Digital pathology and multimodal learning on oncology data
Abstract
Cancer presents a complex tapestry of biological, clinical, and molecular characteristics that collectively influence its diagnosis, progression, and treatment. This review article delves into the recent advancements in integrating multimodal oncology data, a pivotal aspect in the evolving field of digital pathology (DP). The complexity and heterogeneity of cancer, characterized by its multimodal nature, present significant challenges in diagnosis and treatment. Traditional methods of oncology data analysis often fall short of capturing the comprehensive landscape of cancer’s multifaceted characteristics. The advent of artificial intelligence, machine learning, and deep learning has marked a new era in multimodal learning. These technologies have revolutionized how oncologists and researchers understand and approach cancer, allowing for a more nuanced and detailed analysis. In this review article, we attempt to examine and present how DP enriches its methods and analysis with other data modalities, including clinical, radiological, and molecular information. We present opportunities and challenges of multimodal learning in oncology, highlighting the synergistic potential of combining various data types for improving cancer care with a focus on DP. Continuous innovation in multimodal data integration will be instrumental in transforming cancer diagnosis, treatment planning, prognosis, and post-treatment surveillance.
Article type: Review Article
Keywords: artificial intelligence, machine learning, digital pathology, computational pathology, multimodal learning, oncology data integration, cancer research
Affiliations: Department of Cancer Epidemiology, H. Lee Moffitt Cancer Center & Research Institsute, Tampa, FL, 33612, United States; Machine Learning Department, H. Lee Moffitt Cancer Center & Research Institute, Tampa, FL, 33612, United States; Rashid Latif Medical College, Lahore, 54600, Pakistan; Mobile Infirmary Medical Center, Mobile, AL, 36607, United States; Department of Pathology, H. Lee Moffitt Cancer Center & Research Institute, Tampa, FL, 33612, United States; Morsani College of Medicine, University of South Florida, Tampa, FL, 33612, United States; Department of Neuro-Oncology, H. Lee Moffitt Cancer Center & Research Institute, Tampa, FL, 33612, United States
License: © The Author(s) 2024. Published by Oxford University Press on behalf of the British Institute of Radiology. CC BY 4.0 This is an Open Access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted reuse, distribution, and reproduction in any medium, provided the original work is properly cited.
Article links: DOI: 10.1093/bjrai/ubae014 | PubMed: 42064394 | PMC: PMC13045696
Relevance: Moderate: mentioned 3+ times in text
Full text: PDF (1.4 MB)
Introduction
The examination of tissue samples under a microscope by pathologists is the standard procedure for disease diagnosis in clinical settings. However, technological advancements in the last couple of decades have ushered in a new era in the field of pathology called digital pathology (DP). Some of the key factors that have enabled the transformation from traditional pathology to DP include: (1) Whole Slide Imaging (WSI), which scans entire tissue sections on glass slides at high resolution and stores them as digital slides that can be analysed computationally,ref. ubae014-B1 (2) Image analysis algorithms from the fields of computer vision and machine learning (ML) that can detect, segment, classify, and quantify morphological features, cells, tissues, biomarkers from digitized images,ref. ubae014-B2 (3) Advanced imaging techniques such as computational staining and multiplexed imaging,ref. ubae014-B3 (4) Molecular assays that generate quantitative molecular data from tissues using techniques such as polymerase chain reaction, microarrays, and next-generation sequencing,ref. ubae014-B4 and (5) Enterprise imaging that includes pathology informatics.ref. ubae014-B5
As the field of pathology evolves, so too does our understanding of cancer. Cancer is a complex disease that can manifest in different forms, each with its distinct characteristics and implications for screening, diagnosis, prognosis, and treatment. Over time, our understanding of cancer has evolved. Initially thought to be primarily a result of genetic mutations, it is now recognized as a complex interplay of genetic, environmental, and lifestyle factors. Advances in genomics, proteomics, bioinformatics, and imaging have led to a deeper understanding of cancer, revealing that even cancers originating in the same organ can have vastly different characteristics and responses to treatment. This complexity underscores the necessity for personalized medicine approaches tailored to the unique genetic makeup of each cancer and each patient.
Transitioning from the complexity of cancer biology to the use of advanced technologies, the success of artificial intelligence (AI) and ML in different domains of science, especially those focused on quantitative imaging analysis, has attracted anatomical pathology researchers and practitioners. As a result, computational pathology (CPATH) emerged when AI and ML models were introduced for various types of quantitative analysis in DP.ref. ubae014-B6 CPATH uses AI and ML models to extract information from digital images that may extend beyond immediate human capabilities. By offering AI/ML-powered analytical tools, CPATH has the potential to enhance the roles of pathologists in clinical settings by making their work more efficient, accurate, and reproducible. This enhancement spans various aspects, including improving the precision, reproducibility, and scalability of tasks related to disease diagnosis and the quantification of biomarkers for prognosis and prediction.
However, while the integration of AI and ML into DP clinical workflow promises significant advances, it also presents several challenges. These challenges include a shortage of staff trained in both AI and DP that may temporarily lead to increased workloads primarily due to the learning curve in adoption of AI in the DP workflows. Nevertheless, over time, the increased use of AI and its incorporation into the training curriculum will gradually improve the number of AI-qualified individuals (see definition in Box 1) in the DP/CPATH field. The growing complexity of diagnostics due to evolving protocols and new biomarkers, as well as variations in rare diseases and morphological similarities, also pose challenges. Additionally, quality issues arising from tissue artefacts and staining/imaging inconsistencies, along with a lack of standardized practices and protocols, may impede interoperability and pose reproducibility challenges.
Box 1.Key pathology terminologies.PathologyStems from the Greek word pathologia, meaning the study of suffering, it is a medical specialty investigating the origin, progression, and alterations in the structure, function, and natural course of diseases.7 There are two main branches of pathology: anatomical pathology (AP) focuses on studying tissue samples to diagnose diseases such as cancer, and clinical pathology (CP) analyses bodily fluids and conducts laboratory tests.8 Molecular pathology is an emerging branch that studies diseases at the molecular level by examining tissue or body fluids.HistologyMicroscopic examination of tissues and organs by sectioning, applying stains, and observing the prepared sections under a microscope. This process enables the visualization of tissue structures and any distinctive alterations the tissue might have undergone. Haematoxylin (a basic dye that stains cell nuclei in a purplish blue colour) and Eosin (an acidic dye that stains cytoplasm in a pinkish red colour), referred to as H&E, are commonly used together to stain structures of the cells to define intracellular organelles and proteins.9HistopathologyIt extends the utility of histological analysis by focusing on the study of disease indicators through the microscopic examination of processed and fixed glass slides containing sections from biopsies or surgical specimens. To visualize various tissue components under a microscope, these specimens are dyed with one or more stains depending on the investigational question being asked.9,10Immunohistochemistry (IHC)A valuable tool extensively employed in pathology for cell classification and diagnosis. Commonly performed on formalin-fixed paraffin-embedded (FFPE) tissue, IHC targets specific antigens in tissues and cells using antibodies to determine cell type and organ of origin. IHC usage has lately expanded to study predictive and prognostic biomarkers in oncological settings, making it a useful technique in modern pathology practice.11WSIThe process of digitizing glass slides at multiple magnifications and focal levels to create digital images for observation and image analysis. WSI aims to replicate the experience of traditional light microscopy through digital means and is often known as virtual microscopy.12Cytology slidesThese refer to glass slides that contain specimens of individual cells or cell clusters obtained from various bodily fluids, tissues, or fine needle aspiration (FNA) biopsy. Unlike histology, which focuses on the function of tissues, cytology slides are prepared for microscopic examination to analyse cellular morphology, structure, and characteristics.13Digital pathology (DP)DP includes converting histopathology, IHC, or cytology slides into digital formats through whole-slide scanners, followed by applying computer-aided analytical tools for objective analysis. DP has improved pathology diagnoses through multiple techniques as discussed in this review.CPATHCPATH is the sub-branch of DP that involves computational analysis of digital slides. CPATH aims to analyse patient specimens by extracting information from digitized pathology images in combination with their associated meta-data, typically using AI methods to gain valuable insights into disease processes.6,8,14RadiomicsRadiomics is a sub-field of radiology, where radiological images are converted into high-dimensional mine-able data for AI-based analysis to uncover biomarkers for diagnosis, prognosis, and treatment response prediction.15PathomicsPathomics is a sub-field of pathology, that covers a wide variety of data captured from image analyses in the form of features that describe multiple phenotypic features of tissues in WSIs. Using AI, pathomics provides a quantitative assessment of identified structures and features to complement traditional histopathologic evaluation by pathologists.16AI-qualified individualsIn the DP/CPATH context, individuals who not only understand AI at a basic technical level but also have the clinical, regulatory, and practical knowledge to apply AI effectively in pathology. This includes clinical validation, understanding regulatory and ethical implications, and the ability to implement, use, and interpret AI tools and their outcomes in daily practice. While it is optional but valuable that these individuals may also be involved in developing new AI tools, the term more broadly encompasses those who are well-equipped to work with AI in a clinical environment, ensuring its accurate, safe, and effective use.
Box 1.: Key pathology terminologies.
| Pathology | Stems from the Greek word pathologia, meaning the study of suffering, it is a medical specialty investigating the origin, progression, and alterations in the structure, function, and natural course of diseases.ref. ubae014-B7 There are two main branches of pathology: anatomical pathology (AP) focuses on studying tissue samples to diagnose diseases such as cancer, and clinical pathology (CP) analyses bodily fluids and conducts laboratory tests.ref. ubae014-B8 Molecular pathology is an emerging branch that studies diseases at the molecular level by examining tissue or body fluids. |
| Histology | Microscopic examination of tissues and organs by sectioning, applying stains, and observing the prepared sections under a microscope. This process enables the visualization of tissue structures and any distinctive alterations the tissue might have undergone. Haematoxylin (a basic dye that stains cell nuclei in a purplish blue colour) and Eosin (an acidic dye that stains cytoplasm in a pinkish red colour), referred to as H&E, are commonly used together to stain structures of the cells to define intracellular organelles and proteins.ref. ubae014-B9 |
| Histopathology | It extends the utility of histological analysis by focusing on the study of disease indicators through the microscopic examination of processed and fixed glass slides containing sections from biopsies or surgical specimens. To visualize various tissue components under a microscope, these specimens are dyed with one or more stains depending on the investigational question being asked.ref. ubae014-B9,ref. ubae014-B10 |
| Immunohistochemistry (IHC) | A valuable tool extensively employed in pathology for cell classification and diagnosis. Commonly performed on formalin-fixed paraffin-embedded (FFPE) tissue, IHC targets specific antigens in tissues and cells using antibodies to determine cell type and organ of origin. IHC usage has lately expanded to study predictive and prognostic biomarkers in oncological settings, making it a useful technique in modern pathology practice.ref. ubae014-B11 |
| WSI | The process of digitizing glass slides at multiple magnifications and focal levels to create digital images for observation and image analysis. WSI aims to replicate the experience of traditional light microscopy through digital means and is often known as virtual microscopy.ref. ubae014-B12 |
| Cytology slides | These refer to glass slides that contain specimens of individual cells or cell clusters obtained from various bodily fluids, tissues, or fine needle aspiration (FNA) biopsy. Unlike histology, which focuses on the function of tissues, cytology slides are prepared for microscopic examination to analyse cellular morphology, structure, and characteristics.ref. ubae014-B13 |
| Digital pathology (DP) | DP includes converting histopathology, IHC, or cytology slides into digital formats through whole-slide scanners, followed by applying computer-aided analytical tools for objective analysis. DP has improved pathology diagnoses through multiple techniques as discussed in this review. |
| CPATH | CPATH is the sub-branch of DP that involves computational analysis of digital slides. CPATH aims to analyse patient specimens by extracting information from digitized pathology images in combination with their associated meta-data, typically using AI methods to gain valuable insights into disease processes.ref. ubae014-B6,ref. ubae014-B8,ref. ubae014-B14 |
| Radiomics | Radiomics is a sub-field of radiology, where radiological images are converted into high-dimensional mine-able data for AI-based analysis to uncover biomarkers for diagnosis, prognosis, and treatment response prediction.ref. ubae014-B15 |
| Pathomics | Pathomics is a sub-field of pathology, that covers a wide variety of data captured from image analyses in the form of features that describe multiple phenotypic features of tissues in WSIs. Using AI, pathomics provides a quantitative assessment of identified structures and features to complement traditional histopathologic evaluation by pathologists.ref. ubae014-B16 |
| AI-qualified individuals | In the DP/CPATH context, individuals who not only understand AI at a basic technical level but also have the clinical, regulatory, and practical knowledge to apply AI effectively in pathology. This includes clinical validation, understanding regulatory and ethical implications, and the ability to implement, use, and interpret AI tools and their outcomes in daily practice. While it is optional but valuable that these individuals may also be involved in developing new AI tools, the term more broadly encompasses those who are well-equipped to work with AI in a clinical environment, ensuring its accurate, safe, and effective use. |
Amid these challenges, multimodal learning in oncology represents a cutting-edge approach to cancer research and treatment, leveraging the integration of various data types to gain a comprehensive understanding of cancer’s complexities.ref. ubae014-B17 This approach combines diverse datasets, including genomic, proteomic, imaging, clinical, and demographic data, to develop a holistic view of cancer biology. By harnessing advanced computational techniques offered by AI/ML, multimodal learning enables analysing these disparate data types jointly in a unified framework.ref. ubae014-B8,ref. ubae014-B19 This integration is paramount for identifying novel biomarkers, elucidating tumour heterogeneity, predicting treatment responses, and ultimately facilitating the development of more effective, personalized therapies.ref. ubae014-B20 Building upon these developments, DP and CPATH have paved the way for exploring novel methods for cancer diagnosis, prognosis, treatment planning, and also for research into a deeper understanding of cancer biology.ref. ubae014-B21 The innovative algorithms within CPATH, especially those employing differential privacy and federated learning, enable the development of robust analytical models. These models are crucial for safeguarding patient privacy, fostering collaboration among institutions, and paving the way for the adoption of clinical-grade systems.ref. ubae014-B25 Such advancements underscore the critical role of AI and ML in transforming cancer care, making it more precise, personalized, and effective.
From conventional to digital pathology
Conventionally, pathologists observe tissue samples on the glass slides under a microscope.ref. ubae014-B8 Such a traditional approach involves physical preparation and storage of glass slides and local consultations only, limiting the efficiency, reproducibility, scalability, and accessibility of the technique. These limitations, along with the advent of modern computers, imaging systems, and quantitative image processing techniques, have prompted a shift towards DP.ref. ubae014-B26 The journey of digitization in pathology began with the development of the first virtual microscope during 1996-1998, where high-resolution images replicating the glass slides were produced using imaging scanners.ref. ubae014-B27 The system was later advanced to support data caching and precomputed image pyramids.ref. ubae014-B27 In the last two and half decades, the emergence and commercialization of WSI and glass slide scanners have revolutionized clinical pathology practices.ref. ubae014-B27,ref. ubae014-B28 Presently, many commercial WSI scanners are available (Philips, Leica, and Hamamatsu, etc), and many open-source WSI systems (caMicroscope, Digital Slide Archive, Sedeen viewer, QuPath) are supported actively.ref. ubae014-B29 The advent of DP, powered by WSI, has revolutionized medical diagnostics.ref. ubae014-B26,ref. ubae014-B33 Improvements in technology by turning glass slides into digital formats and decreasing storage costs have increased the use of WSIs.ref. ubae014-B34 The adoption of DP by pathologists has seen significant growth, increasing from 30% in 2013 to 64% in 2021, according to a report by the College of American Pathologists.ref. ubae014-B26 In Figure 1, we have presented a template for the future roadmap for DP and CPATH that may have the necessary components and methods in a unified framework. With a multitude of advantages, DP has brought a revolution in high-resolution magnification analysis, electronic storage, automation of processes, quality assurance and control, telepathology and enhanced diagnostic efficiency, streamlined diagnostic workflow, accessible measurements and quantification, cancer prescreening, intraoperative consultation, histopathological second opinion, and education and training.
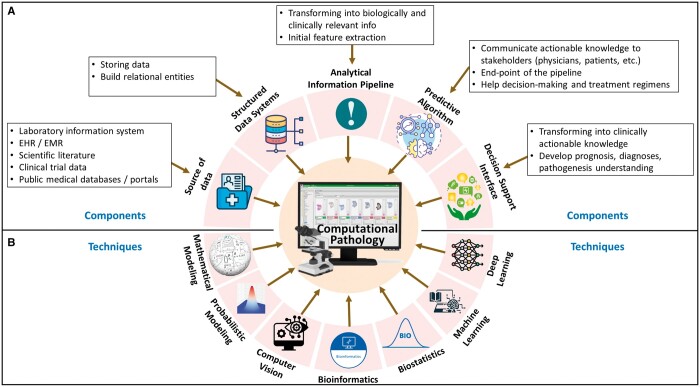
Computational pathology—AI in DP
More recently, the advancements in AI and ML have started revolutionizing DP, which has been a helpful tool in histopathology image analysis in the last decade.ref. ubae014-B26 Digital Pathology Association (DPA) defines CPATH as, “the big-data approach to pathology,” where patient’s pathology images and associated meta-data are combined to extract patterns and analyse image features through AI techniques.ref. ubae014-B6 To address the evolving landscape of pathology practice and the increasing adoption of digital tools in diagnostic workflows, the College of American Pathologists in consultation with the American Medical Association, announced the inclusion of 30 new current procedural terminologies (CPTs) related to DP for use by pathologists, effective from January 1, 2024.ref. ubae014-B35 The integration of AI in DP has facilitated precise diagnosis by processing information that may not be perceived by the human eye at times.ref. ubae014-B8,ref. ubae014-B27 The cardinal tasks of AI in DP include the development and validation of models and algorithms that can detect different patterns and features from data presented in the form of WSIs. The AI models can identify regions of interest (ROIs) from WSIs, extract and classify patches, classify entire WSIs, generate labels, and identify various objects of interest.ref. ubae014-B27 Examples of how various AI and ML models have been proposed to advance various DP tasks include, diagnosis support, enhanced image analysis, improving prognostics and biomarkers predictions, post-treatment analysis and disease surveillance, and cross-modality learning.ref. ubae014-B6,ref. ubae014-B8,ref. ubae014-B19,ref. ubae014-B26,ref. ubae014-B36 The key terminologies used in AI are presented in Box 2.
Box 2.Key AI terminologies.AIInitially started as a simple theory of human intelligence being exhibited by machines, AI aims to simulate human intelligence in machines by developing computers that can engage in learning, reasoning, self-correction, problem-solving, and decision-making.37MLML is a subset of AI that programs computers to process natural data and solve an optimization problem using data samples or past experience. It is an engineering discipline that is closely related to mathematical statistics to build learning models.38Deep Learning (DL)DL is a subfield of ML and refers to models developed using multiple layers of artificial neural networks. The deep neural networks focus on compositional learning using multiple (hence the word “deep”) layers of representations, where each layer is built using multiple artificial neurons.38–42 Various types of DL models have been developed, including Convolutional Neural networks (CNNs), Recurrent Neural networks (RNNs), Graph Neural Networks (GNNs), Transformers, and many others.Convolutional Neural NetworksCNNs are specialized DL models for analysing image or grid data.38,43,44 CNNs use convolution operations and can learn spatial correlations, features, and relationships from the input data.43,45Recurrent Neural NetworksRNNs are a family of DL models used for processing sequential data like speech, text, or time-series data (eg, patient vital signs recorded over time).43,46,47 RNNs are used to model time/space-based relationships between sequences of input data and to learn patterns based on the order (in time or space).43 Sub-categories of RNNs called Long short-term memory network (LSTM) and gated recurrent unit (GRU) models are better at learning long-term dependencies, making them great for real-world use cases, eg, predicting medical events and detecting anomalies in time series data.46Graph Neural NetworksGNNs are DL models designed for analysing graph data, like social relationships or biological pathways.40 GNNs pass messages between neighbouring nodes to aggregate local and global information. By modelling these relationships, GNNs can make predictions about nodes, edges, and sub-graphs in the network, where each entity may represent patients, drugs, proteins, etc GNNs enable new insights in fields like precision medicine and drug discovery.48–50TransformersTransformers are DL models that have shown promising results on language translation and image recognition tasks.51,52 Their key innovation is a technique called self-attention. Unlike RNNs, which process data sequentially, self-attention allows Transformers to learn relationships between any parts of the input, regardless of their position in the sequence. For example, a Transformer can learn that the symptoms “cough” and “fever” are strongly related even if they are distant in the patient’s medical history. The model learns which connections are important by assigning an “attention” score between input sequences (also referred to as tokens).47,51,52Foundation Models (FMs)FMs are a new class of DL models characterized by their large scale and ability to adapt to new tasks.53 FMs are trained on extensive datasets, allowing them to develop a broad understanding of various topics.8,54 The key characteristics of FMs include scalability in learning from diverse data sources, multimodal learning, compositionally in terms of generalization, and the emergence of implicit learning. Transfer learning, adaptation, in-context learning, and fine-tuning make FMs exceptionally useful across numerous fields, including the medical domain.55 A recent review paper covers existing FMs and their prospective use in DP.8Multimodal LearningMultimodal Learning involves integrating data from multiple modalities for an ML task, eg, prediction, classification, or regression. A modality refers to specific data input or source types, such as text, images, audio, video, or other data. The core idea is to leverage the complementary information from different modalities to improve the performance and accuracy of learning algorithms. Multimodal learning aims to achieve a more comprehensive understanding of the problem, leading to better decision-making than using a single modality.19,56
DP, CPATH, and multimodal AI
Advancements such as the digitization of patient health records, the development of new diagnostic technologies, innovative laboratory tests, and the establishment of robust information technology infrastructure in hospitals have led to an exponential increase in the volume of medical data collected for each patient.ref. ubae014-B19 Recent progress in multimodal learning, driven by deep neural networks, has demonstrated a remarkable ability to learn from varied data modalities, such as computer vision and natural language processing.ref. ubae014-B19,ref. ubae014-B53 Multimodal foundation models such as OpenAI’s Contrastive Language-Image Pretraining (CLIP) and Generative Pretraining Transformer (GPT-4) have established new standards in the field, while the Foundational Language And Vision Alignment Model (FLAVA) marks a notable advancement by integrating vision and language representation learning to enable multimodal reasoning.ref. ubae014-B57 The success of multimodal AI models in other domains has inspired cancer researchers to explore the potential of multimodal AI in healthcare.ref. ubae014-B60 The multimodal AI models being developed in the medical domain can integrate diverse modalities, including radiological images, genetic profiles, and clinical information, within the DP workflow, enhancing diagnostic precision and enabling personalized treatments.ref. ubae014-B61,ref. ubae014-B62 Multimodal AI models are poised to empower healthcare teams by offering them a holistic view of the patient’s disease state and paving the way for truly personalized cancer care.ref. ubae014-B9,ref. ubae014-B54,ref. ubae014-B61
In our context, “modality” refers to a specific data source or type. For example, EHR text descriptions of surgical procedures or post-visit reports represent one modality. Lab test values, vital measurements, and demographic information form another. Histopathology slides (stored as WSI), radiological scans (X-ray, MRI, PET, CT), and molecular data (genomics, proteomics) are separate modalities.ref. ubae014-B39,ref. ubae014-B62,ref. ubae014-B63 Some modalities have sub-categories, such as MRI scans with FLAIR, T1, T2, and CT scans with various views and contrast protocols.
In multimodal learning, researchers use various modality-specific data processing methods and data fusion mechanisms. The generally defined 5 main stages are: (1) preprocessing, (2) feature extraction, (3) data fusion, (4) primary learner, and (5) final classifier.ref. ubae014-B19 Data fusion from different modalities can occur at various levels. Early fusion combines features before feeding data into ML models.ref. ubae014-B62,ref. ubae014-B64 Intermediate fusion occurs at the model level, while late fusion merges processed data at the decision level.ref. ubae014-B19 Explainability in deep learning involves techniques to make models’ decisions, behaviours, and outputs transparent and understandable.ref. ubae014-B65,ref. ubae014-B66 This includes building interpretable models or generating post hoc explanations.ref. ubae014-B8,ref. ubae014-B65 In DP, which uses high-resolution WSIs, explainability is crucial. Techniques include heatmaps showing areas of significance, case-based explanations comparing with established predictions, and layer-wise relevance propagation to trace outputs back to inputs.ref. ubae014-B67
In the following, we provide a chronological overview of various applications of multimodal AI/ML techniques with a focus on histopathology data as the essential ingredient integrated with one or more other modalities (eg, radiology, genomics, or clinical data). A schematic layout of the multimodal AI in DP and CPATH is provided in Figure 2. We also provide a summary of references for each category of multimodal learning in Table 1, along with the explainability method of these works where applicable.
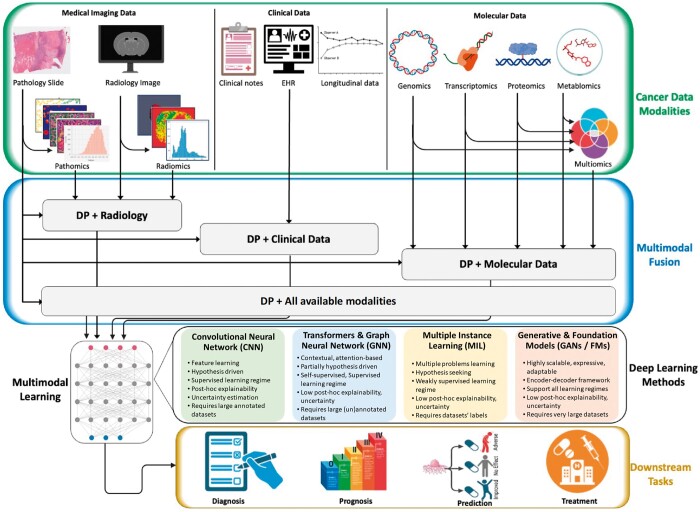
Table 1.: Summary of multimodal AI models for DP reported in this review.
| Modalities | Ref | Data type | Metrics | # patients | Task | Cancer type | Explainability |
|---|---|---|---|---|---|---|---|
| Pathology,Molecular | ref. ubae014-B68 | H&E, Multiomics (mutation status, CNV, RNAseq expr) | 5-fold cv,C-index, AUC | 5720 | Survival outcome prediction and prognostic features correlations. | 14 types (BLCA, BRCA, COADREAD, HNSC, KIRC, KIRP, LGG, LIHC, LUAD, LUSC, PAAD, SKCM, STAD, UCEC) | heatmaps, integrated gradients |
| ref. ubae014-B69 | H&E, Multiomics (mutation status, CNV, RNAseq expr) | 15-fold cv,C-index | Glioma: 769,CCRC: 417 | Survival outcome prediction. | Glioma (TCGA-GBM, LGG), CCRCC (TCGA-KIRC) | heatmaps | |
| ref. ubae014-B18 | WSIs, genomic features | 5-fold cv,C-index | 3523 | OS | 5 types (BLCA, BRCA, GBMLGG, LUAD, UCEC) | heatmaps | |
| ref. ubae014-B70 | WSIs, Multiomics (DNA mutation, mRNA expr) | AUC | 1214 | Microsatellite status prediction | TCGA-COAD, Asian-CRC | correlation heatmaps | |
| ref. ubae014-B71 | H&E slides, spatially resolved gene expression | Accuracy | 7 | Cancer detection, predict spatial variation of individual genes | Prostate cancer | heatmaps | |
| ref. ubae014-B72 | WSI, Multiomics (genomics, proteomics, and molecular subtyping) | C-index | 1888 | Survival prediction, MSI prediction | Colorectal cancer (TCGA-COAD, TCGA-READ) | importance scores, heatmaps | |
| Pathology,Clinical | ref. ubae014-B73 | WSIs, Pathomic features, Clinical | 10-fold cv,Accuracy, AUC | 735 | Tumour grade prediction | TCGA-LGG, TCGA-HGG | no |
| ref. ubae014-B74 | Dermatology images, Clinical | Balanced accuracy, AUC | >3300 | Cancer classification | Skin cancer | no | |
| Pathology,Radiology | ref. ubae014-B75 | WSI, MRI | NPV, PPV, accuracy, AUC, Cohen’s kappa | 400 | Tumour classification1) benign/cancerous, 2) high-risk/low-risk tumour | Prostate cancer | class activation heatmaps |
| ref. ubae014-B76 | Radiomics (from MRI), Pathomic features (from H&E) | Spearman’s correlation | 48 | Association among radiomics and pathomics features | Glioblastoma multiforme (CPTAC-GBM) | no | |
| ref. ubae014-B87 | MRI (T1-GD, T1, T2, and T2-FLAIR), H&E images | 5-fold cv,Regression | 171 | Survival prediction | Gliomas (TCGA-HGG, LGG) | no | |
| ref. ubae014-B88 | MRI (T2, T1-CE, DWI), H&E images | AUC, PPV,NPV | Retrospective study: 933,Prospective study: 100 | Predict pathological complete response | Rectal cancer | no | |
| ref. ubae014-B89 | MRI (T2WI, DWI),H&E images | AUC | 153 | Predict pathological good responses | Rectal cancer | no | |
| ref. ubae014-B77 | CT, H&E images | AUC, accuracy | 171 | Cancer classification | NSCLC | heatmaps | |
| ref. ubae014-B78 | CT, WSIs | AUC, accuracy | 252 | Prediction of LNM stage | Gastric cancer | class activation maps | |
| Pathology,Radiology,Molecular | ref. ubae014-B79 | Radiology (CT), pathology (IHC), Multiomics (somatic mutations, CNAs, gene fusions) | 10-fold cv,AUC | 247 | Immunotherapy response (IHC score, TMB) | NSCLC | no |
| Pathology,Radiology, Clinical | ref. ubae014-B80 | Pathomics, Radiomics features, Immunoscore, Clinical factors | AUC | 103 | Survival prediction | Colorectal cancer (CRC) lung Metastasis | no |
| ref. ubae014-B81 | Pathomics, Radiomics, Clinicopathological features | AUC | 211 | Predicting pathological complete response | Breast cancer | no | |
| Pathology,Molecular,Clinical | ref. ubae014-B82 | WSIs, Multiomics (mRNA, miRNA expr), Clinical | C-index | 11 160 | OS | 20 types (BLCA, BRCA, CESC, COADREAD, HNSC, KICH, KIRC, KIRP, LAML, LGG, LIHC, LUAD, LUSC, OV, PAAD, PRAD, SKCM, STAD, THCA, UCEC) | no |
| ref. ubae014-B83 | WSIs, Multiomics (CNA, somatic genetic mutations), Clinical | AUC | 1281 | Detect malignancy, Diagnose cancer subtypes, Predict survival outcomes | CCRCC | grad-CAM | |
| ref. ubae014-B84 | H&E, Multiomics (genomic and transcriptomic features), Clinical | AUC | 168 | Response to treatment | Breast cancer | no | |
| Pathology, Radiology,Molecular,Clinical | ref. ubae014-B90 | H&E, CT, Multiomics (genomic features), Clinical | C-index | 444 | OS | Ovarian cancer | correlation heatmap |
| ref. ubae014-B85 | H&E, MRI, Multiomics (DNA seq), Clinical | 15-fold Monte Carlo cv, C-index | 176 | OS | Glioma (TCGA-GBM, LGG) | no |
Refer to Table 2 for the list of abbreviations used here.
Table 2.: List of abbreviations used in this article, alphabetically ordered.
| AI | Artificial Intelligence | AP | Anatomical Pathology |
|---|---|---|---|
| AUC | Area Under the Curve | BLCA | Bladder Urothelial Carcinoma |
| BRCA | Breast Invasive Carcinoma | C-Index | Concordance Index |
| CCRCC | Clear Cell Renal Cell Carcinoma | CI | Confidence Interval |
| CNAs | Copy Number Alterations | CNNs | Convolutional Neural Networks |
| CNV | Copy Number Variation | COAD | Colon Adenocarcinoma |
| COADREAD | Colorectal Adenocarcinoma | CP | Clinical Pathology |
| CPATH | Computational Pathology | CPT | Current Procedural Terminologies |
| CPTAC | Clinical Proteomic Tumour Analysis Consortium | CRC | Colorectal Cancer |
| CT | Computed Tomography | CV | Cross Validation |
| DFS | Disease Free Survival | DL | Deep Learning |
| DLRPM | Deep Learning RadioPathomics Model | DNA | Deoxyribonucleic Acid |
| DOF | Deep Orthogonal Fusion | DP | Digital Pathology |
| DPA | Digital Pathology Association | EHR | Electronic Health Records |
| EPLA | Ensembled Patch Likelihood Aggregation | FFPE | Formalin-Fixed Paraffin-Embedded |
| FHIR | Fast Healthcare Interoperability Resources | FLAIR | Fluid Attenuated Inversion Recovery |
| FMs | Foundation Models | FNA | Fine Needle Aspiration biopsy |
| GANs | Generative Adversarial Networks | GBM | Glioblastoma Multiforme |
| GBMLGG | Glioblastoma & Lower Grade Glioma | GNNs | Graph Neural Networks |
| GRU | Gated Recurrent Unit | H&E | Haematoxylin and Eosin |
| HER | Human Epidermal Growth Factor Receptor | HGG | High Grade Gliomas |
| ID | Integrative Diagnostics | ICB | Immune Checkpoint Blockade |
| IHC | Immunohistochemistry | JIF-MMFA | Joint-Individual Fusion with Multi-modal MutualFusion Attention |
| KIRC | Kidney Renal Clear Cell Carcinoma | LGG | Low Grade Gliomas |
| LNM | Lymph Node Metastasis | LSTMs | Long Short-Term Memory Network |
| LUAD | Lung Adenocarcinoma | LUSC | Lung Squamous Cell Carcinoma |
| MCAT | Multimodal Co-Attention Transformer | MIL | Multiple Instance Learning |
| miRNA | Micro-Ribonucleic Acid | ML | Machine Learning |
| MMF | Multimodal Fusion | MMO | Multimodal Orthogonalization |
| MOMA | Multi-Omics Multi-cohort Assessment | MRI | Magnetic Resonance Imaging |
| mRNA | Messenger-Ribonucleic Acid | MSI | Microsatellite Instability |
| nCRT | Neoadjuvant Chemoradiotherapy | NGS | Next-Generation Sequencing |
| NPV | Negative Predictive Value | NSCLC | Non-Small Cell Lung Carcinoma |
| OS | Overall Survival | OV | Ovarian Carcinoma |
| PAAD | Pancreatic Adenocarcinoma | PCR | Polymerase Chain Reaction |
| PD-L1 | Programmed Death-Ligand 1 | PET | Positron Emission Tomography |
| pGR | Pathological Good Response | PI-RADS | Prostate Imaging Reporting And Data System |
| PPV | Positive Predictive Value | QA | Quality Assurance |
| QC | Quality Control | RAPIDS | RAdioPathomics Integrated preDiction System |
| ResNet | Residual Network | RNA-Seq | Ribonucleic Acid Sequence |
| RNNs | Recurrent Neural Networks | SNNs | Self-Normalizing Networks |
| SVM | Support Vector Machine | TCGA | The Cancer Genome Atlas |
| TMB | Tumour Mutation Burden | TME | Tumour Microenvironment |
| UCEC | Uterine Corpus Endometrial Carcinoma | ViT | Vision Transformer |
| WHO | World Health Organization | WSI | Whole Slide Imaging |
2019
Cheerla et al. developed a multimodal model to predict the survival of patients for 20 different cancer types by fusing three different data modalities that included clinical data, mRNA and microRNA expressions, and WSIs.ref. ubae014-B82 The proposed multimodal AI model was able to predict the overall survival (OS) with the concordance index (C-index) of 0.78.ref. ubae014-B82
2020
Chen et al. proposed an end-to-end multimodal framework called “Pathomic Fusion” that predicted survival outcomes from WSIs and genomic (mutations, CNV, RNA-Seq) features of glioma and clear cell renal cell carcinoma (CCRCC).ref. ubae014-B69 For the glioma patients, Pathomic Fusion outperformed the WHO paradigm and previous state-of-the-art with 6.31% and 5.76% improvements, respectively, and reaching a C-index of 0.826.ref. ubae014-B69 For the CCRCC, similar results were observed, with tri-modal Pathomic Fusion achieving a C-index of 0.720.ref. ubae014-B69
Microsatellite Instability (MSI) is an approved pan-cancer biomarker for immune checkpoint blockade (ICB) therapy. A multiple-instance AI model, called Ensembled Patch Likelihood Aggregation (EPLA), was proposed to integrate pathological, genomic, and transcriptomic phenotypes and predict MSI.ref. ubae014-B70 The model also provided an interpretation of its results for effective use in ICB therapy. The proposed EPLA model got an AUC of 0.8848 and 0.8504 in The Cancer Genome Atlas (TCGA)-COAD and Asian-CRC external validation, respectively.ref. ubae014-B70 EPLA was also able to capture correlation between poor differentiation and MSI.ref. ubae014-B70 The model was also able to identify imaging signatures associated with mutations burden, DNA damage, and antitumour immunity.ref. ubae014-B70
Rathore et al. used support vector machine (SVM) model to distinguish high-grade gliomas (HGG) from low-grade gliomas (LGG) by combining textual features with conventional imaging and clinical markers.ref. ubae014-B73 Their model identified the microvascular proliferation level, mitotic activity, presence of necrosis, and nuclear atypia in WSIs.ref. ubae014-B73 The texture features were successfully validated on the glioma patients in 10-fold cross-validation (accuracy = 75.12%, AUC = 0.652). Adding texture features to clinical and conventional imaging features significantly improved grade prediction compared to the models trained on clinical and conventional imaging features alone.ref. ubae014-B73
Classical ML models have been shown to capture the multi-scale association between H&E and CT images, which was used to classify lung adenocarcinoma (LUAD) from squamous cell carcinoma in NSCLC.ref. ubae014-B77 Alvarez et al. identified discriminant pathomic patterns for cancer classification and established correlations between multimodal features of 171 patients, depicting significant cross-scale associations between cell density statistics and CT intensity.ref. ubae014-B77
2021
The Multimodal Co-Attention Transformer (MCAT), an AI model leveraging the Transformer architecture and cross-modality attention, has outperformed all unimodal models and the more commonly used late fusion methods in survival prediction for 5 distinct types of cancers, including Bladder Urothelial Carcinoma (BLCA), Breast Invasive Carcinoma (BRCA), Glioblastoma & Lower Grade Glioma (GBMLGG), LUAD, and Uterine Corpus Endometrial Carcinoma (UCEC).ref. ubae014-B18 MCAT outperformed state-of-the-art methods on all benchmarks, with a C-index increase of 6.35% and 12.4% compared to Attention Multi-Instance Learning (MIL) and Deep Attention Multiple Instance Survival Learning (DeepAttn-MISL) methods, respectively. The success of the MCAT model indicated that multimodal learning could be used in general for any survival outcome prediction task.ref. ubae014-B18
Khosravi et al. proposed AI-biopsy, an AI model based on the Inception-V1 architecture, which was pretrained on a non-medical ImageNet dataset and transfer learned on MRI scans labelled with pathology assessments like Gleason scores and grade groups.ref. ubae014-B75 The proposed early data-fusion technique exhibited a higher agreement with biopsy results when compared to the Prostate Imaging Reporting and Data System (PI-RADS) score, indicating promising advancements in diagnostic precision.ref. ubae014-B75 AI-biopsy achieved AUCs of 0.78 (95% CI: [0.74-0.82]) and 0.89 (95% CI: [0.86-0.92]) in 400 patients on the tasks of distinguishing high-risk from low-risk cancers and cancerous vs benign prostate disease, respectively.
WSI, demographic, genomic, and clinical data collected from 3 data sources, TCGA, Clinical Proteomic Tumour Analysis Consortium (CPTAC), and Brigham and Women’s Hospital, were used to develop an AI model for various tasks related to renal cell carcinoma.ref. ubae014-B83 The trained models were able to detect malignancy (AUC = 0.964-0.985), diagnose renal cancer subtypes (AUCs = 0.953-0.993), and predict survival outcomes.ref. ubae014-B83 The model also identified histopathology image features that were indicative of biomarkers such as copy-number alterations (CNAs) and tumour mutation burden (TMB).ref. ubae014-B83
In general, single-omics data modality may not be able to provide a comprehensive view of tumour-related molecular alterations. In such cases, integrating multiple data modalities even within the -omics regime can help improve diagnosis, personalized treatments, and predict outcomes more accurately.ref. ubae014-B86 Tong et al. developed an encoder-decoder AI model for cancer classification among 4 cancer types [LUAD, Kidney Renal Clear Cell Carcinoma (KIRC), Lung Squamous Cell Carcinoma (LUSC), and pancreatic adenocarcinoma (PAAD)] and patient survival prediction in 2 cancer types (BRCA, Ovarian Carcinoma [OV]).ref. ubae014-B86 Two data integration methods were evaluated for 4 types of -omics data that include gene expression, DNA methylation, miRNA expression, and copy number variations.ref. ubae014-B86 The multimodal AI models outperformed single-omics models for both tasks, cancer classification and survival prediction. Within the 2 data fusion methods, divergence-based consensus networks performed better than concatenation-based integration networks.ref. ubae014-B86
Multimodal learning using clinical data, H&E images, genomics, and transcriptomic features were shown to predict pathological complete response to treatment and improve outcomes in 168 breast cancer patients treated with chemotherapy with or without HER2-targeted therapy before surgery.ref. ubae014-B84 The multiomic AI model predicted pathological complete response in an external validation cohort of 75 patients with an AUC of 0.87. The final AI model consisted of an ensemble of traditional ML models, including logistic regression, SVM, and random forest.ref. ubae014-B84
Chelebian et al. developed an ensemble CNN to provide interpretations that matched pathologists’ manual assessments.ref. ubae014-B71 The resulting interpretations were more informative when the underlying genes were considered.ref. ubae014-B71 The framework paired H&E slides with spatially resolved gene expressions from the spatial transcriptomics technique, predicting the spatial variation of individual genes.ref. ubae014-B71
Braman et al.ref. ubae014-B85 proposed a Deep Orthogonal Fusion (DOF) model to learn from multiparametric MRI scans, H&E slides, DNA sequencing, and clinical features into a multimodal risk score.ref. ubae014-B85 The embeddings (or representations) from individual data modalities were learned and fused through attention-gated tensor fusion. The authors introduced multimodal orthogonalization (MMO) learning loss that incentivized each embedding to be more complementary.ref. ubae014-B85 DOF predicted OS in 176 glioma patients with a median C-index of 0.788 ± 0.067, compared to the best-performing unimodal model (C-index of 0.718 ± 0.064).ref. ubae014-B85
2022
Compared with unimodal data, combining H&E WSIs with the molecular assay improved survival prediction reliability in multiple cancers, including BRCA, COADREAD, LUAD, PAAD, and others.ref. ubae014-B68 Chen et al. proposed a deep learning framework, multimodal fusion (MMF), which achieved an overall C-index of 0.644 across the 14 cancer types, whereas the MIL model using WSI and SNN on molecular data had overall C-Indices of 0.578 and 0.606, respectively. On survival AUC, a similar improvement for the multimodal model was reported with an overall performance of 0.662 compared with unimodal models.ref. ubae014-B68
Often referred to as a “virtual biopsy,” radiomics features enhance conventional diagnostic radiologic workflows when combined with the pathomic features from H&E slides.ref. ubae014-B76 Brancato et al. employed different feature generation techniques to extract 91 radiomics features from MRI scans and 156 pathomic features from H&E images for 48 Glioblastoma multiforme (GBM) patients from the CPTAC data portal. The proposed radiomics-pathomics fusion technique showed significant cross-scale associations of features among the GBM patients.ref. ubae014-B76
Multimodal integration of radiology, pathology, and genomics data modalities improved insights into response to immunotherapy PD-L1 blockade in patients with NSCLC.ref. ubae014-B79 Vanguri et al. explored the multimodal integration of CT images, digitized programmed death ligand-1 IHC slides, genomics data including somatic mutations, copy number alterations and gene fusions, and known outcomes to immunotherapy to predict immunotherapy response among 247 patients with advanced NSCLC. The multimodal model called DyAM showed an AUC of 0.80, 95% CI [0.74-0.86] on immunotherapy response prediction task, that outperformed unimodal methods, including TMB (AUC = 0.61, 95% CI [0.52-0.70]) and programmed death ligand-1 IHC score (AUC = 0.73, 95% CI [0.65-0.81]).ref. ubae014-B79
Ensemble of a range of ML regression models, including support vector regression, AdaBoost, gradient boost, and random forest, were shown to provide better survival outcomes prediction for 171 Glioma patients using integrated features from MRI (T1-GD, T1, T2, and T2-FLAIR) and H&E images.ref. ubae014-B87 The proposed ensemble fusion approach outperformed several other techniques based solely on radiology features, gene expression data, or pathology images.ref. ubae014-B87
Combining pretreatment MRI (T2, T1-CE, DWI) and H&E-stained biopsy slides using AI models can predict pathological complete response in patients with locally advanced rectal cancer.ref. ubae014-B88 The proposed framework, RAdioPathomics Integrated preDiction System (RAPIDS), showed a significant improvement in the prediction of pathological complete response with a combined AUC of 0.812 compared to the unimodal models for prospective validation study.ref. ubae014-B88 RAPIDS also performed better in the retrospective study training cohort (AUC 0.868 [95% CI 0.825-0.912]), validation cohort 1 (0.860 CI [0.828-0.892]), and validation cohort 2 (0.872 CI [0.810-0.934]).ref. ubae014-B88
Harnessing AI-driven insights with radiomics–pathomics has led to personalized treatment plans, optimizing therapies while minimizing adverse effects.ref. ubae014-B80 An innovative fusion of pathomics, radiomics, and immunoscore data identified high-risk patients with colorectal cancer (CRC) lung metastasis. The combined nomogram model predicted the OS with an AUC of 0.860 and disease-free survival (DFS) with an AUC of 0.875. This integration offers insights about aggressive treatment strategies and tailored follow-up plans for high-risk patients.ref. ubae014-B80
Wan et al. developed a multiscale framework that harnessed the power of radiomics features extracted from MRI (T2WI, DWI) data, pathomics features derived from H&E images, and clinical variables to predict pathological good responses (pGR) in 153 patients with locally advanced rectal cancer who underwent neoadjuvant chemoradiotherapy (nCRT).ref. ubae014-B89 This macroscopic–microscopic scale fusion identified the patients with down-staging to stage ypT0-1N0 after nCRT, a critical factor in guiding organ-preserving strategies to minimize invasion and uphold the quality of life. By integrating these diverse modalities, the multiscale AI model outperformed the traditional clinical-radiological model with an AUC of 0.90 (95% CI: [0.78-1.00]) compared to 0.68 (95% CI: [0.46-0.91]) in the validation group. These results highlight the complementary nature of information present in radiomics and pathomics features for assessing treatment response.ref. ubae014-B89
Boehm et al. used a multimodal dataset of 444 patients with ovarian cancer and integrated radiological data (CT scans), histopathological images (H&E), clinical, and genomic modalities to stratify patients based on risk.ref. ubae014-B90 The multimodal AI model stratified patients in the training and the test cohorts with the C-indices of 0.55 (95% CI: [0.549-0.554]) and 0.53 (95% CI: [0.517-0.547]), respectively. The authors also discovered quantitative features, such as tumour nuclear size on H&E and omental texture on CT, associated with prognosis.ref. ubae014-B90
2023
Palmal et al. employed a graph convolutional network (GCN) and Choquet fuzzy ensemble consisting of classical ML models (Logistic Regression, Random Forest, and SVM) to fuse the multiomics and clinical data.ref. ubae014-B91 1980 samples of breast cancer patients were classified as short-term or long-term survivors using different combinations of gene expression (24 000 features), copy number alteration (26 000 features), and clinical data (27 features).ref. ubae014-B91 The performance metrics of accuracy, Matthews correlation coefficient, precision, sensitivity, specificity, balanced accuracy, and F1-measure showed significant result scores as 0.820, 0.528, 0.630, 0.666, 0.871, 0.769, and 0.647, respectively.ref. ubae014-B91
Tsai et al. developed a Multi-omics Multi-cohort Assessment (MOMA) framework to fuse WSIs with genomics, proteomics, and molecular data to predict OS, progression-free survival, and other variables for colorectal cancer patients.ref. ubae014-B72 MOMA was able to identify cross-scale relationships among histologic patterns, multiomics, and clinical profiles in 1888 patients. MOMA predicted OS, progression-free survival, and MSI in held-out test datasets of stage I, II, and III colorectal cancer patients better than other state-of-the-art methods.ref. ubae014-B72
An AI model that fuses CT images with WSIs was shown to accurately predict lymph node metastasis stage (LNMs) in patients with gastric cancer.ref. ubae014-B78 Features extracted from CT images using a CNN (ResNet-50) and from WSI using the Vision Transformer model (ViT) were fused to learn the prediction task of 5 LNM stages.ref. ubae014-B78 The proposed radiopathologic integrated method achieved 84.0% accuracy and significantly improved AUCs in predicting five LNM stages compared to single-modality models.ref. ubae014-B78 The integrated model also achieved an AUC of 0.978 (95% CI: [0.912-1.0]) in metastasis prediction task.ref. ubae014-B78
An AI model called Deep Learning Radio-Pathomics Model (DLRPM) seamlessly integrated radiomics, pathomics, clinical, and pathological features using ResNet-50 and SVM on 211 breast cancer patients who underwent neoadjuvant chemotherapy.ref. ubae014-B81 DLRPM demonstrated remarkable capabilities for predicting pathological complete response in the validation dataset with an AUC of 0.927 (95% CI: [0.858-0.996]), significantly outperforming models that used individual data modalities and features, such as radiomics and pathomics.ref. ubae014-B81
The fusion of radiomics features with pathomics features and molecular data may provide a more comprehensive view of the intricate tumour microenvironment (TME), enabling a deeper understanding of its composition and tumour heterogeneity.ref. ubae014-B92 Fusing radiological and pathomics data may offer precise spatial mapping of histological features, facilitating detailed TME examination. This holistic approach may unravel the complex interactions between tumour cells and the surrounding microenvironment, advancing our understanding of cancer biology.ref. ubae014-B92
Recently, Tang et al. designed a new fusion method to combine dermatological images and patient clinical metadata for skin cancer classification using three public skin lesion datasets with more than 3300 patients.ref. ubae014-B74 The proposed AI model, called joint-individual fusion with multimodal mutual fusion attention (JIF-MMFA), learns the shared features of multimodal data by employing a fusion attention module to enhance the relevant features from both data modalities.ref. ubae014-B74 JIF-MMFA showed the effectiveness of using patients’ clinical data in improving the classification of skin cancer.ref. ubae014-B74
Discussion
The global market for AI-based DP solutions is projected to grow at a compound annual growth rate of 8.59% from 2024 to 2031, with North America leading due to advanced healthcare infrastructure, higher income levels, and research facilities.ref. ubae014-B93 AI patent filings in DP have sharply increased in major patent offices, indicating growing innovation.ref. ubae014-B94 There is a significant push towards open-source and reproducible AI, with initiatives like Allen AI contributing through platforms like Hugging Face.ref. ubae014-B95 AI is increasingly used to predict patient responses to immunotherapy by analysing histopathological and molecular data.ref. ubae014-B96 The integration of AI in DP promotes collaboration among pathologists, oncologists, radiologists, and bioinformaticians, leading to a holistic approach to patient care.ref. ubae014-B97
Large language models (LLMs) like ChatGPT enhance pathology reports by ensuring consistency and translating complex medical data for clinicians and patients. LLMs assist pathologists by providing quick access to relevant literature and summarizing research findings.ref. ubae014-B8,ref. ubae014-B98 While most multimodal works focus on 2D WSIs, incorporating 3D data from techniques like microscopy or medical imaging could provide additional spatial context, improving model performance.ref. ubae014-B99 Combining WSIs with text reports, genomic data, or medical videos in multimodal learning is emerging as a way to enhance diagnostic capabilities.ref. ubae014-B100 Models like CLIP, CONCH, which integrate text and image data, can be adapted for DP tasks.ref. ubae014-B55 Integrating histopathological, radiomic, genomic, and clinical data using a late-fusion approach has been shown to improve cancer patient risk stratification.ref. ubae014-B90 Multimodal AI is used across various domains, including biomedicine for disease diagnosis and treatment planning, autonomous vehicles for real-time decision-making, and augmented generative AI for content creation and multimodal translation.ref. ubae014-B101 Below we discuss some of the opportunities and challenges of multimodal learning in DP.
Opportunities and challenges
Opportunities
Multimodal patient stratification: Integrating various data types, including genomics, transcriptomics, epigenomics, proteomics, histopathology, and radiology, allows more precise patient stratification.ref. ubae014-B17 AI models can identify distinct patient subgroups with different phenotypes and treatment responses. This stratification can lead to more personalized and effective treatment strategies, enhancing patient care.
Non-invasive alternatives, personalized medicine: AI models enable the prediction of histology subtypes or grades from radiomics features.ref. ubae014-B106 This capability will open many opportunities for developing non-invasive surrogates for existing biomarkers, reducing the need for invasive procedures in the future. Additionally, AI models demonstrate promise in predicting clinical outcomes, including survival, treatment response, and recurrence, enabling personalized medicine approaches and better patient management.
Patient empowerment: Integrating different modalities can create user-friendly digital solutions that empower patients to become active partners in managing their health.ref. ubae014-B56 Patients can access their own health records, diagnostic reports, and integrated biomarker information, allowing them to make informed decisions about their care. This level of patient engagement and access to integrated health information can enhance the patient experience, improve communication with healthcare providers, and improve health outcomes.
Improving outcomes: Multimodal data integration, especially fusing histopathology and genomics data, has shown promise in improving the accuracy of outcome predictions for cancer patients.ref. ubae014-B17 Researchers and clinicians can improve overall/progression-free survival analysis by leveraging multimodal AI techniques, enabling more precise prognostication beyond traditional clinical factors. This can facilitate the development of more effective treatment regimens for high-risk patients and improve treatment outcomes.
Morphological associations, biomarker discovery: Multimodal AI models can potentially discover morphological associations spread across data modalities, including the relationship between genetic mutations and tissue morphology.ref. ubae014-B106 Such findings can help exploratory studies and reduce the search space for potential biomarker candidates. By identifying morphological associates, AI can support cost-efficient biomarker discovery and aid in cancer screening and therapeutic target discovery.
Integrative diagnostics: Integrative diagnostics (ID) is defined to be the convergence of radiology and pathology imaging, clinical laboratory medicine, and advanced computational methods.ref. ubae014-B107 ID, especially with a focus on radiology and pathology data fusion, offers many benefits and untapped potential in oncology. Yet, despite the anticipation of numerous benefits, a noticeable scarcity of comprehensive literature surrounds this emerging field. ID can potentially identify appropriate therapies and suggest treatment changes when needed, decreasing morbidity and costs and improving patient outcomes.ref. ubae014-B107
Correlational studies: Multimodal learning may hold enormous potential for conducting correlational studies and improving oncology patient diagnoses. Using histopathological and molecular data, AI models have already been shown to identify novel biomarkers and underlying tuberculosis mechanisms in susceptible individuals.ref. ubae014-B108
Medical report integration: Despite recent advancements related to digitizing medical data in EHR and radiological and laboratory imaging systems, there are apparently no efforts focused on correlating the imaging data (histopathology and radiology) with the textual reports (eg, radiology reports, pathology reports, and other clinical notes) at the data storage level. One obvious reason is related to the siloed storage and data management systems. Modern AI systems based on vector databases can provide the necessary building blocks for data integration at the storage level.ref. ubae014-B109
Challenges
Data availability, integration, missingness: Obtaining and integrating diverse medical data types, such as images and clinical records, is an extremely challenging task.ref. ubae014-B110 Data security and governance issues, data scarcity, unstructured formats, and varying data identifiers pose significant hurdles in this process.ref. ubae014-B17,ref. ubae014-B92 Moreover, the issue of missing or incomplete data within each data modality further complicates the training of AI models.ref. ubae014-B106,ref. ubae014-B110 Strategies like synthetic data generation, dropout-based methods, and embedding models are being explored to address this challenge, but concerns about the accuracy and representativeness of synthetic data still remain.ref. ubae014-B62,ref. ubae014-B64
Interoperability: Ensuring that different medical devices, systems, and data formats can seamlessly communicate and exchange information is a considerable challenge.ref. ubae014-B56,ref. ubae014-B91 Data standards, protocols, and technologies differ across vendors, providers, and sometimes healthcare institutions and systems. Efforts such as Fast Healthcare Interoperability Resources (FHIR) for the EHR to overcome this challenge are essential for integrating diverse modalities and facilitating comprehensive patient care.
Data modelling and analysis: This challenge arises when integrating data from different sources or institutions because of institution-specific biases stemming from variations in data acquisition methods, clinical data standards, and the catchment area demographics.ref. ubae014-B17 Ensuring model generalizability in the presence of heterogeneous data modalities is another aspect of this challenge.ref. ubae014-B17
Transparency and clinical adoption: While multimodal AI models hold great promise in oncology, their adoption in clinical practice faces several hurdles. Ensuring transparency and interpretability of these models is challenging, as they often use abstract feature representations.ref. ubae014-B106 Prospective clinical trials and rigorous validation studies will be needed to demonstrate the added value of AI models in real-world clinical settings. Regulatory guidelines and addressing issues like fairness and dataset shifts are equally important for clinical adoption.ref. ubae014-B106
Personnel training: The primary users of AI systems in DP are clinicians and pathologists. Training the administrative, technical, and professional personnel on AI systems is a secondary but essential challenge that needs to be addressed for the success of AI systems.ref. ubae014-B111
Explainability, trust, reproducibility, and output visualization of AI models: AI models are known to be complex; this impacts the ability of pathologists and clinicians to understand the internal workings of these models.ref. ubae014-B65 Similarly, the reproducibility of model output and performance is another challenge that makes their adoption difficult. Having an interactive graphical user interface is essential in AI systems, but such interfaces are either not available entirely or do not provide output visualizations for the intermediate layers of the model. All these challenges reduce users’ trust in AI systems, especially in the case of DP, where decisions directly impact diagnosis and clinical outcomes.ref. ubae014-B111
Conclusion
The integration of multimodal oncology data using advanced computational methods represents a pivotal advancement in the realm of DP and CPATH. The advancements in AI and ML models have enabled a more nuanced understanding of cancer’s multifaceted nature. By synergistically combining clinical, radiological, pathology, and molecular data, researchers and clinicians can gain unprecedented insights into the complex dynamics of cancer, leading to more effective and personalized therapeutic strategies. However, there are multiple associated challenges to multimodal learning. Looking forward, DP stands on the brink of a new era where continuous innovation and interdisciplinary collaboration are key to harnessing the full potential of multimodal data integration. This will undoubtedly be crucial in advancing our understanding of cancer and opening new avenues for patient care and research.
Box 1.: Key pathology terminologies.
| Pathology | Stems from the Greek word pathologia, meaning the study of suffering, it is a medical specialty investigating the origin, progression, and alterations in the structure, function, and natural course of diseases.ref. ubae014-B7 There are two main branches of pathology: anatomical pathology (AP) focuses on studying tissue samples to diagnose diseases such as cancer, and clinical pathology (CP) analyses bodily fluids and conducts laboratory tests.ref. ubae014-B8 Molecular pathology is an emerging branch that studies diseases at the molecular level by examining tissue or body fluids. |
| Histology | Microscopic examination of tissues and organs by sectioning, applying stains, and observing the prepared sections under a microscope. This process enables the visualization of tissue structures and any distinctive alterations the tissue might have undergone. Haematoxylin (a basic dye that stains cell nuclei in a purplish blue colour) and Eosin (an acidic dye that stains cytoplasm in a pinkish red colour), referred to as H&E, are commonly used together to stain structures of the cells to define intracellular organelles and proteins.ref. ubae014-B9 |
| Histopathology | It extends the utility of histological analysis by focusing on the study of disease indicators through the microscopic examination of processed and fixed glass slides containing sections from biopsies or surgical specimens. To visualize various tissue components under a microscope, these specimens are dyed with one or more stains depending on the investigational question being asked.ref. ubae014-B9,ref. ubae014-B10 |
| Immunohistochemistry (IHC) | A valuable tool extensively employed in pathology for cell classification and diagnosis. Commonly performed on formalin-fixed paraffin-embedded (FFPE) tissue, IHC targets specific antigens in tissues and cells using antibodies to determine cell type and organ of origin. IHC usage has lately expanded to study predictive and prognostic biomarkers in oncological settings, making it a useful technique in modern pathology practice.ref. ubae014-B11 |
| WSI | The process of digitizing glass slides at multiple magnifications and focal levels to create digital images for observation and image analysis. WSI aims to replicate the experience of traditional light microscopy through digital means and is often known as virtual microscopy.ref. ubae014-B12 |
| Cytology slides | These refer to glass slides that contain specimens of individual cells or cell clusters obtained from various bodily fluids, tissues, or fine needle aspiration (FNA) biopsy. Unlike histology, which focuses on the function of tissues, cytology slides are prepared for microscopic examination to analyse cellular morphology, structure, and characteristics.ref. ubae014-B13 |
| Digital pathology (DP) | DP includes converting histopathology, IHC, or cytology slides into digital formats through whole-slide scanners, followed by applying computer-aided analytical tools for objective analysis. DP has improved pathology diagnoses through multiple techniques as discussed in this review. |
| CPATH | CPATH is the sub-branch of DP that involves computational analysis of digital slides. CPATH aims to analyse patient specimens by extracting information from digitized pathology images in combination with their associated meta-data, typically using AI methods to gain valuable insights into disease processes.ref. ubae014-B6,ref. ubae014-B8,ref. ubae014-B14 |
| Radiomics | Radiomics is a sub-field of radiology, where radiological images are converted into high-dimensional mine-able data for AI-based analysis to uncover biomarkers for diagnosis, prognosis, and treatment response prediction.ref. ubae014-B15 |
| Pathomics | Pathomics is a sub-field of pathology, that covers a wide variety of data captured from image analyses in the form of features that describe multiple phenotypic features of tissues in WSIs. Using AI, pathomics provides a quantitative assessment of identified structures and features to complement traditional histopathologic evaluation by pathologists.ref. ubae014-B16 |
| AI-qualified individuals | In the DP/CPATH context, individuals who not only understand AI at a basic technical level but also have the clinical, regulatory, and practical knowledge to apply AI effectively in pathology. This includes clinical validation, understanding regulatory and ethical implications, and the ability to implement, use, and interpret AI tools and their outcomes in daily practice. While it is optional but valuable that these individuals may also be involved in developing new AI tools, the term more broadly encompasses those who are well-equipped to work with AI in a clinical environment, ensuring its accurate, safe, and effective use. |
Box 2.: Key AI terminologies.
| AI | Initially started as a simple theory of human intelligence being exhibited by machines, AI aims to simulate human intelligence in machines by developing computers that can engage in learning, reasoning, self-correction, problem-solving, and decision-making.ref. ubae014-B37 |
| ML | ML is a subset of AI that programs computers to process natural data and solve an optimization problem using data samples or past experience. It is an engineering discipline that is closely related to mathematical statistics to build learning models.ref. ubae014-B38 |
| Deep Learning (DL) | DL is a subfield of ML and refers to models developed using multiple layers of artificial neural networks. The deep neural networks focus on compositional learning using multiple (hence the word “deep”) layers of representations, where each layer is built using multiple artificial neurons.ref. ubae014-B38 Various types of DL models have been developed, including Convolutional Neural networks (CNNs), Recurrent Neural networks (RNNs), Graph Neural Networks (GNNs), Transformers, and many others. |
| Convolutional Neural Networks | CNNs are specialized DL models for analysing image or grid data.ref. ubae014-B38,ref. ubae014-B43,ref. ubae014-B44 CNNs use convolution operations and can learn spatial correlations, features, and relationships from the input data.ref. ubae014-B43,ref. ubae014-B45 |
| Recurrent Neural Networks | RNNs are a family of DL models used for processing sequential data like speech, text, or time-series data (eg, patient vital signs recorded over time).ref. ubae014-B43,ref. ubae014-B46,ref. ubae014-B47 RNNs are used to model time/space-based relationships between sequences of input data and to learn patterns based on the order (in time or space).ref. ubae014-B43 Sub-categories of RNNs called Long short-term memory network (LSTM) and gated recurrent unit (GRU) models are better at learning long-term dependencies, making them great for real-world use cases, eg, predicting medical events and detecting anomalies in time series data.ref. ubae014-B46 |
| Graph Neural Networks | GNNs are DL models designed for analysing graph data, like social relationships or biological pathways.ref. ubae014-B40 GNNs pass messages between neighbouring nodes to aggregate local and global information. By modelling these relationships, GNNs can make predictions about nodes, edges, and sub-graphs in the network, where each entity may represent patients, drugs, proteins, etc GNNs enable new insights in fields like precision medicine and drug discovery.ref. ubae014-B48 |
| Transformers | Transformers are DL models that have shown promising results on language translation and image recognition tasks.ref. ubae014-B51,ref. ubae014-B52 Their key innovation is a technique called self-attention. Unlike RNNs, which process data sequentially, self-attention allows Transformers to learn relationships between any parts of the input, regardless of their position in the sequence. For example, a Transformer can learn that the symptoms “cough” and “fever” are strongly related even if they are distant in the patient’s medical history. The model learns which connections are important by assigning an “attention” score between input sequences (also referred to as tokens).ref. ubae014-B47,ref. ubae014-B51,ref. ubae014-B52 |
| Foundation Models (FMs) | FMs are a new class of DL models characterized by their large scale and ability to adapt to new tasks.ref. ubae014-B53 FMs are trained on extensive datasets, allowing them to develop a broad understanding of various topics.ref. ubae014-B8,ref. ubae014-B54 The key characteristics of FMs include scalability in learning from diverse data sources, multimodal learning, compositionally in terms of generalization, and the emergence of implicit learning. Transfer learning, adaptation, in-context learning, and fine-tuning make FMs exceptionally useful across numerous fields, including the medical domain.ref. ubae014-B55 A recent review paper covers existing FMs and their prospective use in DP.ref. ubae014-B8 |
| Multimodal Learning | Multimodal Learning involves integrating data from multiple modalities for an ML task, eg, prediction, classification, or regression. A modality refers to specific data input or source types, such as text, images, audio, video, or other data. The core idea is to leverage the complementary information from different modalities to improve the performance and accuracy of learning algorithms. Multimodal learning aims to achieve a more comprehensive understanding of the problem, leading to better decision-making than using a single modality.ref. ubae014-B19,ref. ubae014-B56 |
References
- A practical guide to whole slide imaging: a white paper from the digital pathology association.. Arch Pathol Lab Med., 2019. [PubMed]
- Introduction to digital image analysis in whole-slide imaging: a white paper from the Digital Pathology Association.. J Pathol Inform., 2019. [PubMed]
- Deep learning-enabled virtual histological staining of biological samples.. Light Sci Appl., 2023. [PubMed]
- 4 Butz H , PatócsA. Brief summary of the most important molecular genetic methods (PCR, qPCR, microarray, next-generation sequencing, etc.). In: Igaz P, Patócs A, eds. Genetics of Endocrine Diseases and Syndromes. Vol. 111. Springer, Cham; 2019:33-52. 10.1007/978-3-030-25905-1_4
- Technical challenges of enterprise imaging: HIMSS-SIIM collaborative white paper.. J Digit Imaging., 2016. [PubMed]
- Computational pathology definitions, best practices, and recommendations for regulatory guidance: a white paper from the digital pathology association.. J Pathol., 2019. [PubMed]
- 7 Funkhouser WK. Pathology: the clinical description of human disease. Essential Concepts in Molecular Pathology. Elsevier; 2020:177-190.
- Revolutionizing digital pathology with the power of generative artificial intelligence and foundation models.. Lab Invest., 2023. [PubMed]
- 9 Gurina TS , SimmsL. Histology, staining. In: StatPearls. StatPearls Publishing; 2022. http://europepmc.org/books/NBK557663
- Histopathological image analysis: a review.. IEEE Rev Biomed Eng. 2009;2:147-171.
- 11 Magaki S , HojatSA, WeiB, SoA, YongWH. An introduction to the performance of immunohistochemistry. In: Yong W, ed. Biobanking: Methods and Protocols. Humana Press, New York, NY; 2019:289-298. 10.1007/978-1-4939-8935-5_25
- Review of the current state of whole slide imaging in pathology.. J Pathol Inform., 2011. [PubMed]
- Digital cytology part 2: artificial intelligence in cytology a concept paper with review and recommendations from the American Society of Cytopathology Digital Cytology Task Force.. J Am Soc Cytopathol., 2024. [PubMed]
- Application of digital pathology-based advanced analytics of tumour microenvironment organisation to predict prognosis and therapeutic response.. J Pathol., 2023. [PubMed]
- Images are data: challenges and opportunities in the clinical translation of radiomics.. Cancer Res., 2022. [PubMed]
- The emergence of pathomics.. Curr Pathobiol Rep., 2019
- Harnessing multimodal data integration to advance precision oncology.. Nat Rev Cancer., 2022. [PubMed]
- 18 Richard J , ChenMY, LuW-H, et al Multimodal co-attention transformer for survival prediction in gigapixel whole slide images. In: Proceedings of the IEEE/CVF International Conference on Computer Vision, Montreal, QC, Canada. IEEE; 2021:4015-4025. 10.1109/ICCV48922.2021.00398
- Multimodal data integration for oncology in the era of deep neural networks: a review.. Front Artif Intell., 2024. [PubMed]
- Moffitt cancer center: why we are building the first machine learning department in oncology.. Cancer Lett., 2021
- Digital pathology and artificial intelligence in translational medicine and clinical practice.. Mod Pathol., 2022. [PubMed]
- Deep learning in digital pathology for personalized treatment plans of cancer patients.. Semin Diagn Pathol., 2023. [PubMed]
- Artificial intelligence in histopathology: enhancing cancer research and clinical oncology.. Nat Cancer., 2022. [PubMed]
- Role of AI and digital pathology for colorectal immuno-oncology.. Br J Cancer., 2023. [PubMed]
- Federated learning for computational pathology on gigapixel whole slide images.. Med Image Anal., 2022. [DOI | PubMed]
- Digital pathology: Transforming diagnosis in the digital age.. Cureus., 2023. [PubMed]
- Twenty years of digital pathology: an overview of the road travelled, what is on the horizon, and the emergence of vendor-neutral archives.. J Pathol Inform., 2018. [PubMed]
- The status of digital pathology and associated infrastructure within Alzheimer’s disease centers.. J Neuropathol Exp Neurol., 2023. [PubMed]
- The digital slide archive: a software platform for management, integration, and analysis of histology for cancer research.. Cancer Res., 2017. [PubMed]
- An image analysis resource for cancer research: PIIP—Pathology image informatics platform for visualization, analysis, and management.. Cancer Res., 2017. [PubMed]
- A containerized software system for generation, management, and exploration of features from whole slide tissue images.. Cancer Res., 2017. [PubMed]
- Qupath: Open source software for digital pathology image analysis.. Sci Rep., 2017. [PubMed]
- Artificial intelligence in dermatopathology: diagnosis, education, and research.. J Cutan Pathol., 2021. [PubMed]
- A narrative review of digital pathology and artificial intelligence: focusing on lung cancer.. Transl Lung Cancer Res., 2020. [PubMed]
- 35 College of American Pathologists. Digital Pathology Codes. College of American Pathologists; June 2023.
- Multi-modality artificial intelligence in digital pathology.. Brief Bioinform., 2022. [PubMed]
- Artificial intelligence: definition, trends, techniques, and cases.. Artif Intell., 2009
- 38 Chollet F. Deep Learning with Python. Simon and Schuster; 2021.
- Brain tumor segmentation and surveillance with deep artificial neural networks.. Deep Learning for Biomedical Data Analysis., 2021
- Exploring robust architectures for deep artificial neural networks.. Commun Eng., 2022
- Failure detection in deep neural networks for medical imaging.. Front Med Technol., 2022. [DOI | PubMed]
- 42 Khan H , BouaynayaNC, RasoolG. The importance of robust features in mitigating catastrophic forgetting. In: 2023 IEEE Symposium on Computers and Communications (ISCC). IEEE; 2023:752-757.
- 43 Goodfellow I , BengioY, CourvilleA. Deep Learning. MIT Press; 2016.
- Intelligent helipad detection and (grad-cam) estimation using satellite imagery.. Transportation Research Board., 2021
- 45 Dera D , RasoolG, BouaynayaNC, et al Bayes-SAR net: Robust SAR image classification with uncertainty estimation using Bayesian convolutional neural network. In: 2020 IEEE International Radar Conference (RADAR). IEEE; 2020:362-367.
- TRustworthy uncertainty propagation for sequential time-series analysis in RNNs.. IEEE Trans Knowl Data Eng. 2024;. [DOI]
- Transformers in time-series analysis: a tutorial.. Circuits Syst Signal Process., 2023
- 48 Adnan M , KalraS, TizhooshHR. Representation learning of histopathology images using graph neural networks. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops, Seattle, WA, USA. IEEE; 2020:4254-4261. 10.1109/CVPRW50498.2020.00502
- 49 Sureka M , PatilA, AnandD, SethiA. Visualization for histopathology images using graph convolutional neural networks. In: 2020 IEEE 20th International Conference on Bioinformatics and Bioengineering (BIBE). IEEE; 2020:331-335.
- 50 Wang J , ChenRJ, LuMY, BarasA, MahmoodF. Weakly supervised prostate TMA classification via graph convolutional networks. In: 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI), IEEE; 2020:239-243.
- Attention is all you need.. Adv Neural Inf Process Syst., 2017
- 52 Dosovitskiy A , BeyerL, KolesnikovA, et al An image is worth 16×16 words: transformers for image recognition at scale. International Conference on Learning Representations; 2021. https://openreview.net/forum?id=YicbFdNTTy
- 53 Bommasani R , HudsonDA, AdeliE, et al On the opportunities and risks of foundation models. arXiv preprint arXiv:2108.07258. 2021, 10.48550/arXiv.2108.07258, preprint: not peer reviewed.
- 54 Lu MY , ChenB, WilliamsonDFK, et al A Multimodal Generative AI Copilot for Human Pathology. Nature 2024. 10.1038/s41586-024-07618-3
- 55 Lu MY , ChenB, WilliamsonDFK, et al A visual-language foundation model for computational pathology. Nat Med. 2024;30(3):863-874.
- Multimodal analysis and the oncology patient: creating a hospital system for integrated diagnostics and discovery.. Comput Struct Biotechnol J., 2023. [DOI | PubMed]
- 57 Achiam J , AdlerS, AgarwalS, et al Gpt-4 technical report. arXiv preprint arXiv:2303.08774. 2023, 10.48550/arXiv.2303.08774, preprint: not peer reviewed.
- 58 Radford A , KimJW, HallacyC, et al Learning transferable visual models from natural language supervision. In: International Conference on Machine Learning. PMLR; 2021:8748-8763.
- 59 Singh A , HuR, GoswamiV, et al Flava: a foundational language and vision alignment model. In: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, New Orleans, LA, USA. IEEE; 2022:15617-15629. 10.1109/CVPR52688.2022.01519
- 60 Saab K , TuT, WengW-H, et al Capabilities of gemini models in medicine. arXiv preprint arXiv:2404.18416. 2024, 10.48550/arXiv.2404.18416, preprint: not peer reviewed.
- 61 Hartsock I , RasoolG. Vision-language models for medical report generation and visual question answering: a review. arXiv preprint arXiv:2403.02469. 2024, 10.48550/arXiv.2403.02469, preprint: not peer reviewed.
- SeNMo: a self-normalizing deep learning model for enhanced multi-omics data analysis in oncology.. arXiv e-prints., 2024. [DOI]
- Methods for nuclei detection, segmentation, and classification in digital histopathology: a review—current status and future potential.. IEEE Rev Biomed Eng. 2014;7
- 64 Tripathi A , WaqasA, YilmazY, RasoolG. Honeybee: a scalable modular framework for creating multimodal oncology datasets with foundational embedding models. arXiv preprint arXiv:2405.07460. 2024, 10.48550/arXiv.2405.07460, preprint: not peer reviewed.
- Robust explainability: a tutorial on gradient-based attribution methods for deep neural networks.. IEEE Signal Process Mag., 2022
- 66 Tavolara TE , SuZ, GurcanMN, NiaziMKK. One label is all you need: interpretable AI-enhanced histopathology for oncology. Seminars in Cancer Biology. Elsevier; 2023.
- Resolving challenges in deep learning-based analyses of histopathological images using explanation methods.. Sci Rep., 2020. [PubMed]
- Pan-cancer integrative histology-genomic analysis via multimodal deep learning.. Cancer Cell., 2022. [PubMed]
- Pathomic fusion: an integrated framework for fusing histopathology and genomic features for cancer diagnosis and prognosis.. IEEE Trans Med Imaging., 2022. [PubMed]
- Development and interpretation of a pathomics-based model for the prediction of microsatellite instability in colorectal cancer.. Theranostics., 2020. [PubMed]
- Morphological features extracted by AI associated with spatial transcriptomics in prostate cancer.. Cancers (Basel)., 2021. [PubMed]
- Histopathology images predict multi-omics aberrations and prognoses in colorectal cancer patients.. Nat Commun., 2023. [PubMed]
- Glioma grading via analysis of digital pathology images using machine learning.. Cancers (Basel)., 2020. [PubMed]
- 74 Tang P , YanX, NanY, et al Joint-individual fusion structure with fusion attention module for multi-modal skin cancer classification. Pattern Recognit. 2024;154:110604. 10.1016/j.patcog.2024.110604
- A deep learning approach to diagnostic classification of prostate cancer using pathology–radiology fusion.. J Magn Reson Imaging., 2021. [PubMed]
- The relationship between radiomics and pathomics in glioblastoma patients: Preliminary results from a cross-scale association study.. Front Oncol., 2022. [PubMed]
- Identifying cross-scale associations between radiomic and pathomic signatures of non-small cell lung cancer subtypes: preliminary results.. Cancers (Basel)., 2020. [PubMed]
- A radio-pathologic integrated model for prediction of lymph node metastasis stage in patients with gastric cancer.. Abdom Radiol (NY)., 2023. [PubMed]
- Multimodal integration of radiology, pathology and genomics for prediction of response to PD-(L)1 blockade in patients with non-small cell lung cancer.. Nat Cancer., 2022. [PubMed]
- Development of a novel combined nomogram model integrating deep learning-pathomics, radiomics and immunoscore to predict postoperative outcome of colorectal cancer lung metastasis patients.. J Hematol Oncol., 2022. [PubMed]
- Development and validation of a radiopathomic model for predicting pathologic complete response to neoadjuvant chemotherapy in breast cancer patients.. BMC Cancer., 2023. [PubMed]
- Deep learning with multimodal representation for pancancer prognosis prediction.. Bioinformatics., 2019. [PubMed]
- Development of a histopathology informatics pipeline for classification and prediction of clinical outcomes in subtypes of renal cell carcinoma.. Clin Cancer Res., 2021. [PubMed]
- Multi-omic machine learning predictor of breast cancer therapy response.. Nature., 2022. [PubMed]
- 85 Braman N , GordonJWH, GoossensET, WillisC, StumpeMC, VenkataramanJ. Deep orthogonal fusion: multimodal prognostic biomarker discovery integrating radiology, pathology, genomic, and clinical data. In: Medical Image Computing and Computer Assisted Intervention–MICCAI 2021: 24th International Conference. September 27-October 1, 2021, Proceedings, Part V 24, Springer; 2021:667-677.
- Integrating multi-omics data by learning modality invariant representations for improved prediction of overall survival of cancer.. Methods., 2021. [PubMed]
- Survival prediction of glioma patients from integrated radiology and pathology images using machine learning ensemble regression methods.. Appl Sci., 2022
- Development and validation of a radiopathomics model to predict pathological complete response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer: a multicentre observational study.. Lancet Digit Health., 2022. [PubMed]
- Selecting candidates for organ-preserving strategies after neoadjuvant chemoradiotherapy for rectal cancer: development and validation of a model integrating MRI radiomics and pathomics.. J Magn Reson Imaging., 2022. [PubMed]
- Multimodal data integration using machine learning improves risk stratification of high-grade serous ovarian cancer.. Nat Cancer., 2022. [PubMed]
- Breast cancer survival prognosis using the graph convolutional network with choquet fuzzy integral.. Sci Rep., 2023. [PubMed]
- Application of radiomics-based multiomics combinations in the tumor microenvironment and cancer prognosis.. J Transl Med., 2023. [PubMed]
- 93 Insight Ace. AI-Based Digital Pathology Solutions Market Size, Share & Trends Analysis Report. @ONLINE, February 2024.
- Current trend of artificial intelligence patents in digital pathology: a systematic evaluation of the patent landscape.. Cancers (Basel)., 2022. [PubMed]
- Moving beyond the turing test with the Allen AI science challenge.. Commun ACM., 2017
- An AI-based approach for modeling the synergy between radiotherapy and immunotherapy.. Sci Rep., 2024. [PubMed]
- A new era: artificial intelligence and machine learning in prostate cancer.. Nat Rev Urol., 2019. [PubMed]
- Charting new AI education in gastroenterology: cross-sectional evaluation of ChatGPT and perplexity AI in medical residency exam.. Dig Liv Dis., 2024
- Application of artificial intelligence in pathology: trends and challenges.. Diagnostics., 2022. [PubMed]
- Artificial intelligence in digital pathology: a systematic review and meta-analysis of diagnostic test accuracy.. NPJ Digit Med., 2024. [PubMed]
- Artificial intelligence for digital and computational pathology.. Nat Rev Bioeng., 2023
- 102 Cui C , MaY, CaoX, et al survey on multimodal large language models for autonomous driving. In: Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision. 2024:958-979.
- 103 OpenAI. Hello gpt-4o. @ONLINE, May 2024.
- 104 Gozalo-Brizuela R , Garrido-MerchánEC. A survey of generative AI applications. arXiv preprint arXiv:2306.02781. 2023, 10.48550/arXiv.2306.02781, preprint: not peer reviewed.
- 105 Htet A , LianaSR, AungT, BhaumikA. ChatGpT in content creation: techniques, applications, and ethical implications. Advanced Applications of Generative AI and Natural Language Processing Models. IGI Global; 2024:43-68.
- Artificial intelligence for multimodal data integration in oncology.. Cancer Cell., 2022. [PubMed]
- Integrative diagnostics: the time is now—a report from the international society for strategic studies in radiology.. Insights Imaging., 2023. [PubMed]
- Radiomics biomarkers correlate with CD8 expression and predict immune signatures in melanoma patients.. Mol Cancer Res., 2021. [PubMed]
- Interactive multimedia reporting technical considerations: HIMSS-SIIM collaborative white paper.. J Digit Imaging., 2022. [PubMed]
- Building flexible, scalable, and machine learning-ready multimodal oncology datasets.. Sensors., 2024. [PubMed]
- Recommendations for performance evaluation of machine learning in pathology: a concept paper from the college of American pathologists.. Arch Pathol Lab Med., 2023. [DOI]
