| TCGA-Woerl | ref. bb0730 | TCGA | H&E | 407 WSIs | 40× | ROI | S | 4 | I |
| Bla-NHS-LTGU | ref. bb1935 | N/A | IF | 75 ROIs | N/A | Pixel | S | 2 | U |
| CCC-EMN MIBC | ref. bb0730 | N/A | H&E | 16 WSIs | 40× | ROI | S | 4 | I |
| UrCyt | ref. bb3095 | N/A | ThinStrip | 217 WSIs | 40× | Pixel | S | 3 | U |
| AACHEN-BLADDER | ref. bb2460 | N/A | H&E | 183 Patients | N/A | Patient | S | 2 | I |
| Brain | |||||||||
| TCGA-Shirazi | ref. bb0425 | N/A | H&E | 654 WSIs, 849 ROIs | 0.5 | ROI | M | 4 | I |
| TCGA-GBM-Tang | ref. bb3100 | TCGA | N/A | 209 Patients, 424 WSIs | 0.5/20× | Patient | S | 2 | I |
| MICCAI14 | ref. bb0380,ref. bb3105 | N/A | H&E | 45 WSIs | N/A | Slide | S | 2 | B |
| M-Qureshi | ref. bb1740 | N/A | H&E | 320 ROIs | N/A | ROI | S | 4 | B |
| Lai et al. | ref. bb3015 | N/A | Amyloid-β antibody | 30 WSIs | 20× | Slide, Pixel | S | 2, 3 | I, U |
| Vessel | ref. bb3110 | Link | H&E, PAS-H, Masson tri-chome, Jones | 226 WSIs | 0.25/40× | ROI | M | 3 | I |
| WCM | ref. bb3115 | N/A | H&E | 87 WSIs | N/A | Patch | S | 2 | U |
| Esophagus | |||||||||
| ESO-DHMC | ref. bb1475 | N/A | H&E | 180 WSIs, 379 ROIs | 20× | ROI | S | 4 | U |
| Kidney | |||||||||
| AIDPATHA | ref. bb1980,ref. bb3120 | Link | PAS | 31 WSIs | 20× | Pixel | S | 3 | U |
| AIDPATHB | ref. bb1980,ref. bb3120 | Link | PAS | 2 ∼ 340 Patches | 20× | Patch | S | 2 | B |
| M-Gadermayr | ref. bb3125 | N/A | PAS | 24 WSIs | 20× | ROI | S | 2 | U |
| TCGA-RCC-Lu | ref. bb0320 | TCGA | H&E | 884 WSIs | 20×, 40× | Slide | S | 3 | I |
| BWH-CRCC | ref. bb0320 | N/A | H&E | 135 WSIs | 10×, 20× | Slide | S | 3 | I |
| BWH-BRCC | ref. bb0320 | N/A | H&E | 92 WSIs | 40× | Slide | S | 3 | I |
| BWH-RCC | ref. bb0320 | N/A | H&E | 135 WSIs | 40× | Slide | S | 3 | I |
| Kid-Wu | ref. bb1970 | N/A | N/A | 1 216 Patients, 60 800 Patches | N/A | Patch | S | 2 | U |
| UHZ-Fuchs | ref. bb1940 | N/A | MIB-1 | 133 Patients | 0.23/40× | Patient | S | 9 | I |
| WUPAx | ref. bb0420 | N/A | H&E | 48 WSIs | 0.495 | ROI | S | 2 | I |
| Pantomics | ref. bb3130 | N/A | H&E | 21 349 Patches | 0.5/20× | Patch | S | 2 | U |
| RUMC | ref. bb0760 | N/A | PAS | 50 WSIs | 0.24/20× | Pixel | S | 10 | I |
| Mayo | ref. bb0760 | N/A | PAS | 10 WSIs | 0.49/20× | Pixel | S | 10 | I |
| UHZ-RCC | ref. bb1480 | N/A | MIB-1 | 1 272 Patches | N/A | Patch | S | 2 | I |
| Kid-Cicalese | ref. bb3135 | N/A | PAS | 1 503 ROIs | N/A | ROI | S | 2 | I |
| Kid-Yang | ref. bb1975 | N/A | H&E, PAS, Jones | 949 WSIs | 0.25 | ROI | S | 2 | U |
| Kid-BWH-TCGA | ref. bb2270 | N/A | H&E | 1 184 WSIs | 20× | Slide | S | 3 | I |
| WTH | ref. bb2140 | N/A | N/A | 3 734 Patients | N/A | Patient | S | 3 | I |
| TCGA-RCC-Chen | ref. bb3140 | N/A | N/A | 45K ROIs | 20× | ROI | S | 3 | B |
| BWH-RCC-Chen | ref. bb3140 | N/A | N/A | 1 661 ROIs | 20× | ROI | S | 3 | I |
| MC-Gallego | ref. bb1755 | N/A | H&E, PAS | 20 WSIs, 1 184 ROIs | 20×, 40× | ROI | S | 2 | I |
| AACHEN-RCC | ref. bb2460 | N/A | H&E | 249 Patients | N/A | Patient | S | 3 | I |
| ANHIR | ref. bb3145 | Link | H&E, MAS, PAS, and PASM | 50 WSIs | 0.1 to 0.2/40× | Slide | S | 8 | I |
| Glomeruli renal biopsies | ref. bb3150 | N/A | H&E, PAS or Jones | 42 WSIs | 0.25 | ROI | S | 2 | I |
| Hubmap Glom | ref. bb3155 | Link | H&E, PAS, PAS-H, Silver, Jones, Van Gieson, etc | 3712 WSIs | 0.13 to 0.25/40× | ROI | S | 2 | U |
| KPMP | ref. bb3155 | Link | PAS-H | 26 WSIs | 0.25/40× | ROI | M | 2 | U |
| Breast | |||||||||
| BreakHis | ref. bb1165,ref. bb1175 | By Req. | H&E | 82 Patients, 7 909 ROIs | 40×, 100×, 200×, 400× | ROI | S | 2 | I |
| CAMELYON 16 | ref. bb0305,ref. bb1190,ref. bb2570 | link | H&E | 399 WSIs | 0.243/20×, 0.226/40× | Slide, ROI | S | 3 | I |
| BACH18 | ref. bb1135,ref. bb2980 | Link | H&E | 40 WSIs, 400 Patches | 0.42, 0.467 | Patch, Pixel | S | 4 | B |
| TUPAC16 | ref. bb1360 | Link | H&E | 821 WSIs | 40× | Slide | S | 3 | I |
| TUPAC16-Mitoses | ref. bb1360 | Link | H&E | 73 WSIs | 0.25/40× | ROI | S | 2 | U |
| TUPAC16-ROIs | ref. bb1360 | Link | H&E | 148 WSIs | 40× | ROI | S | 2 | U |
| CAMELYON 17 | ref. bb0305,ref. bb1195,ref. bb3160 | Link | H&E | 1 399 WSIs | 0.23, 0.24, 0.25 | Patient, Slide, | S | 5, | I |
| ROI | 4, 3 | ||||||||
| BioImaging | ref. bb1315,ref. bb3165 | N/A | H&E | 285 WSIs | 0.42/200× | Slide | S | 4 | B |
| Ext-BioImaging | ref. bb1320 | N/A | H&E | 1 568 WSIs | 0.42/200× | Slide | S | 4 | I |
| MITOS-ATYPIA14 | ref. bb1905,ref. bb3170 | Link | H&E | 1 696 HPFs | 40× | Pixel | S | 2 | U |
| MITOS12 | ref. bb1905,ref. bb3175 | Link | H&E | 50 HPFs | 0.185/40×, 0.2273/40×, 0.22753/40×, 0.2456/40× | Pixel | S | 2 | U |
| AJ-Lymphocyte | ref. bb1985,ref. bb2750 | Link | H&E | 100 ROIs | 40× | Pixel | S | 2 | U |
| MSK | ref. bb0310,ref. bb3180,ref. bb3185 | Link | H&E | 130 WSIs | 0.5/20× | Slide | S | 2 | I |
| CCB | ref. bb1785 | Link | H&E | 33 Patches | 40× | Pixel | S | 2 | U |
| BIDMC-MGH | ref. bb0130 | Link | H&E | 167 Patients, 167 WSIs | 0.25/40× | Patient | S | 4 | I |
| PUIH | ref. bb1065 | N/A | H&E | 4 020 WSIs | 100×, 200× | Slide | S | 4 | I |
| HASHI | ref. bb1130 | Link | H&E | 584 WSIs | 0.2456/40×, 0.23/40× | ROI | S | 2 | U |
| TNBC-CI | ref. bb1300 | Link | H&E | 50 Patches | 40× | Pixel | S | 2 | U |
| AP | ref. bb3190,ref. bb3195 | Link | H&E | 300 ROIs | 40× | ROI | S | 3 | I |
| KIMIA Path24 | ref. bb3200 | Link | N/A | 28 380 Patches | 0.5/20×, 0.25/40× | Patch | S | 24 | U |
| BCSC | ref. bb1305,ref. bb2225 | By Req. | H&E | 240 WSIs | 40× | Slide, ROI | M | 14 | I |
| AJ-IDC | ref. bb1510,ref. bb2750,ref. bb3205 | Link | H&E | 162 WSIs, 277 524 Patches | 40× | Patch | S | 2 | I |
| PCam | ref. bb2675 | Link | H&E | 327 680 Patches | 10× | Patch | S | 2 | I |
| AJ-N | ref. bb3210 | N/A | H&E | 141 ROIs | 40× | Pixel | S | 2 | U |
| TCGA-Cruz-Roa | ref. bb3215 | TCGA | H&E | 195 WSIs | 0.25/40× | ROI | M | 5 | U |
| TCGA-Jaber | ref. bb3220 | TCGA | H&E | 1 142 WSIs | 20× | Patch | S | 2 | I |
| TCGA-Corvò | ref. bb3225 | TCGA | H&E | 91 WSIs | N/A | ROI | S | 4 | U |
| TCGA-Lu-Xu | ref. bb1985 | TCGA | H&E | 1K WSIs | N/A | ROI | S | 2 | I |
| AMIDA13 | ref. bb1900,ref. bb3230 | N/A | H&E | 606 ROIs | 0.25/40× | Pixel | S | 2 | U |
| MICCAI16/17 | ref. bb1685 | N/A | H&E | 64 WSIs | N/A | Pixel | S | 2 | U |
| MICCAI18 | ref. bb1685 | N/A | H&E | 33 WSIs | N/A | Pixel | S | 2 | U |
| RUMC-Litjens | ref. bb0355 | N/A | H&E | 271 WSIs | 0.24/20× | ROI | S | 2 | U |
| ABCTB | ref. bb2180 | N/A | H&E | 2 531 WSIs | 20× | Patient | S | 3 | U |
| NHO-1 | ref. bb0485 | N/A | H&E | 110 WSIs | 40× | Slide | S | 2 | I |
| RUMC-Bejnordi | ref. bb3235 | N/A | H&E | 221 WSIs | 0.243/20× | Slide, ROI | S | 3 | I |
| UVLCM-UVMC | ref. bb3240 | N/A | H&E | 2 387 WSIs | 0.455/20× | Slide | S | 6 | I |
| HUP | ref. bb3215 | N/A | H&E | 239 WSIs | 0.25/40× | ROI | M | 5 | U |
| UHCMC-CWRU | ref. bb3215 | N/A | H&E | 110 WSIs | 0.23/40× | ROI | M | 5 | U |
| CINJ | ref. bb3215 | N/A | H&E | 40 WSIs | 0.25/40× | ROI | M | 5 | U |
| Bre-Steiner | ref. bb0135 | N/A | H&E, IHC | 70 WSIs | 0.25 | Slide | S | 4 | I |
| NMCSD | ref. bb0360 | N/A | N/A | 108 WSIs | 0.24 | Slide | S | 4 | I |
| BC-Priego-Torres | ref. bb2835 | N/A | H&E | 12 WSIs | 0.2524/40× | Pixel | S | 2 | U |
| BIRL-SRI | ref. bb3245 | N/A | H&E | 65 WSIs, 5 151 Patches | 2/5 ×, 1/10× | ROI | S | 2 | U |
| BWH-Lymph | ref. bb0320 | N/A | H&E | 133 WSIs | 40× | Slide | S | 2 | B |
| NHS-Wetstein | ref. bb1735 | N/A | H&E | 92 WSIs | 0.16/40× | ROI | S | 3 | U |
| BRE-Parvatikar | ref. bb1295 | N/A | H&E | 93 WSIs, 1 441 ROIs | 0.5/20× | ROI | S | 2 | I |
| BWH-TCGA-Breast | ref. bb2270 | N/A | H&E | 2 126 WSIs | 20× | Slide | S | 2 | I |
| Bre-Brieu | ref. bb1935 | N/A | H&E | 30 ROIs | N/A | Pixel | S | 2 | U |
| Duke | ref. bb3010 | N/A | H&E | 140 WSIs | 0.5/20× | ROI | S | 2 | U |
| TransATAC | ref. bb3010 | N/A | H&E | 30 WSIs | 0.45/20× | ROI | S | 2 | U |
| BRACS | ref. 638, ref. 639, ref. 640 | By req. | H&E | 547 WSIs, 4 539 ROIs | 0.25/40× | Slide, ROI | S | 7 | I |
| Post-NAT-BRCA | ref. bb3265 | Link | H&E | 138 Patients | 40× | Patient | S | 3 | I |
| BCSS | ref. bb3270 | Link | H&E | 151 WSIs, 20K ROIs | 0.25 | ROI | M | 20 | I |
| Amgad et al. | ref. bb1870 | N/A | H&E | 151 WSIs, 20 340 ROIs | 0.25/40× | ROI | S | 5 | U |
| SMH+OVC | ref. bb1745 | N/A | Ki67 | 30 TMAs, 660 Patches | 20× | Pixel | S | 2 | U |
| DeepSlides | ref. bb1745 | Link | Ki67 | 452 Patches | 40× | Pixel | S | 2 | U |
| Protein Atlas | ref. bb1745,ref. bb3275 | Link | Ki67 | 56 TMAs | 20× | Slide | S | 3 | U |
| Yale HER2 | ref. bb1745 | N/A | H&E | 188 WSIs | 20× | ROI | S | 3 | U |
| Yale Response | ref. bb1745 | N/A | H&E | 85 WSIs | N/A | ROI | S | 2 | U |
| TCGA-Farahmand | ref. bb1745 | N/A | H&E | 187 WSIs | N/A | ROI | S | 2 | U |
| Breast Histopathology Images | ref. bb3280 | Link | H&E | 162 WSIs | 40× | Patch | S | 2 | I |
| Colsanitas | ref. bb3285 | N/A | H&E | 544 WSIs | 0.46/40× | ROI | M | 4 | I |
| Pancreas | |||||||||
| Pan-Bai | ref. bb1705 | N/A | Ki67 IHC | 203 TMAs | Ma×. of 20× | Pixel | S | 3 | I |
| Liver | |||||||||
| SUMC | ref. bb2485 | N/A | H&E | 80 WSIs | 0.25/40× | Slide | S | 2 | B |
| MGH | ref. bb3290 | N/A | H&E | 10 WSIs | 0.46/20× | ROI | S | 4 | U |
| Liv-Atupelage | ref. bb2805 | N/A | H&E | 305 ROIs | 20× | ROI | S | 5 | I |
| IHC-Seg | ref. bb1715 | N/A | H&E, PD1, CD163/CD68, CD8/CD3, CEA, Ki67/CD3, Ki67/CD8, FoxP3, PRF/CD3 | 77 WSIs | 20× | Pixel | S | 4 | I |
| Lung | |||||||||
| TCGA-Gertych | ref. bb1235 | IDs | H&E | 27 WSIs, 209 ROIs | 0.5/20×, 0.25/40× | ROI, Pixel | S | 4 | I |
| TCGA-Brieu | ref. bb1935 | TCGA | H&E | 142 ROIs | N/A | Pixel | S | 2 | U |
| TCGA-Wang | ref. bb2705,ref. bb3185,ref. bb3295 | TCGA | H&E | 1 337 WSIs | 20×, 40× | ROI | S | 3 | U |
| NLST-Wang | ref. bb2705,ref. bb3300 | By Req. | H&E | 345 WSIs | 40× | ROI | S | 3 | U |
| TCGA-Wang-Rong | ref. bb1070,ref. bb3185,ref. bb3295 | TCGA | H&E | 431 WSIs | 40× | ROI | S | 6 | U |
| NLST-Wang-Rong | ref. bb1070,ref. bb3300 | By Req. | H&E | 208 WSIs | 40× | Pixel | S | 7 | U |
| SPORE | ref. bb2705,ref. bb3305 | N/A | H&E | 130 WSIs | 20× | ROI | S | 3 | U |
| CHCAMS | ref. bb2705 | N/A | H&E | 102 WSIs | 20× | ROI | S | 3 | U |
| TCGA-Hou-2 | ref. bb1205 | TCGA | H&E | 23 356 Patches | 0.5/20× | Patch | S | 2 | I |
| TCGA-LUSC-Tang | ref. bb3100 | TCGA | N/A | 98 Patients, 305 WSIs | 0.5/20× | Patient | S | 2 | I |
| TCGA-CPTAC-Lu | ref. bb0320 | TCGA, CPTAC | H&E | 1 967 WSIs | 20×, 40× | Slide | S | 2 | I |
| DHMC | ref. bb0375 | N/A | H&E | 422 WSIs, 4 161 ROIs, 1 068 | 20× | Slide, | M, | 6 | I |
| Patches | ROI, Patch | S, S | |||||||
| Lung-NHS-LTGU | ref. bb1935 | N/A | IF | 29 ROIs | N/A | Pixel | S | 2 | U |
| CSMC | ref. bb1235 | N/A | H&E | 91 WSIs, 703 ROIs | 0.5/20× | ROI, Pixel | S | 4 | I |
| MIMW | ref. bb1235 | N/A | H&E | 88 WSIs, 1 026 ROIs | 0.389/20× | ROI, Pixel | S | 4 | I |
| NSCLC-Wang | ref. bb0445 | N/A | H&E | 305 Patients | 20× | Patient | S | 2 | I |
| ES-NSCLC | ref. bb3310 | N/A | H&E | 434 Patients, 434 TMAs | 20× | Patient | S | 2 | I |
| BWH-NSCLC-CL | ref. bb0320 | N/A | H&E | 131 WSIs | 20× | Slide | S | 2 | I |
| BWH-NSCLC-BL | ref. bb0320 | N/A | H&E | 110 WSIs | 40× | Slide | S | 2 | B |
| BWH-NSCLC-RL | ref. bb0320 | N/A | H&E | 131 WSIs | 20×, 40× | Slide | S | 2 | I |
| VCCC | ref. bb2850 | N/A | N/A | 472 Patients | N/A | Patient | S | 3 | I |
| Dijon+Caen | ref. bb3315 | N/A | HES | 197 WSIs | 20× | ROI | S | 2 | U |
| PKUCH+TMUCH | ref. bb1760 | N/A | IHC | 239 WSIs, 677 ROIs | 20× | ROI | S | 2 | U |
| Lymph Nodes | |||||||||
| LYON19 | ref. bb0840,ref. bb3320,ref. bb3325 | Link | IHC | 441 ROIs | 0.24 | Pixel | S | 2 | U |
| AJ-Lymph | ref. bb2750,ref. bb3330 | Link | H&E | 374 WSIs | 40× | Slide | S | 3 | I |
| TUCI-DUH | ref. bb2825 | N/A | H&E | 378 WSIs | 0.24/20× | Slide | S | 2 | I |
| Thagaard-2 | ref. bb3335 | N/A | H&E, IHC | 56 Patches | 20× | Patch | S | 2 | I |
| Thagaard-3 | ref. bb3335 | N/A | H&E, IHC | 135 Patches | 20× | Patch | S | 2 | B |
| Thagaard-4 | ref. bb3335 | N/A | H&E, IHC | 81 Patches | 20× | Patch | S | 2 | I |
| Thagaard-5 | ref. bb3335 | N/A | H&E, IHC | 60 Patches | 20× | Patch | S | 2 | I |
| Zhongshan Hospital | ref. bb0830 | N/A | H&E | 595 WSIs | 0.5/20× | ROI | M | 2 | I |
| Mouth/Esophagus | |||||||||
| SKMCH&RC | ref. bb3340 | N/A | H&E | 70 WSIs, 193 ROIs | 0.275/40× | ROI | S | 2 | I |
| SKMCH&RC-M | ref. bb3340 | N/A | H&E | 30 WSIs | 0.275/40× | ROI | S | 4 | U |
| ECMC | ref. bb1050 | N/A | H&E | 143 WSIs | 0.172/40×, 0.345/20×, 0.689/10× | Pixel | S | 7 | U |
| BCRWC | ref. bb1855 | N/A | N/A | 126 WSIs | 1.163/50× | Pixel | S | 4 | U |
| LNM-OSCC | ref. bb0160 | N/A | H&E | 217 WSIs | 0.2467/20×, 0.25/40× | ROI | S | 2 | U |
| OP-SCC-Vanderbilt | ref. bb3310 | N/A | H&E | 50 Patients | 40× | Patient | S | 2 | B |
| Sheffield University | ref. bb1860 | N/A | H&E | 43 WSIs | 0.4952/20× | Slide | S | 4 | I |
| Prostate/Ovary | |||||||||
| PCa-Bulten | ref. bb3350 | Link | H&E, IHC | 102 WSIs, 160 ROIs | 0.24/20× | Pixel | S | 2 | U |
| OV-Kobel | ref. bb3355 | Link | H&E, Ki-67, Mammoglobin B, ER, Mesothelin, MUC5, WT1, p16, p53, Vimentin, HNF-1b | 168 WSIs, 88 TMAs | N/A | Slide | S | 6 | I |
| TCGA-Tolkach | ref. bb3360 | TCGA | H&E | 389 WSIs | 0.25/40× | ROI | S | 3 | U |
| UHZ | ref. bb3365 | Link | H&E | 886 TMAs | 0.23/40× | ROI | S | 5 | U |
| SMS-TCGA | ref. bb2415 | N/A | H&E | 310 WSIs | 20×, 40× | ROI | S | 2 | U |
| TCGA-Arvaniti | ref. bb3370 | TCGA | H&E | 447 WSIs | 20×, 40× | Slide | S | 2 | I |
| TCGA-Yaar | ref. bb2250 | TCGA | H&E | 220 Patients | 20× | Patient | S | 2 | I |
| Pro-RUMC | ref. bb0355 | N/A | H&E | 225 WSIs | 0.16/40× | ROI | S | 2 | B |
| UHZ-PCa | ref. bb1480 | N/A | MIB-1 | 826 Patches | N/A | Patch | S | 2 | I |
| SUH | ref. bb1040 | N/A | H&E | 230 WSIs, 1 103 160 Patches | 10× | ROI | S | 4 | I |
| CSMC | ref. bb1945,ref. bb2245 | N/A | H&E | 513 Patches | 0.5/20× | Pixel | S | 4 | U |
| HUH | ref. bb0365 | N/A | H&E | 28 WSIs | 0.22 | Pixel | S | 2 | U |
| RCINJ | ref. bb3375 | N/A | H&E | 83 WSIs | 20× | Slide | S | 2 | I |
| Pro-Raciti | ref. bb2125 | N/A | H&E | 304 WSIs | 0.5/20× | Slide, ROI | S | 2 | I, U |
| VPC | ref. bb3380 | N/A | H&E | 333 TMAs | 40× | Pixel | S | 4 | U |
| Pro-Campanella | ref. bb0975 | N/A | H&E | 137 376 Patches | 20× | Patch | S | 6 | U |
| UPenn-Yan | ref. bb3385 | N/A | H&E | 43 WSIs | 40× | ROI | S | 2 | U |
| Pro-Doyle | ref. bb3390 | N/A | H&E | 12K ROIs | 0.25/40× | ROI | S | 2 | U |
| UPenn-Doyle | ref. bb3395 | N/A | H&E | 214 WSIs | 40× | ROI | S | 7 | U |
| RUMC-Bulten | ref. bb1370 | N/A | H&E | 1 243 WSIs | 0.24 | Slide | S | 2 | U |
| VGH | ref. bb0595 | N/A | H&E | 305 WSIs | N/A | ROI | S | 5 | U |
| NMCSD+MML+TCGA | ref. bb3400 | N/A | H&E | 1 557 WSIs | 0.25/40×, 0.5/20× | Slide, ROI | S | 4 | I, U |
| OVCARE | ref. bb3405 | N/A | H&E | 354 WSIs | 40× | ROI | S | 5 | U |
| CWU | ref. bb3410 | Link | H&E | 478 WSIs, 120K Patches | 0.504/20× | Patch | S | 3 | I |
| UHC | ref. bb3410 | Link | H&E | 157 WSIs, 120K Patches | 0.231/40× | Patch | S | 3 | I |
| HWN | ref. bb3410 | Link | H&E | 51 WSIs, 120K Patches | 0.264/40× | Patch | S | 3 | I |
| CSMC | ref. bb1770 | N/A | N/A | 625 Patches | N/A | Pixel | S | 4 | U |
| DiagSet-A | ref. bb1365 | By Req. | H&E | 2 604 206 Patches | 5×, 10×, 20×, 40× | Patch | S | 9 | I |
| DiagSet-B | ref. bb1365 | By Req. | H&E | 4 675 WSIs | 0.25/40× | Slide | S | 2 | I |
| DiagSet-C | ref. bb1365 | By Req. | H&E | 46 WSIs | 0.25/40× | Slide | S | 3 | U |
| SICAPv2 | ref. bb1955,ref. bb3415 | Link | H&E | 182 WSIs | 40× | Slide, Pixel | S | 4 | I, U |
| OVCARE-Farahani | ref. bb3420 | N/A | H&E | 485 Patients, 948 WSIs | 40× | Patient, Slide | S | 5 | I |
| University of Calgary | ref. bb3420 | N/A | H&E | 60 Patients, 60 WSIs | 40× | Patient, Slide | S | 5 | I |
| PANDA | ref. bb3425 | Link | H&E | 11 000 WSIs | 40× | ROI | S | 5 | I |
| Thyroid | |||||||||
| UPMC | ref. bb2820 | N/A | Feulgen | 10-20 WSIs | 0.074 | Pixel | S | 3 | U |
| Chen et al. | ref. bb1765 | N/A | N/A | 600 WSIs | 40× | Slide | S | 3 | I |
| TCGA-Hoehne | ref. bb2305 | TCGA | H&E | 482 WSIs | 40× | Slide | S | 4 | I |
| DEC | ref. bb2305 | N/A | H&E | 224 WSIs | 40× | Slide | S | 4 | I |
| ACQ | ref. bb2305 | N/A | H&E | 100 WSIs | 40× | Slide | S | 4 | I |
| Stomach & Colon | |||||||||
| UMCM | ref. bb1135,ref. bb3430 | Link | H&E | 5K Patches | 0.495/20× | Patch | S | 8 | B |
| GLaS | ref. bb0415,ref. bb3435 | Link | H&E | 165 WSIs | 0.62/20× | ROI | S | 5 | I |
| CRCHistoPhenotypes | ref. bb0385,ref. bb1420 | Link | H&E | 10 WSIs, 100 Patches | 0.55/20× | Pixel | S | 4 | I |
| DACHS | ref. bb2175 | N/A | H&E | 3 729 WSIs | 20× | Slide | S | 3 | I |
| NCT-CRC-HE-100K | ref. bb0430,ref. bb2375,ref. bb3430 | Link | H&E | 86 WSIs, 100K Patches | 0.5 | Patch | S | 9 | I |
| NCT-CRC-HE-7K | ref. bb0430,ref. bb2375,ref. bb3430 | Link | H&E | 25 WSIs, 7 180 Patches | 0.5 | Patch | S | 9 | I |
| CoNSeP | ref. bb1420 | Link | H&E | 16 WSIs, 41 Patches | 40× | Pixel | S | 7 | I |
| OSU | ref. bb2195 | Link | H&E, Pan-Cytokeratin | 115 WSIs | 0.061/40× | ROI | S | 2 | U |
| Warwick-CRC | ref. bb0845,ref. bb3440 | Link | H&E | 139 ROIs | 20× | ROI | S | 3 | I |
| HUH | ref. bb3445 | Link | EGFR | 27 TMAs, 1 377 ROIs | 20× | ROI | S | 2 | I |
| CRAG | ref. bb0845 | N/A | H&E | 38 WSIs, 139 Patches | 0.275/20× | Patch | S | 3 | I |
| ULeeds | ref. bb3450,ref. bb3455 | Link | H&E | 27 WSIs | N/A | Slide | S | 3 | B |
| Kather et al. | ref. bb1470 | Link | H&E | 11 977 Patches | 0.5 | Patch | S | 3 | U |
| ZU | ref. bb0710 | By Req. | H&E | 717 ROIs | 0.226/40× | ROI | S | 6 | I |
| KCCH | ref. bb1470 | N/A | H&E | 185 Patients | N/A | Patient | S | 3 | I |
| SC-Takahama | ref. bb1425 | N/A | H&E | 1 019 WSIs | Ma×. of 20× | Pixel | S | 2 | U |
| HUCH | ref. bb0440 | N/A | H&E | 420 Patients | 0.22 | Patient | S | 2 | I |
| RC-Ciompi | ref. bb3460 | N/A | H&E | 74 WSIs | 0.455/200× | ROI | S | 9 | U |
| DHMC-Korbar | ref. bb0390 | N/A | H&E | 1 962 WSIs | 200× | Slide | S | 6 | U |
| CRC-TP | ref. bb1430 | N/A | H&E | 20 WSIs, 280K Patches | 20× | ROI | S | 7 | U |
| CRC-CDC | ref. bb1430 | N/A | H&E | 256 Patches | 20× | Pixel | S | 5 | I |
| SC-Xu | ref. bb2235 | N/A | N/A | 60 WSIs | N/A | ROI | S | 2 | U |
| FAHZU-Xu | ref. bb2240 | N/A | H&E | 13 838 WSIs | 40× | Slide | S | 2 | I |
| Bilkent | ref. bb0415,ref. bb3465 | N/A | H&E | 72 Patches | 20× | Pixel | S | 2 | U |
| DHMC-Wei | ref. bb1500 | N/A | H&E | 1 230 WSIs | 20× | Slide | S | 3 | I |
| Warwick-UHCW | ref. bb1850 | N/A | H&E | 75 WSIs | 0.275/40× | ROI | S | 2 | U |
| Warwick-Osaka | ref. bb1850 | N/A | H&E | 50 WSIs | 0.23/40× | Slide | S | 6 | I |
| GNUCH | ref. bb1675 | N/A | H&E | 94 WSIs, 343 ROIs | N/A | Slide, ROI | S | 4, 2 | I |
| SPSCI | ref. bb3470 | N/A | H&E | 55 WSIs, 251 ROIs | 0.19/40× | ROI | S | 5 | I |
| WSGI | ref. bb2230 | N/A | H&E | 608 WSIs | 0.2517/40× | Slide, Pixel | S | 3, 2 | I, U |
| TBB | ref. bb3475 | N/A | H&E | 44 TMAs | N/A | Slide | S | 3 | I |
| UV | ref. bb3480 | N/A | H&E | 456 WSIs | 40× | Slide | S | 4 | U |
| SC-Sali | ref. bb3485 | N/A | H&E | 1 150 WSIs | N/A | Slide | S | 7 | I |
| SC-Holland | ref. bb1515 | N/A | H&E | 10 WSIs, 1K Patches | 40×, 100× | Slide | S | 2 | B |
| SC-Kong | ref. bb3490 | N/A | H&E | 272 WSIs | 40× | ROI | S | 2 | U |
| SSMH-STAD | ref. bb1215 | N/A | H&E | 50 WSIs | N/A | Slide | S | 2 | B |
| HIUH | ref. bb1155 | N/A | H&E | 8 164 WSIs | 20× | Slide, ROI | S | 3 | I |
| HAH | ref. bb1155 | N/A | H&E | 1K WSIs | 20× | Slide | S | 3 | I |
| SC-Galjart | ref. bb1990 | N/A | H&E | 363 Patients, 1 571 WSIs | 0.25 | Slide | S | 2 | U |
| SC-Zheng | ref. bb2135 | N/A | H&E | 983 WSIs, 10 030 ROIs | 0.96/10× | ROI | S | 5 | U |
| CRC-I-Chikontwe | ref. bb2260 | N/A | H&E | 173 WSIs | 40× | Slide | S | 2 | I |
| CRC-II-Chikontwe | ref. bb2260 | N/A | H&E | 193 WSIs | 40× | Slide | S | 2 | I |
| PLAGH | ref. bb3495 | N/A | H&E | 2 123 WSIs | 0.238/40× | Pixel | S | 4 | U |
| QUASAR | ref. bb3500 | N/A | H&E | 106 268 ROIs | 0.5 | ROI | S | 2 | U |
| CGMH | ref. bb1550 | N/A | H&E | 297 WSIs | 0.229/40× | ROI | S | 2 | U |
| AOEC-RUMC-I | ref. bb2300 | N/A | H&E | 2 131 WSIs | 5×-10× | Slide | M | 5 | I |
| AOEC-RUMC-II | ref. bb2300 | N/A | H&E | 192 WSIs | 5×-10× | Slide, ROI | M, S | 4 | I, U |
| Lizard | ref. bb1335 | Link | H&E | 291 WSIs, | 0.5/20× | Pixel | S | 6 | I |
| YCR-BCIP | ref. bb2175 | N/A | H&E | 889 WSIs | 20× | Slide | S | 2 | I |
| MHIST | ref. bb1310 | By Req. | H&E | 3 152 Patches | 40× | Patch | S | 2 | I |
| YSMH | ref. bb1875 | N/A | H&E | 390 WSIs | 20× | Slide, ROI | S | 5 | I, U |
| ColonPredict-Plus-2 | ref. bb1750 | N/A | H&E | 200 Patients, 2 537 Patches | N/A | Pixel | S | 2 | U |
| PAIP | ref. bb3505 | By Req. | H&E | 47 WSIs | N/A | ROI | S | 2 | U |
| Li et al. | ref. bb1380 | N/A | N/A | 10 894 WSIs, 200 Patches | 0.5/20× | Slide, Pixel | S | 2 | I |
| Stanford Hospital | ref. bb1590 | Link | H&E, p53 IHC | 70 WSIs | 20× | Slide | S | 2 | U |
| IMP Diagnostics Lab. | ref. bb3510,ref. bb3515 | By Req. | H&E | 1133 WSIs | 40× | Slide | S | 3 | I |
| Chaoyang | ref. bb3520 | Link | H&E | 6 160 patches | N/A | Patch | S | 8 | I |
| BERN-GASTRIC-MSI | ref. bb2460 | N/A | H&E | 302 Patients | N/A | Patient | S | 2 | I |
| BERN-GASTRIC-EBV | ref. bb2460 | N/A | H&E | 304 Patients | N/A | Patient | S | 2 | I |
| Bone Marrow | |||||||||
| BM-MICCAI15 | ref. bb3525 | Link | H&E | 11 WSIs | N/A | Pixel | S | 3 | U |
| MICCAI15-Hu | ref. bb2200 | N/A | H&E | 11 WSIs, 1 995 Patches | N/A | Patch | S | 4 | I |
| FAHZU-Hu | ref. bb2200 | N/A | H&E | 24 WSIs, 600 Patches | N/A | Patch | S | 3 | B |
| BM-Hu | ref. bb2200 | N/A | H&E | 84 WSIs | N/A | Slide | S | 2 | I |
| RUMC-Eekelen | ref. bb1805 | N/A | PAS | 24 WSIs | 0.25 | Pixel | S | 6 | U |
| MSKCC | ref. bb1730 | N/A | H&E | 1 578 WSIs | 0.5025, 0.5031/20× | Pixel | S | 7 | I |
| EUH | ref. bb3530 | N/A | Wright’s Stain | 9 230 ROIs | 0.25/40× | ROI | S | 12 | I |
| Frankel et al. | ref. bb3535 | N/A | H&E | 424 WSIs | 40× | Slide | S | 9 | I |
| Internal-STAD | ref. bb3540 | N/A | H&E | 203 WSIs | 0.25/40× | ROI | S | N/A | U |
| MultiCenter-STAD | ref. bb3540 | N/A | H&E | 417 WSIs | 0.46/40× | ROI | S | N/A | U |
| Cervix | |||||||||
| TCGA-Idlahcen | ref. bb1340 | TCGA | H&E | 10 WSIs | 2.5×-40× | Slide | S | 2 | B |
| XH-FMMU | ref. bb3545 | N/A | H&E | 800 WSIs | 4×-40× | ROI | S | 2 | U |
| Pap-Cytology | ref. bb0840 | N/A | N/A | 42 ROIs | 20× | Pixel | S | 2 | U |
| Chen et al. | ref. bb1765 | N/A | N/A | 372 WSIs | 40× | Slide | S | 2 | I |
| OAUTHC | ref. bb2290 | N/A | H&E | 1 331 ROIs | N/A | ROI | S | 2 | I |
| Multi-organ | |||||||||
| MoNuSeg | ref. bb1325 | Link | H&E | 30 WSIs | 40× | Pixel | S | 2 | U |
| UHN | ref. bb1285 | Link | H&E | 1 656 WSIs, 838 644 Patches | 0.504/20× | Slide, Patch | S | 74 | I |
| CPM-15 | ref. bb1420 | Link | H&E | 15 Patches | 20×, 40× | Pixel | S | 2 | U |
| CPM-17 | ref. bb1420 | Link | H&E | 32 Patches | 20×, 40× | Pixel | S | 2 | U |
| ADP | ref. bb0835,ref. bb1535 | By Req. | H&E | 100 WSIs, 17 668 Patches | 0.25/40× | Patch | M | 57 | I |
| Bándi-Dev-Set | ref. bb0990 | Link | H&E, Sirius Red, PAS, Ki-67, AE1AE3, CK8-18 | 100 WSIs | 0.2275, 0.2278, 0.2431, 0.25, 0.2525, 0.5034 | Pixel | S | 6 | U |
| Bándi-Dis-Set | ref. bb0990 | Link | H&E, Alcian Blue, Von Kossa, Perls, CAB, Grocott | 8 WSIs | 0.2431 | ROI | S | 4 | U |
| PanNuke | ref. bb3550,ref. bb3555 | Link | H&E | 20K WSIs, 205 343 ROIs | 40× | ROI | S | 5 | I |
| Salvi-SCAN | ref. bb3340 | Link | H&E | 270 ROIs | 10×, 20×, 40× | Pixel | S | 2 | U |
| TCGA-Nuclei | ref. bb1685,ref. bb3185,ref. bb3560 | Link | H&E | 5 060 WSIs, 1 356 Patches | 0.25/40× | Pixel | S | 14 | U |
| MO-Khoshdeli | ref. bb1815 | Link | H&E | 32 WSIs, 32 Patches | 0.5 | Pixel | S | 2 | U |
| FocusPath | ref. bb0980 | Link | H&E, Trichrome, IRON(FE), Mucicarmine, CR, PAS, AFB, Grocott | 9 WSIs, 8 640 Patches | 0.25/40× | Patch | S | 15 | U |
| Cheng-Jiang | ref. bb3565 | Link | H&E, TCT, IHC | 20 521 WSIs | 10× | Slide | 4 | S | I |
| Stanford-TMA | ref. bb1485,ref. bb3570 | By Req. | H&E, IHC | 6 402 TMAs | N/A | Slide | S | 4 | I |
| TCGA-Courtiol | ref. bb0435 | TCGA | H&E | 56 Patients, 56 WSIs | N/A | Patient, Slide | S | 3 | I |
| BreCaHAD | ref. bb0370 | link | H&E | 170 ROIs | 40× | ROI | S | 2 | U |
| TCGA-Hegde | ref. bb3575 | TCGA | H&E | 60 WSIs | 10× | ROI | S | 10 | U |
| TCGA-Diao | ref. bb3580 | TCGA | H&E | 2 917 WSIs | 20×, 40× | ROI, Pixel | S | 4, 6 | I |
| TCGA-Levine | ref. bb3405 | TCGA | H&E | 668 WSIs | N/A | ROI | S | 5 | U |
| TCGA@Focus | ref. bb0980 | Link | H&E | 1K WSIs, 14 371 Patches | N/A | Patch | S | 2 | I |
| TCGA-Shen | ref. bb3585 | TCGA | H&E | 1 063 WSIs | 20× | Patch | S | 3 | U |
| TCGA-Lerousseau | ref. bb2255 | TCGA | H&E | 6 481 WSIs | 20× | Pixel | S | 3 | U |
| TCGA-Schmauch | ref. bb3590 | TCGA | H&E | 10 514 WSIs | N/A | Slide | S | 28 | I |
| MO-Khan | ref. bb1010 | N/A | H&E | 60 WSIs | 20×, 40× | Pixel | S | 3 | U |
| MESOPATH/MESOBANK | ref. bb0435 | N/A | HES | 2 981 Patients, 2 981 WSIs | 40× | Patient, Slide | S | 3 | U |
| Mo-Campanella | ref. bb0975 | N/A | H&E, SDF-1, TOM20 | 249 600 Patches | 20× | Patch | S | 6 | U |
| BWH-TCGA-MO | ref. bb2275 | N/A | H&E | 25 547 WSIs | N/A | Slide | S | 18 | I |
| BWH-Lu | ref. bb1545 | N/A | H&E | 19 162 WSIs | 20×, 40× | Slide | 2 | I | |
| Feng et al. | ref. bb2390 | N/A | H&E, IHC | 500 WSIs | 20× | Slide | S | 10 | B |
| SegSet | ref. bb1365 | N/A | H&E | 30 WSIs | 0.25/40× | Pixel | S | 2 | U |
| LC25000 | ref. bb3595 | Link | H&E | 25 000 Patches | N/A | Patch | S | 5 | B |
| OCELOT-CELL | ref. bb3600 | Link | H&E | 306 WSIs, 673 Patches | 0.2 | ROI | S | 2 | I |
| OCELOT-TISSUE | ref. bb3600 | Link | H&E | 306 WSIs, 673 Patches | 0.2 | Pixel | S | 3 | I |
| Other | |||||||||
| MUH | ref. bb2375,ref. bb3605 | N/A | N/A | 18 365 Patches | 14.14/100× | Patch | S | 15 | I |
| UPenn | ref. bb3610 | N/A | H&E | 209 Patients | 20× | Patient | S | 2 | I |
| CMTHis | ref. bb1530 | N/A | H&E | 352 ROIs | 40×, 100×, 200×, 400× | ROI | S | 2 | I |
| Heidelberg University | ref. bb3615 | N/A | H&E | 431 WSIs | N/A | ROI | S | 2 | U |
| CHOA | ref. bb3620 | N/A | H&E | 43 WSIs | 10× | Slide | S | 4 | I |
| Han-Wistar Rats | ref. bb2295 | N/A | H&E, ISH | 349 WSIs | 40× | Slide | S | 2 | U |
| Osteosarcoma | ref. bb2430 | Link | H&E | 1 144 ROIs | 10× | ROI | S | 3 | I |
| UPenn+OSU+UH | ref. bb3625 | N/A | H&E | 2 358 WSIs | 40× | Slide | S | 4 | I |
| Kaggle 2018 Data Science Bowl | ref. bb3630 | Link | DAPI, Hoechst, H&E | 670 WSIs | N/A | Pixel | S | 2 | U |
| ALL-IDB2 | ref. bb3635,ref. bb3640 | By Req. | N/A | 260 ROIs | 300× – 500× | Slide | S | 2 | B |
Compilation of tasks found in different Co-Path papers categorized by organ (see 9.10)
| References | Tasks | Disease Specification | Methods |
|---|---|---|---|
| Basal/Epithelium | |||
| ref. bb3025 | Detection | Metastasis | End-to-end classifier using cascaded CNNs |
| ref. bb3090 | Detection | Metastasis | Unsupervised learning via auto-encoder |
| ref. bb3080 | Disease diagnosis | Melanoma, intra-dermal, compound, junctional nevus | CNN-based patch classifier |
| ref. bb3645 | Nuclei subtype classification | lymphocyte, stromal, artefact, cancer | CRImage and TTG/CNx for cell identification and classification |
| ref. bb3650 | Tissue subtype classification | Epithelial, stromal tissues, Spitz, conventional melanocytic lesions | Integration of CNN and HFCM segmentation |
| ref. bb3085 | Tissue subtype classification | Epithelial, stromal tissues, Spitz, conventional melanocytic lesions | CNN-based classifier with transfer learning |
| ref. bb0435 | Patient prognosis | Epithelioid, sarcomatoid, biphasic in mesothelium, distant metastatic recurrence | ResNet classifier with transfer learning |
| ref. bb1330 | Patient prognosis | Epithelioid, sarcomatoid, biphasic in mesothelium, distant metastatic recurrence | Combination of DNN and RNN for feature processing |
| ref. bb1385 | Tumor segmentation | Tumor, epidermis, dermis, background | FCN based segmentation |
| ref. bb1520 | Classification | Nevi, melanoma | CNN-based classifier |
| Bladder | |||
| ref. bb2110 | Classification | Papillary urothelial carcinoma LG/HG | Combination of CNN and LSTM |
| ref. bb1935 | Segmentation | Voronoi objects, edges, background regions | CycleGAN with U-Net segmentation |
| ref. bb1325 | Nuclei Segmentation | Nuclear, Non-nuclear, Boundary | CNN-based classifier with AJI evaluation |
| ref. bb0730 | Tissue Subtype Classification | Double negative, basal, luminal, luminal p53-like | ResNet variation classifier |
| Brain | |||
| ref. bb0380 | Classification | Glioblastoma multiforme, LG glioma | Elastic net classifier with weighted voting |
| ref. bb3655 | Classification | LGG Grade II/III, GBM | Modular CNN-ensemble network |
| ref. bb0710 | Classification | LGG and GBM | CNN-based classifier with transfer learning |
| ref. bb3660 | Classification | Glioma grading III, IV, V | SVM classifier |
| ref. bb1285 | Classification | Tissue feature correlation analysis | CNN-based classifier with transfer learning |
| ref. bb3115 | Patient prognosis | Tissue feature correlation analysis | Densenet121 classifiers, initialized with imageNet pre-trained weights |
| ref. bb1225 | Patient prognosis | IDH mutation | Survival CNN with genetic biomarker data integration |
| ref. bb0425 | Patient prognosis | Survival period for glioblastoma | CNN-based patch classifier |
| ref. bb2420 | Patient prognosis | GBM prognostic index | Fusion network of genome, histopathology, and demography |
| ref. bb3100 | Patient prognosis | Glioblastoma Multiforme | Custom CNN classifier |
| ref. bb2815 | Patient prognosis/Tissue subtype classification | Oligodendroglioma, IDH-mutant/wild type astrocytoma | CNN-based classifier |
| ref. bb3015 | Segmentation | Superior Middle Temporal Gyri in the temporal cortex | Semi-supervised active learning(SSL) |
| Mouth/Esophagus | |||
| ref. bb3345 | Tissue subtype classification | Stroma, lymphocytes, tumor, mucosa, kerClassificationatin pearls, blood, adipose | Modified AlexNet patch classifier with active learning |
| ref. bb1475 | Disease Diagnosis | Barrett esophagus no dysplasia, esophageal adenocarcinoma, normal, Barrett esophagus with dysplasia | Attention based classifier |
| ref. bb3665 | Patient prognosis | Oropharyngeal squamous cell carcinoma | Computational cell cluster graph |
| ref. bb1860 | Segmentation | Oral epithelial dysplasia (OED) | HoVer-Net+, a deep learning framework consists of an encoder branch, and three decoder branches |
| Breast | |||
| ref. bb0995,ref. bb2480 | Detection | Benign, malignant | CNN-based patch classifier |
| ref. bb1530,ref. bb3670,ref. bb3675 | Detection | Benign, malignant | CNN classifier with transfer learning |
| ref. bb3215 | Detection | Benign, malignant | CNN-based pixel classifier |
| ref. bb2575 | Detection | Benign, malignant | Pre-trained AlexNet with automatic label query |
| ref. bb3680 | Detection | Benign, malignant | Pre-trained AlexNet with Bi-LSTM classifier |
| ref. bb1650 | Detection | Benign, malignant | Combination of CNN classifier and U-Net segmentation |
| ref. bb3685 | Detection | Benign, malignant | CNN-based classifier |
| ref. bb2120 | Detection | Benign, malignant | 3 stage LSTM-RNN classifier |
| ref. bb2220 | Detection | Benign, malignant | Attention-based MIL model |
| ref. bb1510 | Detection | Benign, malignant | Tri-branched ResNet model |
| ref. bb1170 | Detection | Benign, malignant | Combination of CNN and hand-crafted features |
| ref. bb1130 | Detection | Custom CNN classifier with Quasi-Monte Carlo sampling | |
| ref. bb1985 | Detection, Patient Prognosis | Tumor, Normal/ Tumor-infiltrating lymphocytes | U-Net based classifier |
| ref. bb1725 | Detection | Mitosis | Multi-scale custom CNN classifier |
| ref. bb3280 | Detection | Invasive Ductal Carcinoma | Bayesian Convolution Neural Networks |
| ref. bb2985 | Binary classification | Breast cancer to axillary lymph nodes (ALNs) | Pre-trained architectures: DenseNet121, ResNet50, VGG16, Xception and lightweight convolutional neural network (LCNN) |
| ref. bb1865 | Tumor Segmentation and Classification | Breast Cancer Metastases | SOTA methods, designed MLVDeepLabV3+ |
| ref. bb3265 | Segmentation | Segmentation of the malignant nuclei within each tumor bed | Mask regional convolutional neural network (Mask R-CNN) |
| ref. bb3690 | Segmentation | Segmentation of multiple subtypes on breast images | Deep Multi Magnification Network (DMMN), CNN architecture |
| ref. bb1525 | Detection | Metastasis/ Micro, Macro | CNN-based pixel classifier |
| ref. bb2130 | Detection | Metastasis/ Micro, Macro | Resnet with transfer learning |
| ref. bb3695 | Detection | Metastasis/ Micro, Macro | Combination of CNNs with LSTM-RNN, DCNN-based classifier |
| ref. bb1645 | Detection | Cancer metastasis detection | MIL+RNN classifier, Neural conditional random field |
| ref. bb3700 | Detection | Cancer metastasis detection | CNN with attention mechanism |
| ref. bb2465 | Detection | Metastasis in sentinel lymph node | CNN with Random Forest classifier |
| ref. bb3705 | Detection | Invasive ductal carcinoma | CNN-based patch classifier |
| ref. bb1490 | Detection | Invasive ductal carcinoma | ResNet with transfer learning |
| ref. bb3205 | Detection | Invasive ductal carcinoma | CNN-based random forest classifier |
| ref. bb3330 | Detection | Invasive ductal carcinoma | Autoencoder network |
| ref. bb0360 | Detection | Macrometastasis, micrometastasis, isolated tumor cells, negative | Customized InceptionV3 classifier |
| ref. bb1125 | Detection | Mitosis detection | CNN classifier with two-phase training |
| ref. bb1345 | Detection | Mitosis detection | Task-based CNN ensemble |
| ref. bb1890 | Detection | Mitosis detection | CNN-based random forest classifier |
| ref. bb1720 | Detection | Mitosis detection | CNN classifier with transfer learning |
| ref. bb1895 | Detection | Mitosis detection | Multi-stage RCNN classifier |
| ref. bb1900 | Detection | Mitosis detection | FCN classifier |
| ref. bb1905 | Detection | Mitosis detection | Adaptive Mask RCNN |
| ref. bb1910 | Detection | Mitosis detection | CNN-based patch classifier |
| ref. bb1000 | Detection | Mitosis detection | Combination of DCNN network |
| ref. bb1920 | Detection | Mitosis detection | R2U-Net based regression model |
| ref. bb1175 | Classification | Epithelium, Stroma | Magnification invariant CNN classifier |
| ref. bb1180 | Classification | Benign, malignant | CNN classifier interleaved with squeeze-excitation modules (SENet) |
| ref. bb3710 | Classification | Adenosis, fibroadenoma, phyllodes tumors, tubular adenoma, ductal, lobular, mucinous, papillary | Inception Recurrent Residual Convolutional Neural Network (IRRCNN) |
| ref. bb3715 | Classification | Adenosis, fibroadenoma, phyllodes tumors, tubular adenoma, ductal, lobular, mucinous, papillary | Custom DenseNet classifier |
| ref. bb3285 | Classification | normal tissue, benign lesion, in situ carcinoma, and invasive carcinoma | Custom multiclass dense layer classifier based on Xception network |
| ref. bb3720 | Disease diagnosis | Adenosis, fibroadenoma, phyllodes tumors, tubular adenoma, ductal, lobular, mucinous, papillary | Two-stage ResNet classifier (MuDeRN) |
| ref. bb1655 | Disease diagnosis | Benign, malignant | Ensemble of CNN classifiers |
| ref. bb1505 | Disease diagnosis | Adenosis, fibroadenoma, phyllodes tumors, tubular adenoma, ductal, lobular, mucinous, papillary | CNN-based classifier with transfer learning |
| ref. bb3725 | Disease diagnosis | Adenosis, fibroadenoma, phyllodes tumors, tubular adenoma, ductal, lobular, mucinous, papillary | Class structured DCNN |
| ref. bb2145 | Disease diagnosis | Adenosis, fibroadenoma, phyllodes tumors, tubular adenoma, ductal, lobular, mucinous, papillary | Two stage classification and selection network |
| ref. bb3730 | Disease diagnosis | Adenosis, fibroadenoma, phyllodes tumors, tubular adenoma, ductal, lobular, mucinous, papillary | Domain adaptation based on representation learning |
| ref. bb3735 | Disease diagnosis | Benign, in-situ, invasive carcinoma | CNN-based classifier with gravitational loss |
| ref. bb3740 | Disease diagnosis | Benign, in-situ, invasive carcinoma | CNN ensemble with LightGBM |
| ref. bb3745 | Disease diagnosis | Benign, in-situ, invasive carcinoma | InceptionV3 classifier using dual path network |
| ref. bb0130 | Disease diagnosis | Usual ductal hyperplasia, ductal carcinoma in situ | Logistic regression with Lasso regularization |
| ref. bb1845 | Disease diagnosis | Proliferation score (1, 2, or 3) | Encoder-decoder with Gaussian Mixture model |
| ref. bb1295 | Disease diagnosis | Low risk/High risk | Logistic regression using morphological features |
| ref. bb2475 | Disease diagnosis | normal, benign, in situ carcinoma, invasive carcinoma | Hybrid CNN classifier |
| ref. bb3240 | Disease diagnosis | Proliferative without atypia, atypical hyperplasia, ductal / lobular carcinoma in situ, invasive carcinoma | Cascade of VGG-Net like classifier |
| ref. bb3750 | Disease diagnosis | Benign, malignant | CNN-based classifier with fourier pre-processing |
| ref. bb2105 | Disease diagnosis | Benign, malignant | Combination of CNN and LSTM classifiers |
| ref. bb3755 | Disease diagnosis | Tumour, normal | Metric learning using similarities |
| ref. bb1135 | Classification | Clinically relevant classes | CNN-based patch classifier with aggregation |
| ref. bb1315 | Classification | Benign, in-situ, invasive carcinoma | Scale-based CNN classifier |
| ref. bb2980 | Classification | Normal, benign, in situ, invasive carcinoma | Combination of patch and image level CNN |
| ref. bb3235 | Classification | Normal, benign, DCIS, invasive ductal carcinoma (IDC) | Context-aware stacked CNN |
| ref. bb3760 | Classification | Benign, in-situ, invasive carcinoma | CNN-based classifier with dimensionality reduction |
| ref. bb0315 | Classification | Benign, in-situ, invasive carcinoma | MIL with auto-regression |
| ref. bb2115 | Classification | Benign, in-situ, invasive carcinoma | Parallel network with CNN-RNN |
| ref. bb1320 | Classification | Benign, in-situ, invasive carcinoma | Hybrid Convolutional and Recurrent NN |
| ref. bb1540 | Classification | Benign, in-situ, invasive carcinoma | CNN-based patch classifier |
| ref. bb3765 | Classification | Benign, in-situ, invasive carcinoma | Convolutional capsule network |
| ref. bb3165 | Classification | Benign, in-situ, invasive carcinoma | Combination of residual and spatial model |
| ref. bb3770 | Classification | Benign, in-situ, invasive carcinoma | Custom CNN patch classifier |
| ref. bb1065 | Classification | Benign, in-situ, invasive carcinoma | CNN classifier with bidirectional LSTM |
| ref. bb3775 | Classification | Tumor, non-tumor | Custom CNN-based classifier |
| ref. bb1685 | Nuclei segmentation | N/A | UNet segmentation with GAN patch refinement |
| ref. bb1935 | Nuclei segmentation | N/A | UNet segmentation with CycleGAN domain transfer |
| ref. bb1785 | Nuclei segmentation | N/A | Ensemble of several CNNs with different architectures |
| ref. bb1325 | Nuclei segmentation | Normal, malignant, dysplastic epithelial, fibroblast, muscle, inflammatory, endothelial, miscellaneous | Sequential CNN network |
| ref. bb3210 | Nuclei segmentation | Normal, malignant, dysplastic epithelial, fibroblast, muscle, inflammatory, endothelial, miscellaneous | Custom encoder-decoder model |
| ref. bb3010 | Detection, Segmentation | DCIS and invasive cancers | IM-Net for DCIS detection and segmentation |
| ref. bb1425 | Tumour segmentation | Tumor, Normal | U-Net segmentation with GoogleNet patch level feature extraction |
| ref. bb1840 | Tumour segmentation | Normal, benign, in situ carcinoma or invasive carcinoma | Global and local ResNet feature extractors, FCN with auto zoom |
| ref. bb1800 | Tumour segmentation | Normal, benign, in situ carcinoma or invasive carcinoma | Global and local ResNet feature extractors, FCN with auto zoom |
| ref. bb1870 | Segmentation | Breast cancer | U-Net, Residual Multi-Scale (RMS) |
| ref. bb1745 | Segmentation | Ki67 detection for breast cancer | U-NET, piNET |
| ref. bb2835 | Tumour segmentation | Non-tumor, ductal carcinoma in situ (DCIS), invasive ductal carcinoma (IDC), lobular carcinoma in situ (LCIS), invasive lobular carcinoma (ILC) | Ensemble of CNN with atrous spatial pyramid encoding |
| ref. bb1560 | Classification | Prediction of HER2 status and trastuzumab response | CNN classifier with Inception v3, Transfer learning |
| ref. bb1575 | Tissue subtype classification | Classifying cancerous tissues | weakly supervised approach, Multiple Instance Learning (MIL) model, Transfer learning pre-trained models (Trans-AMlL), VGG, DenseNet, ResNe |
| ref. bb3200 | Tissue subtype classification | 24 different tissues | Ensemble of different CNN architectures with transfer learning |
| ref. bb2225 | Tissue subtype classification | Proliferative without atypia, atypical hyperplasia, ductal / lobular carcinoma in situ, invasive carcinoma | Multi-class MIL |
| ref. bb3250 | Disease Diagnosis | Normal, Benign, Atypical, Ductal Carcinoma In Situ, Invasive | CNN-based classifier using graphical representation |
| ref. bb2180 | Tissue subtype classification | Estrogen, Progesterone, Her2 receptor | Style invariant ResNet classifier |
| ref. bb0135 | Tissue subtype classification | Negative, micrometastasis, macrometastasis, isolated tumor cell cluster (ITC) | Custom CNN-based classifier |
| ref. bb1735 | Tissue subtype classification | Adipose regions, TDLU regions, acini centroid | UNet based CNN classifier |
| ref. bb3255 | Classification, Segmentation | Benign, Atypical (flat epithelial atypia, atypical ductal hyperplasia), Malignant (ductal, in situ, invasive) | Graphical neural networks |
| ref. bb3220 | Tissue subtype classification | Basal-like, HER2-enriched, Luminal A, and Luminal B | CNN-based classifier with PCA |
| ref. bb3780 | Classification | Malignant, normal | CNN classifier with transfer learning |
| ref. bb1045 | Segmentation | Stain normalization | Style transfer using CycleGAN, Relevance vector machine |
| ref. bb2160 | Segmentation | Realistic patch generation | GAN based architecture |
| ref. bb3785 | Classification | Processing technique comparison | Comparison of color normalization methods |
| ref. bb1660 | Classification | 19 histological types, HER2-, HER2+, PR+, PR- | Graph CNN slide level classifier |
| ref. bb1670 | Classification | normal, benign, in situ, and invasive | Dynamic Deep Ensemble CNN |
| ref. bb2315 | Binary Classification | Breast cancer | Transformer based MIL (TransMIL) |
| Liver | |||
| ref. bb2805 | Disease diagnosis, Nuclei segmentation | G0, G1, G2, G3, G4 (HCC grade) | BoF-based classifier |
| ref. bb2485 | Disease diagnosis | Hepatocellular/cholangio carcinoma | CNN-based end to end diagnostic tool |
| ref. bb3095 | Nuclei segmentation | N/A | CycleGAN based segmentation |
| ref. bb1715 | Tissue segmentation | Background, tumor, tissue and necrosis | UNet with color deconvolution |
| ref. bb3790 | Tissue segmentation | Steatosis droplet | Mask-RCNN segmentation |
| ref. bb3290 | Classification | Stain normalization | Relevance vector machine |
| ref. bb2185 | Classification | Hematoxylin, eosin, unstained RBC | Linear discriminant classifier |
| ref. bb2190 | Classification | Stain style transfer | CycleGAN based architecture, CycleGAN with perceptual embedding consistency loss |
| Lymph Nodes | |||
| ref. bb1025 | Detection | Detection and quantification of Lymphocytes | U-Net and SegNet with VGG16 and Resnet50 and pre-trained weights of ImageNet |
| ref. bb1190 | Detection | Metastasis | Ensemble of CNNs with different architectures |
| ref. bb3795 | Detection | Metastasis | Custom CNN for discrimitive feature learning |
| ref. bb1355 | Detection | Metastasis | DCNN classifier |
| ref. bb0485 | Detection | Metastasis | CNN-based patch classifier |
| ref. bb3800 | Detection | Metastasis (Isolated / micro / macro) | Variants of ResNet/GoogleNet |
| ref. bb2825 | Disease diagnosis | Hyperplasia, small B cell lymphomas | Bayesian NN with dropout variance |
| ref. bb1855 | Tissue segmentation | Keratin, subepithelial, epithelial, background | Custom CNN model |
| ref. bb1835 | Tumor segmentation | normal, metastatic | Representation-Aggregation Network with LSTM |
| ref. bb3805 | Classification, Segmentation | Domain shift analysis for breast tumour | Comparison of CNN models, data augmentation, and normalization techniques |
| ref. bb1015 | Classification, Segmentation | Stain normalization | Deep Gaussian mixture color normalization model |
| ref. bb2170 | Classification, Segmentation | Stain normalization | GAN, stain-style transfer network |
| ref. bb3810 | Classification, Segmentation | Similar image retrieval | Siamese network |
| ref. bb2380 | Classification | metastatic tissue | Contrastive predictive coding, Autoregressor PixelCNN |
| ref. bb0830 | Prognosis of lymph node metastasis | lymph node metastasis of papillary thyroid carcinoma | Transformer-Guided Multi-instance Learning, Attention-based mutual knowledge distillation |
| Prostate/Ovary | |||
| ref. bb2990 | Tissue subtype classification | Gland, Gland border region, or Stroma | SVM with RBF kernel, CNN classifiers |
| ref. bb1770 | Tissue subtype classification | stromal areas (ST), benign/normal (BN), low-grade pattern (G3) and high-grade pattern (G4) cancer | CNN classifiers using a modified U-Net architecture |
| ref. bb1775 | Detection | Tumor, No-tumor | Generative Adversarial Network named GAN-CS |
| ref. bb0365,ref. bb1370 | Detection | Tumor, Normal | CNN-based classifier with transfer learning, MIL model |
| ref. bb1350 | Detection | Pancreatic adenocarcinoma | Custom CNN classifier |
| ref. bb3390 | Detection | Probabilistic boosting tree with active learning | |
| ref. bb3410 | Detection | benign glandular, nonglandular, tumor tissue | Pre-trained and validated model based on InceptionResNetV2 convolutional architecture |
| ref. bb1945 | Disease diagnosis | Prostate cancer (benign, LG, HG) | Region based CNN classifier |
| ref. bb3815 | Disease diagnosis | Gleason Score (G6-G10) | Comparison of commonly used CNN models |
| ref. bb3815 | Disease diagnosis | Gleason Score (0-5) | Active Learning Framework |
| ref. bb3360 | Disease diagnosis | Invasive carcinoma, Benign (glandular, non-glandular, stromal, seminal vesicles, ejaculatory ducts, high-grade prostatic intraepithelial neoplasia, HGPIN), intraductal carcinoma | NASNetLarge classifier with transfer learning |
| ref. bb3370 | Disease diagnosis | Gleason grading (3, 4, 5) | ResNet classifier with symmetric domain adaptation |
| ref. bb3400 | Disease diagnosis | Gleason grading (3, 4, 5) | Two-stage deep learning system |
| ref. bb3820 | Disease diagnosis | Gleason grading (3, 4, 5) | Multi-scale U-Net for pixel-wise Gleason score prediction |
| ref. bb1040 | Disease diagnosis | Gleason grading (3, 4, 5) | CNN classifier with CycleGAN |
| ref. bb2245 | Disease diagnosis | Gleason grading (3, 4, 5) | Attention-based MIL classifier |
| ref. bb3380 | Disease diagnosis | Benign, Gleason Grades 3-5 | Ensemble of CNN classifiers |
| ref. bb3385 | Disease diagnosis | Gleason grades 1-5 | K-NN classifier using statistical representation of Homology Profiles |
| ref. bb0595 | Disease diagnosis | High-grade serous ovarian, clear cell ovarian, endometrioid (ENOC), low-grade serous, mucinous carcinoma | Two-staged CNN with RF classifier |
| ref. bb3825 | Disease diagnosis | Low risk Gleason score (6-7), high risk (8-10) | Information retrieval using TF-IDF |
| ref. bb2840 | Gland segmentation | Gleason Score (1-5) | Image analysis using mathematical morphology |
| ref. bb1365 | Tissue subtype classification | Diagset A : scan background (BG), tissue background (T), normal, healthy tissue (N), acquisition artifact (A), or one of the 1-5 Gleason grades (R1-R5), Diagset B: presence of cancerous tissue on the scan (C) or lack thereof (NC), Diagset C:containing cancerous tissue (C), not containing cancerous tissue (NC), or uncertain and requiring further medical examination (IHC) | CNNs, a variant of fully-convolutional VDSR networks, AlexNet, VGG16/19, ResNet50, InceptionV3 |
| ref. bb1020 | Tissue subtype classification | Epithelial, stromal | Multiresolution segmentation |
| ref. bb3395 | Tissue subtype classification | Gleason grade 3-5, Benign Epithelium (BE), Benign stroma (BS), Tissue atrophy (AT), PIN | Cascaded approach |
| ref. bb2125 | Clinical validation | Cancer, non-cancerous | Two-stage MIL-RNN classifier |
| ref. bb2250 | Prediction of treatment response | Positive, negative (response to platinum chemotherapy) | Ensemble of RBF+SVM and MIL-based CNN classifier |
| Kidney | |||
| ref. bb3130 | Detection | Tumor, normal | DCNN based classifier |
| ref. bb3135 | Detection | Antibody mediated rejection | CNN-based classifier |
| ref. bb2140 | Classification | Abnormalities of blood chemistry, Kidney function and dehydration | k Nearest Neighbour (kNN), Long short-term memory (LSTM) |
| ref. bb1755 | Classification, Segmentation | Cancer | Modified U-Net CNN model |
| ref. bb1975 | Detection | Glomeruli boundaries | CNN-based classifier with center point localization |
| ref. bb3830 | Detection | Glomeruli and Nuclei | Anchor Free Backbone + center point localization |
| ref. bb1940 | Prognosis | Survival rate for renal cell carcinoma | Random forest classifier for nuclei detection |
| ref. bb3835 | Prognosis | Survival rate | Kaplan-Meier analysis |
| ref. bb0420 | Classification | Sclerosed glomeruli, tubulointerstitium | Laplacian-of-Gaussian method for blob detection |
| ref. bb1970 | Classification | Glomerulus, lymphocytes | Different architectures of standard CNN with patient privacy preservation |
| ref. bb1980 | Classification | Non-glomerular tissue, normal glomeruli, sclerosed glomeruli | U-Net based classifier |
| ref. bb1610 | WSI representation and classification | Kidney Chromophobe Renal Cell Carcinoma (KICH), Kidney Renal Clear Cell Carcinoma (KIRC) and Kidney Renal Papillary Cell Carcinoma(KIRP) | hierarchical global-to-local clustering, weakly-supervised |
| ref. bb3125 | Segmentation | Glomeruli | Cascaded UNet model |
| ref. bb0760 | Segmentation | Glomeruli, sclerotic glomeruli, empty Bowman’s capsules, proximal tubuli, distal tubuli, atrophic tubuli, undefined tubuli, capsules, arteries, interstitium | Ensemble of U-Net |
| ref. bb2315 | Multiple Classification | 3 cancer types | Transformer based MIL (TransMIL) |
| Lung | |||
| ref. bb1205 | Detection | Lymphocyte richness | Unsupervised classifier using convolutional autoencoder |
| ref. bb1695 | Segmentation, Classification | Mitosis, ND, LUAD, LUSC | Deep residual aggregation network with U-Net |
| ref. bb3315 | Classification | Squamous and nonsquamous nonsmall cell | Inception V3 |
| ref. bb1760 | Detection | Tumor, cell Detection | U-Net |
| ref. bb2850 | Classification | Cancer | Decision Tree, AdaBoost and XGBoost |
| ref. bb2490 | Disease diagnosis | LUAD, LUSC | Deep CNN with transfer learning |
| ref. bb3840 | CNN ensemble with random forest aggregation | ||
| ref. bb1290 | ML models with Cox hazard model | ||
| ref. bb1695 | Deep residual aggregation with U-Net | ||
| ref. bb2265 | Prognosis | Malignant, Normal | MIL-based CNN classifier |
| ref. bb0375 | Disease diagnosis | Lepidic, acinar, papillary, micropapillary, solid, benign | CNN-based patch classifer |
| ref. bb1235 | Disease diagnosis | Acinar, micropapillary, solid, cribriform, non-tumor | CNN-based classifier |
| ref. bb2490 | Prognosis | STK11, EGFR, SETBP1, TP53, FAT1, KRAS, KEAP1, LRP1B, FAT4, NF1 | Deep CNN with transfer learning |
| ref. bb3845 | Segmentation | Characterizing spatial arrangement features of the immune response | Watershed-based model |
| ref. bb1070,ref. bb1685,ref. bb1935 | Segmentation | N/ANuclei of tumor cells, stromal cells, lymphocytes, macrophages, blood cells, karyorrhexis | UNet segmentation with GAN patch refinement, UNet segmentation with CycleGAN domain transfer Mask-RCNN based classifier |
| ref. bb2705 | Classification | Tumor cell, stromal cell, lymphocyte | CNN |
| ref. bb3850 | Classification | LUAD, LUSC | Pre-trained DenseNet |
| ref. bb1610 | WSI representation and classification | Lung Adenocarcinoma (LUAD) and Lung Squamous Cell Carcinoma (LUSC) | hierarchical global-to-local clustering, weakly-supervised |
| ref. bb1290 | Prognosis | Pathology grade, non-small cancer Recurrence prediction for non-small cell cancer Lung squamous cell carcinoma Tumor cell, stromal cell, lymphocyte | Various ML models with Cox hazard model |
| ref. bb0445 | CNN-based classifier | ||
| ref. bb1290 | Custom CNN | ||
| ref. bb1695 | Cox regression model | ||
| ref. bb2855 | Prognosis | Squamous cell | Self-supervised pre-trained model, HANet |
| ref. bb1625 | Segmentation | Stain normalization | CNN, LSTM based feature aware normalization |
| ref. bb3855 | Classification | Cancer, normal | High resolution heatmaps from CNN |
| ref. bb2315 | Binary Classification | LUSC/LUAD subtypes classification | Transformer based MIL (TransMIL) |
| ref. bb2315 | Detection | carcinomas and benign tissue | Transfer Learning |
| Paneras | |||
| ref. bb1285 | Classification | Feature correlation analysisNuclei, antigen, cytoplasm, blood, ECM | CNN-based classifier with transfer learningNon-linear tissue component discrimination |
| ref. bb1705 | Classification | Immunopositive tumor, immunonegative tumor, non-tumor | U-Net |
| ref. bb1665 | Tissue Classification | benign lung tissues (LN), lung adenocarcinomas (LAC), lung squamous cell carcinomas (LSCC), benign colonic tissues (CN), and colon adenocarcinomas (CAC) | Pyramid Deep-Broad Learning |
| Thyroid | |||
| ref. bb2820 | Classification | Follicular lesion (FA, FTC), normal | Radial based SVM classifier |
| ref. bb2305 | Classification, Detection | Unknown/have mutation(BRAF+), dont have mutation (BRAF-)/ have fusion(NTRK+), dont have fusion (NTRK-) | Attention-based deep multiple instance learning classifier, DenseNet121 |
| ref. bb2310 | Classification, Detection | Tumor, Healthy/papillary, follicular, poorly differentiated, anaplastic/have mutation(BRAF+), dont have mutation (BRAF-) | Multi Instance Learning (MIL) |
| Stomach/Colon | |||
| ref. bb2195,ref. bb2235,ref. bb2240 | Detection | Cancer | Combination of CNN and MIL, InceptionV3 classifier with conditional GANs, Generalized mean model with parallel MIL |
| ref. bb1470 | Detection | Microsatellite instability | ResNet with transfer learning |
| ref. bb2265 | Prognosis | Malignant, normal | MIL-based CNN classifier |
| ref. bb0385 | Detection | Nucleus | Space-constrained CNN |
| ref. bb1215 | Detection | Adenocarcinoma, normal | CNN-based classifier |
| ref. bb1515 | Detection | Carcinoma, benign | CNN-based classifier |
| ref. bb3495 | Detection | High-grade intraepithelial neoplasia | Deep learning classifier with ResNet backbone |
| ref. bb1380 | Detection | Gastric cancer | DLA34, Hybrid and Weak supervision Learning method |
| ref. bb2175 | Detection | Tumor-bearing tissue, Non-tumor tissue | SuffleNet with end-to-end learning method |
| ref. bb1140 | Detection | BRAF mutational status and microsatellite instability | Swarm Learning |
| ref. bb2375 | Classification | N/A | Encoder, Resnet18 |
| ref. bb1750 | Classification | Colorectal carcinoma, Colorectal cancer | Weakly supervised neural network named comparative segmentation network (CompSegNet), U-Net |
| ref. bb1665 | Classification of tissue | Cadipose (ADI), background (BACK), debris (DEB), lymphocytes (LYM), mucus (MUC), smooth muscle (MUS), normal colon mucosa (NORM), cancer-associated stroma (STR), and colorectal adenocarcinoma epithelium (TUM) | Pyramid Deep-Broad Learning |
| ref. bb1875 | Segmentation | colorectal cancer | CNN, Sliding window method, U-Net+ + |
| ref. bb1555 | Segmentation, Classification | Colorectal cancer | U-Net16/19 network with a VGG-16/19 net as backbone |
| ref. bb3470 | Disease diagnosis | Non-epithelial normal, normal gastric epithelium, neoplastic gastric epithelium/tubular gastric adenocarcinoma, solid-type gastric adenocarcinoma, diffuse/discohesive gastric carcinoma | Custom CNN classifier |
| ref. bb0710 | Disease diagnosis | Adenocarcinoma, mucinous carcinoma, serrated carcinoma, papillary carcinoma, and cribriform comedo-type carcinoma | CNN-based classifier with transfer learning |
| ref. bb1155 | Disease diagnosis | Adenocarcinoma, adenoma, non-neoplastic | CNN classifier with RNN aggregation |
| ref. bb3440 | Disease diagnosis | Colorectal cancer | CNN |
| ref. bb1500 | Disease diagnosis | Celiac disease, nonspecific duodenitis | ResNet patch classifier |
| ref. bb2260 | Disease diagnosis | Cancer | Multiple instance learning |
| ref. bb1675 | Disease diagnosis | Adenocarcinoma, poorly cohesive carcinoma, normal gastric mucosa | Multi-scale receptive field model |
| ref. bb2230 | Disease diagnosis | Dysplasia, Cancer | Multi-instance deep learning classifier |
| ref. bb0410 | Disease diagnosis | Healthy, adenomatous, moderately differentiated, moderately-to- poorly differentiated, and poorly differentiated | Multi-task classifier |
| ref. bb3455 | Disease diagnosis | Adenocarcinoma (AC), tubulovillous adenoma (AD), healthy (H) | Downstream classifiers (ResNet18, SVM) |
| ref. bb0400,ref. bb0415,ref. bb0420 | Segmentation | Benign, malignant | Deep contour-aware networks using transfer learning, CNN |
| ref. bb0405,ref. bb1830 | Segmentation | Carcinoma | Random polygon model, Multi-scale CNN with minimal information loss |
| ref. bb0410,ref. bb3860 | Segmentation | Lumen, cytoplasm, nuclei | SVM classifier with RBF kernel, Regions containing glandular structures, Multi-task classifier |
| ref. bb1965,ref. bb3865 | Segmentation | Colorectal Cancer | Two-parallel-branch DNN |
| ref. bb1875 | Segmentation | Colon: adenocarcinoma, high-grade adenoma with dysplasia, low-grade adenoma with dysplasia, carcinoid, and hyperplastic polyp | Ensemble method, wavelet transform (WWE) |
| ref. bb1335 | Segmentation, Classification | Epithelial cell, Connective tissue cell, Lymphocytes, Plasma cells, Neutrophils, Eosinophils | ResNet-34 network with contrastive learning, HoVerNet |
| ref. bb3505 | Segmentation, Classification | high/ low mutation density, microsatellite instability/ stability, chromosomal instability/ genomic stability, CIMP-high/ low, BRAF mutation/wild-type, TP53 mutation/ wild-type, KRAS wild-type/ mutation | ResNet-18 network/ Adapted ResNet34/ HoVerNet, Weakly supervised learning |
| ref. bb1550 | Classification | Colorectal Cancer | customized CNNs, pretrained model VGG, ResNet, Inception, IRV2 |
| ref. bb1950 | Segmentation | Epithelial, inflammatory, fibroblast, miscellaneous, unassigned | Spatially Constrained CNN |
| ref. bb1425 | Segmentation | Tumour | U-Net segmentation with GoogleNet patch level feature extraction, Custom CNN with random forest regression |
| ref. bb1495 | Segmentation | Gastric cancer | CNN models with transfer learning |
| ref. bb1850 | Segmentation | colon/ adenoma, adenocarcinoma, signet, and healthy cases | combination of PHPs, CNN features |
| ref. bb3500 | Classification | tumor, stroma | random forests |
| ref. bb0430 | Prognosis | Stroma | CNN based on neuronal activation in tissues |
| ref. bb0440 | Prognosis | Five year survival rate | CNN/LSTM bassed regression classifier |
| ref. bb3540 | Prognosis | EBV-associated gastric cancer | deep convolutional neural network backboned by ResNet50 |
| ref. bb2300 | Classification | cancerous, high-grade dysplasia, low-grade dysplasia, hyperplastic polyp, normal glands | CNN based classifier with a Multi-Scale Task Multiple Instance Learning (MuSTMIL) |
| ref. bb0390,ref. 762, ref. 763, ref. 764, ref. 765 | Classification | Hyperplastic polyp, sessile serrated polyp, traditional serrated / tubular / tubulovillous / villous adenoma | Radial based SVM classifier, CNN classifier with dropout variance and active learning, SqueezeNet with transfer learning ResNet patch classifier |
| ref. bb3875,ref. bb3880 | Classification | tumor epithelium, simple stroma, complex stroma, immune cell conglomerates, debris and mucus, normal mucosal glands, adipose tissue, background | Bilinear CNN classifier, Convolutional networks (ConvNets) |
| ref. bb3475 | Classification | Normal epithelium, normal stroma, tumor | VGG16, hierarchical neural network |
| ref. bb1155 | Classification | Adenocarcinoma, adenoma, or non-neoplastic | InceptionV3 patch classifier |
| ref. bb1430 | Classification | Epithelial, Spindle-shaped, Necrotic, Inammatory | SC-CNN with Delaunay Triangulation |
| ref. bb3890 | Segmentation | Colorectal cancer, Gastroesophageal junction (dysplasic) lesion, Head and neck carcinoma | DCNN with residual blocks |
| ref. bb3455 | Classification | Adenocarcinoma, corresponding to noticeable CRC, Tubulovillous adenoma, a precursive lesion of CRC, Healthy tissue | Bayesian CNNs (B-CNNs), |
| ref. bb1590 | Classification | cancer, non-cancer | Transfer Learning |
| ref. bb2135 | Disease diagnosis | Gastric cancer | GCN-RNN based feature extraction and encoding |
| ref. bb3895 | Segmentation | Tumor | GAN |
| ref. bb3900 | Prognosis | Adenocarcinoma, disease-specific survival time | ECA histomorphometric-based image classifier |
| ref. bb2165 | Synthesis of large highresolution images | Colorectal Cancer | Novel framework called SAFRON (Stitching Across the FROntier Network) |
| Multi-Organ | |||
| ref. bb1480 | Classification | Benign, malignant | CNN based classifier |
| ref. bb0320 | Classification | Kidney, Lymph nodes, Lung/ Chromophobe, clear cell carcinoma, papillary | Weakly supervised ResNet50 with transfer learning |
| ref. bb1420 | Segmentation | Bladder, Breast, Kidney, Liver, Prostate, Stomach/ Normal, malignant, dysplastic epithelial, fibroblast, muscle, inflammatory, endothelial, miscellaneous | Modified Preact-ResNet50 |
| ref. bb1325 | Segmentation | Nuclear, non-nuclear, boundary | CNN-based classifier with AJI evaluation |
| ref. bb2415 | Detection | Liver, Prostate/ Tumor, normal | Custom CNN architecture |
| ref. bb3445 | Classification | Epithelial, stromal tissues | DCNN classifier |
| ref. bb3905 | Classification | Brain, Breast, Kidney/ DCIS, ERBB2+, triple negative | Transfer learning using multi-scale convolutional sparse coding |
| ref. bb1485 | Detection | Bladder, Breast, Lymph nodes, Lung | CNN-based classifier with transfer learning |
| ref. bb0310 | Detection | Basal, Breast, Prostate/ Cell carcinoma, Metastasis | RNN classifier with multiple instance learning |
| ref. bb0355 | Classification | Micro/Macro metastasis | RNN classifier with MIL |
| ref. bb3490 | Detection | Breast, Stomach/ Tumor, normal | Custom CNN classifier |
| ref. bb1885 | Segmentation | Bladder, Breast, Liver, Prostate, Kidney, Stomach/ Edge, foreground, background | Domain-Adversarial Neural Network |
| ref. bb1695 | Segmentation, Classification | Brain, Lung, Esophagus/ Mitosis, Lymphocyte richness, LUAD, LUSC | Deep residual aggregation network with U-Net segmentation |
| ref. bb1010 | Segmentation | Breast, Esophagus, Liver/ Stain normalization | Relevance vector machine |
| ref. bb3910 | Classification | colon, kidney, ovarian cancer, lung adenocarcinoma, gastric mucosa, astrocytoma, skin cutaneous melanoma, breast cancer/ Nuclei, antigen, cytoplasm, blood, ECM | Non-linear tissue component discrimination |
| ref. bb2295 | Classification | Different organs of rats/Exploring the morphological changes in tissue to biomarker level | CNN, MIL, multi-task learning |
| ref. bb1570 | Classification | Adrenal gland, Bladder, Breast, Liver, Lung, Ovary, Pancreas, Prostate, Testis, Thyroid, Uterus, Heart | CNN models with transfer learning/ ResNet-152 pretrained on ImageNet and GTEx |
| ref. bb1030 | Classification | Colon, Breast/molecular fingerprint of a deficient mismatch (Microsatellite stability(MSS)/ Microsatellite instability (MSI) | CNN models with transfer learning method |
| ref. bb1565 | Classification, Segmentation | blood, breast, lymph, colon, bone, prostate, liver, pancreas, bladder, cervix, esophagus, head, neck, kidney, lung, thyroid, uterus, bone marrow, skin, brain, stomach, and ovary | Unsupervised contrastive learning, residual networks pretrained with self-supervised learning |
| ref. bb1815 | Segmentation | Brain, Breast/ Nuclei | Custom encoder-decoder model |
| ref. bb1390 | Prognosis | Bladder/ Lung, Low TMB, Medium TMB, High TMB | Deep transfer learning, SVM with Gaussian kernel |
| ref. bb3915 | Classification | Bladder, Brain, Breast, Bronchus and lung, Connective, subcutaneous and other soft tissues, Kidney, Liver and intrahepatic bile ducts, Pancreas, Prostate gland, Thyroid gland/ Cancer | ResNet18, self-supervised BYOL method, Clustering tiles using k-means clustering |
| ref. bb1780 | Segmentation, Classification | Colon, Liver, lymph node sections | An ensemble of FCNs architectures/U-Net with DenseNet, ResNet |
| ref. bb3920 | Segmentation cellular nuclei | Multiple | Cross-patch Dense Contrastive Learning |
| ref. bb2845 | Prognosis, Cancer Grade Classification | brain and kidney | CNN, GCN, SNN |
| ref. bb3925 | Segmentation | Colon, Lymph Node | Kullback-Leibler (KL) divergence with classifier |
| ref. bb2430 | Segmentation | Breast, Bone, Tissue | Performing Neural architecture search(NAS) |
| ref. bb3930 | Segmentation of nuclei and cytoplasm | Lung, Bladder | Multi-task model |
| ref. bb3935 | Classification | Axillary lymph nodes, Breast/Metastasis, Colorectal cancer | Adversarial autoencoder, Progressive growing algorithm for GANs, Resnet18 with pre-trained ImageNet weights |
| ref. bb3060 | Classification, Detection | Skin/ cutaneous melanoma(SKCM), Stomach/ adenocarcinoma(STAD), Breast/ cancer(BRCA), Lung/ adenocarcinoma(LUAD), Lung/ squamous cell carcinoma (LUSC) | Convolutional neural networks(CNNs), Birch clustering |
| ref. bb3940 | Classification of tissue | Stomach, Colon, Rectum | CNN+ Pathology Deformable Conditional Random Field |
| ref. bb2395 | Classification of tissue | Liver, Lung, Colon, Rectum | Contrastive learning (CL) with latent augmentation (LA) |
| ref. bb3945 | Classification | Stomach, Intestine, Lymph node, Colon | label correction + NSHE scheme |
| ref. bb3950 | Detection | Renal/ cell carcinoma (RCC), lung/nonsmall cell cancer (NSCLC), Breast/cancer lymph node metastasis | Attention-based learning, Instance-level clustering |
| ref. bb3955 | Classification | Breast, Colon /Tumor metastasis and Tumor cellularity quantification | ResNet-18 |
| ref. bb1930 | Classification, Detection | Breast cancer, Colorectal adenocarcinoma, Colorectal cancer | Co-representation learning (CoReL), Neighborhood-aware multiple similarity sampling strategy |
| ref. bb1740 | segmentations | Nuclei in pancreatic, tubules in colorectal, epithelium in breast | U-net |
| ref. bb2285 | Classification | Brain, Endocrine, Gastro, Gynaeco, Liver, pancreas, Urinary tract, Melanocytic, Pulmonary, Prostate Cancer | DenseNet121, KimiaNet |
| ref. bb3960 | Detection | cell nuclear | Robust Self-Trained Network(RSTN) trained on distance maps(DMs) |
| ref. bb1955 | Segmentation, Classification | Nuclei in the breast, prostate/Benign, ADH, DCIS | GNN models |
| ref. bb1580 | Classification, Segmentation | Lung and Skin/nuclei | ResGANet |
| ref. bb3310 | Prognosis | HPV+, HPV-, survival class | MIL classifier with discriminant analysis |
| ref. bb2275 | Detection | 18 primary organ/Tumor | MIL with attention pooling |
| ref. bb1230 | Detection | Tumor, normal | Sparse coding and transfer learning |
| ref. bb0370 | Detection | Tumor, normal from 23 cohorts | CNN-based classifier with transfer learning |
| ref. bb3585 | Detection | Loose non-tumor tissue, dense non-tumor tissue, normal tumor tissue | Custom CNN classifier |
| ref. bb1915 | Detection | Mitosis centroid | G-CNN for rotational invariance |
| ref. bb1595 | Detection, Segmentation | Colon, Rectum | Concept Contrastive Learning |
| ref. bb1545 | Disease diagnosis | Lung, Breast, Colorectal, Glioma, Renal, Endometrial, Skin, Head and neck, Prostate, Bladder, Thyroid, Ovarian, Liver, Germ cell, Cervix, Adrenal/metastatic tumors and Cancer | MIL |
| ref. bb1400 | Segmentation | Breast, Pancreatic, Colon/Cell Nuclei, Tubules, Epithelium | u-net |
| ref. bb1035 | Segmentation | prostate, colon, breast, kidney, liver, bladder, stomach/Nuclei | U-Net |
| ref. bb2390 | Segmentation | Bladder, Breast, Colorectal, Endometrial, Ovarian, Pancreatic, Prostate/Nuclei | Hovernet on tiles, Nuc2Vec with a ResNet34 with contrastive learning method |
| ref. bb3155 | Segmentation | Brain, Kidney, | semantic segmentation+ Xception |
| ref. bb2830 | Segmentation | Breast, liver, kidney, prostate, bladder, colon, stomach/Cell boundary pixels, Nuclei | Hard-boundary Attention Network (HBANet) with background weaken module (BWM) |
| ref. bb1880 | Segmentation, Classification | Bladder, Breast, Colorectal, Endometrial, Ovarian, Pancreatic, Prostate/Nucleus boundaries/Normal epithelial, malignant/dysplastic epithelial, fibroblast, muscle, inflammatory, endothelial, miscellaneous | CNN pretrained on ImageNe/End-to-end learning |
| ref. bb1765 | Classification | Thyroid frozen sections, Colonoscopy tissue, Cytological cervical pap smear/ benign, non-benign | VGG16bn, ResNet50, U-net, with stochastic selection and attention fusion |
| ref. bb3965 | Classification | Colon, Breast/ non-discriminative and discriminative regions | CNN classifier, ResNet18, Weakly supervised learning, Max-Min uncertainty |
| ref. bb1915 | Classification, Segmentation | Nuclear boundaries, Benign, malignant | G-CNN for rotational invariance |
| ref. bb0860 | Classification | Basaloid, Melanocytic, Squamous | Multi-stage CNN classifier |
| ref. bb3970 | Classification | Colorectal glands, Tumor, normal | Dense steerable filter CNN for rotational invariance |
| ref. bb2255 | Segmentation | Contoured tumor regions | Resnet classifier with transfer learning |
| ref. bb3580 | Classification | Skin melanoma, stomach adenocarcinoma, breast cancer, lung adenocarcinoma, lung squamous cell carcinoma | Custom CNN using human-interpretable image features (HIF) |
| ref. bb3975 | Classification | 20 classes for muscle, epithelial, connective tissue | Inception Residual Recurrent CNN |
| ref. bb0160 | Tissue subtype classification | Adipose (ADI), background (BACK), debris (DEB), lymphocytes (LYM), mucus (MUC), smooth muscle (MUS), normal colon mucosa (NORM), cancer-associated stroma (STR), colorectal adenocarcinoma epithelium (TUM) | ResNet based classifier |
| ref. bb1960 | Nuclei segmentation | Breast, Colon, Liver, Prostate, Kidney, Stomach, Colorectal, Bladder, Ovarian | CNN model, VGG-19 network |
| ref. bb1375 | Segmentation | Lung, Breast | Multiple Instance Learning (MIL), self-supervised contrastive learning in SimCLR setting, feature vector aggregation |
| ref. bb1615 | Prognosis | Prediction of cancer rate survival in the Bladder, Breast, Lung, Uterus, Brain | Graph Convolutional Neural Net(GCN) |
| ref. bb1035 | Nuclei segmentation | prostate, colon, breast, kidney, liver, bladder, stomach | A convolutional U-Net architecture |
| ref. bb1700 | Nuclei segmentation | Nuclear boundaries | U-net based architecture |
| ref. bb2425 | Nuclei segmentation | Nuclear boundaries | Modified HoVer-Net segmentation |
| ref. bb1790 | Nuclei segmentation | Nuclear boundaries | CNN-based attention network |
| ref. bb1535 | Segmentation | 3-level hierarchy of histological types | Pixel level semantic segmentation |
| ref. bb0990 | Segmentation | Tissue, background, edge artifacts, inner artifacts, inner/external margin | Custom FCNN |
| ref. bb1330 | Segmentation | Lymphocytes, necrosis | Semantic segmentation CNN classifier |
| ref. bb0845 | Disease diagnosis | usual ductal hyperplasia, ductal carcinoma in-situ | Deep-learning based CAD tool for pathologists |
| ref. bb2155 | Nuclei segmentation | Positive/negative in nuclei boundaries | Conditional GAN |
| ref. bb1825 | Nuclei segmentation | Nuclear boundaries | CNN-based Boundary-assisted Region Proposal Network |
| ref. bb1820 | Nuclei segmentation | Nuclei, other | CNN-based multi-branch network classifier |
| ref. bb0995 | Detection | Mitosis and metastasis detection | U-Net based normalization |
| ref. bb0840,ref. bb1690 | Nuclei segmentation | normal epithelial, myoepithelial, invasive carcinoma, fibroblasts endothelial, adipocytes, macrophages, inflammatory | U-Net with regression loss |
| ref. bb1710 | Nuclei segmentation | Nuclei body, nuclei boundary, background Background removal Nuclei boundary | UNet based classifier with self-supervised learning UNet with transfer learning Custom encoder-decoder network |
| ref. bb0600 | Tissue subtype classification | 60 types of tissues from a various datasets | ResNet50 feature encoder/decoder for 11 tasks |
| ref. bb3335 | Detection | Tumor, normal | ResNet based patch classifier |
| ref. bb3140 | Classification | Skin/Skin lesions, Chest/ Benign, malignant, Kidney/Chromophobe, clear cell, papillary carcinoma | Conditional Progressive Growing GAN (PG-GAN/ResNet-50) |
| ref. bb1060 | Classification | Neural image compression for Rectal carcinoma | CNN classifier with encoder compression network |
| ref. bb1995 | Pathology report information extraction | Tumor description relating to primary cancer site, laterality, behavior, histological type, and histological grade | Ensemble of multi-task CNN |
| ref. bb0975 | Detection | Blur detection | Combination of CNN and Random Forest regressor |
| ref. bb0980 | Classification | 15 types based on focus level | Lightweight CNN |
| ref. bb3590 | Segmentation | Molecular feature extraction | Multi-layer perceptron with aggregation |
| ref. bb3565 | Classification | Deblurring | Encoder-decoder with VGG-16 blur type classifier |
| ref. bb3510 | Classification | WSI Classification | Multi-scale Context-aware MIL, Multi-level Zooming |
| ref. bb0325 | Classification | BRCA subtyping, NSCLC subtyping, RCC Subtyping | Vision Transformer |
| ref. bb3980 | Classification | Classification of glioma and non-small-cell lung carcinoma cases into subtypes | Two-level model consisting of an Expectation Maximization based method combined with CNN and a decision fusion model |
| ref. bb0325 | Prognosis | Survival prediction of IDC, CCRCC, PRCC, LUAD, CRC, and STAD cancer types | Vision Transformer |
| ref. bb3985 | WSI Processing | Stain normalization | Combination of segmentation and clustering for nuclear/stroma detection |
| ref. bb1050 | WSI Processing | Stain normalization | Self-supervised cycleGAN |
| ref. bb1795 | WSI Processing | Stain normalization | Modified Wasserstein Barycenter approach for multiple referencing |
| ref. bb3405 | WSI Processing | Patch synthesis | Progressive GAN model |
| ref. bb3990 | WSI Processing | Similar image retrieval | Classifier based on ANN with K-means clustering |
| Other | |||
| ref. bb3995 | Detection | Heart/rejection and nonrejection tissue tiles | Progressive Generative Adversarial Network + Inspirational Image Generation with a VGG-19 as a classifier |
| ref. bb3610 | Detection | Heart failure | CNN based patch classifier |
| ref. bb3625 | Classification | Heart/Endomyocardial disease | CACHE-Grader, SVM and K-means clustering |
| ref. bb3615 | Classification | Skin/Cancer | random forest ensemble learning method, feature extractor using ResNeXt50 |
| ref. bb1810 | Detection | Eye/Macular edema | Fully convolutional neural network (FCN), Improved attention U-Net architecture (IAUNet) |
| ref. bb2270 | Prognosis | N/A | Custom MIL framework with attention modules |
| ref. bb2200 | Classification | Bone marrow/Neutrophil, myeloblast, monocyte, lymphocyte | GAN-based classifier |
| ref. bb3530 | Classification, Detection | Bone marrow/nonneoplastic, myeloid leukemia, myeloma | Two-stage detection and classification model |
| ref. bb4000 | Detection | Bone marrow/aspirate pathology synopses | BERT-based NLP mode, Active learning |
| ref. bb1730 | Segmentation | Viable tumor, necrosis with/without bone, normal bone, normal tissue, cartilage, blank | UNet-based multi-magnification network |
| ref. bb4005 | Diseases diagnosis | Bacterial disease | CNN-based classifier |
| ref. bb1805 | Segmentation | Bone marrow/Myelopoietic cells, erythropoietic cells, matured erythrocytes, megakaryocytes, bone, lipocytes | Custom CNN |
| ref. bb3560 | Segmentation | 10 Cancer types | U-Net, Mask R-CNN for quality control |
| ref. bb0835 | Classification | Level 1 (Epithelial, Connective Proper, Blood, Skeletal, Muscular, Adipose, Nervous, Glandular), Level 2 (23 sub-classes from Level 1), Level 3 (36 sub-classes from Level 2 classes) | Ensemble of different CNN architectures |
| ref. bb3575 | Classification | Arteries, nerves, smooth muscle, fat | InceptionV3, Deep ranking network |
| ref. bb3480 | Disease diagnosis | Duodenum/Celiac | CNN-based classifier |
| ref. bb1340 | Disease diagnosis | Cervical cancer, Squamous cell carcinoma, adenocarcinoma | CNN-based patch classifier |
| ref. bb3485 | Disease diagnosis | Duodenum/Celiac, environmental enteropathy, Esophagus/EoE, Ileum/Crohns disease | Hierarchical CNN classifier |
| ref. bb3545 | Disease diagnosis | Squamous carcinoma | Combinations of CNN classifiers |
| ref. bb2290 | Classification | Normal, Cervicitis, Squamous Intra-epithelial Lesion- Low and High, Cancer | Deep multiple instance learning |
| ref. bb1805 | Classification, Segmentation | Bone marrow/Aplasia | SVM classifier with BoW |
| ref. bb3225 | Detection | Tumor cells, stromal cells, lymphocytes, stromal fibroblasts | k-means, Hierarchical Clustering |
| ref. bb1990 | Classification | colorectal liver metastasis | Ensemble of 4 MLP and an encoder, supervised multitask learning (MTL) |
| ref. bb1925 | Detection | Mitosis | Feature pyramid network |
| ref. bb3640 | Detection | Acute Lymphoblastic (or Lymphocytic) Leukemia (ALL), normal/lymphoblast | Transfer Learning, CNN pretrained on a histopathology dataset, ResNet18 and VGG16 as the backbone |
Compilation of information and Neural Network architectures found in different Co-Path papers categorized by task (see 9.11)
| References | Disease/Organ Specification | Architecture | Datasets |
|---|---|---|---|
| Detection Task | |||
| ref. bb0160,ref. bb0310,ref. bb0320,ref. bb0355,ref. bb0360,ref. bb0485,ref. bb0995,ref. bb1060,ref. bb1130,ref. bb1190,ref. bb1355,ref. bb1490,ref. bb1525,ref. bb1545,ref. bb1560,ref. bb1645,ref. bb1865,ref. bb1915,ref. bb2130,ref. bb2170,ref. bb2425,ref. bb2465,ref. bb2985,ref. bb3205,ref. bb3235,ref. bb3330,ref. bb3490,ref. bb3585,ref. bb3695,ref. bb3700,ref. bb3705,ref. bb3755,ref. bb3780,ref. bb3795,ref. bb3800,ref. bb3970 | Breast cancer | Custom CNN {15}, Inception {6}, ResNet {14}, VGG {4}, U-Net {2}, Multi-stage CNN {1}, DenseNet {4}, GAN {1}, AlexNet {1}, E-D CNN {1}, CAS-CNN {1}, Attention CNN {3}, HoVer-Net {1}, MLV-DeepLabV3+ {1}, Xception {1}, Lightweight-CNN {1} | RUMC, CAMELYON16, CAMELYON17, MSK, HUP+CINJ, NHO-1, IDC-Moh, AJ-IDC, PCam, NMCSD, HASHI, TCGA, Cancer Imaging Archive, TCGA-BRCA, Yale HER2 dataset, Yale response dataset |
| ref. bb0310,ref. bb0355,ref. bb0365,ref. bb1370,ref. bb1545,ref. bb2125,ref. bb2990,ref. bb3390,ref. bb3360 | Prostate cancer | Custom CNN {2}, Res Net {4}, Inception {1}, Non-DL {1}, NASNetLarge {1} | RUMC, MSK, HUH, Pro-Raciti, Pro-Doyle, CUH, UHB, Gleason 2019 |
| ref. bb0310,ref. bb1330,ref. bb1545,ref. bb3025,ref. bb3090,ref. bb3615 | Skin cancer | ResNet {3}, Inception {1}, Custom CNN {1}, E-D CNN {1}, ResNeXt {1} | SCMOI, YSM, GHS, MIP, MSK, BE-Cruz-Roa, Private |
| ref. bb1140,ref. bb1155,ref. bb1550,ref. bb1555,ref. bb2195,ref. bb2235,ref. bb2240,ref. bb2260,ref. bb2300,ref. bb3585,ref. bb3895 | Colon cancer | Custom CNN {2}, Inception {1}, GAN {1}, Novel algorithm {1}, DenseNet {1}, ResNet {2}, Inception-ResNet {1}, U-Net {1}, VGG {1}, Swarm Learning {1} | SC-Xu, FAHZU, OSU, TCGA, CRC-Chikontwe, Novel Dataset, DigestPath 2019, Epi700, DACHS, TCGA-CRC, QUASAR trial, YCR-BCIP |
| ref. bb1155,ref. bb1215,ref. bb1380,ref. bb1725,ref. bb3490,ref. bb3495,ref. bb3585, | Stomach cancer | AlexNet {1}, ResNet {3}, Inception {3}, DenseNet {1}, DeepLab {1}, VGG {1}, DLA {1}, Custom CNN {1} | TCGA, SSMH-STAD, SC-Kong |
| ref. bb1390,ref. bb1545 | Bladder cancer | Inception {1}, ResNet {2} | TCGA |
| ref. bb1545,ref. bb3545 | Cervix cancer | Inception {1}, ResNet {2}, Inception-ResNet {1} | XH-FMMU |
| ref. bb3130 | Kidney cancer | Custom CNN {1} | Pantomics |
| ref. bb1545,ref. bb1760,ref. bb2850,ref. bb3315,ref. bb3855 | Lung cancer | Inception {2}, ResNet {1}, DT {1}, Ad-aBoost {1}, XGBoost {1}, U-Net {1} | TCGA(-LUAD,-LUSC), MedicineInsight, 22c3, Ventana PD-L1, Private |
| ref. bb0160 | Oral cancer | Custom CNN {1} | LNM-OSCC |
| ref. bb0995,ref. bb1000,ref. bb1125,ref. bb1345,ref. bb1720,ref. 370, ref. 371, ref. 372, ref. 373, ref. 374, ref. 375, ref. 376, ref. 377, ref. 378 | Mitosis | Custom CNN {7}, AlexNet {1}, U-Net {2}, Multi-stage CNN {2}, FCN {1}, R-CNN {1}, ResNet {2} | TUPAC16, RUMC, MITOS12, TNBC-JRC, AMIDA13, MITOS-ATYPIA14, CWRU |
| ref. bb0385,ref. bb1860,ref. bb1880,ref. bb1935,ref. bb1940,ref. bb1945,ref. bb1950,ref. bb1960,ref. bb2385,ref. bb2390,ref. bb3845,ref. bb3960 | Nuclei | U-Net {2}, GAN {1}, Non-DL {2}, Custom CNN {2}, Hover-Net {2}, SC-CNN {1}, Robust-Self Trained Network (RSTN) {1}, RCNN {1}, VGG {1}, ResNet {2}, E-D CNN {1} | NHS-LTGU, TNBC-CI, MoNuSeg, UHZ, CRCHistoPhenotypes, TCGA, Private, BCFM, PanNuke, NuCLS, CoNSeP, NCT-CRC-HE-100K, Cleveland Clinic (CC) |
| ref. bb0400,ref. bb1965 | Colorectal gland | FCN {2} | GLaS |
| ref. bb0995 | Epithelial cell | Custom CNN {1} | PCa-Bulten, RUMC |
| ref. 387, ref. 388, ref. 389 | Glomeruli | ResNet {1}, VGG {1}, AlexNet {1}, MobileNet {1} | Kid-Wu, Kid-Yang |
| ref. bb3610,ref. bb3625,ref. bb3995 | Heart failure, Heart Transplant | Custom CNN {1}, K-Means {1}, SVM {1}, VGG {1}, PG-GAN {1} | UPenn, CHOA |
| ref. bb1855 | Keratin pearl | Custom CNN {1} | BCRWC |
| ref. bb2805,ref. bb3020 | Liver, Liver fibrous region | Non-DL {1}, Autoencoder CNN {1} | Liv-Atupelage, PAIP |
| ref. bb1205 | Lymphocyte-richness | Autoencoder CNN {1} | TCGA |
| ref. bb1470 | Microsatellite instability | ResNet {1} | TCGA, DACHS, KCCH |
| ref. bb1985,ref. bb3010 | Tumor-infiltrating lymphocyte | U-Net {1}, IM-Net {1}, DRDIN {1} | TCGA, DUKE |
| ref. bb0370,ref. bb1230,ref. bb1545,ref. bb1580,ref. bb2285,ref. bb3060 | Multi-organ tumor | KimiaNet {1}, Novel algorithm {1}, ResNet {3}, Inception {1}, DenseNet {1}, Custom CNN {1}, MLV-DeepLabV3+ {1} | AJ-Lymph, TCGA, ISIC2017, LUNA, COVID19-CT |
| ref. bb0975,ref. bb0980,ref. bb4010 | WSI defect | ResNet {2}, DenseNet {1}, Novel algorithm {1}, Custom CNN {1} | Pro-Campanella, MO-Campanella, MGH, TCGA@Focus, FocusPath |
| ref. bb3640 | Acute Lymphoblastic (or Lymphocytic) Leukemia (ALL) | Custom CNN {1}, ResNet {1}, VGG {1} | ADP, ALL-IDB2 |
| Tissue Subtype Classification Task | |||
| ref. bb0160,ref. bb0390,ref. bb0430,ref. bb0995,ref. bb1430,ref. bb1590,ref. bb3445,ref. bb3460,ref. bb3475,ref. 762, ref. 763, ref. 764, ref. 765 | Colorectal cancer | Non-DL {1}, FCN {1}, ResNet {3}, VGG {3}, AlexNet {1}, Inception {1}, SqueezeNet {2}, BCNN {1}, Capsule CNN {1}, Custom CNN {5}, U-Net {2} | NCT-CRC-HE-100K, NCT-CRC-HE-7K, RUMC, RC-Ciompi, GLaS, CRC-TP, CRC-CDC, UMCM, DHMC-Korbar, TBB, HUH, Stanford Hospital, TCGA |
| ref. bb0135,ref. bb1670,ref. bb2180,ref. bb2225,ref. bb3220,ref. bb3445,ref. bb3905 | Breast cancer | Custom CNN {1}, ResNet {1}, Novel algorithm {2}, Inception {2}, Novel CNN {1} | US-Biomax, ABCTB, TCGA, Bre-Chang, Bre-Steiner, BCSC, NKI-VGH, BACH |
| ref. bb1285,ref. bb2815,ref. bb3905 | Brain cancer | VGG {1}, Novel algorithm {1}, ResNet {1} | UHN, TCGA |
| ref. bb0375,ref. bb1235,ref. bb2705 | Lung cancer | ResNet {2}, AlexNet {1}, Inception {1}, Custom CNN {1} | DHMC, CSMC, MIMW, TCGA, NLST, SPORE, CHCAMS |
| ref. bb1020,ref. bb1480,ref. bb3395 | Prostate cancer | Non-DL {2}, Custom CNN {1}, ResNet {1} | CPCTR, UHZ-PCa, UPenn |
| ref. bb0320,ref. bb1480,ref. bb3905 | Kidney cancer | Novel algorithm {1}, ResNet {2} | TCGA, UHZ-RCC, BWH |
| ref. bb0730 | Bladder cancer | ResNet {1} | CCC-EMN MIBC |
| ref. bb3470 | Stomach cancer | Custom CNN {1} | SPSCI |
| ref. bb0325,ref. bb0835,ref. bb1010,ref. bb1480,ref. bb2275,ref. bb3200,ref. bb3580,ref. bb3915,ref. bb3975 | Multi-organ | Non-DL {1}, Custom CNN {2}, VGG {2}, Inception {3}, ResNet {4}, K-Means {2}, XGBoost {1}, ViT {1} | MO-Khan, KIMIA Path24, ADP, UHZ, TCGA, KIMIA Path960, MO-Diao, BWH-TCGA-MO, CRC-100K, BCSS, BreastPathQ |
| ref. bb0385,ref. bb1350,ref. bb1420,ref. bb1860,ref. bb2385,ref. bb2805 | Nuclei | Custom CNN {3}, ResNet {1}, Non-DL {1}, Hover-Net {2} | CRCHistoPhenotypes, CoNSeP, Liv-Atupelage, PHI, Private, PanNuke, NuCLS |
| ref. bb1155,ref. bb1860,ref. bb3650 | Epithelial | Inception {1}, Custom CNN {1}, Hover-Net+ {1} | HUH-HH, NKI-VGH, TCGA, Private |
| ref. bb1980 | Glomeruli | AlexNet {1} | AIDPATHA, AIDPATHB |
| ref. bb1805,ref. bb2200,ref. bb3530 | Bone marrow | VGG {1}, GAN {1}, FCN {1} | BM-MICCAI15, BM-Hu, FAHZU, RUMC, EUH |
| ref. bb1295,ref. bb2820,ref. bb3085 | Lesion | Non-DL {2}, Inception {1} | UPMC, BE-Hart, Bre-Parvatikar |
| ref. bb3345 | Oral cavity | AlexNet {1} | ECMC |
| Disease Diagnosis Task | |||
| ref. bb0130,ref. bb0315,ref. bb1065,ref. bb1135,ref. 227, ref. 228, ref. 229,ref. bb1315,ref. bb1320,ref. bb1485,ref. bb1490,ref. bb1505,ref. bb1510,ref. bb1525,ref. bb1530,ref. bb1540,ref. bb1640,ref. bb1650,ref. bb1655,ref. bb2105,ref. bb2115,ref. bb2120,ref. bb2145,ref. bb2220,ref. bb2270,ref. bb2475,ref. bb2480,ref. bb2575,ref. bb2980,ref. bb3165,ref. bb3205,ref. bb3235,ref. bb3240,ref. bb3250,ref. bb3255,ref. 722, ref. 723, ref. 724, ref. 725,ref. 730, ref. 731, ref. 732, ref. 733, ref. 734, ref. 735, ref. 736, ref. 737, ref. 738,ref. 740, ref. 741, ref. 742,ref. bb3805 | Breast cancer | ResNet {14}, VGG {7}, Inception {9}, Custom CNN {12}, AlexNet {3}, XGBoost {1}, MobileNet {1}, Xception {1}, DenseNet {7}, Multi-stage CNN {3}, Capsule CNN {1}, SENet {1}, Inception-ResNet {1}, VGGNet {2}, Attention CNN {3}, RCNN {1}, CaffeNet {1}, TriResNet {1}, Class Structured Deep CNN {1}, Non-DL {1} | BACH18, BreakHis, BioImaging, Ext-BioImaging, CAMELYON16, CAMELYON17, CMTHis, AP, AJ-IDC, BIDMC-MGH, PUIH, BRACS, TCGA |
| ref. bb1000,ref. bb1040,ref. bb1945,ref. bb2245,ref. bb3360,ref. 661, ref. 662, ref. 663, ref. 664,ref. bb3400,ref. 751, ref. 752, ref. 753 | Prostate cancer | Custom CNN {3}, U-Net {1}, ResNet {2}, VGG {2}, Inception {1}, AlexNet {2}, Non-DL {1}, MobileNet {1}, DenseNet {1}, DCNN {1}, NASNetLarge {1} | SUH, CSMC, TCGA, NMCSD-MML-TCGA, VPC, UPenn, RCINJ |
| ref. bb0410,ref. bb0710,ref. bb1515,ref. bb3440,ref. bb3455 | Colon cancer | Inception {1}, ResNet {4}, SqueezeNet {1}, AlexNet {2}, MobileNet {1}, Xception {1} | Warwick-CRC, Ext-Warwick-CRC, SC-Holland, GLaS, ZU, ULeeds |
| ref. bb0320,ref. bb1290,ref. bb1695,ref. bb2490,ref. bb3310,ref. bb3840,ref. bb3850 | Lung cancer | Inception {1}, ResNet {3}, Non-DL {2}, DenseNet {1}, GCNN {1} | TCGA, MICCAI17, Stanford-TMA, NYU LMC, BWH, DHMC, ES-NSCLC |
| ref. bb0380,ref. bb0710,ref. bb3655,ref. bb3660 | Brain cancer | Non-DL {2}, Custom CNN {1}, AlexNet {1} | TCGA, MICCAI14 |
| ref. bb2485,ref. bb2805 | Liver cancer | DenseNet {1}, Non-DL {1} | TCGA, SUMC, Liv-Atupelage |
| ref. bb1675,ref. bb2230 | Stomach cancer | Custom CNN {1}, Multi-stage CNN {1} | GNUCH, WSGI |
| ref. bb0860,ref. bb1520,ref. bb3080 | Skin cancer | ResNet {2}, VGG {1}, Multi-stage CNN {1} | DKI, Y/CSUXH-TCGA, DLCS, BE-TF-Florida-MC |
| ref. bb2110 | Bladder cancer | Custom CNN {1}, Inception {1}, Multi-stage CNN {1} | TCGA+UFHSH |
| ref. bb1340 | Cervix cancer | VGG {1} | TCGA |
| ref. bb1475 | Esophagus cancer | ResNet {1}, Attention CNN {1} | DHMC |
| ref. bb2270 | Kidney cancer | Custom CNN {1} | TCGA |
| ref. bb1485,ref. bb1765 | Multi-organ cancer | Inception {1}, ResNet {2}, Custom CNN {1}, U-Net {1}, VGG {1} | Stanford-TMA, BIDMC-MGH, Private |
| ref. bb3310 | Oral cancer | Custom CNN {1} | OP-SCC-Vanderbilt |
| ref. bb0595 | Ovarian cancer | VGG {1}, Multi-stage CNN {1} | VGH |
| ref. bb1500,ref. bb3480,ref. bb3485 | Non-cancer GI tract disorder | ResNet {2}, VGG {1} | DHMC-Wei, UV, SC-Sali |
| ref. bb2825,ref. bb3330 | Lymphoma | E-D CNN {1}, Custom CNN {1} | TUCI-DUH, AJ-Lymph |
| Segmentation Task | |||
| ref. bb0840,ref. bb1035,ref. bb1070,ref. bb1205,ref. bb1325,ref. bb1420,ref. 330, ref. 331, ref. 332, ref. 333, ref. 334, ref. 335,ref. 350, ref. 351, ref. 352,ref. 356, ref. 357, ref. 358,ref. bb1880,ref. bb1885,ref. bb1915,ref. bb2155,ref. bb2200,ref. bb2385,ref. bb2390,ref. bb2425,ref. bb2805,ref. bb2830,ref. bb3095,ref. bb3210,ref. bb3560,ref. bb3845,ref. bb3970 | Nuclei | U-Net {6}, Custom CNN {8}, FCN {1}, ResNet {5}, GAN {4}, Non-DL {2}, Multistage CNN {1}, E-D CNN {4}, Autoencoder CNN {1}, PangNet {1}, DeconvNet {1}, Hover-Net {3}, multi-branch CNN {1}, Attention(EP, SM) CNN {1}, HBANet {1} | MICCAI15-18, TCGA, TNBC-CI, MoNuSeg, CPM-15, CPM-17, CCB, CRCHistoPhenotypes, CoNSeP, BM-Hu, FAHZU, Liv-Atupelage, DHMC, MO-Khoshdeli, AJ-N, Kumar-TCGA, TCGA-Nuclei, SOX10, UrCyt, NLST, Pan-Bai, PanNuke, NuCLS, Cleveland Clinic (CC) |
| ref. 80, ref. 81, ref. 82, ref. 83, ref. 84,ref. bb0840,ref. bb1830,ref. bb1965,ref. bb3860,ref. bb3865,ref. bb3970 | Gland | FCN {2}, Non-DL {1}, Custom CNN {5}, ResNet {2}, VGG {1}, multi-branch CNN {1}, E-D CNN {1} | GLaS, Bilkent, CRAG, Priv-IHC |
| ref. bb1425,ref. bb1560,ref. bb1650,ref. bb1800,ref. 360, ref. 361, ref. 362,ref. bb1865,ref. bb1870,ref. bb2835,ref. bb3215,ref. bb3690 | Breast tumor | Custom CNN {2}, Inception {2}, U-Net {3}, FCN 1, E-D CNN {1}, RAN {1}, DA-RefineNet {1}, DeepLab {1}, MLV-DeepLabV3 {1} | CAMELYON16, CAMELYON17, BACH18, TCGA, UHCMC-CWRU, BC-Priego-Torres, TUPAC16, AMGrad, TCGA-BRCA, Yale HER2 dataset, Yale response dataset |
| ref. bb0710,ref. bb1555,ref. bb1850,ref. bb1875 | Colon tumor | Custom CNN {1}, VGG {1}, U-Net {2}, Non-DL {1}, AlexNet {1} | Warwick-UHCW, Warwick-Osaka, ZU, DigestPath 2019, Yeouido |
| ref. bb1715,ref. bb2415,ref. bb3020 | Liver tumor | PlexusNet {1}, U-Net {1}, Autoencoder CNN {1} | TCGA, IHC-Seg, PAIP |
| ref. bb1760 | Lung tumor | U-Net {1} | 22c3, Ventana PD-L1 |
| ref. bb1770,ref. bb2415,ref. bb2840 | Prostate tumor | PlexusNet {1}, Non-DL {1}, U-Net {1} | SMS-TCGA, UUH, Private |
| ref. bb1425,ref. bb1495 | Stomach tumor | Inception {1}, U-Net {1}, ResNet {1} | SC-Takahama, SC-Liu |
| ref. bb0710 | Brain tumor | AlexNet {1} | MICCAI14 |
| ref. bb1730 | Bone tumor | U-Net {1} | MSKCC |
| ref. bb1385,ref. bb1580 | Skin tumor | FCN {1}, ResNet {1}, ResGANet {1} | TCGA, ISIC2018 |
| ref. bb2255 | Multi-organ tumor | ResNet {1} | TCGA |
| ref. bb0420,ref. bb0760,ref. bb1755,ref. bb3125 | Kidney tissue structure | Custom CNN {1}, U-Net {1}, cascaded CNN {1} | WUPAX, M-Gadermayr, RUMC, Mayo, AIDPATH |
| ref. bb1805 | Bone marrow cell | FCN {1} | RUMC |
| ref. bb1735 | Breast tissue subtype | U-Net {1} | NHS |
| ref. bb1535 | Histological tissue type | Custom CNN {1} | ADP |
| ref. bb3790 | Liver steatosis | ResNet {1} | Liv-Guo |
| ref. bb1720,ref. bb1725 | Mitosis | U-Net {1}, FCN {1} | MITOS12, MITOS-ATYPIA14, AMIDA13 |
| ref. bb1855,ref. bb1860 | Oral mucosa, Oral Epithelial Dysplasia | Custom CNN {1}, Hover-Net+ {1} | BCRWC,Private |
| ref. bb1025,ref. bb1330,ref. bb3010 | Lymphocytes(Tumor-infiltrating,segmentation) | E-D CNN {1}, IM-Net {1}, DRDIN {1}, U-Net {1}, SegNet {1} | TCGA, DUKE, Lymphocyte Detection(from Andrew Janowczyk and Anant Madabhushi) |
| ref. bb0990,ref. bb1810 | Tissue region, Fluid Lesions | Custom CNN {1}, FCN {1}, IAUNet {1} | Bándi-Dev-Set, Bándi-Dis-Set, RETOUCH |
| WSI Processing Task | |||
| ref. bb1000,ref. bb1045,ref. bb1935,ref. bb2185,ref. bb2190,ref. bb3095,ref. bb3375 | Domain adaptation | GAN {5}, U-Net {1}, AlexNet {1}, Custom CNN {1}, ResNet {5} | NHS-LTGU, MITOS-ATYPIA14, RCINJ, Roche, Liv-Lahiani, TU-PAC16, TCGA, DHMC, SOX10, UrCyt |
| ref. bb1010,ref. bb1015,ref. bb1040,ref. bb1050,ref. bb1625,ref. bb1795,ref. bb3290,ref. bb3985 | Stain normalization | Custom CNN {2}, ResNet {1}, VGG {1}, U-NET {1}, GAN {3}, E-D CNN {1}, Multistage CNN {1}, Non-DL {4} | SUH, Leica Biosystems, MO-Khan, MGH, Lym-Bejnordi, Salvi-SCAN, MITOS-ATYPIA14 |
| ref. bb1685,ref. 424, ref. 425, ref. 426,ref. bb2480,ref. bb3140,ref. bb3405 | Patch synthesis | Custom CNN {2}, GAN {5}, PG-GAN {2}, ResNet {1}, VGG {1}, U-Net {2} | MICCAI16/17/18, Kumar-TCGA, BreakHis, NKI-VGH, TCGA, OVCARE, ISIC 2020, ChestXray-NIHCC, CRAG, Digestpath |
| ref. bb0995,ref. bb1795,ref. bb2170,ref. bb3785 | Processing technique comparison | U-Net {1}, Custom CNN {2}, GAN {2} | RUMC, CAMELYON16, MITOS-ATYPIA14 |
| ref. bb1060,ref. bb1990 | WSI compression | Custom CNN {2}, E-D CNN {1}, GAN {1} | N/A |
| ref. bb3455 | Data cleaning | ResNet {1} | ULeeds |
| ref. bb3910 | Stain augmentation | VGG {1} | Kid-Cicalese |
| ref. bb3910 | Tissue component discrimination | Non-DL {1} | TCGA, MO-JHU/US/UB |
| ref. bb3750 | WSI transformations | Custom CNN {1} | BreakHis |
| ref. bb3510 | WSI Classification | MIL {1} | IMP Diagnostics Lab., BRIGHT, CAMELYON16 |
| Patient Prognosis Task | |||
| ref. bb0425,ref. bb1225,ref. bb2420,ref. bb2815,ref. bb3100 | Brain cancer | Inception {1}, VGG {1}, Custom CNN {2}, Capsule CNN {1}, ResNet {1} | TCGA |
| ref. bb1070,ref. bb1290,ref. bb2265,ref. bb2705,ref. bb2855,ref. bb3100,ref. bb3310 | Lung cancer | Custom CNN {2}, Non-DL {2}, AttentionMIL {1}, MI-FCN {1}, HANet {1} | Stanford-TMA, TCGA(-LUSC), CHCAMS, NLST, ES-NSCLC |
| ref. bb0430,ref. bb0440,ref. bb2265,ref. bb3900 | Colon cancer | VGG {2}, Inception {1}, ResNet {1}, Non-DL {1}, AlexNet {1}, SqueezeNet {1}, AttentionMIL {1}, MI-FCN {1} | NCT-CRC-HE-100K, NCT-CRC-HE-7K, HUCH, WRH-WCH, MCO |
| ref. bb1940,ref. bb2270,ref. bb3835 | Kidney cancer | Non-DL {2}, AttentionMIL {1} | UHZ, TCGA, HPA |
| ref. bb1985,ref. bb2270 | Breast cancer | U-Net {1}, AttentionMIL {1} | AJ-Lymph, TCGA |
| ref. bb3400 | Prostate cancer | Inception {1} | NMCSD+MML+TCGA |
| ref. bb1330 | Melanoma | Multi-stage CNN {1} | MIP, YSM, GHS |
| ref. bb0435 | Mesothelioma | ResNet {1} | MESOPATH, TCGA |
| ref. bb0325,ref. bb1615 | Multi-Organ | AttentionMIL {1}, GCN {1}, ViT {1} | TCGA, CRC-100K, BCSS, BreastPathQ |
| ref. bb0445,ref. bb3665 | Recurrence prediction | Custom CNN {1}, Non-DL {1} | NSCLC-Wang, ROOHNS |
| Other Tasks | |||
| ref. bb0135,ref. bb1520,ref. bb1750,ref. bb2125,ref. bb2485,ref. bb3410 | Clinical validation, Stress Test, Quality Control, Explainability | DenseNet {1}, ResNet {2}, Inception {1}, Inception-ResNet {1}, Extended U-Net {1} | SUMC, Bre-Steiner, Pro-Raciti, DKI, TCGA, Private |
| ref. bb0835,ref. bb1030,ref. bb1065,ref. bb1335,ref. bb1365,ref. bb1400,ref. bb1530,ref. bb1570,ref. bb1775,ref. bb1780,ref. bb1955,ref. bb2175,ref. bb3260,ref. bb3560 | Dataset creation/curation and annotation, IntegratedAPI and (End-to-End) Toolkits | Custom CNN {1}, GNN {1}, VGG {3}, Inception {3}, AlexNet {1}, FCN {2}, U-Net {4}, ResNet {6}, Hover-Net {1}, DenseNet {1}, MobileNet {1}, DeepLab {1}, SLAM {1} | CMTHis, ADP, TCGA, TCGA-Nuclei, PUIH, BRACS, BACH, UZH, SICAPv2, Lizard, GTEx Dataset(V8), BreaKHis, HF(Heart failure) Dataset, DACHS, YCR-BCIP, Diagset-A, Diagset-B, Diagset-C, Painter by Numbers, miniImageNet, CRC(DX), CAMELYON(16,17), DigestPath, PAIP, Private |
| ref. bb3335,ref. bb3855,ref. bb3890 | Data deficiency study | Inception {1}, Custom CNN {1}, ResNet {1} | TCGA, GLaS, CAMELYON16, CAMELYON17, Thagaard |
| ref. bb1875,ref. bb2135,ref. bb2280,ref. bb2295,ref. bb3575,ref. bb3810,ref. bb3825,ref. bb3915 | Image retrieval/compression, Representation Learning | ResNet {1}, Inception {1}, Non-DL {2}, GCN {1}, AttnMIL {1}, Custom CNN {2}, U-Net++ {1}, Barcodes {1}, XGBoost {1}, K-Means {1} | TCGA, CAMELYON16, SC-Zheng, CRA, Han-Wistar Rats |
| ref. bb0600,ref. bb1375,ref. bb1380,ref. bb1565,ref. bb1575,ref. bb1615,ref. bb1990,ref. bb2175,ref. 451, ref. 452, ref. 453, ref. 454, ref. 455, ref. 456,ref. bb2375,ref. bb2385,ref. bb2390,ref. bb3015,ref. bb3050,ref. bb3505,ref. bb3950,ref. bb3955,ref. bb3965,ref. bb4015 | Multi-(task,instance) learning(MT,MIL), (Weak,Semi,Self)-Supervised Learning, Contrastive Learning | ResNet {9}, GCN {1}, AttentionMIL {2}, MuSTMIL {1}, SimCLR {3}, MIL {2}, D(S)MIL {1}, Pretext-RSP {1}, MoCo {1}, MLP {1}, CLAM {1}, GAN {1}, VGG {2}, DenseNet {2}, Hover-Net {2}, Custom CNN {2}, SLAM {1}, DLA {1}, TransMIL {1} | UHC-WNHST, PanNuke, AJ-Epi-Seg, OSCC, TCGA(-CRC-DX, – THCA, -NSCLC, -RCC), CAMELYON16, CAMELYON17, NCT-CRC-HE-100K, NCT-CRC-HE-7K, CPM-17, AJ-Lymph, M-Qureshi, SKMCH&RC, SKMCH&RC-M, OV-Kobel, MT-Tellez, TUPAC16, SC-Galjart, CT-CRC-HE-100K, Munich AML, MSK, MHIST, CPTAC, Kather multi-class, BreastPathQ, CRC, Novel, PanNuke, NuCLS, GlaS, OAUTHC, DACHS, YCR-BCIP, BreakHis, DEC, TH-TMA17 |
| ref. bb1060,ref. bb1345,ref. bb1745 | Proliferation scoring | Custom CNN {2}, piNET {1} | TUPAC16, DeepSlides |
| ref. bb2200 | Cell clustering | ResNet {1} | MICCAI15, BM-Hu, FAHZU |
| ref. bb2200 | Chemosensitivity prediction | Non-DL {1} | TCGA |
| ref. bb2430 | Neural Architecture Search | DARTS {1} | ADP,BCCS,BACH, Osteosarcoma |
| ref. bb1285,ref. bb2380,ref. bb2390,ref. bb3020,ref. bb3060,ref. bb3810 | Feature extraction/analysis, | VGG {1}, Custom CNN {2}, Hover-Net {1}, | CAMELYON16, UHN, TCGA, Private |
| Unsupervised Learning | PixelCNN {1}, AutoEncoder CNN {1} | ||
| ref. bb2305,ref. bb2310,ref. bb2490,ref. bb3505 | Gene mutation prediction | VGG {1}, Inception {1}, AttentionMIL {2}, DenseNet {1}, ResNet {1}, Hover-Net {1} | TCGA(-CRC-DX, -THCA), DEC, PAIP, TH-TMA17, Private |
| ref. bb1555,ref. bb1930,ref. bb1960,ref. bb1970,ref. bb3735 | Novel loss function, Novel optimizer | ResNet {3}, VGG {3}, MobileNet {1}, DenseNet {1}, U-Net {1} | BACH18, AJ-Lymphocyte, CRCHistoPhenotypes, CoNSeP, ICPR12, AMIDA13, Kather Multi-class, DigestPath 2019, CT-CRC-HE-100K |
| ref. bb3050,ref. bb3445 | Patch triaging | Non-DL {1}, ResNet {1}, Pretext-RSP {1}, MoCo {1}, MLP {1} | BIRL-SRI, CAMEYLON16, MSK, MHIST |
| ref. bb1995,ref. bb4000 | Pathology report information extraction | BERT {1}, Custom CNN {1} | LTR |
| ref. bb1660 | Receptor status prediction | Custom CNN {1} | TCGA |
| ref. bb1390 | TMB prediction | Inception {1} | TCGA |
| ref. bb1345,ref. bb3265 | Tumor grading | Mask R-CNN {1}, Custom CNN {1}, Non-DL {1} | TUPAC16, Post-NAT-BRCA, ILC |
| ref. bb3225 | Visual analytic tool | Non-DL {1} | TCGA |
| Keywords | comma separated list |
| Organ Application | Organ: |
| Task: | |
| Dataset Compilation | Name: |
| Availability: | |
| Dataset Size: (#patches/#slides/#images) | |
| Image Resolution: | |
| Staining Type: | |
| Annotation Type: (region/patch/slide-level) | |
| Histological Type: (cellular/tissue ROI/etc, I.e., on what basis is it labeled) | |
| Label Structure: (single label/multi label) | |
| Class Balance: (is size of dataset balanced across each classes) | |
| Technicality | Model: (architecture/transfer learning/output format) |
| Training Algorithm: (end-to-end/separately staged) | |
| Code Availability: (give source) | |
| Data Processing | Image Pre-processing: (patching, data augmentation, color normalization) |
| Output Processing: | |
| Performance Summary | Evaluation Metrics: |
| Notable Results: Numerical result for strongest performing model. | |
| Comparison to Other Works: Comparison to state-of-the-art models (one sentence) | |
| Novelty | Medical Applications/Perspectives: |
| Technical Innovation: (algorithms for processing or deep learning, new metrics) | |
| Explainability | Visual Representations: (feature distribution, heatmaps, tsne, gradCAM, pseudocode, etc.) |
| Clinical Validation | Usage in Clinical Settings: Has the work been used by pathologists in clinical setting? |
| Suggested Usage: How can the work be used by pathologists? | |
| Performance Comparison: Has the model performance been compared to that of pathologists? | |
| Caveats and Recommendations | • Personal comments on the paper |
| • Relevant info from other papers | |
| • Criticism and limitations of the work |
| Keywords | Deep learning, convolutional neural networks, lung adenocarcinoma, multi class, ResNet |
| Organ Application | Organ: Lung |
| Task: Histologic pattern classification in lung adenocarcinoma | |
| Dataset Compilation | Name: Dartmouth-Hitchcock Medical Centre in Lebanon, New Hampshire |
| Availability: Unavailable due to patient privacy constraints. Anonymized version available upon request. | |
| Dataset Size: 422 WSIs, 4 161 training ROIs, 1 068 validation patches | |
| Image Resolution: 20 magnification | |
| Staining Type: H&E | |
| Annotation Type: Region-level training set, patch-level validation set, slide-level test set | |
| Histological Type: Lepidic, acinar, papillary, micropapillary, solid, and benign. | |
| Label Structure: Single label | |
| Class Balance: Imbalanced, with significantly fewer papillary patterns in all data. | |
| Technicality | Model: ResNet model with 18 layers |
| Training Algorithm: | |
| • Multi-class cross-entropy loss | |
| • Initial learning rate of 0.001 | |
| • Learning rate decay by factor of 0.9 per epoch | |
| Code Availability:https://github.com/BMIRDS/deepslide | |
| Data Processing | Image Pre-processing: |
| • Created training ROIs by selectively cropping regions of 245 WSIs. | |
| • Spliced 34 validation WSIs into 1 068 224×224 patches. | |
| • Colour channel normalization to mean and standard deviation of entire training set. | |
| • Data augmentation by rotation; flipping; and random colour jittering on brightness, contrast, hue, and saturation. | |
| Output Processing: Low-confidence predictions filtered out for predictions below a threshold. Thresholds are determined by a grid search over classes, optimizing for similarity between the trained model and the validation data. | |
| Performance Summary | Evaluation Metrics: F1-Score, AUC |
| Notable Results: F1-Score of 0.904 on validation set, AUC greater than 0.97 for all classes. | |
| Comparison to Other Works: ResNet18, 34, 50, 101, 152 compared for performance to choose optimal depth. All had similar accuracies on validation set, so chose ResNet18 for lower model complexity. | |
| Novelty | Medical Applications/Perspectives: Potential platform for quality assurance of diagnosis and slide analysis. |
| Technical Innovation: First paper to attempt to classify based on histological lung adenocarcinoma subtypes. | |
| Explainability | Visual Representations: Heatmaps for patterns detected, AUC curve for each class |
| Clinical Validation | Usage in Clinical Settings: N/A |
| Suggested Usage: | |
| • Could be integrated into existing lab information management systems to provide second opinions to diagnoses. | |
| • Visualization of a slide could highlight important tissue structures. | |
| • Could help facilitate tumour diagnosis process by automatically requesting genetic testing based on histological data for patient. | |
| Performance Comparison: | |
| • On par with pathologists for all evaluated metrics | |
| • Model in agreement 66.6% of the time with pathologists on average, with robust agreement (agreement with 2/3 of the pathologists) 76.7% of the time. | |
| • WSI region annotation differences between pathologist and model are compared for a sample slide. | |
| Caveats and Recommendations | • Data taken from one medical centre, so may not be representative of lung adenocarcinoma morphology• Dataset relatively small compared to other deep learning datasets, with some classes having very few instances |
| Keywords | Non-small cell lung cancer, histology image classification, computational pathology, deep learning |
| Organ Application | Organ: Lung |
| Task: Classification between non-small cell lung cancer types | |
| Dataset Compilation | Name: Computational Precision Medicine at MICCAI 2017 |
| Availability: Unavailable, link on MICCAI 2017 website unreachable | |
| Dataset Size: 64 WSIs | |
| Image Resolution: 20 magnification | |
| Staining Type: H&E | |
| Annotation Type: Pixel-level and Slide-level | |
| Histological Type: | |
| • At pixel-level, classifies as lung adenocarcinoma (LUAD), lung squamous cell carcinoma (LUSC), and non-diagnostic (ND) | |
| • At slide-level, LUAD or LUSC | |
| Label Structure: Single label | |
| Class Balance: Balanced dataset at the slide level, 32 LUAD and 32 LUSC | |
| Technicality | Model: |
| • Ensemble ML model | |
| • Variant of ResNet50, called ResNet32 with 32 layers and 3×3 kernel, as compared to 7×7 kernel with ResNet50. | |
| • 50 statistical and morphological features extracted from probability maps generated by ResNet32. The top 25 are selected for best class separability and used as input to a random forest. | |
| Training Algorithm: Separately staged, ResNet32 creates probability maps, then random forest generates final prediction for each WSI | |
| Code Availability: Unavailable | |
| Data Processing | Image Pre-processing: |
| • Splicing of slides into 256256 patches, then random cropping into 224224 patches | |
| • Reinhard stain normalization | |
| • Random crop, flip, rotation data augmentation | |
| Output Processing: N/A | |
| Performance Summary | Evaluation Metrics: Accuracy |
| Notable Results: | |
| • ResNet32 with Random Forest achieves 0.81 accuracy over WSI | |
| • Results superior to ResNet32 with Maximum Vote, which had 0.78 accuracy. Features for the random forest are tailored for WSI classification, and so can achieve higher performance. | |
| Comparison to Other Works: Compared ResNet32 to VGG, GoogLeNet, and ResNet50, with higher average classification accuracy. | |
| Novelty | Medical Applications/Perspectives: Automated distinguishing of LUAD tissue from LUSC could be done at scale to assist pathologists in diagnosis and treatment planning for patients. |
| Technical Innovation: | |
| • First 3-class network for classification of WSI into diagnostic/nondiagnostic areas | |
| • Ensemble method resulted in greatest accuracy at the MICCAI 2017 competition. | |
| Explainability | Visual Representations: Probability maps for each pixel-level class |
| Clinical Validation | Usage in Clinical Settings: N/A |
| Suggested Usage: Automated distinguishing of LUAD and LUSC slides could aid pathologists in treatment planning. | |
| Performance Comparison: N/A | |
| Caveats and Recommendations | • Because features for random forest training are chosen based on categorization of lung tissue samples, may not be able to generalize well to other tissue types. |
References
- FDA News Release. Fda allows marketing of first whole slide imaging system for digital pathology. 2017
- Andrew J. Evans, Thomas W. Bauer, Marilyn M. Bui. Us food and drug administration approval of whole slide imaging for primary diagnosis: a key milestone is reached and new questions are raised. Arch Pathol Lab Med, 2018. [PubMed]
- Anna Luíza Damaceno Araújo, Lady Paola Aristizábal Arboleda, Natalia Rangel Palmier. The performance of digital microscopy for primary diagnosis in human pathology: a systematic review. Virchows Arch, 2019. [PubMed]
- Bethany Jill Williams, Andrew Hanby, Rebecca Millican-Slater, Anju Nijhawan, Eldo Verghese, Darren Treanor. Digital pathology for the primary diagnosis of breast histopathological specimens: an innovative validation and concordance study on digital pathology validation and training. Histopathology, 2018. [PubMed]
- Frederik Großerueschkamp, Hendrik Jütte, Klaus Gerwert, Andrea Tannapfel. Advances in digital pathology: from artificial intelligence to label-free imaging. Visceral Med, 2021
- Kuo-Hsing Kuo, Joyce M. Leo. Optical versus virtual microscope for medical education: a systematic review. Anat Sci Educ, 2019. [PubMed]
- Robert Pell, Karin Oien, Max Robinson. UK National Cancer Research Institute (NCRI) Cellular-Molecular Pathology (CM-Path) quality assurance working group, Owen J Driskell, et al. The use of digital pathology and image analysis in clinical trials. J Pathol Clin Res, 2019. [PubMed]
- Shaimaa Al-Janabi, André Huisman, Paul J. Van Diest. Digital pathology: current status and future perspectives. Histopathology, 2012
- Jon Griffin, Darren Treanor. Digital pathology in clinical use: where are we now and what is holding us back?. Histopathology, 2017. [PubMed]
- Adela Saco, Jose Antoni Bombi, Adriana Garcia, Jose Ramírez, Jaume Ordi. Current status of whole-slide imaging in education. Pathobiology, 2016. [PubMed]
- Rajiv Kumar Kaushal, Sathyanarayanan Rajaganesan, Vidya Rao, Akash Sali, Balaji More, Sangeeta B. Desai. Validation of a portable whole-slide imaging system for frozen section diagnosis. J Pathol Inform, 2021
- Jeroen Van der Laak, Geert Litjens, Francesco Ciompi. Deep learning in histopathology: the path to the clinic. Nat Med, 2021. [PubMed]
- Miao Cui, David Y. Zhang. Artificial intelligence and computational pathology. Lab Invest, 2021. [PubMed]
- Amelie Echle, Niklas Timon Rindtorff, Titus Josef Brinker, Tom Luedde, Alexander Thomas Pearson, Jakob Nikolas Kather. Deep learning in cancer pathology: a new generation of clinical biomarkers. Br J Cancer, 2021. [PubMed]
- Balázs Acs, Mattias Rantalainen, Johan Hartman. Artificial intelligence as the next step towards precision pathology. J Intern Med, 2020. [PubMed]
- U. Massimo Salvi, Rajendra Acharya, Filippo Molinari, Kristen M. Meiburger. The impact of pre- and post-image processing techniques on deep learning frameworks: A comprehensive review for digital pathology image analysis. Comput Biol Med, 2021
- Chetan L. Srinidhi, Ozan Ciga, Anne L. Martel. Deep neural network models for computational histopathology: A survey. Med Image Anal, 2021
- Giovani Lujan, Zaibo Li, Anil V. Parwani. Challenges in implementing a digital pathology workflow in surgical pathology. Human Pathol Rep, 2022
- Yingci Liu, Liron Pantanowitz. Digital pathology: Review of current opportunities and challenges for oral pathologists. J Oral Pathol Med, 2019. [PubMed]
- Ana Richelia Jara-Lazaro, Thomas Paulraj Thamboo, Ming Teh, Puay Hoon Tan. Digital pathology: exploring its applications in diagnostic surgical pathology practice. Pathology, 2010. [PubMed]
- Julie Smith, Sys Johnsen, Mette Christa Zeuthen. On the road to digital pathology in denmark—national survey and interviews. J Digit Imaging, 2022
- Sandhya Sundar, Pratibha Ramani, Herald J. Sherlin, Gheena Ranjith, Abilasha Ramasubramani, Gifrina Jayaraj. Awareness about whole slide imaging and digital pathology among pathologists-cross sectional survey. Indian. J For Med Toxicol, 2020
- Ali Jasem Buabbas, Tareq Mohammad, Adel K. Ayed, Hawraa Mallah, Hamza Al-Shawaf, Abdulwahed Mohammed Khalfan. Evaluating the success of the tele-pathology system in governmental hospitals in kuwait: an explanatory sequential mixed methods design. BMC Med Inform Decis Mak, 2021. [PubMed]
- Alexi Baidoshvili, Anca Bucur, Jasper van Leeuwen, Jeroen van der Laak, Philip Kluin, Paul J. van Diest. Evaluating the benefits of digital pathology implementation: time savings in laboratory logistics. Histopathology, 2018. [PubMed]
- T. Dennis, R.D. Start, Simon S. Cross. The use of digital imaging, video conferencing, and telepathology in histopathology: a national survey. J Clin Pathol, 2005. [PubMed]
- Fei Dong, Humayun Irshad, Oh. Eun-Yeong. Computational pathology to discriminate benign from malignant intraductal proliferations of the breast. PloS One, 2014
- David F. Steiner, Robert MacDonald, Yun Liu. Impact of deep learning assistance on the histopathologic review of lymph nodes for metastatic breast cancer. Am J Surg Pathol, 2018. [PubMed]
- A.D. Lee, Alexis B. Carter, Alton B. Farris. Proceedings of the IEEE, 2012. [PubMed]
- Inho Kim, Kyungmin Kang, Youngjae Song, Tae-Jung Kim. Application of artificial intelligence in pathology: Trends and challenges. Diagnostics, 2022. [PubMed]
- Esther Abels, Liron Pantanowitz, Famke Aeffner. Computational pathology definitions, best practices, and recommendations for regulatory guidance: a white paper from the digital pathology association. J Pathol, 2019. [PubMed]
- Neeta Kumar, Ruchika Gupta, Sanjay Gupta. Whole slide imaging (wsi) in pathology: current perspectives and future directions. J Digit Imaging, 2020. [PubMed]
- Navid Alemi Koohbanani, Balagopal Unnikrishnan, Syed Ali Khurram, Pavitra Krishnaswamy, Nasir Rajpoot. Self-path: Self-supervision for classification of pathology images with limited annotations. IEEE Trans Med Imaging, 2021. [PubMed]
- Kaustav Bera, Kurt A. Schalper, David L. Rimm, Vamsidhar Velcheti, Anant Madabhushi. Artificial intelligence in digital pathology—new tools for diagnosis and precision oncology. Nat Rev Clin Oncol, 2019. [PubMed]
- Muhammad Khalid Khan Niazi, Anil V. Parwani, Metin N. Gurcan. Digital pathology and artificial intelligence. Lancet Oncol, 2019. [PubMed]
- Faranak Sobhani, Ruth Robinson, Azam Hamidinekoo, Ioannis Roxanis, Navita Somaiah, Yinyin Yuan. Artificial intelligence and digital pathology: Opportunities and implications for immuno-oncology. Biochim Biophys Acta (BBA)-Rev Cancer, 2021
- Christophe Klein, Qinghe Zeng, Floriane Arbaretaz. Artificial intelligence for solid tumour diagnosis in digital pathology. Br J Pharmacol, 2021. [PubMed]
- Heounjeong Go. Digital pathology and artificial intelligence applications in pathology. Brain Tumor Res Treat, 2022. [PubMed]
- Jérôme Rony, Soufiane Belharbi, Jose Dolz, Ismail Ben Ayed, Luke McCaffrey, Eric Granger. Deep weakly-supervised learning methods for classification and localization in histology images: a survey. arXiv preprint arXiv, 2019
- K. Abinaya, B. Sivakumar. 2022 6th International Conference on Trends in Electronics and Informatics (ICOEI), 2022
- Mohsin Bilal, Mohammed Nimir, David Snead, Graham S. Taylor, Nasir Rajpoot. Role of ai and digital pathology for colorectal immuno-oncology. Br J Cancer, 2022
- Lucas Schneider, Sara Laiouar-Pedari, Sara Kuntz. Integration of deep learning-based image analysis and genomic data in cancer pathology: A systematic review. Eur J Cancer, 2022. [PubMed]
- Yahui Jiang, Meng Yang, Shuhao Wang, Xiangchun Li, Yan Sun. Emerging role of deep learning-based artificial intelligence in tumor pathology. Cancer Commun, 2020
- Cesare Lancellotti, Pierandrea Cancian, Victor Savevski. Artificial intelligence & tissue biomarkers: advantages, risks and perspectives for pathology. Cells, 2021. [PubMed]
- Richard Colling, Helen Pitman, Karin Oien, Nasir Rajpoot, Philip Macklin. CM-Path AI in Histopathology Working Group, Velicia Bachtiar, Richard Booth, Alyson Bryant, Joshua Bull, et al. Artificial intelligence in digital pathology: a roadmap to routine use in clinical practice. J Pathol, 2019. [PubMed]
- Vipul Baxi, Robin Edwards, Michael Montalto, Saurabh Saha. Digital pathology and artificial intelligence in translational medicine and clinical practice. Mod Pathol, 2022. [PubMed]
- Taro Sakamoto, Tomoi Furukawa, Kris Lami. A narrative review of digital pathology and artificial intelligence: focusing on lung cancer. Trans Lung Cancer Res, 2020
- Jerome Y. Cheng, Jacob T. Abel, Ulysses G.J. Balis, David S. McClintock, Liron Pantanowitz. Challenges in the development, deployment, and regulation of artificial intelligence in anatomic pathology. Am J Pathol, 2021. [PubMed]
- Romain Brixtel, Sébastien Bougleux, Olivier Lézoray. Whole slide image quality in digital pathology: review and perspectives. IEEE Access, 2022
- Artem Shmatko, Narmin Ghaffari Laleh, Moritz Gerstung, Jakob Nikolas Kather. Artificial intelligence in histopathology: enhancing cancer research and clinical oncology. Nat Cancer, 2022. [PubMed]
- Yasmine Makhlouf, Manuel Salto-Tellez, Jacqueline James, Paul O’Reilly, Perry Maxwell. General roadmap and core steps for the development of ai tools in digital pathology. Diagnostics, 2022. [PubMed]
- Yuankai Huo, Ruining Deng, Quan Liu, Agnes B. Fogo, Haichun Yang. Ai applications in renal pathology. Kidney Int, 2021. [PubMed]
- Ahmed Serag, Adrian Ion-Margineanu, Hammad Qureshi. Translational ai and deep learning in diagnostic pathology. Front Med, 2019
- Alex Ngai Nick Wong, Zebang He, Ka Long Leung. Current developments of artificial intelligence in digital pathology and its future clinical applications in gastrointestinal cancers. Cancers, 2022. [PubMed]
- Manal AlAmir, Manal AlGhamdi. The role of generative adversarial network in medical image analysis: An in-depth survey. ACM Comput Surv, 2022
- Didem Cifci, Sebastian Foersch, Jakob Nikolas Kather. Artificial intelligence to identify genetic alterations in conventional histopathology. J Pathol, 2022. [PubMed]
- Sarah Haggenmüller, Roman C. Maron, Achim Hekler. Skin cancer classification via convolutional neural networks: systematic review of studies involving human experts. Eur J Cancer, 2021. [PubMed]
- 57Margaret Mitchell, Simone Wu, Andrew Zaldivar, Parker Barnes, Lucy Vasserman, Ben Hutchinson, Elena Spitzer, Inioluwa Deborah Raji, and Timnit Gebru. Model cards for model reporting. In Proceedings of the conference on fairness, accountability, and transparency, pages 220 – 229, 2019.
- Gu Hongyan, Jingbin Huang, Lauren Hung, Xiang Anthony Chen. Proceedings of the ACM on Human-Computer Interaction, 2021. [PubMed]
- John E. Tomaszewski. Artificial Intelligence and Deep Learning in Pathology, 2021
- Jia Deng, Wei Dong, Richard Socher, Li-Jia Li, Kai Li, Li Fei-Fei. IEEE Conference on Computer Vision and Pattern Recognition, 2009
- Geert Litjens, Peter Bandi, Babak Ehteshami Bejnordi. 1399 h&e-stained sentinel lymph node sections of breast cancer patients: the camelyon dataset. Gigascience, 2018. [PubMed]
- Gabriele Campanella, Matthew G. Hanna, Luke Geneslaw. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat Med, 2019. [PubMed]
- Ming Y. Lu, Richard J. Chen, Faisal Mahmood. Medical Imaging: Digital Pathology, 2020
- Ming Y. Lu, Drew F.K. Williamson, Tiffany Y. Chen, Richard J. Chen, Matteo Barbieri, Faisal Mahmood. Data-efficient and weakly supervised computational pathology on whole-slide images. Nature. Biomed Eng, 2021
- J. Richard, Chengkuan Chen, Yicong Li. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2022
- Ting Chen, Simon Kornblith, Mohammad Norouzi, Geoffrey Hinton. International conference on machine learning, 2020
- Mathilde Caron, Hugo Touvron, Ishan Misra. Proceedings of the IEEE/CVF International Conference on Computer Vision, 2021
- Sarah W. Njoroge, James H. Nichols. Risk management in the clinical laboratory. Ann Lab Med, 2014. [PubMed]
- Andrew A. Renshaw, Mercy Mena-Allauca, Edwin W. Gould, S. Joseph Sirintrapun. Synoptic reporting: Evidence-based review and future directions. JCO Clinical. Cancer Inform, 2018
- Ekkehard Hewer. The oncologist’s guide to synoptic reporting: a primer. Oncology, 2020. [PubMed]
- Geert Litjens, Clara I. Sánchez, Nadya Timofeeva. Deep learning as a tool for increased accuracy and efficiency of histopathological diagnosis. Sci Rep, 2016. [PubMed]
- Yun Liu, Timo Kohlberger, Mohammad Norouzi. Artificial intelligence–based breast cancer nodal metastasis detection: Insights into the black box for pathologists. Arch Pathol Lab Med, 2019. [PubMed]
- Umair Akhtar Hasan Khan, Carolin Stürenberg, Oguzhan Gencoglu. European Congress on Digital Pathology, 2019
- Javad Noorbakhsh, Saman Farahmand, Sandeep Namburi. Deep learning-based cross-classifications reveal conserved spatial behaviors within tumor histological images. Nat Commun, 2020. [PubMed]
- Jason W. Wei, Laura J. Tafe, Yevgeniy A. Linnik, Louis J. Vaickus, Naofumi Tomita, Saeed Hassanpour. Pathologist-level classification of histologic patterns on resected lung adenocarcinoma slides with deep neural networks. Sci Rep, 2019. [PubMed]
- Jocelyn Barker, Assaf Hoogi, Adrien Depeursinge, Daniel L. Rubin. Automated classification of brain tumor type in whole-slide digital pathology images using local representative tiles. Med Image Anal, 2016. [PubMed]
- Korsuk Sirinukunwattana, Shan E. Ahmed, Yee-Wah Tsang Raza, David R.J. Snead, Ian A. Cree, Nasir M. Rajpoot. Locality sensitive deep learning for detection and classification of nuclei in routine colon cancer histology images. IEEE Trans Med Imaging, 2016. [PubMed]
- Bruno Korbar, Andrea M. Olofson, Allen P. Miraflor. Deep learning for classification of colorectal polyps on whole-slide images. J Pathol Inform, 2017
- Lyndon Chan, Mahdi S. Hosseini, Konstantinos N. Plataniotis. A comprehensive analysis of weakly-supervised semantic segmentation in different image domains. Int J Comput Vision, 2021
- Hao Chen, Xiaojuan Qi, Lequan Yu, Pheng-Ann Heng. IEEE Conference on Computer Vision and Pattern Recognition, 2016
- Ramin Nateghi, Habibollah Danyali, Mohammad-Sadegh Helfroush. Iranian Conference on Electrical Engineering (ICEE), 2016
- Chaoyang Yan, Xu Jun, Jiawei Xie, Chengfei Cai, Lu. Haoda. IEEE International Symposium on Biomedical Imaging, 2020
- Korsuk Sirinukunwattana, David R.J. Snead, Nasir M. Rajpoot. A stochastic polygons model for glandular structures in colon histology images. IEEE Trans Med Imaging, 2015. [PubMed]
- Jon N. Marsh, Matthew K. Matlock, Satoru Kudose. Deep learning global glomerulosclerosis in transplant kidney frozen sections. IEEE Trans Med Imaging, 2018. [PubMed]
- Amin Zadeh Shirazi, Eric Fornaciari, Narjes Sadat Bagherian, Lisa M. Ebert, Barbara Koszyca, Guillermo A. Gomez. Deepsurvnet: deep survival convolutional network for brain cancer survival rate classification based on histopathological images. Med Biol Eng Comput, 2020. [PubMed]
- Jakob Nikolas Kather, Johannes Krisam, Pornpimol Charoentong. Predicting survival from colorectal cancer histology slides using deep learning: A retrospective multicenter study. PLoS Med, 2019
- Pierre Courtiol, Charles Maussion, Matahi Moarii. Deep learning-based classification of mesothelioma improves prediction of patient outcome. Nat Med, 2019. [PubMed]
- Dmitrii Bychkov, Nina Linder, Riku Turkki. Deep learning based tissue analysis predicts outcome in colorectal cancer. Sci Rep, 2018. [PubMed]
- Xiangxue Wang, Yu Andrew Janowczyk, Rajat Thawani Zhou. Prediction of recurrence in early stage non-small cell lung cancer using computer extracted nuclear features from digital h&e images. Sci Rep, 2017. [PubMed]
- Rebecca L. Siegel, Kimberly D. Miller, Ahmedin Jemal. Cancer statistics, 2020. CA Cancer J Clin, 2020. [PubMed]
- 91American Cancer Society. What is breast cancer? 2023. Available at: https://www.cancer.org/cancer/breast-cancer/about/what-is-breast-cancer.html. (accessed Jan 22, 2023).
- Connolly Fitzgibbons. Protocol for the examination of resection specimens from patients with ductal carcinoma in situ (dcis) of the breast. College Am Pathol, 2021
- Kimberly H. Allison, M. Elizabeth, H. Hammond. Estrogen and progesterone receptor testing in breast cancer: Asco/cap guideline update. J Clin Oncol, 2020. [PubMed]
- Antonio C. Wolff, M. Elizabeth Hale Hammond, Kimberly H. Allison, Brittany E. Harvey, Lisa M. McShane, Mitchell Dowsett. Her2 testing in breast cancer: American society of clinical oncology/college of american pathologists clinical practice guideline focused update summary. J Oncol Pract, 2018. [PubMed]
- Mitch Dowsett, Torsten O. Nielsen, Roger A’Hern. Assessment of ki67 in breast cancer: recommendations from the international ki67 in breast cancer working group. J Natl Cancer Inst, 2011. [PubMed]
- Megan A. Healey, Kelly A. Hirko, Andrew H. Beck. Assessment of ki67 expression for breast cancer subtype classification and prognosis in the nurses’ health study. Breast Cancer Res Treat, 2017. [PubMed]
- Yun Liu, Krishna Gadepalli, Mohammad Norouzi. Detecting cancer metastases on gigapixel pathology images. arXiv preprint arXiv, 2017
- 98American Cancer Society. What is prostate cancer? 2023. Available at: https://www.cancer.org/cancer/prostate-cancer/about/what-is-prostate-cancer.html. (accessed Jan 21, 2023).
- Yujiro Ito, Emily A. Vertosick, Daniel D. Sjoberg. In organ-confined prostate cancer, tumor quantitation not found to aid in prediction of biochemical recurrence. Am J Surg Pathol, 2019
- Jonathan I. Epstein. Prognostic significance of tumor volume in radical prostatectomy and needle biopsy specimens. J Urol, 2011. [PubMed]
- Laurent Salomon, Olivier Levrel, Aristotelis G. Anastasiadis. Prognostic significance of tumor volume after radical prostatectomy: a multivariate analysis of pathological prognostic factors. Eur Urol, 2003. [PubMed]
- Thomas A. Stamey, John E. McNeal, Cheryl M. Yemoto, Bronislava M. Sigal, Iain M. Johnstone. Biological determinants of cancer progression in men with prostate cancer. Jama, 1999. [PubMed]
- 103J Joy Lee, I-Chun Thomas, Rosalie Nolley, Michelle Ferrari, James D Brooks, and John T Leppert. Biologic differences between peripheral and transition zone prostate cancer. Prostate, 75(2):183–190, 2015.
- Srigley Paner. Nov 2021
- Jonathan L. Wright, Bruce L. Dalkin, Lawrence D. True. Positive surgical margins at radical prostatectomy predict prostate cancer specific mortality. J Urol, 2010. [PubMed]
- Murali Varma. Intraductal carcinoma of the prostate: a guide for the practicing pathologist. Adv Anat Pathol, 2021. [PubMed]
- Rodolfo Montironi, Ming Zhou, Cristina Magi-Galluzzi, Jonathan I. Epstein. Features and prognostic significance of intraductal carcinoma of the prostate. European Urology. Oncology, 2018
- Ming Zhou. Intraductal carcinoma of the prostate: the whole story. Pathology, 2013. [PubMed]
- Ronald J. Cohen, Thomas M. Wheeler, Helmut Bonkhoff, Mark A. Rubin. A proposal on the identification, histologic reporting, and implications of intraductal prostatic carcinoma. Arch Pathol Lab Med, 2007. [PubMed]
- Charles C. Guo, Jonathan I. Epstein. Intraductal carcinoma of the prostate on needle biopsy: histologic features and clinical significance. Mod Pathol, 2006. [PubMed]
- 111American Cancer Society. Tests to diagnose and stage prostate cancer. 2023. Available at: https://www.cancer.org/cancer/prostate-cancer/detection-diagnosis-staging/how-diagnosed.html. (accessed Jan 21, 2023).
- Charlotte F. Kweldam, Mark F. Wildhagen, Ewout W. Steyerberg, Chris H. Bangma, Theodorus H. Van Der Kwast, Geert Jlh Van Leenders. Cribriform growth is highly predictive for postoperative metastasis and disease-specific death in gleason score 7 prostate cancer. Mod Pathol, 2015. [PubMed]
- Thomas K. Lee, Jae Y. Ro. Spectrum of cribriform proliferations of the prostate: from benign to malignant. Arch Pathol Lab Med, 2018. [PubMed]
- S. Emily Bachert, Anthony McDowell Jr, Dava Piecoro, Lauren Baldwin Branch. Serous tubal intraepithelial carcinoma: a concise review for the practicing pathologist and clinician. Diagnostics, 2020. [PubMed]
- 115American Cancer Society. What is ovarian cancer? Available at: https://www.cancer.org/cancer/ovarian-cancer/about/what-is-ovarian-cancer.html. (accessed Jan 21, 2023).
- Shi-Ping Yang, Su Hui-Luan, Xiu-Bei Chen. Long-term survival among histological subtypes in advanced epithelial ovarian cancer: population-based study using the surveillance, epidemiology, and end results database. JMIR Public Health Surveill, 2021
- Lisa Vermij, Alicia Léon-Castillo, Naveena Singh. p53 immunohistochemistry in endometrial cancer: clinical and molecular correlates in the portec-3 trial. Mod Pathol, 2022. [PubMed]
- Yu Zhang, Lan Cao, Daniel Nguyen, Lu. Hua. Tp53 mutations in epithelial ovarian cancer. Transl Cancer Res, 2016. [PubMed]
- Yiping Wang, David Farnell, Hossein Farahani. Medical Imaging with Deep Learning, 2020
- Jevgenij Gamper, Navid Alemi Kooohbanani, Nasir Rajpoot. Multi-task learning in histo-pathology for widely generalizable model. arXiv preprint arXiv, 2020
- 121National Cancer Institute. Common cancer types. Available at: https://www.cancer.gov/types/common-cancers#:∼:text=The most common type of,are combined for the list/. (accessed June 10, 2021).
- 122American Cancer Society. Tests to diagnose and stage prostate cancer. Available at: https://www.cancer.org/cancer/lung-cancer/about/what-is.html. (accessed Jan 21, 2023).
- Akihiko Yoshizawa, Noriko Motoi, Gregory J. Riely. Impact of proposed iaslc/ats/ers classification of lung adenocarcinoma: prognostic subgroups and implications for further revision of staging based on analysis of 514 stage i cases. Mod Pathol, 2011. [PubMed]
- Andre L. Moreira, Paolo S.S. Ocampo, Yuhe Xia. A grading system for invasive pulmonary adenocarcinoma: a proposal from the international association for the study of lung cancer pathology committee. J Thorac Oncol, 2020. [PubMed]
- Yasuhiro Tsutani, Yoshihiro Miyata, Haruhiko Nakayama. Prognostic significance of using solid versus whole tumor size on high-resolution computed tomography for predicting pathologic malignant grade of tumors in clinical stage ia lung adenocarcinoma: a multicenter study. J Thorac Cardiovasc Surg, 2012. [PubMed]
- Tatsuo Maeyashiki, Kenji Suzuki, Aritoshi Hattori, Takeshi Matsunaga, Kazuya Takamochi, Oh. Shiaki. The size of consolidation on thin-section computed tomography is a better predictor of survival than the maximum tumour dimension in resectable lung cancer. Eur J Cardiothorac Surg, 2013. [PubMed]
- B. Mahul, Frederick L. Greene, Stephen B. Edge. The eighth edition ajcc cancer staging manual: continuing to build a bridge from a population-based to a more “personalized” approach to cancer staging. CA Cancer J Clin, 2017. [PubMed]
- Liang Wang, Xuejun Dou, Tao Liu, Lu Weiqiang, Yunlei Ma, Yue Yang. Tumor size and lymph node metastasis are prognostic markers of small cell lung cancer in a chinese population. Medicine, 2018
- Jianjun Zhang, Kathryn A. Gold, Heather Y. Lin. Relationship between tumor size and survival in non–small-cell lung cancer (nsclc): an analysis of the surveillance, epidemiology, and end results (seer) registry. J Thorac Oncol, 2015. [PubMed]
- Yina Gao, Yangyang Dong, Yingxu Zhou. Peripheral tumor location predicts a favorable prognosis in patients with resected small cell lung cancer. Int J Clin Pract, 2022
- Mahul B. Amin, Stephen B. Edge, Frederick L. Greene. 2017
- American Cancer Society. 2020
- 133American Cancer Society. What is colorectal cancer? 2023. Available at: https://www.cancer.org/cancer/colon-rectal-cancer/about/what-is-colorectal-cancer.html. (accessed Jan 21, 2023).
- Y. Nancy You, Karin M. Hardiman, Andrea Bafford. The american society of colon and rectal surgeons clinical practice guidelines for the management of rectal cancer. Dis Colon Rectum, 2020. [PubMed]
- Seok-Byung Lim, Yu Chang Sik, Se Jin Jang, Tae Won Kim, Jong Hoon Kim, Jin Cheon Kim. Prognostic significance of lymphovascular invasion in sporadic colorectal cancer. Dis Colon Rectum, 2010. [PubMed]
- C. Santos, A. López-Doriga, M. Navarro. Clinicopathological risk factors of stage ii colon cancer: results of a prospective study. Colorectal Dis, 2013. [PubMed]
- Dhanwant Gomez, Abed M. Zaitoun, Antonella De Rosa. Critical review of the prognostic significance of pathological variables in patients undergoing resection for colorectal liver metastases. HPB, 2014. [PubMed]
- Catherine Liebig, Gustavo Ayala, Jonathan Wilks. Perineural invasion is an independent predictor of outcome in colorectal cancer. J Clin Oncol, 2009. [PubMed]
- H. Ueno, K. Shirouzu, Y. Eishi. Study group for perineural invasion projected by the japanese society for cancer of the colon and rectum (jsccr). characterization of perineural invasion as a component of colorectal cancer staging. Am J Surg Pathol, 2013. [PubMed]
- Amanda I. Phipps, Noralane M. Lindor, Mark A. Jenkins. Colon and rectal cancer survival by tumor location and microsatellite instability: the colon cancer family registry. Dis Colon Rectum, 2013. [PubMed]
- Marco Vacante, Antonio Maria Borzì, Francesco Basile, Antonio Biondi. Biomarkers in colorectal cancer: Current clinical utility and future perspectives. World J Clin Cases, 2018. [PubMed]
- Xu Yan, Zhipeng Jia, Liang-Bo Wang. Large scale tissue histopathology image classification, segmentation, and visualization via deep convolutional activation features. BMC Bioinforma, 2017
- American Cancer Society. What is bladder cancer?. 2023
- Venu Chalasani, Joseph L. Chin, Jonathan I. Izawa. Histologic variants of urothelial bladder cancer and nonurothelial histology in bladder cancer. Can Urol Assoc J, 2009. [PubMed]
- Yair Lotan, Amit Gupta, Shahrokh F. Shariat. Lymphovascular invasion is independently associated with overall survival, cause-specific survival, and local and distant recurrence in patients with negative lymph nodes at radical cystectomy. J Clin Oncol, 2005. [PubMed]
- Ann-Christin Woerl, Markus Eckstein, Josephine Geiger. Deep learning predicts molecular subtype of muscle-invasive bladder cancer from conventional histopathological slides. Eur Urol, 2020. [PubMed]
- American Cancer Society. What is kidney cancer?. 2023
- R. John, M. Zhou, R. Allan. Protocol for the examination of resection specimens from patients with invasive carcinoma of renal tubular origin. College American Pathologists (CAP) Cancer Protocols, 2020
- Stephen M. Bonsib. Renal lymphatics, and lymphatic involvement in sinus vein invasive (pt3b) clear cell renal cell carcinoma: a study of 40 cases. Mod Pathol, 2006. [PubMed]
- Valdair F. Muglia, Adilson Prando. Renal cell carcinoma: histological classification and correlation with imaging findings. Radiol Bras, 2015. [PubMed]
- Shruti Kannan, Laura A. Morgan, Benjamin Liang. Segmentation of glomeruli within trichrome images using deep learning. Kidney Int Rep, 2019. [PubMed]
- Meyke Hermsen, Thomas de Bel, Marjolijn Den Boer. Deep learning–based histopathologic assessment of kidney tissue. J Am Soc Nephrol, 2019. [PubMed]
- Edmund A. Gehan, Michael D. Walker. Prognostic factors for patients with brain tumors. Nat Cancer Inst Monogr, 1977. [PubMed]
- Yao Li, Zuo-Xin Zhang, Guo-Hao Huang. A systematic review of multifocal and multicentric glioblastoma. J Clin Neurosci, 2021. [PubMed]
- American Cancer Society. Brain tumors – classifications, symptoms, diagnosis and treatments. 2023
- David N. Louis, Arie Perry, Pieter Wesseling. The 2021 who classification of tumors of the central nervous system: a summary. Neuro-oncology, 2021. [PubMed]
- WHO Classification of Tumours Editorial Board. 2021
- 158Canadian Cancer Society. Survival statistics for brain and spinal cord tumours. Available at: https://www.cancer.ca/en/cancer-information/cancer-type/brain-spinal/prognosis-and-survival/survival-statistics/?region=on#:∼:text=In Canada, the 5-year,survive at least 5 years. (accessed June 10, 2021).
- World Health Organization. Cancer
- Haeryoung Kim, Mi Jang, Young Nyun Park. Histopathological variants of hepatocellular carcinomas: an update according to the 5th edition of the who classification of digestive system tumors. J Liver Cancer, 2020. [PubMed]
- Gregory Y. Lauwers, Benoit Terris, Ulysses J. Balis. Prognostic histologic indicators of curatively resected hepatocellular carcinomas: a multi-institutional analysis of 425 patients with definition of a histologic prognostic index. Am J Surg Pathol, 2002. [PubMed]
- Gaya Spolverato, Yuhree Kim, Sorin Alexandrescu. Is hepatic resection for large or multifocal intrahepatic cholangiocarcinoma justified? results from a multi-institutional collaboration. Ann Surg Oncol, 2015. [PubMed]
- Ian R. Wanless. Terminology of nodular hepatocellular lesions. Hepatology, 1995. [PubMed]
- Sebastiao N. Martins-Filho, Caterina Paiva, Raymundo Soares Azevedo, Venancio Avancini Ferreira Alves. Histological grading of hepatocellular carcinoma—a systematic review of literature. Front Med, 2017
- American Cancer Society. Lymph nodes and cancer. 2023
- Zhihua Wang, Yu Lequan, Xin Ding, Xuehong Liao, Liansheng Wang. Lymph node metastasis prediction from whole slide images with transformer-guided multi-instance learning and knowledge transfer. IEEE Trans Med Imaging, 2022. [PubMed]
- S. Mahdi, Lyndon Chan, Gabriel Tse. IEEE Conference on Computer Vision and Pattern Recognition, 2019
- Navid Alemi Koohbanani, Mostafa Jahanifar, Neda Zamani Tajadin, Nasir Rajpoot. Nuclick: a deep learning framework for interactive segmentation of microscopic images. Med Image Anal, 2020
- Ruqayya Awan, Korsuk Sirinukunwattana, David Epstein. Glandular morphometrics for objective grading of colorectal adenocarcinoma histology images. Sci Rep, 2017. [PubMed]
- Wenyuan Li, Jiayun Li, Zichen Wang. Pathal: An active learning framework for histopathology image analysis. IEEE Trans Med Imaging, 2021
- Farzad Ghaznavi, Andrew Evans, Anant Madabhushi, Michael Feldman. Digital imaging in pathology: whole-slide imaging and beyond. Ann Rev Pathol Mech Dis, 2013
- Julianna D. Ianni, Rajath E. Soans, Sivaramakrishnan Sankarapandian. Tailored for real-world: a whole slide image classification system validated on uncurated multi-site data emulating the prospective pathology workload. Sci Rep, 2020. [PubMed]
- Geoffrey Rolls. Biosystems, 2016
- S. Kim Suvarna, Christopher Layton, John D. Bancroft. 2019
- R. Stephen. 2016
- Yukako Yagi. Diagnostic pathology, 2011. [PubMed]
- Babak Ehteshami Bejnordi, Geert Litjens, Nadya Timofeeva. Stain specific standardization of whole-slide histopathological images. IEEE Trans Med Imaging, 2015. [PubMed]
- Babak Ehteshami Bejnordi, Nadya Timofeeva, Irene Otte-Höller, Nico Karssemeijer, Jeroen A.W.M. van der Laak. Medical Imaging: Digital Pathology, 2014
- Daisuke Komura, Shumpei Ishikawa. Machine learning methods for histopathological image analysis. Comput Struct Biotechnol J, 2018. [PubMed]
- Mark D. Zarella, Douglas Bowman, Famke Aeffner. A practical guide to whole slide imaging: a white paper from the digital pathology association. Arch Pathol Lab Med, 2019. [PubMed]
- Liron Pantanowitz. Digital images and the future of digital pathology. J Pathol Inform, 2010
- Liron Pantanowitz, Ashish Sharma, Alexis B. Carter, Tahsin Kurc, Alan Sussman, Joel Saltz. Twenty years of digital pathology: an overview of the road travelled, what is on the horizon, and the emergence of vendor-neutral archives. J Pathol Inform, 2018
- Md Shakhawat Hossain, Toyama Nakamura, Fumikazu Kimura, Yukako Yagi, Masahiro Yamaguchi. Biomedical Imaging and Sensing Conference, 2018
- Kazuhiro Tabata, Naohiro Uraoka, Jamal Benhamida. Validation of mitotic cell quantification via microscopy and multiple whole-slide scanners. Diagn Pathol, 2019. [PubMed]
- Wei-Chung Cheng, Firdous Saleheen, Aldo Badano. Assessing color performance of whole-slide imaging scanners for digital pathology. Color Res Appl, 2019
- Paul Lemaillet, Kazuyo Takeda, Andrew C. Lamont, Anant Agrawal. Colorimetrical uncertainty estimation for the performance assessment of whole slide imaging scanners. Journal of Medical. Imaging, 2021
- M. Indu, R. Rathy, M.P. Binu. “slide less pathology”: Fairy tale or reality?. J Oral Maxillofacial Pathol, 2016
- Markus D. Herrmann, David A. Clunie, Andriy Fedorov. Implementing the dicom standard for digital pathology. J Pathol Inform, 2018
- Tiago Marques Godinho, Rui Lebre, Luís Bastião Silva, Carlos Costa. An efficient architecture to support digital pathology in standard medical imaging repositories. J Biomed Inform, 2017. [PubMed]
- David A. Clunie. Dicom format and protocol standardization—a core requirement for digital pathology success. Toxicol Pathol, 2021. [PubMed]
- 191Fusheng Wang, Tae W Oh, Cristobal Vergara-Niedermayr, Tahsin Kurc, and Joel Saltz. Managing and querying whole slide images. In Medical Imaging 2012: Advanced PACS-Based Imaging Informatics and Therapeutic Applications, volume 8319, pages 137–148. SPIE, 2012.
- 192Daniel E Lopez Barron, Dig Vijay Kumar Yarlagadda, Praveen Rao, Ossama Tawfik, and Deepthi Rao. Scalable storage of whole slide images and fast retrieval of tiles using apache spark. In Medical Imaging: Digital Pathology, volume 10581, page 1058113 International Society for Optics and Photonics, 2018.
- Rajendra Singh, Lauren Chubb, Liron Pantanowitz, Anil Parwani. Standardization in digital pathology: Supplement 145 of the dicom standards. J Pathol Inform, 2011
- Neel Kanwal, Fernando Pérez-Bueno, Arne Schmidt, Kjersti Engan, Rafael Molina. The devil is in the details: Whole slide image acquisition and processing for artifacts detection, color variation, and data augmentation: A review. IEEE Access, 2022
- Gabriele Campanella, Arjun R. Rajanna, Lorraine Corsale, Peter J. Schüffler, Yukako Yagi, Thomas J. Fuchs. Towards machine learned quality control: A benchmark for sharpness quantification in digital pathology. Comput Med Imaging Graph, 2018. [PubMed]
- Zhongling Wang, Mahdi S. Hosseini, Adyn Miles, Konstantinos N. Plataniotis, Zhou Wang. Medical Image Computing and Computer-Assisted Intervention – MICCAI, volume 12265 of Lecture Notes in Computer Science, 2020
- Simon Cross, Peter Furness, Laszlo Igali, David Snead, Darren Treanor. Technical Report G162, The Royal College of Pathologists, 4th Floor 21 Prescott Street, London, United Kingdom E1 8BB, 2018
- Péter Bándi, Maschenka Balkenhol, Bram van Ginneken, Jeroen van der Laak, Geert Litjens. Resolution-agnostic tissue segmentation in whole-slide histopathology images with convolutional neural networks. PeerJ, 2019
- David Tellez, Geert Litjens, Péter Bándi. Quantifying the effects of data augmentation and stain color normalization in convolutional neural networks for computational pathology. Med Image Anal, 2019
- Sebastian Otálora, Manfredo Atzori, Vincent Andrearczyk, Amjad Khan, Henning Müller. Staining invariant features for improving generalization of deep convolutional neural networks in computational pathology. Front Bioeng Biotechnol, 2019. [PubMed]
- Savannah R. Duenweg, Samuel A. Bobholz, Allison K. Lowman. Whole slide imaging (wsi) scanner differences influence optical and computed properties of digitized prostate cancer histology. J Pathol Inform, 2023
- Adnan Mujahid Khan, Nasir Rajpoot, Darren Treanor, Derek Magee. A nonlinear mapping approach to stain normalization in digital histopathology images using image-specific color deconvolution. IEEE Trans Biomed Eng, 2014. [PubMed]
- Farhad Ghazvinian Zanjani, Svitlana Zinger, Babak E. Bejnordi. 1st Conference on Medical Imaging with Deep Learning (MIDL), 2018
- Amit Sethi, Lingdao Sha, Abhishek Ramnath Vahadane. Empirical comparison of color normalization methods for epithelial-stromal classification in h and e images. J Pathol Inform, 2016
- Rodrigo Escobar Díaz Guerrero, José Luís Oliveira. 2021
- Rikiya Yamashita, Jin Long, Snikitha Banda, Jeanne Shen, Daniel L. Rubin. Learning domain-agnostic visual representation for computational pathology using medically-irrelevant style transfer augmentation. IEEE Trans Med Imaging, 2021. [PubMed]
- Abhishek Vahadane, B. Atheeth, Shantanu Majumdar. 2021
- Ida Arvidsson, Niels Christian Overgaard, Kalle Åström, Anders Heyden. IEEE International Symposium on Biomedical Imaging, 2019
- M. Tarek Shaban, Christoph Baur, Nassir Navab, Shadi Albarqouni. IEEE International Symposium on Biomedical Imaging, 2019
- Dwarikanath Mahapatra, Behzad Bozorgtabar, Jean-Philippe Thiran, Ling Shao. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2020
- Shahid Mehmood, Taher M. Ghazal, Muhammad Adnan Khan. Malignancy detection in lung and colon histopathology images using transfer learning with class selective image processing. IEEE Access, 2022
- David Tellez, Geert Litjens, Jeroen van der Laak, Francesco Ciompi. Neural image compression for gigapixel histopathology image analysis. IEEE Trans Pattern Anal Mach Intell, 2019
- Rui Yan, Fei Ren, Zihao Wang. Breast cancer histopathological image classification using a hybrid deep neural network. Methods, 2020. [PubMed]
- Shidan Wang, Ruichen Rong, Donghan M. Yang. Computational staining of pathology images to study the tumor microenvironment in lung cancer. Cancer Res, 2020. [PubMed]
- Yiyang Lin, Bowei Zeng, Yifeng Wang. 2022
- Jerry Wei, Arief Suriawinata, Bing Ren. IEEE/CVF Conference on Computer Vision, 2021
- Marc D. Kohli, Ronald M. Summers, J. Raymond, Geis. Medical image data and datasets in the era of machine learning—whitepaper from the 2016 c-mimi meeting dataset session. J Digit Imaging, 2017. [PubMed]
- Noorul Wahab, Asifullah Khan, Yeon Soo Lee. Two-phase deep convolutional neural network for reducing class skewness in histopathological images based breast cancer detection. Comput Biol Med, 2017. [PubMed]
- Angel Cruz-Roa, Hannah Gilmore, Ajay Basavanhally. High-throughput adaptive sampling for whole-slide histopathology image analysis (hashi) via convolutional neural networks: Application to invasive breast cancer detection. PloS One, 2018
- Guilherme Aresta, Teresa Araújo, Scotty Kwok. Bach: Grand challenge on breast cancer histology images. Med Image Anal, 2019. [PubMed]
- Oliver Lester Saldanha, Philip Quirke, Nicholas P. West. Swarm learning for decentralized artificial intelligence in cancer histopathology. Nat Med, 2022. [PubMed]
- Martin J. Willemink, Wojciech A. Koszek, Cailin Hardell. Preparing medical imaging data for machine learning. Radiology, 2020. [PubMed]
- Sharib Ali, Nasullah Khalid Alham, Clare Verrill, Jens Rittscher. IEEE International Symposium on Biomedical Imaging, 2019
- Osamu Iizuka, Fahdi Kanavati, Kei Kato, Michael Rambeau, Koji Arihiro, Masayuki Tsuneki. Deep learning models for histopathological classification of gastric and colonic epithelial tumours. Sci Rep, 2020. [PubMed]
- J. Deng, W. Dong, R. Socher, L.-J. Li, K. Li, L. Fei-Fei. CVPR09, 2009
- Fabio A. Spanhol, Luiz S. Oliveira, Caroline Petitjean, Laurent Heutte. A dataset for breast cancer histopathological image classification. IEEE Trans Biomed Eng, 2015. [PubMed]
- Dalal Bardou, Kun Zhang, Sayed Mohammad Ahmad. Classification of breast cancer based on histology images using convolutional neural networks. IEEE Access, 2018
- Neslihan Bayramoglu, Juho Kannala, Janne Heikkilä. International Conference on Pattern Recognition, 2016
- Xia Li, Xi Shen, Yongxia Zhou, Xiuhui Wang, Tie-Qiang Li. Classification of breast cancer histopathological images using interleaved densenet with senet (idsnet). PloS One, 2020
- Douglas Joseph Hartman, Jeroen A.W.M. Van Der Laak, Metin N. Gurcan, Liron Pantanowitz. Value of public challenges for the development of pathology deep learning algorithms. J Pathol Inform, 2020
- Babak Ehteshami Bejnordi, Mitko Veta, Paul Johannes Van Diest. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA Netw Open, 2017
- Péter Bándi, Oscar Geessink, Quirine Manson. Bram van Ginneken, Jeroen van der Laak, and Geert Litjens. From detection of individual metastases to classification of lymph node status at the patient level: The camelyon17 challenge. IEEE Trans Med Imaging, 2019. [PubMed]
- Korsuk Sirinukunwattana, Josien P.W. Pluim, Hao Chen. Gland segmentation in colon histology images: The glas challenge contest. Med Image Anal, 2017. [PubMed]
- Vu Le Hou, Ariel B. Nguyen, Dimitris Samaras Kanevsky. Sparse autoencoder for unsupervised nucleus detection and representation in histopathology images. Pattern Recogn Lett, 2019
- Robert L. Grossman, Allison P. Heath, Vincent Ferretti. Toward a shared vision for cancer genomic data. N Engl J Med, 2016. [PubMed]
- Kyung-Ok Cho, Sung Hak Lee, Hyun-Jong Jang. Feasibility of fully automated classification of whole slide images based on deep learning. Korean J Physiol & Pharmacol, 2020. [PubMed]
- Le Hou, Rajarsi Gupta, John S. Van Arnam. Dataset of segmented nuclei in hematoxylin and eosin stained histopathology images of ten cancer types. Sci Data, 2020. [PubMed]
- Pooya Mobadersany, Safoora Yousefi, Mohamed Amgad. Predicting cancer outcomes from histology and genomics using convolutional networks. Natl Acad Sci, 2018
- Will Fischer, Sanketh S. Moudgalya, Judith D. Cohn, Nga T.T. Nguyen, Garrett T. Kenyon. Sparse coding of pathology slides compared to transfer learning with deep neural networks. BMC Bioinform, 2018
- Arkadiusz Gertych, Zaneta Swiderska-Chadaj, Zhaoxuan Ma. Convolutional neural networks can accurately distinguish four histologic growth patterns of lung adenocarcinoma in digital slides. Sci Rep, 2019. [PubMed]
- Keisuke Nakagawa, Lama Moukheiber, Leo A. Celi. Ai in pathology: What could possibly go wrong?. Semin Diagn Pathol, 2023. [PubMed]
- Chhavi Chauhan, Rama Gullapalli. Ethics of ai in pathology: Current paradigms and emerging issues. Am J Pathol, 2021. [PubMed]
- Zeyu Wang, Klint Qinami, Ioannis Christos Karakozis. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), June 2020
- Xu Tian, Jennifer White, Sinan Kalkan, Hatice Gunes. Computer Vision – ECCV 2020 Workshops, 2020
- Simone Fabbrizzi, Symeon Papadopoulos, Eirini Ntoutsi, Ioannis Kompatsiaris. A survey on bias in visual datasets. Comput Vis Image Underst, 2022
- Markos Georgopoulos, Yannis Panagakis, Maja Pantic. Investigating bias in deep face analysis: The kanface dataset and empirical study. Image Vis Comput, 2020
- Ninareh Mehrabi, Fred Morstatter, Nripsuta Saxena, Kristina Lerman, Aram Galstyan. A survey on bias and fairness in machine learning. ACM Comput Surv, Jul 2021
- Taher Dehkharghanian, Azam Asilian Bidgoli, Abtin Riasatian. Biased data, biased ai: Deep networks predict the acquisition site of tcga images. Diagn Pathol, May 2023
- Frederick M. Howard, James Dolezal, Sara Kochanny. The impact of site-specific digital histology signatures on deep learning model accuracy and bias. Nat Commun, 2021
- Kevin Faust, Sudarshan Bala, Randy Van Ommeren. Intelligent feature engineering and ontological mapping of brain tumour histomorphologies by deep learning. Nat Mach Intell, 2019
- Yu Kun-Hsing, Ce Zhang, Gerald J. Berry. Predicting non-small cell lung cancer prognosis by fully automated microscopic pathology image features. Nat Commun, 2016
- Akash Parvatikar, Om Choudhary, Arvind Ramanathan. Medical Image Computing and Computer Assisted Intervention – MICCAI, volume 12265 of Lecture Notes in Computer Science, 2020
- Peter Naylor, Marick Laé, Fabien Reyal, Thomas Walter. Segmentation of nuclei in histopathology images by deep regression of the distance map. IEEE Trans Med Imaging, 2018
- Breast Cancer Surveillance Consortium. 2024
- Jerry Wei, Arief Suriawinata, Bing Ren. International Conference on Artificial Intelligence in Medicine, 2021
- Teresa Araújo, Guilherme Aresta, Eduardo Castro. Classification of breast cancer histology images using convolutional neural networks. PloS One, 2017
- Rui Yan, Fei Ren, Zihao Wang. IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 2018
- Neeraj Kumar, Ruchika Verma, Sanuj Sharma, Surabhi Bhargava, Abhishek Vahadane, Amit Sethi. A dataset and a technique for generalized nuclear segmentation for computational pathology. IEEE Trans Med Imaging, 2017. [PubMed]
- Joel Saltz, Rajarsi Gupta, Le Hou. Spatial organization and molecular correlation of tumor-infiltrating lymphocytes using deep learning on pathology images. Cell Rep, 2018. [PubMed]
- Simon Graham, Mostafa Jahanifar, Ayesha Azam. IEEE/CVF International Conference on Computer Vision, 2021
- Ferdaous Idlahcen, Mohammed Majid Himmi, Abdelhak Mahmoudi. Cnn-based approach for cervical cancer classification in whole-slide histopathology images. arXiv preprint, 2020
- David Tellez, Maschenka Balkenhol, Irene Otte-Höller. Whole-slide mitosis detection in h&e breast histology using phh3 as a reference to train distilled stain-invariant convolutional networks. IEEE Trans Med Imaging, 2018
- Young Hwan Chang, Guillaume Thibault, Owen Madin. International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), 2017
- Peter Bandi, Oscar Geessink, Quirine Manson. From detection of individual metastases to classification of lymph node status at the patient level: the camelyon17 challenge. IEEE Trans Med Imaging, 2018
- Mitko Veta, Yujing J. Heng, Nikolas Stathonikos. Predicting breast tumor proliferation from whole-slide images: the tupac16 challenge. Med Image Anal, 2019. [PubMed]
- Michał Koziarski, Bogusław Cyganek, Bogusław Olborski. Diagset: a dataset for prostate cancer histopathological image classification. arXiv preprint arXiv, 2021
- Hans Pinckaers, Wouter Bulten, Jeroen van der Laak, Geert Litjens. Detection of prostate cancer in whole-slide images through end-to-end training with image-level labels. IEEE Trans Med Imaging, 2021. [PubMed]
- Bin Li, Yin Li, Kevin W. Eliceiri. IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2021
- Jiahui Li, Wen Chen, Xiaodi Huang. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2021
- Adon Phillips, Iris Teo, Jochen Lang. Fully convolutional network for melanoma diagnostics. arXiv preprint arXiv, 2018
- Xu Hongming, Sunho Park, Jean René Clemenceau, Nathan Radakovich, Sung Hak Lee, Tae Hyun Hwang. 2020
- James A. Diao, Richard J. Chen, Joseph C. Kvedar. Efficient cellular annotation of histopathology slides with real-time ai augmentation. NPJ Digit Medi, 2021
- Runtian Miao, Yu Robert Toth, Anant Madabhushi Zhou, Andrew Janowczyk. Quick annotator: an open-source digital pathology based rapid image annotation tool. J Pathol, 2021
- Ziyu Zhang, Sanja Fidler, Raquel Urtasun. IEEE Conference on Computer Vision and Pattern Recognition, 2016
- Yuxuan Zhang, Huan Ling, Jun Gao. IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2021
- Bowen Chen, Huan Ling, Xiaohui Zeng, Jun Gao, Xu Ziyue, Sanja Fidler. European Conference on Computer Vision, 2020
- Simon Graham, Vu Quoc Dang, Shan E. Ahmed. Hover-net: Simultaneous segmentation and classification of nuclei in multi-tissue histology images. Med Image Anal, 2019
- Shusuke Takahama, Yusuke Kurose, Yusuke Mukuta. IEEE/CVF International Conference on Computer Vision, 2019
- Sajid Javed, Arif Mahmood, Muhammad Moazam Fraz. Cellular community detection for tissue phenotyping in colorectal cancer histology images. Med Image Anal, 2020
- Yuji Roh, Geon Heo, Steven Euijong Whang. A survey on data collection for machine learning: a big data-ai integration perspective. IEEE Trans Knowl Data Eng, 2019
- Noorul Wahab, Islam M. Miligy, Katherine Dodd. Semantic annotation for computational pathology: Multidisciplinary experience and best practice recommendations. J Pathol Clin Res, 2022. [PubMed]
- Nithya Sambasivan, Shivani Kapania, Hannah Highfill, Diana Akrong, Praveen Kumar Paritosh, Lora Mois Aroyo. Everyone wants to do the model work, not the data work. Data Cascades in High-Stakes Ai, 2021
- Cathy O’Neil. 2016
- Michael A. Lones. How to avoid machine learning pitfalls: a guide for academic researchers. arXiv preprint arXiv, 2021
- Yoshua Bengio, Aaron Courville, Pascal Vincent. 2012
- Yann LeCun, Yoshua Bengio, Geoffrey Hinton. Deep learning. Nature, 2015. [PubMed]
- Jakob Nikolas Kather, Alexander T. Pearson, Niels Halama. Deep learning can predict microsatellite instability directly from histology in gastrointestinal cancer. Nat Med, 2019. [PubMed]
- Naofumi Tomita, Behnaz Abdollahi, Jason Wei, Bing Ren, Arief Suriawinata, Saeed Hassanpour. Attention-based deep neural networks for detection of cancerous and precancerous esophagus tissue on histopathological slides. JAMA Netw Open, 2019
- Stefan Bauer, Nicolas Carion, Peter Schüffler, Thomas Fuchs, Peter Wild, Joachim M. Buhmann. Multi-organ cancer classification and survival analysis. arXiv preprint arXiv, 2016
- Mehdi Habibzadeh Motlagh, Mahboobeh Jannesari, HamidReza Aboulkheyr. Breast cancer histopathological image classification: A deep learning approach. BioRxiv, 2018
- Yusuf Celik, Muhammed Talo, Ozal Yildirim, Murat Karabatak, U. Rajendra, Acharya. Automated invasive ductal carcinoma detection based using deep transfer learning with whole-slide images. Pattern Recogn Lett, 2020
- Bo Liu, Kelu Yao, Mengmeng Huang, Jiahui Zhang, Yong Li, Rong Li. IEEE Annual Computer Software and Applications Conference (COMPSAC), 2018
- Jason W. Wei, Jerry W. Wei, Christopher R. Jackson, Bing Ren, Arief A. Suriawinata, Saeed Hassanpour. Automated detection of celiac disease on duodenal biopsy slides: A deep learning approach. J Pathol Inform, 2019
- Shweta Saxena, Sanyam Shukla, Manasi Gyanchandani. Pre-trained convolutional neural networks as feature extractors for diagnosis of breast cancer using histopathology. Int J Imaging Syst Technol, 2020
- Rene Bidart, Alexander Wong. International Conference on Image Analysis and Recognition, 2019
- Lorne Holland, Dongguang Wei, Kristin A. Olson. Limited number of cases may yield generalizable models, a proof of concept in deep learning for colon histology. J Pathol Inform, 2020
- Achim Hekler, Jochen S. Utikal, Alexander H. Enk. Deep learning outperformed 11 pathologists in the classification of histopathological melanoma images. Eur J Cancer, 2019. [PubMed]
- Byungjae Lee, Kyunghyun Paeng. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2018
- Abhinav Kumar, Sanjay Kumar Singh, K. Sonal Saxena. Deep feature learning for histopathological image classification of canine mammary tumors and human breast cancer. Inform Sci, 2020
- Lyndon Chan, Mahdi S. Hosseini, Corwyn Rowsell, Konstantinos N. Plataniotis, Savvas Damaskinos. IEEE/CVF International Conference on Computer Vision, 2019
- Zeya Wang, Nanqing Dong, Wei Dai, Sean D. Rosario, Eric P. Xing. International Conference Image Analysis and Recognition, 2018
- Ming Y. Lu, Tiffany Y. Chen, Drew F.K. Williamson. Ai-based pathology predicts origins for cancers of unknown primary. Nature, 2021. [PubMed]
- Pushpanjali Gupta, Yenlin Huang, Prasan Kumar Sahoo. Colon tissues classification and localization in whole slide images using deep learning. Diagnostics, 2021. [PubMed]
- Ruiwei Feng, Xuechen Liu, Jintai Chen, Danny Z. Chen, Honghao Gao, Wu. Jian. A deep learning approach for colonoscopy pathology wsi analysis: accurate segmentation and classification. IEEE J Biomed Health Inform, 2020
- Saman Farahmand, Aileen I. Fernandez, Fahad Shabbir Ahmed. Deep learning trained on hematoxylin and eosin tumor region of interest predicts her2 status and trastuzumab treatment response in her2+ breast cancer. Mod Pathol, 2022. [PubMed]
- Ozan Ciga, Xu Tony, Anne Louise Martel. Self supervised contrastive learning for digital histopathology. Mach Learn Appl, 2022
- Nicole Bussola, Alessia Marcolini, Valerio Maggio, Giuseppe Jurman, Cesare Furlanello. International Conference on Pattern Recognition, 2021
- Afaf Alharbi, Yaqi Wang, Qianni Zhang. International Conference on Biomedical Signal and Image Processing, 2021
- Junlong Cheng, Shengwei Tian, Yu Long. Resganet: Residual group attention network for medical image classification and segmentation. Med Image Anal, 2022
- Eu Wern Teh, Graham W. Taylor. Learning with less labels in digital pathology via scribble supervision from natural images. arXiv preprint arXiv, 2022
- Su Andrew, HoJoon Lee, Xiao Tan. A deep learning model for molecular label transfer that enables cancer cell identification from histopathology images. NPJ Precis Oncol, 2022. [PubMed]
- Jiawei Yang, Hanbo Chen, Yuan Liang, Junzhou Huang, Lei He, Jianhua Yao. European Conference on Computer Vision, 2022
- Chuanqi Tan, Fuchun Sun, Tao Kong, Wenchang Zhang, Chao Yang, Chunfang Liu. International conference on artificial neural networks, 2018
- Thomas N. Kipf, Max Welling. International Conference on Learning Representations, 2017
- Yonghang Guan, Jun Zhang, Kuan Tian. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2022
- Richard J. Chen, Ming Y. Lu, Muhammad Shaban. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2021
- Alexey Dosovitskiy, Lucas Beyer, Alexander Kolesnikov. International Conference on Learning Representations, 2024
- Daniel Bug, Steffen Schneider, Anne Grote. Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support, 2017
- Zeyu Gao, Bangyang Hong, Xianli Zhang. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2021
- Kelei He, Chen Gan, Zhuoyuan Li. Transformers in medical image analysis. Intell Med, 2023
- Yun Jiang, Li Chen, Hai Zhang, Xiao Xiao. Breast cancer histopathological image classification using convolutional neural networks with small se-resnet module. PloS One, 2019
- Yi Li, Wei Ping. Cancer metastasis detection with neural conditional random field. arXiv preprint arXiv, 2018
- Artem Pimkin, Gleb Makarchuk, Vladimir Kondratenko, Maxim Pisov, Egor Krivov, Mikhail Belyaev. International Conference Image Analysis and Recognition, 2018
- Sara Hosseinzadeh Kassani, Peyman Hosseinzadeh Kassani, Michal J. Wesolowski, Kevin A. Schneider, Ralph Deters. International Conference on Computer Science and Software Engineering, 2019
- Wenqi Lu, Simon Graham, Mohsin Bilal, Nasir Rajpoot, Fayyaz Minhas. IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops, 2020
- Jiatai Lin, Guoqiang Han, Xipeng Pan. Pdbl: Improving histopathological tissue classification with plug-and-play pyramidal deep-broad learning. IEEE Transactions on Medical Imaging, 2022. [PubMed]
- Zakaria Senousy, Mohammed M. Abdelsamea, Mohamed Medhat Gaber. Mcua: Multi-level context and uncertainty aware dynamic deep ensemble for breast cancer histology image classification. IEEE Trans Biomed Eng, 2021
- Sai Chandra Kosaraju, Jie Hao, Hyun Min Koh, Mingon Kang. Deep-hipo: Multi-scale receptive field deep learning for histopathological image analysis. Methods, 2020. [PubMed]
- Olaf Ronneberger, Philipp Fischer, Thomas Brox. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2015
- Le Hou, Ayush Agarwal, Dimitris Samaras, Tahsin M. Kurc, Rajarsi R. Gupta, Joel H. Saltz. 2019
- Hoo-Chang Shin, Holger R. Roth, Mingchen Gao. Deep convolutional neural networks for computer-aided detection: Cnn architectures, dataset characteristics and transfer learning. IEEE Trans Med Imaging, 2016. [PubMed]
- Vu Quoc Dang, Simon Graham, Tahsin Kurc. Methods for segmentation and classification of digital microscopy tissue images. Front Bioeng Biotechnol, 2019. [PubMed]
- Kemeng Chen, Ning Zhang, Linda Powers, Janet Roveda. Spring Simulation Conference (SpringSim), 2019
- Tian Bai, Xu Jiayu, Fuyong Xing. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2020
- Xinpeng Xie, Jiawei Chen, Yuexiang Li, Linlin Shen, Kai Ma, Yefeng Zheng. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2020
- Amal Lahiani, Jacob Gildenblat, Irina Klaman, Nassir Navab, Eldad Klaiman. Generalizing multistain immunohistochemistry tissue segmentation using one-shot color deconvolution deep neural networks. arXiv preprint arXiv, 2018
- Gabriel Jiménez, Daniel Racoceanu. Deep learning for semantic segmentation versus classification in computational pathology: Application to mitosis analysis in breast cancer grading. Front Bioeng Biotechnol, 2019. [PubMed]
- Tasleem Kausar, Wang MingJiang, M. Adnan Ashraf, Adeeba Kausar. Smallmitosis: Small size mitotic cells detection in breast histopathology images. IEEE Access, 2020
- David Joon Ho, Narasimhan P. Agaram, Peter J. Schüffler. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2020
- Suzanne C. Wetstein, Allison M. Onken, Christina Luffman. Deep learning assessment of breast terminal duct lobular unit involution: Towards automated prediction of breast cancer risk. PloS One, 2020
- Hammad Qureshi, Olcay Sertel, Nasir Rajpoot, Roland Wilson, Metin Gurcan. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2008
- Rokshana Stephny Geread, Abishika Sivanandarajah, Emily Rita Brouwer. pinet–an automated proliferation index calculator framework for ki67 breast cancer images. Cancers, 2021
- D Schuhmacher, S Schörner, C Küpper. A framework for falsifiable explanations of machine learning models with an application in computational pathology. Medical Image Analysis, 2022 Nov 1. [PubMed]
- Jaime Gallego, Zaneta Swiderska-Chadaj, Tomasz Markiewicz, M. Michifumi Yamashita, Alejandra Gabaldon, Arkadiusz Gertych. A u-net based framework to quantify glomerulosclerosis in digitized pas and h&e stained human tissues. Comput Med Imaging Graph, 2021
- Wu Jianghua, Changling Liu, Xiaoqing Liu. Artificial intelligence-assisted system for precision diagnosis of pd-l1 expression in non-small cell lung cancer. Mod Pathol, 2021
- Pingjun Chen, Yun Liang, Xiaoshuang Shi, Lin Yang, Paul Gader. Automatic whole slide pathology image diagnosis framework via unit stochastic selection and attention fusion. Neurocomputing, 2021. [PubMed]
- Abdala Nour, Sherif Saad, Boubakeur Boufama. ACM Conference on Bioinformatics, Computational Biology, and Health Informatics, 2021
- Akram Bayat, Connor Anderson, Pratik Shah. 2021
- Mahendra Khened, Avinash Kori, Haran Rajkumar, Ganapathy Krishnamurthi, Balaji Srinivasan. A generalized deep learning framework for whole-slide image segmentation and analysis. Sci Rep, 2021. [PubMed]
- Peter Naylor, Marick Laé, Fabien Reyal, Thomas Walter. 2017
- Mihir Sahasrabudhe, Stergios Christodoulidis, Roberto Salgado. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2020
- Saad Nadeem, Travis Hollmann, Allen Tannenbaum. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2020
- Nanqing Dong, Michael Kampffmeyer, Xiaodan Liang, Zeya Wang, Wei Dai, Eric Xing. Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support, 2018
- Leander van Eekelen, Hans Pinckaers, Konnie M. Hebeda, Geert Litjens. 2020
- Gang Xing, Jianqin Lei, Xiayu Xu. The Fifth International Conference on Biological Information and Biomedical Engineering, 2021
- Mina Khoshdeli, Garrett Winkelmaier, Bahram Parvin. Fusion of encoder-decoder deep networks improves delineation of multiple nuclear phenotypes. BMC Bioinform, 2018
- Mohamed Abdel-Nasser, Adel Saleh, Domenec Puig. VISIGRAPP, 2020
- Shengcong Chen, Changxing Ding, Dacheng Tao. 2020
- Simon Graham, Hao Chen, Jevgenij Gamper. Mild-net: Minimal information loss dilated network for gland instance segmentation in colon histology images. Med Image Anal, 2019. [PubMed]
- Abhinav Agarwalla, Muhammad Shaban, Nasir M. Rajpoot. Representation-aggregation networks for segmentation of multi-gigapixel histology images. ArXiv, 2017
- Ziqiang Li, Rentuo Tao, Qianrun Wu, Bin Li. Da-refinenet: A dual input whole slide image segmentation algorithm based on attention. arXiv, 2019
- Pushpak Pati, Sonali Andani, Matthew Pediaditis. Medical Imaging: Digital Pathology, 2018
- Talha Qaiser, Yee-Wah Tsang, Daiki Taniyama. Fast and accurate tumor segmentation of histology images using persistent homology and deep convolutional features. Med Image Anal, 2019. [PubMed]
- Dev Kumar Das, Surajit Bose, Asok Kumar Maiti, Bhaskar Mitra, Gopeswar Mukherjee, Pranab Kumar Dutta. Automatic identification of clinically relevant regions from oral tissue histological images for oral squamous cell carcinoma diagnosis. Tissue Cell, 2018. [PubMed]
- J. Adam, Simon Graham Shephard, Saad Bashir. IEEE/CVF International Conference on Computer Vision, 2021
- Liuan Wang, Li Sun, Mingjie Zhang. ACM International Conference on Multimedia, 2021
- Mostafa Jahanifar, Neda Zamani Tajeddin, Navid Alemi Koohbanani, Nasir M. Rajpoot. IEEE/CVF International Conference on Computer Vision, 2021
- Hyeongsub Kim, Hongjoon Yoon, Nishant Thakur. Deep learning-based histopathological segmentation for whole slide images of colorectal cancer in a compressed domain. Sci Rep, 2021. [PubMed]
- G. Murtaza Dogar, Muhammad Moazam Fraz, Sajid Javed. 2021
- Maxime W. Lafarge, Josien P.W. Pluim, Koen A.J. Eppenhof, Mitko Veta. Learning domain-invariant representations of histological images. Front Med, 2019
- Haibo Wang, Angel Cruz Roa, Ajay N. Basavanhally. Mitosis detection in breast cancer pathology images by combining handcrafted and convolutional neural network features. J Med Imag, 2014
- Chao Li, Xinggang Wang, Wenyu Liu, Longin Jan Latecki. Deepmitosis: Mitosis detection via deep detection, verification and segmentation networks. Med Image Anal, 2018. [PubMed]
- Chao Li, Xinggang Wang, Wenyu Liu, Longin Jan Latecki, Bo Wang, Junzhou Huang. Weakly supervised mitosis detection in breast histopathology images using concentric loss. Med Image Anal, 2019. [PubMed]
- Meriem Sebai, Xinggang Wang, Tianjiang Wang. Maskmitosis: a deep learning framework for fully supervised, weakly supervised, and unsupervised mitosis detection in histopathology images. Med Biol Eng Comput, 2020. [PubMed]
- Saad Ullah Akram, Talha Qaiser, Simon Graham, Juho Kannala, Janne Heikkilä, Nasir Rajpoot. 2018
- Maxime W. Lafarge, Erik J. Bekkers, Josien P.W. Pluim, Remco Duits, Mitko Veta. Roto-translation equivariant convolutional networks: Application to histopathology image analysis. Med Image Anal, 2021
- Md Zahangir Alom, Theus Aspiras, Tarek M. Taha, Tj Bowen, Vijayan K. Asari. Mitosisnet: End-to-end mitotic cell detection by multi-task learning. IEEE Access, 2020
- Hansheng Li, Xin Han, Yuxin Kang. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2020
- Pushpak Pati, Antonio Foncubierta-Rodríguez, Orcun Goksel, Maria Gabrani. Reducing annotation effort in digital pathology: A co-representation learning framework for classification tasks. Med Image Anal, 2021
- Nicolas Brieu, Armin Meier, Ansh Kapil. Domain adaptation-based augmentation for weakly supervised nuclei detection. arXiv preprint arXiv, 2019
- J. Fuchs Thomas, Peter J. Wild, Holger Moch, Joachim M. Buhmann. 2008
- Wenyuan Li, Jiayun Li, Karthik V. Sarma. Path r-cnn for prostate cancer diagnosis and gleason grading of histological images. IEEE Trans Med Imaging, 2018. [PubMed]
- Muhammad Nasim Kashif, Shan E. Ahmed, Korsuk Sirinukunwattana Raza, Muhammmad Arif, Nasir Rajpoot. IEEE International Symposium on Biomedical Imaging, 2016
- Guillaume Jaume, Pushpak Pati, Valentin Anklin, Antonio Foncubierta, Maria Gabrani. MICCAI Workshop on Computational Pathology, 2021
- Sajid Javed, Arif Mahmood, Jorge Dias, Naoufel Werghi, Nasir Rajpoot. Spatially constrained context-aware hierarchical deep correlation filters for nucleus detection in histology images. Med Image Anal, 2021
- Linbo Wang, Hui Zhen, Xianyong Fang, Shaohua Wan, Weiping Ding, Yanwen Guo. A unified two-parallel-branch deep neural network for joint gland contour and segmentation learning. Futur Gener Comput Syst, 2019
- Bingzhe Wu, Shiwan Zhao, Guangyu Sun. IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2019
- Haichun Yang, Ruining Deng, Lu Yuzhe. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2020
- M. Gloria Bueno, Milagro Fernandez-Carrobles, Lucia Gonzalez-Lopez, Oscar Deniz. Glomerulosclerosis identification in whole slide images using semantic segmentation. Comput Methods Programs Biomed, 2020
- Lu Zixiao, Xu Siwen, Wei Shao. Deep-learning–based characterization of tumor-infiltrating lymphocytes in breast cancers from histopathology images and multiomics data. JCO Clin Cancer Inform, 2020. [PubMed]
- David Tellez, Diederik Höppener, Cornelis Verhoef. 2020
- Mohammed Alawad, Shang Gao, John X. Qiu. Automatic extraction of cancer registry reportable information from free-text pathology reports using multitask convolutional neural networks. J Am Med Inform Assoc, 2020. [PubMed]
- Zunlei Feng, Zhonghua Wang, Xinchao Wang. Proceedings of the IEEE/CVF International Conference on Computer Vision, 2021
- Ozan Sener, Vladlen Koltun. Advances in Neural Information Processing Systems, 2018
- Amelie Royer, Tijmen Blankevoort, Babak Ehteshami Bejnordi Bejnordi. Advances in Neural Information Processing Systems, 2023
- Jiquan Ngiam, Aditya Khosla, Mingyu Kim, Juhan Nam, Honglak Lee, Andrew Y. Ng. ICML, 2011
- Haoyang Mi, Trinity J. Bivalacqua, Max Kates. Predictive models of response to neoadjuvant chemotherapy in muscle-invasive bladder cancer using nuclear morphology and tissue architecture. Cell Rep Med, 2021
- Sebastian Foersch, Christina Glasner, Ann-Christin Woerl. Multistain deep learning for prediction of prognosis and therapy response in colorectal cancer. Nat Med, 2023. [PubMed]
- Zhi Huang, Wei Shao, Zhi Han. Artificial intelligence reveals features associated with breast cancer neoadjuvant chemotherapy responses from multi-stain histopathologic images. NPJ Precis Oncol, 2023. [PubMed]
- Chunyuan Li, Xinliang Zhu, Jiawen Yao, Junzhou Huang. In 2022 26th International Conference on Pattern Recognition (ICPR), 2022
- 401Yawen Wu, Michael Cheng, Shuo Huang, Zongxiang Pei, Yingli Zuo, Jianxin Liu, Kai Yang, Qi Zhu, Jie Zhang, Honghai Hong, Daoqiang Zhang, Kun Huang, Liang Cheng, and Wei Shao. Recent advances of deep learning for computational histopathology: principles and applications. Cancers (Basel), 14(5):1199, February 2022.
- 402Xinrui Huang, Zhaotong Li, Minghui Zhang, and Song Gao. Fusing hand-crafted and deep-learning features in a convolutional neural network model to identify prostate cancer in pathology images. Front Oncol, 12:994950, September 2022.
- Mobeen Ur Rehman, Suhail Akhtar, Muhammad Zakwan, Muhammad Habib Mahmood. Novel architecture with selected feature vector for effective classification of mitotic and non-mitotic cells in breast cancer histology images. Biomed Signal Process Control, 2022
- I. Onur Sigirci, Abdulkadir Albayrak, Gokhan Bilgin. Detection of mitotic cells in breast cancer histopathological images using deep versus handcrafted features. Multimed Tools Appl, 2021. [PubMed]
- Wei-Hung Weng, Yuannan Cai, Angela Lin, Fraser Tan, Po-Hsuan Cameron Chen. 2019
- Kevin M. Boehm, Pegah Khosravi, Rami Vanguri, Jianjiong Gao, Sohrab P. Shah. Harnessing multimodal data integration to advance precision oncology. Nat Rev Cancer, 2021. [PubMed]
- Wei Shao, Tongxin Wang, Liang Sun. Multi-task multi-modal learning for joint diagnosis and prognosis of human cancers. Med Image Anal, 2020
- Zhiqin Wang, Ruiqing Li, Minghui Wang, Ao Li. Gpdbn: deep bilinear network integrating both genomic data and pathological images for breast cancer prognosis prediction. Bioinformatics, 2021. [PubMed]
- Zhi Huang, Federico Bianchi, Mert Yuksekgonul, Thomas J. Montine, James Zou. A visual–language foundation model for pathology image analysis using medical twitter. Nat Med, 2023. [PubMed]
- Wisdom Oluchi Ikezogwo, Mehmet Saygin Seyfioglu, Fatemeh Ghezloo. 2023
- Qu Linhao, Xiaoyuan Luo, Kexue Fu, Manning Wang, Zhijian Song. Conference on Computer Vision and Pattern Recognition (CVPR), 2023
- Ming Y. Lu, Bowen Chen, Andrew Zhang. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), June 2023
- Yann LeCun, Yoshua Bengio, Geoffrey Hinton. Deep learning. Nature, 2015. [PubMed]
- AA Nahid, MA Mehrabi, Y Kong. Histopathological Breast Cancer Image Classification by Deep Neural Network Techniques Guided by Local Clustering. Biomed Research International, 2018 Mar 7
- Zizhao Zhang, Pingjun Chen, Mason McGough. Pathologist-level interpretable whole-slide cancer diagnosis with deep learning. Nat Mach Intell, 2019
- Hongdou Yao, Xuejie Zhang, Xiaobing Zhou, Shengyan Liu. Parallel structure deep neural network using cnn and rnn with an attention mechanism for breast cancer histology image classification. Cancers, 2019. [PubMed]
- Xu Bolei, Jingxin Liu, Xianxu Hou. IEEE International Symposium on Biomedical Imaging, 2019
- Patricia Raciti, Jillian Sue, Rodrigo Ceballos. Novel artificial intelligence system increases the detection of prostate cancer in whole slide images of core needle biopsies. Mod Pathol, 2020. [PubMed]
- Aïcha BenTaieb, Ghassan Hamarneh. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2018
- Yushan Zheng, Zhiguo Jiang, Haopeng Zhang, Fengying Xie, Jun Shi. Medical Image Computing and Computer Assisted Intervention – MICCAI, Volume 12265 of Lecture Notes in Computer Science, 2020
- Jing Qi, Girvan Burnside, Paul Charnley, Frans Coenen. Proceedings of the 13th International Joint Conference On Knowledge Discovery, Knowledge Engineering and Knowledge Management (KDIR), 2021
- Xu Bolei, Jingxin Liu, Xianxu Hou. Attention by selection: a deep selective attention approach to breast cancer classification. IEEE Trans Med Imaging, 2019. [PubMed]
- Ashish Vaswani, Noam Shazeer, Niki Parmar. Attention is all you need. Adv Neural Inform Process Syst, 2017
- Faisal Mahmood, Daniel Borders, Richard J. Chen. Deep adversarial training for multi-organ nuclei segmentation in histopathology images. IEEE Trans Med Imaging, 2019
- Adalberto Claudio Quiros, Roderick Murray-Smith, Ke Yuan. Pathologygan: learning deep representations of cancer tissue. J Mach Learn Biomed Imaging, 2021
- Srijay Deshpande, Fayyaz Minhas, Simon Graham, Nasir Rajpoot. Safron: stitching across the frontier network for generating colorectal cancer histology images. Med Image Anal, 2022
- 427Hyungjoo Cho, Sungbin Lim, Gunho Choi, and Hyunseok Min. Neural stain-style transfer learning using gan for histopathological images. arXiv preprint arXiv:1710.08543, 2017.
- Peter Leonard Schrammen, Narmin Ghaffari Laleh, Amelie Echle. Weakly supervised annotation-free cancer detection and prediction of genotype in routine histopathology. J Pathol, 2022. [PubMed]
- Rishi R. Rawat, Itzel Ortega, Preeyam Roy. Deep learned tissue “fingerprints” classify breast cancers by er/pr/her2 status from h&e images. Sci Rep, 2020. [PubMed]
- Amal Lahiani, Jacob Gildenblat, Irina Klaman, Shadi Albarqouni, Nassir Navab, Eldad Klaiman. European Congress on Digital Pathology, 2019
- Amal Lahiani, Nassir Navab, Shadi Albarqouni, Eldad Klaiman. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2019
- Thomas E. Tavolara, M. Khalid Khan Niazi, Vidya Arole, Wei Chen, Wendy Frankel, Metin N. Gurcan. A modular cgan classification framework: application to colorectal tumor detection. Sci Rep, 2019. [PubMed]
- Hu Bo, I. Ye Tang, Chao Chang Eric, Yubo Fan, Maode Lai, Xu. Yan. Unsupervised learning for cell-level visual representation in histopathology images with generative adversarial networks. IEEE J Biomed Health Inform, 2018. [PubMed]
- Zhaoyang Xu, Carlos Fernández Moro, Béla Bozóky, Qianni Zhang. 2019
- Shahira Abousamra, Rajarsi Gupta, Tahsin Kurc, Dimitris Samaras, Joel Saltz, Chao Chen. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2023
- Marco Aversa, Gabriel Nobis, Miriam Hägele. 2023
- Abhijeet Patil, Dipesh Tamboli, Swati Meena, Deepak Anand, Amit Sethi. IEEE International WIE Conference on Electrical and Computer Engineering (WIECON-ECE), 2019
- Caner Mercan, Selim Aksoy, Ezgi Mercan, Linda G. Shapiro, Donald L. Weaver, Joann G. Elmore. Multi-instance multi-label learning for multi-class classification of whole slide breast histopathology images. IEEE Trans Med Imaging, 2017. [PubMed]
- Shujun Wang, Yaxi Zhu, Yu Lequan. Rmdl: recalibrated multi-instance deep learning for whole slide gastric image classification. Med Image Anal, 2019
- Xu Yan, Tao Mo, Qiwei Feng, Peilin Zhong, I. Maode Lai, Eric, and Chao Chang. IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), 2014
- Xu Yan, Yeshu Li, Zhengyang Shen. Parallel multiple instance learning for extremely large histopathology image analysis. BMC Bioinforma, 2017
- Jiayun Li, Wenyuan Li, Arkadiusz Gertych, Beatrice S. Knudsen, William Speier, Corey W. Arnold. 2019
- Asfand Yaar, Amina Asif, Shan E. Ahmed Raza, Nasir Rajpoot, Fayyaz Minhas. IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops, 2023
- Marvin Lerousseau, Maria Vakalopoulou, Marion Classe. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2020
- Philip Chikontwe, Meejeong Kim, Soo Jeong Nam, Heounjeong Go, Sang Hyun Park. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2020
- Jiawen Yao, Xinliang Zhu, Jitendra Jonnagaddala, Nicholas Hawkins, Junzhou Huang. Whole slide images based cancer survival prediction using attention guided deep multiple instance learning networks. Med Image Anal, 2020
- Ming Y. Lu, Richard J. Chen, Dehan Kong. Federated learning for computational pathology on gigapixel whole slide images. Med Image Anal, 2022
- Ming Y. Lu, Tiffany Y. Chen, Drew F.K. Williamson. Ai-based pathology predicts origins for cancers of unknown primary. Nature, 2021. [PubMed]
- Robert Jewsbury, Abhir Bhalerao, Nasir M. Rajpoot. IEEE/CVF International Conference on Computer Vision, 2021
- Abtin Riasatian, Morteza Babaie, Danial Maleki. Fine-tuning and training of densenet for histopathology image representation using tcga diagnostic slides. Med Image Anal, 2021
- Anabik Pal, Zhiyun Xue, Kanan Desai. Deep multiple-instance learning for abnormal cell detection in cervical histopathology images. Comput Biol Med, 2021
- Christophe A.C. Freyre, Stephan Spiegel, Caroline Gubser Keller. Biomarker-based classification and localization of renal lesions using learned representations of histology—a machine learning approach to histopathology. Toxicol Pathol, 2021. [PubMed]
- Niccolò Marini, Sebastian Otálora, Francesco Ciompi. MICCAI Workshop on Computational Pathology, 2021
- Johannes Höhne, Jacob de Zoete, Arndt A. Schmitz, Tricia Bal, Emmanuelle di Tomaso, Matthias Lenga. MICCAI Workshop on Computational Pathology, 2021
- Deepak Anand, Kumar Yashashwi, Neeraj Kumar, Swapnil Rane, Peter H. Gann, Amit Sethi. Weakly supervised learning on unannotated h&e-stained slides predicts braf mutation in thyroid cancer with high accuracy. J Pathol, 2021. [PubMed]
- Zhuchen Shao, Hao Bian, Yang Chen. Transmil: transformer based correlated multiple instance learning for whole slide image classification. Adv Neural Inform Process Syst, 2021
- Kailu Li, Ziniu Qian, Yingnan Han. Weakly supervised histopathology image segmentation with self-attention. Med Image Anal, 2023
- Fei Li, Mingyu Wang, Bin Huang. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2023
- Ramin Nakhli, Allen Zhang, Ali Mirabadi. Proceedings of the IEEE/CVF International Conference on Computer Vision (ICCV), October 2023
- Syed Farhan Abbas, Trinh Thi Le Vuong, Kyungeun Kim, Boram Song, Jin Tae Kwak. Multi-cell type and multi-level graph aggregation network for cancer grading in pathology images. Med Image Anal, 2023
- Jaeung Lee, Keunho Byeon, Jin Tae Kwak. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2023
- Puria Azadi, Jonathan Suderman, Ramin Nakhli. Lecture Notes in Computer Science, 2023
- Ramin Nakhli, Puria Azadi Moghadam, Haoyang Mi. In 2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2023
- Gabriele Campanella, Vitor Werneck Krauss Silva, Thomas J. Fuchs. 2018
- Hongrun Zhang, Yanda Meng, Yitian Zhao. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2022
- Raia Hadsell, Sumit Chopra, Yann LeCun. IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR’06), 2006
- Florian Schroff, Dmitry Kalenichenko, James Philbin. IEEE Conference on Computer Vision and Pattern Recognition, 2015
- Manuel Tran, Sophia J. Wagner, Melanie Boxberg, Tingying Peng. Medical Image Computing and Computer Assisted Intervention – MICCAI 2022, 2022
- Jacob Carse, Frank Carey, Stephen McKenna. IEEE International Symposium on Biomedical Imaging, 2021
- Muhammad Dawood, Kim Branson, Nasir M. Rajpoot, Fayyaz Minhas. IEEE/CVF International Conference on Computer Vision, 2021
- Chao Feng, Chad Vanderbilt, Thomas Fuchs. Medical Imaging with Deep Learning, 2021
- Jiawei Yang, Hanbo Chen, Jiangpeng Yan, Xiaoyu Chen, Jianhua Yao. 2022
- Jiang Cheng, Xinhai Hou, Akhil Kondepudi. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), June 2023
- Jacob Gildenblat, Anil Yüce, Samaneh Abbasi-Sureshjani, Konstanty Korski. Proceedings of the International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI), October 2023
- Hritam Basak, Zhaozheng Yin. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), June 2023
- Okyaz Eminaga, Mahmoud Abbas, Christian Kunder. 2019
- Jie Hao, Sai Chandra Kosaraju, Nelson Zange Tsaku, Dae Hyun Song, Mingon Kang. Pacific Symposium on Biocomputing, 2019
- Ting-An Yen, Hung-Chun Hsu, Pushpak Pati, Maria Gabrani, Antonio Foncubierta-Rodríguez, Pau-Choo Chung. 2020
- Sheyang Tang, Mahdi S. Hosseini, Lina Chen. IEEE/CVF International Conference on Computer Vision, 2021
- Edgar Galván, Peter Mooney. Neuroevolution in deep neural networks: current trends and future challenges. IEEE Trans Artif Intell, 2021
- Yuqiao Liu, Yanan Sun, Bing Xue, Mengjie Zhang, Gary G. Yen, Kay Chen Tan. IEEE Transactions on Neural Networks and Learning Systems, 2021
- Prasanna Balaprakash, Romain Egele, Misha Salim. International Conference for High Performance Computing, Networking, Storage and Analysis, 2019
- David H. Wolpert, William G. Macready. No free lunch theorems for optimization. IEEE Trans Evol Comput, 1997
- Stavros P. Adam, Stamatios-Aggelos N. Alexandropoulos, Panos M. Pardalos, Michael N. Vrahatis. No free lunch theorem: a review. Approx Optim, 2019
- Narmin Ghaffari Laleh, Hannah Sophie Muti, Chiara Maria Lavinia Loeffler. Benchmarking weakly-supervised deep learning pipelines for whole slide classification in computational pathology. Med Image Anal, 2022
- Richard Chen, Yating Jing, Hunter Jackson. 2016
- Forrest N. Iandola, Song Han, Matthew W. Moskewicz, Khalid Ashraf, William J. Dally, Kurt Keutzer. 2016
- Laith Alzubaidi, Omran Al-Shamma, Mohammed A. Fadhel, Laith Farhan, Jinglan Zhang, Ye Duan. Optimizing the performance of breast cancer classification by employing the same domain transfer learning from hybrid deep convolutional neural network model. Electronics, 2020
- Mai Bui Huynh Thuy, Vinh Truong Hoang. International Conference on Computer Science, Applied Mathematics and Applications, 2019
- Amirhossein Kiani, Bora Uyumazturk, Pranav Rajpurkar. Impact of a deep learning assistant on the histopathologic classification of liver cancer. NPJ Digit Med, 2020. [PubMed]
- Nicolas Coudray, Paolo Santiago Ocampo, Theodore Sakellaropoulos. Classification and mutation prediction from non–small cell lung cancer histopathology images using deep learning. Nat Med, 2018. [PubMed]
- Stan Benjamens, Pranavsingh Dhunnoo, Bertalan Meskó. The state of artificial intelligence-based fda-approved medical devices and algorithms: an online database. NPJ Digit Med, 2020. [PubMed]
- Alec Radford, Jong Wook Kim, Chris Hallacy. International Conference on Machine Learning, 2021
- Xiaohua Zhai, Xiao Wang, Basil Mustafa. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2022
- Amanpreet Singh, Ronghang Hu, Vedanuj Goswami. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2022
- Michiel Bakker, Martin Chadwick, Hannah Sheahan. Advances in Neural Information Processing Systems, 2022
- Alexey Dosovitskiy, German Ros, Felipe Codevilla, Antonio Lopez, Vladlen Koltun. Conference on Robot Learning, 2017
- Steve Borkman, Adam Crespi, Saurav Dhakad. 2021
- Viktor Makoviychuk, Lukasz Wawrzyniak, Yunrong Guo. 2021
- Gregory Griffin, Alex Holub, Pietro Perona. 2007
- Alex Krizhevsky, Ilya Sutskever, Geoffrey E. Hinton. Imagenet classification with deep convolutional neural networks. Adv Neural Inform Process Syst, 2012
- 502Guansong Pang, Chunhua Shen, Longbing Cao, and Anton Van Den Hengel. Deep learning for anomaly detection: a review. ACM Comput Surv, 54(2), mar 2021.
- Mahdi S. Hosseini, Lyndon Chan, Weimin Huang. European Conference on Computer Vision, 2020
- Lu Yuchen, Xu. Peng. 2018
- Andre Esteva, Brett Kuprel, Roberto A. Novoa. Dermatologist-level classification of skin cancer with deep neural networks. Nature, 2017. [PubMed]
- Dimitris K. Iakovidis, Spiros V. Georgakopoulos, Michael Vasilakakis, Anastasios Koulaouzidis, Vassilis P. Plagianakos. Detecting and locating gastrointestinal anomalies using deep learning and iterative cluster unification. IEEE Trans Med Imaging, 2018. [PubMed]
- CAMELYON16 ISBI challenge on cancer metastasis detection in lymph node. 2016
- Qi Qi, Yanlong Li, Jitian Wang. Label-efficient breast cancer histopathological image classification. IEEE J Biomed Health Inform, 2018. [PubMed]
- Mathilde Caron, Hugo Touvron, Ishan Misra. Proceedings of the IEEE/CVF International Conference on Computer Vision, 2021
- Chen Sun, Abhinav Shrivastava, Saurabh Singh, Abhinav Gupta. Proceedings of the IEEE International Conference on Computer Vision, 2017
- Alhanoof Althnian, Duaa AlSaeed, Heyam Al-Baity. Impact of dataset size on classification performance: an empirical evaluation in the medical domain. Appl Sci, 2021
- Alireza Abdollahi, Hiva Saffar, Hana Saffar. Types and frequency of errors during different phases of testing at a clinical medical laboratory of a teaching hospital in tehran, iran. North Am J Med Sci, 2014
- Frederick A. Meier, Ruan C. Varney, Richard J. Zarbo. Study of amended reports to evaluate and improve surgical pathology processes. Adv Anat Pathol, 2011. [PubMed]
- Teresa P. Darcy, Samuel P. Barasch, Rhona J. Souers, Peter L. Perrotta. Test cancellation: a college of american pathologists q-probes study. Arch Pathol Lab Med, 2016. [PubMed]
- Raouf E. Nakhleh. A prelude to error reduction in anatomic pathology. Am J Clin Pathol, 2005. [PubMed]
- Raouf E. Nakhleh. Error reduction in surgical pathology. Arch Pathol Lab Med, 2006. [PubMed]
- Raouf E. Nakhleh. Patient safety and error reduction in surgical pathology. Arch Pathol Lab Med, 2008. [PubMed]
- Raouf E. Nakhleh, Vania Nosé, Carol Colasacco. Interpretive diagnostic error reduction in surgical pathology and cytology: guideline from the college of american pathologists pathology and laboratory quality center and the association of directors of anatomic and surgical pathology. Arch Pathol Lab Med, 2016. [PubMed]
- Raouf E. Nakhleh. Role of informatics in patient safety and quality assurance. Surg Pathol Clin, 2015. [PubMed]
- Anobel Y. Odisho, Briton Park, Nicholas Altieri. Natural language processing systems for pathology parsing in limited data environments with uncertainty estimation. JAMIA Open, 2020. [PubMed]
- Pilar López-Úbeda, Teodoro Martín-Noguerol, José Aneiros-Fernández, Antonio Luna. Natural language processing in pathology: Current trends and future insights. Am J Pathol, 2022. [PubMed]
- Yoojoong Kim, Jeong Hyeon Lee, Sunho Choi. Validation of deep learning natural language processing algorithm for keyword extraction from pathology reports in electronic health records. Sci Rep, 2020. [PubMed]
- John X. Qiu, Hong-Jun Yoon, Paul A. Fearn, Georgia D. Tourassi. Deep learning for automated extraction of primary sites from cancer pathology reports. IEEE J Biomed Health Inform, 2017. [PubMed]
- Amir R. Zamir, Alexander Sax, William Shen, Leonidas J. Guibas, Jitendra Malik, Silvio Savarese. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2018
- Yu Zhang, Qiang Yang. A survey on multi-task learning. IEEE Trans Knowl Data Eng, 2021
- Keith Bonawitz, Hubert Eichner, Wolfgang Grieskamp. Proceedings of Machine Learning and Systems, 2019
- H. Peter Kairouz, Brendan McMahan, Brendan Avent. Advances and open problems in federated learning. Found Trends Mach Learn, 2021
- Bastiaan S. Veeling, Jasper Linmans, Jim Winkens, Taco Cohen, Max Welling. Medical Image Computing and Computer Assisted Intervention – MICCAI, volume 11071 of Lecture Notes in Computer Science, 2018
- Peter J. Schüffler, D. Luke Geneslaw, Vijay K. Yarlagadda. Integrated digital pathology at scale: a solution for clinical diagnostics and cancer research at a large academic medical center. J Am Med Inform Assoc, 2021. [PubMed]
- Matthew G. Hanna, Victor E. Reuter, Meera R. Hameed. Whole slide imaging equivalency and efficiency study: experience at a large academic center. Mod Pathol, 2019. [PubMed]
- Matthew G. Hanna, Victor E. Reuter, Jennifer Samboy. Implementation of digital pathology offers clinical and operational increase in efficiency and cost savings. Arch Pathol Lab Med, 2019. [PubMed]
- Jonhan Ho, Stefan M. Ahlers, Curtis Stratman. Can digital pathology result in cost savings? A financial projection for digital pathology implementation at a large integrated health care organization. J Pathol Inform, 2014. [PubMed]
- Rudenko Ekaterina Evgenievna, Demura Tatiana Alexandrovna, Vekhova Ksenia Andreevna. Analysis of the three-year work of a digital pathomorphological laboratory built from the ground. J Pathol Inform, 2022
- Shidan Wang, Tao Wang, Lin Yang. Convpath: a software tool for lung adenocarcinoma digital pathological image analysis aided by a convolutional neural network. EBioMedicine, 2019. [PubMed]
- Mike Isaacs, Jochen K. Lennerz, Stacey Yates, Walter Clermont, Joan Rossi, John D. Pfeifer. Implementation of whole slide imaging in surgical pathology: a value added approach. J Pathol Inform, 2011. [PubMed]
- Guy Pare, Julien Meyer, Marie-Claude Trudel, Bernard Tetu. Impacts of a large decentralized telepathology network in canada. Telemed e-Health, 2016
- Sten Thorstenson, Jesper Molin, Claes Lundström. Implementation of large-scale routine diagnostics using whole slide imaging in sweden: digital pathology experiences 2006-2013. J Pathol Inform, 2014. [PubMed]
- Chee Leong Cheng, Rafay Azhar, Shi Hui Adeline Sng. Enabling digital pathology in the diagnostic setting: navigating through the implementation journey in an academic medical centre. J Clin Pathol, 2016. [PubMed]
- Nikolas Stathonikos, Mitko Veta, André Huisman, Paul J. van Diest. Going fully digital: perspective of a dutch academic pathology lab. J Pathol Inform, 2013. [PubMed]
- Liron Pantanowitz, John H. Sinard, Walter H. Henricks. Validating whole slide imaging for diagnostic purposes in pathology: guideline from the college of american pathologists pathology and laboratory quality center. Arch Pathol Lab Med, 2013. [PubMed]
- Esther Abels, Liron Pantanowitz. Current state of the regulatory trajectory for whole slide imaging devices in the usa. J Pathol Inform, 2017. [PubMed]
- Anil V. Parwani, Lewis Hassell, Eric Glassy, Liron Pantanowitz. Regulatory barriers surrounding the use of whole slide imaging in the united states of america. J Pathol Inform, 2014
- Andrew Janowczyk, Anant Madabhushi. Deep learning for digital pathology image analysis: a comprehensive tutorial with selected use cases. J Pathol Inform, 2016
- Sivaramakrishnan Sankarapandian, Saul Kohn, Vaughn Spurrier. Proceedings of the IEEE/CVF International Conference on Computer Vision, 2021
- Saeed Alshieban, Khaled Al-Surimi. Reducing turnaround time of surgical pathology reports in pathology and laboratory medicine departments. BMJ Open Qual, 2015
- Raouf E. Nakhleh, Patrick L. Fitzgibbons. 2005
- David M. Metter, Terence J. Colgan, Stanley T. Leung, Charles F. Timmons, Jason Y. Park. Trends in the us and canadian pathologist workforces from 2007 to 2017. JAMA Netw Open, 2019. [PubMed]
- Wouter Bulten, Kimmo Kartasalo, Po-Hsuan Cameron Chen. Artificial intelligence for diagnosis and gleason grading of prostate cancer: the panda challenge. Nat Med, 2022. [PubMed]
- Martyn Peck, David Moffat, Bruce Latham, Tony Badrick. Review of diagnostic error in anatomical pathology and the role and value of second opinions in error prevention. J Clin Pathol, 2018. [PubMed]
- John H.F. Smith. Cytology, liquid-based cytology and automation. Best Pract Res Clin Obstet Gynaecol, 2011. [PubMed]
- Erin Brender, Alison Burke, Richard M. Glass. Frozen section biopsy. Jama, 2005. [PubMed]
- Sudha Ayyagari, Anusha Potnuru, Sk Aamer Saleem, Pavani Marapaka. Analysis of frozen section compared to permanent section: a 2 year study in a single teritiary care hospital. J Pathol Nepal, 2021
- Stephan W. Jahn, Markus Plass, Farid Moinfar. Digital pathology: advantages, limitations and emerging perspective. J Clin Med, 2020. [PubMed]
- Chamidu Atupelage, Hiroshi Nagahashi, Fumikazu Kimura. International Conference on Advances in ICT for Emerging Regions (ICTer), 2013
- Peter Bankhead, Maurice B. Loughrey, José A. Fernández. Qupath: open source software for digital pathology image analysis. Sci Rep, 2017. [PubMed]
- S. Rathore, M. Iftikhar, M. Nasrallah, M. Gurcan, N. Rajpoot, Z. Mourelatos. Tmod-35. prediction of overall survival, and molecular markers in gliomas via analysis of digital pathology images using deep learning. Neuro-Oncology, 2019
- Wei Wang, John A. Ozolek, Gustavo K. Rohde. Detection and classification of thyroid follicular lesions based on nuclear structure from histopathology images. Cytometry A, 2010. [PubMed]
- Charlotte Syrykh, Arnaud Abreu, Nadia Amara. Accurate diagnosis of lymphoma on whole-slide histopathology images using deep learning. NPJ Digit Med, 2020. [PubMed]
- Yalu Cheng, Pengchong Qiao, Hongliang He, Guoli Song, Jie Chen. ACM Multimedia Asia, 2021
- Blanca Maria Priego-Torres, Daniel Sanchez-Morillo, Miguel Angel Fernandez-Granero, Marcial Garcia-Rojo. Automatic segmentation of whole-slide h&e stained breast histopathology images using a deep convolutional neural network architecture. Expert Syst Appl, 2020
- Christophe Avenel, Anna Tolf, Anca Dragomir, Ingrid B. Carlbom. Glandular segmentation of prostate cancer: an illustration of how the choice of histopathological stain is one key to success for computational pathology. Front Bioeng Biotechnol, 2019. [PubMed]
- Richard J. Chen, Ming Y. Lu, Jingwen Wang. Pathomic fusion: an integrated framework for fusing histopathology and genomic features for cancer diagnosis and prognosis. IEEE Trans Med Imaging, 2020
- Goce Ristanoski, Jon Emery, Javiera Martinez Gutierrez, Damien McCarthy, Uwe Aickelin. Australasian Computer Science Week Multiconference, 2021
- Jia-Ren Chang, Ching-Yi Lee, Chi-Chung Chen, Joachim Reischl, Talha Qaiser, Chao-Yuan Yeh. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2021
- 565National Cancer Institute. Cancer stat facts: Common cancer sites. Available at: https://seer.cancer.gov/statfacts/html/common.html. (accessed Feb 12, 2023).
- Stacey A. Kenfield, Esther K. Wei, Meir J. Stampfer, Bernard A. Rosner, Graham A. Colditz. Comparison of aspects of smoking among the four histological types of lung cancer. Tob Control, 2008. [PubMed]
- 567American Cancer Society. Survival rates for breast cancer. Available at: https://www.cancer.org/cancer/breast-cancer/understanding-a-breast-cancer-diagnosis/breast-cancer-survival-rates.html. (accessed Feb 9, 2023).
- 568Cancer Center. Bladder cancer types. Available at: https://www.cancercenter.com/cancer-types/bladder-cancer/types. (accessed Feb 10, 2023).
- 569American Cancer Society. Survival rates for bladder cancer. m: https://www.cancer.org/cancer/bladder-cancer/detection-diagnosis-staging/survival-rates.html. (accessed Feb 10, 2023).
- B.K. Andreassen, B. Aagnes, R. Gislefoss, M. Andreassen, R. Wahlqvist. Incidence and survival of urothelial carcinoma of the urinary bladder in norway 1981-2014. BMC Cancer, 2016
- 571American Cancer Society. Liver cancer survival rates. Available at: https://www.cancer.org/cancer/liver-cancer/detection-diagnosis-staging/survival-rates.html. (accessed Feb 10, 2023).
- Zachary D. Goodman. Neoplasms of the liver. Mod Pathol, 2007. [PubMed]
- L.A. Gloeckler Ries, J.L. Young, G.E. Keel. 2007
- 574National Cancer Institute. Cancer stat facts: Ovarian cancer. Available at: https://seer.cancer.gov/statfacts/html/ovary.html. (accessed Feb 14, 2023).
- American Cancer Society. Survival Rates for Kidney Cancer
- Donald P. Bottaro, W. Marston, Linehan. Multifocal renal cancer: genetic basis and its medical relevance. Clin Cancer Res, 2005. [PubMed]
- Victor Srougi, Raphael B. Kato, Fernanda A. Salvatore, Pedro P.M. Ayres, F. Marcos, Dall’Oglio, and Miguel Srougi. Incidence of benign lesions according to tumor size in solid renal masses. Int Braz J Urol, 2009. [PubMed]
- 578American Cancer Society. Survival rates for colorectal cancer. Available at: https://www.cancer.org/cancer/colon-rectal-cancer/detection-diagnosis-staging/survival-rates.html. (accessed Feb 9, 2023).
- Andrea Remo, Matteo Fassan, Alessandro Vanoli. Morphology and molecular features of rare colorectal carcinoma histotypes. Cancers, 2019. [PubMed]
- 580American Cancer Society. Survival rates for prostate cancer. Available at: https://www.cancer.org/cancer/prostate-cancer/detection-diagnosis-staging/survival-rates.html. (accessed Feb 8, 2023).
- Abul K. Abbas, Jon C. Aster, Vinay Kumar. 2010
- 582American Cancer Society. Lung cancer survival rates. Available at: https://www.cancer.org/cancer/lung-cancer/detection-diagnosis-staging/survival-rates.html. (accessed Feb 11, 2023).
- 583World Health Organization. Cancer. Available at: https://www.who.int/news-room/fact-sheets/detail/cancer. (accessed Feb 13, 2023).
- David Clunie, Dan Hosseinzadeh, Mikael Wintell. Digital imaging and communications in medicine whole slide imaging connectathon at digital pathology association pathology visions 2017. J Pathol Inform, 2018
- Kamyar Nazeri, Azad Aminpour, Mehran Ebrahimi. International Conference Image Analysis and Recognition, 2018
- Turki Turki, Anmar Al-Sharif, Yh. Taguchi. 2021
- Zihan Xiong, Yixuan Zheng, Jiayue Qiu. International Conference on Bioinformatics Research and Applications, 2021
- 588Xiangyu Zhang, Xinyu Zhou, Mengxiao Lin, and Jian Sun. Shufflenet: An extremely efficient convolutional neural network for mobile devices. In 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition, pages 6848–6856, 2018.
- Metin N. Gurcan, Anant Madabhushi, Nasir Rajpoot. International Conference on Pattern Recognition, 2010
- Liron Pantanowitz, Walter H. Henricks, Bruce A. Beckwith. Medical laboratory informatics. Clin Lab Med, 2007. [PubMed]
- Priya Lakshmi Narayanan, Shan E. Ahmed, Allison H. Raza. Unmasking the tissue microecology of ductal carcinoma in situ with deep learning. BioRxiv, 2019
- Zhengfeng Lai, Chao Wang, Luca Cerny Oliveira, Brittany N. Dugger, Sen-Ching Cheung, Chen-Nee Chuah. IEEE/CVF International Conference on Computer Vision, 2021
- Mousumi Roy, Jun Kong, Satyananda Kashyap. Convolutional autoencoder based model histocae for segmentation of viable tumor regions in liver whole-slide images. Sci Rep, 2021. [PubMed]
- Y.Q. Jiang, J.H. Xiong, H.Y. Li. Recognizing basal cell carcinoma on smartphone-captured digital histopathology images with a deep neural network. Br J Dermatol, 2020. [PubMed]
- Shidan Wang, Donghan M. Yang, Ruichen Rong, Xiaowei Zhan, Guanghua Xiao. Pathology image analysis using segmentation deep learning algorithms. Am J Pathol, 2019. [PubMed]
- Hiroki Tokunaga, Yuki Teramoto, Akihiko Yoshizawa, Ryoma Bise. IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2019
- Hye Yoon Chang, Chan Kwon Jung, Junwoo Isaac Woo. Artificial intelligence in pathology. J Pathol Transl Med, 2019. [PubMed]
- Tiange Xiang, Song Yang, Chaoyi Zhang. Dsnet: A dual-stream framework for weakly-supervised gigapixel pathology image analysis. IEEE Transactions on Medical ImagingarXiv, 2022
- 599Chetan L Srinidhi and Anne L Martel. Improving self-supervised learning with hardness-aware dynamic curriculum learning: An application to digital pathology. In IEEE/CVF International Conference on Computer Vision, pages 562–571, 2021.
- Syed Hamad Shirazi, Saeeda Naz, Muhammad Imran Razzak, Arif Iqbal Umar, Ahmad Zaib. Soft Computing Based Medical Image Analysis, 2018
- James A. Diao, Jason K. Wang, Wan Fung Chui. Human-interpretable image features derived from densely mapped cancer pathology slides predict diverse molecular phenotypes. Nat Commun, 2021. [PubMed]
- Andrew H. Beck, Ankur R. Sangoi, Samuel Leung. Systematic analysis of breast cancer morphology uncovers stromal features associated with survival. Sci Transl Med, 2011
- Prathamesh M. Kulkarni, Eric J. Robinson, Jaya Sarin Pradhan. Deep learning based on standard h&e images of primary melanoma tumors identifies patients at risk for visceral recurrence and death. Clin Cancer Res, 2020. [PubMed]
- Peizhen Xie, Ke Zuo, Yu Zhang, Fangfang Li, Mingzhu Yin, Kai Lu. Interpretable classification from skin cancer histology slides using deep learning: A retrospective multicenter study. arXiv, 2019
- Steven N. Hart, William Flotte, Andrew P. Norgan. Classification of melanocytic lesions in selected and whole-slide images via convolutional neural networks. J Pathol Inform, 2019
- Angel Alfonso Cruz-Roa, John Edison Arevalo Ovalle, Anant Madabhushi, Fabio Augusto González Osorio. 2013
- Joshua J. Levy, Christopher R. Jackson, Aravindhan Sriharan, Brock C. Christensen, Louis J. Vaickus. Preliminary evaluation of the utility of deep generative histopathology image translation at a mid-sized NCI cancer center. bioRxiv, 2020
- Bo Tang, Ao Li, Bin Li, Minghui Wang. Capsurv: capsule network for survival analysis with whole slide pathological images. IEEE Access, 2019
- Justin Kirby. MICCAI 2014 Grand Challenges
- Petteri Teikari, Marc Santos, Charissa Poon, Kullervo Hynynen. Deep learning convolutional networks for multiphoton microscopy vasculature segmentation. arXiv, 2016
- Benjamin Liechty, Xu Zhuoran, Zhilu Zhang. Machine learning can aid in prediction of idh mutation from h&e-stained histology slides in infiltrating gliomas. Sci Rep, 2022. [PubMed]
- Gloria Bueno, Lucia Gonzalez-Lopez, Marcial Garcia-Rojo, Arvydas Laurinavicius, Oscar Deniz. Data for glomeruli characterization in histopathological images. Data Brief, 2020
- Michael Gadermayr, Ann-Kathrin Dombrowski, Barbara Mara Klinkhammer, Peter Boor, Dorit Merhof. Cnn cascades for segmenting sparse objects in gigapixel whole slide images. Comput Med Imaging Graph, 2019. [PubMed]
- Gabriel Tjio, Xulei Yang, Jia Mei Hong. Accurate tumor tissue region detection with accelerated deep convolutional neural networks. arXiv, 2020
- Pietro Antonio Cicalese, Aryan Mobiny, Pengyu Yuan, Jan Becker, Chandra Mohan, Hien Van Nguyen. Medical Image Computing and Computer Assisted Intervention – MICCAI, Sep 2020
- Richard J. Chen, Ming Y. Lu, Tiffany Y. Chen, Drew F.K. Williamson, Faisal Mahmood. Synthetic data in machine learning for medicine and healthcare. Nat Biomed Eng, 2021. [PubMed]
- Jiří Borovec, Jan Kybic, Ignacio Arganda-Carreras. Anhir: Automatic non-rigid histological image registration challenge. IEEE Trans Med Imaging, 2020. [PubMed]
- Fei Wu, Pei Liu, Bo Fu, Feng Ye. 2022 14th International Conference on Machine Learning and Computing (ICMLC), 2022
- Brendon Lutnick, David Manthey, Jan U. Becker. A user-friendly tool for cloud-based whole slide image segmentation with examples from renal histopathology. Commun Med, 2022. [PubMed]
- CAMELYON17. 2017
- Yongxiang Huang and Albert Chi-shing Chung. Computational Pathology and Ophthalmic Medical Image Analysis, 2018
- MITOS-ATYPIA-14 Grand Challenge
- Ludovic Roux, Daniel Racoceanu, Nicolas Loménie. Mitosis detection in breast cancer histological images an icpr 2012 contest. J Pathol Inform, 2013
- Gabriele Campanella, Matthew G. Hanna, Edi Brogi, Thomas J. Fuchs. Breast metastases to axillary lymph nodes. Cancer Imaging Arch, 2019
- Kenneth Clark, Bruce Vendt, Kirk Smith. The cancer imaging archive (tcia): maintaining and operating a public information repository. J Digit Imaging, 2013. [PubMed]
- Kosmas Dimitropoulos, Panagiotis Barmpoutis, Christina Zioga, Athanasios Kamas, Kalliopi Patsiaoura, Nikos Grammalidis. Grading of invasive breast carcinoma through grassmannian vlad encoding. PloS One, 2017
- C. Zioga, A. Kamas, K. Patsiaoura, K. Dimitropoulos, P. Barmpoutis, N. Grammalidis. July 2017
- Brady Kieffer, Morteza Babaie, Shivam Kalra, Hamid R. Tizhoosh. International Conference on Image Processing Theory, Tools and Applications (IPTA), 2017
- Puspanjali Mohapatra, Baldev Panda, Samikshya Swain. Enhancing histopathological breast cancer image classification using deep learning. Int J Innov Technol Explor Eng, 2019
- Andrew Janowczyk, Scott Doyle, Hannah Gilmore, Anant Madabhushi. A resolution adaptive deep hierarchical (radhical) learning scheme applied to nuclear segmentation of digital pathology images. Comput Methods Biomech Biomed Eng Imaging Vis, 2018. [PubMed]
- Angel Cruz-Roa, Hannah Gilmore, Ajay Basavanhally. Accurate and reproducible invasive breast cancer detection in whole-slide images: a deep learning approach for quantifying tumor extent. Sci Rep, 2017. [PubMed]
- Mustafa I. Jaber, Bing Song, Clive Taylor. A deep learning image-based intrinsic molecular subtype classifier of breast tumors reveals tumor heterogeneity that may affect survival. Breast Cancer Res, 2020
- Alberto Corvo, Humberto Simon Garcia Caballero, Michel A. Westenberg, Marc A. van Driel, Jarke van Wijk. 2020
- Mitko Veta, Paul J. Van Diest, Stefan M. Willems. Assessment of algorithms for mitosis detection in breast cancer histopathology images. Med Image Anal, 2015. [PubMed]
- Babak Ehteshami Bejnordi, Guido Zuidhof, Maschenka Balkenhol. Context-aware stacked convolutional neural networks for classification of breast carcinomas in whole-slide histopathology images. J Med Imaging, 2017
- Babak Ehteshami Bejnordi, Maeve Mullooly, Ruth M. Pfeiffer. Using deep convolutional neural networks to identify and classify tumor-associated stroma in diagnostic breast biopsies. Mod Pathol, 2018. [PubMed]
- Mohammad Peikari, Mehrdad J. Gangeh, Judit Zubovits, Gina Clarke, Anne L. Martel. Triaging diagnostically relevant regions from pathology whole slides of breast cancer: A texture based approach. IEEE Trans Med Imaging, 2015. [PubMed]
- Guillaume Jaume, Pushpak Pati, Antonio Foncubierta-Rodríguez. ICML, 2020
- Guillaume Jaume, Pushpak Pati, Behzad Bozorgtabar. IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2021
- N Brancati, AM Anniciello, P Pati. A dataset for breast carcinoma subtyping in h&e histology images. Database, 2022 Jan 1. [PubMed]
- Andrew Lagree, Audrey Shiner, Marie Angeli Alera. Assessment of digital pathology imaging biomarkers associated with breast cancer histologic grade. Curr Oncol, 2021. [PubMed]
- Mohamed Amgad, Habiba Elfandy, Hagar Hussein. Structured crowdsourcing enables convolutional segmentation of histology images. Bioinformatics, 2019. [PubMed]
- Mathias Uhlen, Cheng Zhang, Sunjae Lee. A pathology atlas of the human cancer transcriptome. Science, 2017
- Ponkrshnan Thiagarajan, Pushkar Khairnar, Susanta Ghosh. Explanation and use of uncertainty quantified by bayesian neural network classifiers for breast histopathology images. IEEE Trans Med Imaging, 2021
- Zabit Hameed, Begonya Garcia-Zapirain, José Javier Aguirre, Mario Arturo Isaza-Ruget. Multiclass classification of breast cancer histopathology images using multilevel features of deep convolutional neural network. Sci Rep, 2022. [PubMed]
- Pinky A. Bautista, Yukako Yagi. Staining correction in digital pathology by utilizing a dye amount table. J Digit Imaging, 2015. [PubMed]
- B. Albertina, M. Watson, C. Holback. Radiology data from the cancer genome atlas lung adenocarcinoma [tcga-luad] collection. Cancer Imaging Arch, 2016
- National Lung Screening Trial Research Team. The national lung screening trial: overview and study design. Radiology, 2011. [PubMed]
- The Lung Cancer SPORE. 2023
- Lu Cheng, Can Koyuncu, German Corredor. Feature-driven local cell graph (FLocK): New computational pathology-based descriptors for prognosis of lung cancer and hpv status of oropharyngeal cancers. Med Image Anal, 2021
- Anne Laure Le Page, Elise Ballot, Caroline Truntzer. Using a convolutional neural network for classification of squamous and non-squamous non-small cell lung cancer based on diagnostic histopathology hes images. Sci Rep, 2021. [PubMed]
- Zaneta Swiderska-Chadaj, Francesco Ciompi. Lyon19- lymphocyte detection test set (version v1) [data set]. 2019
- Zaneta Swiderska-Chadaj, Hans Pinckaers, Mart van Rijthoven. Learning to detect lymphocytes in immunohistochemistry with deep learning. Med Image Anal, 2019
- Nadia Brancati, Giuseppe De Pietro, Maria Frucci, Daniel Riccio. A deep learning approach for breast invasive ductal carcinoma detection and lymphoma multi-classification in histological images. IEEE Access, 2019
- Jeppe Thagaard, Søren Hauberg, Bert van der Vegt, Thomas Ebstrup, Johan D. Hansen, Anders B. Dahl. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2020
- Muhammad Shaban, Syed Ali Khurram, Muhammad Moazam Fraz. A novel digital score for abundance of tumour infiltrating lymphocytes predicts disease free survival in oral squamous cell carcinoma. Sci Rep, 2019. [PubMed]
- Wouter Bulten, Péter Bándi, Jeffrey Hoven. Epithelium segmentation using deep learning in h&e-stained prostate specimens with immunohistochemistry as reference standard. Sci Rep, 2019. [PubMed]
- Martin Köbel, Steve E. Kalloger, Patricia M. Baker. Diagnosis of ovarian carcinoma cell type is highly reproducible: A transcanadian study. Am J Surg Pathol, 2010. [PubMed]
- Yuri Tolkach, Tilmann Dohmgörgen, Marieta Toma, Glen Kristiansen. High-accuracy prostate cancer pathology using deep learning. Nat Mach Intell, 2020
- Eirini Arvaniti, Kim S. Fricker, Michael Moret. Automated gleason grading of prostate cancer tissue microarrays via deep learning. Sci Rep, 2018. [PubMed]
- Eirini Arvaniti, Manfred Claassen. Coupling weak and strong supervision for classification of prostate cancer histopathology images. ArXiv, 2018
- Jian Ren, Ilker Hacihaliloglu, Eric A. Singer, David J. Foran, Xin Qi. Unsupervised domain adaptation for classification of histopathology whole-slide images. Front Bioeng Biotechnol, 2019. [PubMed]
- Davood Karimi, Guy Nir, Ladan Fazli, Peter C. Black, Larry Goldenberg, Septimiu E. Salcudean. Deep learning-based gleason grading of prostate cancer from histopathology images—role of multiscale decision aggregation and data augmentation. IEEE J Biomed Health Inform, 2019. [PubMed]
- Chaoyang Yan, Kazuaki Nakane, Xiangxue Wang. Automated gleason grading on prostate biopsy slides by statistical representations of homology profile. Comput Methods Programs Biomed, 2020
- Scott Doyle, James Monaco, Michael Feldman, John Tomaszewski, Anant Madabhushi. An active learning based classification strategy for the minority class problem: application to histopathology annotation. BMC Bioinforma, 2011
- Scott Doyle, Michael D. Feldman, Natalie Shih, John Tomaszewski, Anant Madabhushi. Cascaded discrimination of normal, abnormal, and confounder classes in histopathology: Gleason grading of prostate cancer. BMC Bioinforma, 2012
- Kunal Nagpal, Davis Foote, Yun Liu. Development and validation of a deep learning algorithm for improving gleason scoring of prostate cancer. NPJ Digit Med, 2019. [PubMed]
- Adrian B. Levine, Jason Peng, David Farnell. Synthesis of diagnostic quality cancer pathology images by generative adversarial networks. J Pathol, 2020. [PubMed]
- Birgid Schömig-Markiefka, Alexey Pryalukhin, Wolfgang Hulla. Quality control stress test for deep learning-based diagnostic model in digital pathology. Mod Pathol, 2021. [PubMed]
- Julio Silva-Rodríguez, Adrián Colomer, María A. Sales, Rafael Molina, Valery Naranjo. Going deeper through the gleason scoring scale: An automatic end-to-end system for histology prostate grading and cribriform pattern detection. Comput Methods Programs Biomed, 2020
- Hossein Farahani, Jeffrey Boschman, David Farnell. Deep learning-based histotype diagnosis of ovarian carcinoma whole-slide pathology images. Mod Pathol, 2022. [PubMed]
- Wouter Bulten, Geert Litjens, Hans Pinckaers. March 2020
- Jakob Nikolas Kather, F.G. Zöllner, F. Bianconi. Collection of textures in colorectal cancer histology. Zenodo, 2016
- Korsuk Sirinukunwattana, Josien P.W. Pluim, Hao Chen. Gland segmentation in colon histology images: The glas challenge contest. Med Image Anal, 2017. [PubMed]
- Muhammad Shaban, Ruqayya Awan, Muhammad Moazam Fraz. Context-aware convolutional neural network for grading of colorectal cancer histology images. IEEE Trans Med Imaging, 2020. [PubMed]
- Xu Jun, Xiaofei Luo, Guanhao Wang, Hannah Gilmore, Anant Madabhushi. A deep convolutional neural network for segmenting and classifying epithelial and stromal regions in histopathological images. Neurocomputing, 2016. [PubMed]
- University of Leeds. Welcome to the university of leeds virtual pathology project website. 2023
- Francesco Ponzio, Giacomo Deodato, Enrico Macii, Santa Di Cataldo, Elisa Ficarra. IEEE International Symposium on Biomedical Imaging, 2020
- Francesco Ciompi, Oscar Geessink, Babak Ehteshami Bejnordi. IEEE International Symposium on Biomedical Imaging, 2017
- Cigdem Gunduz-Demir, Melih Kandemir, Akif Burak Tosun, Cenk Sokmensuer. Automatic segmentation of colon glands using object-graphs. Med Image Anal, 2010. [PubMed]
- Jonas Kloeckner, Tatiana K. Sansonowicz, Áttila L. Rodrigues, Tatiana W.N. Nunes. Multi-categorical classification using deep learning applied to the diagnosis of gastric cancer. J Bras Patol Med Lab, 2020
- Huu-Giao Nguyen, Annika Blank, Alessandro Lugli, Inti Zlobec. IEEE International Symposium on Biomedical Imaging, 2020
- Rasoul Sali, Lubaina Ehsan, Kamran Kowsari. IEEE International Conference on Bioinformatics and Biomedicine (BIBM), 2019
- Rasoul Sali, Sodiq Adewole, Lubaina Ehsan. IEEE International Conference on Healthcare Informatics (ICHI), 2020
- Bin Kong, Shanhui Sun, Xin Wang, Qi Song, Shaoting Zhang. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2018
- Zhigang Song, Shuangmei Zou, Weixun Zhou. Clinically applicable histopathological diagnosis system for gastric cancer detection using deep learning. Nat Commun, 2020. [PubMed]
- Alexander Ian Wright, Catriona Marie Dunn, Michael Hale, Gordon Hutchins, Darren Treanor. The effect of quality control on accuracy of digital pathology image analysis. IEEE J Biomed Health Inform, 2020
- Mohsin Bilal, Shan E. Ahmed, Ayesha Azam Raza. Development and validation of a weakly supervised deep learning framework to predict the status of molecular pathways and key mutations in colorectal cancer from routine histology images: a retrospective study. Lancet Digit Health, 2021. [PubMed]
- K Thandiackal, B Chen, P Pati. 17th European Conference on Computer Vision (ECCV), OCT 23-27, 2022, Tel Aviv, ISRAEL, 2022
- Sara P. Oliveira, Pedro C. Neto, João Fraga. Cad systems for colorectal cancer from wsi are still not ready for clinical acceptance. Sci Rep, 2021. [PubMed]
- 691Chuang Zhu, Wenkai Chen, Ting Peng, Ying Wang, and Mulan Jin. Hard sample aware noise robust learning for histopathology image classification. IEEE Trans Med Imaging.
- Philipp Kainz, Martin Urschler, Samuel Schulter, Paul Wohlhart, Vincent Lepetit. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2015
- Ramraj Chandradevan, Ahmed A. Aljudi, Bradley R. Drumheller. Machine-based detection and classification for bone marrow aspirate differential counts: initial development focusing on nonneoplastic cells. Lab Invest, 2020. [PubMed]
- Arthur O. Frankel, Melvin Lathara, Celine Y. Shaw. Machine learning for rhabdomyosarcoma histopathology. Mod Pathol, 2022. [PubMed]
- Xueyi Zheng, Ruixuan Wang, Xinke Zhang. A deep learning model and human-machine fusion for prediction of ebv-associated gastric cancer from histopathology. Nat Commun, 2022. [PubMed]
- Ye Tian, Li Yang, Wei Wang. Computer-aided detection of squamous carcinoma of the cervix in whole slide images. arXiv, 2019
- Jevgenij Gamper, Navid Alemi Koohbanani, Ksenija Benet, Ali Khuram, Nasir Rajpoot. European Congress on Digital Pathology, 2019
- Jevgenij Gamper, Navid Alemi Koohbanani, Simon Graham. Pannuke dataset extension, insights and baselines. CoRR, 2020
- Le Hou, Rajarsi Gupta, John S. Van Arnam. Dataset of segmented nuclei in hematoxylin and eosin stained histopathology images of ten cancer types. Sci Data, 2020. [PubMed]
- Cheng Jiang, Jun Liao, Pei Dong. Blind deblurring for microscopic pathology images using deep learning networks. CoRR, 2020
- Robert J. Marinelli, Kelli Montgomery, Chih Long Liu. The stanford tissue microarray database. Nucleic Acids Res, 2007. [PubMed]
- Narayan Hegde, Jason D. Hipp, Yun Liu. Similar image search for histopathology: Smily. NPJ Digit Med, 2019. [PubMed]
- James A. Diao, Wan Fung Chui, Jason K. Wang. Dense, high-resolution mapping of cells and tissues from pathology images for the interpretable prediction of molecular phenotypes in cancer. bioRxiv, 2020
- Yiqing Shen, Jing Ke. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2020
- Benoît Schmauch, Alberto Romagnoni, Elodie Pronier. A deep learning model to predict rna-seq expression of tumours from whole slide images. Nat Commun, 2020. [PubMed]
- Andrew A. Borkowski, Marilyn M. Bui, L. Brannon Thomas, Catherine P. Wilson, Lauren A. DeLand, Stephen M. Mastorides. Lung and colon cancer histopathological image dataset (lc25000). arXiv, 2019
- Jeongun Ryu, Aaron Valero Puche, JaeWoong Shin. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2023
- Christian Matek, Simone Schwarz, Karsten Spiekermann, Carsten Marr. Human-level recognition of blast cells in acute myeloid leukaemia with convolutional neural networks. Nat Mach Intell, 2019
- Jeffrey J. Nirschl, Andrew Janowczyk, Eliot G. Peyster. A deep-learning classifier identifies patients with clinical heart failure using whole-slide images of h&e tissue. PloS One, 2018
- Julia Höhn, Eva Krieghoff-Henning, Tanja B. Jutzi. Combining cnn-based histologic whole slide image analysis and patient data to improve skin cancer classification. Eur J Cancer, 2021. [PubMed]
- Felipe Giuste, Mythreye Venkatesan, Conan Zhao. Proceedings of the 11th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics, 2020
- Eliot G. Peyster, Sara Arabyarmohammadi, Andrew Janowczyk. An automated computational image analysis pipeline for histological grading of cardiac allograft rejection. Eur Heart J, 2021. [PubMed]
- Juan C. Caicedo, Allen Goodman, Kyle W. Karhohs. Nucleus segmentation across imaging experiments: the 2018 data science bowl. Nat Methods, 2019. [PubMed]
- Ruggero Donida Labati, Vincenzo Piuri, Fabio Scotti. 2011 18th IEEE International Conference on Image Processing, 2011
- Angelo Genovese, Mahdi S. Hosseini, Vincenzo Piuri, Konstantinos N. Plataniotis, Fabio Scotti. 2021 IEEE International Conference on Computational Intelligence and Virtual Environments for Measurement Systems and Applications (CIVEMSA), 2021
- Henrik Failmezger, Sathya Muralidhar, Antonio Rullan, Carlos E. de Andrea, Erik Sahai, Yinyin Yuan. Topological tumor graphs: A graph-based spatial model to infer stromal recruitment for immunosuppression in melanoma histology. Cancer Res, 2020. [PubMed]
- Zahraa Al-Milaji, Ilker Ersoy, Adel Hafiane, Kannappan Palaniappan, Filiz Bunyak. Integrating segmentation with deep learning for enhanced classification of epithelial and stromal tissues in H&E images. Pattern Recogn Lett, 2019
- Mehmet Günhan Ertosun, Daniel L. Rubin. AMIA Annual Symposium Proceedings, 2015. [PubMed]
- Saima Rathore, Tamim Niazi, Muhammad Aksam Iftikhar, Ahmad Chaddad. Glioma grading via analysis of digital pathology images using machine learning. Cancers, 2020. [PubMed]
- Jonathan Folmsbee, Xulei Liu, Margaret Brandwein-Weber, Scott Doyle. IEEE International Symposium on Biomedical Imaging, 2018
- James S. Lewis Jr, Sahirzeeshan Ali, Jingqin Luo, Wade L. Thorstad, Anant Madabhushi. A quantitative histomorphometric classifier (quhbic) identifies aggressive versus indolent p16-positive oropharyngeal squamous cell carcinoma. Am J Surg Pathol, 2014. [PubMed]
- Sara Hosseinzadeh Kassani, Peyman Hosseinzadeh Kassani, Michal J. Wesolowski, Kevin A. Schneider, Ralph Deters. International Conference on Information and Communication Technology Convergence (ICTC), 2019
- SanaUllah Khan, Naveed Islam, Zahoor Jan, Ikram Ud Din, Joel J.P.C. Rodrigues. A novel deep learning based framework for the detection and classification of breast cancer using transfer learning. Pattern Recogn Lett, 2019
- Ümit Budak, Zafer Cömert, Zryan Najat Rashid, Abdulkadir Şengür, Musa Çıbuk. Computer-aided diagnosis system combining fcn and bi-lstm model for efficient breast cancer detection from histopathological images. Appl Soft Comput, 2019
- Fabio A. Spanhol, Luiz S. Oliveira, Paulo R. Cavalin, Caroline Petitjean, Laurent Heutte. IEEE International Conference on Systems, Man, and Cybernetics (SMC), 2017
- David Joon Ho, Dig V.K. Yarlagadda, Timothy M. D’Alfonso. Deep multi-magnification networks for multi-class breast cancer image segmentation. Comput Med Imaging Graph, 2021
- Mira Valkonen, Kimmo Kartasalo, Kaisa Liimatainen, Matti Nykter, Leena Latonen, Pekka Ruusuvuori. IEEE International Conference on Computer Vision Workshops, 2017
- Yu Liang, Jinglong Yang, Xiongwen Quan, Han Zhang. Chinese Automation Congress (CAC), 2019
- Angel Cruz-Roa, Ajay Basavanhally, Fabio González. Medical Imaging: Digital Pathology, 2014
- Md Zahangir Alom, Chris Yakopcic, Mst Nasrin. Breast cancer classification from histopathological images with inception recurrent residual convolutional neural network. J Digit Imaging, 2019. [PubMed]
- Majid Nawaz, Adel A. Sewissy, A. Taysir Hassan, Soliman. Multi-class breast cancer classification using deep learning convolutional neural network. Int J Adv Comput Sci Appl, 2018
- Ziba Gandomkar, Patrick C. Brennan, Claudia Mello-Thoms. Mudern: Multi-category classification of breast histopathological image using deep residual networks. Artif Intell Med, 2018. [PubMed]
- Zhongyi Han, Benzheng Wei, Yuanjie Zheng, Yilong Yin, Kejian Li, Shuo Li. Breast cancer multi-classification from histopathological images with structured deep learning model. Sci Rep, 2017. [PubMed]
- Pendar Alirezazadeh, Behzad Hejrati, Alireza Monsef-Esfahani, Abdolhossein Fathi. Representation learning-based unsupervised domain adaptation for classification of breast cancer histopathology images. Biocybernet Biomed Eng, 2018
- Zhu Meng, Zhicheng Zhao, Su. Fei. IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), 2019
- Alexander Rakhlin, Alexey Shvets, Vladimir Iglovikov, Alexandr A. Kalinin. International Conference Image Analysis and Recognition, 2018
- Yeeleng S. Vang, Zhen Chen, Xiaohui Xie. International Conference Image Analysis and Recognition, 2018
- Abdullah-Al Nahid, Yinan Kong. Histopathological breast-image classification using local and frequency domains by convolutional neural network. Information, 2018
- Eu Wern Teh, Graham W. Taylor. International Conference on Medical Imaging with Deep Learning–Extended Abstract Track, 2019
- Ruqayya Awan, Navid Alemi Koohbanani, Muhammad Shaban, Anna Lisowska, Nasir Rajpoot. International Conference Image Analysis and Recognition, 2018
- Tomas Iesmantas, Robertas Alzbutas. International Conference Image Analysis and Recognition, 2018
- Kaushiki Roy, Debapriya Banik, Debotosh Bhattacharjee, Mita Nasipuri. Patch-based system for classification of breast histology images using deep learning. Comput Med Imaging Graph, 2019. [PubMed]
- Gaoyi Lei, Yuanqing Xia, Di-Hua Zhai, Wei Zhang, Duanduan Chen, Defeng Wang. Staincnns: An efficient stain feature learning method. Neurocomputing, 2020
- Amjad Khan, Manfredo Atzori, Sebastian Otálora, Vincent Andrearczyk, Henning Müller. Medical Imaging: Digital Pathology, 2020
- Dorsa Ziaei, Weizhe Li, Samuel Lam, Wei-Chung Cheng, Weijie Chen. Medical Imaging: Digital Pathology, 2020
- X. Guo, F. Wang, G. Teodoro, A.B. Farris, J. Kong. IEEE International Symposium on Biomedical Imaging, 2019
- Yongxiang Huang, Albert Chung. International Conference on Medical Image Computing and Computer-Assisted Intervention, 2019
- Şaban Öztürk, Bayram Akdemir. Hic-net: A deep convolutional neural network model for classification of histopathological breast images. Comput Electric Eng, 2019
- K Stacke, G Eilertsen, J Unger, C Lundström. Measuring domain shift for deep learning in histopathology. IEEE journal of biomedical and health informatics, 2020 Oct 20
- Jacob Gildenblat, Eldad Klaiman. Self-supervised similarity learning for digital pathology. arXiv, 2019
- Sebastian Otálora, Manfredo Atzori, Amjad Khan, Oscar Jimenez-del Toro, Vincent Andrearczyk, Henning Müller. Medical Imaging: Digital Pathology, 2020
- Jiayun Li, Karthik V. Sarma, King Chung Ho, Arkadiusz Gertych, Beatrice S. Knudsen, Corey W. Arnold. 2017
- Juan S. Lara, H. Victor, O. Contreras, Sebastián Otálora, Henning Müller, Fabio A. González. Medical Image Computing and Computer Assisted Intervention – MICCAI, Sep 2020
- Ethan H. Nguyen, Haichun Yang, Ruining Deng. Circle representation for medical object detection. IEEE Trans Med Imaging, 2021
- Saif Almansouri, Susan Zwyea. 2020
- Simon Graham, Muhammad Shaban, Talha Qaiser, Navid Alemi Koohbanani, Syed Ali Khurram, Nasir Rajpoot. Medical Imaging: Digital Pathology, 2018
- Germán Corredor, Paula Toro, Kaustav Bera. Computational pathology reveals unique spatial patterns of immune response in H&E images from covid-19 autopsies: preliminary findings. J Med Imaging, 2021
- Mohammed Adnan, Shivam Kalra, Hamid R. Tizhoosh. IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops, 2020
- Miriam Hägele, Philipp Seegerer, Sebastian Lapuschkin. Resolving challenges in deep learning-based analyses of histopathological images using explanation methods. Sci Rep, 2020. [PubMed]
- Yves-Rémi Van Eycke, Cédric Balsat, Laurine Verset, Olivier Debeir, Isabelle Salmon, Christine Decaestecker. Segmentation of glandular epithelium in colorectal tumours to automatically compartmentalise ihc biomarker quantification: A deep learning approach. Med Image Anal, 2018. [PubMed]
- Xu Yan, Yang Li, Yipei Wang. Gland instance segmentation using deep multichannel neural networks. IEEE Trans Biomed Eng, 2017. [PubMed]
- Jakob Kather, Cleo-Aron Weis, Francesco Bianconi. Multi-class texture analysis in colorectal cancer histology. Sci Rep, 2016. [PubMed]
- Chaofeng Wang, Jun Shi, Qi Zhang, Shihui Ying. International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), 2017
- Łukasz Rączkowski, Marcin Możejko, Joanna Zambonelli, Ewa Szczurek. Ara: accurate, reliable and active histopathological image classification framework with bayesian deep learning. Sci Rep, 2019. [PubMed]
- Srinath Jayachandran, Ashlin Ghosh. IAPR Workshop on Artificial Neural Networks in Pattern Recognition, 2020
- Adrien Foucart, Olivier Debeir, Christine Decaestecker. IEEE International Symposium on Biomedical Imaging, 2019
- Amal Lahiani, Irina Klaman, Nassir Navab, Shadi Albarqouni, Eldad Klaiman. Seamless virtual whole slide image synthesis and validation using perceptual embedding consistency. IEEE J Biomed Health Inform, 2020
- Meng-Yao Ji, Lei Yuan, Lu Shi-Min. Glandular orientation and shape determined by computational pathology could identify aggressive tumor for early colon carcinoma: a triple-center study. J Transl Med, 2020. [PubMed]
- Ju Hang Chang, Cheng Zhong Han, Antoine M. Snijders, Jian-Hua Mao. Unsupervised transfer learning via multi-scale convolutional sparse coding for biomedical applications. IEEE Trans Pattern Anal Mach Intell, 2017. [PubMed]
- Jacob S. Sarnecki, Kathleen H. Burns, Laura D. Wood. A robust nonlinear tissue-component discrimination method for computational pathology. Lab Invest, 2016. [PubMed]
- Corentin Gueréndel, Phil Arnold, Ben Torben-Nielsen. MICCAI Workshop on Computational Pathology, 2021
- Huisi Wu, Zhaoze Wang, Youyi Song, Yang Lin, Jing Qin. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2022
- Soufiane Belharbi, Jérôme Rony, Jose Dolz, Ismail Ben Ayed, Luke McCaffrey, Eric Granger. Deep interpretable classification and weakly-supervised segmentation of histology images via max-min uncertainty. IEEE Trans Med Imaging, 2021
- Parmida Ghahremani, Joseph Marino, Ricardo Dodds, Saad Nadeem. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2022
- Pushpak Pati, Antonio Foncubierta-Rodríguez, Orcun Goksel, Maria Gabrani. Reducing annotation effort in digital pathology: A co-representation learning framework for classification tasks. Med Image Anal, 2021
- Yiqing Shen, Dinggang Shen, Jing Ke. Identify representative samples by conditional random field of cancer histology images. IEEE Trans Med Imaging, 2022. [PubMed]
- Chuang Zhu, Wenkai Chen, Ting Peng, Ying Wang, Mulan Jin. Hard sample aware noise robust learning for histopathology image classification. IEEE Trans Med Imaging, 2021
- Ming Y. Lu, Drew F.K. Williamson, Tiffany Y. Chen, Richard J. Chen, Matteo Barbieri, Faisal Mahmood. Data-efficient and weakly supervised computational pathology on whole-slide images. Nat Biomed Eng, 2021. [PubMed]
- Chetan L. Srinidhi, Seung Wook Kim, Fu-Der Chen, Anne L. Martel. Self-supervised driven consistency training for annotation efficient histopathology image analysis. Med Image Anal, 2022
- Yu’Ang Zhu, Yuxin Zheng, Zhao Chen. International Conference on Pattern Recognition and Intelligent Systems, 2021
- Soufiane Belharbi, Jérôme Rony, Jose Dolz, Ismail Ben Ayed, Luke McCaffrey, Eric Granger. IEEE Transactions on Medical Imaging, 2021
- Simon Graham, David Epstein, Nasir Rajpoot. Dense steerable filter cnns for exploiting rotational symmetry in histology images. IEEE Trans Med Imaging, 2020. [PubMed]
- Md Shamima Nasrin, Zahangir Alom, Tarek M. Taha, Vijayan K. Asari. Medical Imaging: Digital Pathology, 2020
- Le Hou, Dimitris Samaras, Tahsin M. Kurc, Yi Gao, James E. Davis, Joel H. Saltz. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2016
- Massimo Salvi, Nicola Michielli, Filippo Molinari. Stain color adaptive normalization (scan) algorithm: Separation and standardization of histological stains in digital pathology. Comput Methods Programs Biomed, 2020
- Shivam Kalra, Hamid R. Tizhoosh, Sultaan Shah. Pan-cancer diagnostic consensus through searching archival histopathology images using artificial intelligence. NPJ Digit Med, 2020. [PubMed]
- Ali Mirzazadeh, Arshawn Mohseni, Sahar Ibrahim. IEEE EMBS International Conference on Biomedical and Health Informatics (BHI), 2021
- Mu Youqing, Hamid R. Tizhoosh, Rohollah Moosavi Tayebi. A bert model generates diagnostically relevant semantic embeddings from pathology synopses with active learning. Commun Med, 2021. [PubMed]
- Tathagato Rai Dastidar, Renu Ethirajan. Whole slide imaging system using deep learning-based automated focusing. Biomed Opt Express, 2020. [PubMed]
- Pinky A. Bautista, Yukako Yagi. 2009 Annual International Conference of the IEEE Engineering in Medicine and Biology Society, 2009
- Joseph Boyd, Mykola Liashuha, Eric Deutsch, Nikos Paragios, Stergios Christodoulidis, Maria Vakalopoulou. IEEE/CVF International Conference on Computer Vision, 2021
